We narrowed to 19,218 results for: REV;
-
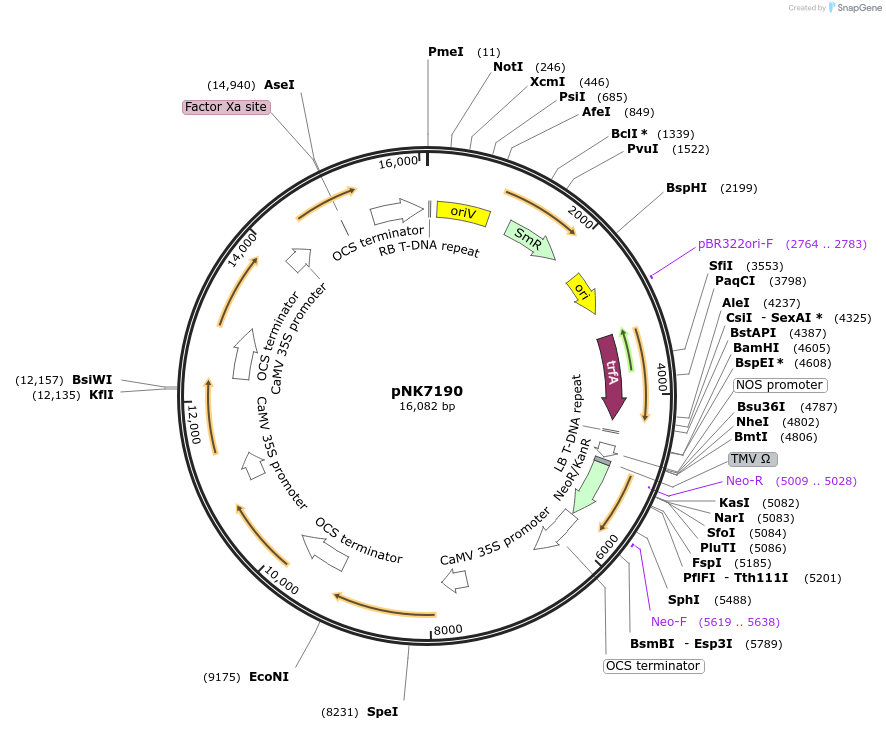
Plasmid#219763PurposeMoClo-compatible Level M for expression nnLuz, nnH3H, nnCPH in plants (N.benthamiana, BY2 cells)DepositorInsertnnLuz, nnH3H, nnCPH
ExpressionPlantAvailable SinceOct. 17, 2024AvailabilityAcademic Institutions and Nonprofits only -
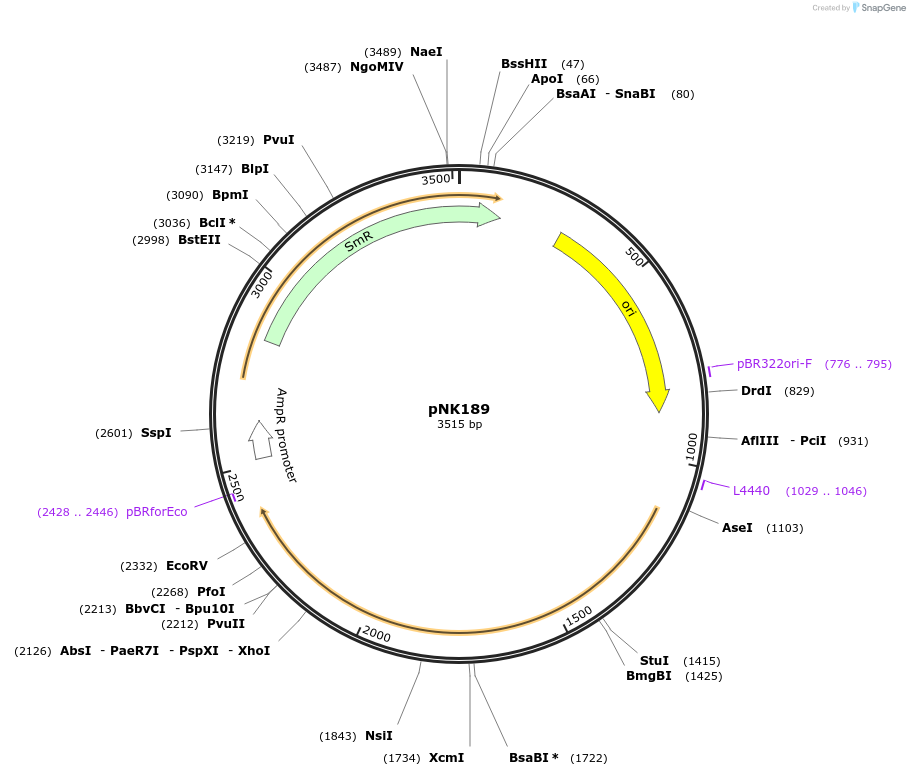
pNK189
Plasmid#219743PurposeMoClo-compatible Level 0 promoterless vector encoding mutant of Neonothopanus nambi hispidin-3-hydroxylase nnH3H_v2 codon-optimised for expression in Homo sapiensDepositorInsertmutant of fungal hispidin-3-hydroxylase
UseSynthetic BiologyMutationD37E, V181I, S323M, M385KAvailable SinceJuly 12, 2024AvailabilityAcademic Institutions and Nonprofits only -
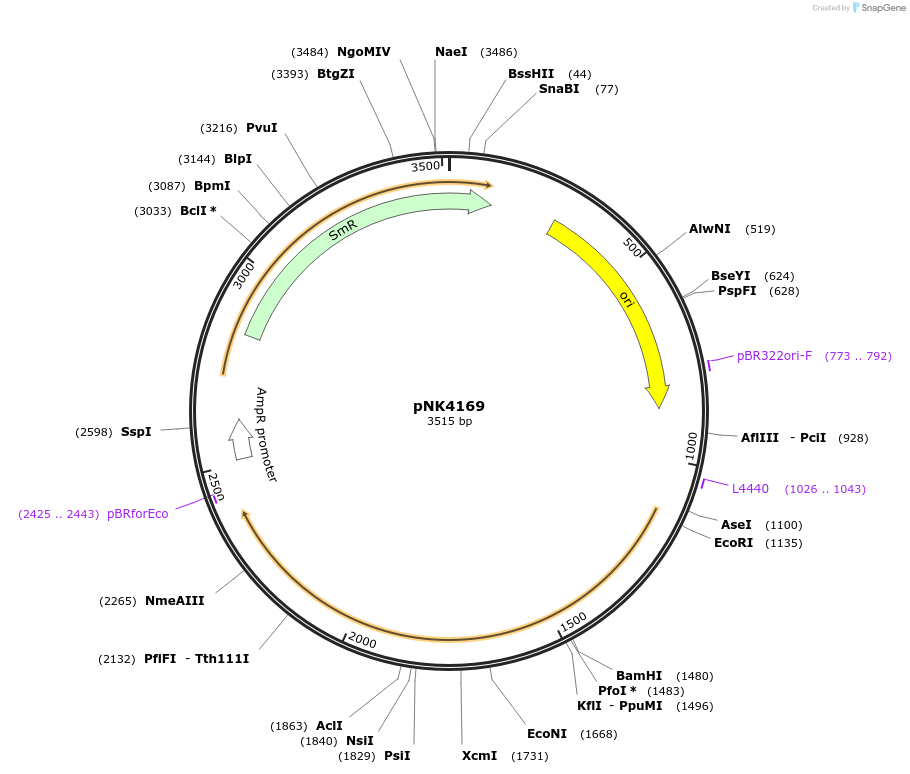
pNK4169
Plasmid#219744PurposeMoClo-compatible Level 0 promoterless vector encoding mutant of Neonothopanus nambi hispidin-3-hydroxylase nnH3H_v2 codon-optimised for expression in Pichia pastorisDepositorInsertmutant of fungal hispidin-3-hydroxylase
