We narrowed to 3,538 results for: cgas
-
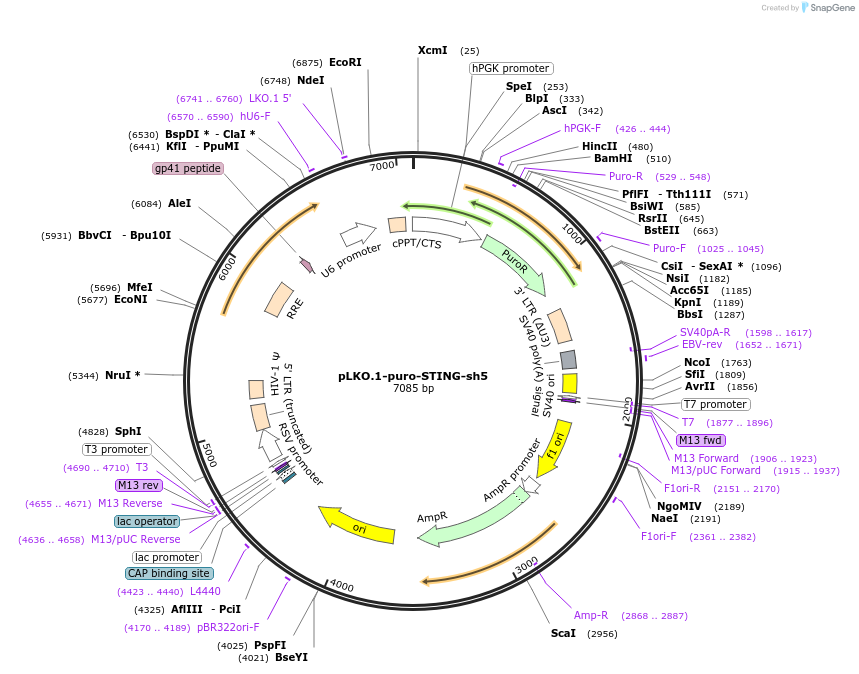
Plasmid#127647PurposeKnock-down of human STINGDepositorInsertSTING shRNA (STING1 Human)
UseLentiviralAvailable SinceJuly 9, 2019AvailabilityAcademic Institutions and Nonprofits only -
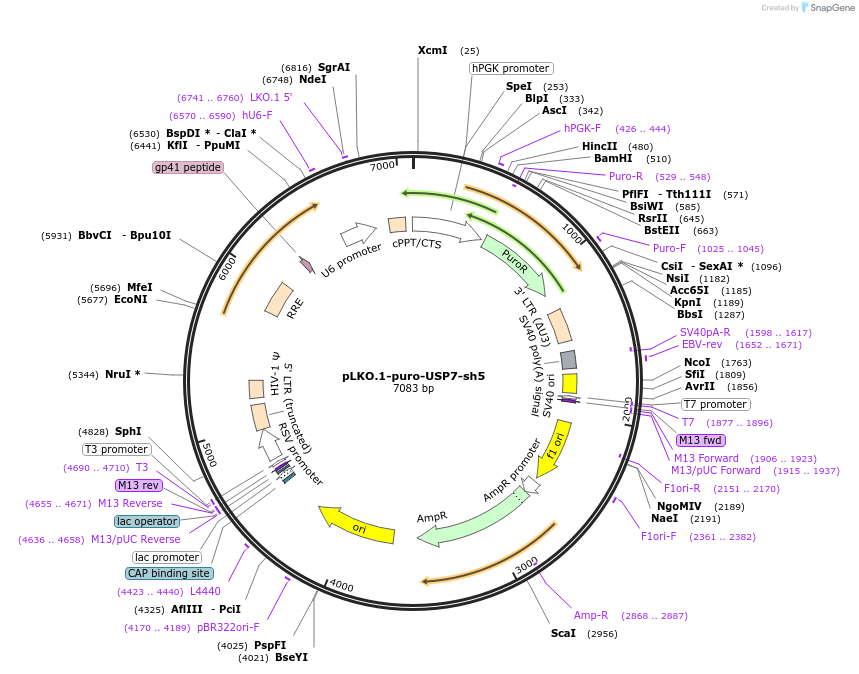
pLKO.1-puro-USP7-sh5
Plasmid#235528PurposeshRNA against human USP7DepositorInsertUSP7 (USP7 Human)
UseLentiviralAvailable SinceJuly 17, 2025AvailabilityAcademic Institutions and Nonprofits only -
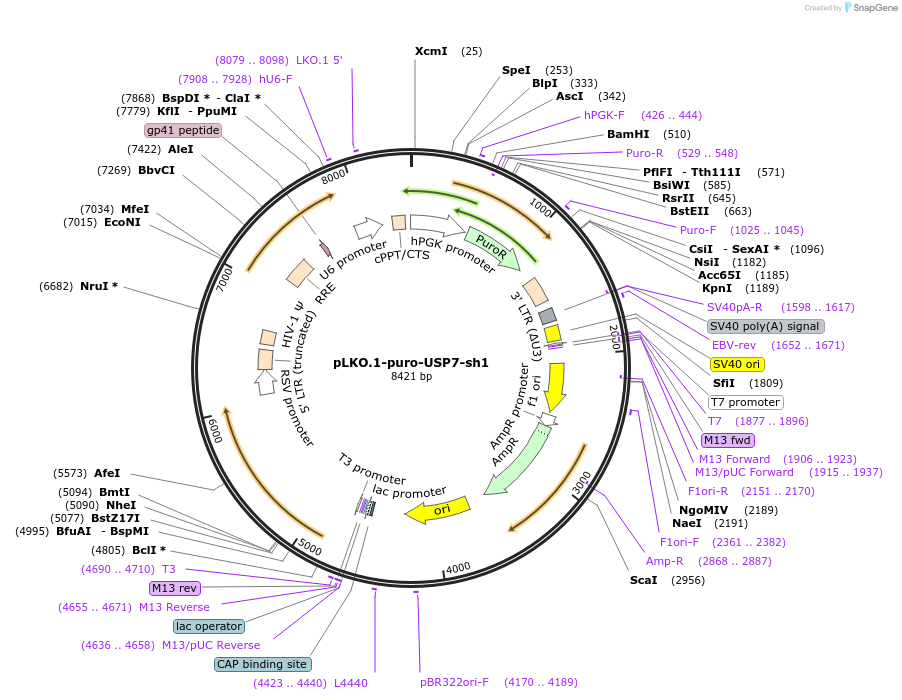
pLKO.1-puro-USP7-sh1
Plasmid#235527PurposeshRNA against human USP7DepositorInsertUSP7 (USP7 Human)
UseLentiviralAvailable SinceJuly 17, 2025AvailabilityAcademic Institutions and Nonprofits only -
