BioBloom Pooled Libraries
(Pooled Library #1000000273)
-
Purpose
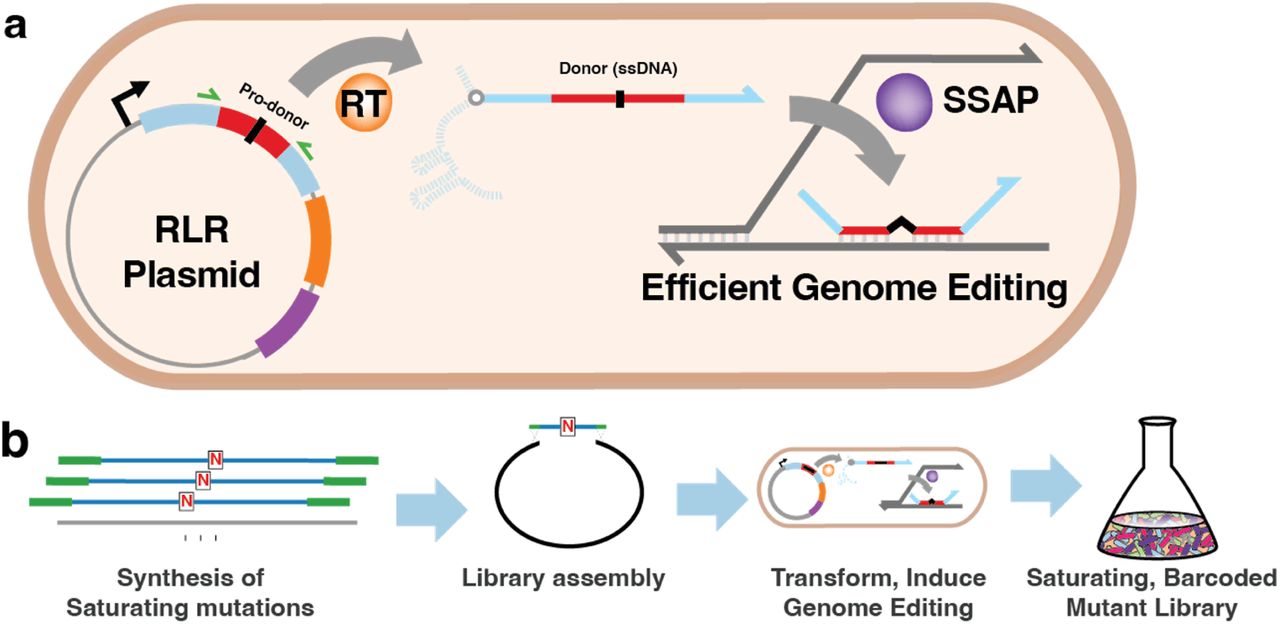
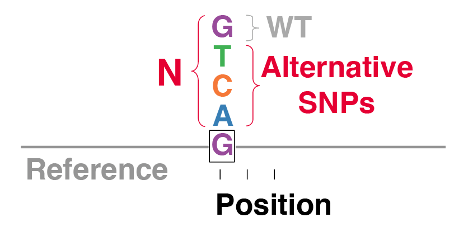
BioBloom is a method for barcoded saturation mutagenesis of bacterial genomes. The BioBloom Pooled Libraries contain nearly every possible single-base transition and transversion mutation of the Escherichia coli genome, with an identifying “pro-donor” barcode sequence to facilitate pooled selection experiments.
BioBloom can be used to quickly identify beneficial mutations in a given selection condition that improve E. coli growth, and produce genome-wide datasets describing many such mutations.
This kit contains 3 tubes, each an independently-prepared replicate of the BioBloom-Ec-2.0 library. Best results are achieved by performing experiments in triplicate, using these three replicate libraries to seed each replicate.
-
Vector Backbone
-
Depositing Labs
Ordering
| Item | Catalog # | Description | Quantity | Price (USD) | |||
|---|---|---|---|---|---|---|---|
| Pooled Library | 1000000273 | BioBloom Pooled Libraries | 1 | $1544 | Add to Cart | ||
Library Details
-
SpeciesE. coli
-
Number of genes~20 million
-
Insert infoThe unique sequences used to identify each mutant are 82 bp in length. Each mutant has a unique retron “pro-donor” sequence that can be used to identify the mutant as a barcode, analogous to a CRISPR guide sequence.
Library Shipping
The BioBloom Libraries will be shipped frozen on dry ice.
-
Aliquots1 mL of confluent glycerol stock (~1 billion cells)*
*See depositor comments for additional details
Resource Information
-
Protocols
-
Depositor Data
-
Scripts
-
Terms and Licenses
Academic/Nonprofit TermsIndustry Terms- Not Available to Industry
Trademarks- Zeocin® is an InvivoGen trademark.
Depositor Comments
*The 1 mL glycerol stocks should cover the library well. We recommend recovery/outgrowth of these stocks and immediate use for experiments, but you may wish to cryopreserve material from this outgrowth, to produce “T2” stocks which may not sample a full saturation library as completely, but still be useful for some experiments.
Please visit https://doi.org/10.64898/2026.02.04.703572 (Link opens in a new window)for bioRxiv report.
These pooled libraries were created by your colleagues. Please acknowledge the Principal Investigator, cite the article in which the plasmids were described, and include Addgene in the Materials and Methods of your future publications.
-
For your Materials & Methods section:
BioBloom Pooled Libraries were a gift from Pioneer Labs (Addgene #1000000273 ; http://n2t.net/addgene:1000000273 ; RRID:Addgene_1000000273) -
For your References section:
BioBloom, a method for barcoded saturation mutagenesis of an entire bacterial genome. Schubert M, Sukarto E, Martin FR, Mancuso JE, Spens A, Pedersen T, Isaev K, Greene H, Hicks ND, Nattermann U, Liu J, Stork DA, DeBenedictis EA. bioRxiv. 2026 Feb 04. doi: 10.64898/2026.02.04.703572. https://doi.org/10.64898/2026.02.04.703572


