We narrowed to 20,363 results for: ren
-
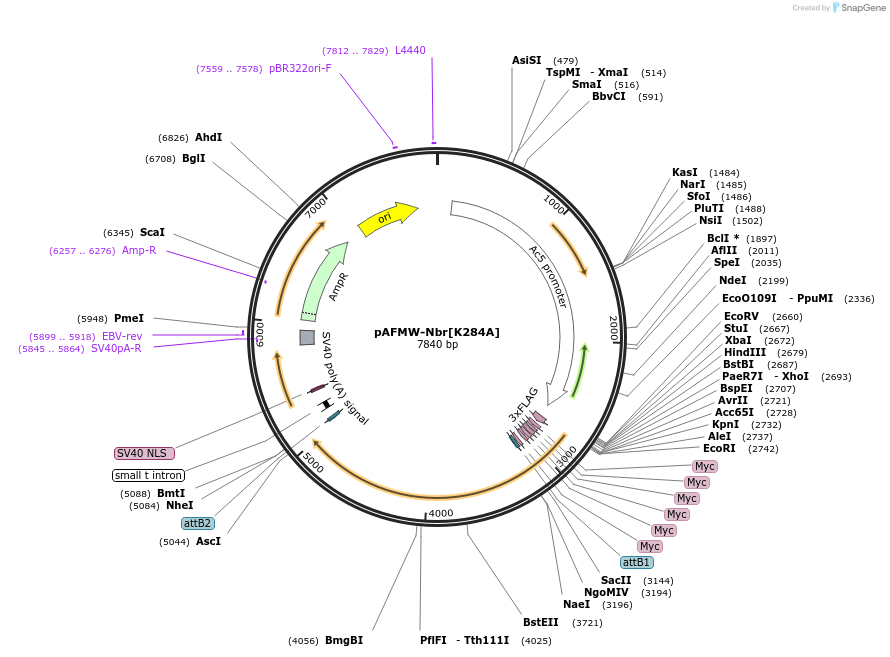
Plasmid#188595PurposeFlag-tagged Nibbler variants expression in Drosophila S2 cellsDepositorAvailable SinceOct. 27, 2022AvailabilityAcademic Institutions and Nonprofits only
-
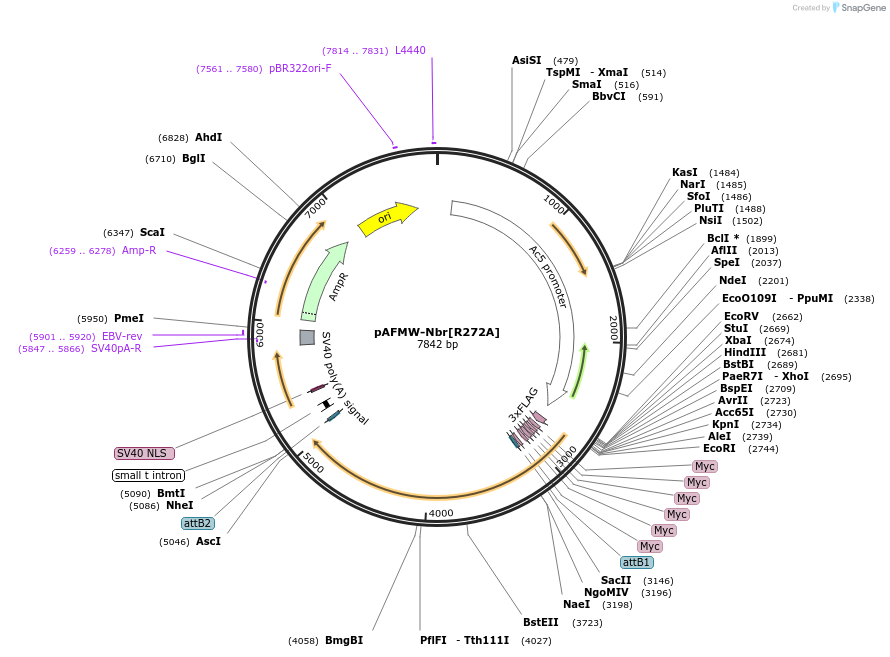
pAFMW-Nbr[R272A]
Plasmid#188594PurposeFlag-tagged Nibbler variants expression in Drosophila S2 cellsDepositorAvailable SinceOct. 27, 2022AvailabilityAcademic Institutions and Nonprofits only -
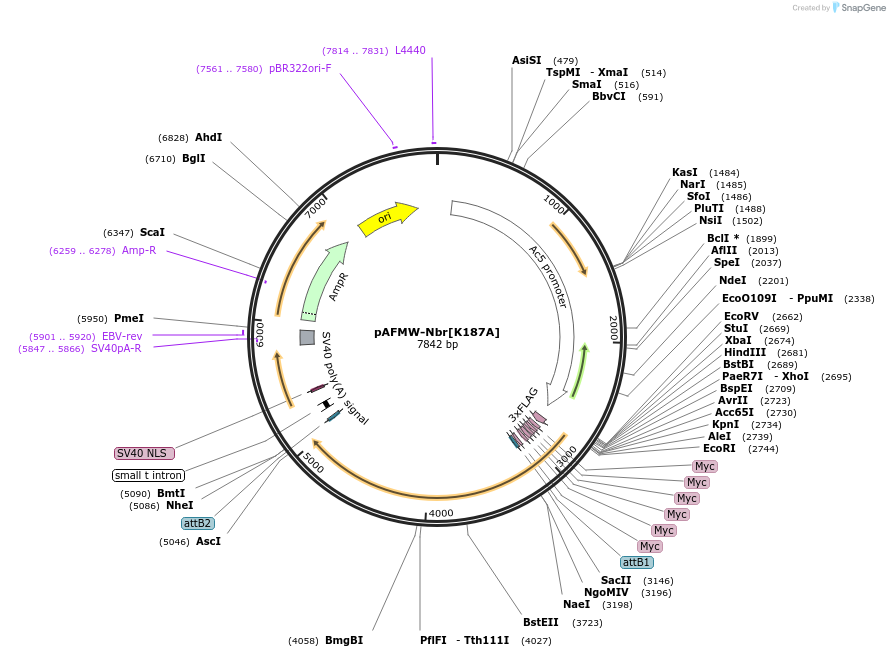
pAFMW-Nbr[K187A]
Plasmid#188593PurposeFlag-tagged Nibbler variants expression in Drosophila S2 cellsDepositorAvailable SinceOct. 27, 2022AvailabilityAcademic Institutions and Nonprofits only -
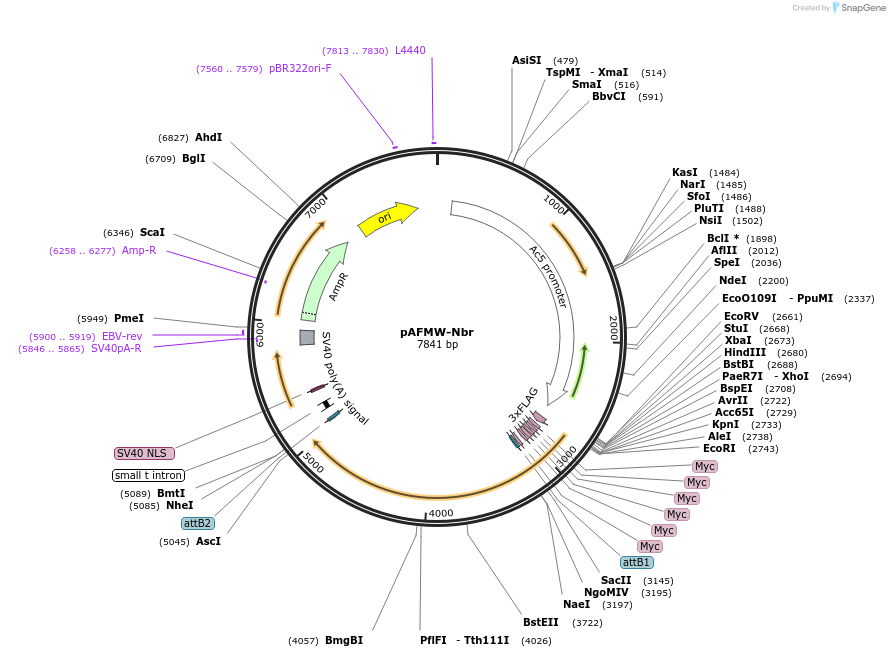
pAFMW-Nbr
Plasmid#188584PurposeFlag-tagged Nibbler variants expression in Drosophila S2 cellsDepositorAvailable SinceOct. 27, 2022AvailabilityAcademic Institutions and Nonprofits only -
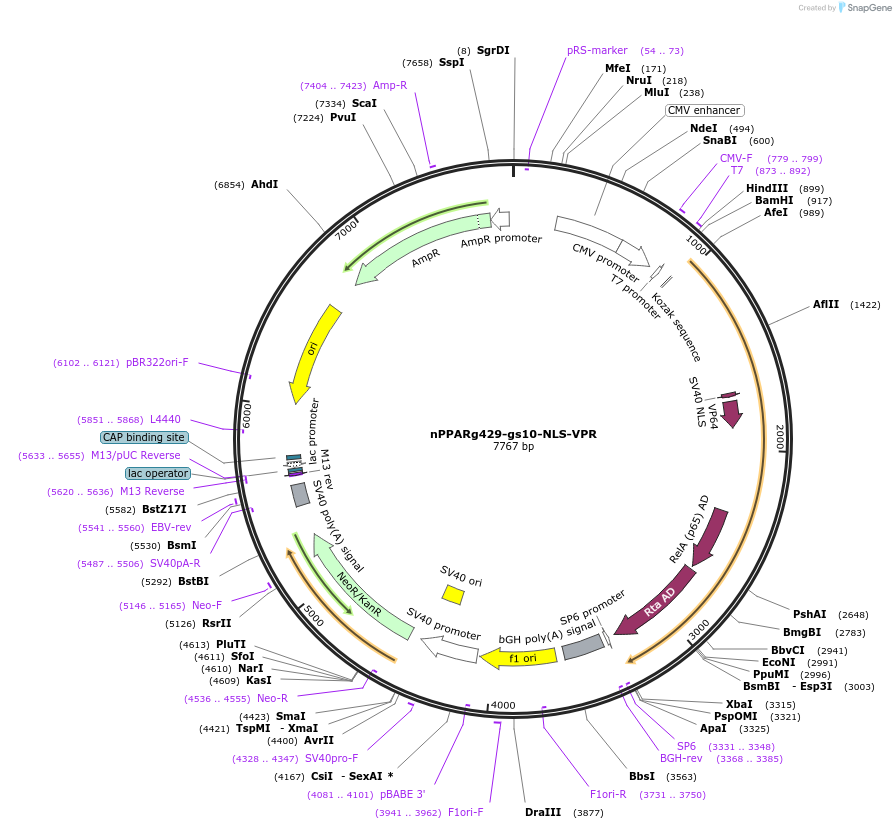
nPPARg429-gs10-NLS-VPR
Plasmid#188266PurposePlasmid ensures constitutive expression of the N-terminal fragment of split PPARgamma (aa 200-429), fused to the VPR activation domain, a flexible GS linker and the SV40 NLS signal at the C-terminusDepositorInsertnPPARg429 (PPARG Human)
ExpressionMammalianAvailable SinceOct. 17, 2022AvailabilityAcademic Institutions and Nonprofits only -
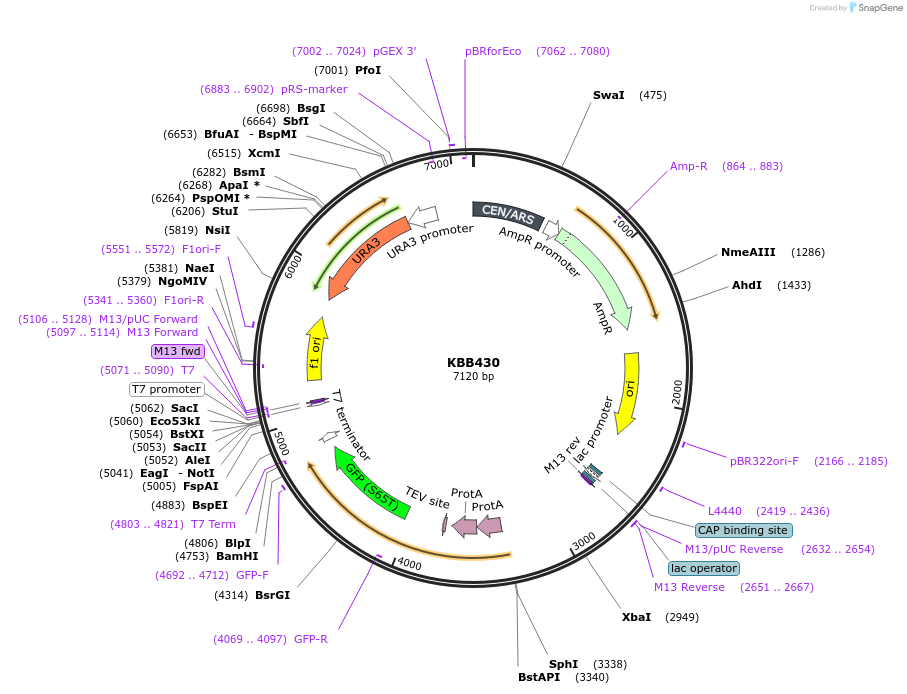
KBB430
Plasmid#185091PurposeNMA111-GFP with first nuclear localization signal mutated (amino acids 28 - 30, KRK -> AAA)DepositorInsertNMA111