UseSynthetic BiologyMutationD37E, V181I, S323M, M385KAvailable SinceJuly 12, 2024AvailabilityAcademic Institutions and Nonprofits only -
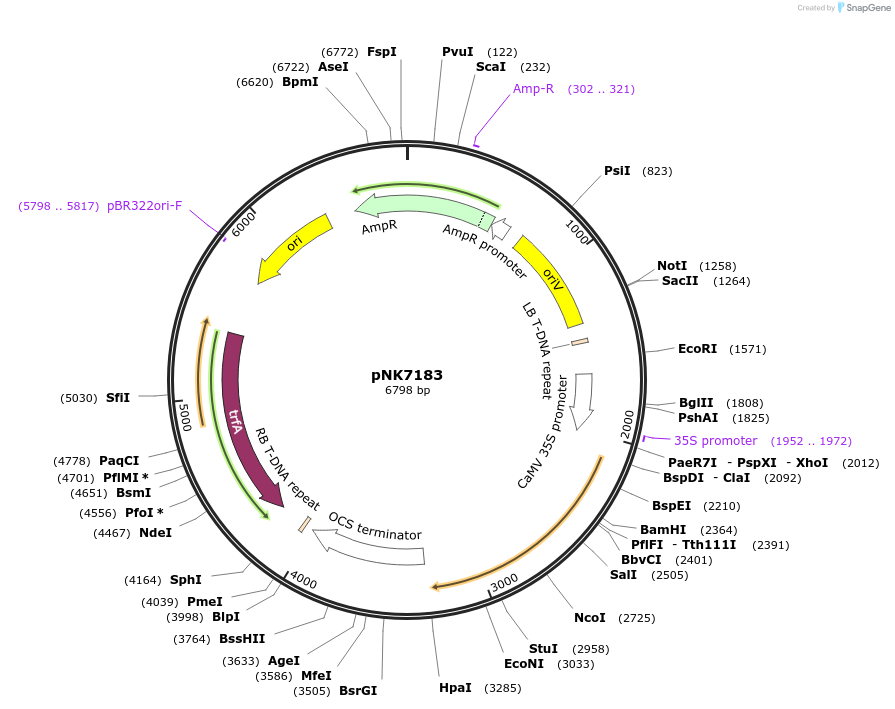
pNK7183
Plasmid#219760PurposeMoClo-compatible Level 1 for expression PzPKS2 in plants (N.benthamiana, BY2 cells)DepositorInsertPzPKS2
ExpressionPlantPromoterp35s_0.4kb - 5'UTR TMV omegaAvailable SinceJuly 12, 2024AvailabilityAcademic Institutions and Nonprofits only -
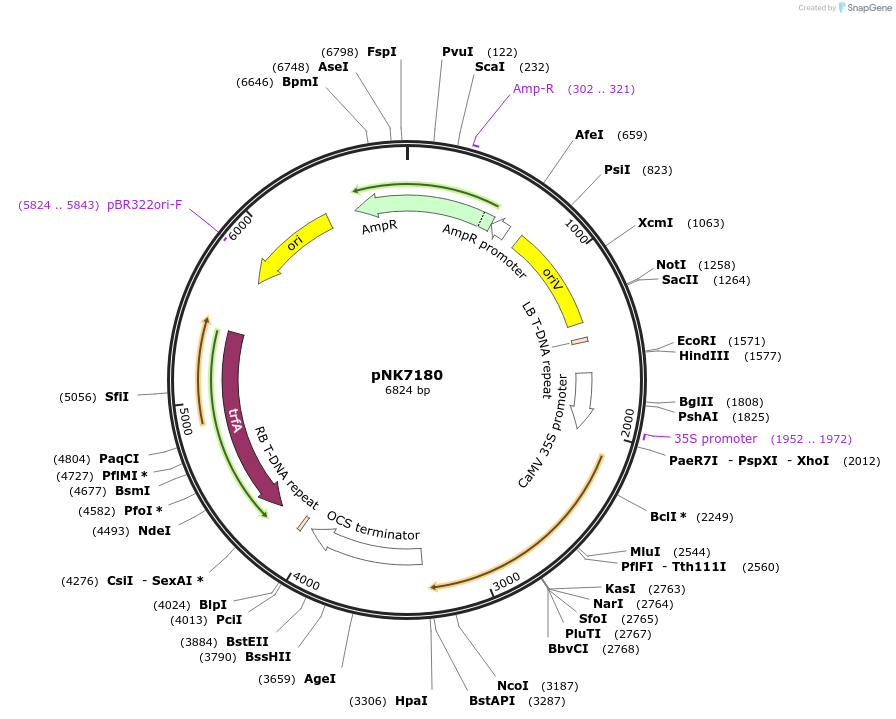
pNK7180
Plasmid#219762PurposeMoClo-compatible Level 1 for expression HmS in plants (N.benthamiana, BY2 cells)DepositorInsertHmS
ExpressionPlantPromoterp35s_0.4kb - 5'UTR TMV omegaAvailable SinceJuly 12, 2024AvailabilityAcademic Institutions and Nonprofits only -
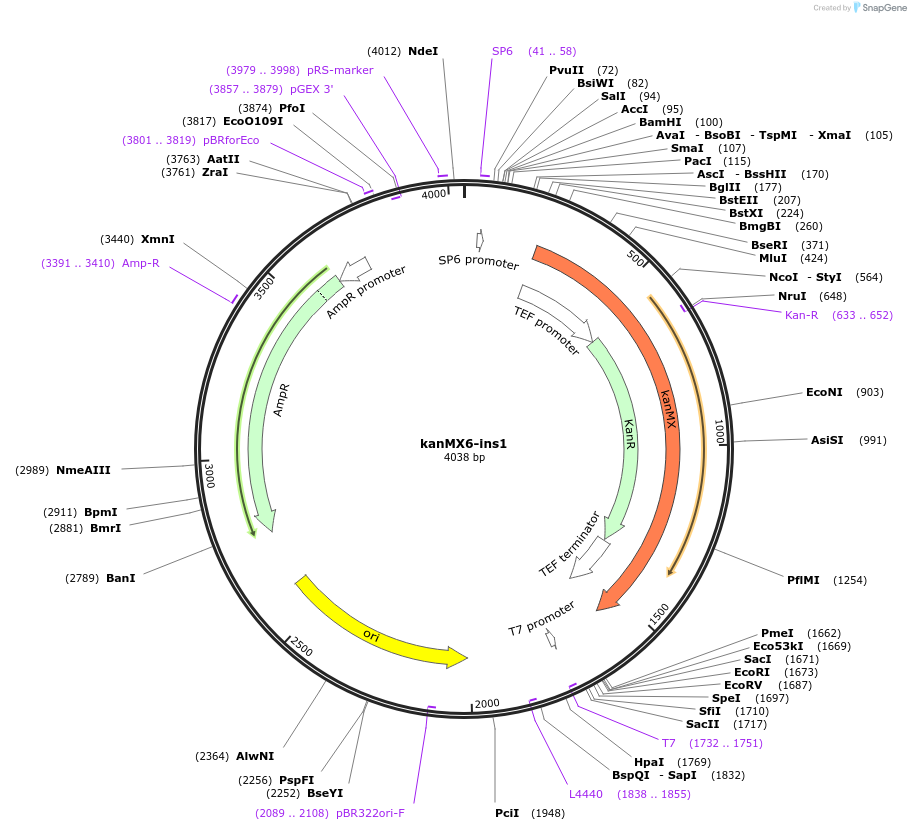
kanMX6-ins1
Plasmid#195038PurposepFA6a derived selection cassette flanked with transcription terminators (tDEG1/tDEG1), allows genome modification without disruption of insertion neighboring genes by transcription interferenceDepositorInsertKanR
