We narrowed to 9,870 results for: SUB;
-
Plasmid#187139PurposeEntry cloneDepositorAvailable SinceAug. 15, 2022AvailabilityAcademic Institutions and Nonprofits only
-
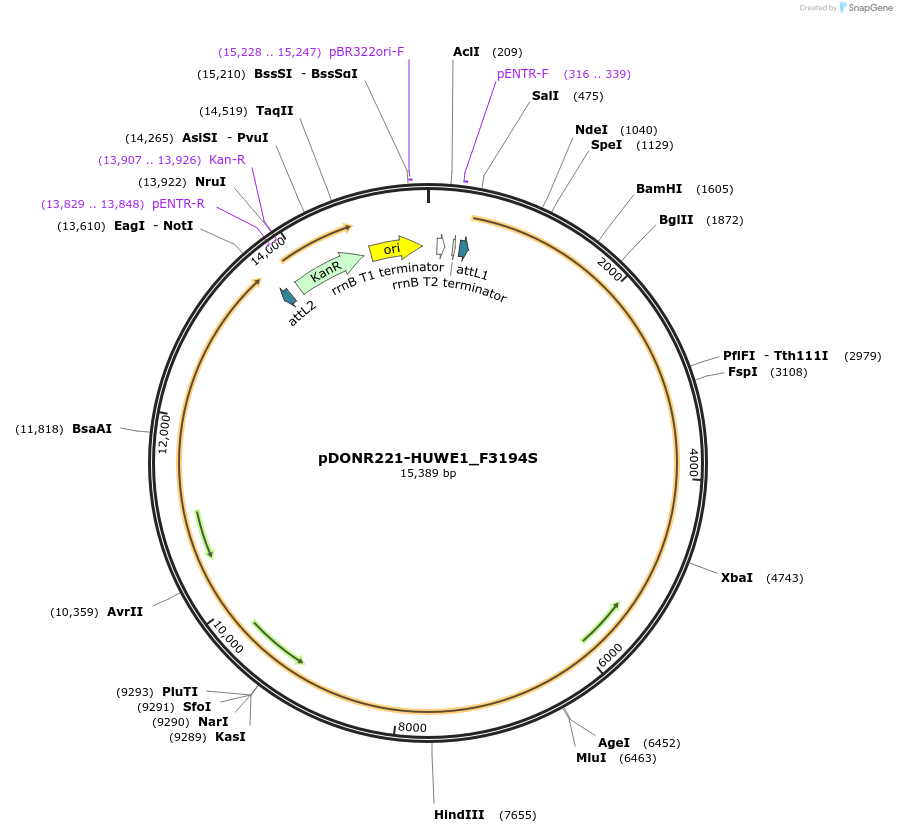
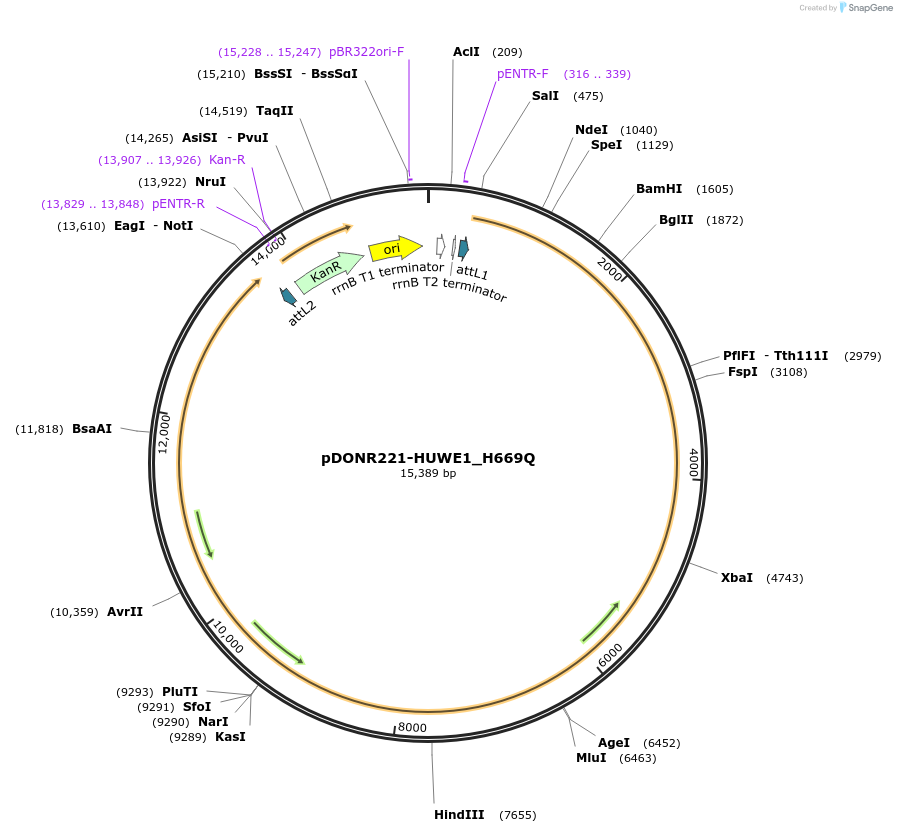
pDONR221-HUWE1_H669Q
Plasmid#187137PurposeEntry cloneDepositorAvailable SinceAug. 15, 2022AvailabilityAcademic Institutions and Nonprofits only -
pDONR221-HUWE1_H3962D
Plasmid#187130PurposeEntry cloneDepositorAvailable SinceAug. 12, 2022AvailabilityAcademic Institutions and Nonprofits only -
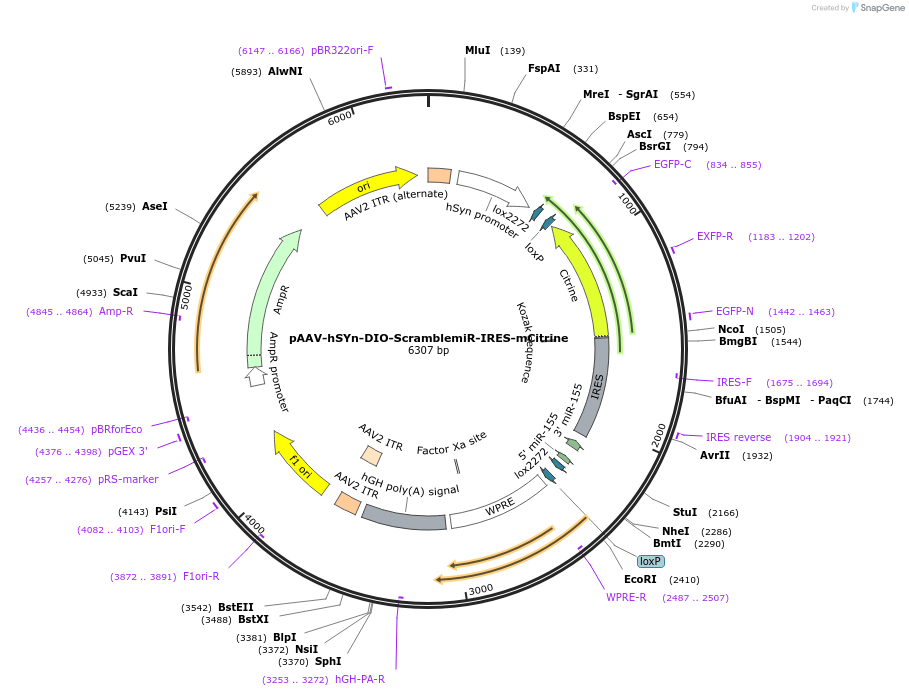
pAAV-hSYn-DIO-ScramblemiR-IRES-mCitrine
Plasmid#186418PurposeExpression of scrambled shRNA used as a control for knock down experimentsDepositorInsertScramblemiR
UseAAVTagsmCitrinePromoterhuman Synapsin1Available SinceAug. 1, 2022AvailabilityAcademic Institutions and Nonprofits only -
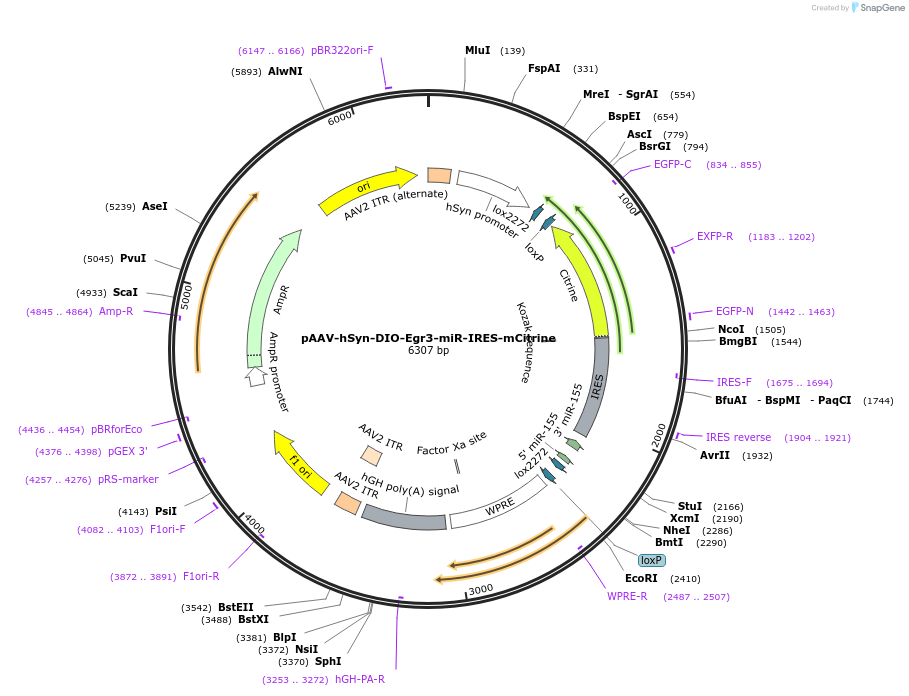
pAAV-hSyn-DIO-Egr3-miR-IRES-mCitrine
Plasmid#186419PurposeExpression of shRNA for the knock down of mouse Egr3DepositorInsertmiREgr3-5D
UseAAVTagsmCitrinePromoterhuman Synapsin1Available SinceAug. 1, 2022AvailabilityAcademic Institutions and Nonprofits only -
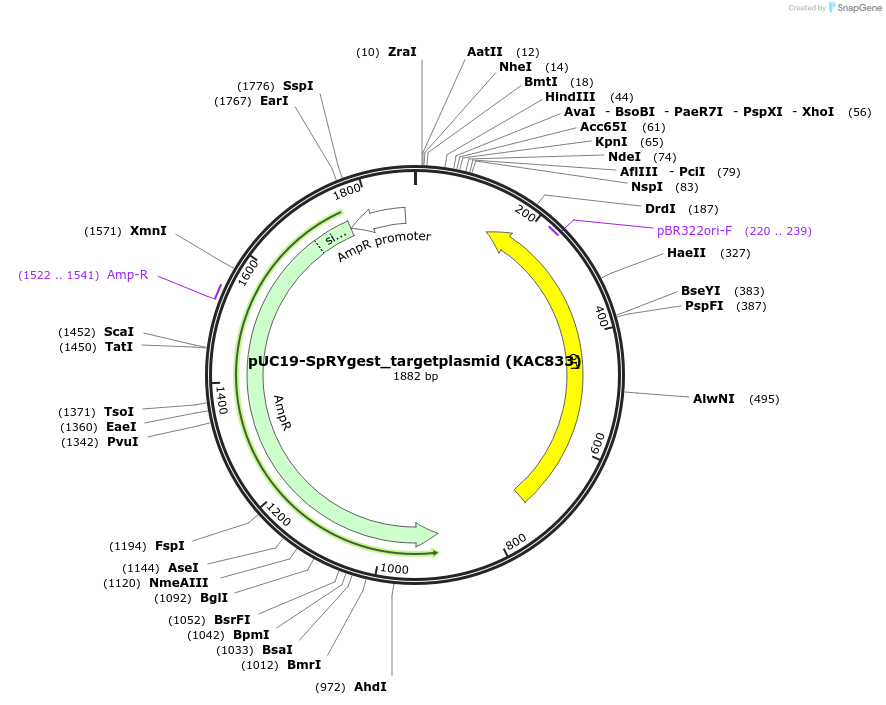
pUC19-SpRYgest_targetplasmid (KAC833)
