We narrowed to 9,329 results for: APP
-
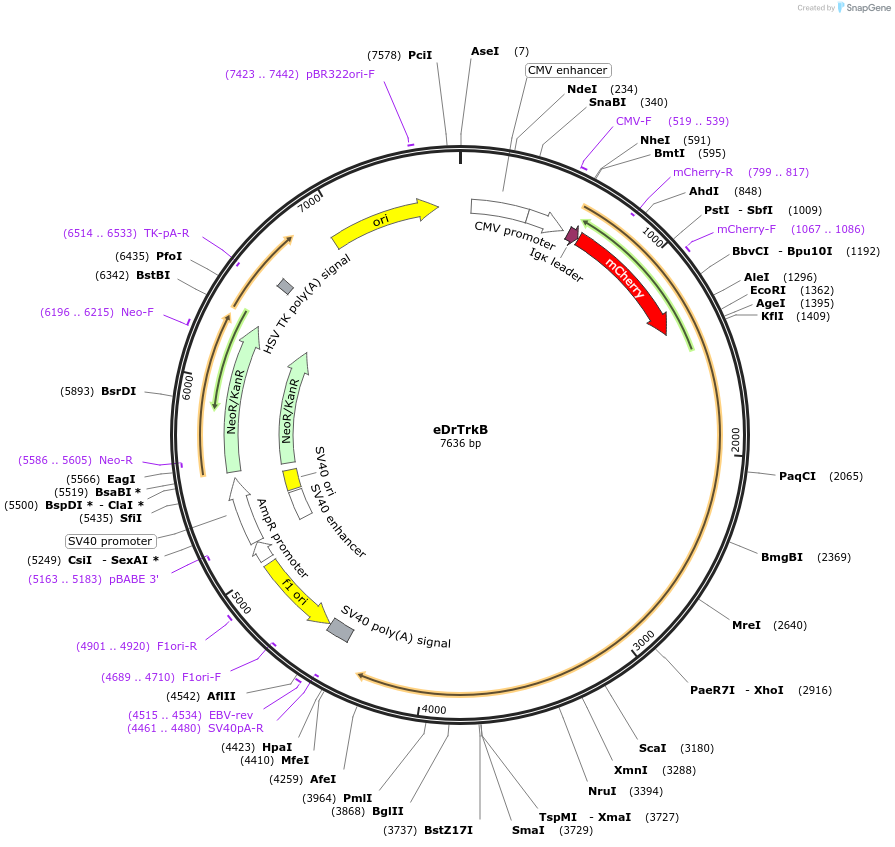
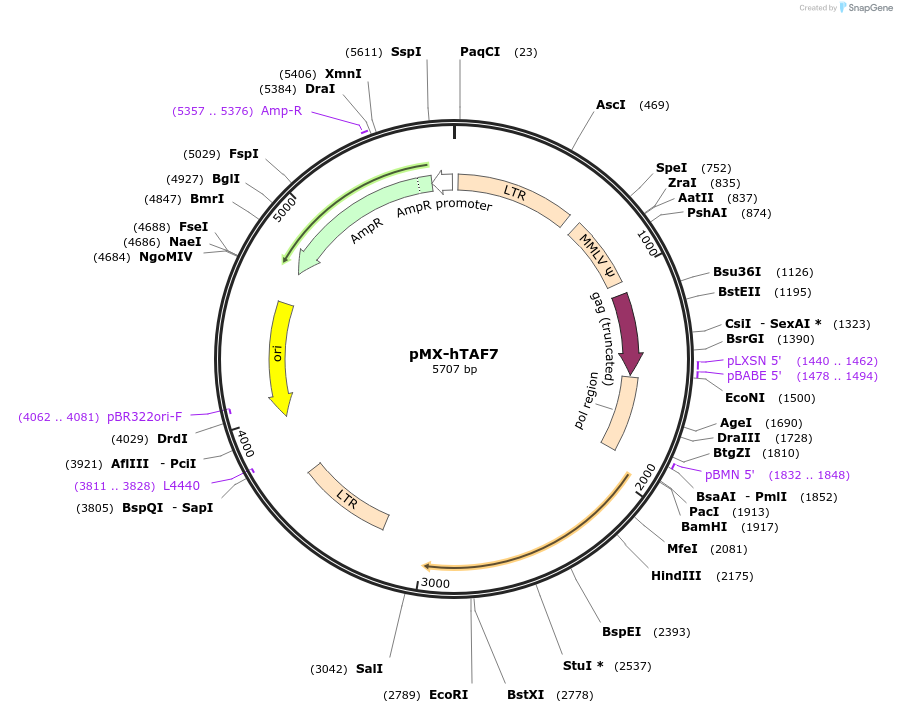
Plasmid#44338DepositorAvailable SinceMay 15, 2013AvailabilityAcademic Institutions and Nonprofits only
-
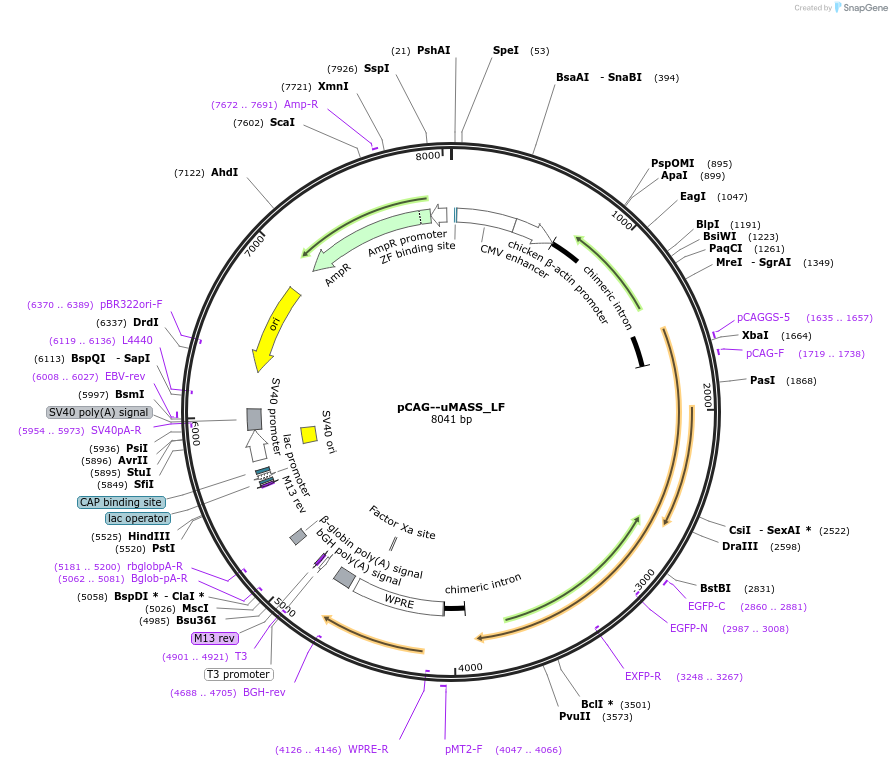
pCAG--uMASS_LF
Plasmid#209763PurposeExpression of uMASS_LF (loss-of-function) in mammalian cells under CAG promoterDepositorInsertuMASS_LF
ExpressionMammalianAvailable SinceNov. 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
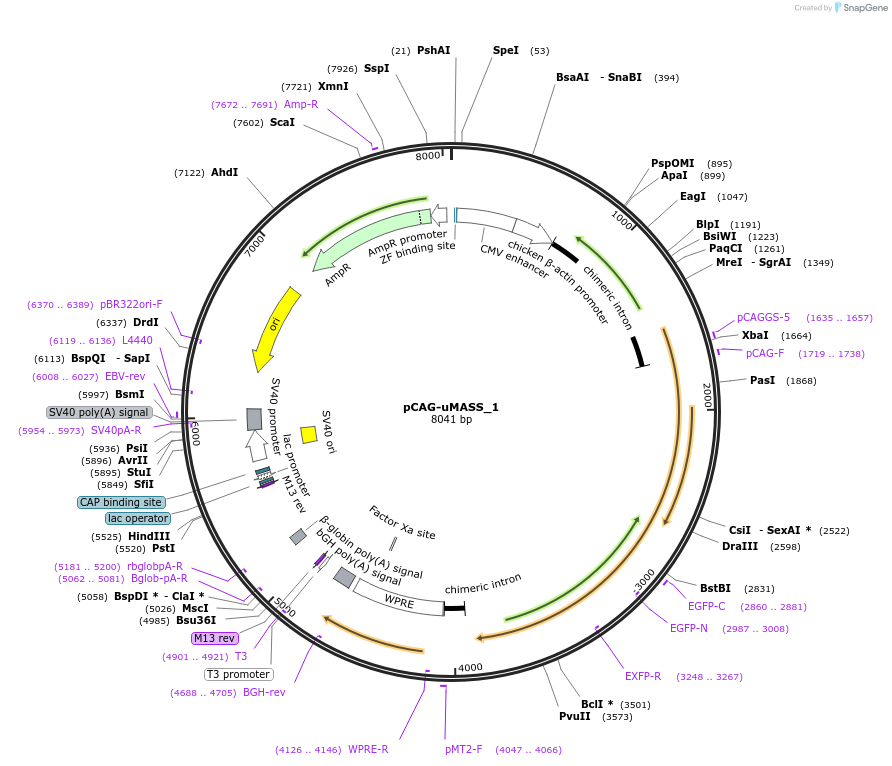
pCAG-uMASS_1
Plasmid#209762PurposeExpression of uMASS_1 in mammalian cells under CAG promoterDepositorInsertuMASS_1
ExpressionMammalianAvailable SinceNov. 13, 2023AvailabilityAcademic Institutions and Nonprofits only -
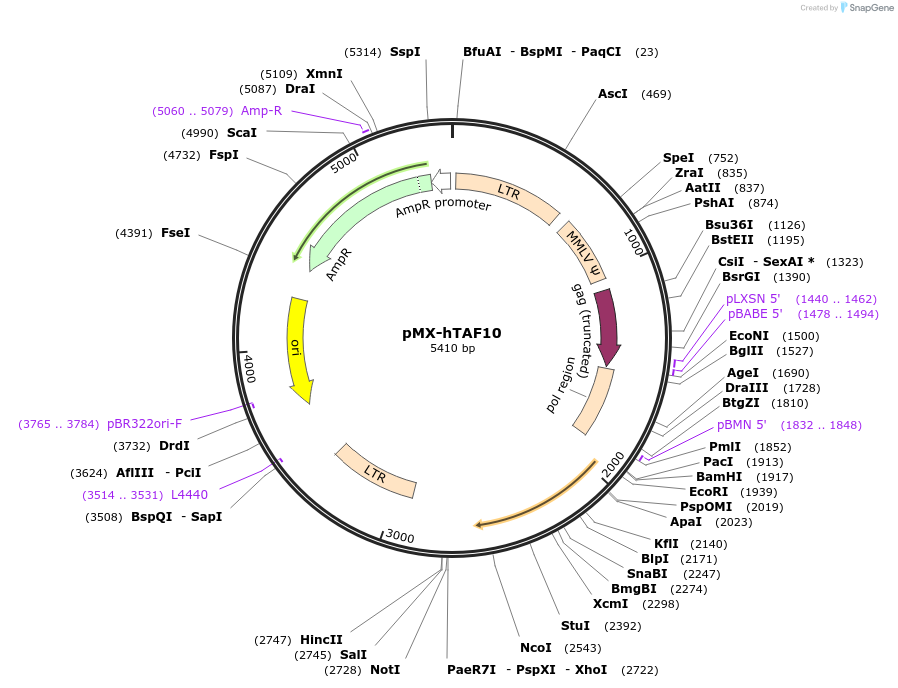
pMX-hTAF10
Plasmid#44340DepositorAvailable SinceJuly 19, 2013AvailabilityAcademic Institutions and Nonprofits only -
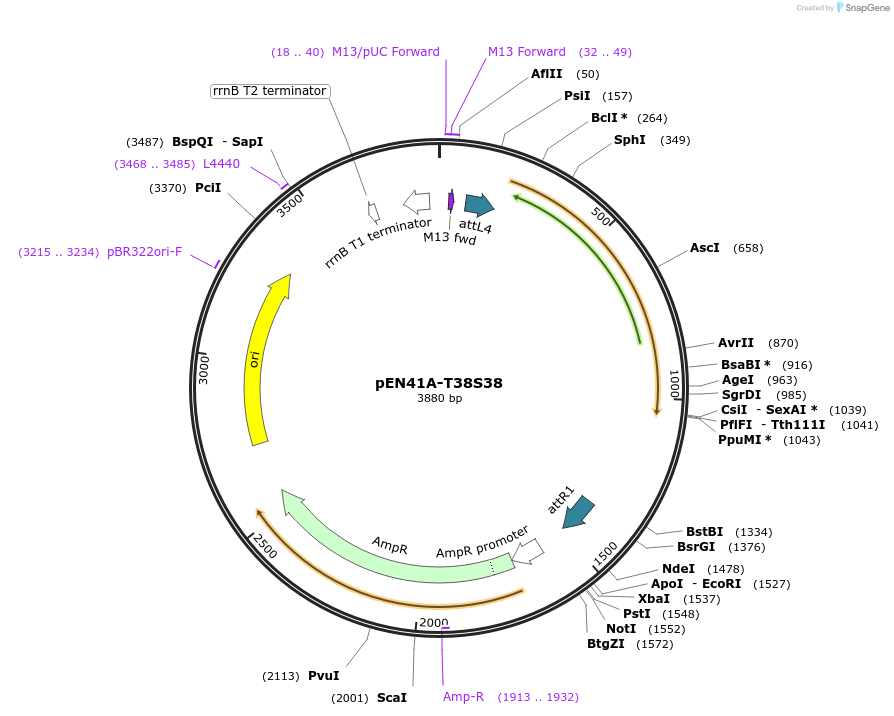
pEN41A-T38S38
Plasmid#49524PurposeAmpicillin resistant; entry clone encoding reverse TetR (tetR38) and contains attL4 and attL1 sites for recombination into a Gateway destination vector at the attR4 and attR1 sitesDepositorInsertconstitutive TetR38
PromoterPtb38Available SinceJan. 17, 2014AvailabilityAcademic Institutions and Nonprofits only -
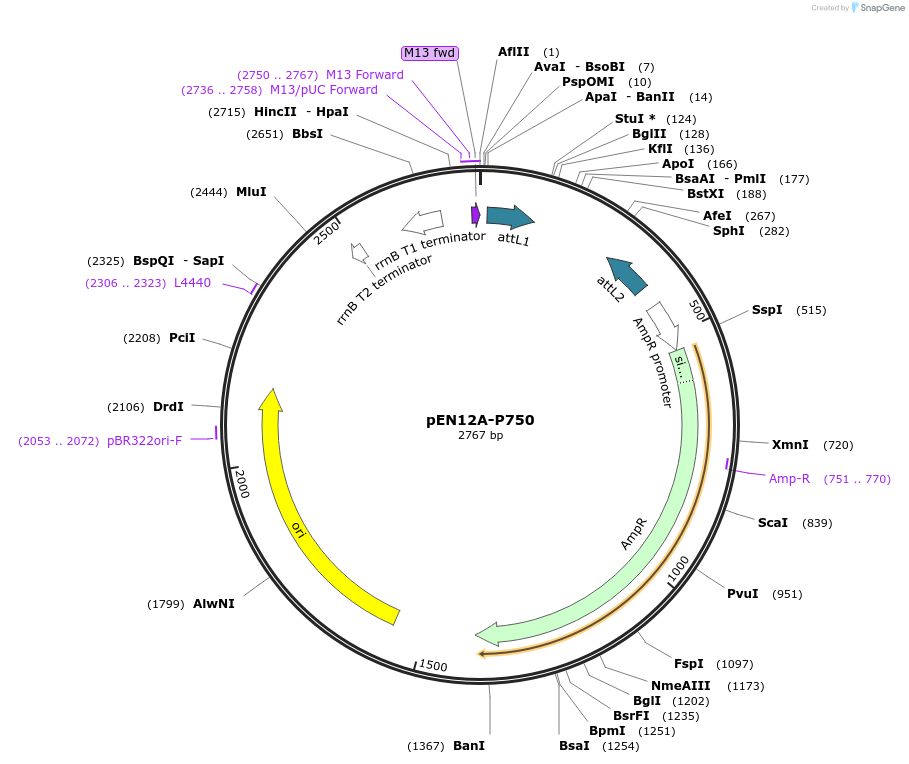
pEN12A-P750
Plasmid#49525PurposeAmpicillin resistant; entry clone containing PtetO-4C5G and attL1 and attL2 sites for recombination into a Gateway destination vector at the attR4 and attR1 sitesDepositorInsertP750
PromoterP750Available SinceJan. 17, 2014AvailabilityAcademic Institutions and Nonprofits only -
pGEX-6p-hDcp2
Plasmid#72216Purposebacterial expression of GST-tagged human Dcp2DepositorInsertDcp2 (DCP2 Human)
TagsGSTExpressionBacterialMutationF146Y and E198K (please see depositor's comm…PromotertacAvailable SinceDec. 21, 2015AvailabilityAcademic Institutions and Nonprofits only -
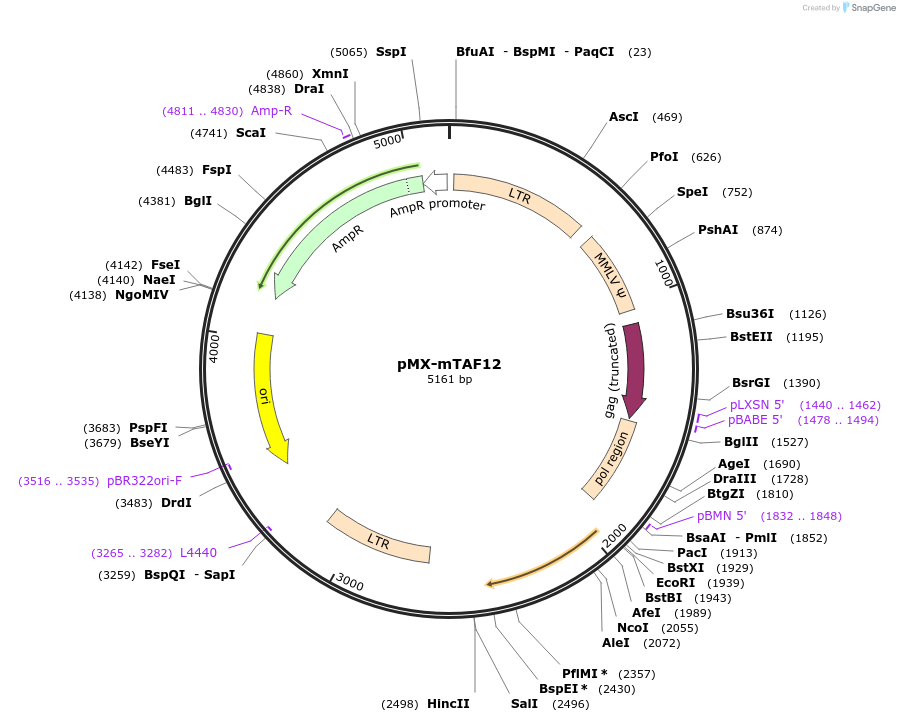
pMX-mTAF12
Plasmid#44342DepositorAvailable SinceMay 14, 2013AvailabilityAcademic Institutions and Nonprofits only -