TagsGFPExpressionYeastMutationFirst nuclear localization signal mutated (amino …Available SinceJuly 29, 2022AvailabilityAcademic Institutions and Nonprofits only -
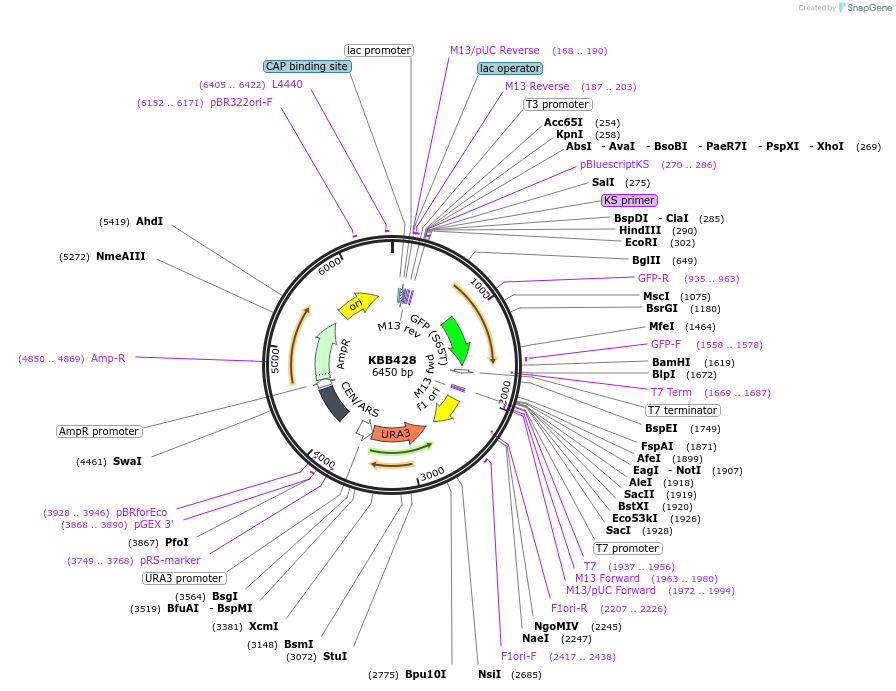
KBB428
Plasmid#185090PurposeNMA111-GFP with first nuclear localization signal mutated (amino acids 9 - 11, KKR -> AAA)DepositorInsertNMA111
TagsGFPExpressionYeastMutationFirst nuclear localization signal mutated (amino …Available SinceJuly 27, 2022AvailabilityAcademic Institutions and Nonprofits only -
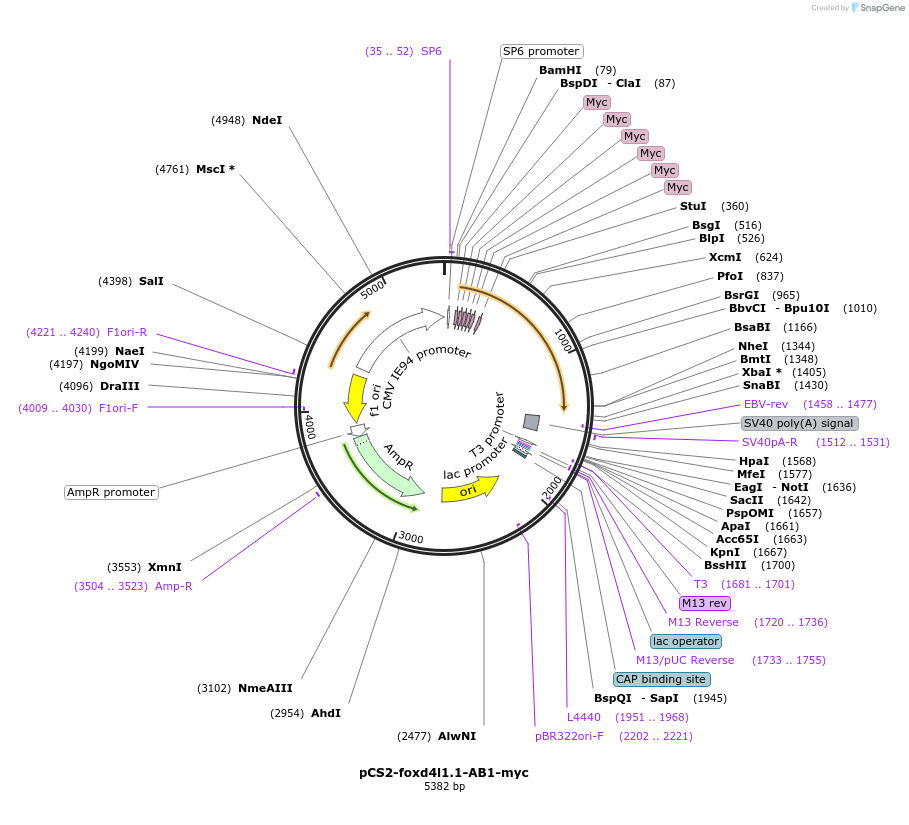
pCS2-foxd4l1.1-AB1-myc
Plasmid#185511PurposeExpression of Xenopus laevis foxd4DepositorInsertfoxd4l1.1
UseIn vitro transcriptionTagsmycMutationdeletion of IDILGE in acidic blobAvailable SinceJuly 18, 2022AvailabilityAcademic Institutions and Nonprofits only -
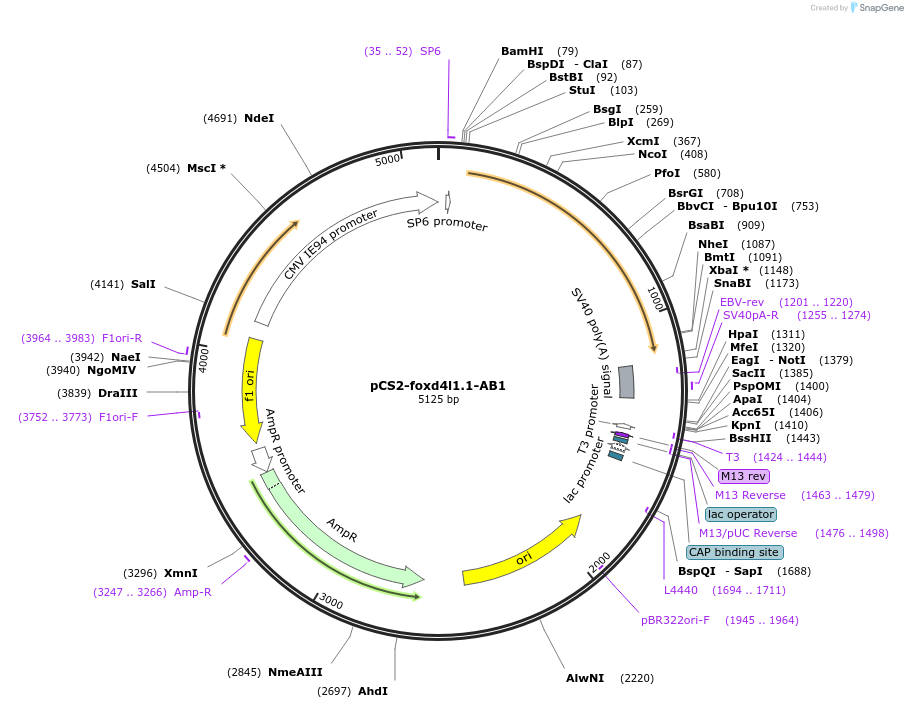
pCS2-foxd4l1.1-AB1
Plasmid#185510PurposeExpression of Xenopus laevis foxd4DepositorInsertfoxd4l1.1
UseIn vitro transcriptionMutationdeletion of IDILGE in acidic blobAvailable SinceJuly 18, 2022AvailabilityAcademic Institutions and Nonprofits only -
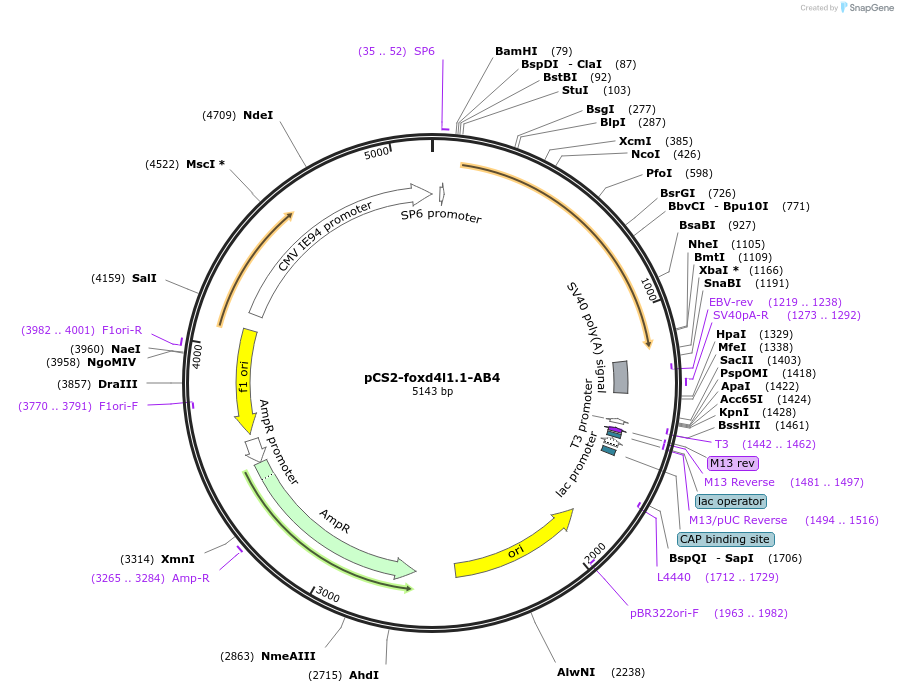
pCS2-foxd4l1.1-AB4
Plasmid#185513PurposeExpression of Xenopus laevis foxd4DepositorInsertfoxd4l1.1
UseIn vitro transcriptionMutationIDILGE replaced with AAAAAAAvailable SinceJuly 18, 2022AvailabilityAcademic Institutions and Nonprofits only -
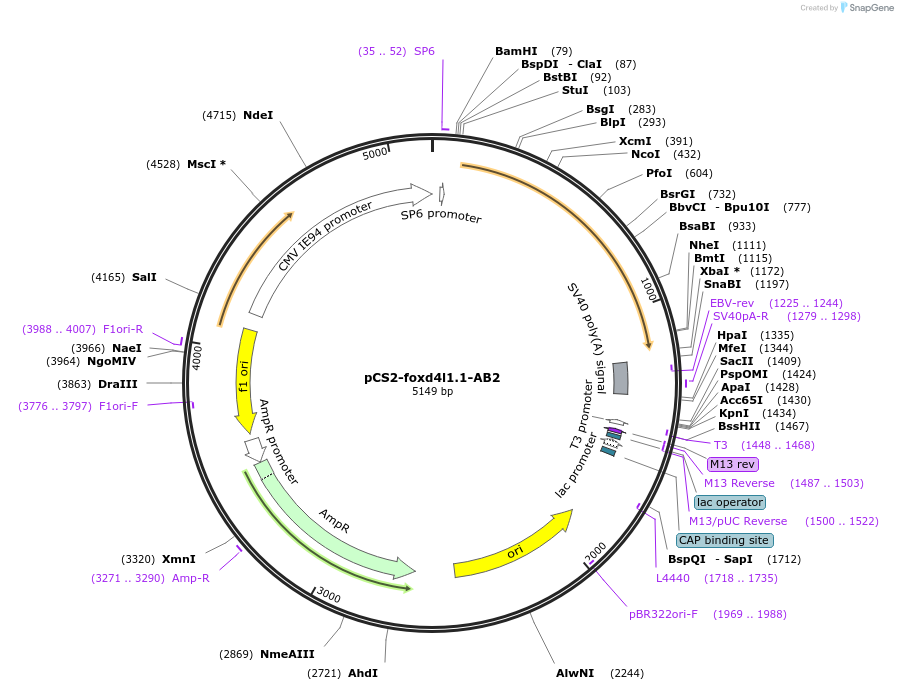
pCS2-foxd4l1.1-AB2
Plasmid#185512PurposeExpression of Xenopus laevis foxd4DepositorInsertfoxd4l1.1
UseIn vitro transcriptionMutationIDIL replaced with AAAAAAAvailable SinceJuly 18, 2022AvailabilityAcademic Institutions and Nonprofits only -