Plasmid#181748PurposepUC19 derivative plasmid used as a substrate for SpRYgest, restriction enzyme, and TtAgo digests. Encodes an SpCas9 EMX1-site1-NGG-PAM target site between NheI and HindIII restriction sites.DepositorTypeEmpty backboneUseSubstrate plasmidAvailable SinceFeb. 16, 2022AvailabilityAcademic Institutions and Nonprofits only -
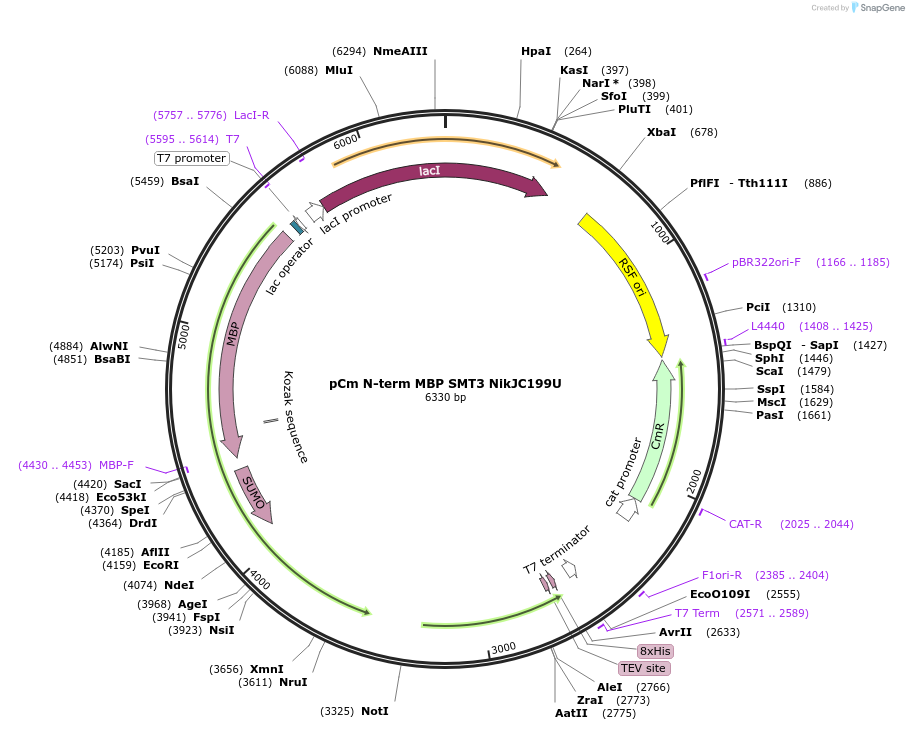
pCm N-term MBP SMT3 NikJC199U
Plasmid#174363PurposeExpression plasmid with a ULP1 removable N-term MBP SUMO tag, a TEV removable C-term 8xHis tag, and Chloramphenicol resistance. NikJ C199U cloned in the cloning site.DepositorInsertnikJ, nikkomycin biosynthesis protein P1 [Streptomyces tendae]
Tags8xHis, MBP, SMT3, and TEVExpressionBacterialMutationC199UPromoterT7Available SinceDec. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
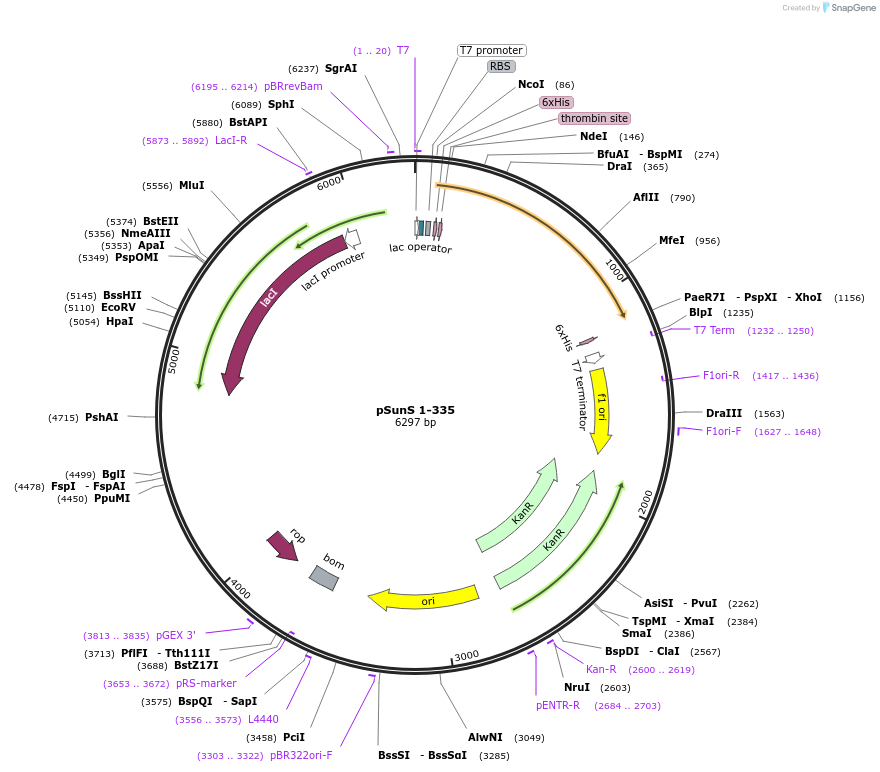
pSunS 1-335
Plasmid#171673PurposeSunS 1-335 expression in E. coliDepositorInsertGlycosyltransferase SunS (sunS Bacillus subtilis (strain 168))
TagsHis-tagExpressionBacterialPromoterT7Available SinceAug. 12, 2021AvailabilityAcademic Institutions and Nonprofits only -
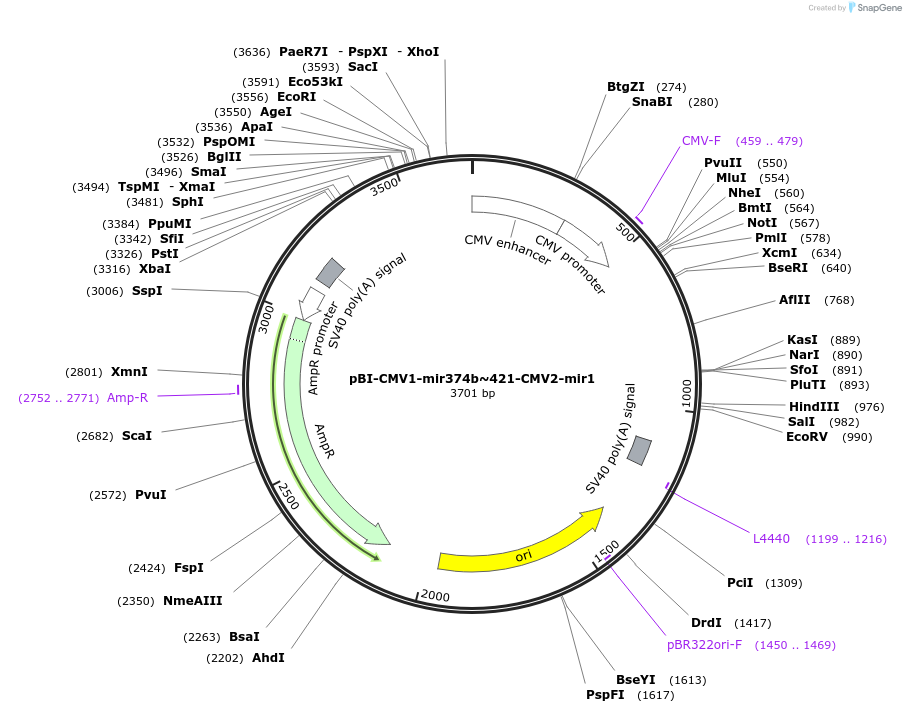
pBI-CMV1-mir374b~421-CMV2-mir1
Plasmid#164674PurposeExpress miR-374b~421 and miR-1-1 in mammalian cellsDepositorAvailable SinceMarch 14, 2021AvailabilityAcademic Institutions and Nonprofits only -
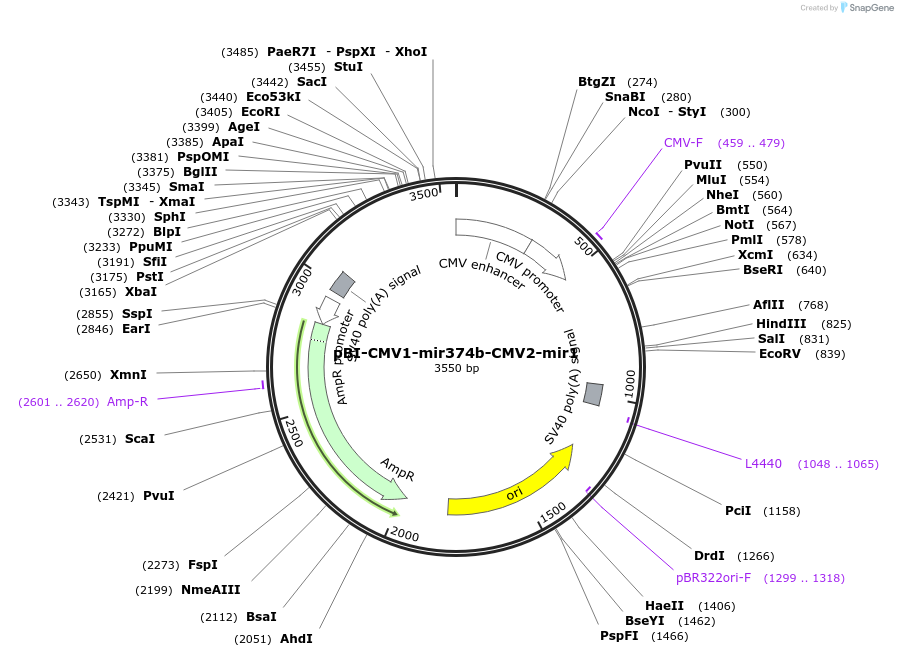
pBI-CMV1-mir374b-CMV2-mir1
Plasmid#164672PurposeExpress miR-374b and miR-1-1 in mammalian cellsDepositorAvailable SinceMarch 14, 2021AvailabilityAcademic Institutions and Nonprofits only -
