We narrowed to 12,331 results for: NSI
-
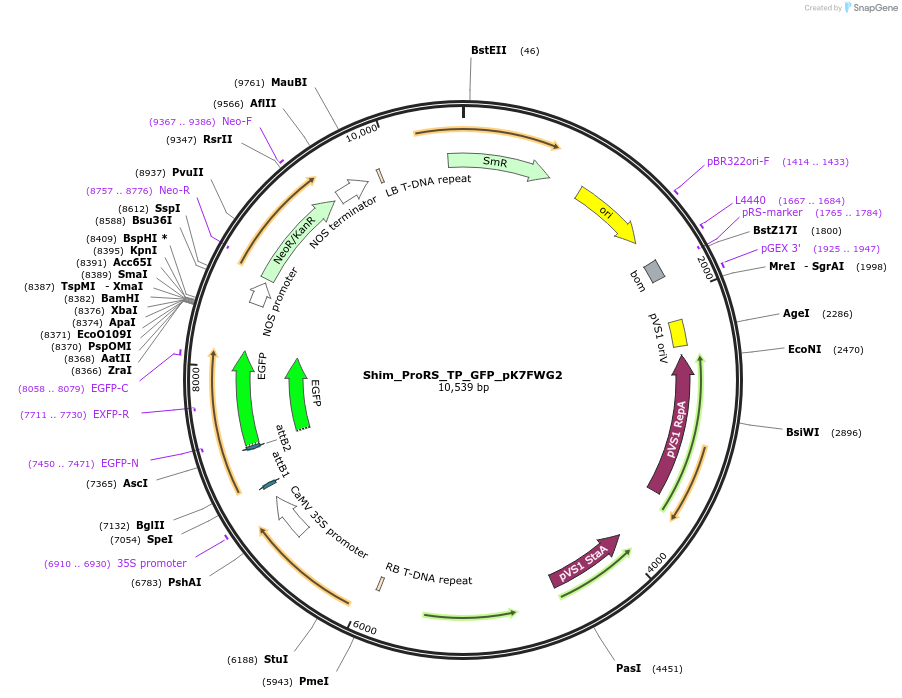
Plasmid#202651PurposeDetermine where transit peptide targets GFP in N. benthamianaDepositorInsertN-terminal transit peptide from putative cytosolic ProRS
TagsGreen Florescent ProtienExpressionPlantAvailable SinceAug. 10, 2023AvailabilityIndustry, Academic Institutions, and Nonprofits -
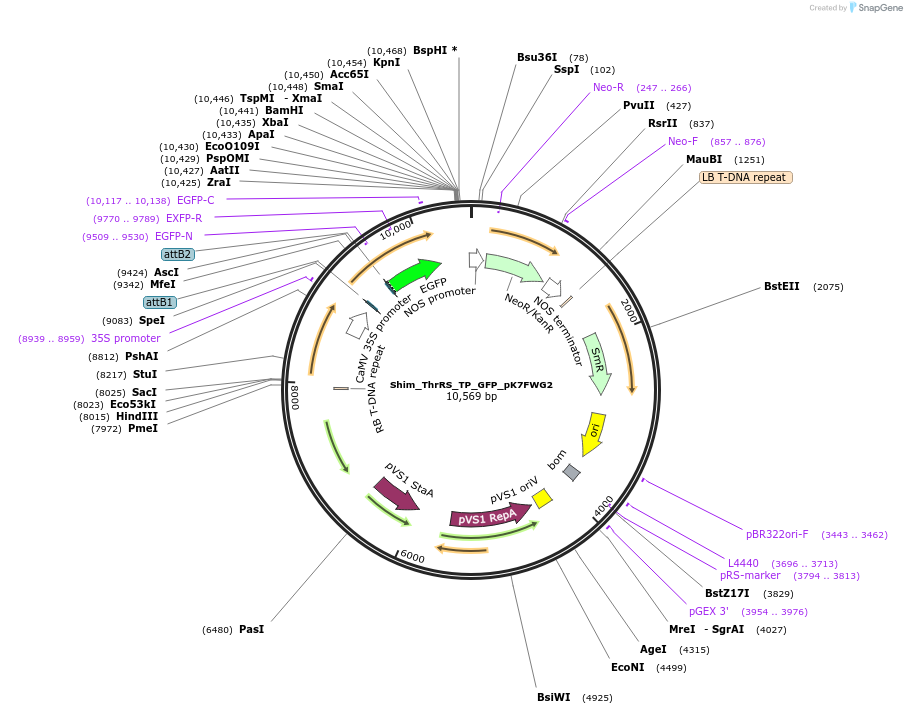
Shim_ThrRS_TP_GFP_pK7FWG2
Plasmid#202652PurposeDetermine where transit peptide targets GFP in N. benthamianaDepositorInsertN-terminal transit peptide from putative cytosolic ThrRS
TagsGreen Florescent ProtienExpressionPlantAvailable SinceAug. 10, 2023AvailabilityIndustry, Academic Institutions, and Nonprofits -
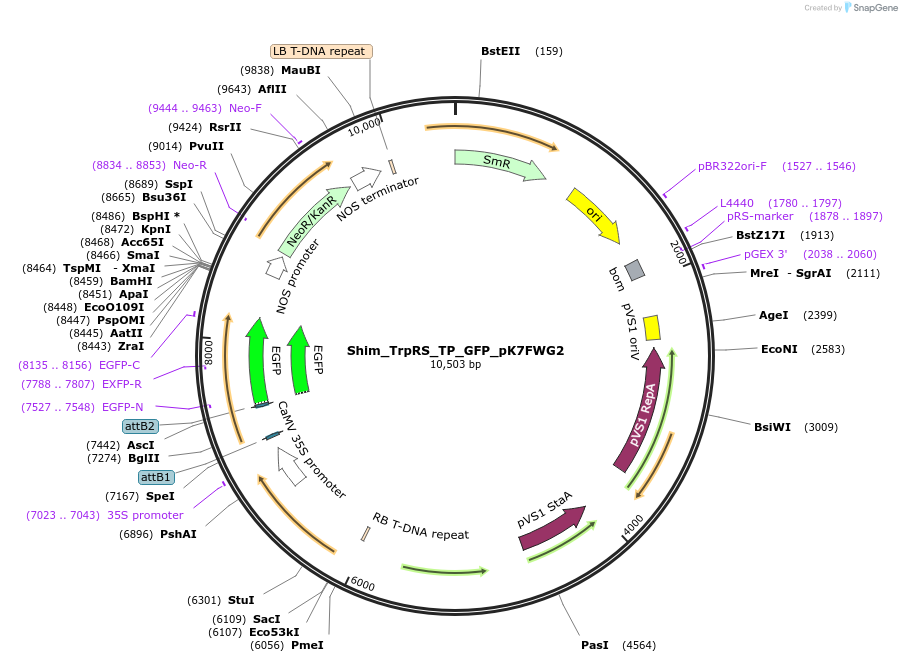
Shim_TrpRS_TP_GFP_pK7FWG2
Plasmid#202653PurposeDetermine where transit peptide targets GFP in N. benthamianaDepositorInsertN-terminal transit peptide from putative cytosolic TrpRS
