-
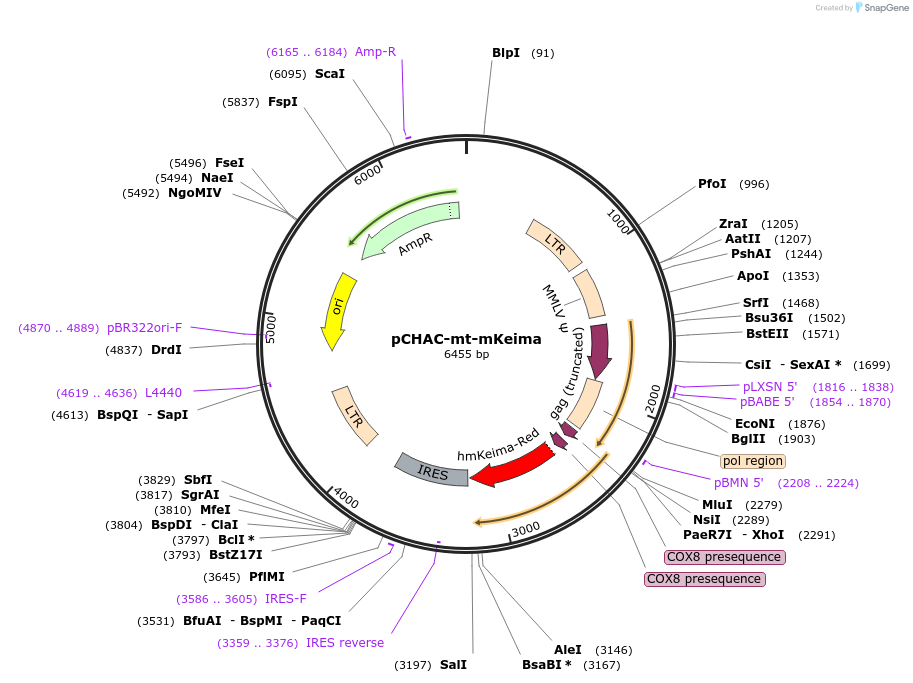
Purposeretrovirus construct for stable expressing mt-mKeima
-
Depositing Lab
-
Sequence Information
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |
|---|---|---|---|---|---|
| Plasmid | 72342 | Standard format: Plasmid sent in bacteria as agar stab | 1 | $94 | |
Backbone
-
Vector backbonepCHAC-MCS1-IRES-MCS2
-
Backbone manufacturerAlleleBiotechnology
- Backbone size w/o insert (bp) 5632
- Total vector size (bp) 6517
-
Vector typeRetroviral
-
Selectable markersnone
Growth in Bacteria
-
Bacterial Resistance(s)Ampicillin, 100 μg/mL
-
Growth Temperature37°C
-
Growth Strain(s)NEB Stable
-
Copy numberHigh Copy
Gene/Insert
-
Gene/Insert namemt-mKeima
-
Speciesstony coral Montipora
-
Insert Size (bp)894
-
GenBank IDAB209969
Cloning Information
- Cloning method Restriction Enzyme
- 5′ cloning site XhoI (not destroyed)
- 3′ cloning site SalI (not destroyed)
- 5′ sequencing primer TGACCTGGGAAGCCTTGGCT
- 3′ sequencing primer TTGCCAAAAGACGGCAATAT
- (Common Sequencing Primers)
Resource Information
-
A portion of this plasmid was derived from a plasmid made byThe insert mt-mKeima was cut out from the original construct pIND(SP1)-mt-mKeima obtained from Dr. Atsushi Miyawaki of Brain Science Institute, RIKEN, Japan
-
Articles Citing this Plasmid
Terms and Licenses
-
Academic/Nonprofit Terms
-
Industry Terms
- Not Available to Industry
Trademarks:
- Zeocin® is an InvivoGen trademark.
Depositor Comments
Please also cite the original paper describing the use of mt-mkeima by Dr. Atsushi Miyawaki's lab at RIKEN, Japan.
Hiroyuki Katayama et al, 2011. A sensitive and quantitative technique for detecting autophagic events based on lysosomal delivery. Chemistry and Bioogy 18(8): 1042-1052
These plasmids were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
pCHAC-mt-mKeima was a gift from Richard Youle (Addgene plasmid # 72342 ; http://n2t.net/addgene:72342 ; RRID:Addgene_72342) -
For your References section:
The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy. Lazarou M, Sliter DA, Kane LA, Sarraf SA, Wang C, Burman JL, Sideris DP, Fogel AI, Youle RJ. Nature. 2015 Aug 20;524(7565):309-14. doi: 10.1038/nature14893. Epub 2015 Aug 12. 10.1038/nature14893 PubMed 26266977