-
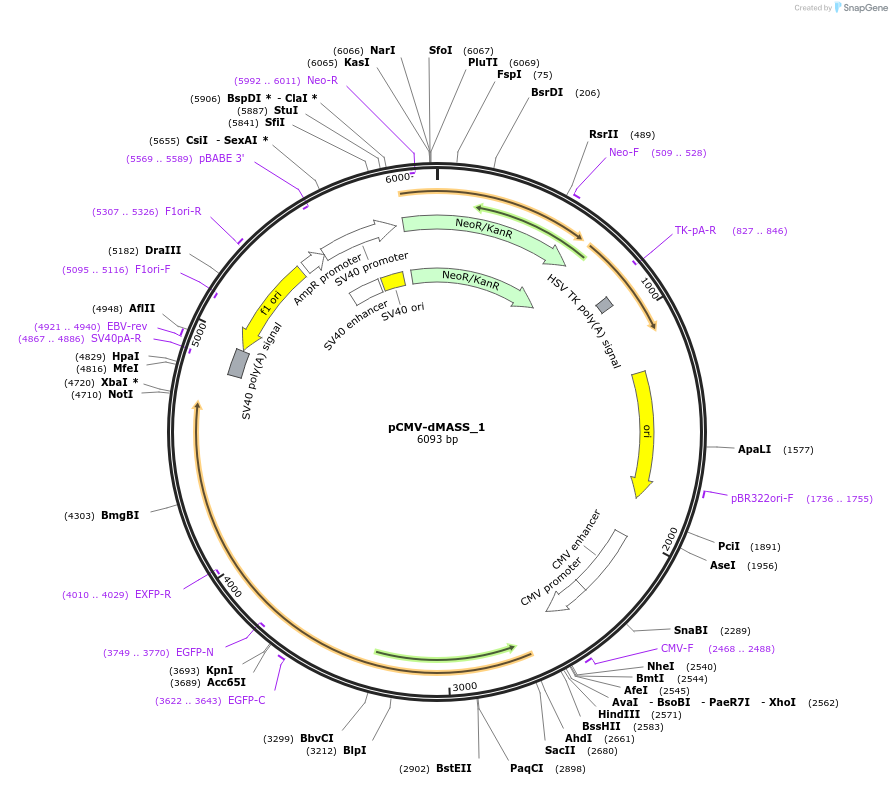
pCMV-dMASS_1
Plasmid#209764PurposeExpression of dMASS_1 in mammalian cells under CMV promoterDepositorInsertdMASS_1
ExpressionMammalianAvailable SinceNov. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
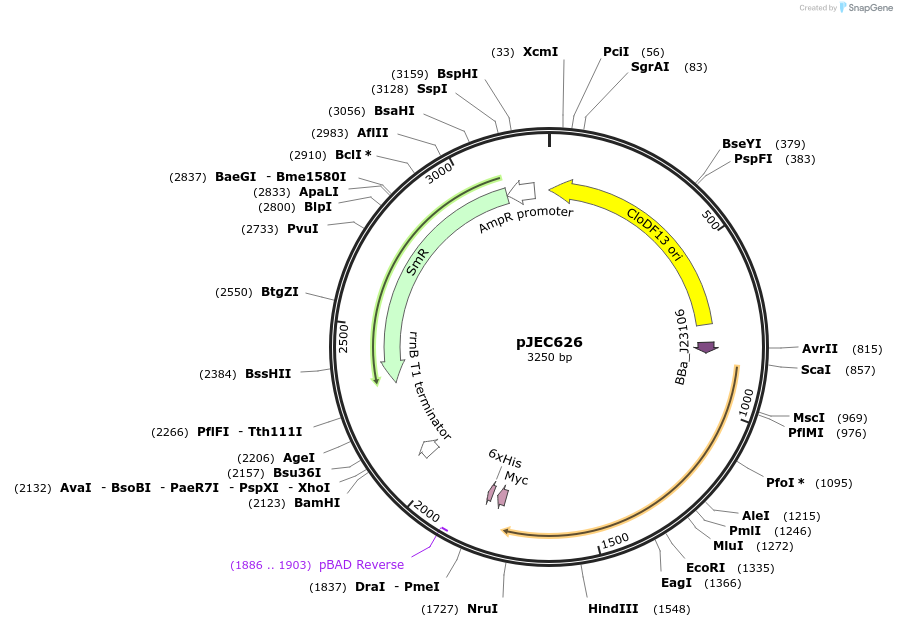
pJEC626
Plasmid#174367PurposeActivation domain for modular CRISPRa. AD-SYNZIP. Expresses alphaNTD from the E coli RNAP with a SYNZIP17 domain fused to the N terminus.DepositorInsertAlpha N-terminal domain from the E. coli RNA polymerase fused to SYNZIP17
UseSynthetic BiologyExpressionBacterialPromoterJ23106Available SinceOct. 20, 2021AvailabilityAcademic Institutions and Nonprofits only -
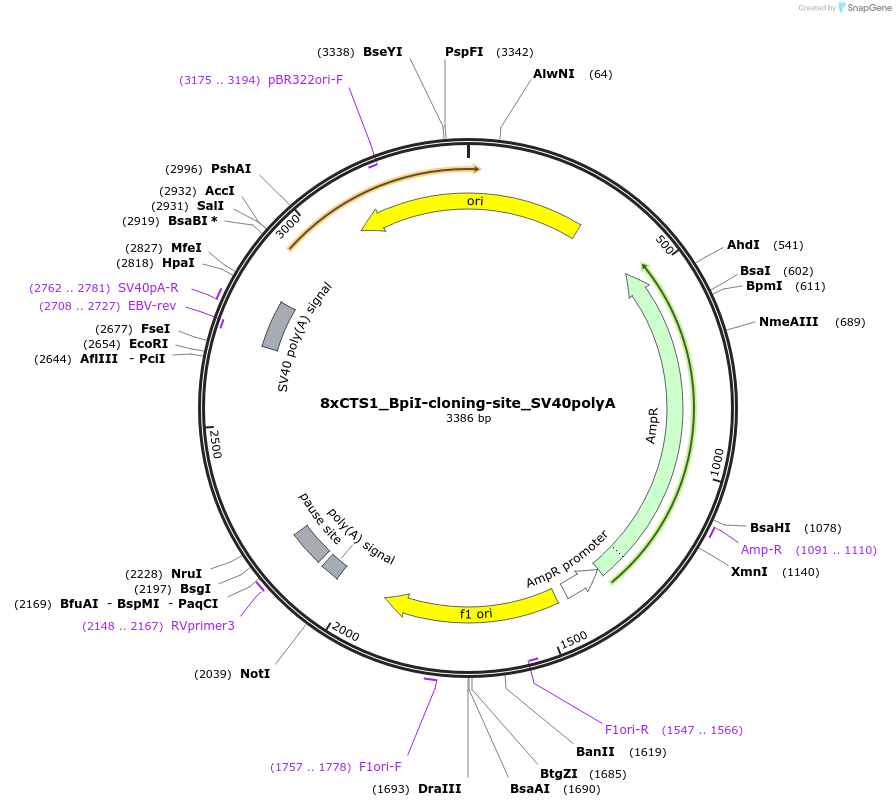
8xCTS1_BpiI-cloning-site_SV40polyA
Plasmid#124547PurposeDestination cloning vector for tRNA-flanked sgRNA. A BpiI placeholder is present between a minimal ADE promoter and SV40 polyA sequence.DepositorTypeEmpty backboneExpressionMammalianAvailable SinceApril 18, 2019AvailabilityAcademic Institutions and Nonprofits only -
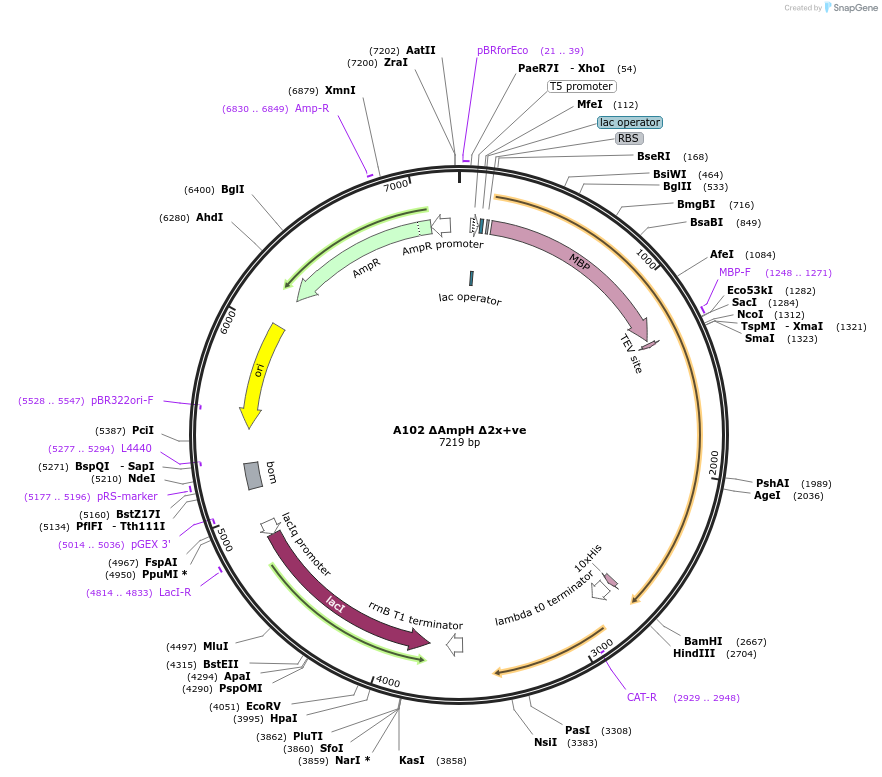
A102 ΔAmpH Δ2x+ve
Plasmid#129484PurposeExpression of Glo3 ΔAmpH lacking the COPI-binding motif in E.ColiDepositorInsertADP-ribosylation factor GTPase-activating protein (GLO3 Budding Yeast)
TagsHis and MBPExpressionBacterialAvailable SinceSept. 25, 2020AvailabilityAcademic Institutions and Nonprofits only -
pet 3d alpha tentacle delete CP
Plasmid#13299DepositorInsertcapping protein alpha 1 (deletion 28 amino acids) beta 1 subunits (CAPZB Chicken)
ExpressionBacterialAvailable SinceDec. 15, 2006AvailabilityAcademic Institutions and Nonprofits only -
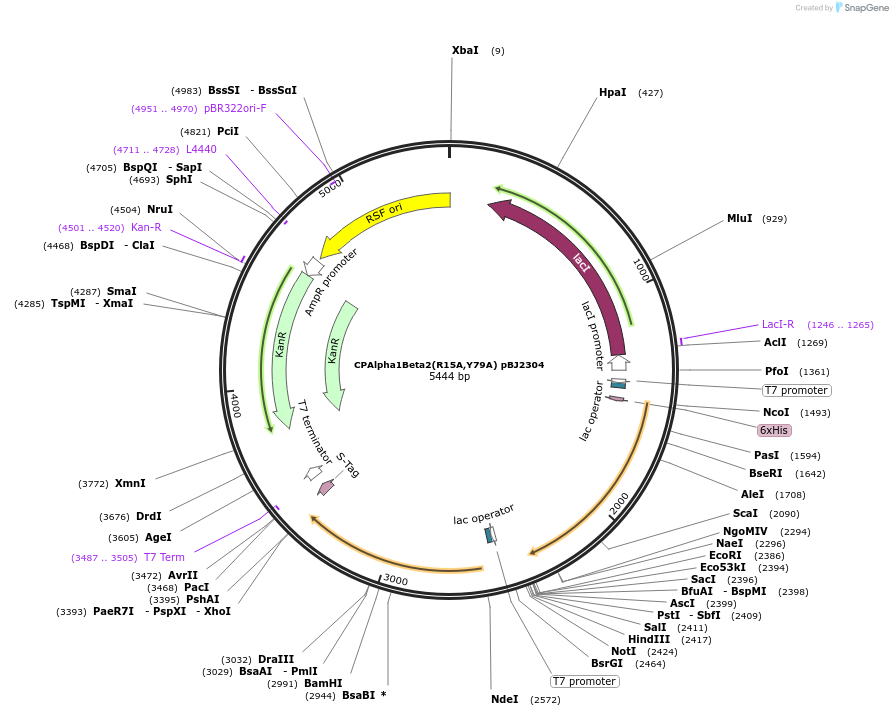
CPAlpha1Beta2(R15A,Y79A) pBJ2304
Plasmid#253114PurposepBJ2304 Cooper Lab. Bacterial expression of His-tagged mouse CP heterodimer (alpha1, beta2 isoforms). Two point mutations of the beta2 subunit (R15A, Y79A). Little to no binding of CPI motifs.DepositorInsertMouse capping protein alpha1 beta2 (R15A,Y79A). His-tagged.