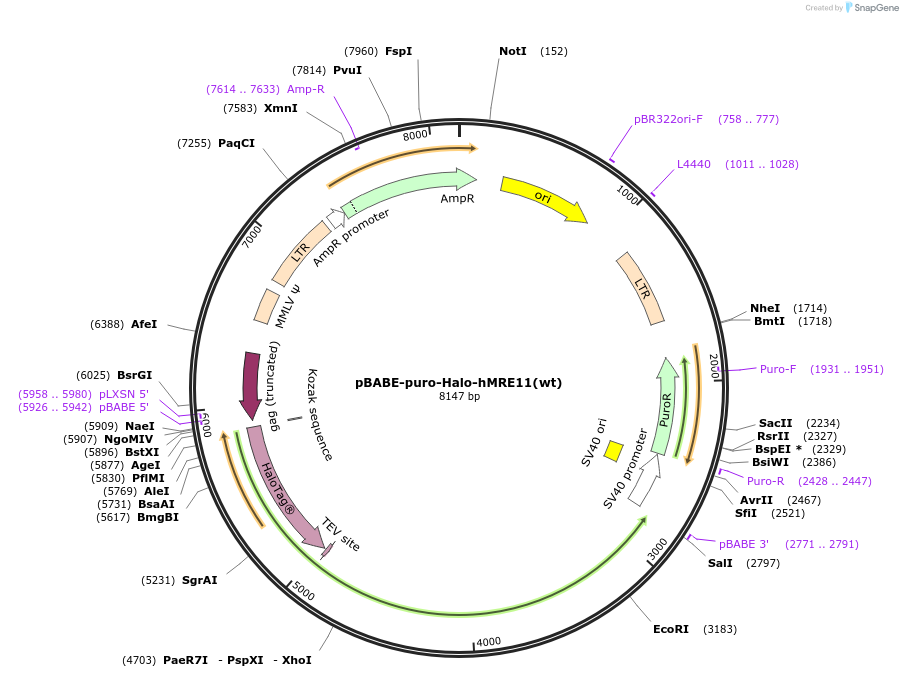
pBABE-puro-Halo-hMRE11(wt)
Plasmid#208393PurposeTo generate stable cell line expressing Halo-human MRE11(WT) by Retroviral infection.DepositorAvailable SinceJan. 11, 2024AvailabilityAcademic Institutions and Nonprofits only -
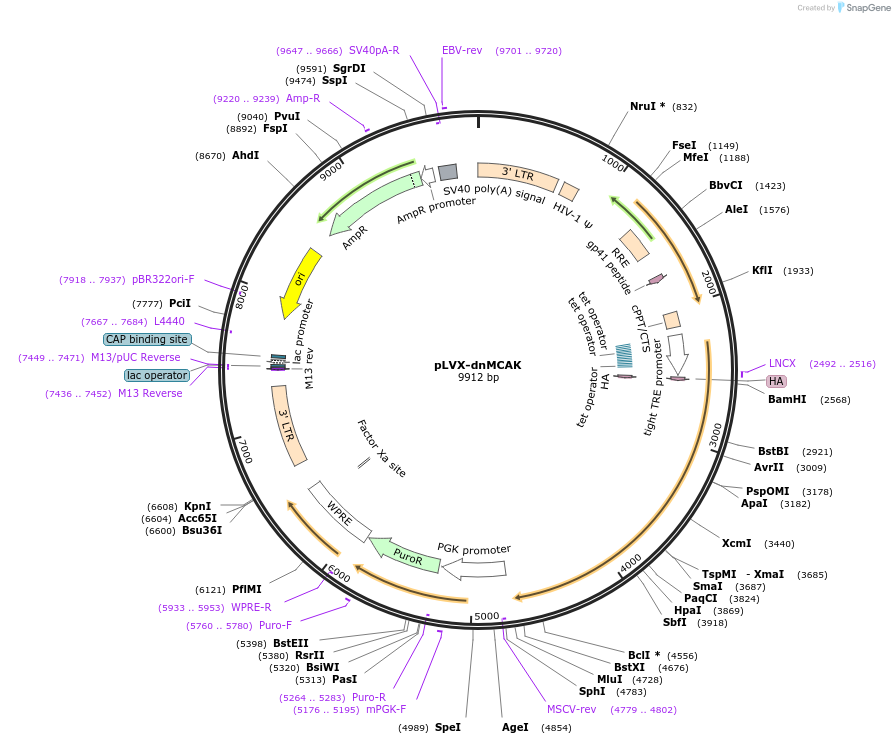
pLVX-dnMCAK
Plasmid#205995Purposeinducible expression of HA-dnMCAKDepositorInsertdnMCAK
TagsHAExpressionMammalianPromoterpTightAvailable SinceNov. 20, 2023AvailabilityAcademic Institutions and Nonprofits only -
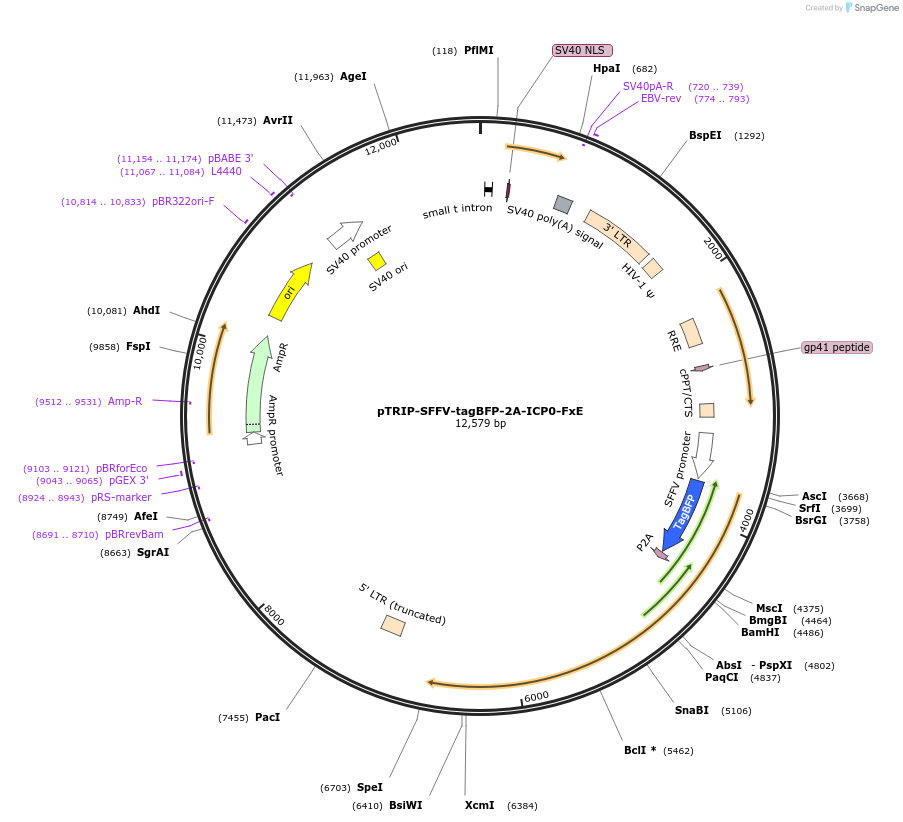
pTRIP-SFFV-tagBFP-2A-ICP0-FxE
Plasmid#235539PurposeTo express tagBFP-2A-ICP0-FXE (ICP0 ∆106-149)DepositorInsertICP0-FxE
UseLentiviralMutationDeletion of the RING domain, ∆106-149.Available SinceJuly 17, 2025AvailabilityAcademic Institutions and Nonprofits only -