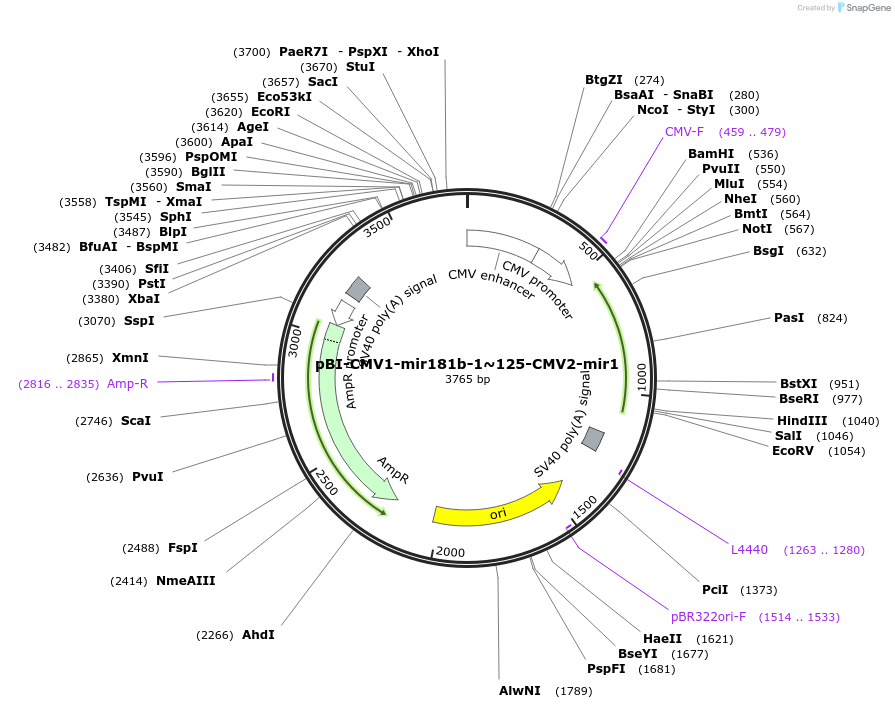
pBI-CMV1-mir181b-1~125-CMV2-mir1
Plasmid#164671PurposeExpress miR-181b-1~125a and miR-1-1 in mammalian cellsDepositorAvailable SinceMarch 14, 2021AvailabilityAcademic Institutions and Nonprofits only -
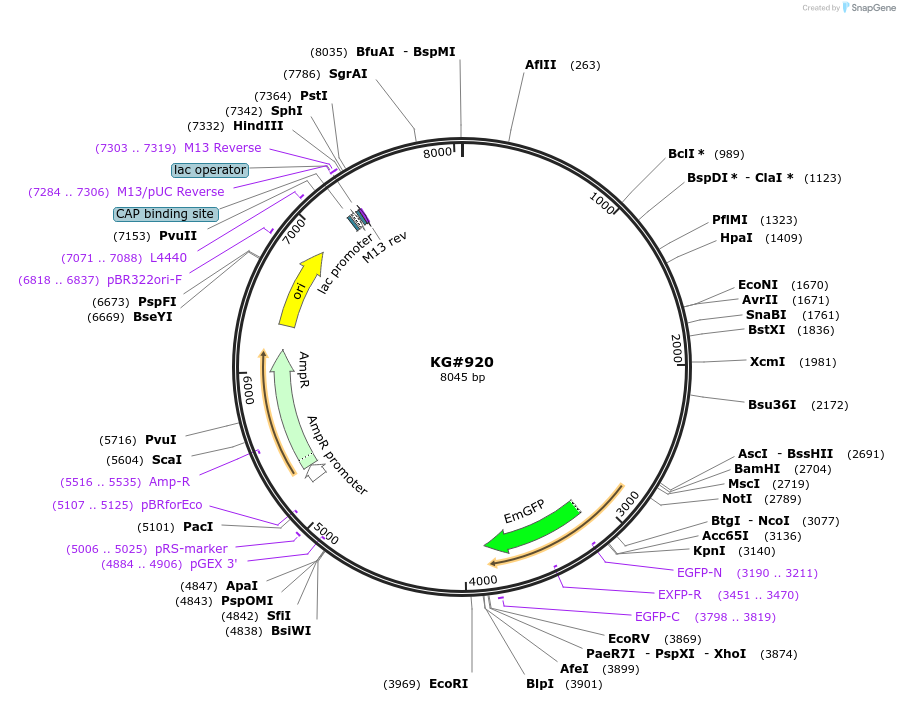
KG#920
Plasmid#110937PurposeExpresses INS-22-Emerald in the cholinergic motor neuron DA9 for imaging Dense Core Vesicles in a single neuron in a living animalDepositorInsertsmig-13 promoter
INS-22-Emerald
unc-54 3' control region with 1 artificial intron just upstream and 1 artificial intron in control region
ExpressionBacterialAvailable SinceMarch 11, 2021AvailabilityAcademic Institutions and Nonprofits only -
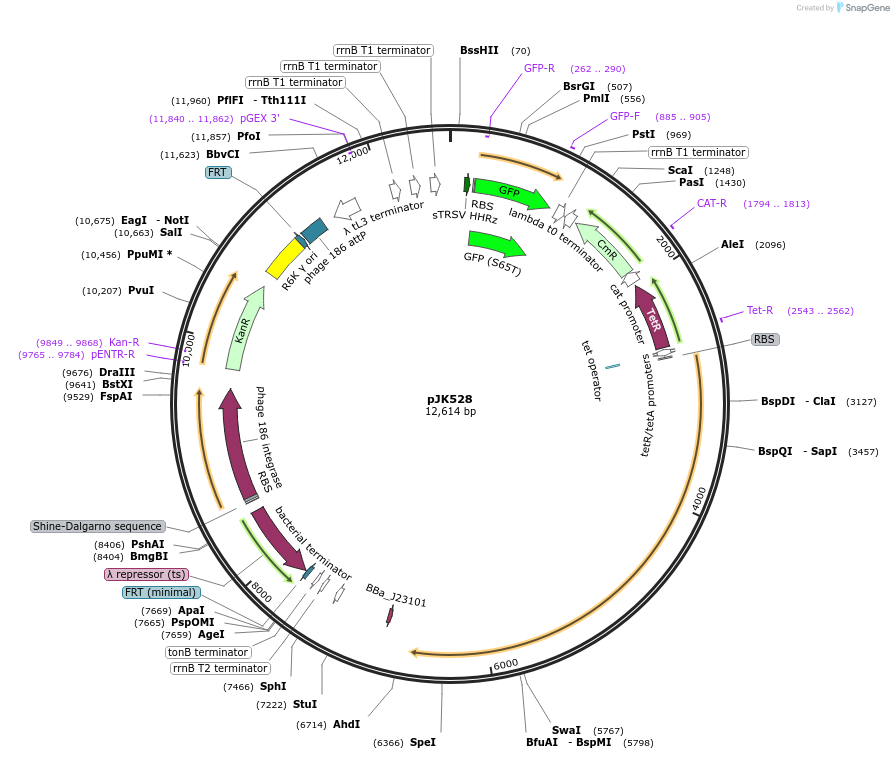
pJK528
Plasmid#155388PurposedCas12a oscillator, reverse crRNAs + PA4-mVenus. (Integration of 511; pOSIP)DepositorInsertsdCas12a (F. novicida)
PA4-mVenus
dCas12a oscillator crRNAs
UseCRISPRExpressionBacterialMutationD917A (nuclease-deactivating)Available SinceFeb. 12, 2021AvailabilityAcademic Institutions and Nonprofits only -
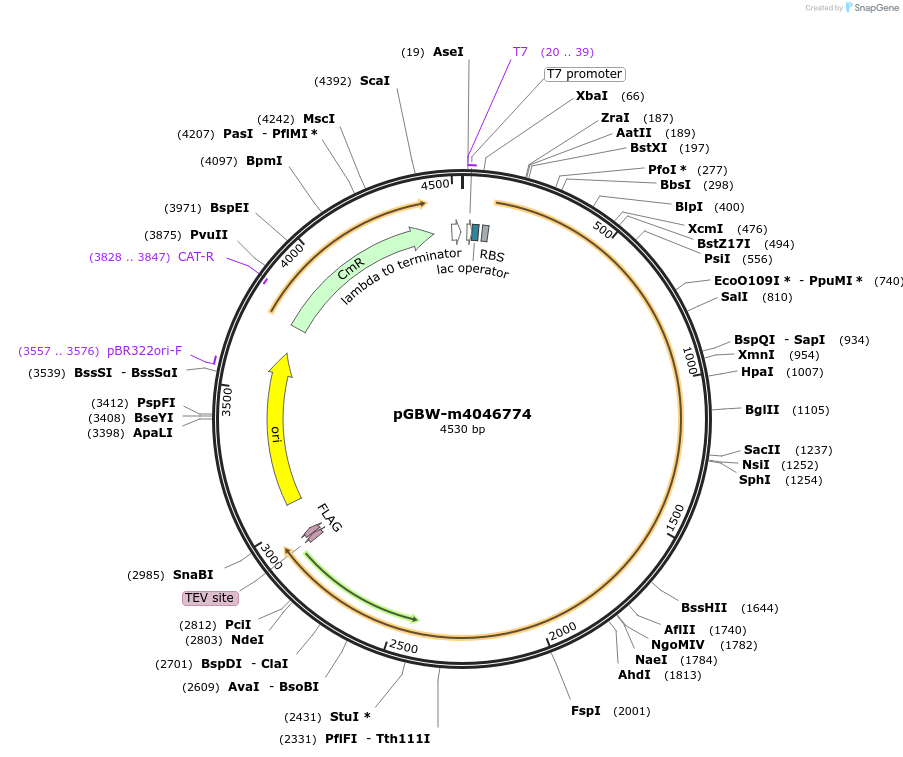
pGBW-m4046774
Plasmid#145574PurposeBacterial Expression plasmid for SARS-CoV-2 RNA-dependent RNA polymeraseDepositorInsertSARS-CoV-2 RNA-dependent RNA polymerase (ORF1ab Escherichia coli str. K-12 substr. MG1655; Severe acute respiratory syndrome coronavirus 2)