UseYeast genomic targetingTagsS. cerevisiae DEG1 terminator, A. gossypii TEF pr…Available SinceFeb. 9, 2023AvailabilityAcademic Institutions and Nonprofits only -
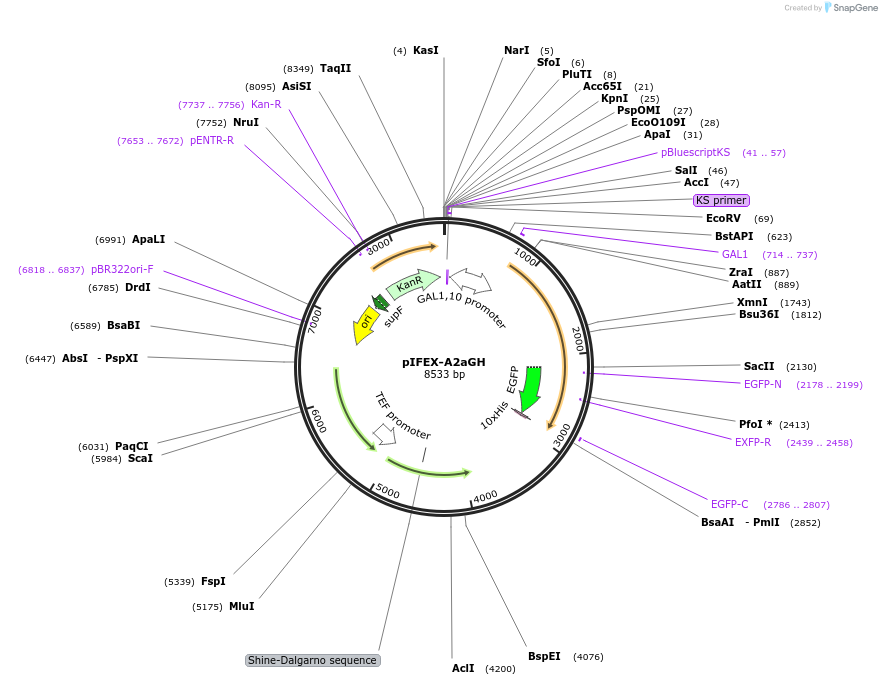
pIFEX-A2aGH
Plasmid#160542PurposepITY4 derivative reporter plasmid encoding Ashbya gossypii PTEF1-Saccharomyces cerevisiae FEX1-MFaT; as well as PGAL1-Homo sapiens ADORA2A-EGFP-10HIS-MFaT.DepositorInsertExpressionYeastMutationN/AAvailable SinceSept. 30, 2022AvailabilityAcademic Institutions and Nonprofits only -
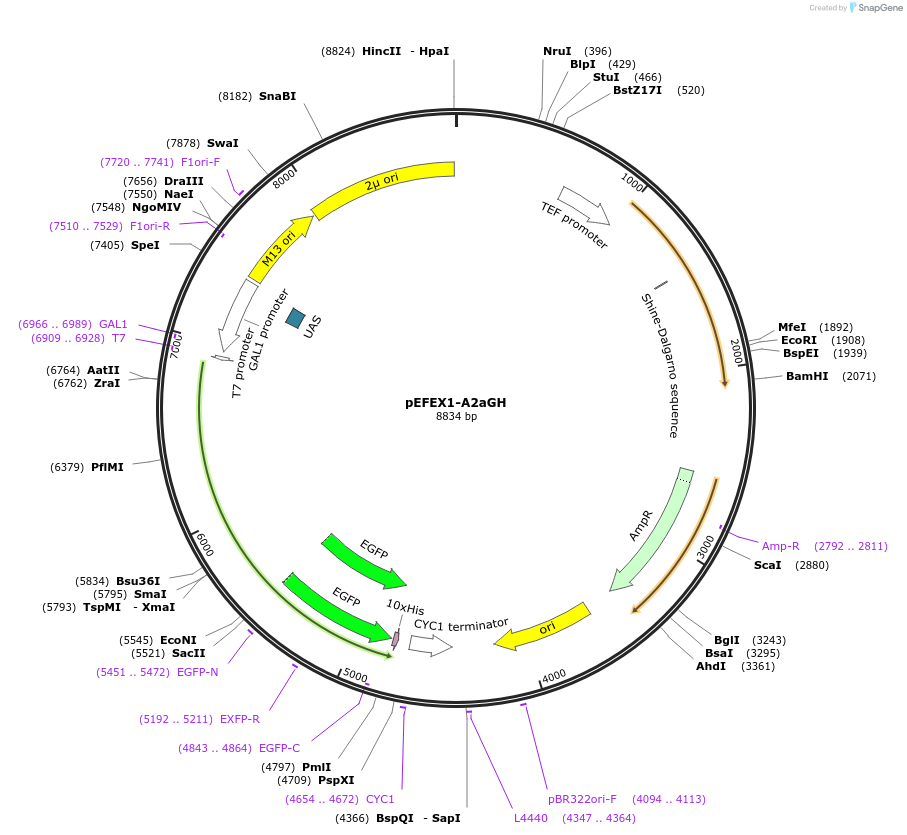
pEFEX1-A2aGH
Plasmid#160543PurposepYES2 derivative reporter plasmid encoding Ashbya gossypii PTEF1-Saccharomyces cerevisiae FEX1-MFaT; as well as PGAL1-Homo sapiens ADORA2A-EGFP-10HIS-MFaT.DepositorInsertExpressionYeastMutationN/AAvailable SinceSept. 30, 2022AvailabilityAcademic Institutions and Nonprofits only -
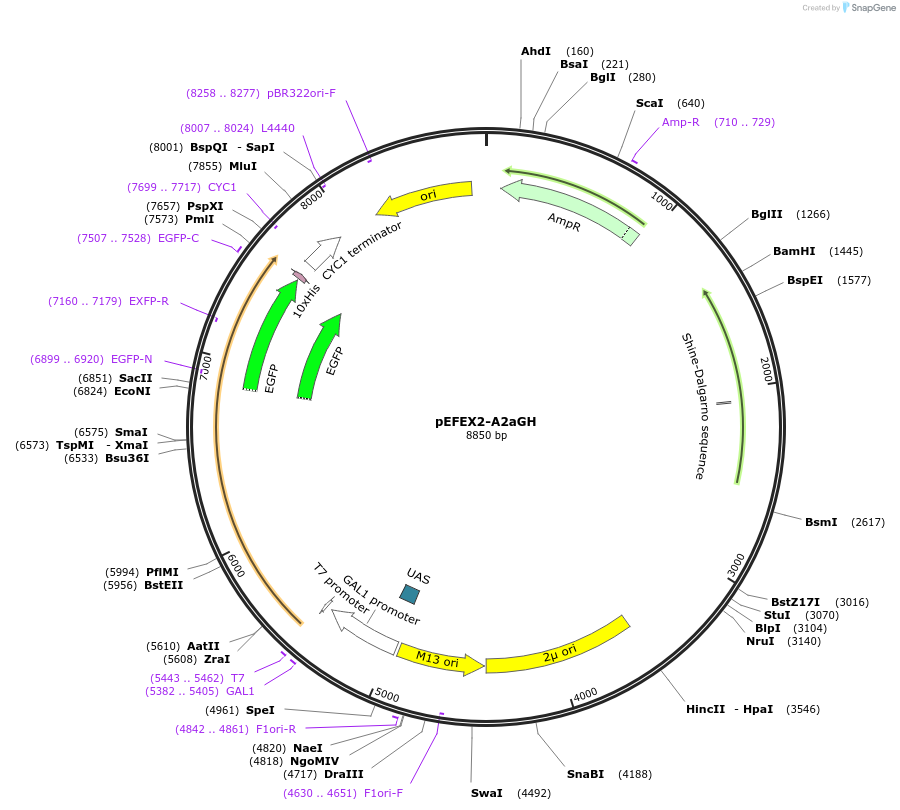