TagsGreen Florescent ProtienExpressionPlantAvailable SinceAug. 10, 2023AvailabilityIndustry, Academic Institutions, and Nonprofits -
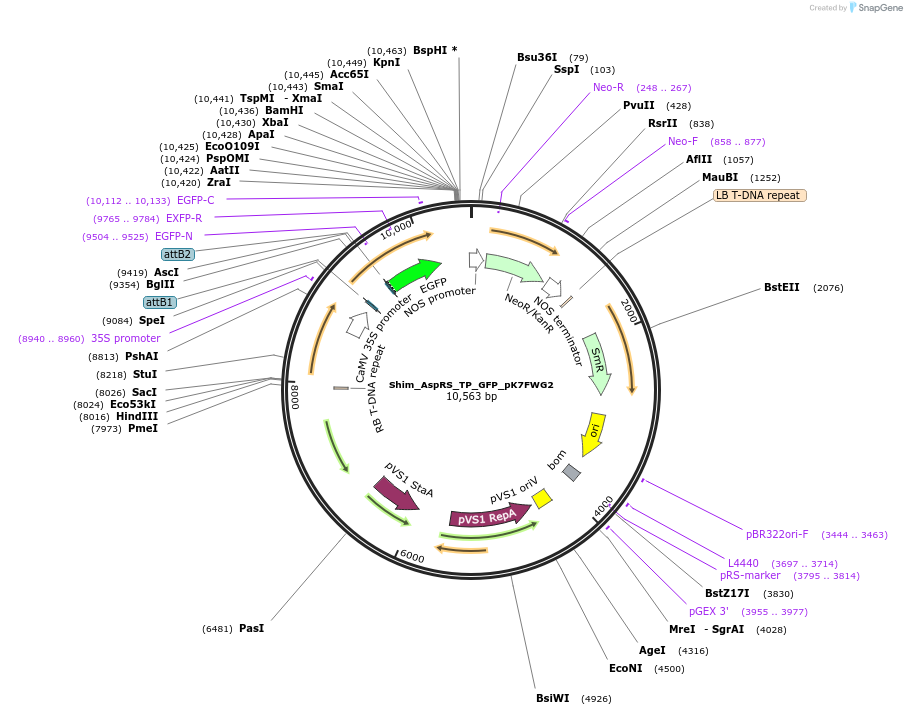
Shim_AspRS_TP_GFP_pK7FWG2
Plasmid#202647PurposeDetermine where transit peptide targets GFP in N. benthamianaDepositorInsertN-terminal transit peptide from putative cytosolic AspRS
TagsGreen Florescent ProtienExpressionPlantAvailable SinceAug. 10, 2023AvailabilityIndustry, Academic Institutions, and Nonprofits -
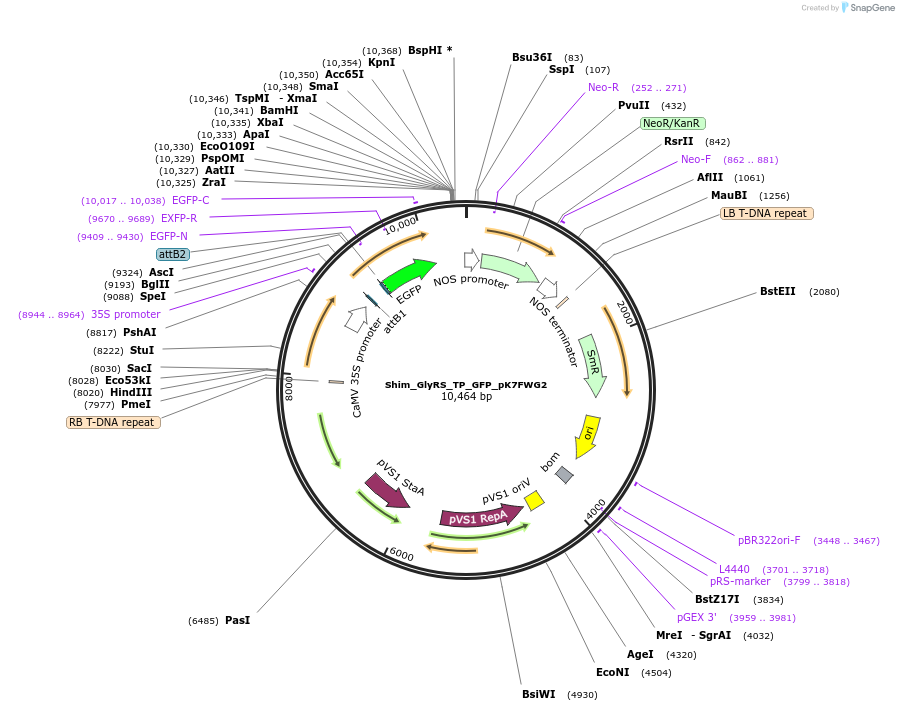
Shim_GlyRS_TP_GFP_pK7FWG2
Plasmid#202648PurposeDetermine where transit peptide targets GFP in N. benthamianaDepositorInsertN-terminal transit peptide from putative cytosolic GlyRS
TagsGreen Florescent ProtienExpressionPlantAvailable SinceAug. 10, 2023AvailabilityIndustry, Academic Institutions, and Nonprofits -