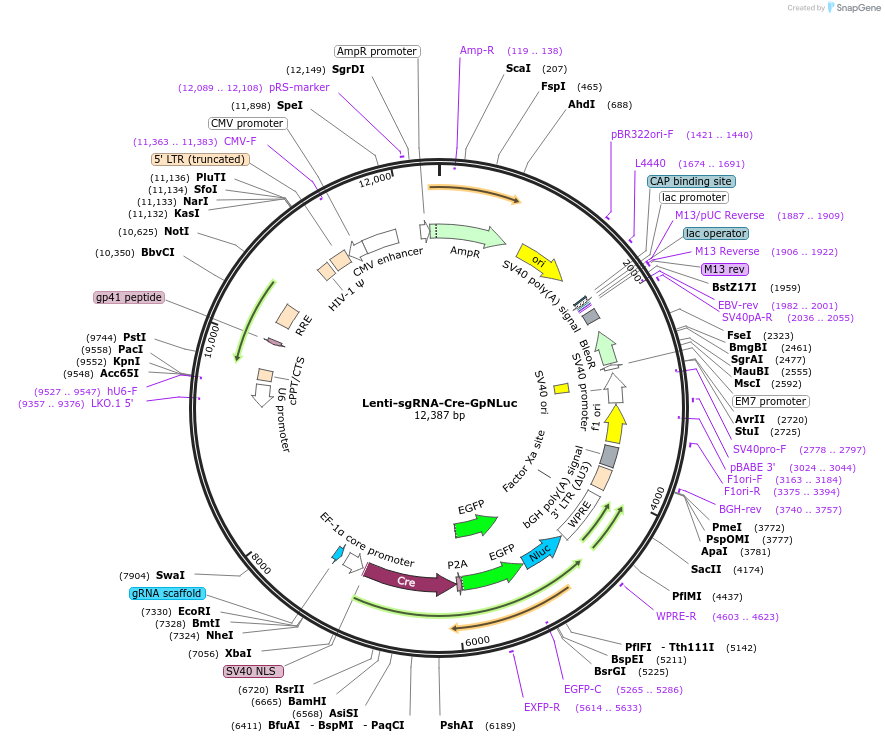
Lenti-sgRNA-Cre-GpNLuc
Plasmid#208382PurposeDerived from LentiCRISPRV2 by replacing Cas9 with Cre recombinase and puromycin resistance with GpNLuc. Contains a stuffer sequence for cloning of sgRNA of interest, as per LentiCRISPRV2.DepositorTypeEmpty backboneUseLentiviral and LuciferaseExpressionMammalianMutationWTAvailable SinceJan. 11, 2024AvailabilityAcademic Institutions and Nonprofits only -
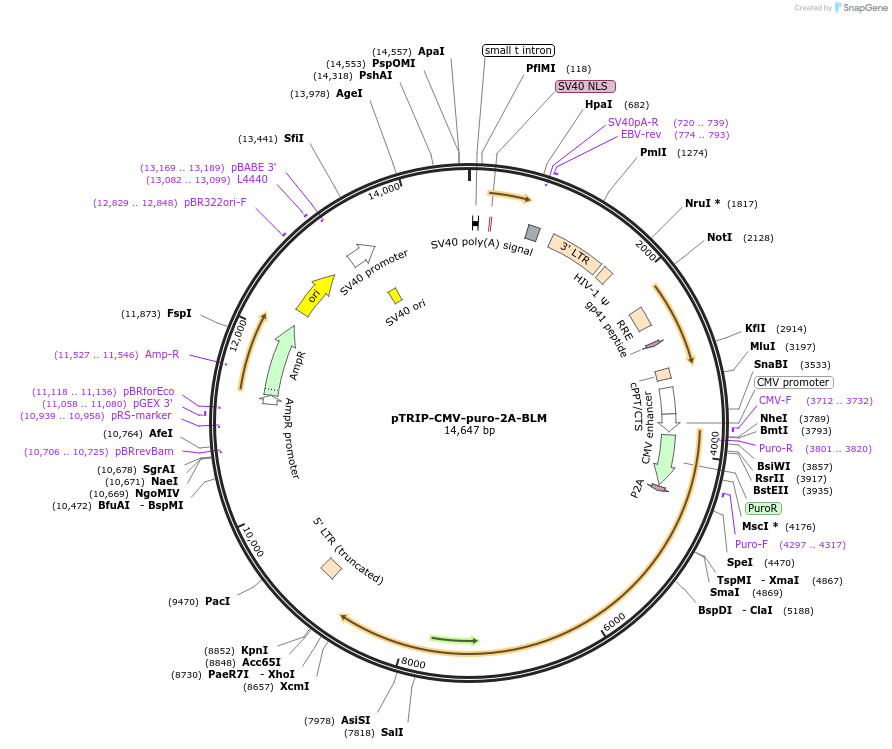
pTRIP-CMV-puro-2A-BLM
Plasmid#127641PurposeOver-expression of human BLMDepositorInsertBLM (BLM Human)
UseLentiviralAvailable SinceJuly 11, 2019AvailabilityAcademic Institutions and Nonprofits only -
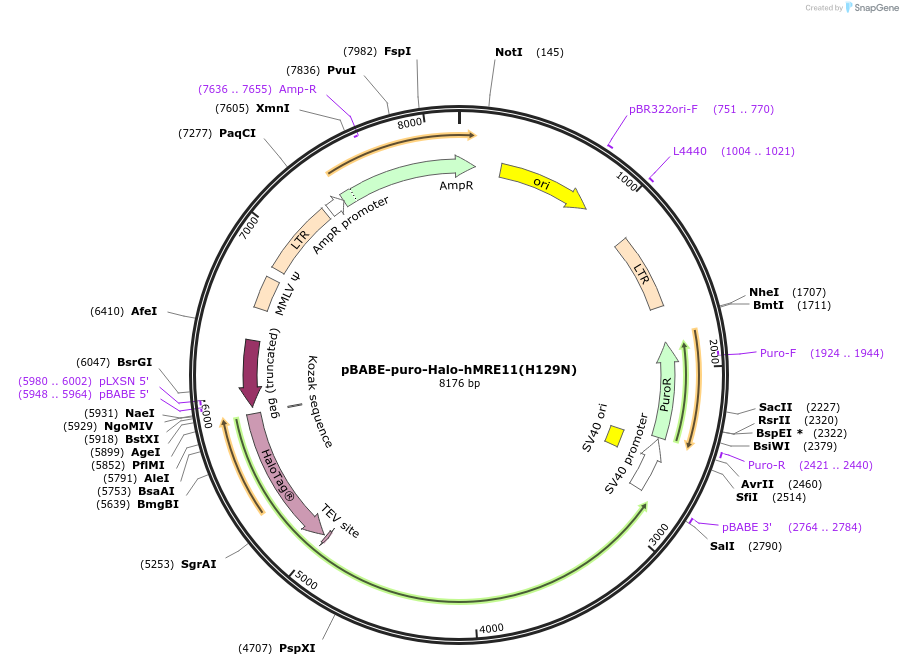