TagsCleavable TEV;FLAG_2ExpressionBacterialMutationEscherichia coli recode 1PromoterPromoter | pT7 ; Operator | lacO ; RBS | T7_rbs ;…Available SinceApril 29, 2020AvailabilityIndustry, Academic Institutions, and Nonprofits -
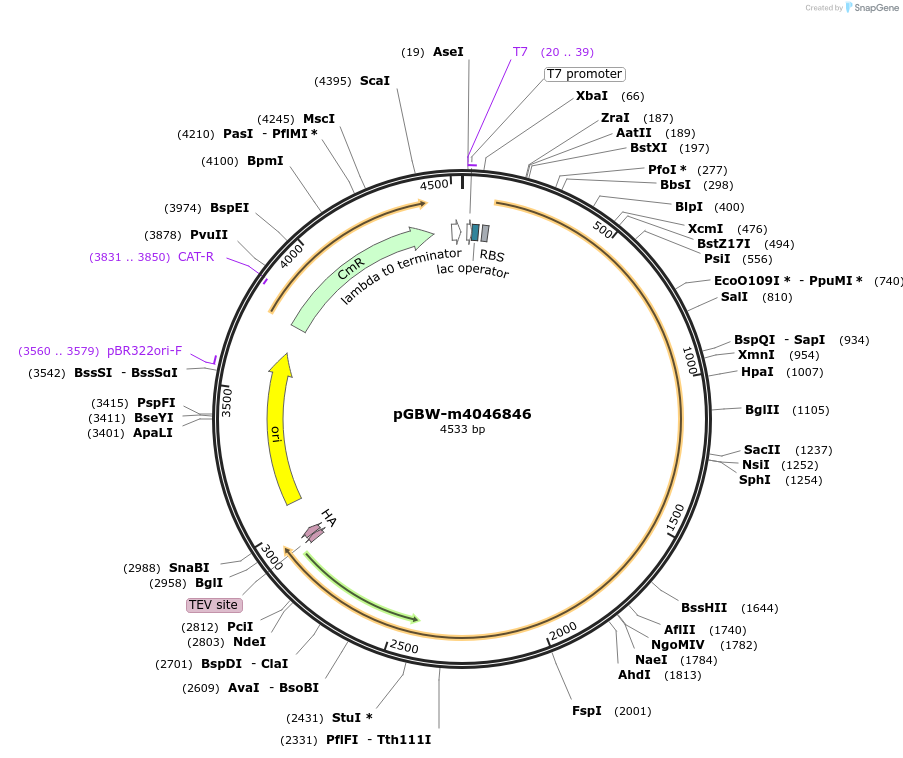
pGBW-m4046846
Plasmid#145575PurposeBacterial Expression plasmid for SARS-CoV-2 RNA-dependent RNA polymeraseDepositorInsertSARS-CoV-2 RNA-dependent RNA polymerase (ORF1ab Escherichia coli str. K-12 substr. MG1655; Severe acute respiratory syndrome coronavirus 2)
TagsCleavable TEV;HAExpressionBacterialMutationEscherichia coli recode 1PromoterPromoter | pT7 ; Operator | lacO ; RBS | T7_rbs ;…Available SinceApril 29, 2020AvailabilityIndustry, Academic Institutions, and Nonprofits -
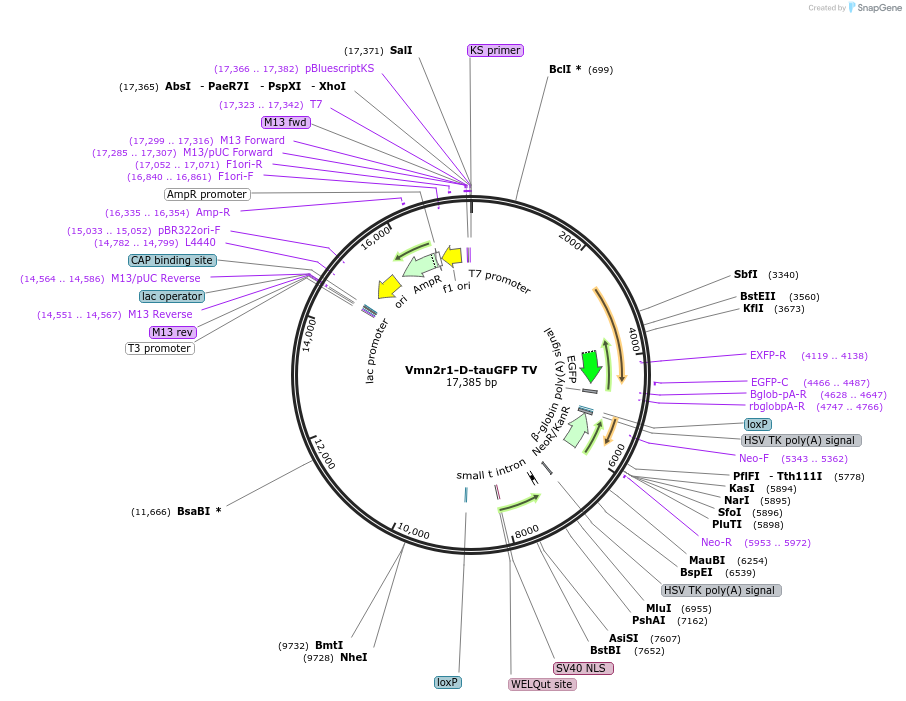
Vmn2r1-D-tauGFP TV
Plasmid#108080PurposeTargeting vector for Vmn2r1 gene knockout mice (∆C1-GFP strain)DepositorInsertVmn2r1-D-tauGFP-pA-ACNF (Vmn2r1 Mouse)
UseMouse TargetingAvailable SinceMay 17, 2019AvailabilityAcademic Institutions and Nonprofits only -
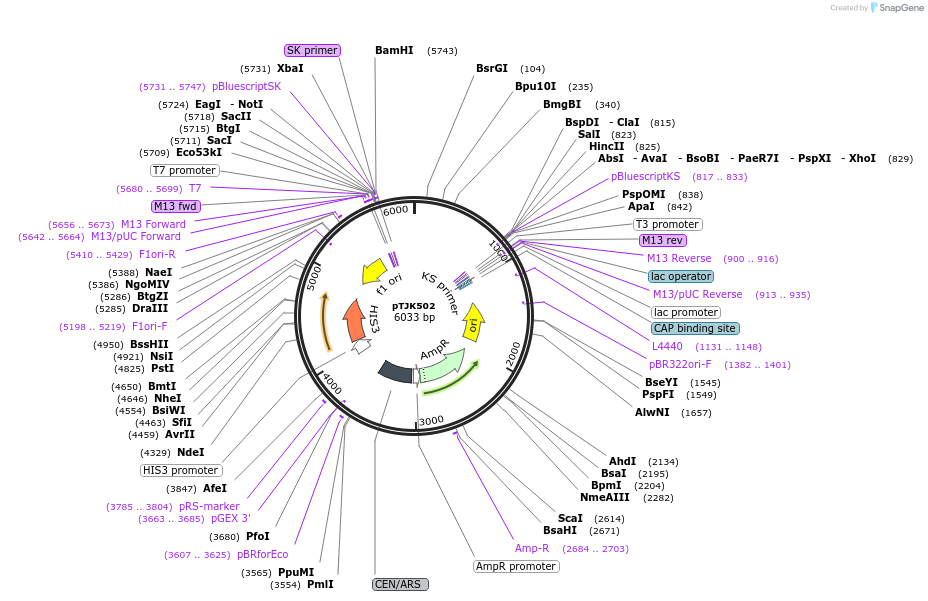
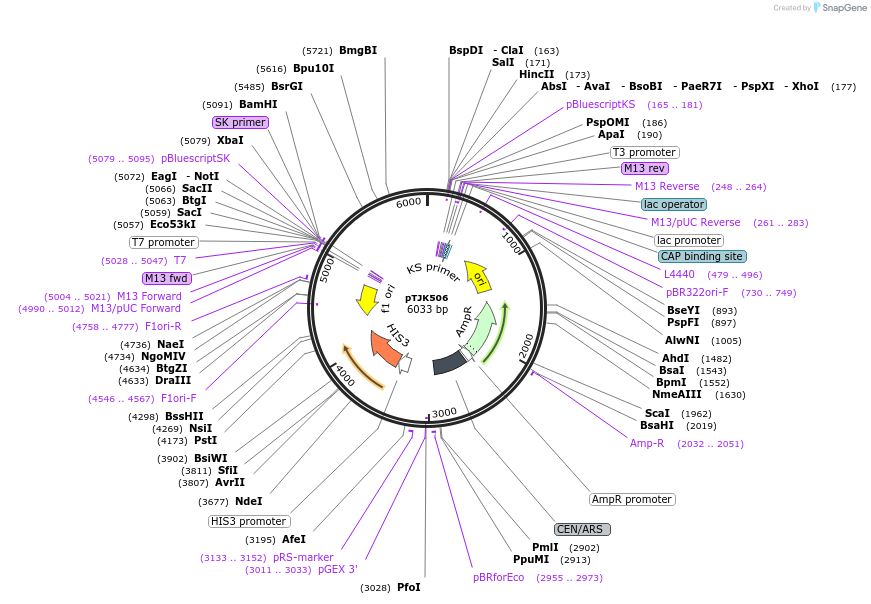
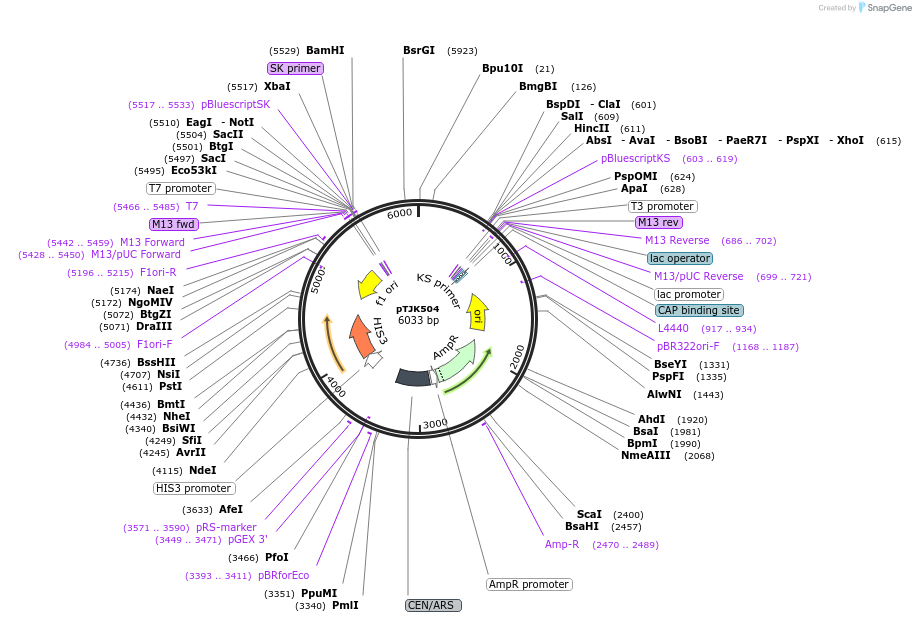
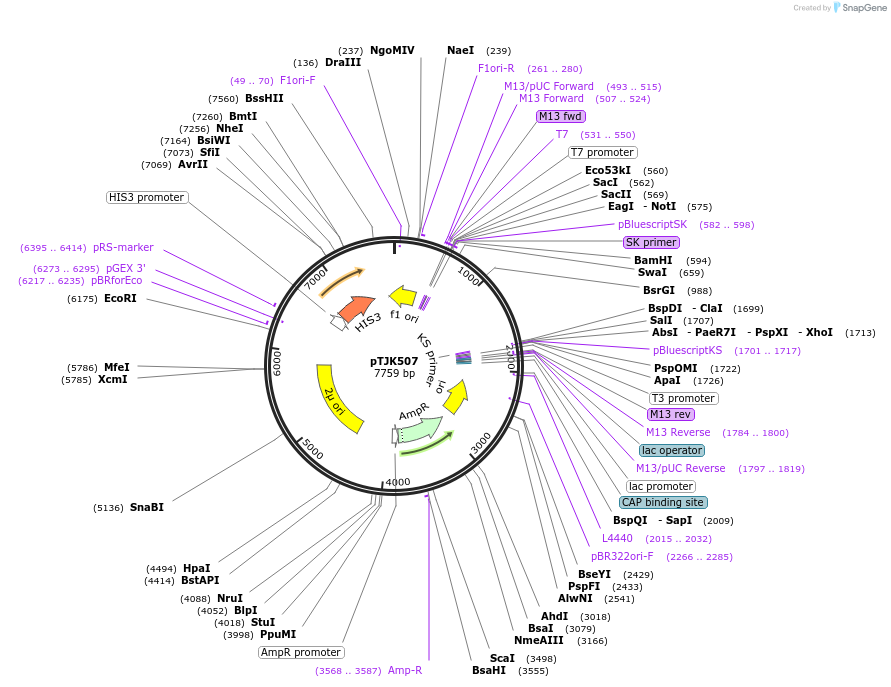
pTJK502
Plasmid#121457PurposeCEN HIS3 CNB1DepositorAvailable SinceMay 2, 2019AvailabilityAcademic Institutions and Nonprofits only -
-
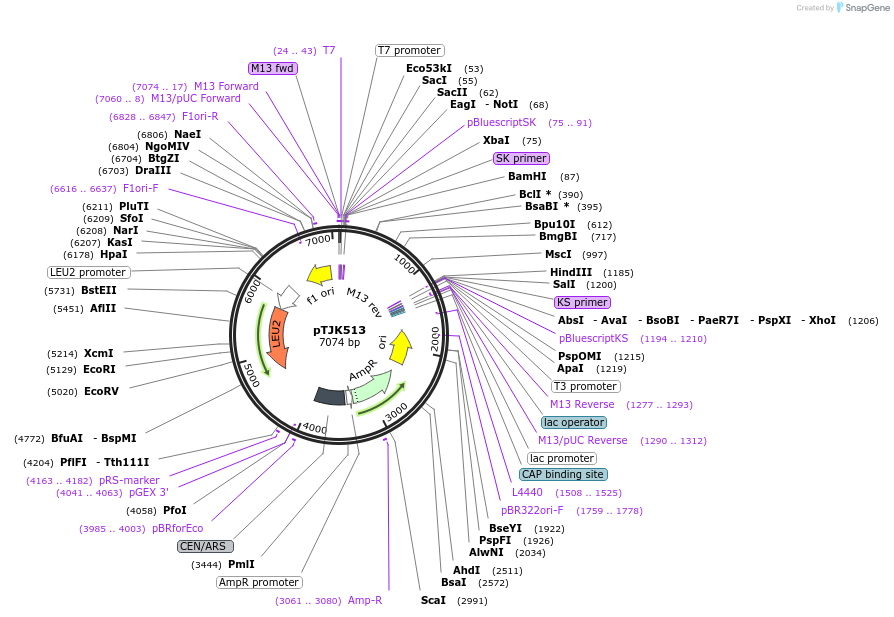