TagsHisExpressionBacterialMutationCP beta2 (R15 to A, Y79 to A).PromoterT7 lacAvailable SinceMarch 30, 2026AvailabilityAcademic Institutions and Nonprofits only -
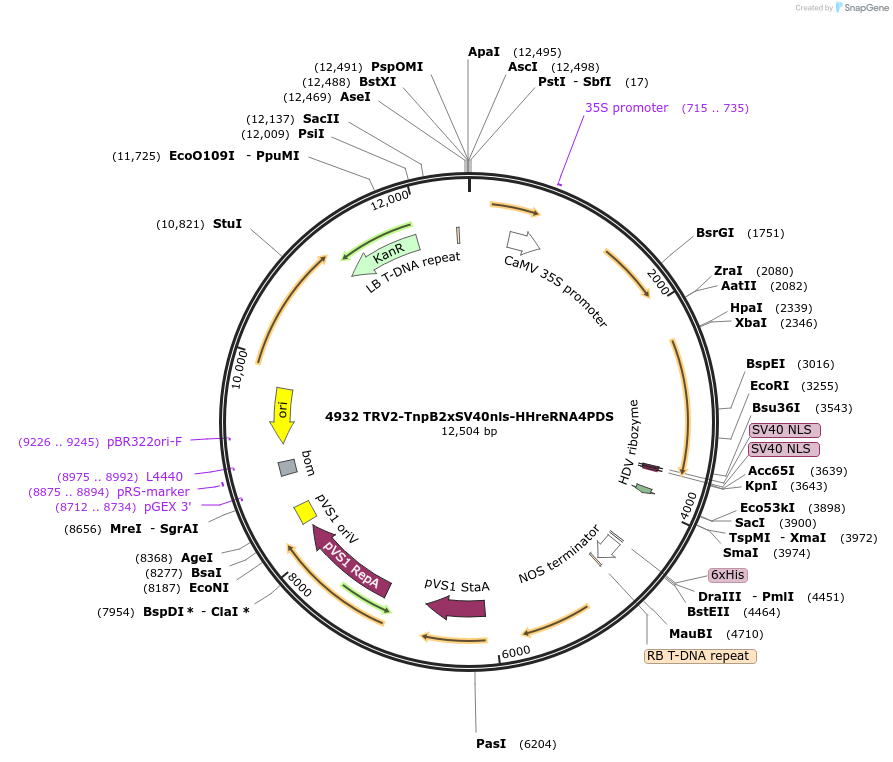
4932 TRV2-TnpB2xSV40nls-HHreRNA4PDS
Plasmid#251821PurposeEditing Photene desaturase (PDS) genes in Nicotiana benthamianaDepositorInsertISDra2 TnpB with guide targeting NbPDS flanked by HH-HDV ribozymes
Tags2xSV40nlsExpressionPlantAvailable SinceMarch 9, 2026AvailabilityAcademic Institutions and Nonprofits only -
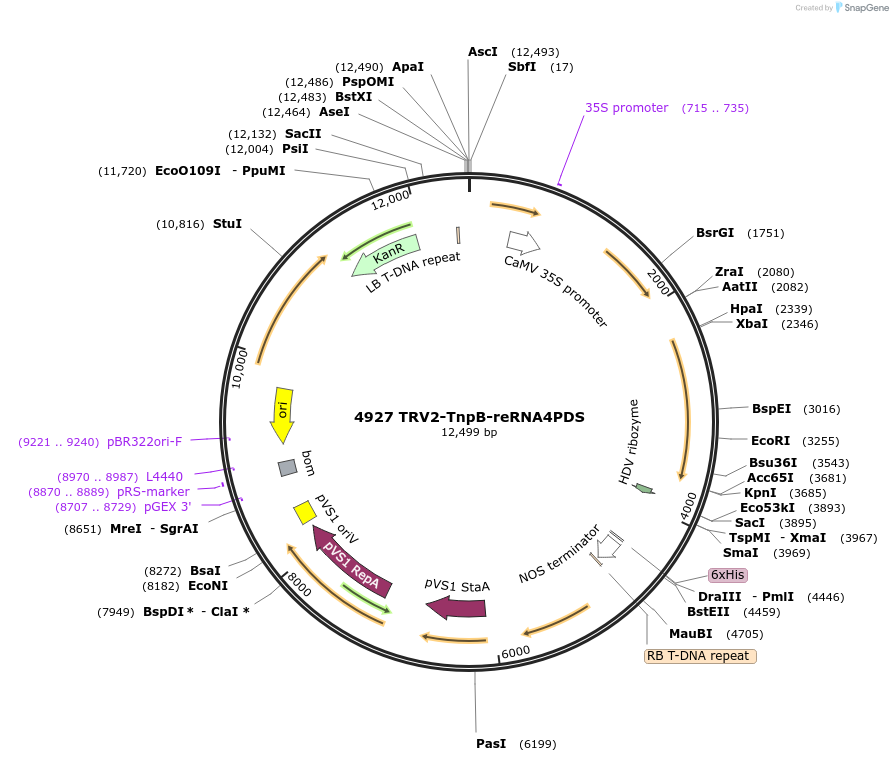
4927 TRV2-TnpB-reRNA4PDS
Plasmid#251814PurposeEditing Photene desaturase (PDS) genes in Nicotiana benthamianaDepositorInsertISDra2 TnpB with guide targeting NbPDS with HDV at 3' end
TagsAtFDnlsExpressionPlantAvailable SinceMarch 9, 2026AvailabilityAcademic Institutions and Nonprofits only -
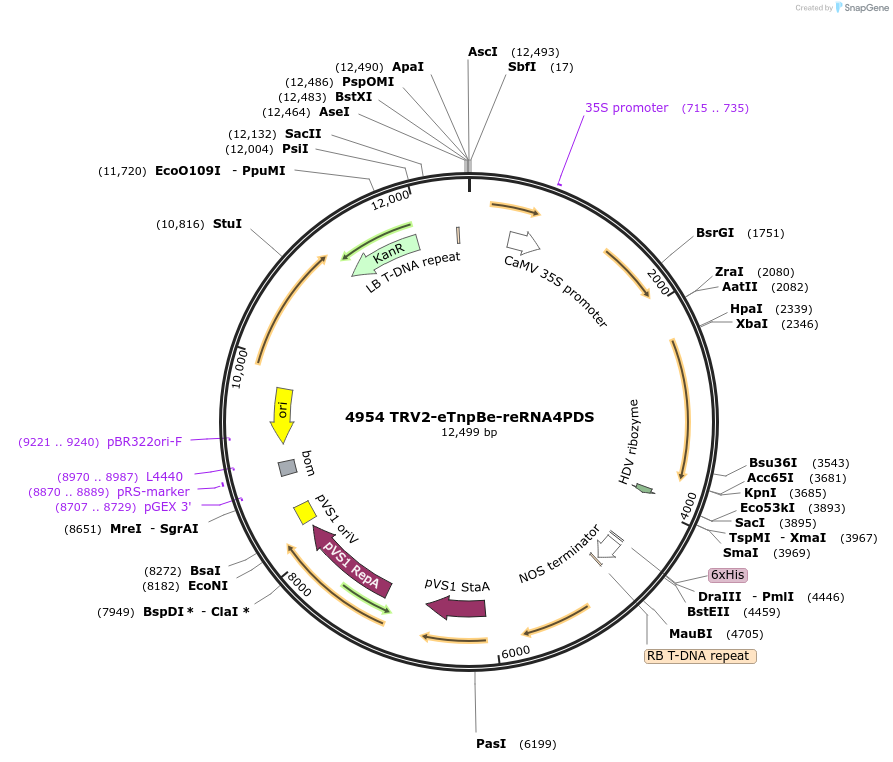
4954 TRV2-eTnpBe-reRNA4PDS
Plasmid#251818PurposeEditing Photene desaturase (PDS) genes in Nicotiana benthamianaDepositorInsertEnhanced variant of ISDra2 eTnpBe with guide targeting NbPDS with HDV at 3' end
TagsAtFDnlsExpressionPlantMutationL172G/V192L/L222I/P282V/I304RAvailable SinceMarch 9, 2026AvailabilityAcademic Institutions and Nonprofits only -
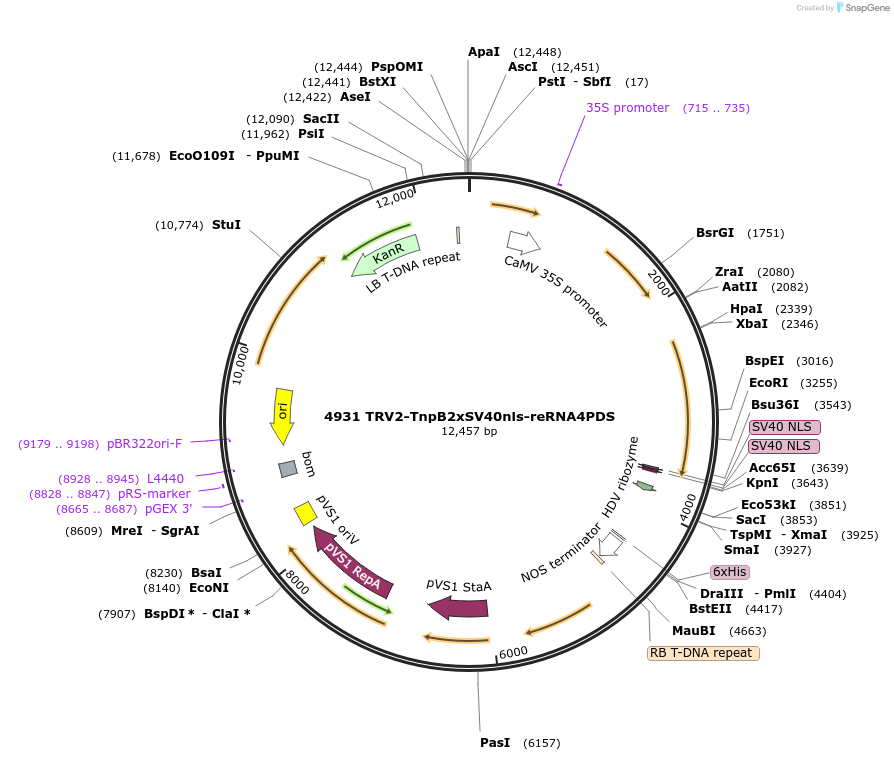
4931 TRV2-TnpB2xSV40nls-reRNA4PDS
Plasmid#251820PurposeEditing Photene desaturase (PDS) genes in Nicotiana benthamianaDepositorInsertISDra2 TnpB with guide targeting NbPDS with HDV at 3' end
Tags2xSV40nlsExpressionPlantAvailable SinceMarch 9, 2026AvailabilityAcademic Institutions and Nonprofits only -
pMKR07
Plasmid#240097PurposeA CRISPR‑Cas9 system for knock‑out and knock‑in of high molecular weight DNA enables module‑swapping of the pikromycin synthase in its native hostDepositorTypeEmpty backboneUseCRISPR and Synthetic BiologyExpressionBacterialPromoterermE* promoterAvailable SinceJuly 14, 2025AvailabilityAcademic Institutions and Nonprofits only -
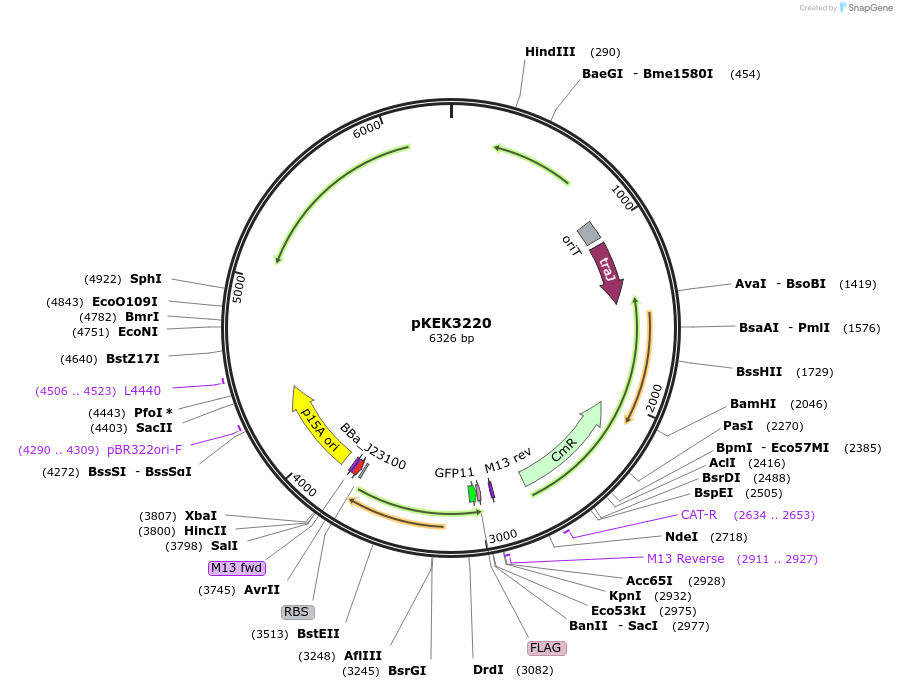
pKEK3220