pEFEX2-A2aGH
Plasmid#160544PurposepYES2 derivative reporter plasmid encoding PPGI1-Saccharomyces cerevisiae FEX1-MFaT; as well as PGAL1-Homo sapiens ADORA2A-EGFP-10HIS-MFaT.DepositorInsertExpressionYeastMutationN/AAvailable SinceSept. 30, 2022AvailabilityAcademic Institutions and Nonprofits only -
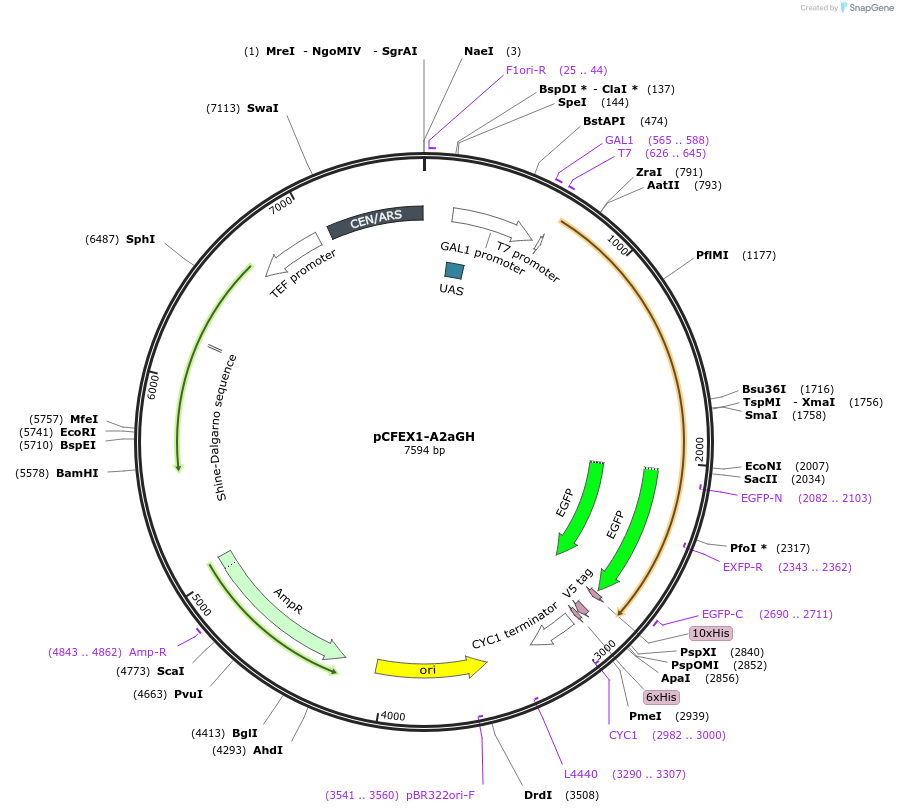
pCFEX1-A2aGH
Plasmid#160546PurposepYC2/CT derivative reporter plasmid encoding Ashbya gossypii PTEF1-Saccharomyces cerevisiae FEX1-MFaT; as well as PGAL1-Homo sapiens ADORA2A-EGFP-10HIS-MFaT.DepositorInsertExpressionYeastMutationN/AAvailable SinceSept. 30, 2022AvailabilityAcademic Institutions and Nonprofits only -
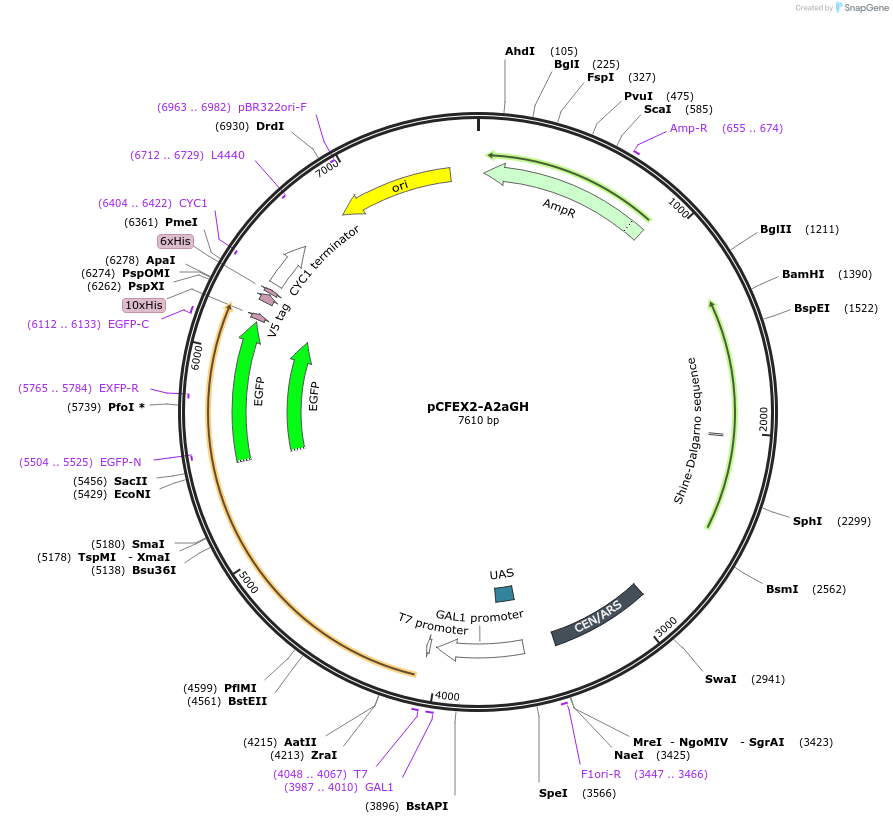
pCFEX2-A2aGH
Plasmid#160547PurposepYC2/CT derivative reporter plasmid encoding PPGI1-Saccharomyces cerevisiae FEX1-MFaT; as well as PGAL1-Homo sapiens ADORA2A-EGFP-10HIS-MFaT.DepositorInsertExpressionYeastMutationN/AAvailable SinceSept. 30, 2022AvailabilityAcademic Institutions and Nonprofits only -