pBABE-puro-Halo-hMRE11(H129N)
Plasmid#208394PurposeTo generate stable cell line expressing Halo-human MRE11(H129N) by Retroviral infection.DepositorAvailable SinceJan. 11, 2024AvailabilityAcademic Institutions and Nonprofits only -
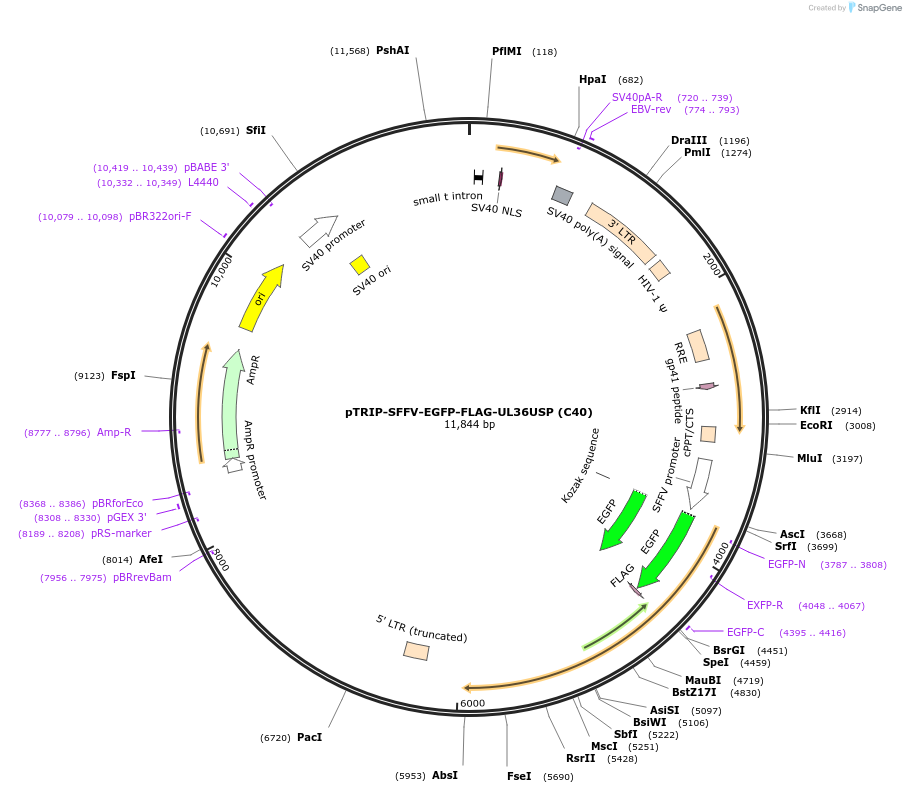
pTRIP-SFFV-EGFP-FLAG-UL36USP (C40)
Plasmid#235548PurposeTo express GFP-FLAG-UL36USP (C40) - region 25-520 of UL36DepositorInsertUL36USP (C40)
UseLentiviralTagsEGFP-FLAGMutationN-terminal part of UL36 starting at the second AT…Available SinceJuly 17, 2025AvailabilityAcademic Institutions and Nonprofits only -
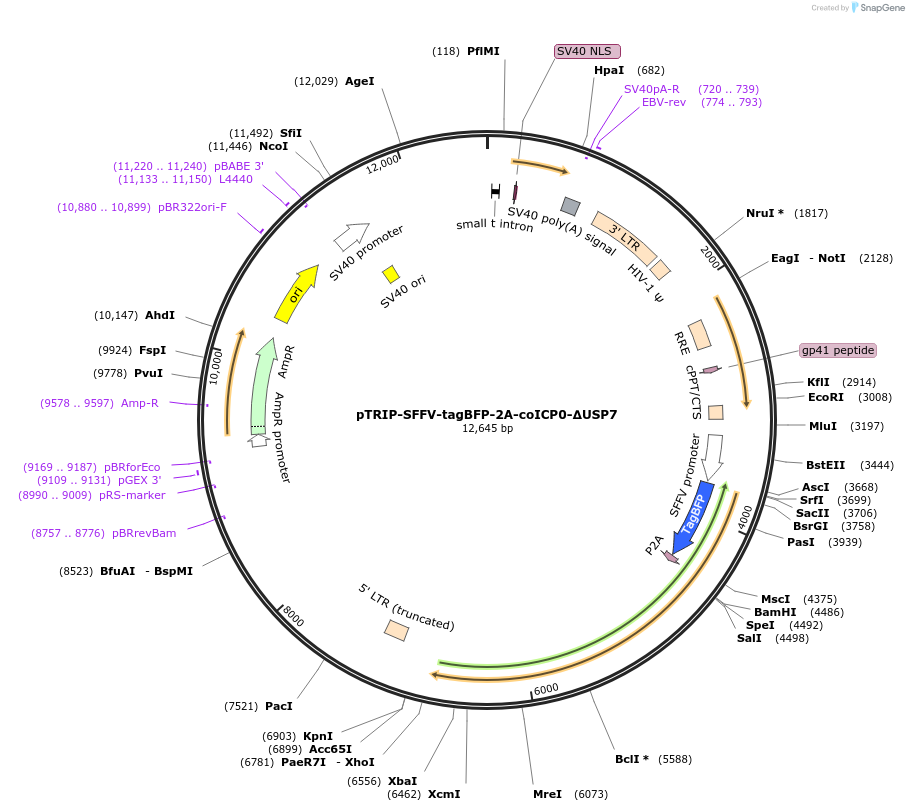
pTRIP-SFFV-tagBFP-2A-coICP0-∆USP7
Plasmid#235540PurposeTo express tagBFP-2A-coICP0-∆USP7 (ICP0 ∆618-633, co = codon-optimized)DepositorInsertICP0-∆USP7