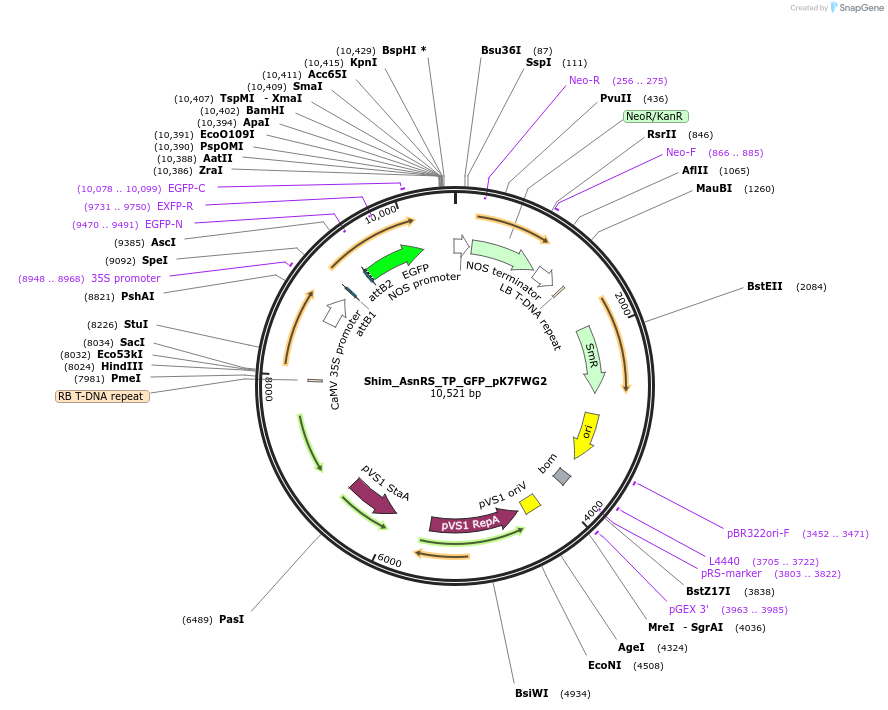
Shim_AsnRS_TP_GFP_pK7FWG2
Plasmid#202649PurposeDetermine where transit peptide targets GFP in N. benthamianaDepositorInsertN-terminal transit peptide from putative cytosolic AsnRS
TagsGreen Florescent ProtienExpressionPlantAvailable SinceAug. 10, 2023AvailabilityIndustry, Academic Institutions, and Nonprofits -
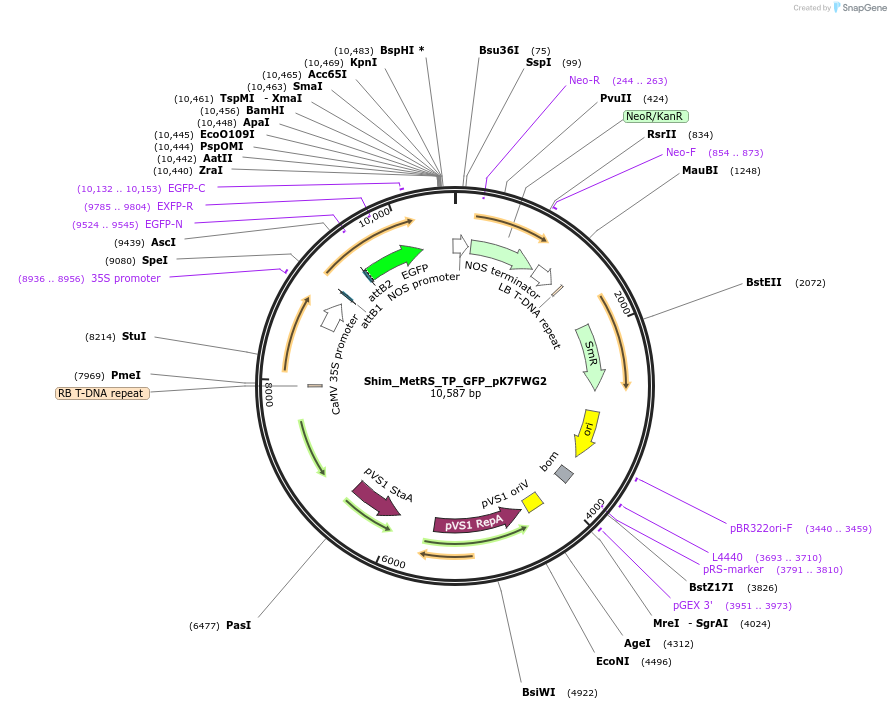
Shim_MetRS_TP_GFP_pK7FWG2
Plasmid#202650PurposeDetermine where transit peptide targets GFP in N. benthamianaDepositorInsertN-terminal transit peptide from putative cytosolic MetRS
TagsGreen Florescent ProtienExpressionPlantAvailable SinceAug. 10, 2023AvailabilityIndustry, Academic Institutions, and Nonprofits -
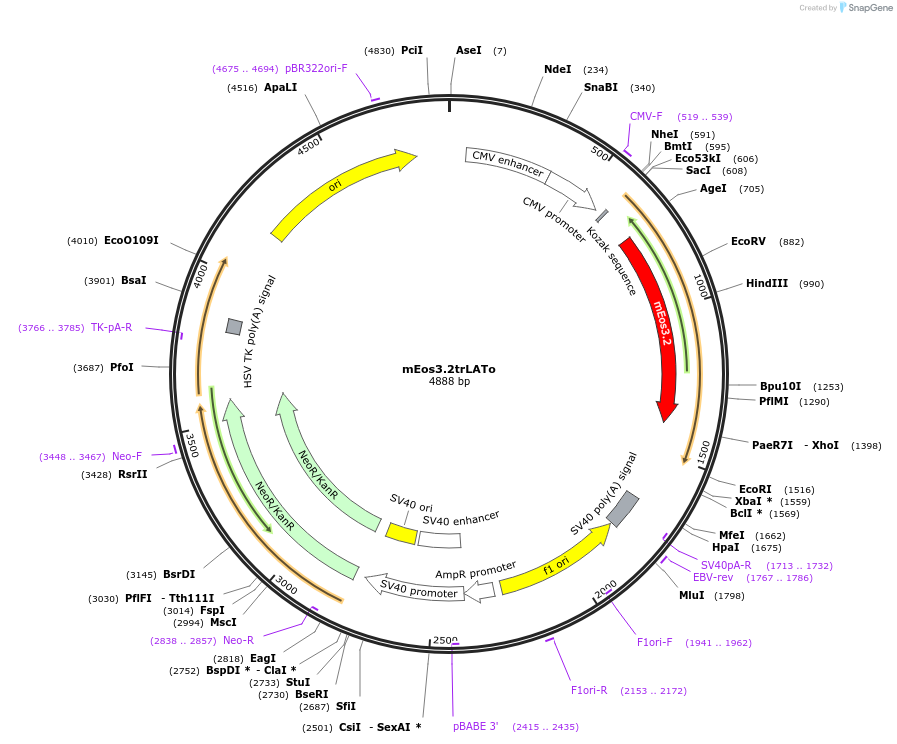
mEos3.2trLATo