Plasmid#234099PurposeCCP expression in diverse gram-negative bacteria, with pre-cloned Golden Gate Compatible (BsaI) restriction sites to facilitate promoter swapping.DepositorInsertsfGFP
TagsFLAGExpressionBacterialMutationS30R/Y39N/N105T/Y145F/I171V/A206VPromoterpJ23100-BsaIAvailable SinceApril 7, 2025AvailabilityAcademic Institutions and Nonprofits only -
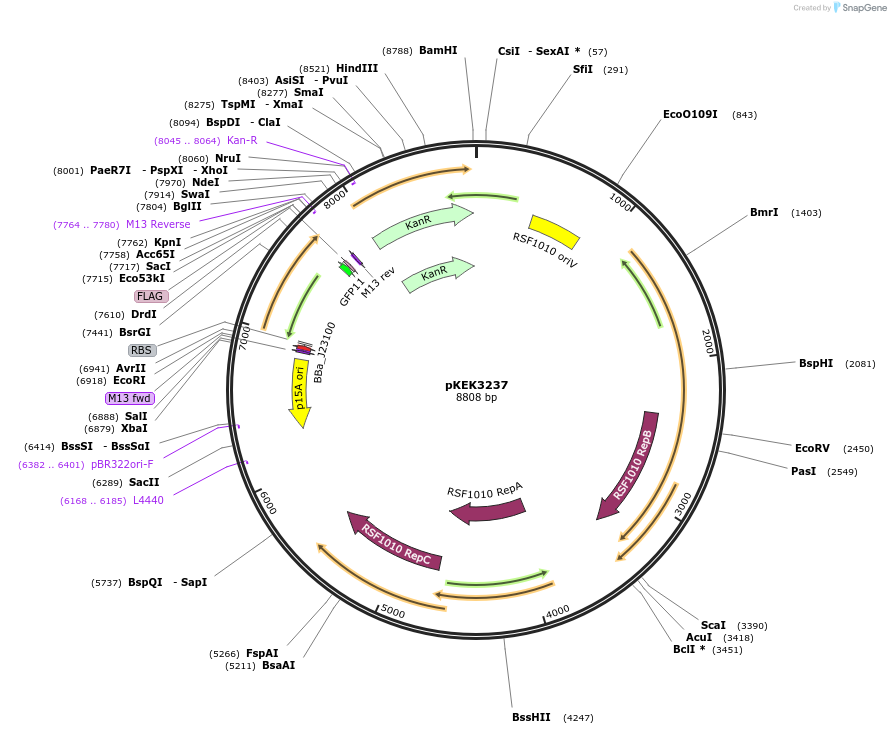
pKEK3237
Plasmid#234091PurposeCCP expression in diverse gram-negative bacteria, with pre-cloned Golden Gate Compatible (BsaI) restriction sites to facilitate promoter swapping.DepositorInsertsfGFP
TagsFLAGExpressionBacterialMutationS30R/Y39N/N105T/Y145F/I171V/A206VPromoterpJ23100-BsaIAvailable SinceApril 7, 2025AvailabilityAcademic Institutions and Nonprofits only -
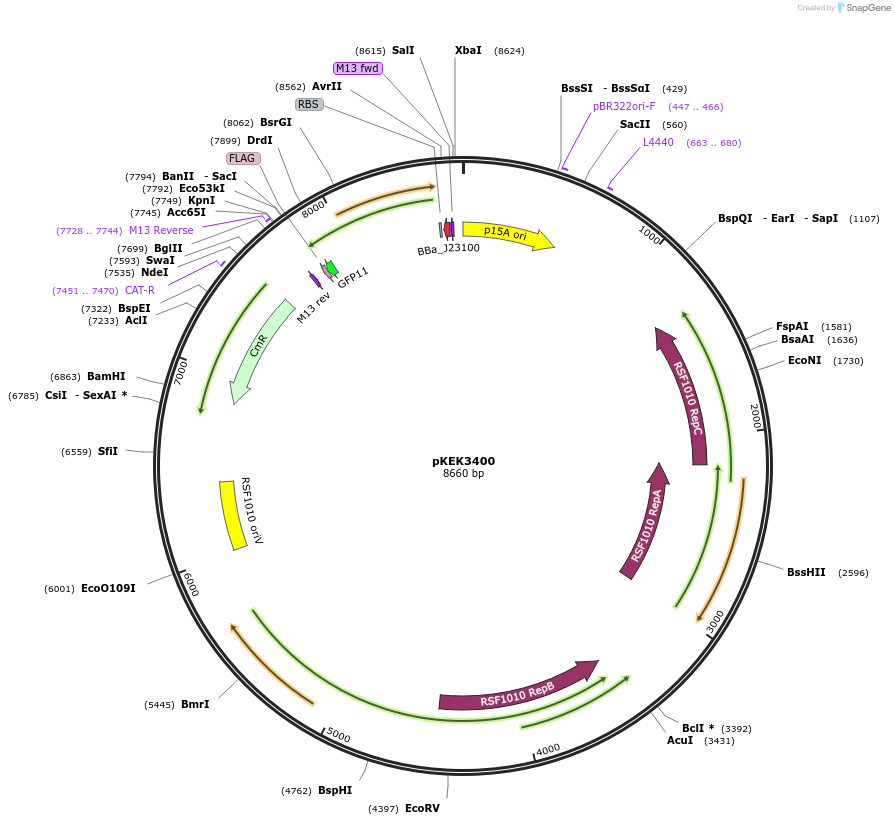
pKEK3400
Plasmid#234084PurposeCCP expression in diverse gram-negative bacteria, with pre-cloned Golden Gate Compatible (BsaI) restriction sites to facilitate promoter swapping.DepositorInsertsfGFP
TagsFLAGExpressionBacterialMutationS30R/Y39N/N105T/Y145F/I171V/A206VPromoterpJ23100-BsaIAvailable SinceMarch 25, 2025AvailabilityAcademic Institutions and Nonprofits only -
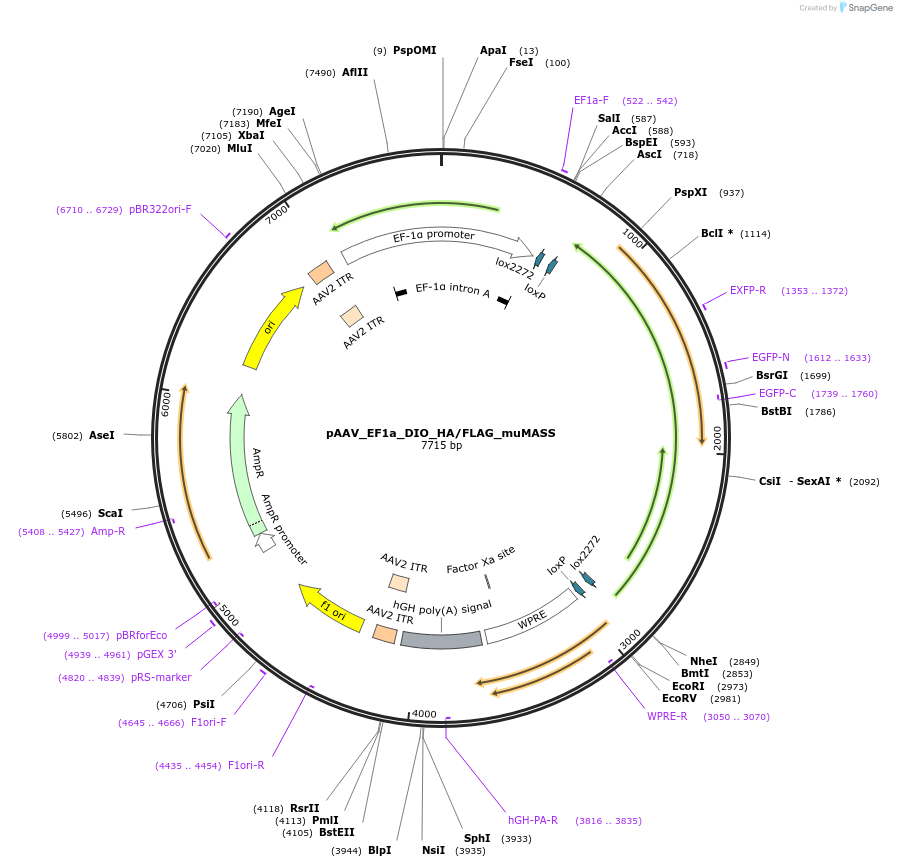
pAAV_EF1a_DIO_HA/FLAG_muMASS
Plasmid#213393PurposeuMASS_1 opioid sensor. AAV Virus production for Cre dependent expressionDepositorInsertuMASS_1
UseAAV and Cre/LoxExpressionMammalianAvailable SinceFeb. 13, 2024AvailabilityAcademic Institutions and Nonprofits only -
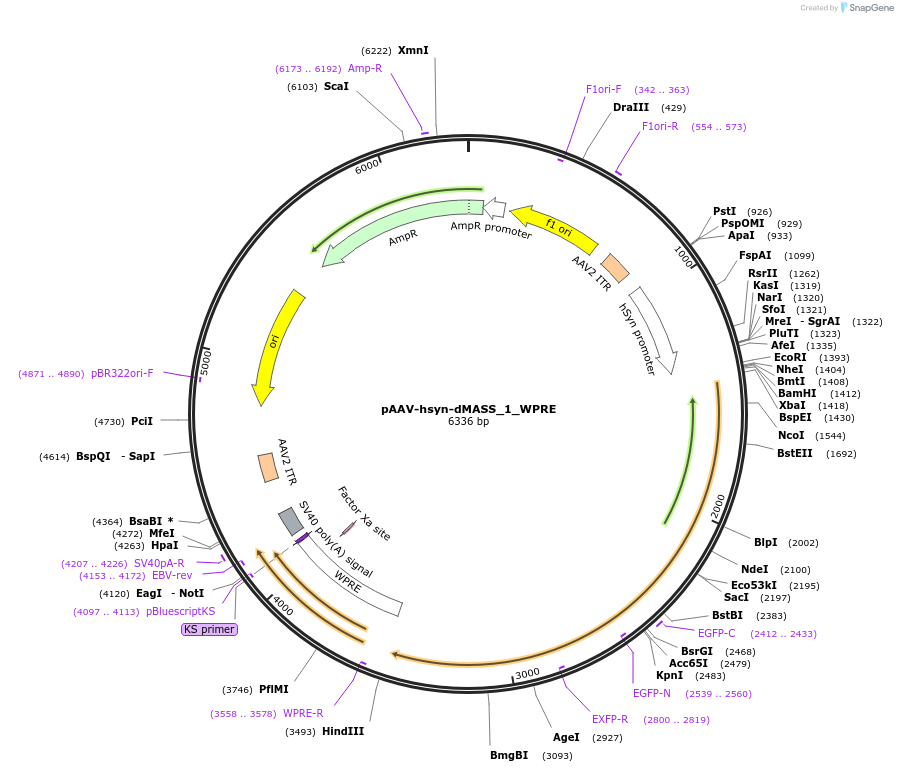
pAAV-hsyn-dMASS_1_WPRE
Plasmid#209765PurposeAAV virus production for neuronal expression of dMASS_1 under hSyn promoterDepositorInsertdMASS_1
UseAAVExpressionMammalianAvailable SinceNov. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
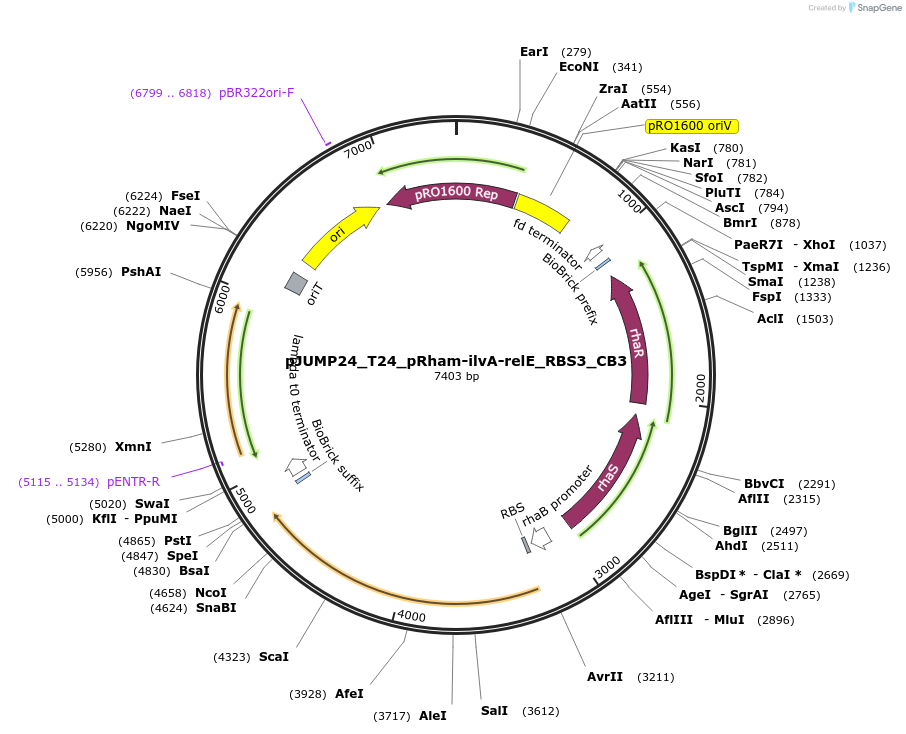
pJUMP24_T24_pRham-ilvA-relE_RBS3_CB3
Plasmid#201534PurposeExpressing synthetic overlapping gene, ilvA-relE with internal RBS modification. Evolved strain with lower ilvA-relE expression. Origin pRO1600/ColE1 (E.coli - Pseudomonas shuttle)DepositorInsertilvA-relE_RBS3_CB3