UseLentiviralMutationDeletion of region ∆618-633.Available SinceJuly 17, 2025AvailabilityAcademic Institutions and Nonprofits only -
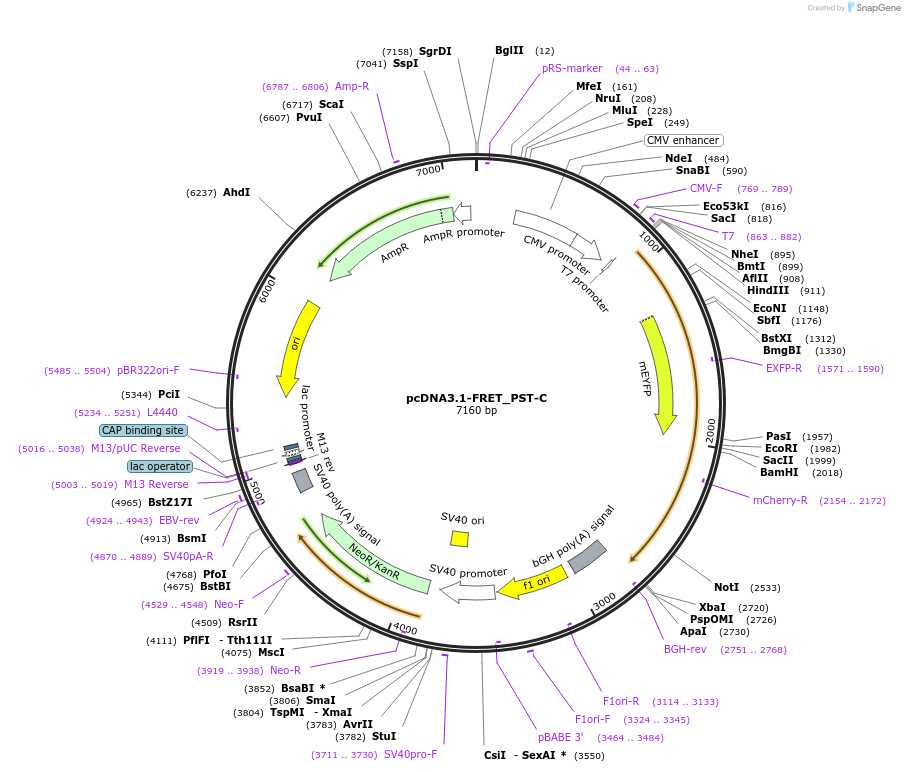
pcDNA3.1-FRET_PST-C
Plasmid#228570PurposeExpressed a FRET sensor for determining proteolytic cleavage of the C-terminal side of PancreastatinDepositorInsertChromogranin-A (CHGA Human)
TagsmCitrine and mScarlet-IExpressionMammalianMutationFRET sensor; see explanatory notePromotercmvAvailable SinceJuly 1, 2025AvailabilityAcademic Institutions and Nonprofits only -
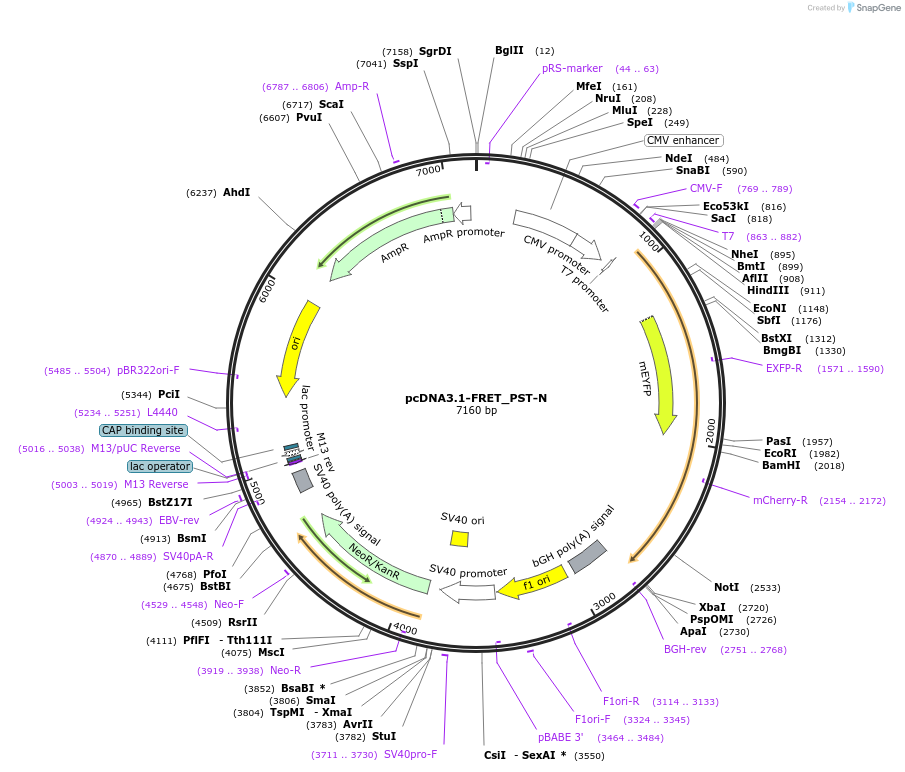
pcDNA3.1-FRET_PST-N
Plasmid#228571PurposeExpressed a FRET sensor for determining proteolytic cleavage of the N-terminal side of PancreastatinDepositorInsertChromogranin-A (CHGA Human)