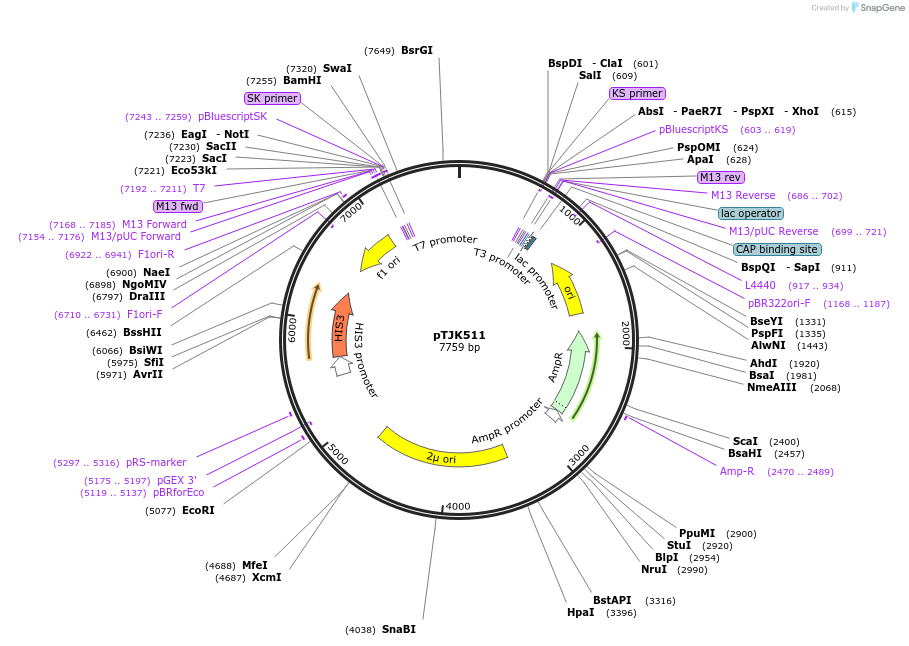
pTJK511
Plasmid#121468Purpose2µm HIS3 CNB1mutEF4DepositorInsertCNB1mutEF4 (CNB1 Budding Yeast)
UseYeast high copy 2 micron vectorMutationChanged aspartic acid 144 to alaninePromoterEndogenous genomic promoter regionAvailable SinceApril 23, 2019AvailabilityAcademic Institutions and Nonprofits only -
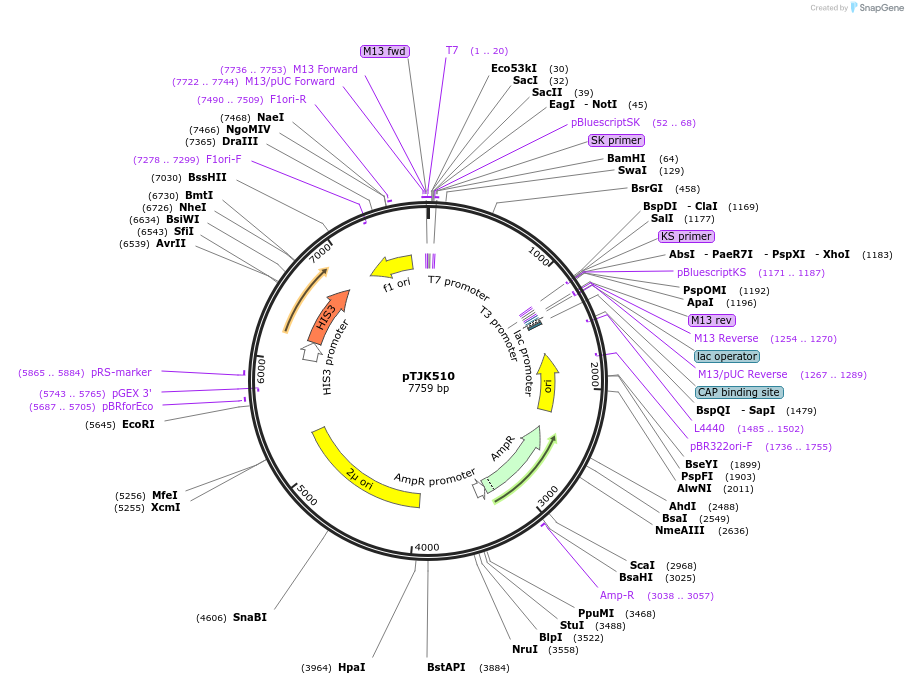
pTJK510
Plasmid#121465Purpose2µm HIS3 CNB1mutEF3DepositorInsertCNB1mutEF3 (CNB1 Budding Yeast)
UseYeast high copy 2 micron vectorMutationChanged aspartic acid 103 to alaninePromoterEndogenous genomic promoter regionAvailable SinceApril 23, 2019AvailabilityAcademic Institutions and Nonprofits only -
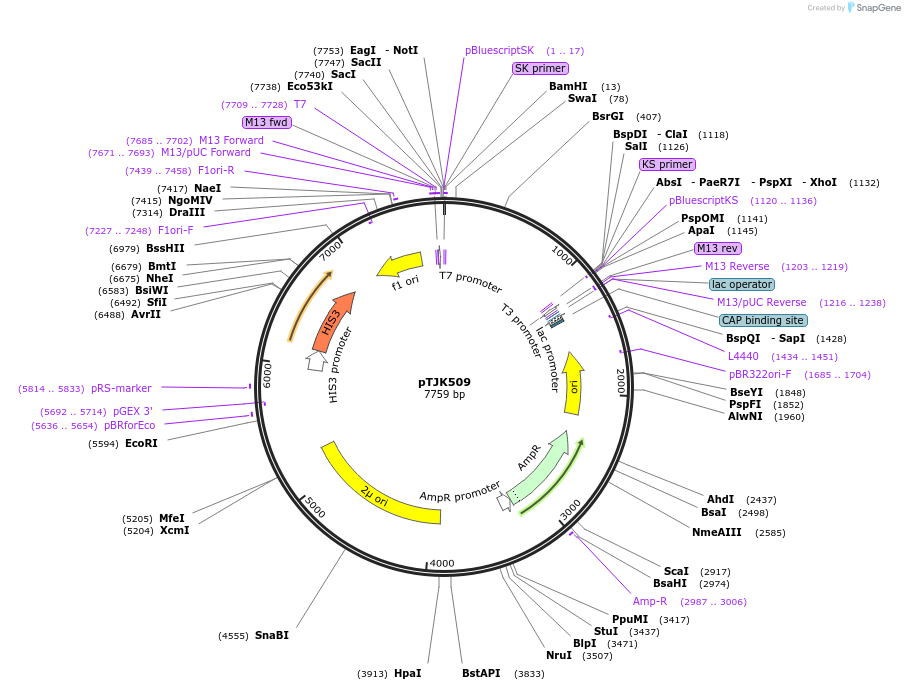
pTJK509
Plasmid#121464Purpose2µm HIS3 CNB1mutEF2DepositorInsertCNB1mutEF2 (CNB1 Budding Yeast)
UseYeast high copy 2 micron vectorMutationChanged aspartic acid 66 to alaninePromoterEndogenous genomic promoter regionAvailable SinceApril 23, 2019AvailabilityAcademic Institutions and Nonprofits only -
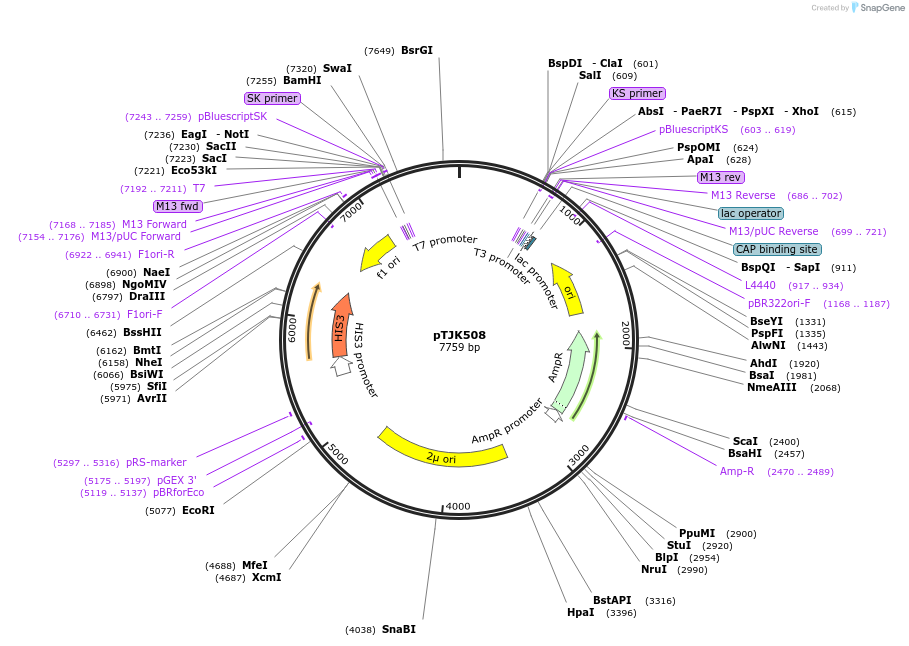
pTJK508
Plasmid#121463Purpose2µm HIS3 CNB1mutEF1DepositorInsertCNB1mutEF1 (CNB1 Budding Yeast)
UseYeast high copy 2 micron vectorMutationChanged aspartic acid 34 to alaninePromoterEndogenous genomic promoter regionAvailable SinceApril 23, 2019AvailabilityAcademic Institutions and Nonprofits only -
-
-
-
-
-
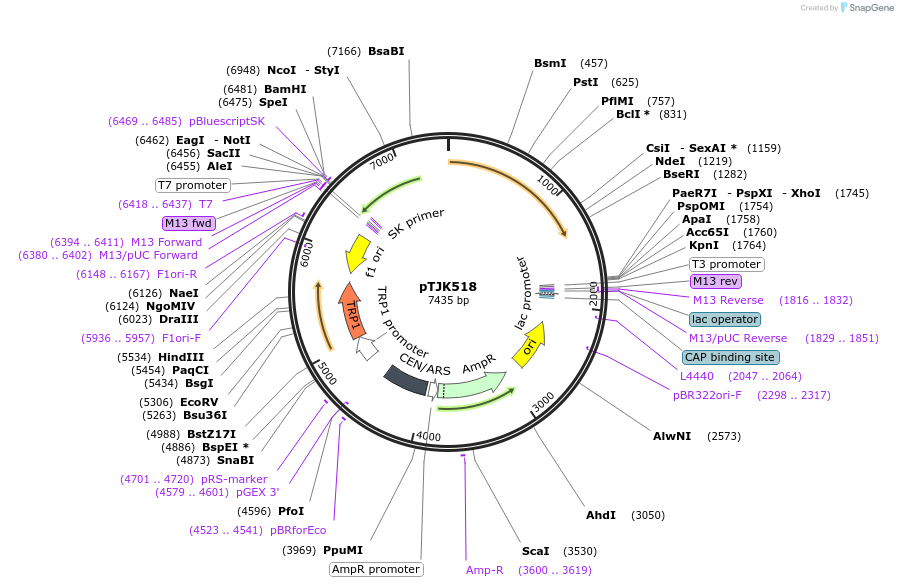