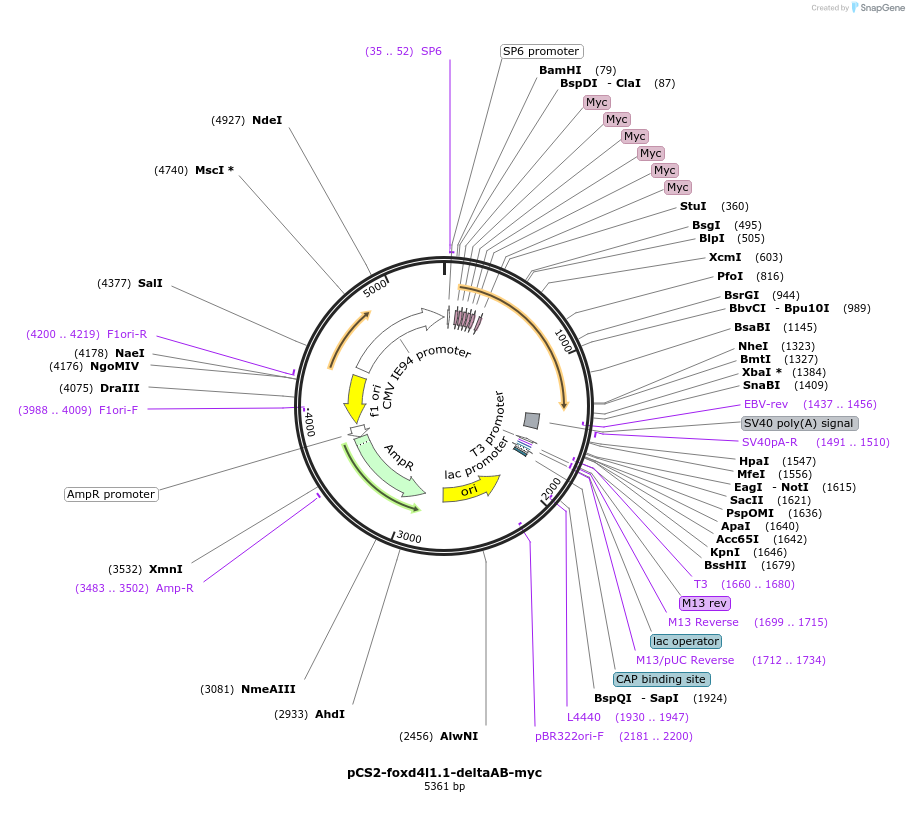
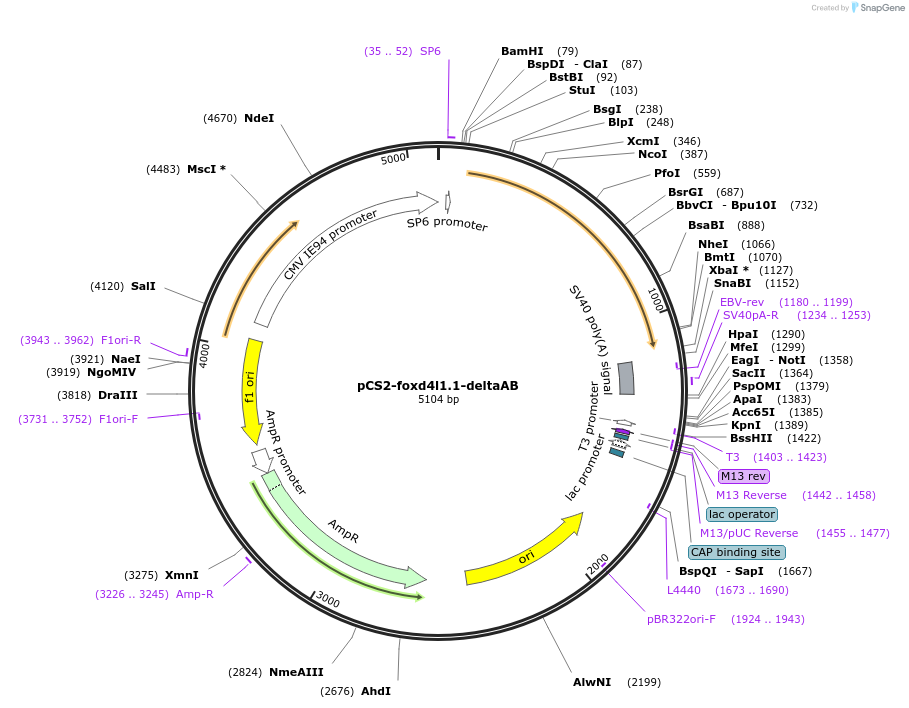
pCS2-foxd4l1.1-deltaAB-myc
Plasmid#185503PurposeExpression of Xenopus laevis foxd4DepositorInsertfoxd4l1.1
UseIn vitro transcriptionTagsmycMutationacidic blob (aa 21-33) deletedAvailable SinceJuly 18, 2022AvailabilityAcademic Institutions and Nonprofits only -
pCS2-foxd4l1.1-deltaAB
Plasmid#185502PurposeExpression of Xenopus laevis foxd4DepositorInsertfoxd4l1.1
UseIn vitro transcriptionMutationacidic blob (aa 21-33) deletedAvailable SinceJuly 18, 2022AvailabilityAcademic Institutions and Nonprofits only -
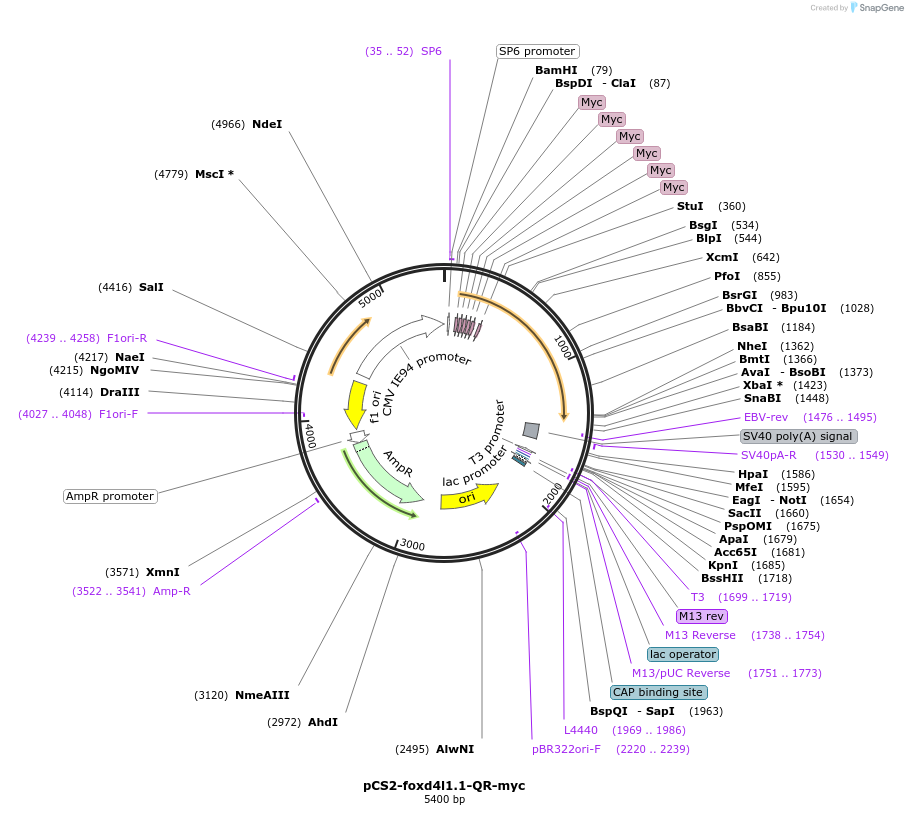
pCS2-foxd4l1.1-QR-myc
Plasmid#185521PurposeExpression of Xenopus laevis foxd4DepositorInsertfoxd4l1.1
UseIn vitro transcriptionTagsmycMutationreplaces Q>R upstream of GARQYLNIAvailable SinceJuly 12, 2022AvailabilityAcademic Institutions and Nonprofits only -
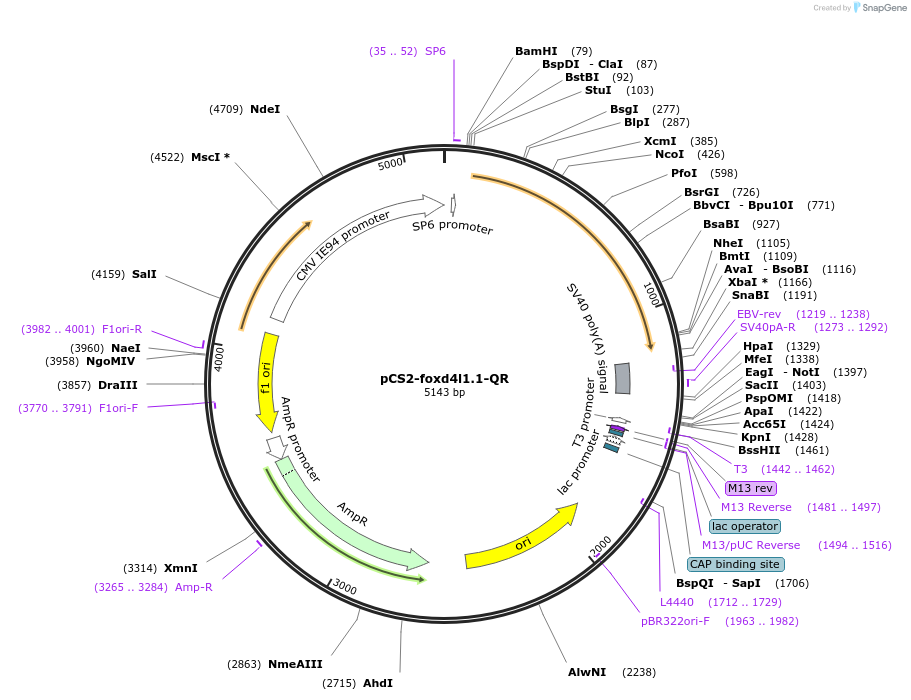
pCS2-foxd4l1.1-QR
Plasmid#185520PurposeExpression of Xenopus laevis foxd4DepositorInsertfoxd4l1.1
UseIn vitro transcriptionMutationreplaces Q>R upstream of GARQYLNIAvailable SinceJuly 12, 2022AvailabilityAcademic Institutions and Nonprofits only -
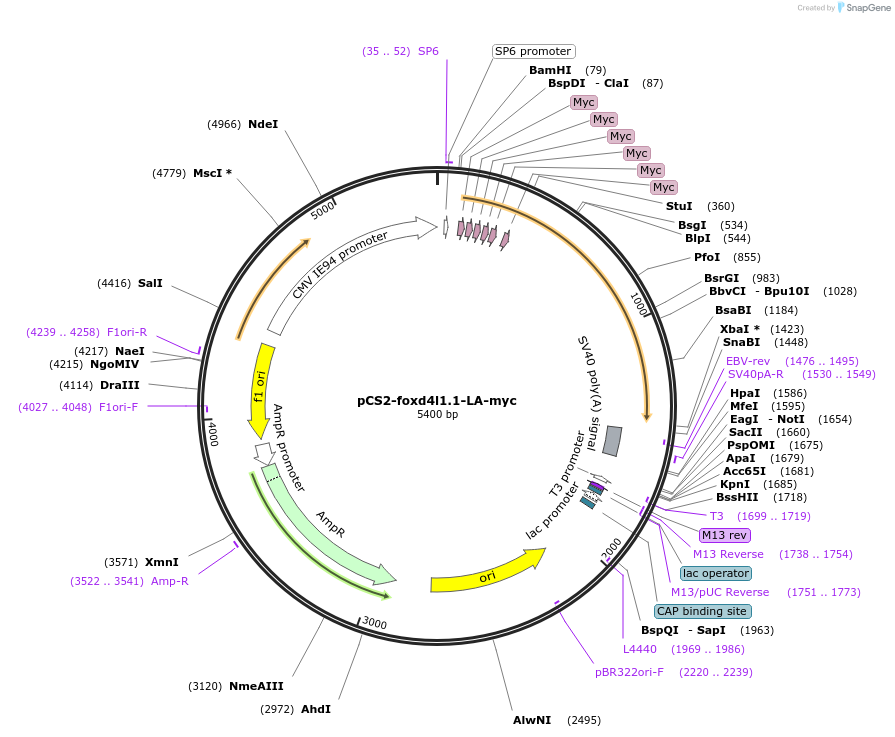
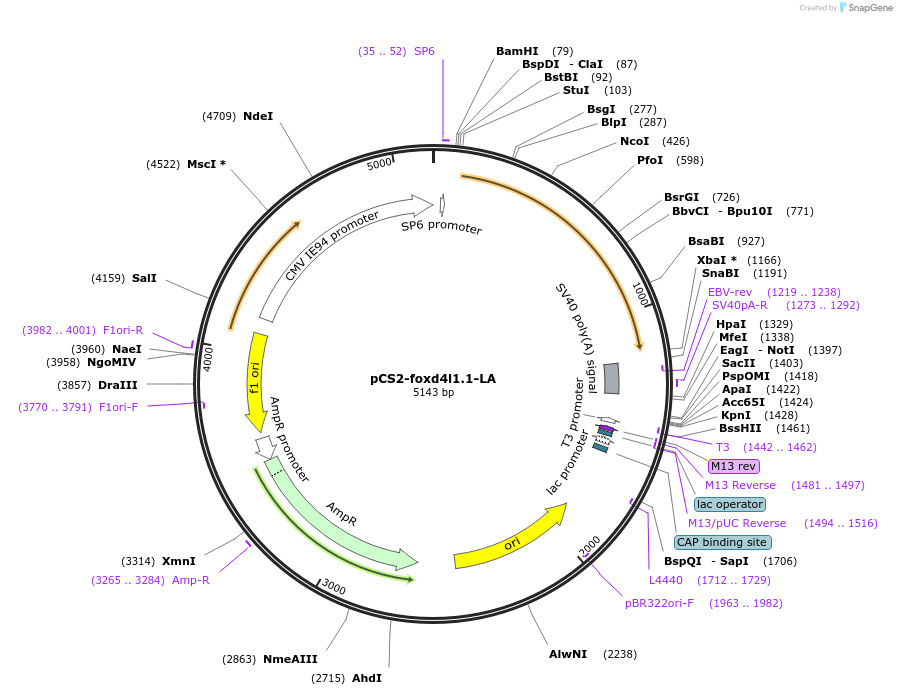
pCS2-foxd4l1.1-LA-myc
Plasmid#185519PurposeExpression of Xenopus laevis foxd4DepositorInsertfoxd4l1.1
UseIn vitro transcriptionTagsmycMutationreplaces L>A upstream of GARQYLNAvailable SinceJuly 12, 2022AvailabilityAcademic Institutions and Nonprofits only -
pCS2-foxd4l1.1-LA