TagsmCitrine and mScarlet-IExpressionMammalianMutationFRET sensor; see explanatory notePromotercmvAvailable SinceJuly 1, 2025AvailabilityAcademic Institutions and Nonprofits only -
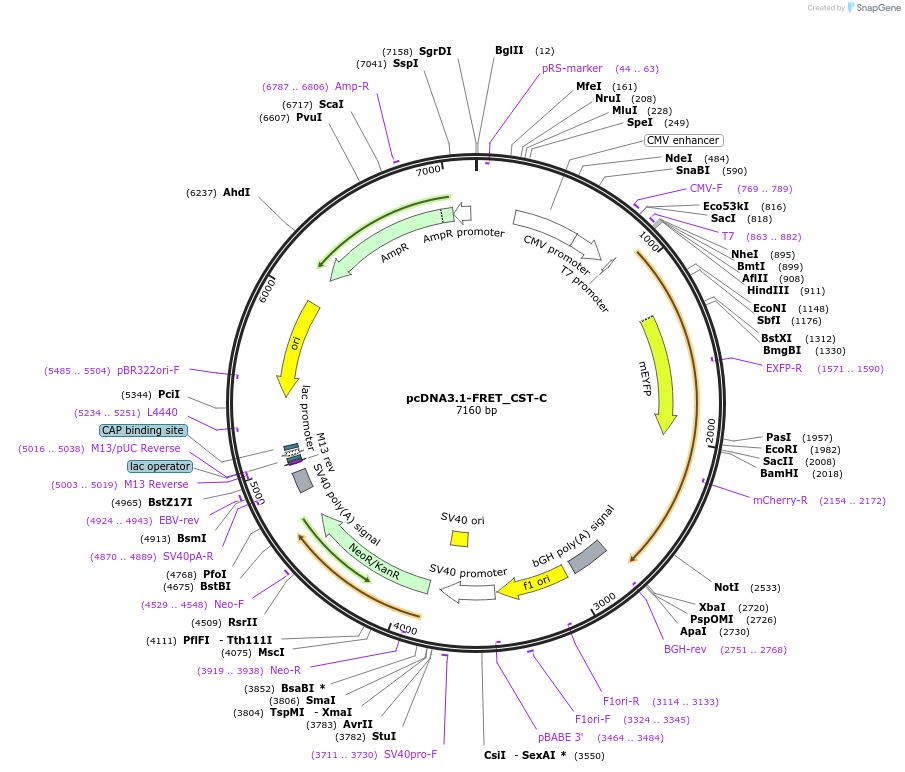
pcDNA3.1-FRET_CST-C
Plasmid#228572PurposeExpressed a FRET sensor for determining proteolytic cleavage of the C-terminal side of CatestatinDepositorInsertChromogranin-A (CHGA Human)
TagsmCitrine and mScarlet-IExpressionMammalianMutationFRET sensor; see explanatory notePromotercmvAvailable SinceJuly 1, 2025AvailabilityAcademic Institutions and Nonprofits only -
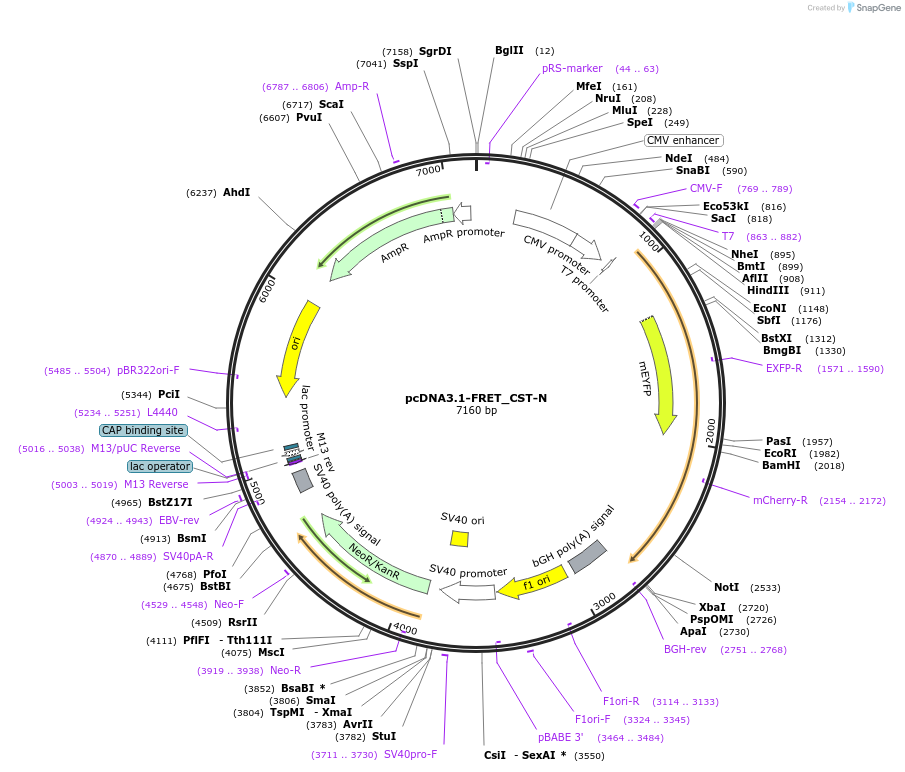
pcDNA3.1-FRET_CST-N
Plasmid#228573PurposeExpressed a FRET sensor for determining proteolytic cleavage of the N-terminal side of CatestatinDepositorInsertChromogranin-A (CHGA Human)