pTJK518
Plasmid#121475PurposeCEN TRP1 CNA1ΔAIDDepositorInsertCNA1deltaAID
UseYeast cen vectorMutationTruncated after amino acid 508PromoterEndogenous genomic promoter regionAvailable SinceApril 23, 2019AvailabilityAcademic Institutions and Nonprofits only -
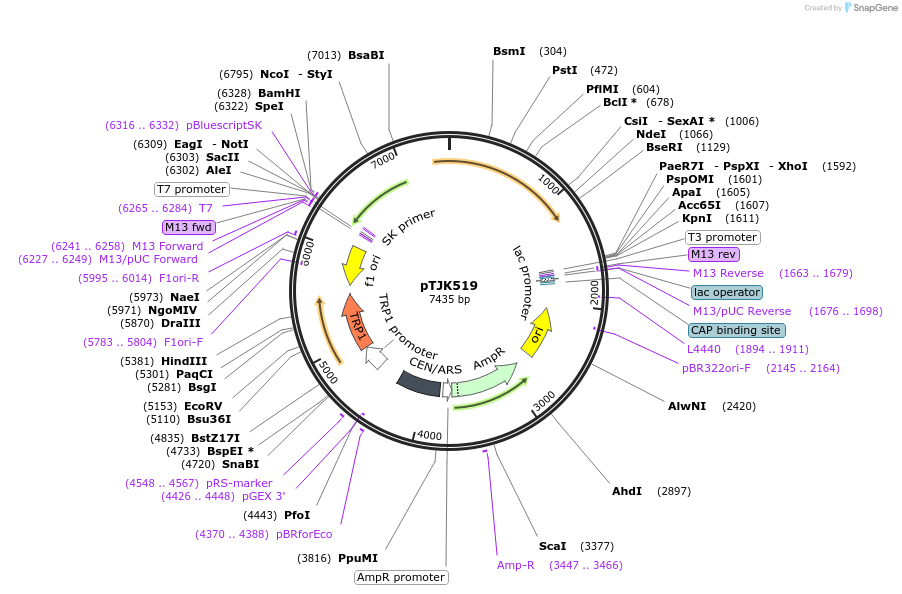
pTJK519
Plasmid#121476PurposeCEN TRP1 CNA1ΔCBDDepositorInsertCNA1deltaCBD
UseYeast cen vectorMutationTruncated after amino acid 453PromoterEndogenous genomic promoter regionAvailable SinceMarch 22, 2019AvailabilityAcademic Institutions and Nonprofits only -
-
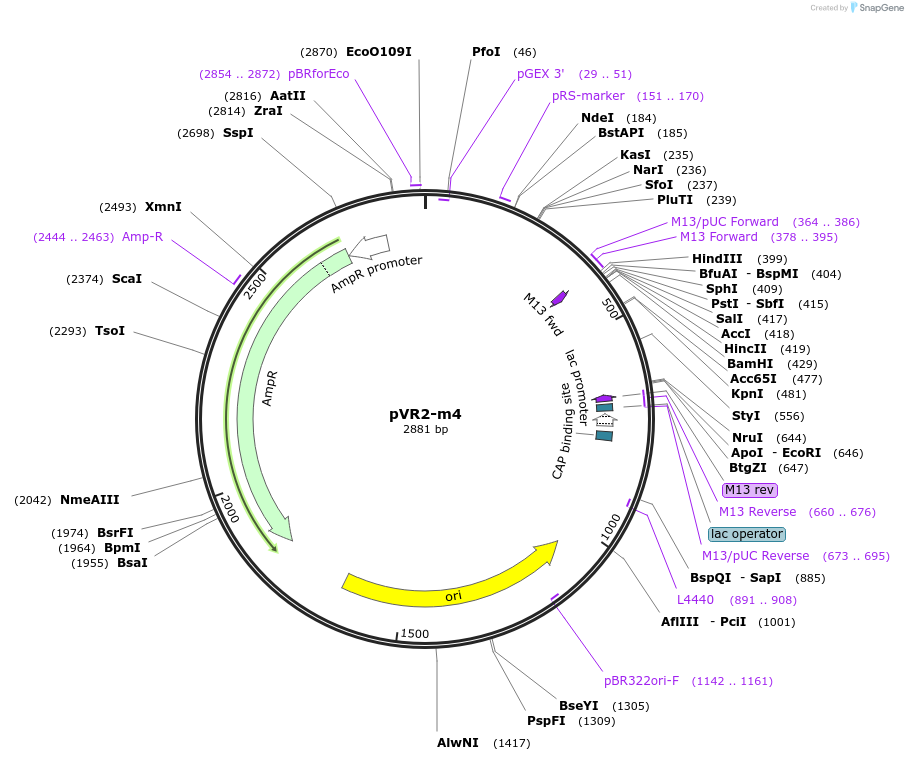
pVR2-m4
Plasmid#84723PurposepVR2 but with G to U and A to C mutations following the riboswitch aptamerDepositorInsertsE. coli RecA promoter
stretch lacking A's followed by 5' UTR of queC gene
ExpressionBacterialMutationG to U mutation at the first nucleotide after the…Available SinceMarch 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
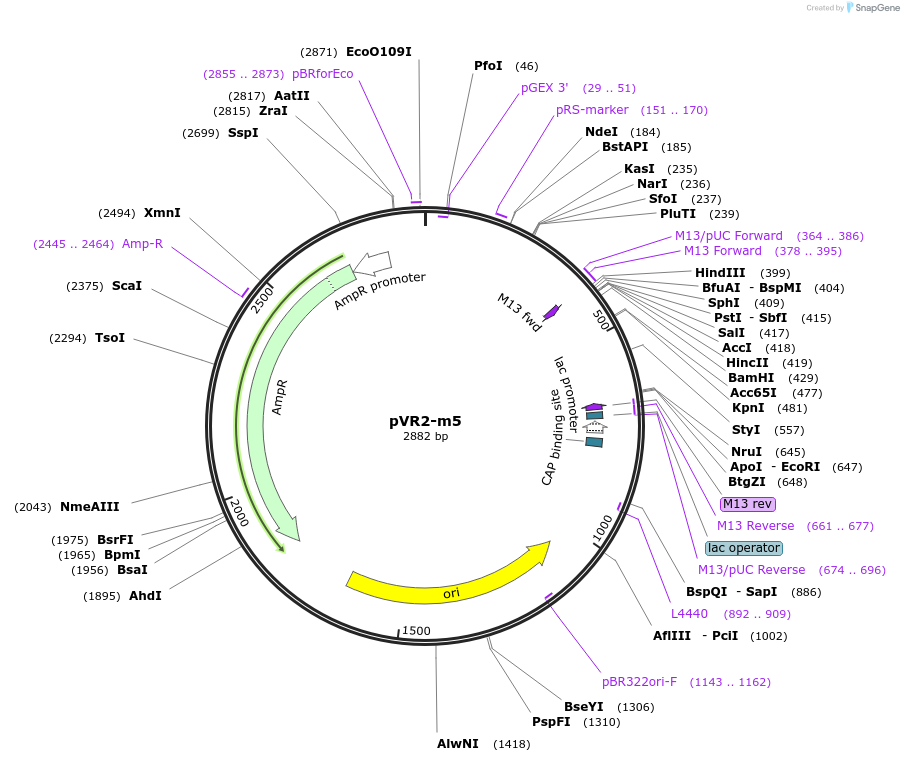
pVR2-m5
Plasmid#84724PurposepVR2 but with an insertion of 1 U immediately following the riboswitch aptamerDepositorInsertsE. coli RecA promoter
stretch lacking A's followed by 5' UTR of queC gene
ExpressionBacterialMutationinsertion of 1 U between positions 39 and 40 of a…Available SinceMarch 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
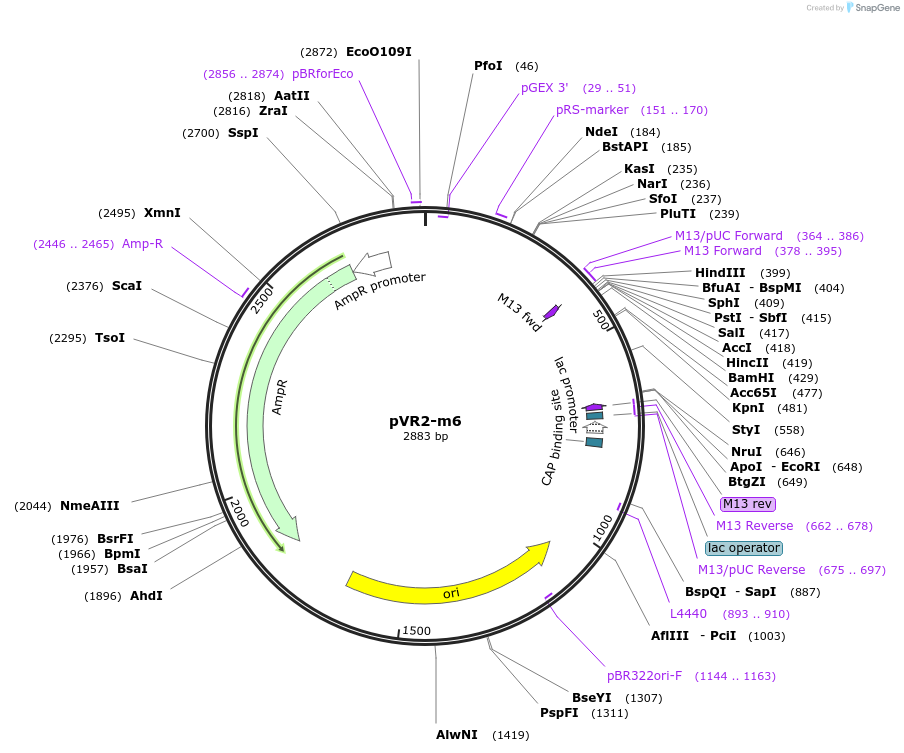
pVR2-m6