Plasmid#185518PurposeExpression of Xenopus laevis foxd4DepositorInsertfoxd4l1.1
UseIn vitro transcriptionMutationreplaces L>A upstream of GARQYLNIAvailable SinceJuly 12, 2022AvailabilityAcademic Institutions and Nonprofits only -
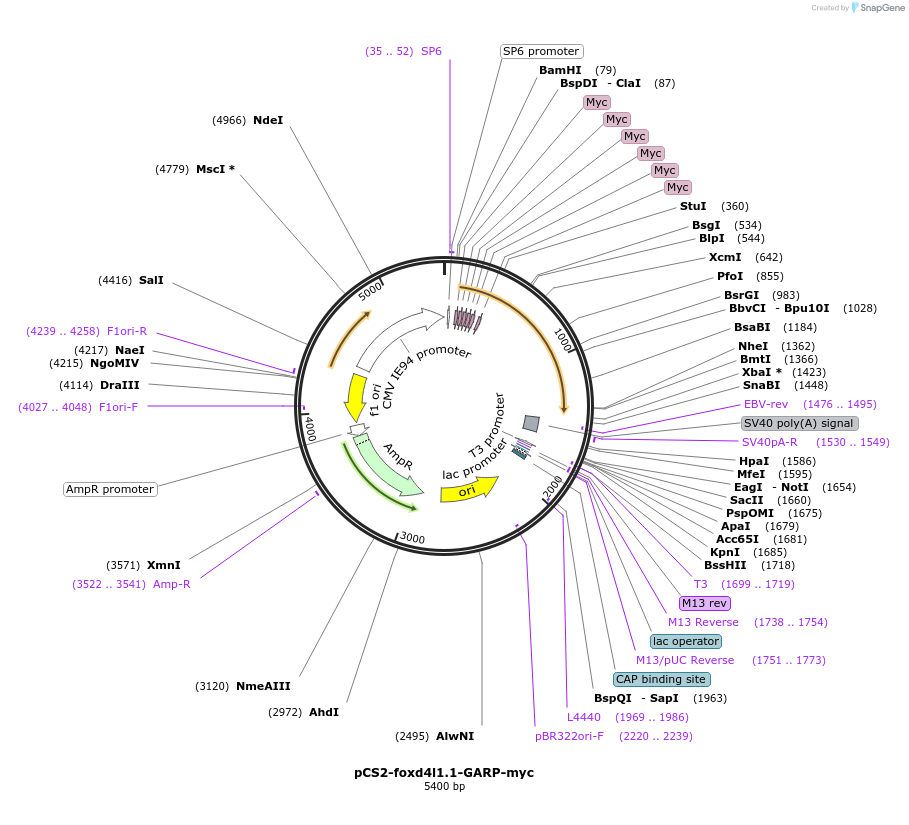
pCS2-foxd4l1.1-GARP-myc
Plasmid#185517PurposeExpression of Xenopus laevis foxd4DepositorInsertfoxd4l1.1
UseIn vitro transcriptionTagsmycMutationReplaces Q to P in GARQYNLIAvailable SinceJuly 12, 2022AvailabilityAcademic Institutions and Nonprofits only -
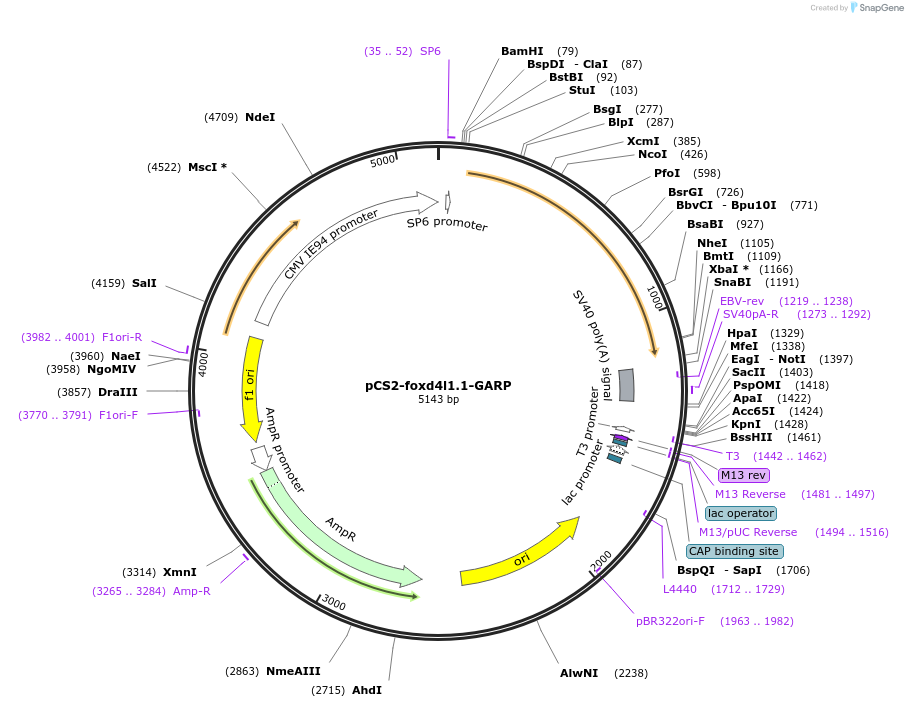
pCS2-foxd4l1.1-GARP
Plasmid#185516PurposeExpression of Xenopus laevis foxd4DepositorInsertfoxd4l1.1
UseIn vitro transcriptionMutationReplaces Q to P in GARQYNLIAvailable SinceJuly 12, 2022AvailabilityAcademic Institutions and Nonprofits only -
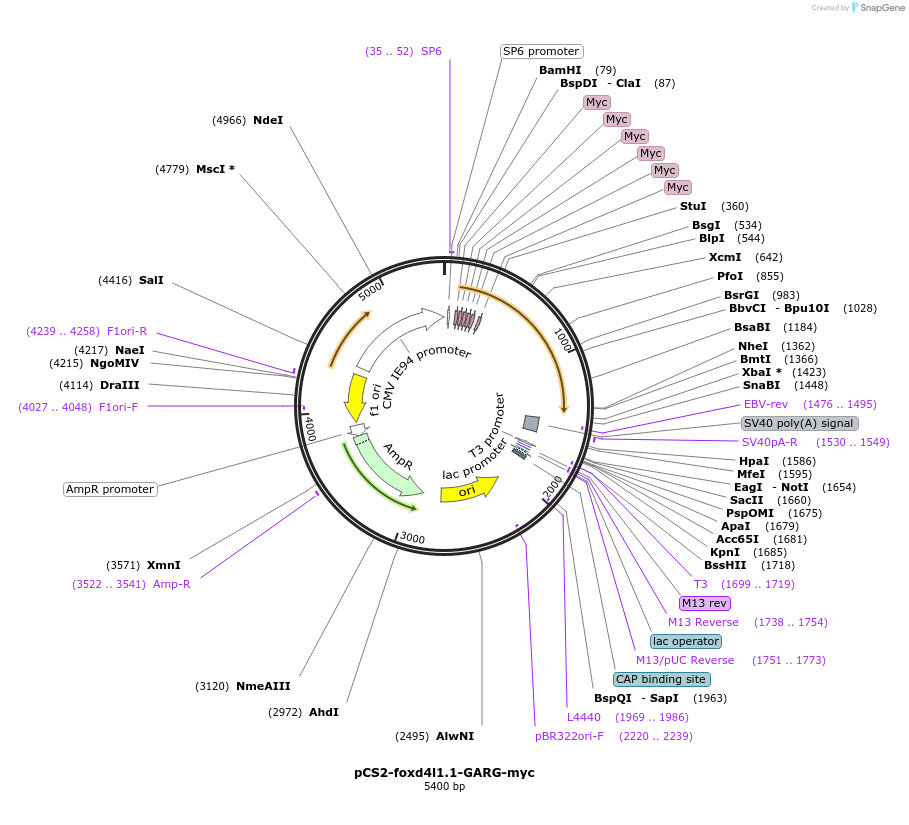
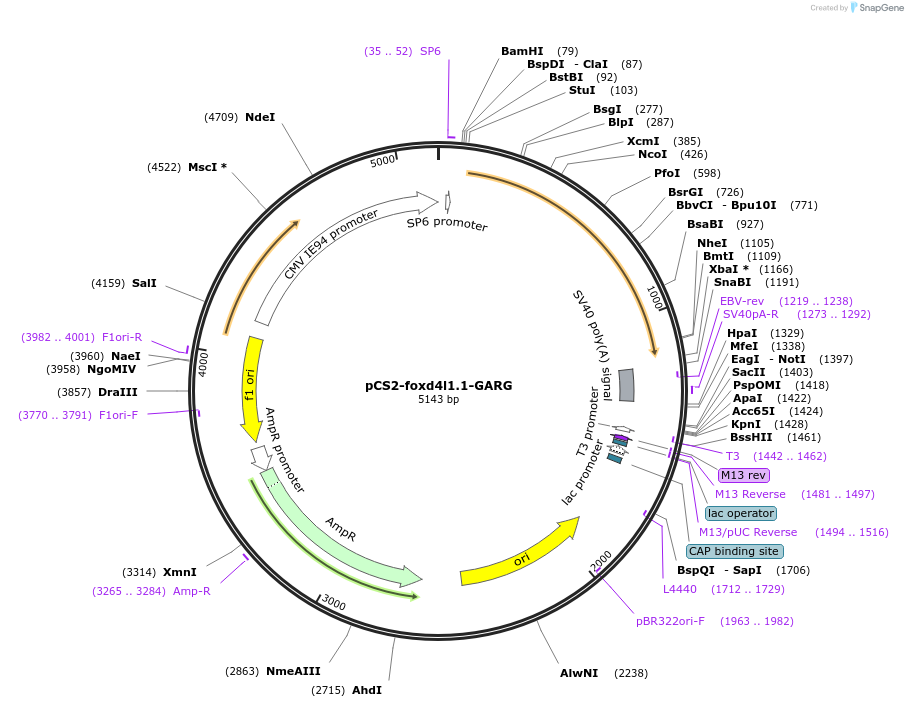
pCS2-foxd4l1.1-GARG-myc
Plasmid#185515PurposeExpression of Xenopus laevis foxd4DepositorInsertfoxd4l1.1
UseIn vitro transcriptionTagsmycMutationReplaces Q to G in GARQYNLIAvailable SinceJuly 12, 2022AvailabilityAcademic Institutions and Nonprofits only -
pCS2-foxd4l1.1-GARG
Plasmid#185514PurposeExpression of Xenopus laevis foxd4DepositorInsertfoxd4l1.1
UseIn vitro transcriptionMutationReplaces Q to G in GARQYNLIAvailable SinceJuly 12, 2022AvailabilityAcademic Institutions and Nonprofits only -
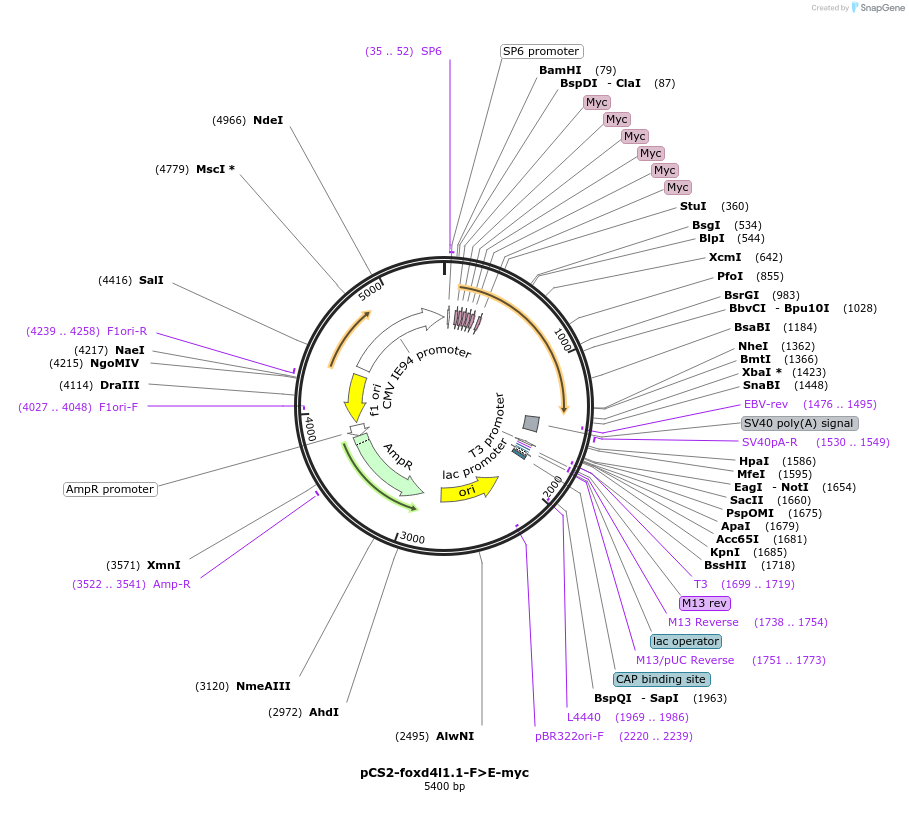
pCS2-foxd4l1.1-F>E-myc
Plasmid#185509PurposeExpression of Xenopus laevis foxd4DepositorInsertfoxd4l1.1
UseIn vitro transcriptionTagsMycMutationF>EAvailable SinceJuly 12, 2022AvailabilityAcademic Institutions and Nonprofits only -