TagsmCitrine and mScarlet-IExpressionMammalianMutationFRET sensor; see explanatory notePromotercmvAvailable SinceJuly 1, 2025AvailabilityAcademic Institutions and Nonprofits only -
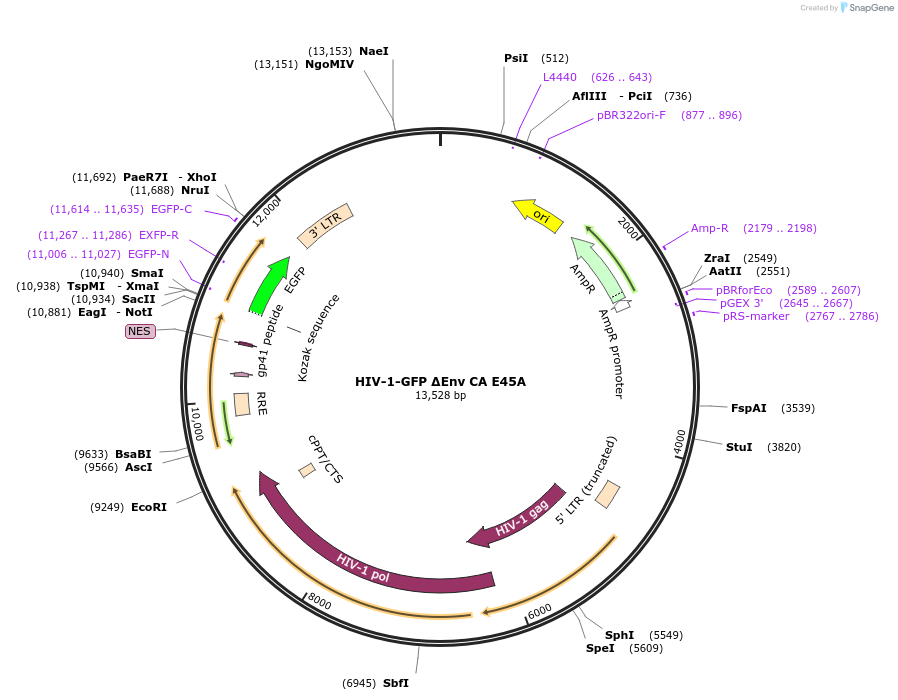
HIV-1-GFP ΔEnv CA E45A
Plasmid#217438PurposeHIV-1 reporter vector that is env-deficient and expresses eGFP in place of nef; contains 5' and 3' LTRs, gag, pol, tat, and rev, but does not express vif, vpr, or vpu (with E45A mutation in CA).DepositorInsertgag/pol
UseLentiviralMutationCA E45AAvailable SinceJan. 28, 2025AvailabilityAcademic Institutions and Nonprofits only -
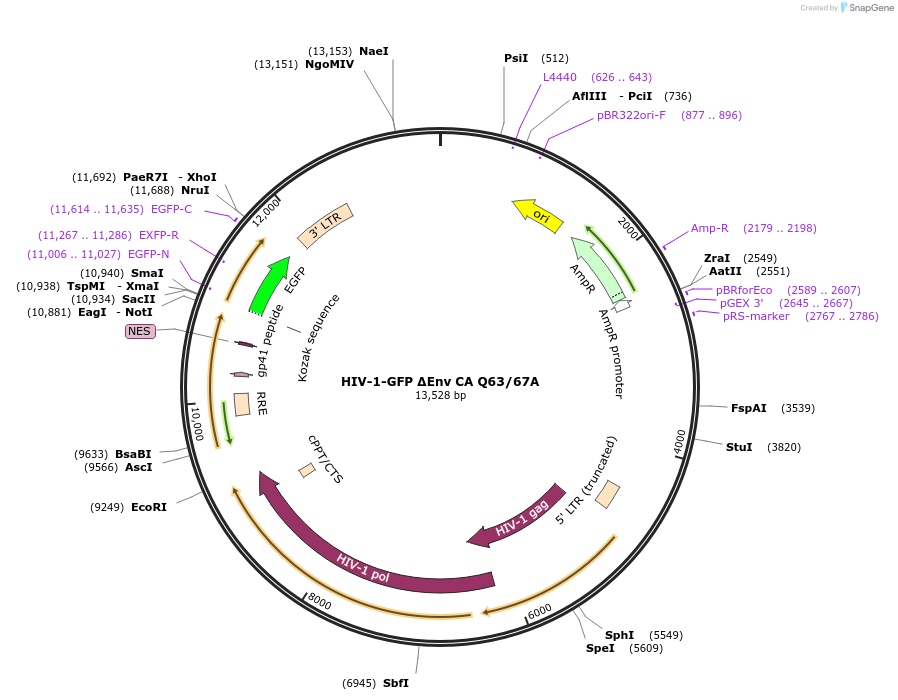
HIV-1-GFP ΔEnv CA Q63/67A