Plasmid#84725PurposepVR2 but with an insertion of 2 U's immediately following the riboswitch aptamerDepositorInsertsE. coli RecA promoter
stretch lacking A's followed by 5' UTR of queC gene
ExpressionBacterialMutationinsertion of 2 Us between positions 39 and 40 of …Available SinceMarch 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
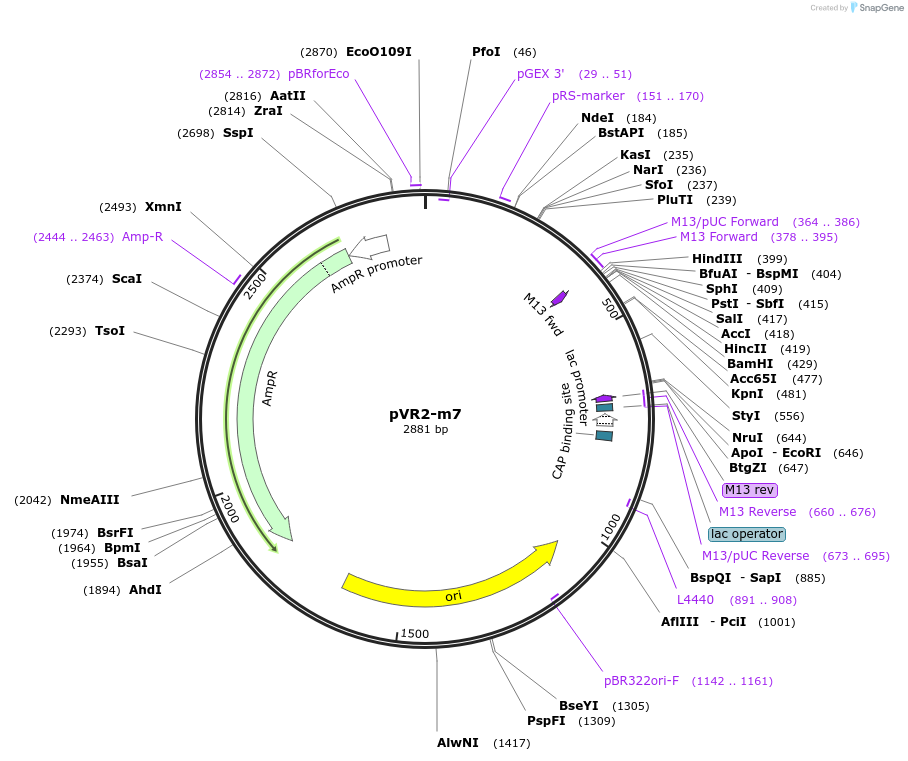
pVR2-m7
Plasmid#84726PurposepVR2 but with a G to U mutation immediately following the riboswitch aptamerDepositorInsertsE. coli RecA promoter
stretch lacking A's followed by 5' UTR of queC gene
ExpressionBacterialMutationG to U mutation at position 41 of annotated "…Available SinceMarch 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
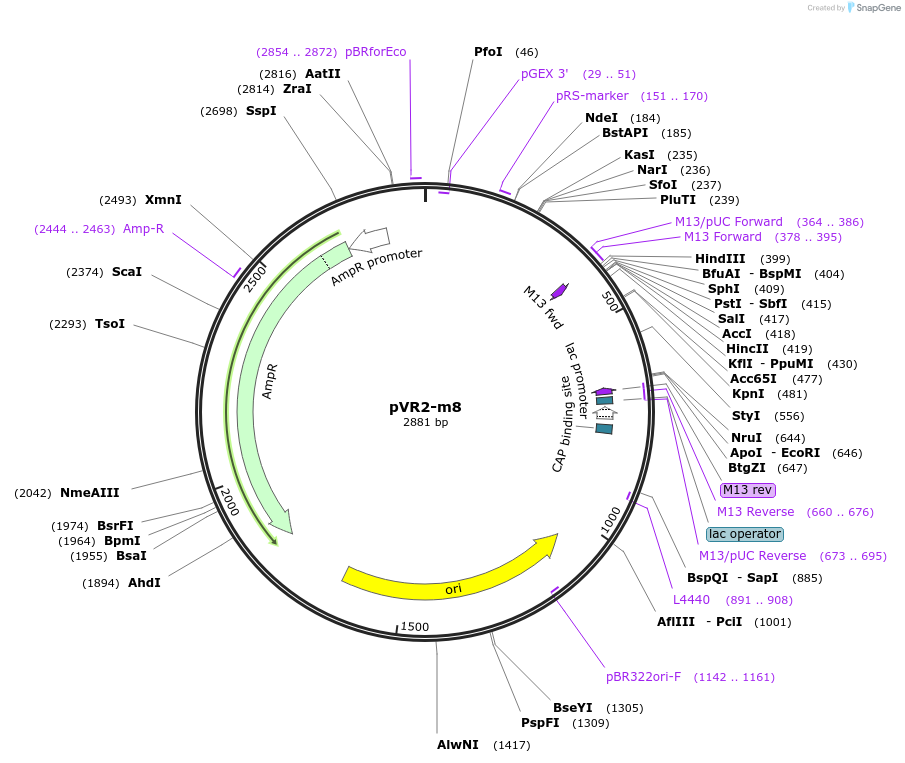
pVR2-m8
Plasmid#84727PurposepVR2 but with two G to U mutations immediately following the riboswitch aptamerDepositorInsertsE. coli RecA promoter
stretch lacking A's followed by 5' UTR of queC gene
ExpressionBacterialMutationG to U mutations at positions 40 and 41 of annota…Available SinceMarch 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
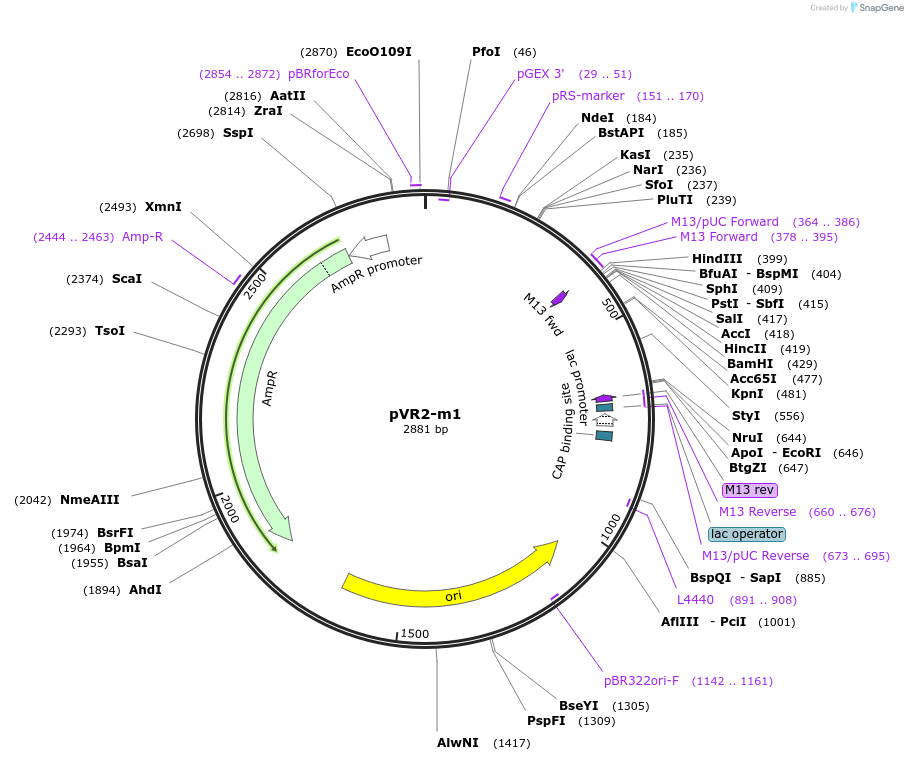
pVR2-m1
Plasmid#84720PurposepVR2 but with a C to U mutation in the riboswitch aptamerDepositorInsertsE. coli RecA promoter
stretch lacking A's followed by 5' UTR of queC gene
ExpressionBacterialMutationC to U mutation at position 13 of annotated "…Available SinceMarch 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