Plasmid#203746PurposeEncodes the transmembrane domain from mouse LAT with mEos3.2 fused on the N terminus. Used as a fluorescent plasma membrane probe.DepositorAvailable SinceAug. 7, 2023AvailabilityAcademic Institutions and Nonprofits only -
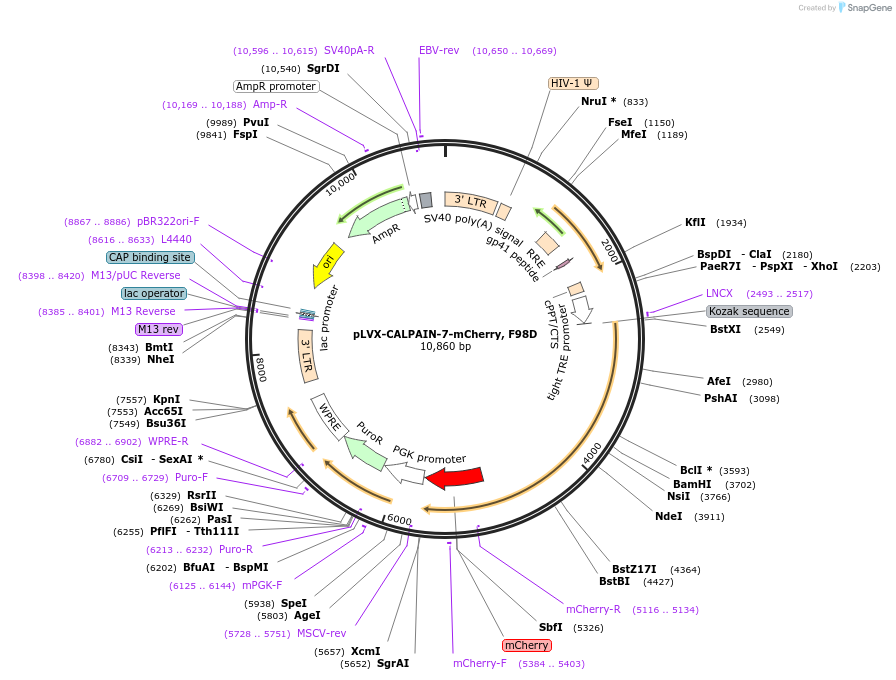
pLVX-CALPAIN-7-mCherry, F98D
Plasmid#180653PurposeLentiviral expression vector for CALPAIN-7. Used for generating cell lines. F98D mutation. Has C-terminal mCherry tag. Dox-inducible. Internal ID: WISP20-53.DepositorInsertCALPAIN-7
UseLentiviralTagsmCherryExpressionMammalianMutationsiRNA resistant to: GCACCCAUACCUUUACAUUAvailable SinceJuly 12, 2023AvailabilityAcademic Institutions and Nonprofits only -
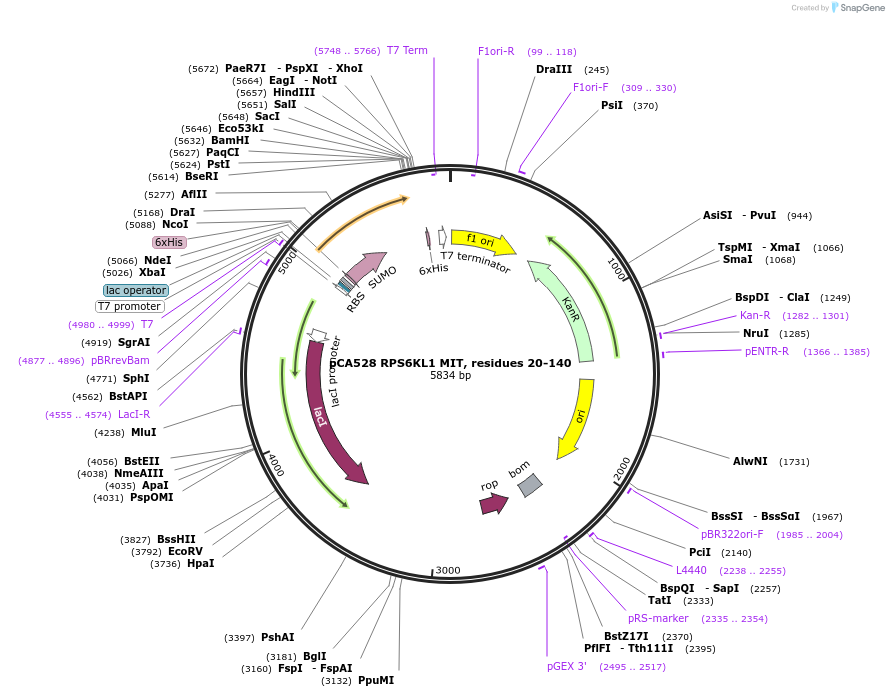
pCA528 RPS6KL1 MIT, residues 20-140
Plasmid#180623PurposeBacterial expression for RPS6KL1 MIT domain, residues 20-140. Has N-terminal HIS-SUMO tag. Internal ID: WISP20-32.DepositorInsertRPS6KL1 (RPS6KL1 Human)