UseSynthetic BiologyMutationrhaS W190GPromoterPrhaBAD (rhamnose)Available SinceSept. 7, 2023AvailabilityAcademic Institutions and Nonprofits only -
-
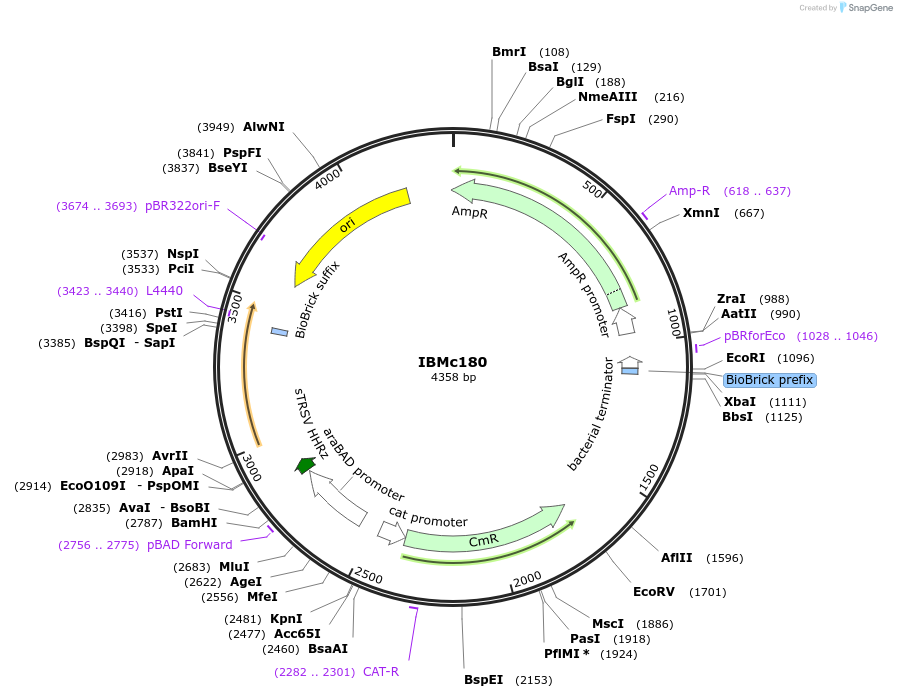
IBMc180
Plasmid#161942PurposeCarries Golden Gate substitution insert for acVHH-assisted bisection mappingDepositorInsertpSB1A3-BbsI-10aa-acVHH-T-cat-PBAD(SapI)-RiboJ-BCD22-acVHH-10aa-SapI
ExpressionBacterialPromoterPBADAvailable SinceMarch 30, 2021AvailabilityAcademic Institutions and Nonprofits only -
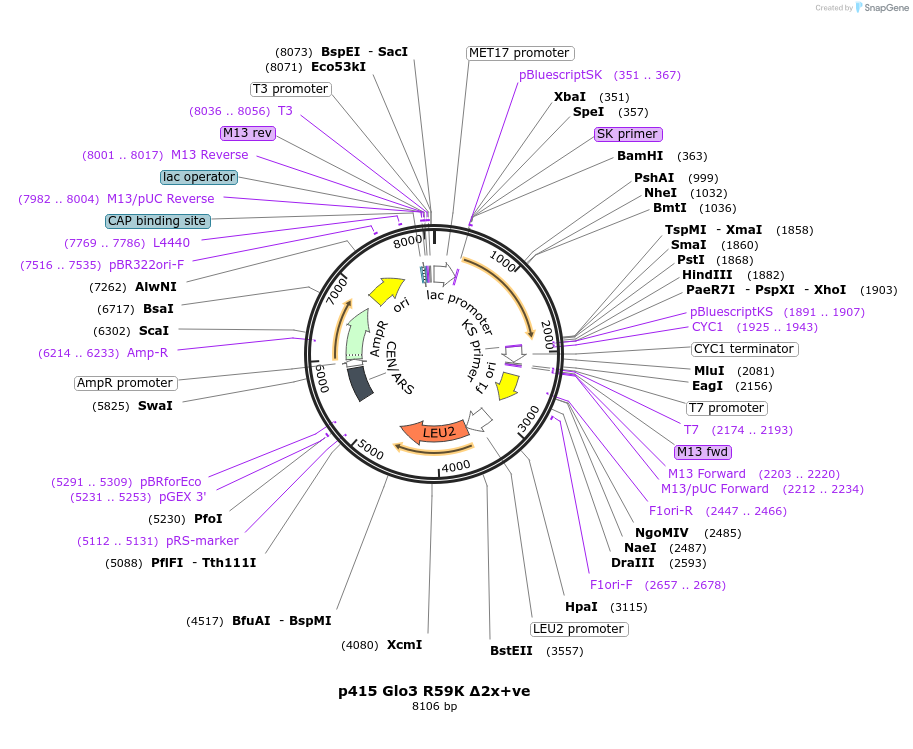
p415 Glo3 R59K Δ2x+ve
Plasmid#112655PurposeExpression S.cerevisiaeDepositorInsertGlo3 R59K Δ2x+ve
ExpressionYeastMutationR59K, mutations (R223E, K224E, K225E, K233E, K234…PromoterMet25Available SinceSept. 25, 2020AvailabilityAcademic Institutions and Nonprofits only -
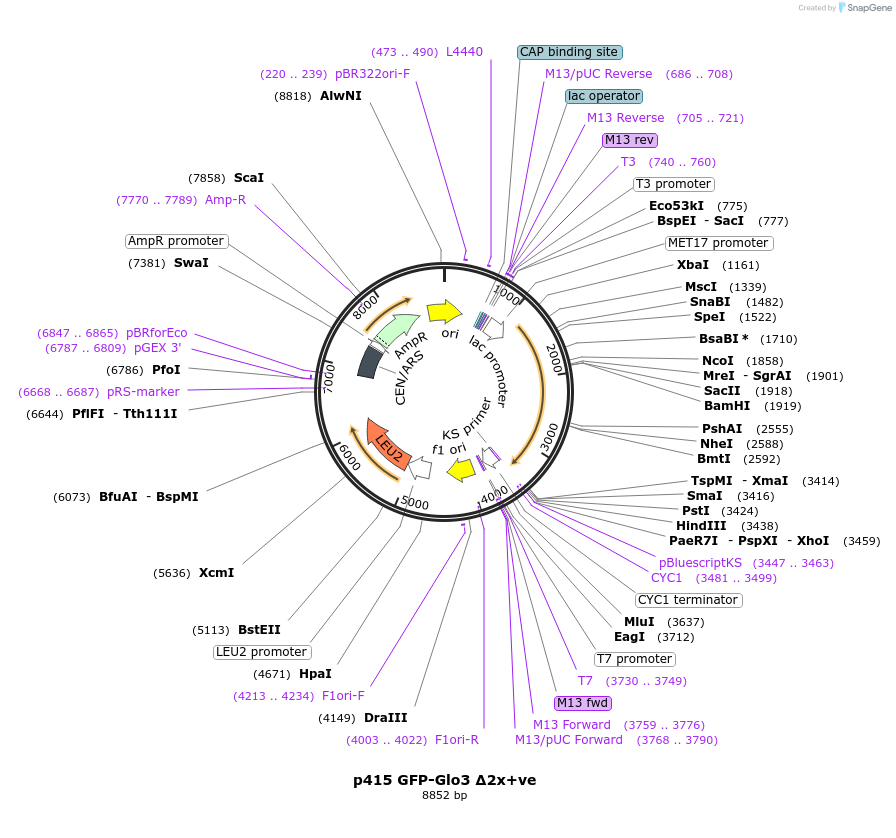
p415 GFP-Glo3 Δ2x+ve
Plasmid#112662PurposeExpression S.cerevisiaeDepositorInsertGFP- Glo3 Δ2x+ve
TagssfGFPExpressionYeastMutationMutations (R223E, K224E, K225E, K233E, K234E, K23…PromoterMet25Available SinceSept. 25, 2020AvailabilityAcademic Institutions and Nonprofits only -
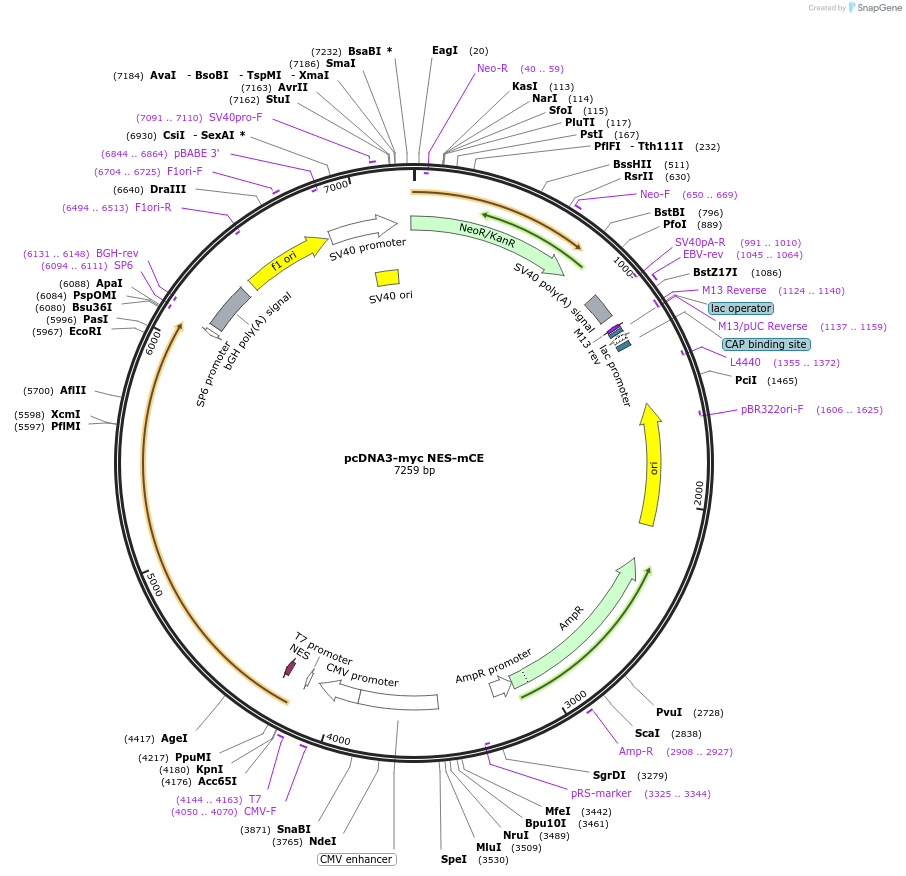
pcDNA3-myc NES-mCE
Plasmid#82463PurposeExpression of myc epitope and murine mRNA Capping Enzyme with Terminal nuclear export signal in mammalian cellsDepositorAvailable SinceSept. 9, 2016AvailabilityAcademic Institutions and Nonprofits only -
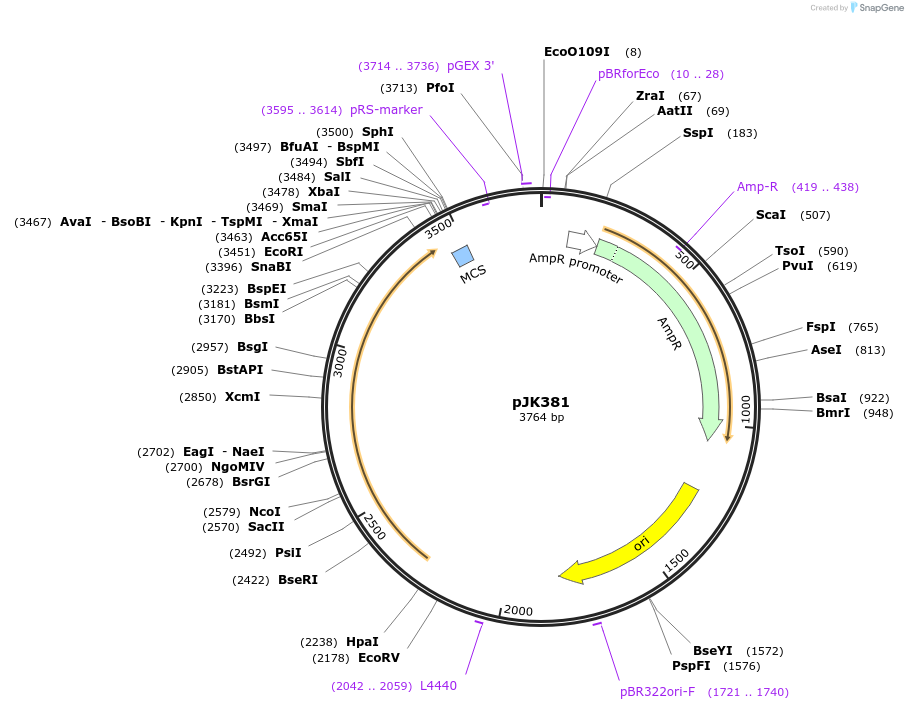
pJK381
Plasmid#71573PurposeProduces Thermoplasma acidophilum citrate synthase, Asp317>Gly mutant (TpCS-D317G)DepositorInsertcitrate synthase
ExpressionBacterialMutationchanges aspartate-317 to glycinePromoterrecAAvailable SinceApril 8, 2016AvailabilityAcademic Institutions and Nonprofits only -
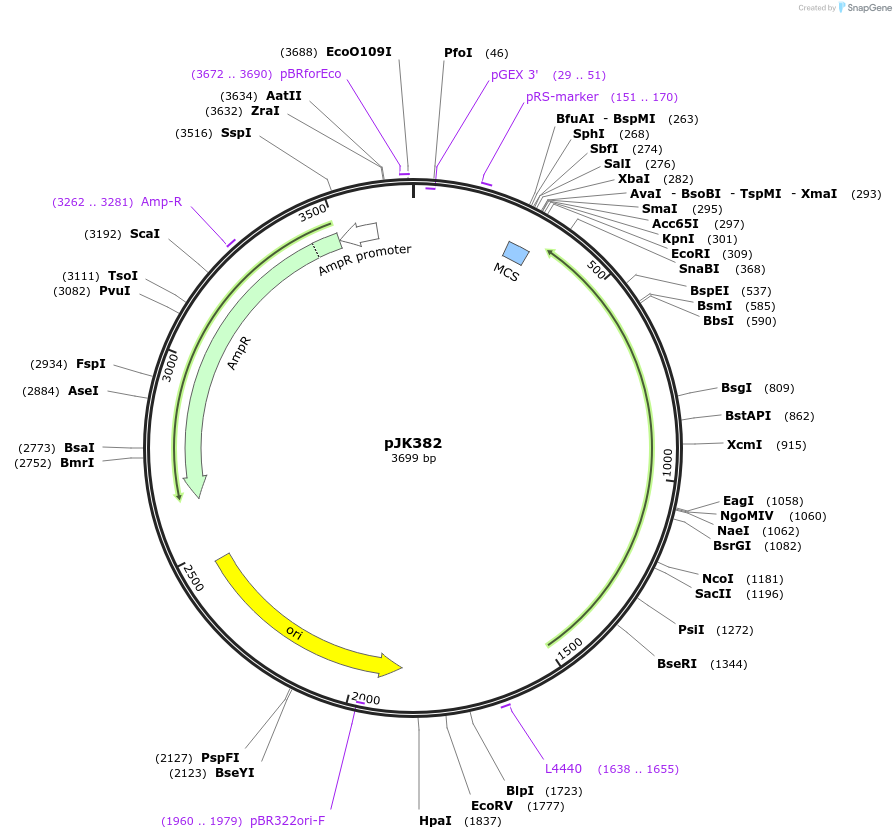
pJK382
Plasmid#71574PurposeProduces Thermoplasma acidophilum citrate synthase, Asp317>Asn mutant (TpCS-D317N)DepositorInsertcitrate synthase
ExpressionBacterialMutationchanges aspartate-317 to asparagine; silent mutat…PromoterrecAAvailable SinceMarch 25, 2016AvailabilityAcademic Institutions and Nonprofits only -