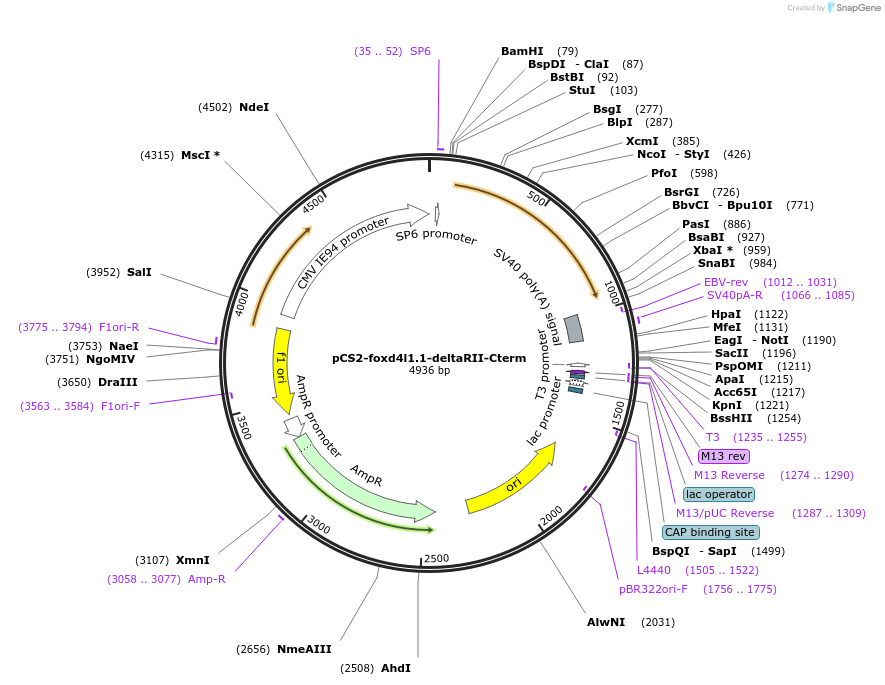
pCS2-foxd4l1.1-deltaRII-Cterm
Plasmid#185504PurposeExpression of Xenopus laevis foxd4DepositorInsertfoxd4l1.1
UseIn vitro transcriptionMutationRII domain and C-term deletedAvailable SinceJuly 12, 2022AvailabilityAcademic Institutions and Nonprofits only -
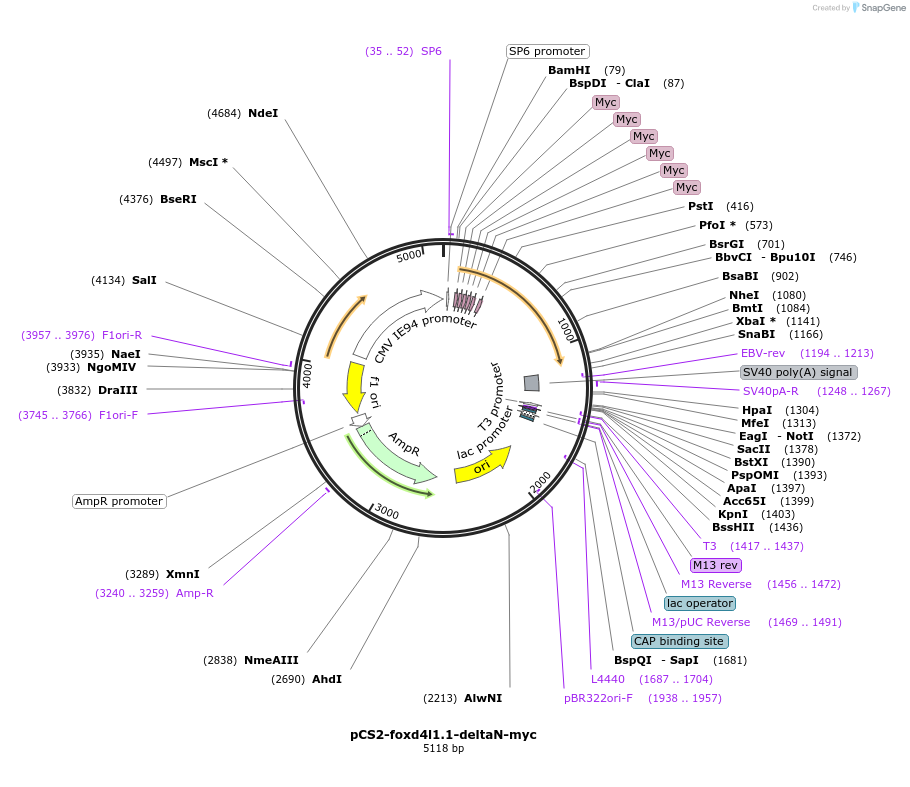
pCS2-foxd4l1.1-deltaN-myc
Plasmid#185500PurposeExpression of Xenopus laevis foxd4DepositorInsertfoxd4l1.1
UseIn vitro transcriptionTagsmycMutationN-terminus deletedAvailable SinceJuly 12, 2022AvailabilityAcademic Institutions and Nonprofits only -
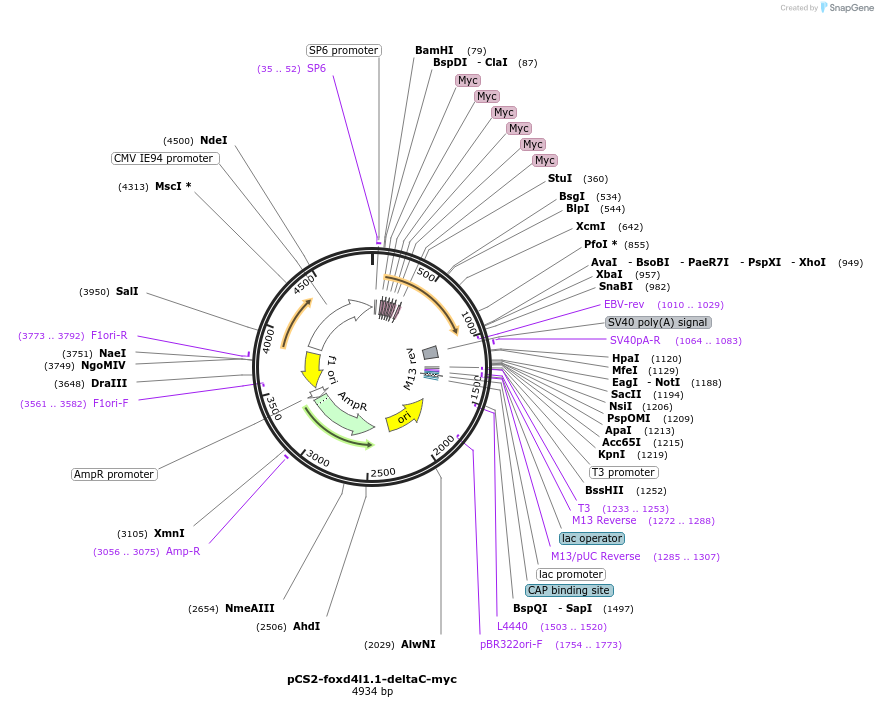
pCS2-foxd4l1.1-deltaC-myc
Plasmid#185501PurposeExpression of Xenopus laevis foxd4DepositorInsertfoxd4l1.1
UseIn vitro transcriptionTagsmycMutationC-terminus deletedAvailable SinceJuly 12, 2022AvailabilityAcademic Institutions and Nonprofits only -
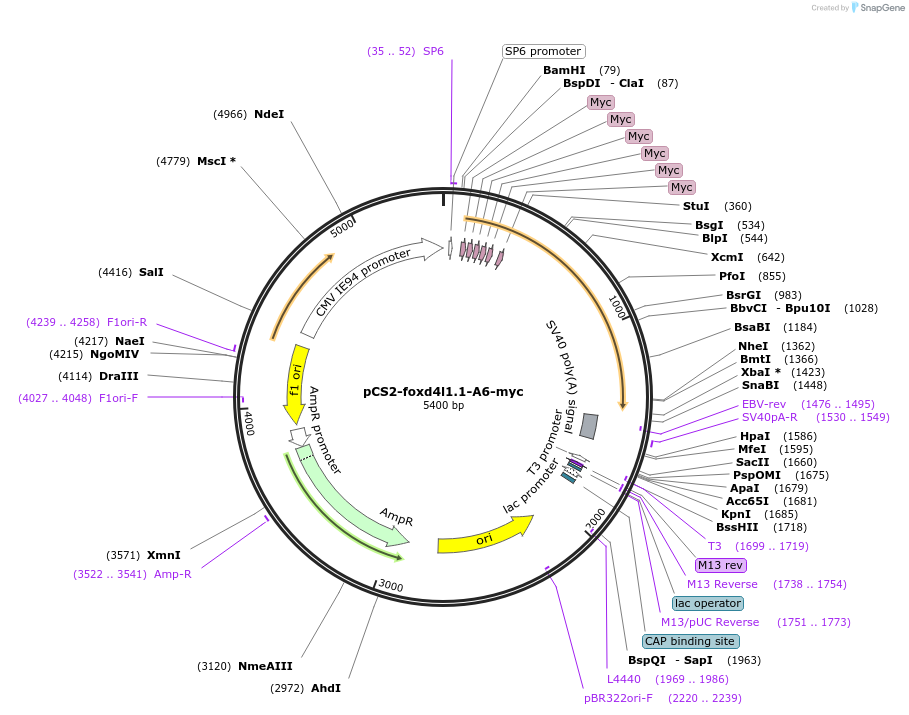
pCS2-foxd4l1.1-A6-myc
Plasmid#185507PurposeExpression of Xenopus laevis foxd4DepositorInsertfoxd4l1.1
UseIn vitro transcriptionTagsmycMutationFSIENIM to AAAAAAMAvailable SinceJuly 8, 2022AvailabilityAcademic Institutions and Nonprofits only -
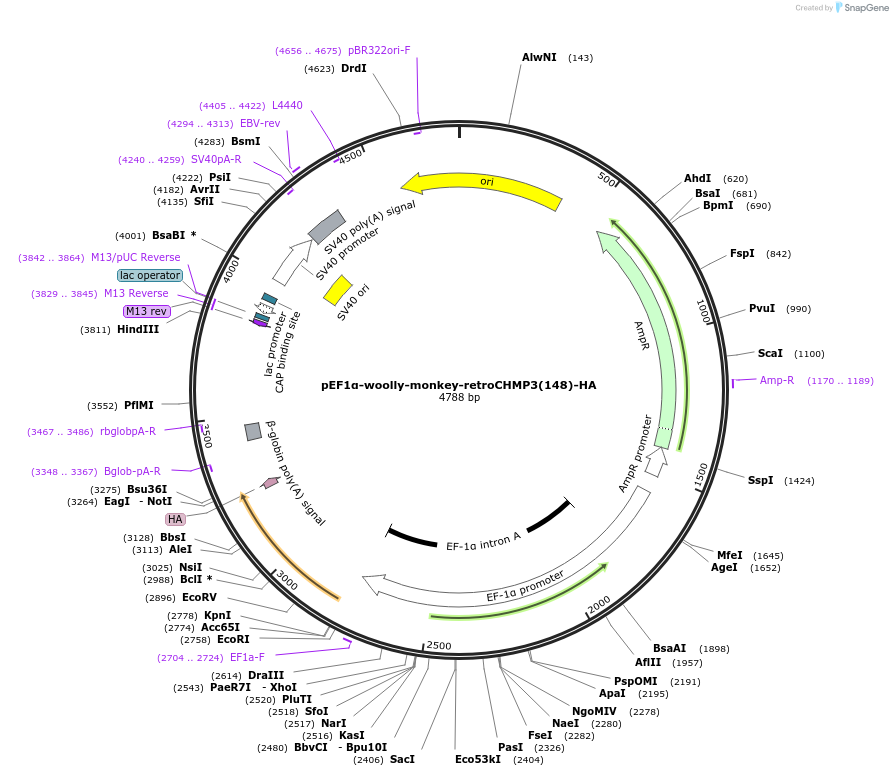
pEF1α-woolly-monkey-retroCHMP3(148)-HA
Plasmid#154250Purposeexpresses prematurely truncated woolly monkey retroCHMP3 in mammalian cellsDepositorInsertretroCHMP3
TagsHAExpressionMammalianMutationC-terminal truncation of retroCHMP3 at amino acid…PromoterEF1-alphaAvailable SinceApril 12, 2022AvailabilityAcademic Institutions and Nonprofits only -