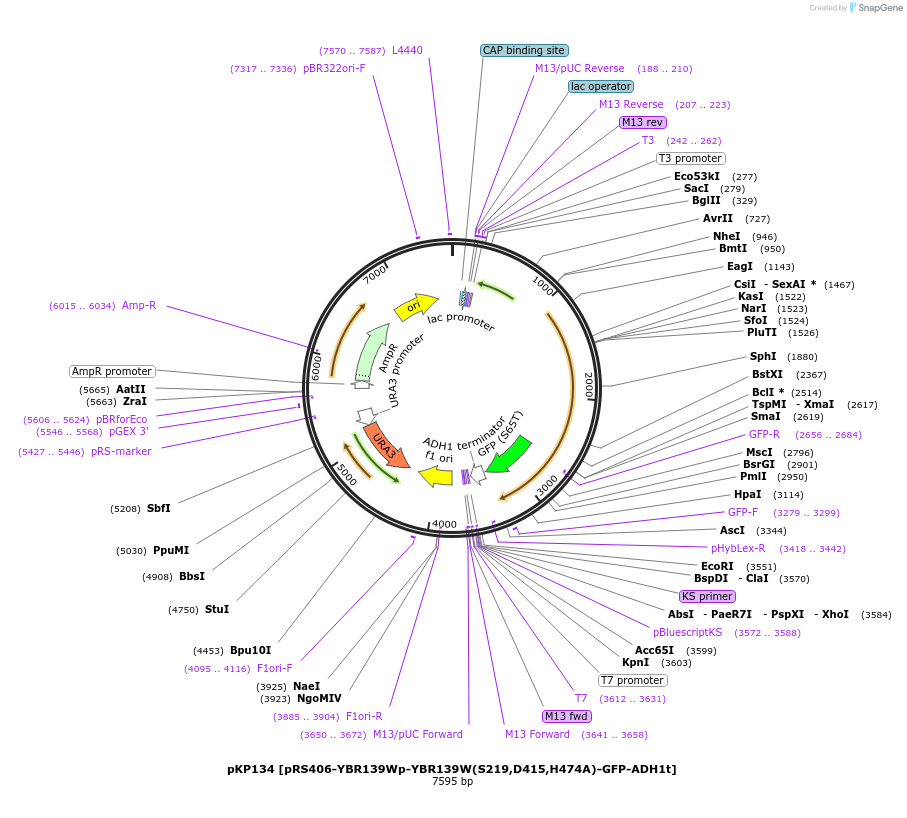
pKP134 [pRS406-YBR139Wp-YBR139W(S219,D415,H474A)-GFP-ADH1t]
Plasmid#106469PurposeExpresses Atg42/Ybr139w(S219,D415,H474A) with a C-terminal GFP tag in yeast cellsDepositorInsertYBR139W(S219,D415,H474A)
TagsGFPExpressionBacterial and YeastMutationChanged Serine 219, Aspartate 415 and Histidine 4…PromoterYBR139WAvailable SinceJune 5, 2018AvailabilityAcademic Institutions and Nonprofits only -
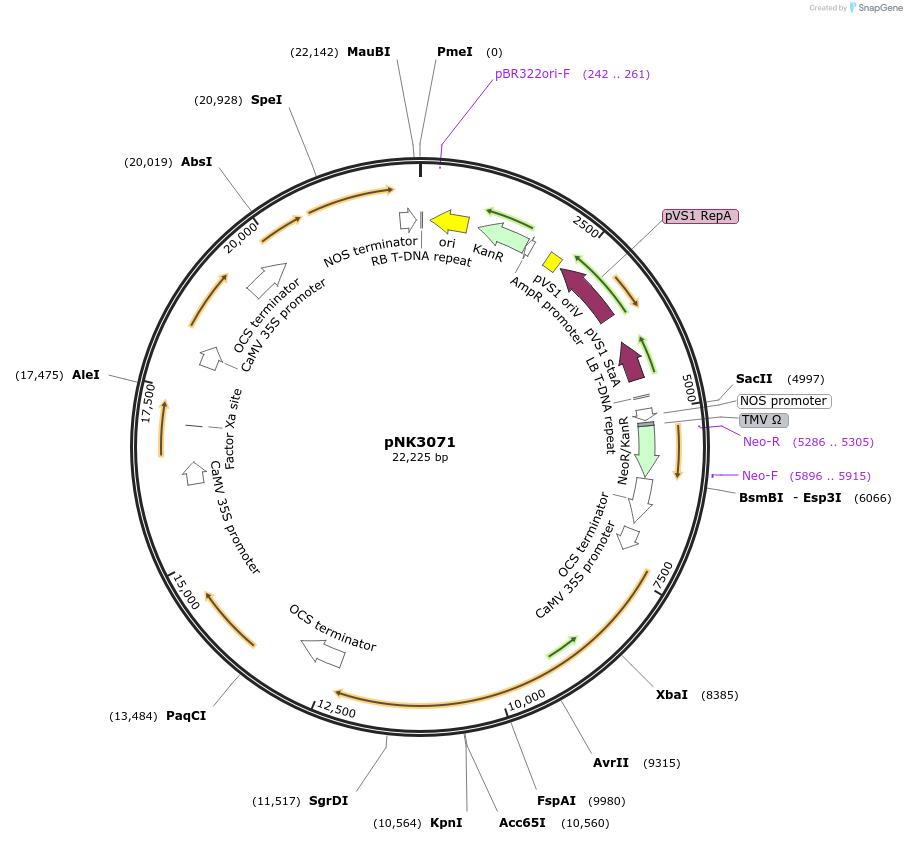
pNK3071
Plasmid#219755PurposeMoClo-compatible Level P vector for improved autonomous bioluminescence in plants encoding kanamycin resistance cassette, mcitHispS, NpgA, nnH3H_v2, nnLuz_v4 and nnCPH.DepositorInsertmcitHispS, NpgA, nnH3H_v2, nnLuz_v4 and nnCPH
ExpressionPlantAvailable SinceOct. 30, 2024AvailabilityAcademic Institutions and Nonprofits only -
LCV2 STING KO
Plasmid#217445PurposeLentiviral vector expressing Cas9 and an sgRNA targeting human STINGDepositorAvailable SinceJan. 28, 2025AvailabilityAcademic Institutions and Nonprofits only -