TagsHIS-SUMOExpressionBacterialMutationResidues 20-140PromoterT7Available SinceJuly 12, 2023AvailabilityAcademic Institutions and Nonprofits only -
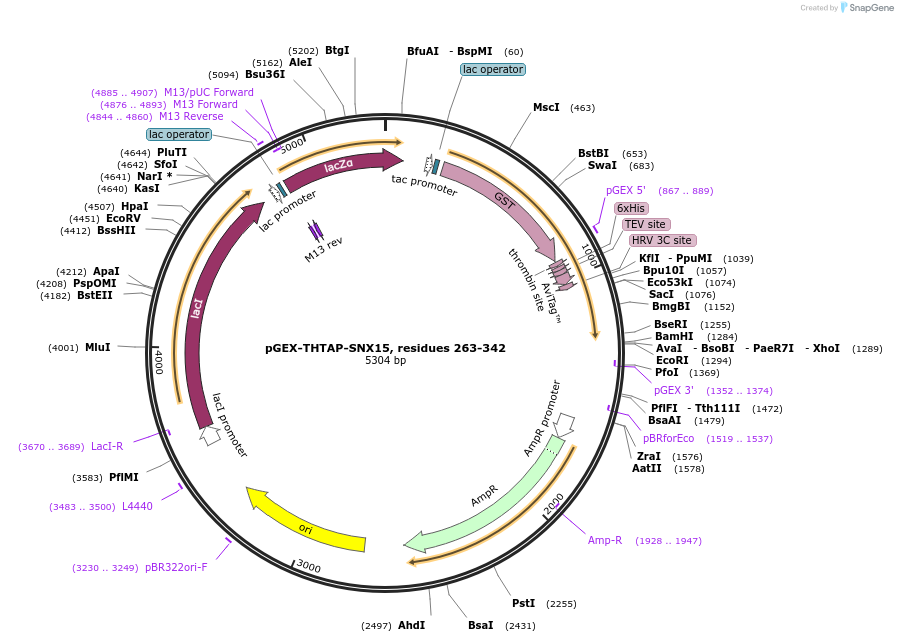
pGEX-THTAP-SNX15, residues 263-342
Plasmid#180624PurposeBacterial expression for SNX15 MIT residues 263-342. Has N-terminal HIS-GST tag. Internal ID:WISP21-130.DepositorAvailable SinceJuly 12, 2023AvailabilityAcademic Institutions and Nonprofits only -
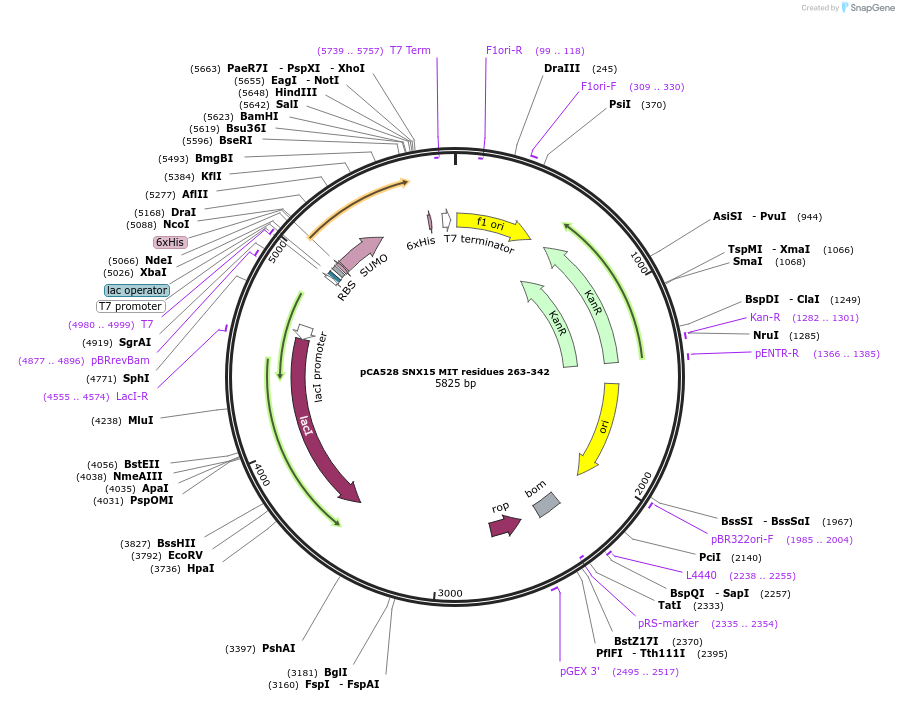
pCA528 SNX15 MIT residues 263-342
Plasmid#180625PurposeBacterial expression for SNX15 MIT residues 263-342. Has N-terminal HIS-SUMO tag. Internal ID:WISP21-14DepositorAvailable SinceJuly 12, 2023AvailabilityAcademic Institutions and Nonprofits only -
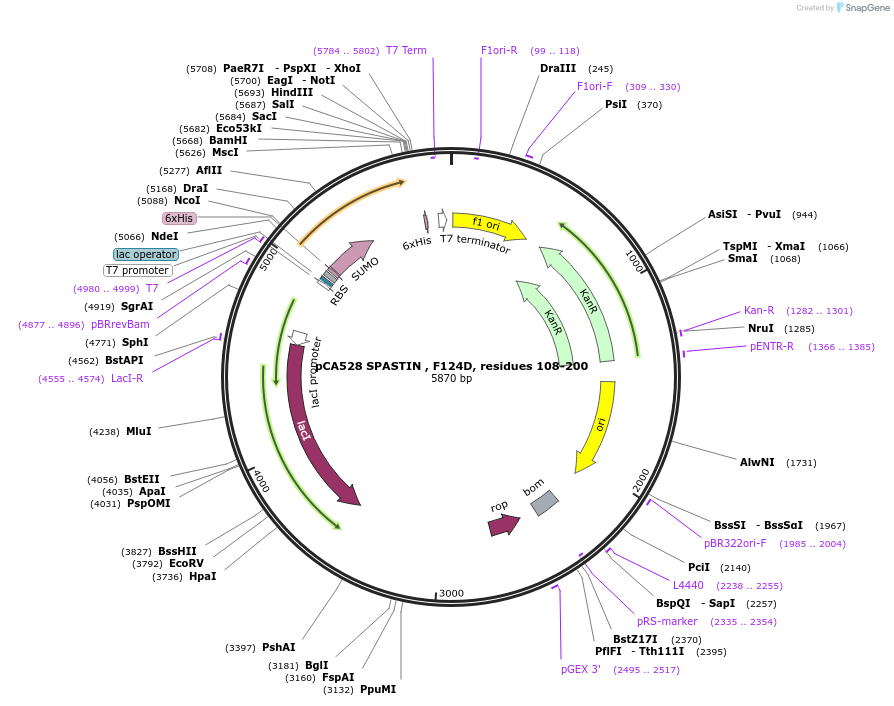
pCA528 SPASTIN , F124D, residues 108-200
Plasmid#180629PurposeBacterial expression for SPASTIN MIT, residues 108-200. Has F124D mutation. Has N-term HIS-SUMO tag. Used for FP experiments. Internal ID: WISP21-12.DepositorInsertSPASTIN