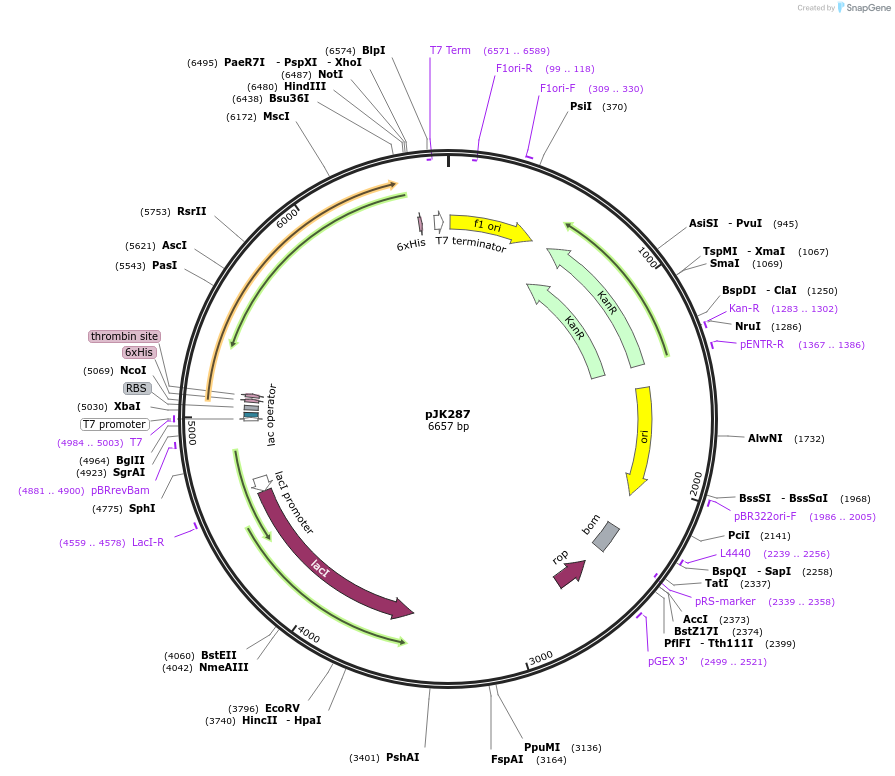
pJK287
Plasmid#71120PurposeProduces Acetobacter aceti 1023 formyl-CoA:oxalate CoA-transferase (H6UctB) in E. coliDepositorInsertformyl-CoA:oxalate CoA-transferase
TagsHis6ExpressionBacterialPromoterT7Available SinceFeb. 26, 2016AvailabilityAcademic Institutions and Nonprofits only -
pJK360
Plasmid#71571PurposeProduces Acetobacter aceti 1023 acetyl-CoA:oxalate CoA-transferase (UctC) with Thr2>Ala mutant (UctC-T2A)DepositorInsertacetyl-CoA:oxalate CoA-transferase
ExpressionBacterialMutationchanges threonine-2 to alaninePromoterT7Available SinceJan. 19, 2016AvailabilityAcademic Institutions and Nonprofits only -
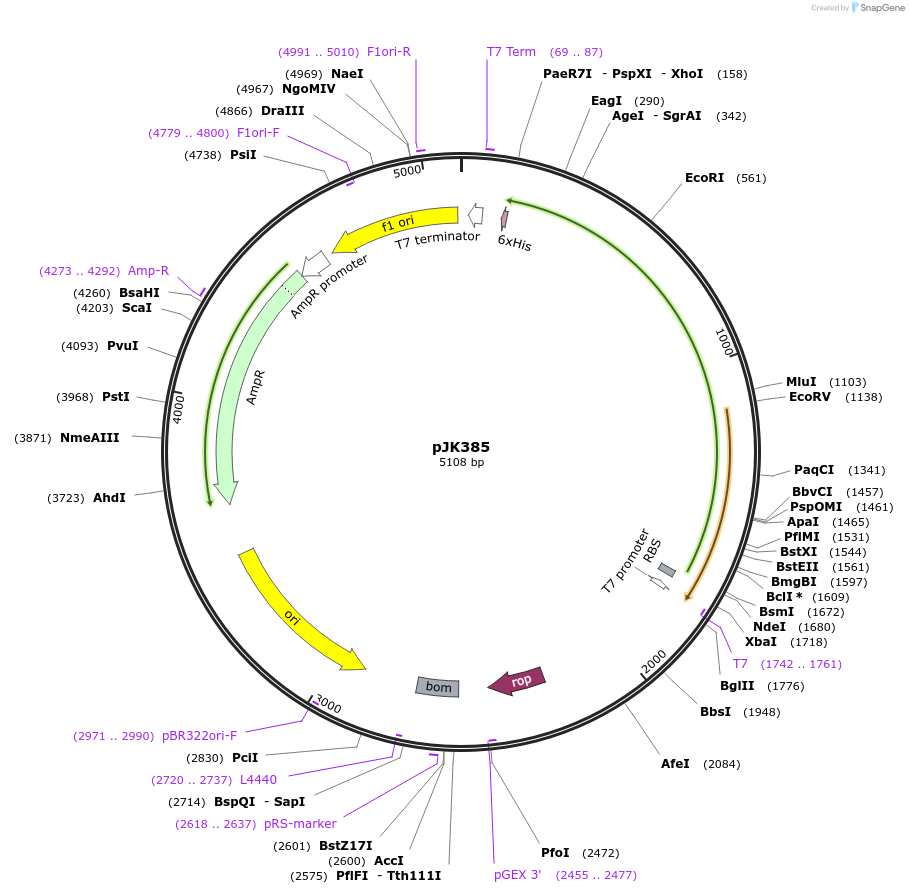
pJK385
Plasmid#71572PurposeProduces Acetobacter aceti 1023 succinyl-CoA:acetate CoA-transferase with C-terminal His6 tag (AarCH6)DepositorInsertsuccinyl-CoA:acetate CoA-transferase
TagsHis6ExpressionBacterialPromoterT7Available SinceJan. 19, 2016AvailabilityAcademic Institutions and Nonprofits only -
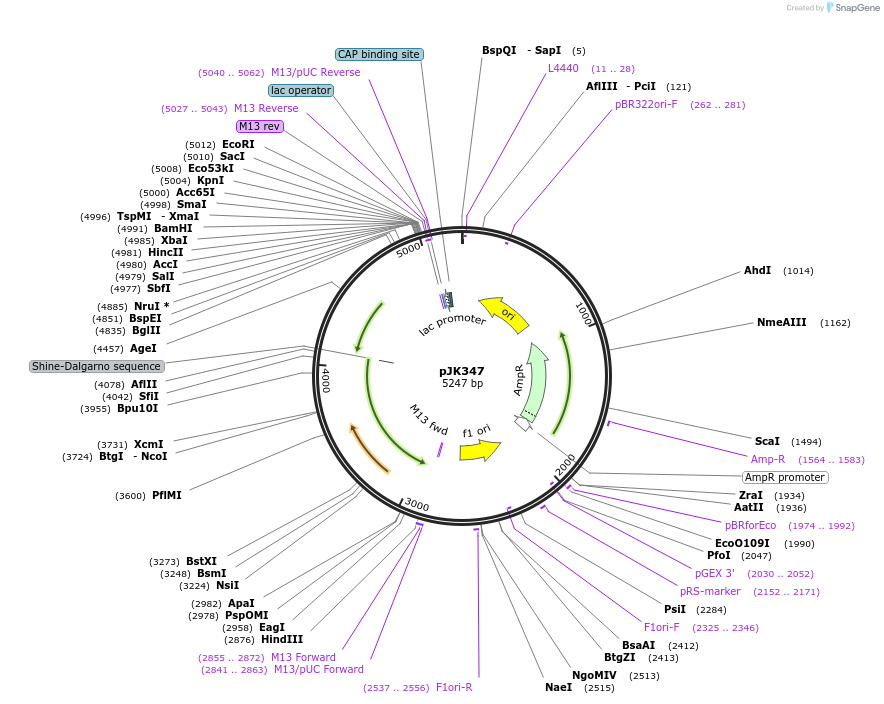
pJK347
Plasmid#71833PurposeCloned region contains Acetobacter aceti 1023 hypothetical protein AZ09_02685, purE(H59D), and purK genesDepositorInsertshypothetical protein
N5-carboxyaminoimidazole ribonucleotide mutase
N5-carboxyaminoimidazole ribonucleotide synthetase
ExpressionBacterialMutationchanges histidine-59 to aspartateAvailable SinceJan. 6, 2016AvailabilityAcademic Institutions and Nonprofits only -
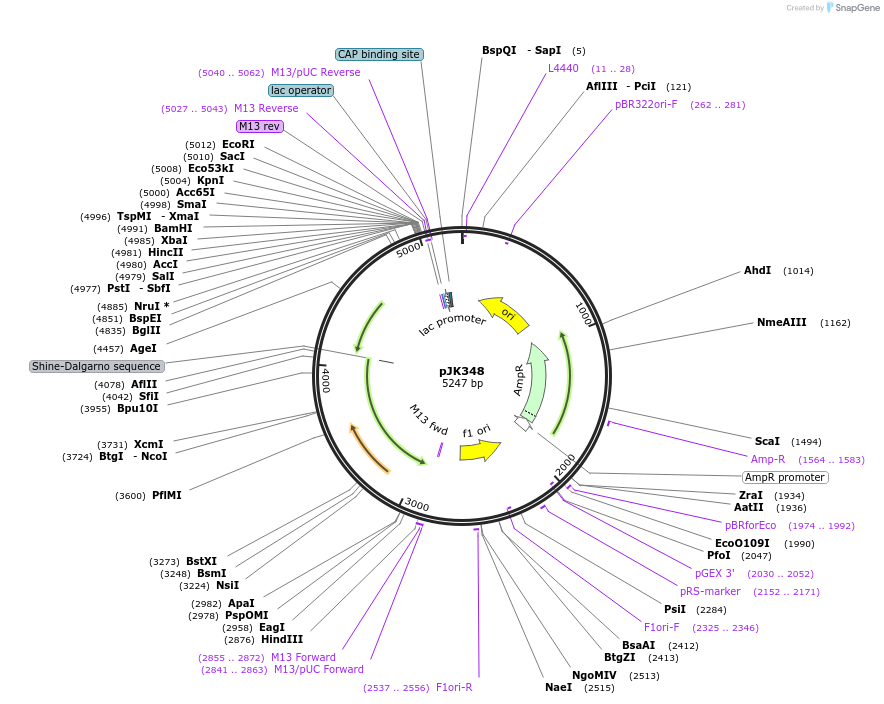
pJK348
Plasmid#71832PurposeCloned region contains Acetobacter aceti 1023 hypothetical protein AZ09_02685, purE(H59N), and purK genesDepositorInsertshypothetical protein
N5-carboxyaminoimidazole ribonucleotide mutase
N5-carboxyaminoimidazole ribonucleotide synthetase
ExpressionBacterialMutationchanges histidine-59 to asparagineAvailable SinceJan. 6, 2016AvailabilityAcademic Institutions and Nonprofits only -
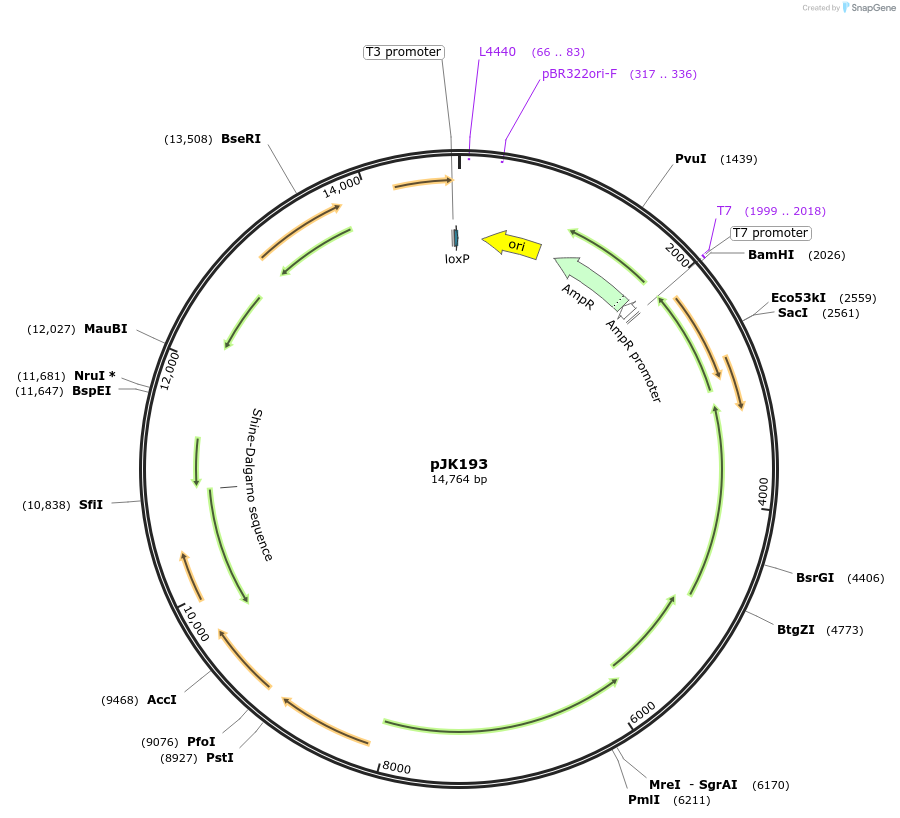
pJK193
Plasmid#71834PurposeCloned region contains Acetobacter aceti 1023 rpmE-radC(and beyond) regionDepositorInsertsN5-carboxyaminoimidazole ribonucleotide mutase