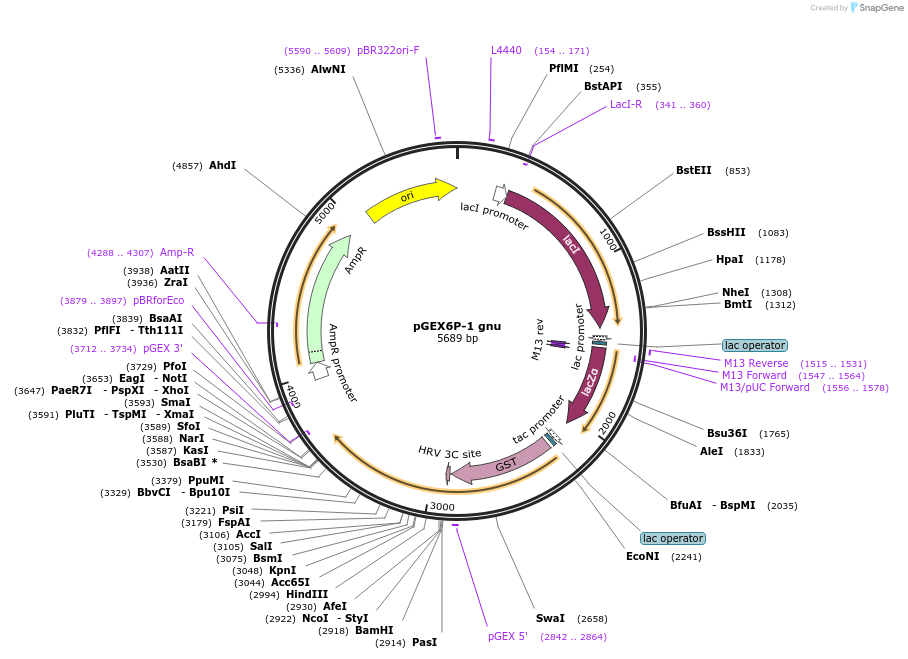
pGEX6P-1 gnu
Plasmid#112999PurposeFor protein expression and purification of full-length Drosophila gnuDepositorInsertfull length gnu (gnu Fly)
ExpressionBacterialAvailable SinceAug. 2, 2018AvailabilityAcademic Institutions and Nonprofits only -
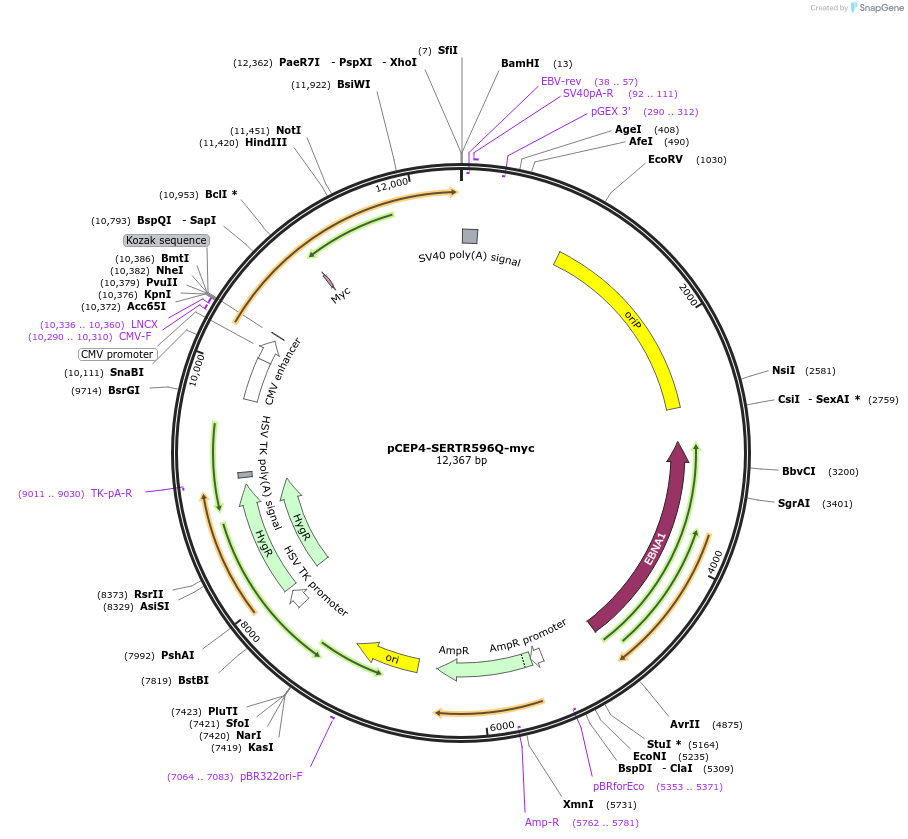
pCEP4-SERTR596Q-myc
Plasmid#107483PurposeMammalian expression plasmid for codon optimized mutant hSERT with internal MYC tag inserted in ECL2, mutation R596Q conferring gain of function phenotypeDepositorAvailable SinceMay 1, 2018AvailabilityAcademic Institutions and Nonprofits only -
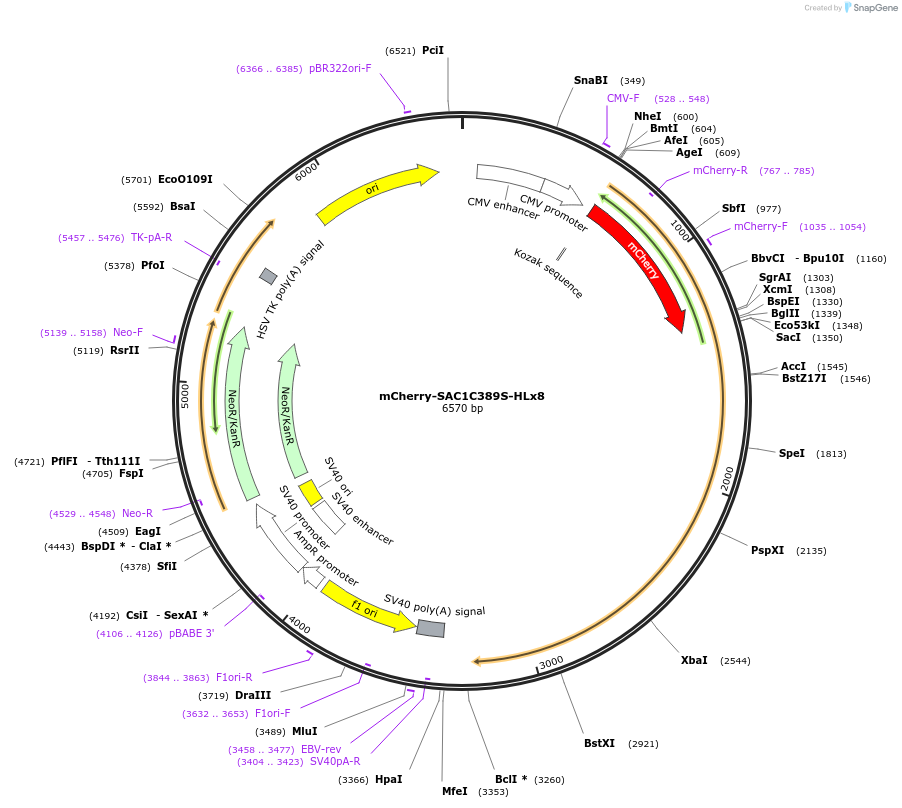
mCherry-SAC1C389S-HLx8
Plasmid#108140PurposeInactive SAC1 with extended linkerDepositorInsertmCherry:SACM1LC389S(1-520):[EAAAR]8:SACM1L(521-587) (SACM1L Human)
ExpressionMammalianMutationSACM1LC389S(1-520):[EAAAR]8:SACM1L(521-587)PromoterCMVAvailable SinceApril 6, 2018AvailabilityAcademic Institutions and Nonprofits only -
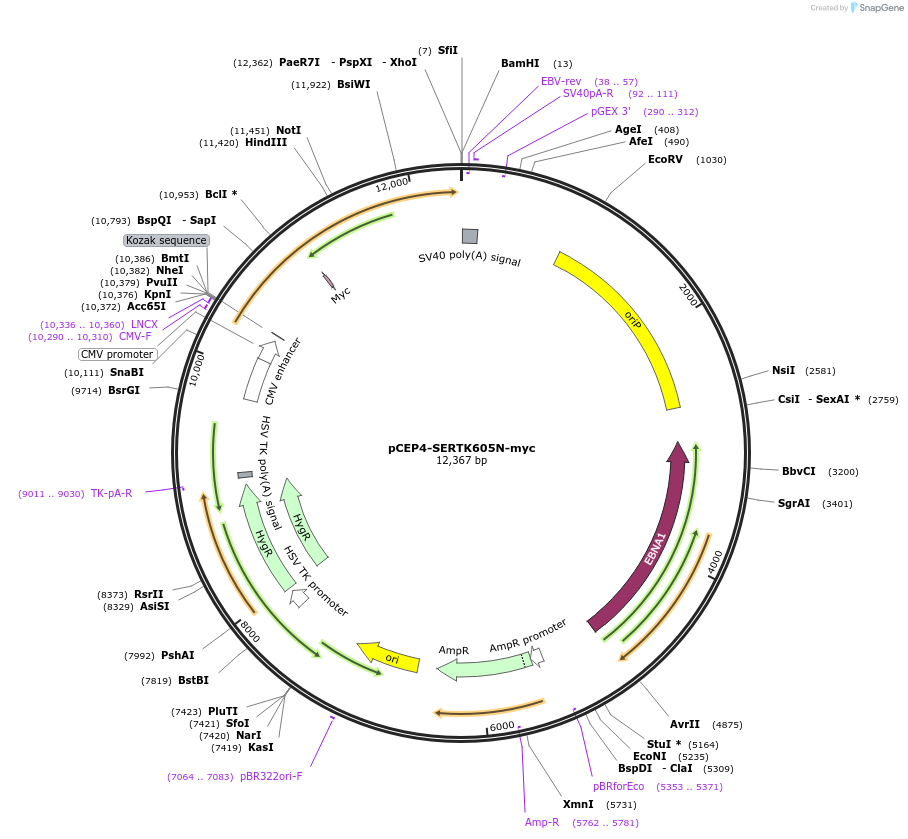
pCEP4-SERTK605N-myc
Plasmid#107475PurposeMammalian expression plasmid for codon optimized mutant hSERT with internal MYC tag inserted in ECL2, mutation K605N approxmately wildtype activity, decreased surface expressionDepositorAvailable SinceMarch 28, 2018AvailabilityAcademic Institutions and Nonprofits only