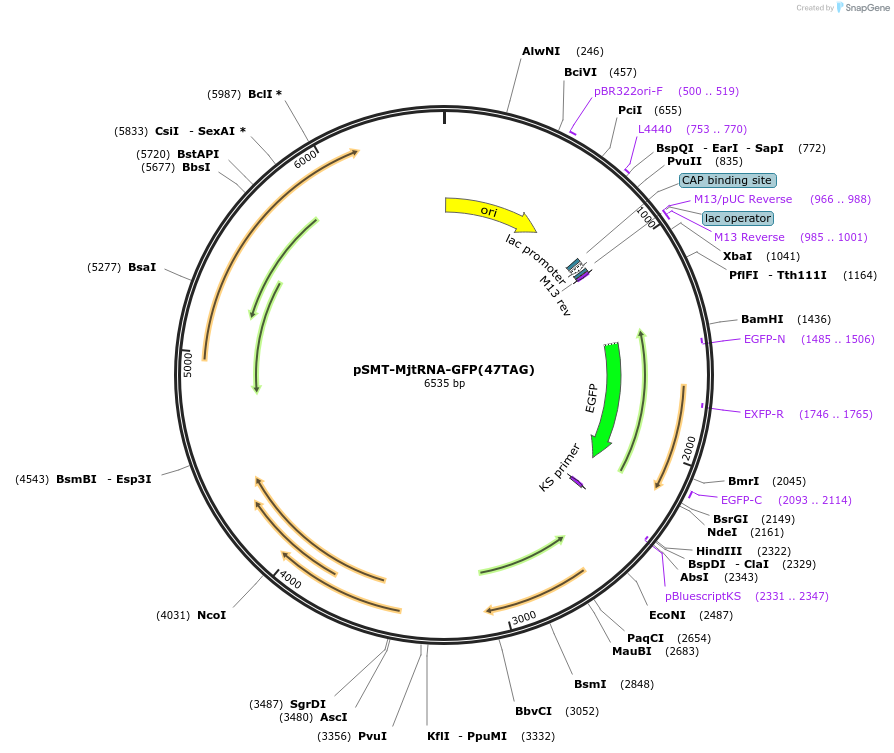
pSMT-MjtRNA-GFP(47TAG)
Plasmid#193251PurposeExpresses the M. jannaschii tRNATyr (MjtRNA) and GFP(47TAG) (aka GFP151TAG)DepositorInsertsGFP(47TAG)
MjtRNA
ExpressionBacterialMutationCodon for Tyr47 replaced with amber stop codon TA…Promoterhsp60 and metUAvailable SinceFeb. 3, 2023AvailabilityAcademic Institutions and Nonprofits only -
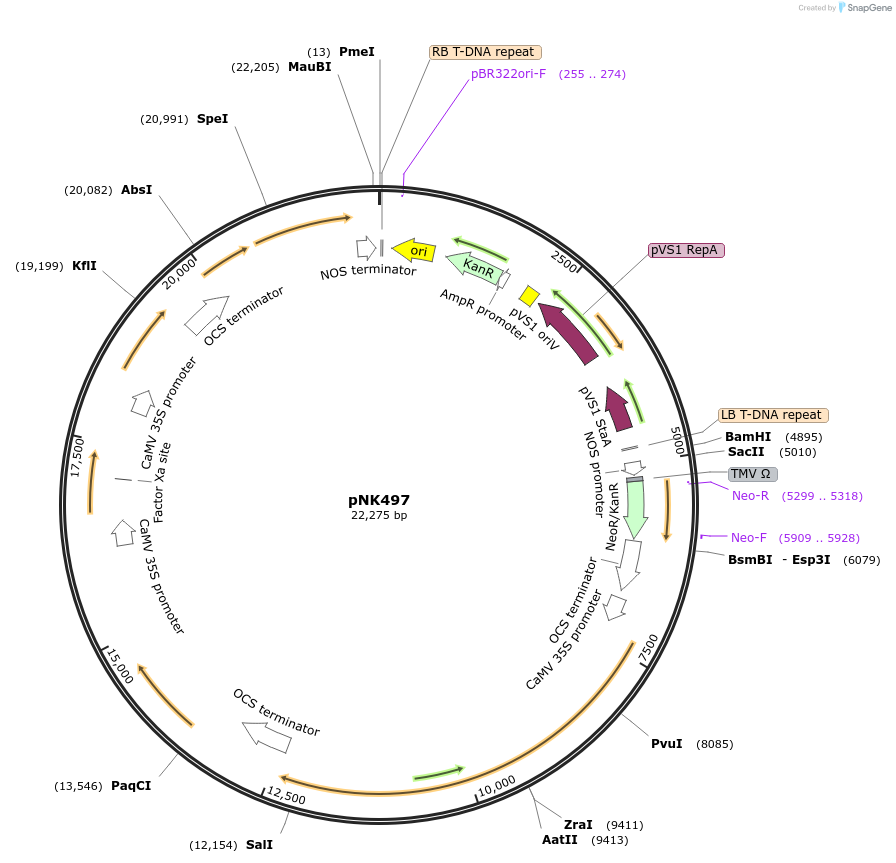
pNK497
Plasmid#219754PurposeMoClo-compatible Level P vector for improved autonomous bioluminescence in plants encoding kanamycin resistance cassette, nnHispS, NpgA, nnH3H_v2, nnLuz_v4 and nnCPH.DepositorInsertnnHispS, NpgA, nnH3H_v2, nnLuz_v4 and nnCPH
ExpressionPlantAvailable SinceJan. 21, 2025AvailabilityAcademic Institutions and Nonprofits only -
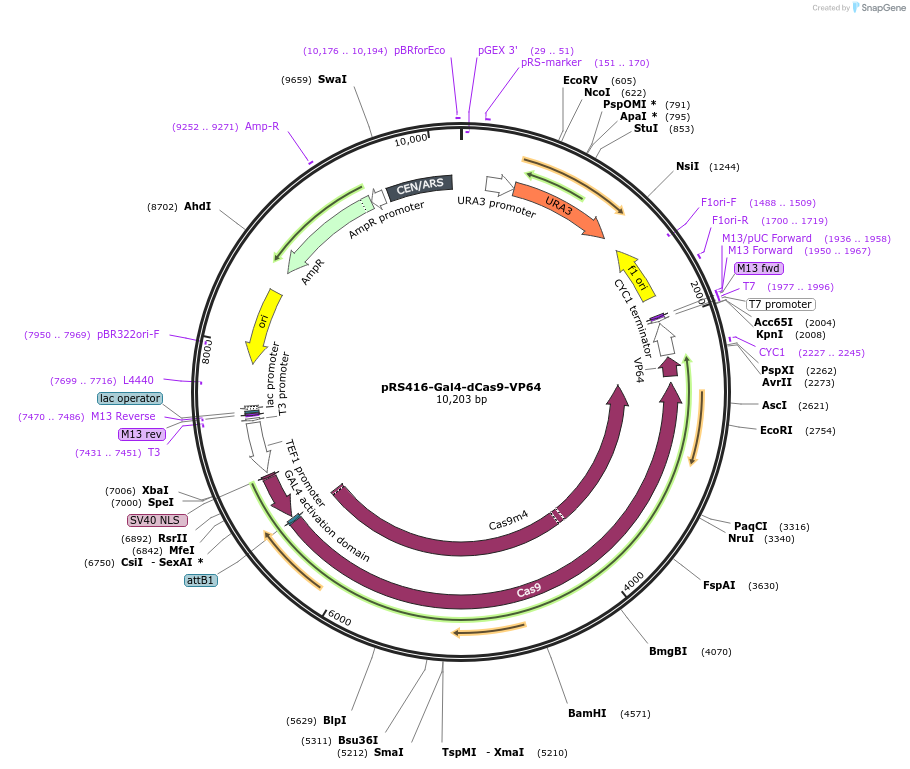
pRS416-Gal4-dCas9-VP64
Plasmid#71128PurposeThis plasmid contains a Cas9 Activator for yeast. The activator is about 1.2-1.8 X more potent than just dCas9-VP64 fusion in yeast. It is on a single copy CEN/ARS plasmid with Ura marker, pRS416.DepositorInsertGal4-dCas9-VP64