N5-carboxyaminoimidazole ribonucleotide synthetase
UsePhage lambda cloning vector with genomic dna inse…Mutationadditional N-terminal methionine encoded by this …Available SinceJan. 5, 2016AvailabilityAcademic Institutions and Nonprofits only -
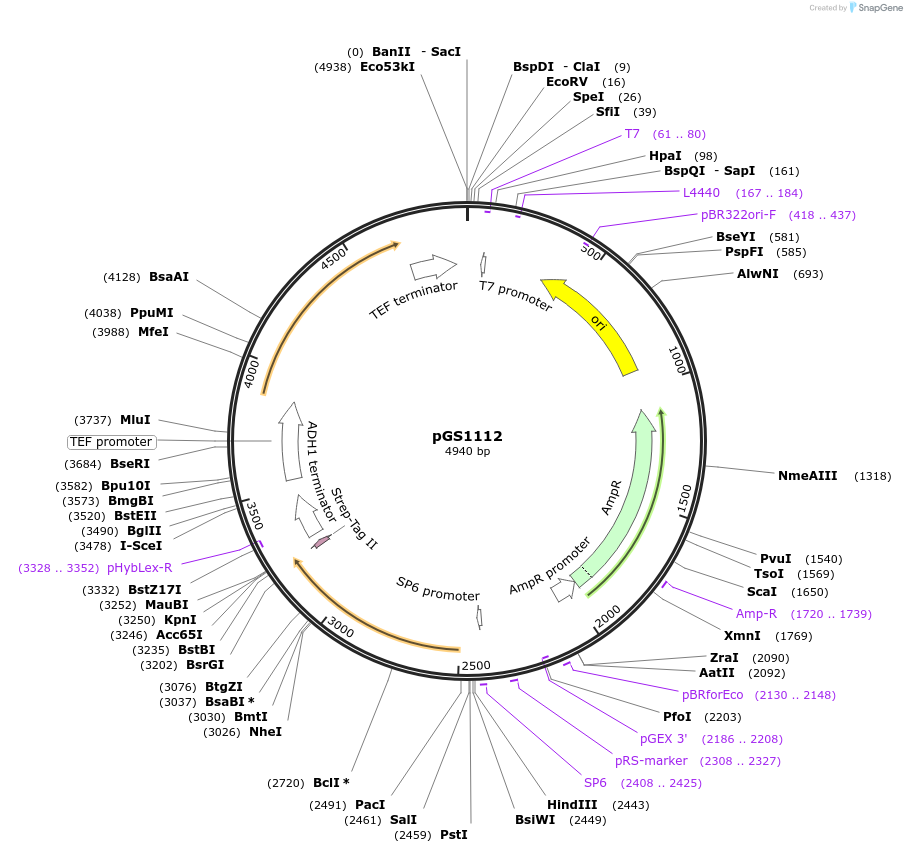
pGS1112
Plasmid#63784PurposeVector to create a cassette for homologous recombination in S. cerevisiae; adds a C-terminal eSapphire Strep-tag II followed by URA3 selection markerDepositorInserteSapphire-Strep-tag II followed by URA3
UseE. coli cloning vectorTagsSapphire-Strep-tag IIPromoternoneAvailable SinceApril 22, 2015AvailabilityAcademic Institutions and Nonprofits only -
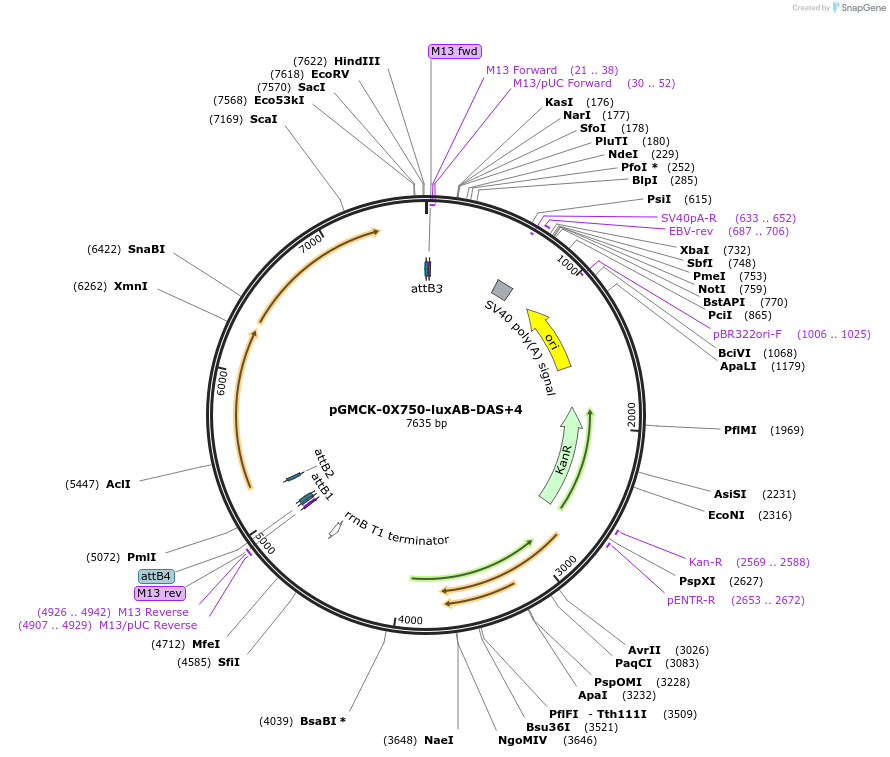
pGMCK-0X750-luxAB-DAS+4
Plasmid#49517PurposeKanamycin resistant; expresses luxAB-DAS+4 under the control of a promoter containing PtetO-4C5G; integrates into the mycobacteriophage att-L5 siteDepositorInsertP750-luxAB-DAS+4
TagsDAS+4 (AANDENYSENYADAS)ExpressionBacterialPromoterP750Available SinceJan. 17, 2014AvailabilityAcademic Institutions and Nonprofits only -
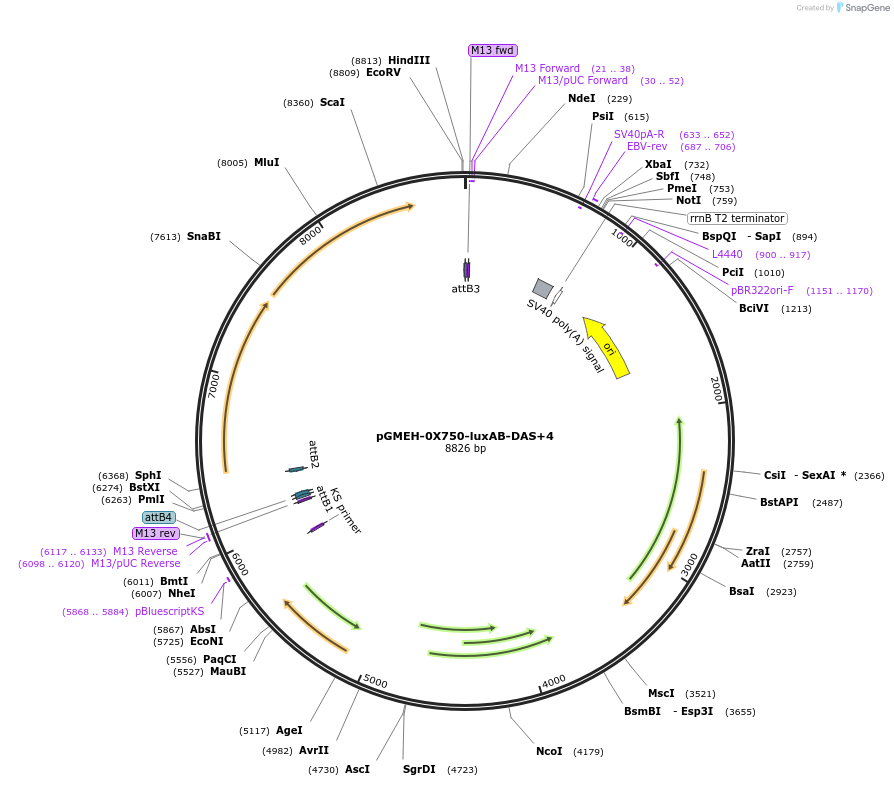
pGMEH-0X750-luxAB-DAS+4
Plasmid#49519PurposeHygromycin resistant; expresses luxAB-DAS+4 under the control of a promoter containing PtetO-4C5G; replicates episomallyDepositorInsertP750-luxAB-DAS+4
TagsDAS+4 (AANDENYSENYADAS)ExpressionBacterialPromoterP750Available SinceJan. 17, 2014AvailabilityAcademic Institutions and Nonprofits only -
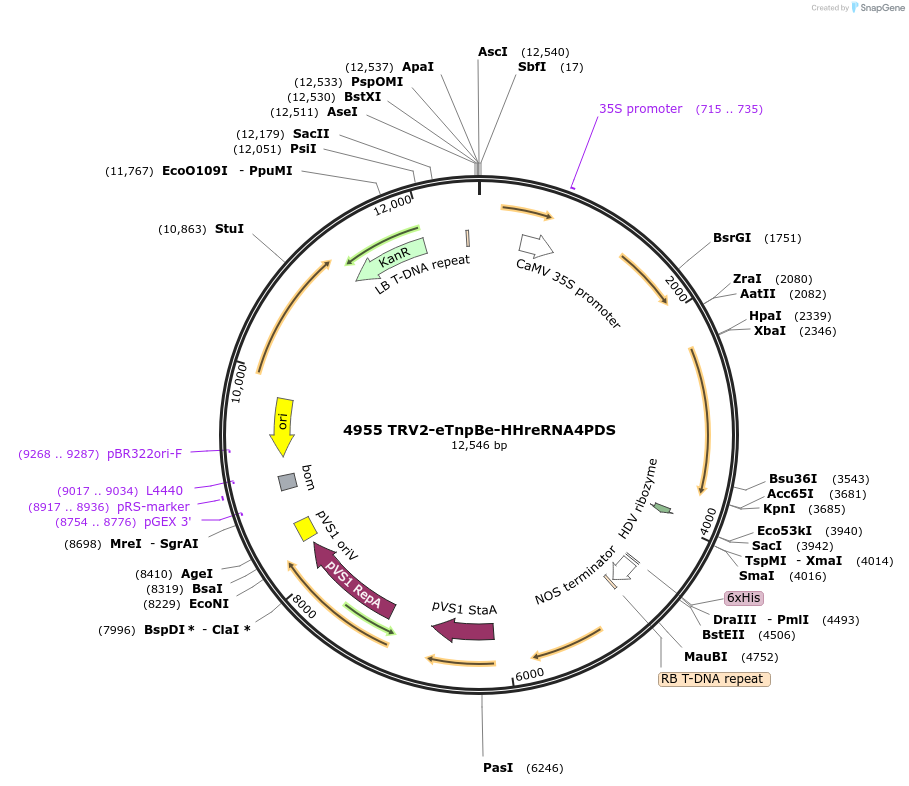
4955 TRV2-eTnpBe-HHreRNA4PDS
Plasmid#251819PurposeEditing Photene desaturase (PDS) genes in Nicotiana benthamianaDepositorInsertEnhanced variant of ISDra2 eTnpBe with guide targeting NbPDS flanked by HH-HDV ribozymes
TagsAtFDnlsExpressionPlantMutationL172G/V192L/L222I/P282V/I304RAvailable SinceMarch 9, 2026AvailabilityAcademic Institutions and Nonprofits only -
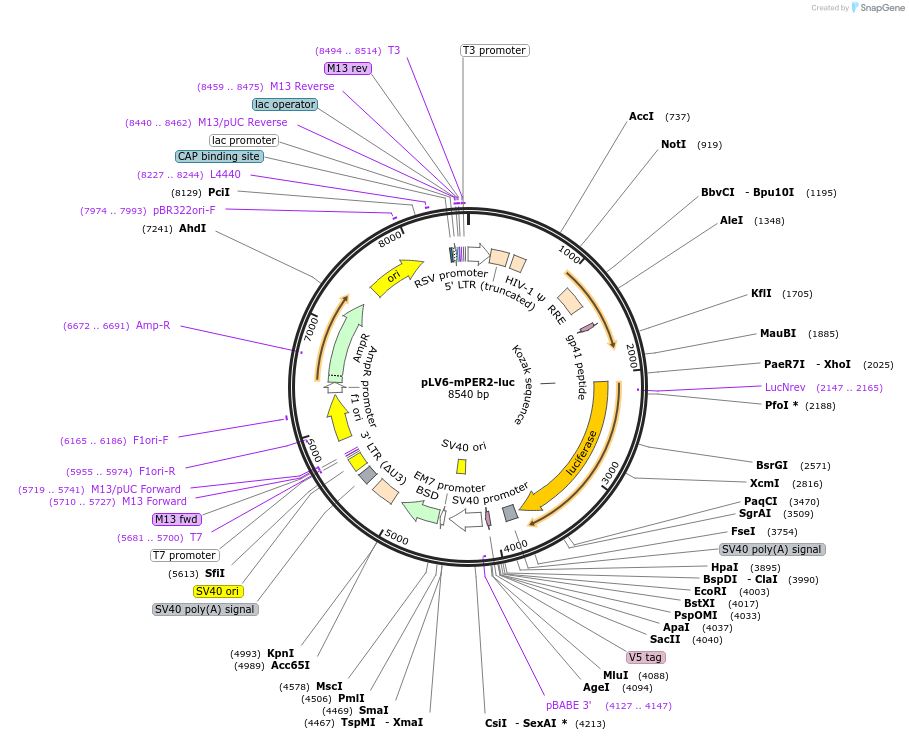
pLV6-mPER2-luc
Plasmid#212035PurposeExpression of murine Per2-luciferase in mammalian cellsDepositorInsertmPer2-luciferase (Per2 Mouse)