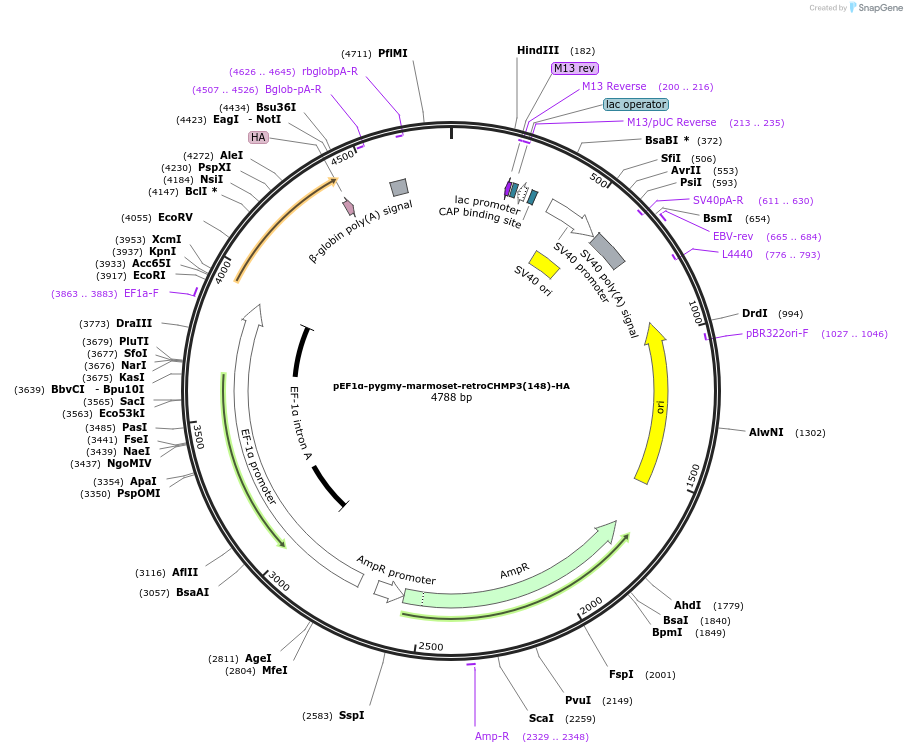
pEF1α-pygmy-marmoset-retroCHMP3(148)-HA
Plasmid#154251Purposeexpresses prematurely truncated pygmy marmoset retroCHMP3 in mammalian cellsDepositorInsertretroCHMP3
TagsHAExpressionMammalianMutationC-terminal truncation of retroCHMP3 at amino acid…PromoterEF1-alphaAvailable SinceApril 12, 2022AvailabilityAcademic Institutions and Nonprofits only -
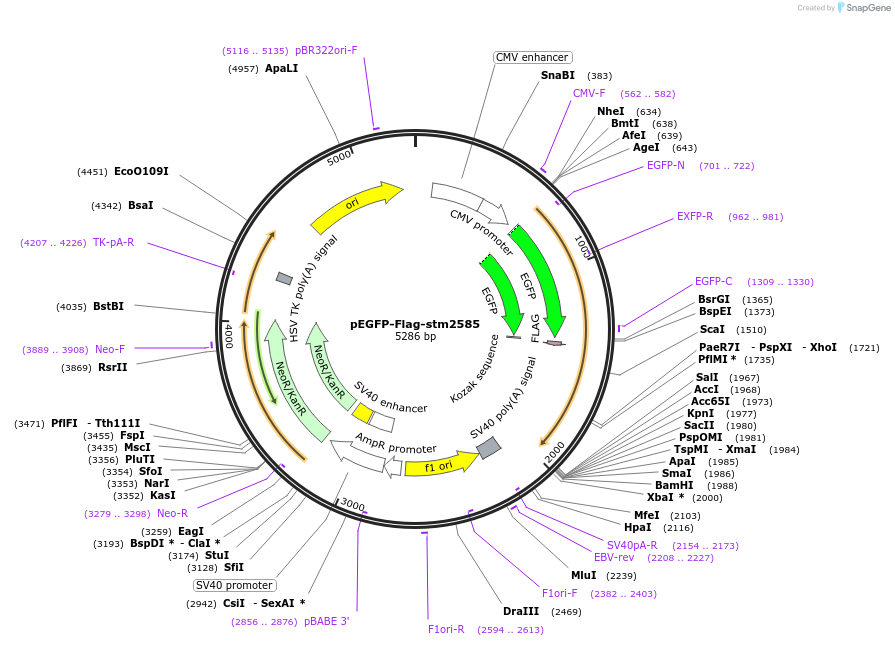
pEGFP-Flag-stm2585
Plasmid#174397PurposeNative S. Typhimurium gene stm2585 (sarA/steE) with N-terminal GFP and Flag tags.DepositorInsertsarA
TagsEGFP and FlagExpressionMammalianAvailable SinceSept. 13, 2021AvailabilityAcademic Institutions and Nonprofits only -
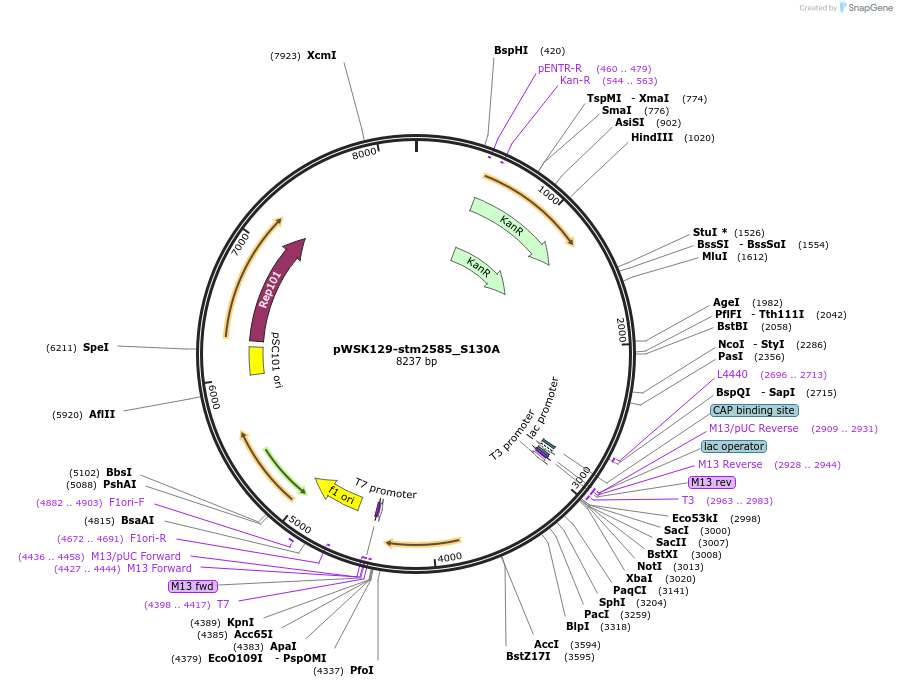
pWSK129-stm2585_S130A
Plasmid#174403PurposeBacterial expression of sarA with its GSK3-binding SxxxSP motif mutated to SxxxAP.DepositorInsertsarA
ExpressionBacterialMutationsarA with S130A mutationAvailable SinceSept. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
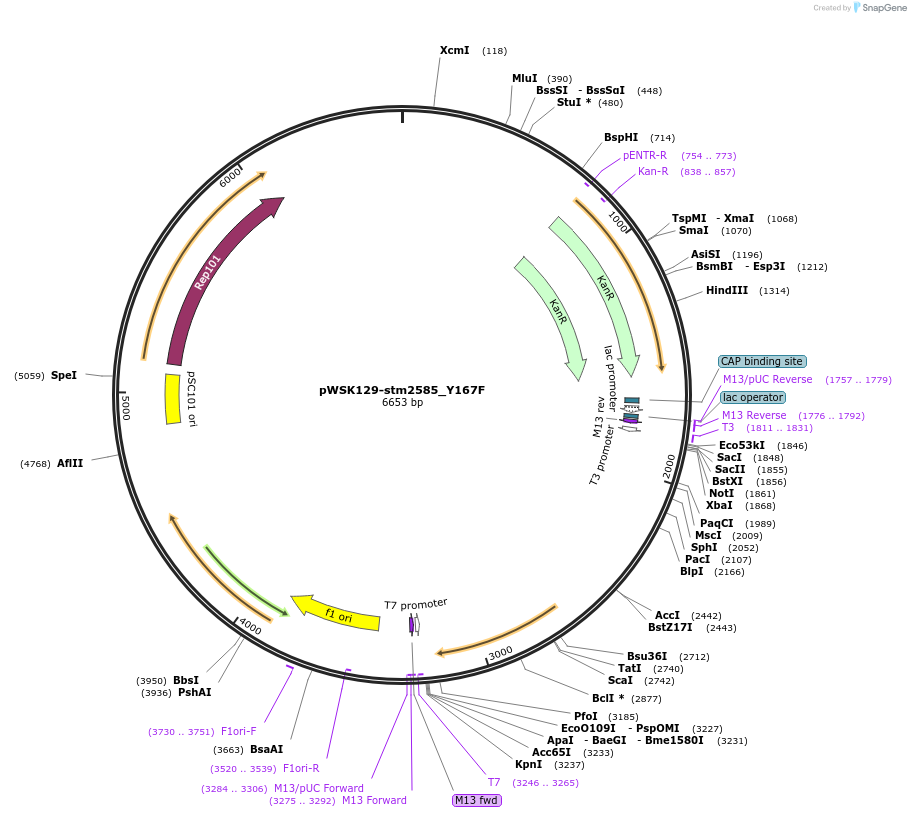
pWSK129-stm2585_Y167F
Plasmid#174402PurposeBacterial expression of sarA with its STAT3-binding YxxQ motif mutated to FxxQDepositorInsertsarA
ExpressionBacterialMutationsarA with Y167F mutationPromoterEndogenousAvailable SinceSept. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
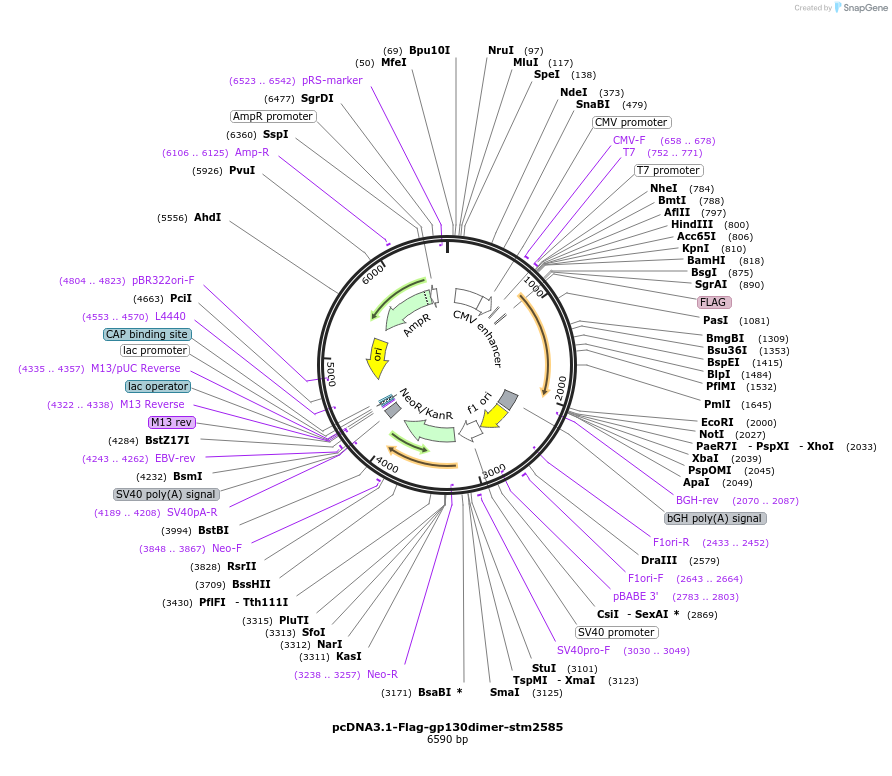
pcDNA3.1-Flag-gp130dimer-stm2585
Plasmid#174395PurposeConstitutively active gp130 dimer with codon-optimized stm2585 insert (replacing amino acids 140-179)DepositorInsertIL6ST cytosolic domain and sarA chimera