TagsNLS n-terminal, N terminal Gal4 Activator domain,…ExpressionYeastPromoterTef1Available SinceMarch 7, 2016AvailabilityAcademic Institutions and Nonprofits only -
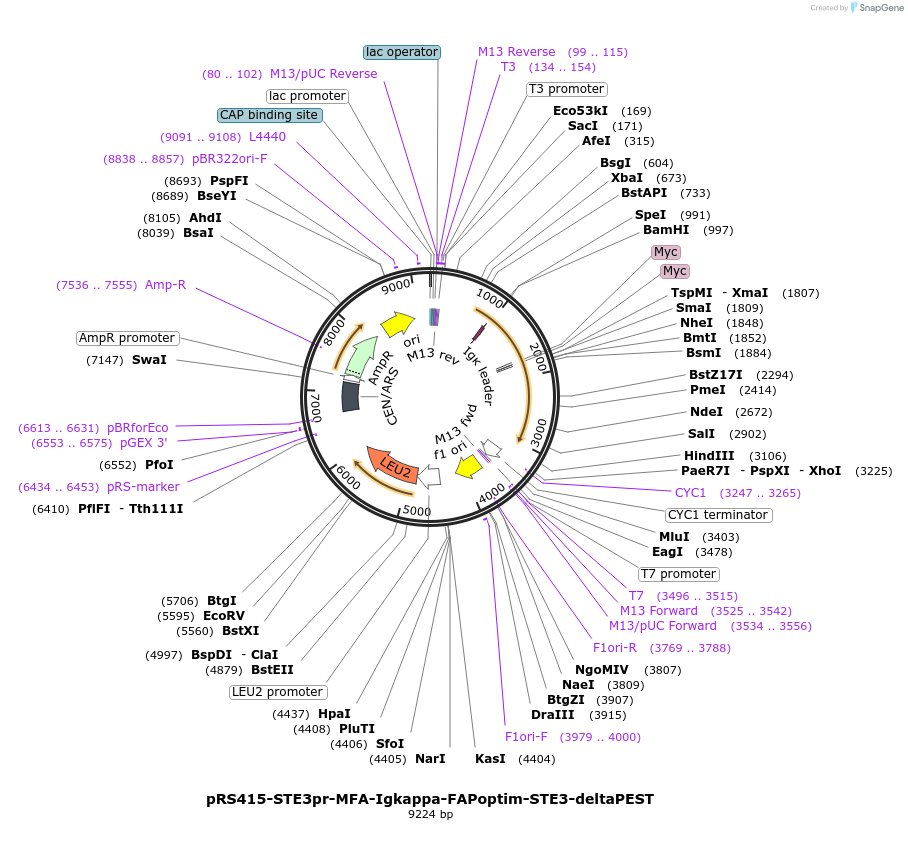
pRS415-STE3pr-MFA-Igkappa-FAPoptim-STE3-deltaPEST
Plasmid#221118PurposeFAP tagged STE3-PEST mutant (delta413-470 - N-terminally tagged, optimized) under the Ste3 promoter with 2xMYC tag and MFA1 signal sequence to help target construct to the ERDepositorInsertMFA-IgKappa-FAPoptim-STE3-deltaPEST
ExpressionYeastMutationSTE3 mutant that lacks the PEST domain in the C-t…PromoterSTE3Available SinceMarch 11, 2025AvailabilityAcademic Institutions and Nonprofits only -
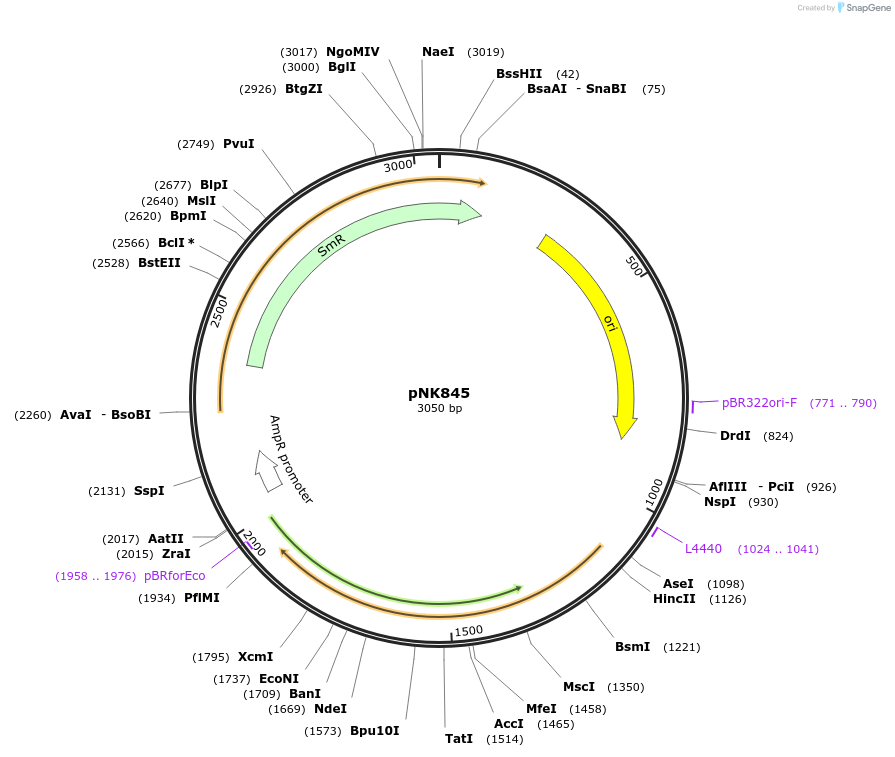
pNK845
Plasmid#219746PurposeMoClo-compatible Level 0 promoterless vector encoding mutant of Neonothopanus nambi luciferase nnLuz_v4 codon-optimised for expression in Pichia pastoris, Homo sapiensDepositorInsertmutant of fungal luciferase