UseLentiviral and LuciferaseTagsluciferaseExpressionMammalianAvailable SinceDec. 20, 2024AvailabilityAcademic Institutions and Nonprofits only -
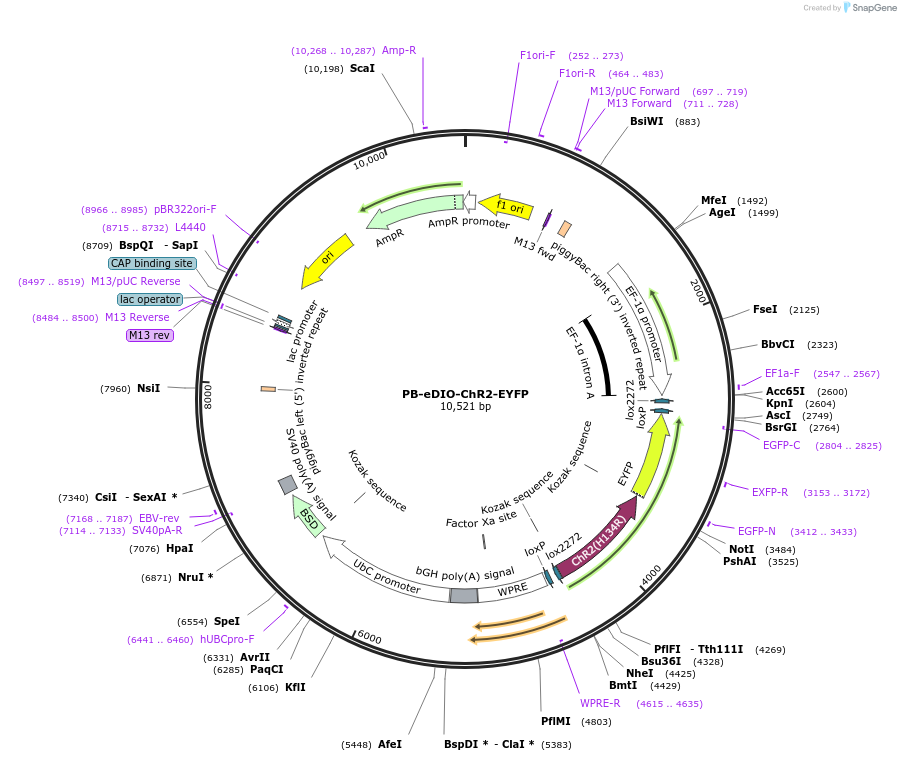
PB-eDIO-ChR2-EYFP
Plasmid#104387PurposePiggyBac plasmid encoding channelrhodopsin 2 fused to EYFP in an inverse orientation flanked by loxP and lox2722 sites under the control of the EF1a promotor.DepositorInsertChannelrhodopsin 2
UseCre/Lox; PiggybacTagsEYFPExpressionMammalianMutationE134RAvailable SinceFeb. 15, 2018AvailabilityAcademic Institutions and Nonprofits only -
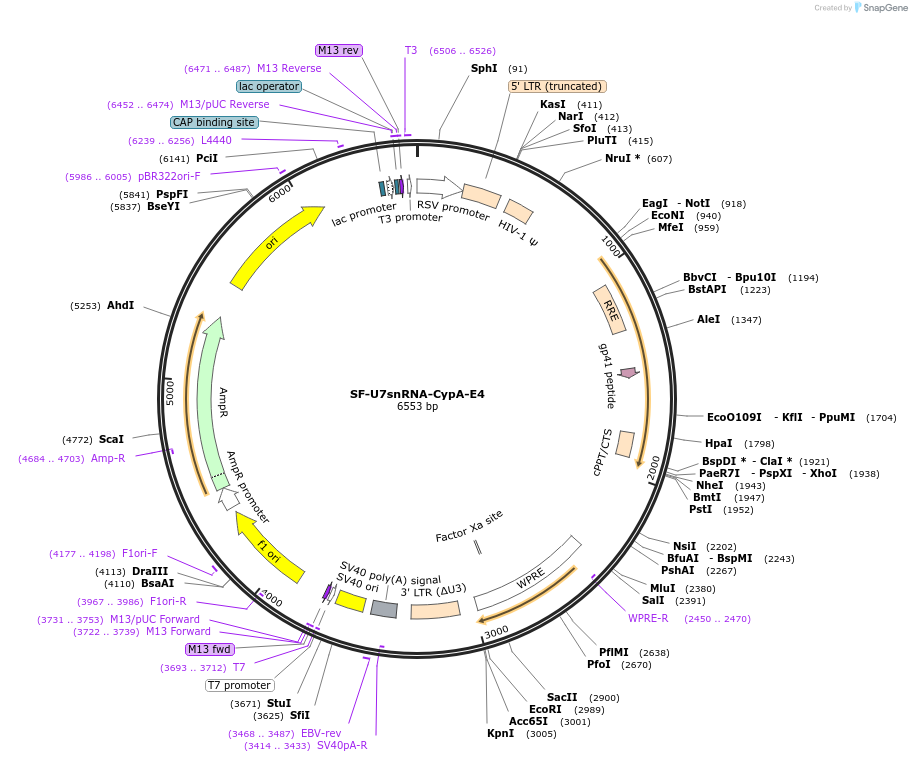
SF-U7snRNA-CypA-E4
Plasmid#190695PurposeControl vector expressing a U7-based antisense oligonucleotide targeting CypA exon 4DepositorInsertU7 snRNA antisense CypA Exon 4
UseLentiviralExpressionMammalianPromoterU7 promoterAvailable SinceApril 11, 2023AvailabilityAcademic Institutions and Nonprofits only -
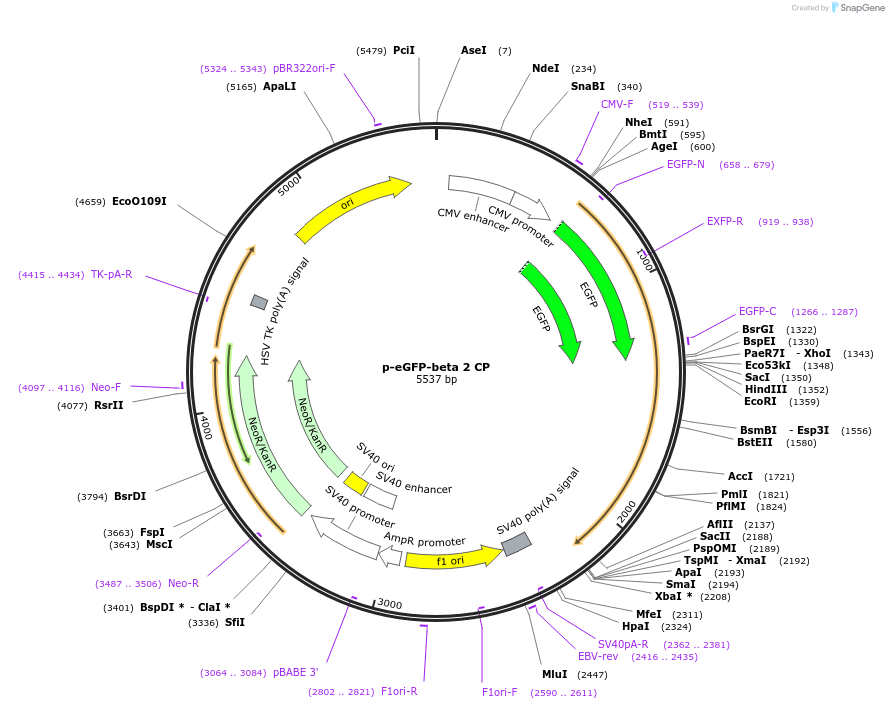
p-eGFP-beta 2 CP
Plasmid#13298DepositorAvailable SinceDec. 15, 2006AvailabilityAcademic Institutions and Nonprofits only -
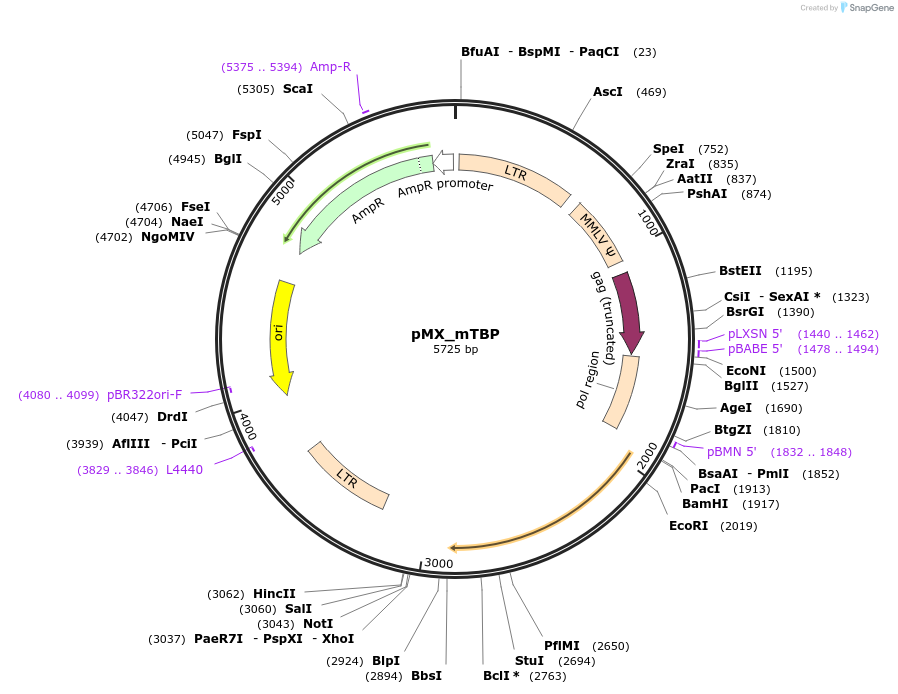
pMX_mTBP
Plasmid#44333DepositorAvailable SinceMay 14, 2013AvailabilityAcademic Institutions and Nonprofits only -
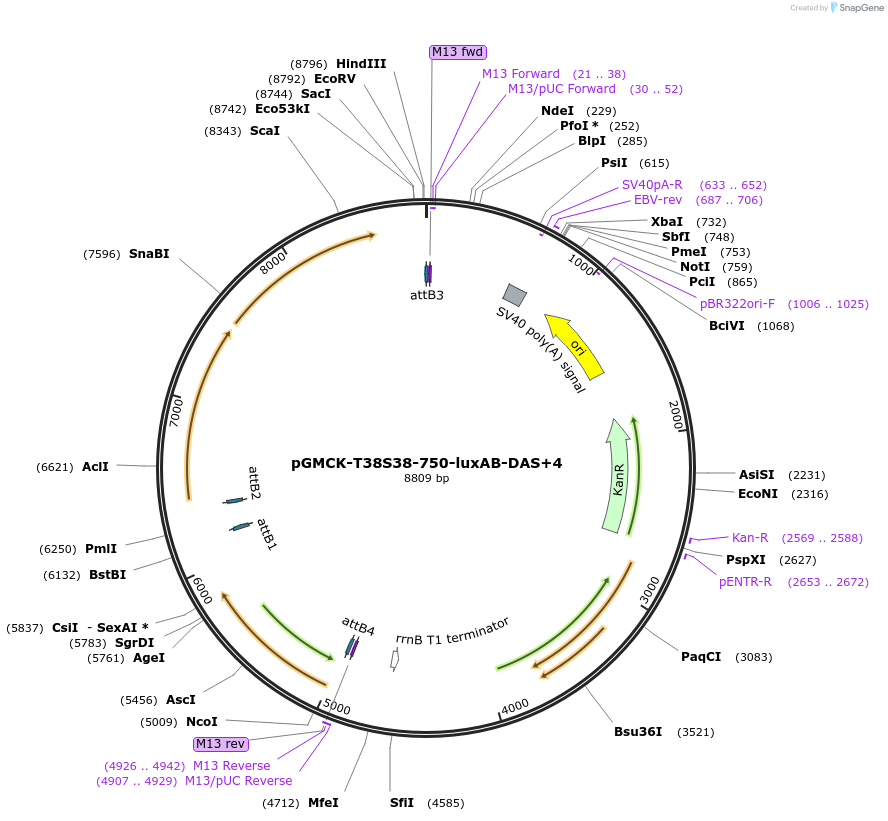
pGMCK-T38S38-750-luxAB-DAS+4
Plasmid#49518PurposeKanamycin resistant; expresses luxAB-DAS+4 under the control of a promoter containing PtetO-4C5G; constitutively expresses reverse TetR (tetR38); integrates into the mycobacteriophage att-L5 siteDepositorInsertT38S8-P750-luxAB-DAS+4
TagsDAS+4 (AANDENYSENYADAS)ExpressionBacterialPromoterP750Available SinceJan. 17, 2014AvailabilityAcademic Institutions and Nonprofits only -
COC053
Plasmid#209131PurposeEnzymatic biotinylation of proteins at a specific lysine residue in a recognition sequence. This has multiple applications, especially in the study of protein-protein interactions.DepositorInsertEnzyme BirA (birA E. coli)
ExpressionBacterialAvailable SinceMarch 6, 2024AvailabilityAcademic Institutions and Nonprofits only