TagsHIS-SUMOExpressionBacterialMutationResidues 108-200, F124DPromoterT7Available SinceJuly 12, 2023AvailabilityAcademic Institutions and Nonprofits only -
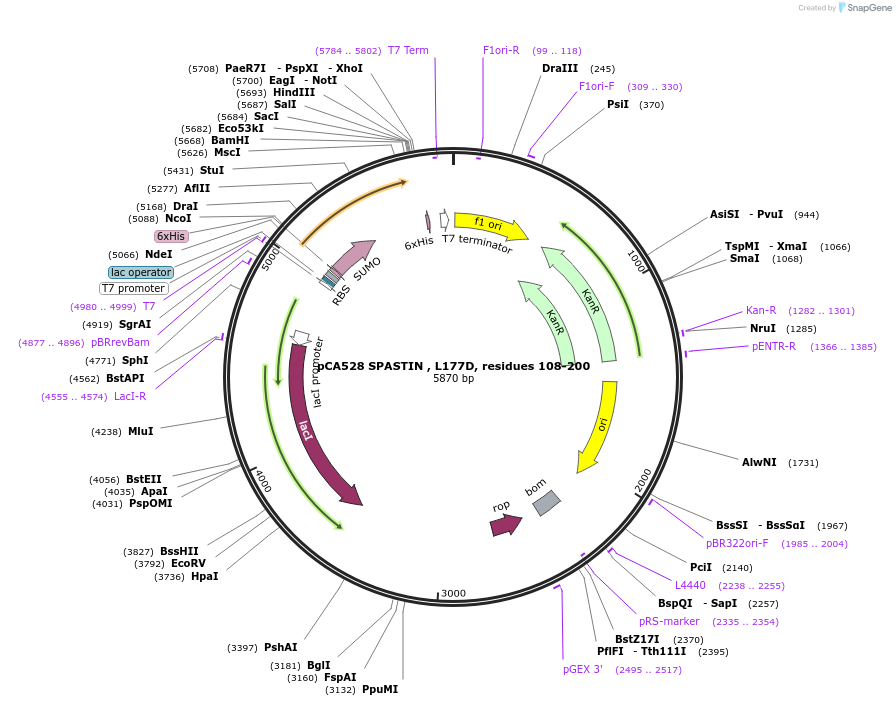
pCA528 SPASTIN , L177D, residues 108-200
Plasmid#180634PurposeBacterial expression for SPASTIN MIT, residues 108-200. Has L177D mutation. Has N-term HIS-SUMO tag. Used for FP experiments. Internal ID: WISP21-13.DepositorInsertSPASTIN
TagsHIS-SUMOExpressionBacterialMutationResidues 108-200, L177DPromoterT7Available SinceJuly 12, 2023AvailabilityAcademic Institutions and Nonprofits only -
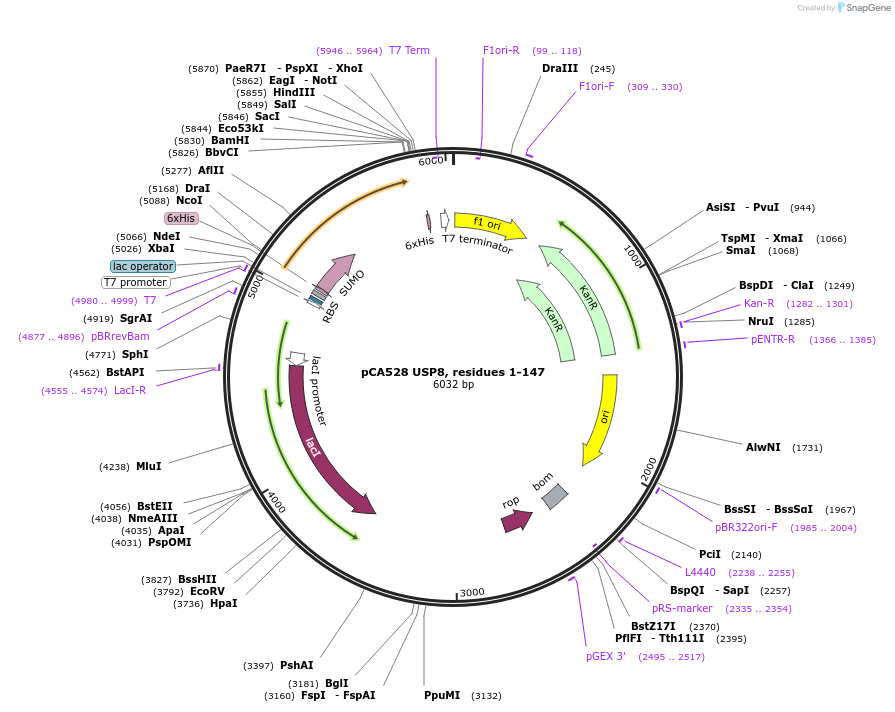
pCA528 USP8, residues 1-147
Plasmid#180636PurposeBacterial expression for USP8 MIT domain, residues 1-147. Has N-terminal HIS-SUMO tag. Internal ID: WISP20-86.DepositorAvailable SinceJuly 12, 2023AvailabilityAcademic Institutions and Nonprofits only -
