-
Depositing Lab
-
Publication
-
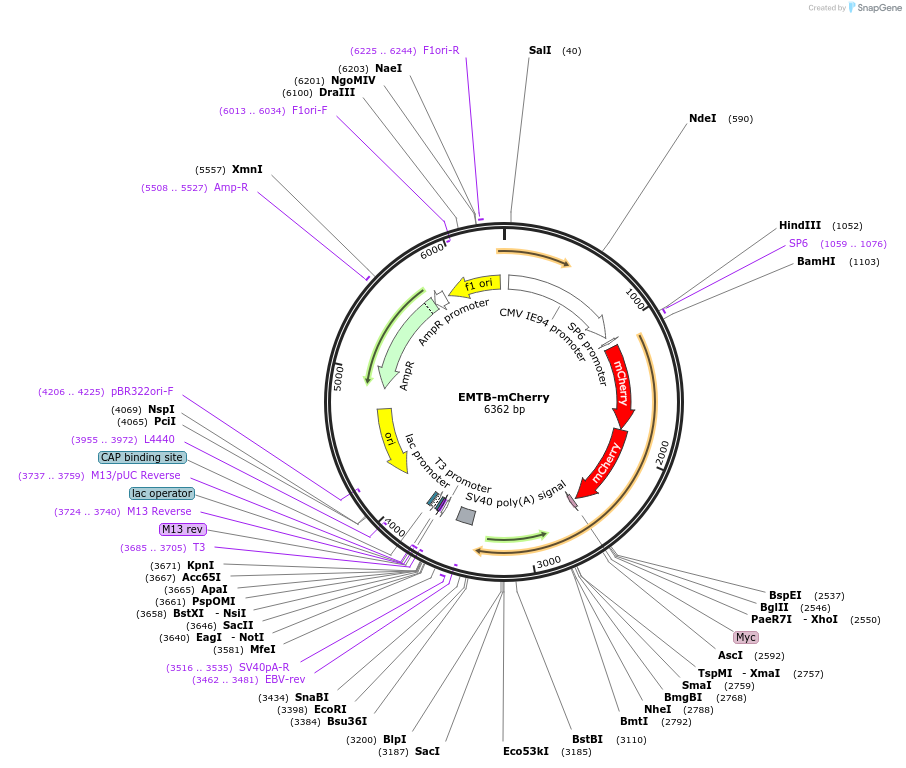
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 26742 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepCS2+
- Backbone size w/o insert (bp) 4097
-
Vector typeMammalian Expression ; probe for microtubules
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)DH5alpha
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert nameensconsin
-
Alt nameE-MAP-115
-
Alt nameEMTB
-
SpeciesH. sapiens (human)
-
Insert Size (bp)816
-
MutationMicrotubule binding-domain of ensconsin (amino acids 18-282 of GenBank reference sequence X73882)
-
GenBank IDX73882
-
Entrez GeneMAP7 (a.k.a. E-MAP-115, EMAP115)
-
Tags
/ Fusion Proteins
- 2x mCherry (N terminal on backbone)
- Myc (N terminal on insert)
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site XhoI (not destroyed)
- 3′ cloning site EcoRI (not destroyed)
- 5′ sequencing primer SP6
- 3′ sequencing primer T7
- (Common Sequencing Primers)
Resource Information
-
A portion of this plasmid was derived from a plasmid made byA EMTB construct was a gift from Chloë Bulinski (Columbia University)
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Note: It is difficult to sequence entirely through 2 tandem mCherrys. Users are encouraged to perform a double-digest on this plasmid with BamHI and EcoRI to verify the presence of a ~2.2kb band corresponding to the EMTB insert + 2XmCherry. Please see above attachment.
The microtubule binding domain of ensconsin (EMTB) was fused to multiple mCherry molecules in order to visualize microtubules. Please note that this plasmid has recombined and is currently only 2X mCherry.
mCherry sequences have point mutation at R154 (CGG to AGA) for optimal codon usage in Xenopus.
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
EMTB-mCherry was a gift from William Bement (Addgene plasmid # 26742 ; http://n2t.net/addgene:26742 ; RRID:Addgene_26742) -
For your References section:
Regulation of cytokinesis by Rho GTPase flux. Miller AL, Bement WM. Nat Cell Biol. 2009 Jan . 11(1):71-7. 10.1038/ncb1814 PubMed 19060892