ExpressionMammalianAvailable SinceSept. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
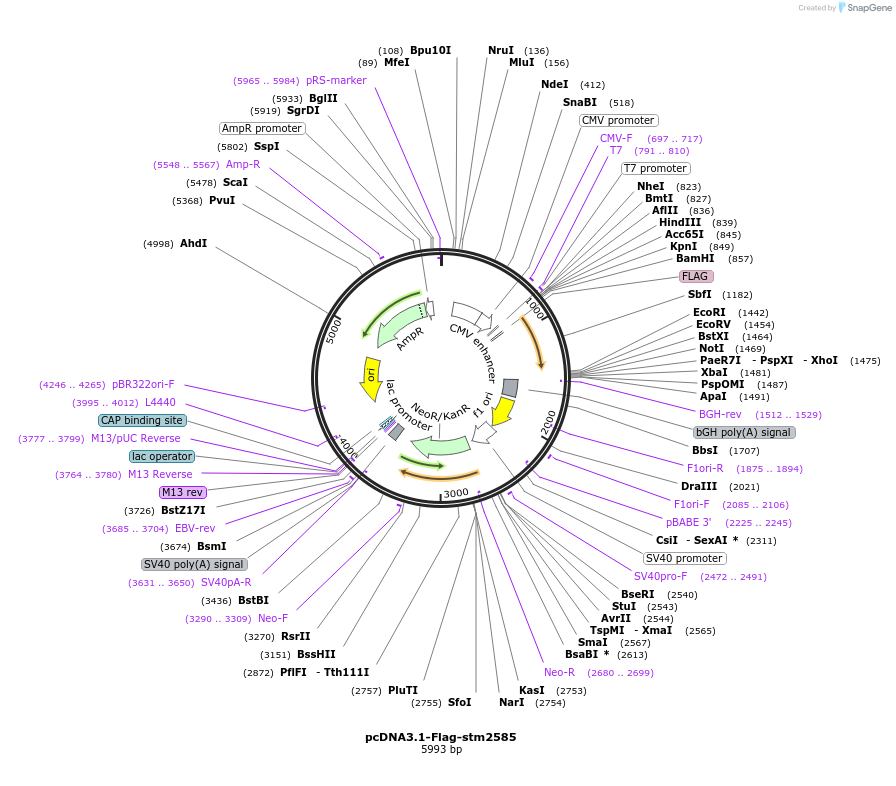
pcDNA3.1-Flag-stm2585
Plasmid#174391PurposeS. Typhimurium gene stm2585 (sarA/steE) codon-optimized for mammalian expression with N-terminal Flag tagDepositorInsertstm2585
TagsFlagExpressionMammalianAvailable SinceSept. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
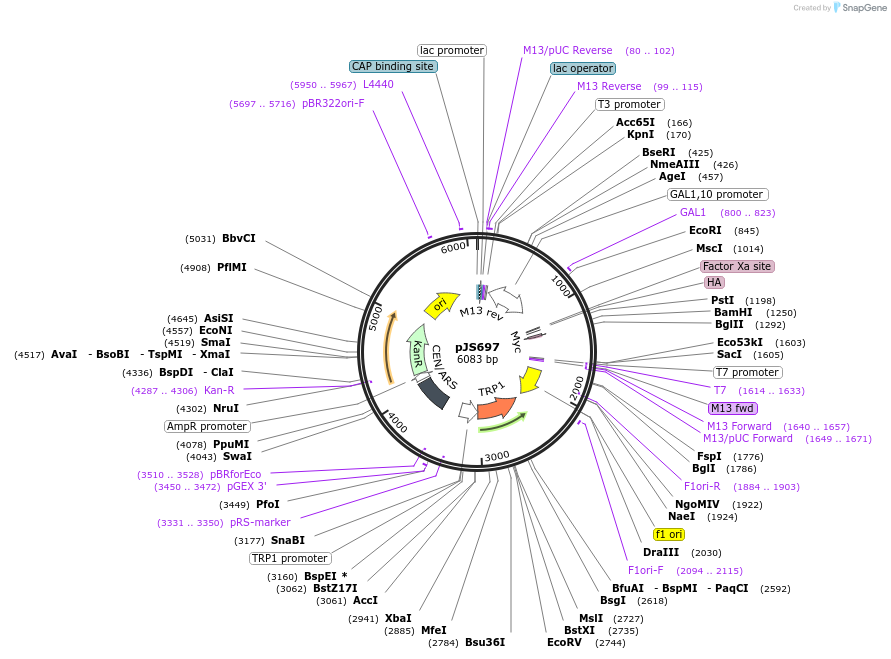
pJS697
Plasmid#168778PurposeYeast surface display plasmid backbone that can be combined with pJS699 through homologous recombinationDepositorTypeEmpty backboneTagsCmyc and HAExpressionYeastAvailable SinceAug. 30, 2021AvailabilityAcademic Institutions and Nonprofits only -
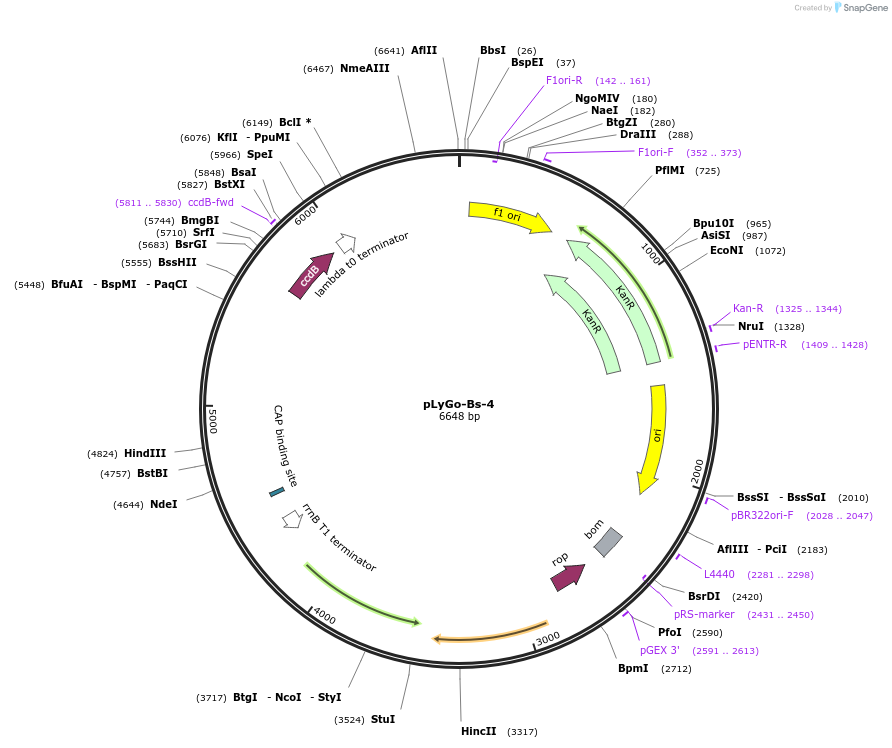
pLyGo-Bs-4
Plasmid#163142PurposepLyGo cloning vector for a sequence of interest (LPMO) in B. subtilis. Vector encoding the BatLPMO10 signal peptide and the LyGo cassette (SapI-ccdB-SapI)DepositorTypeEmpty backboneUseSynthetic BiologyExpressionBacterialPromoterPamyL-PamyQ-PcryIIIA-cryIIIAstab49Available SinceJuly 30, 2021AvailabilityAcademic Institutions and Nonprofits only -
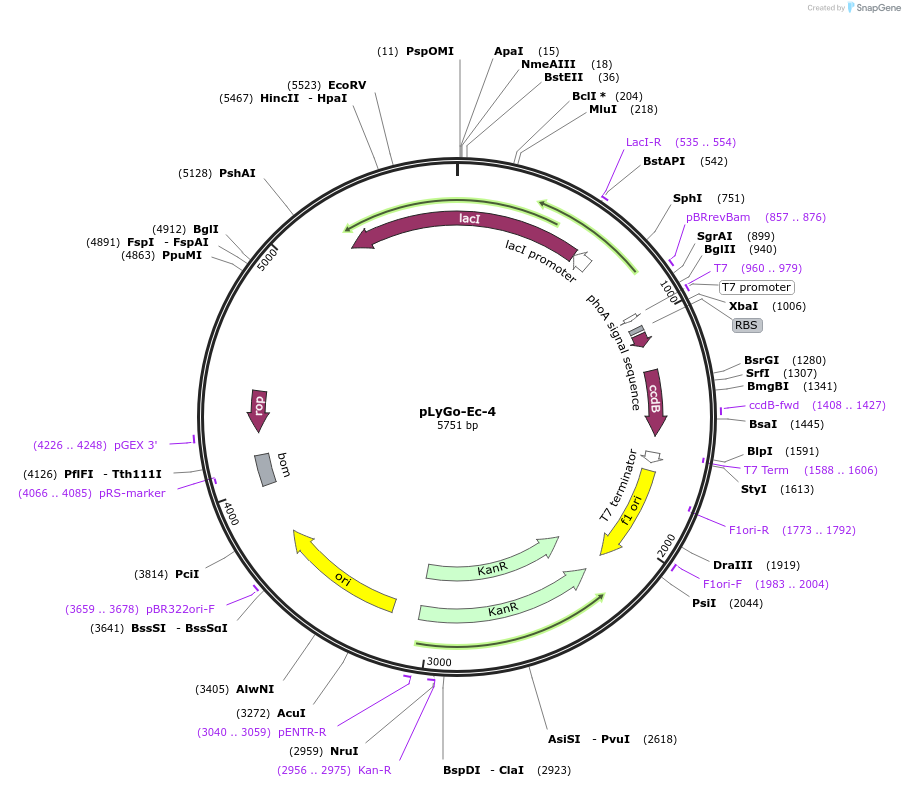
pLyGo-Ec-4
Plasmid#163130PurposepLyGo cloning vector for periplasmic expression in E. coli of a sequence of interest (LPMO). Vector encoding PhoA signal peptide and the LyGo cassette (SapI-ccdB-SapI)DepositorTypeEmpty backboneUseSynthetic BiologyExpressionBacterialPromoterT7Available SinceJuly 30, 2021AvailabilityAcademic Institutions and Nonprofits only -
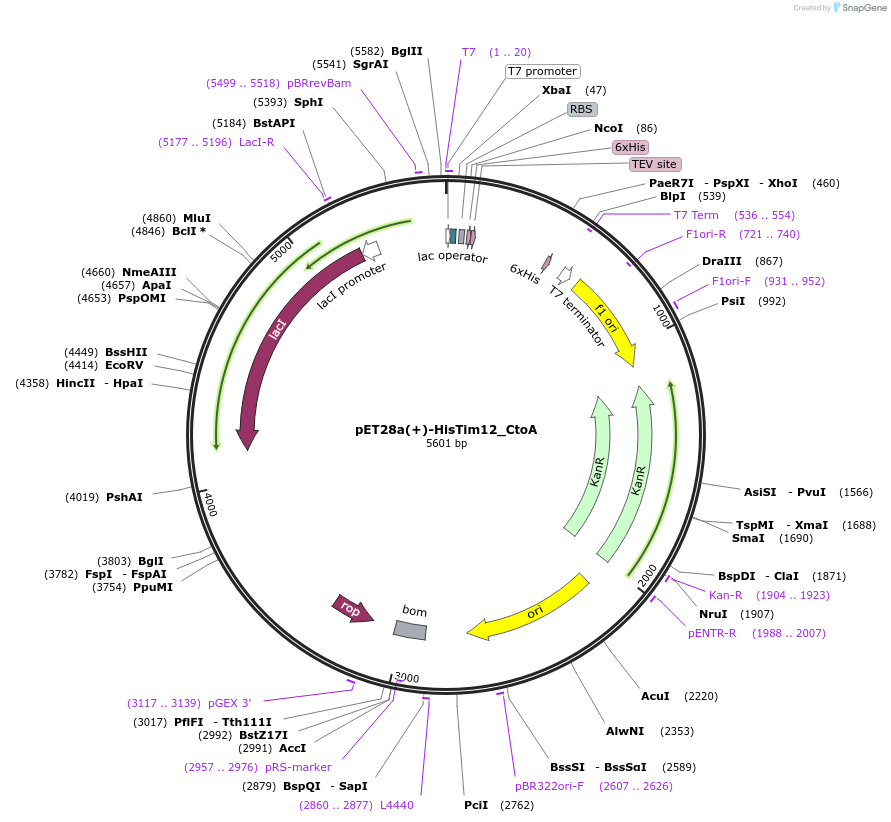