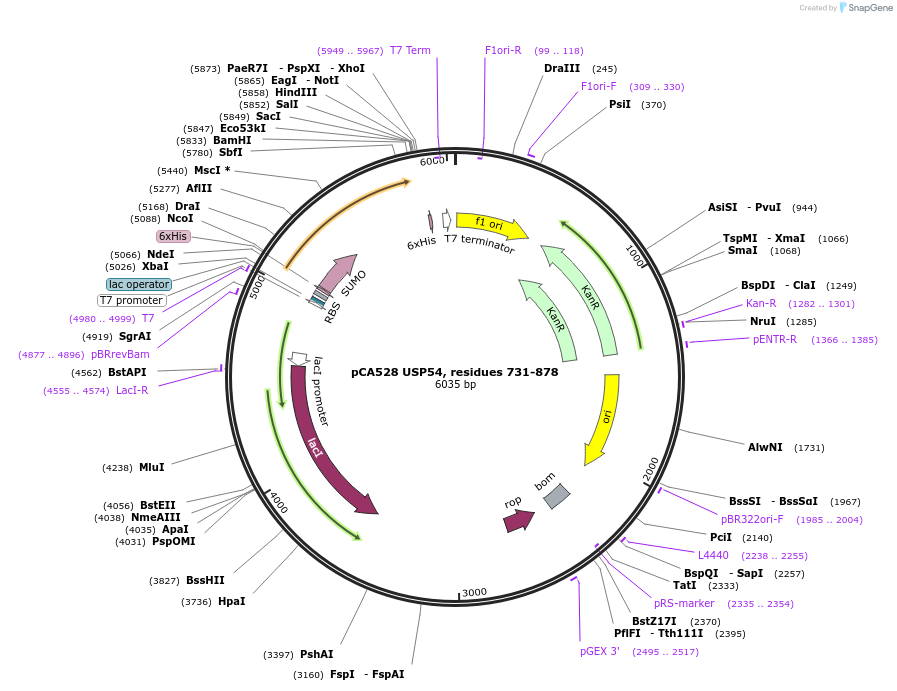
pCA528 USP54, residues 731-878
Plasmid#180637PurposeBacterial expression for USP54 MIT domain, residues 731-878. Has N-terminal HIS-SUMO tag. Internal ID:WISP20-85DepositorAvailable SinceJuly 12, 2023AvailabilityAcademic Institutions and Nonprofits only -
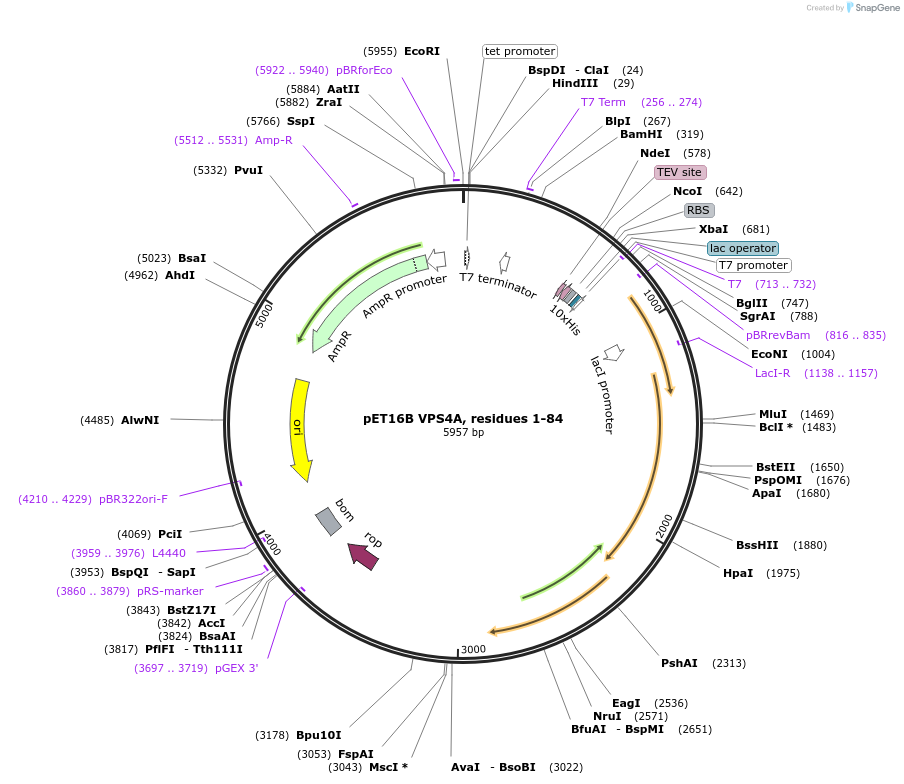
pET16B VPS4A, residues 1-84
Plasmid#180638PurposeBacterial expression for VPS4A MIT domain residues 1-84. Has N-terminal HIS-TAG. Internal ID: WISP05-43DepositorAvailable SinceJuly 12, 2023AvailabilityAcademic Institutions and Nonprofits only -
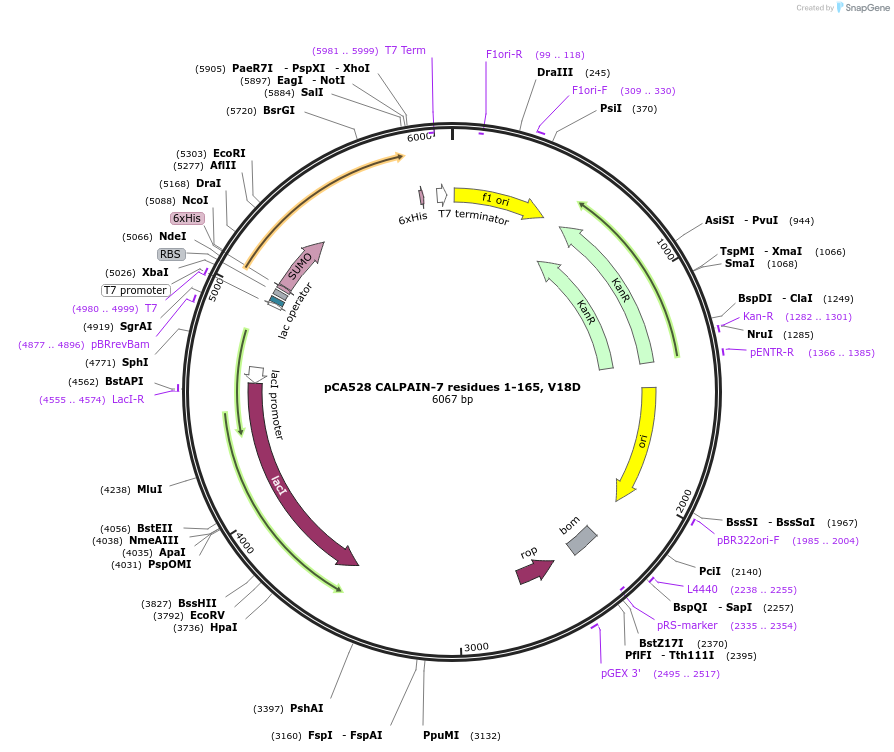
pCA528 CALPAIN-7 residues 1-165, V18D
Plasmid#180612PurposeBacterial expression for CALPAIN-7 MIT domains, residues 1-165. Has V18D mutation. Has N-terminal HIS-SUMO tag. Internal ID: WISP21-26.DepositorInsertCALPAIN-7