UseLuciferase and Synthetic BiologyMutationI3S, N4T, F11L, I63T, T99P, T192S, A199PAvailable SinceJuly 12, 2024AvailabilityAcademic Institutions and Nonprofits only -
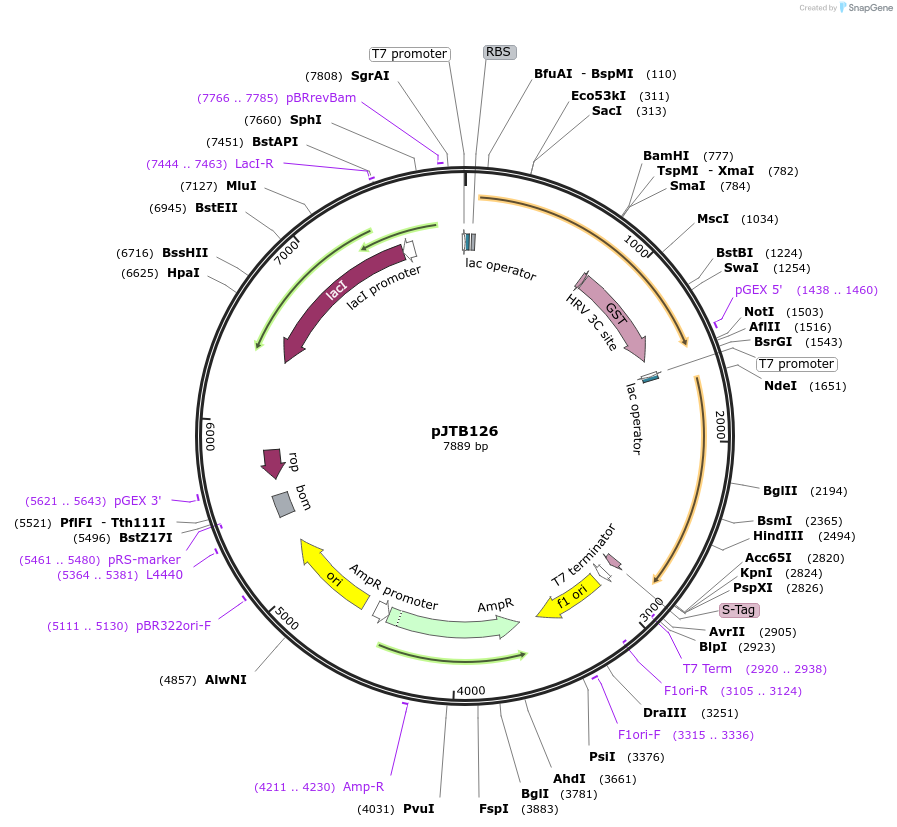
pJTB126
Plasmid#248910Purposeexpresses the yeast analog-sensitive Smk1-Q120A and Ssp2 fused to glutathione-S-transferase in bacteriaDepositorInsertsSMK1 Mitogen-activated Protein Kinase -analog-sensitive form (SMK1 Budding Yeast, Sequence is from the SK1 isolate of S. cerevisiae and will differ by 1 non-synonomous/several synonymous changes from S288C form)
Ssp2 Activator of the Smk1 MAPK (SSP2 Budding Yeast, Sequence is from the SK1 isolate of S. cerevisiae and may differ by synonymous changes from S288C form)
Tagsglutathione-S-transferaseExpressionBacterialMutationcodon 120 has been changed from Q to A (Smk1-Q120…PromoterT7Available SinceJan. 16, 2026AvailabilityAcademic Institutions and Nonprofits only