pET28a(+)-HisTim12_CtoA
Plasmid#170281PurposeExpresses yeast Tim12 (the latter with cleavable N-ter His-tag), codon-optimized for E. coli productionDepositorInsertyeast Tim12 (TIM12 Budding Yeast)
TagsTEV-cleavable N-terminal His tag (on backbone)ExpressionBacterialMutationmutated two non-conserved Cys to AlaAvailable SinceJune 15, 2021AvailabilityAcademic Institutions and Nonprofits only -
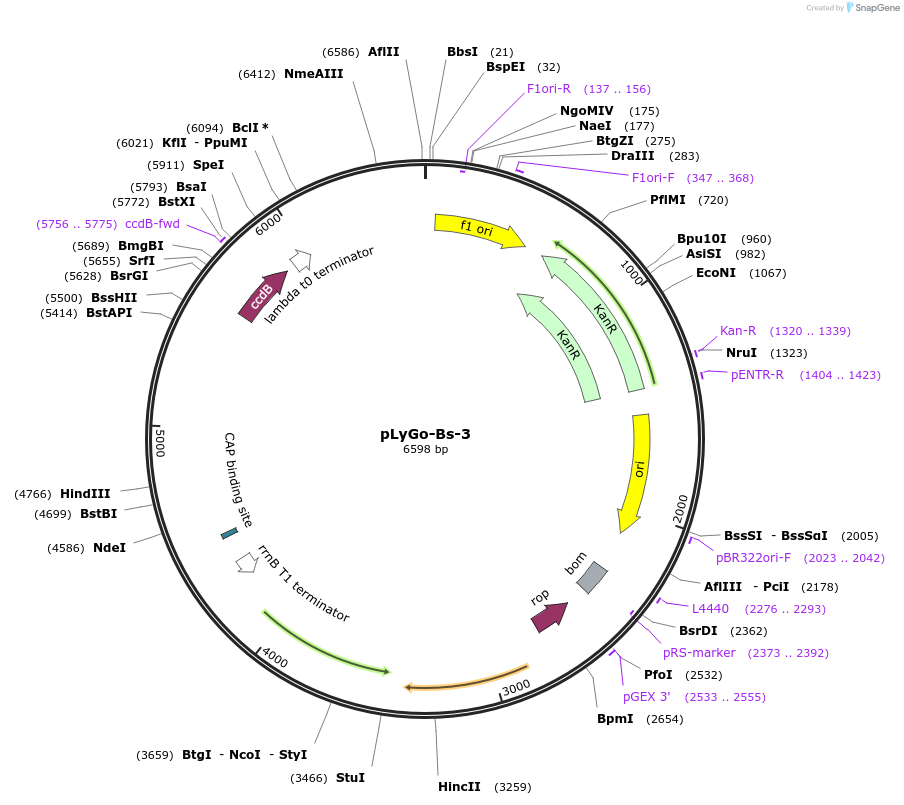
pLyGo-Bs-3
Plasmid#163141PurposepLyGo cloning vector for a sequence of interest (LPMO) in B. subtilis. Vector encoding the AprE signal peptide and the LyGo cassette (SapI-ccdB-SapI)DepositorTypeEmpty backboneUseSynthetic BiologyExpressionBacterialPromoterPamyL-PamyQ-PcryIIIA-cryIIIAstab49Available SinceMay 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
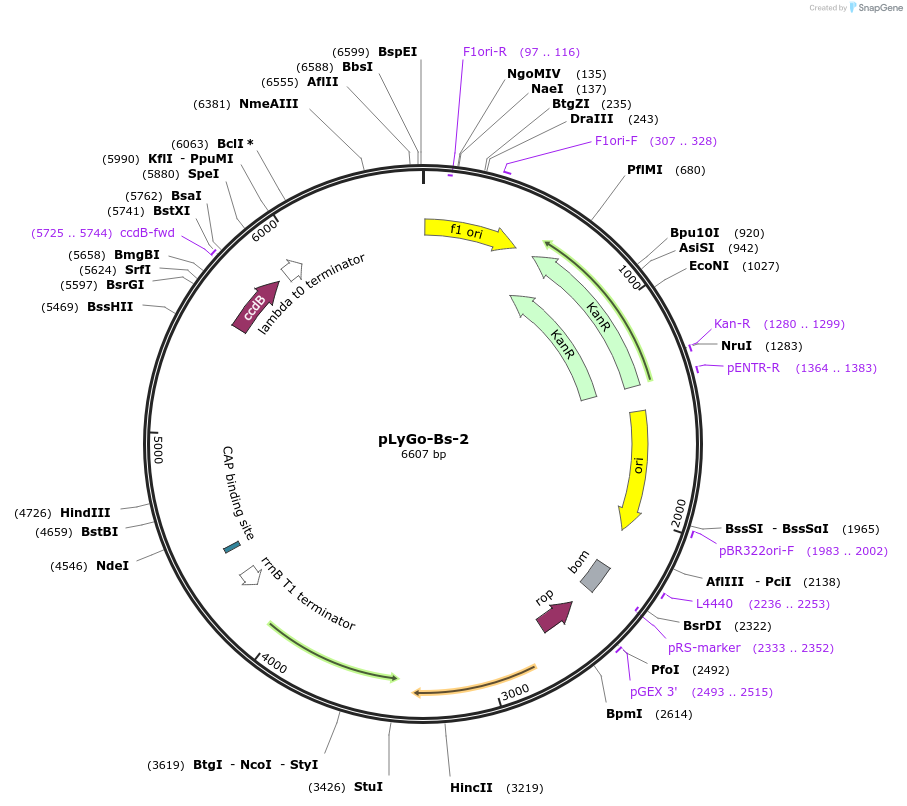
pLyGo-Bs-2
Plasmid#163140PurposepLyGo cloning vector for a sequence of interest (LPMO) in Bacillus subtilis. Vector encoding the AmyQ signal peptide and the LyGo cassette (SapI-ccdB-SapI)DepositorTypeEmpty backboneUseSynthetic BiologyExpressionBacterialPromoterPamyL-PamyQ-PcryIIIA-cryIIIAstab49Available SinceMay 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
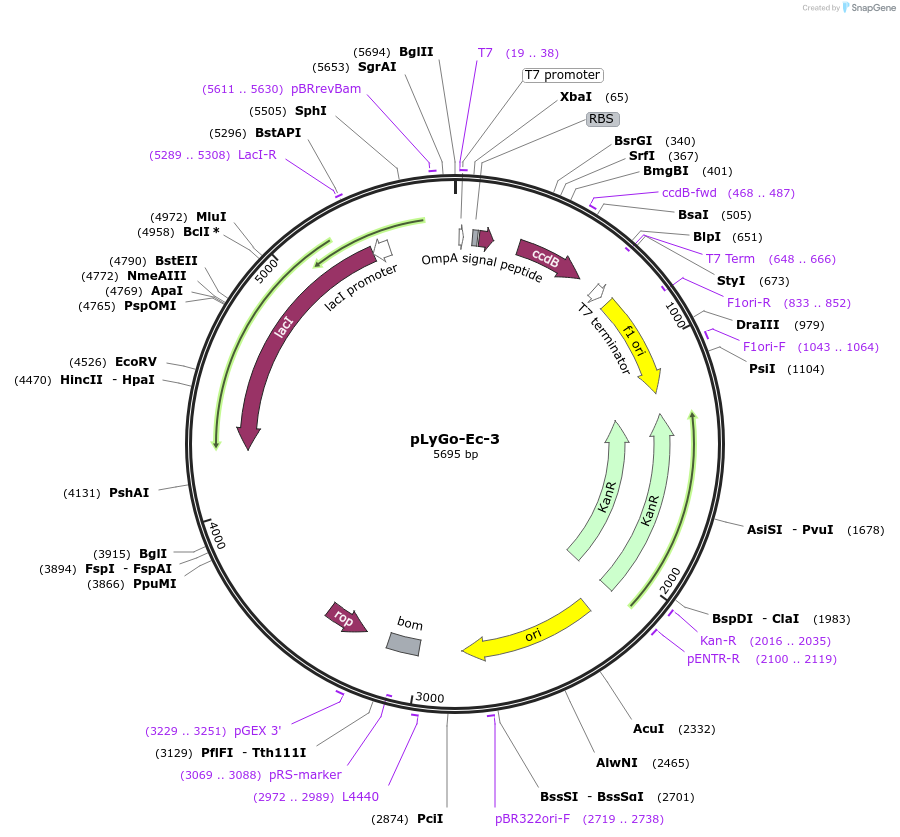
pLyGo-Ec-3
Plasmid#163129PurposepLyGo cloning vector for periplasmic expression in E. coli of a sequence of interest (LPMO). Vector encoding OmpA signal peptide and the LyGo cassette (SapI-ccdB-SapI)DepositorTypeEmpty backboneUseSynthetic BiologyExpressionBacterialPromoterT7Available SinceMay 6, 2021AvailabilityAcademic Institutions and Nonprofits only