TagsHIS-SUMOExpressionBacterialMutationResidues 1-165, V18D mutationPromoterT7Available SinceJuly 12, 2023AvailabilityAcademic Institutions and Nonprofits only -
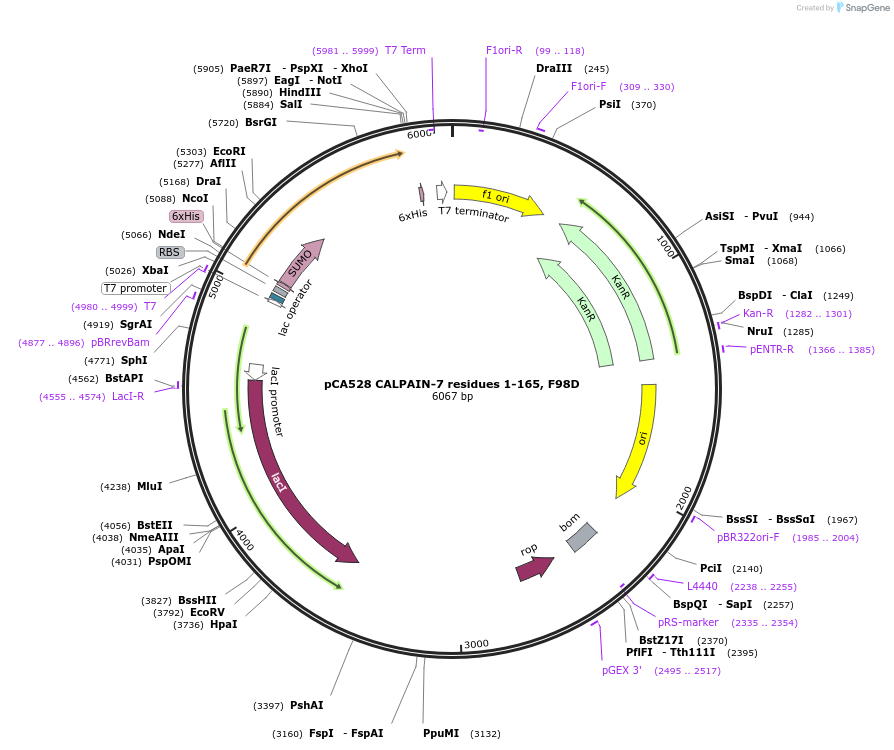
pCA528 CALPAIN-7 residues 1-165, F98D
Plasmid#180613PurposeBacterial expression for CALPAIN-7 MIT domains, residues 1-165. Has F98D mutation. Has N-terminal HIS-SUMO tag. Internal ID: WISP21-27.DepositorInsertCALPAIN-7
TagsHIS-SUMOExpressionBacterialMutationResidues 1-165, F98D mutationPromoterT7Available SinceJuly 12, 2023AvailabilityAcademic Institutions and Nonprofits only -
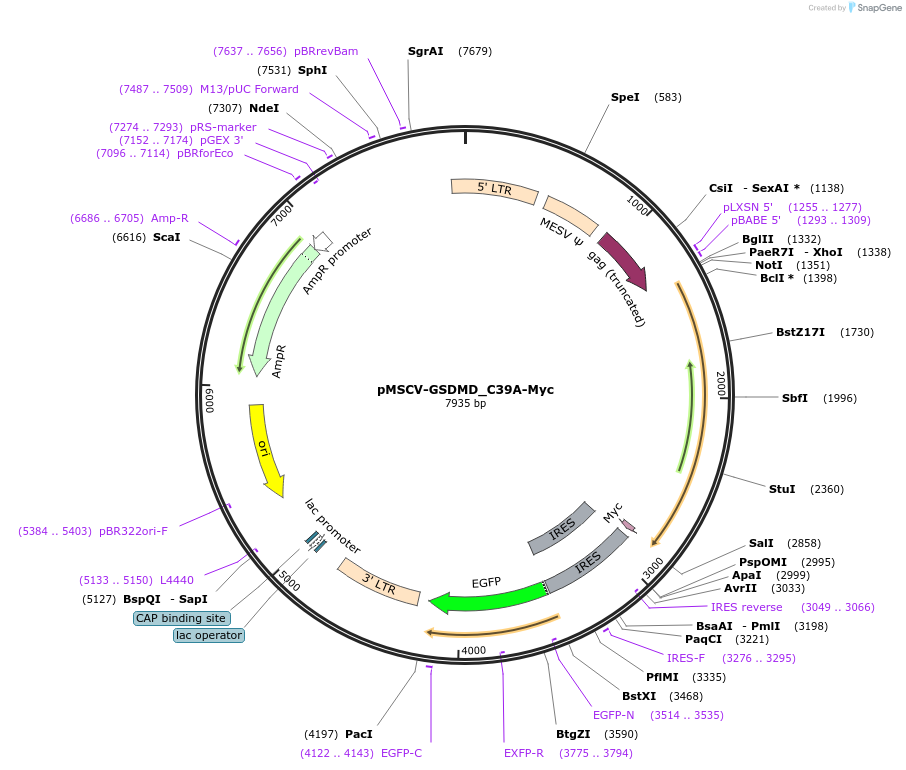
pMSCV-GSDMD_C39A-Myc
Plasmid#201167PurposeExpression of GSDMD gene in mammalian cells by retroviral transductionDepositorAvailable SinceJuly 5, 2023AvailabilityAcademic Institutions and Nonprofits only