Plasmid#217439PurposeHIV-1 reporter vector that is env-deficient and expresses eGFP in place of nef; contains 5' and 3' LTRs, gag, pol, tat, and rev, but does not express vif, vpr, or vpu (with Q63A,Q67A mutations in CA).DepositorInsertgag/pol
UseLentiviralMutationCA Q63A,Q67AAvailable SinceJan. 28, 2025AvailabilityAcademic Institutions and Nonprofits only -
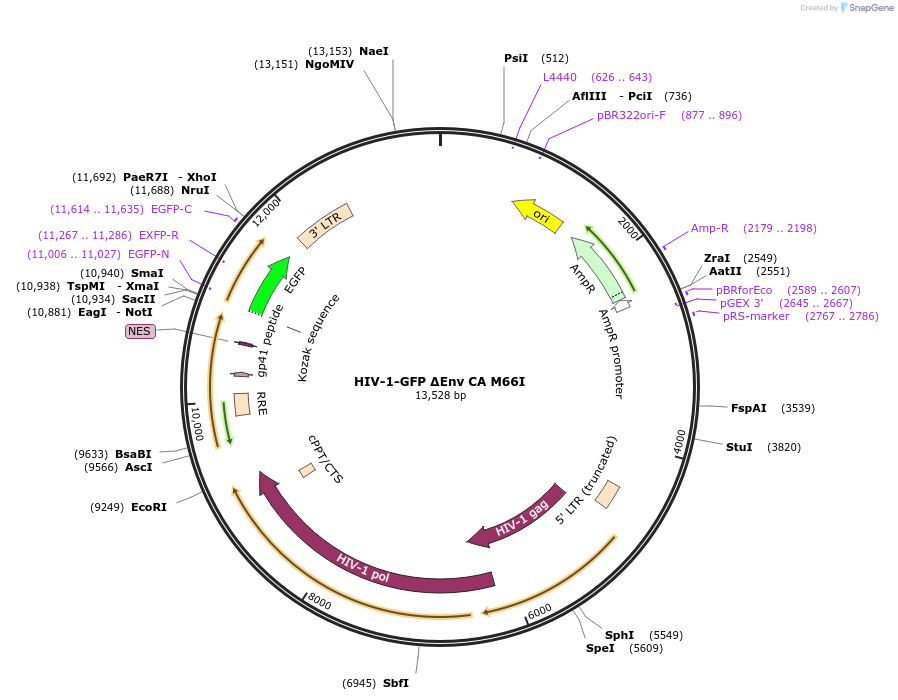
HIV-1-GFP ΔEnv CA M66I
Plasmid#217440PurposeHIV-1 reporter vector that is env-deficient and expresses eGFP in place of nef; contains 5' and 3' LTRs, gag, pol, tat, and rev, but does not express vif, vpr, or vpu (with M66I mutation in CA).DepositorInsertgag/pol
UseLentiviralMutationCA M66IAvailable SinceJan. 28, 2025AvailabilityAcademic Institutions and Nonprofits only -
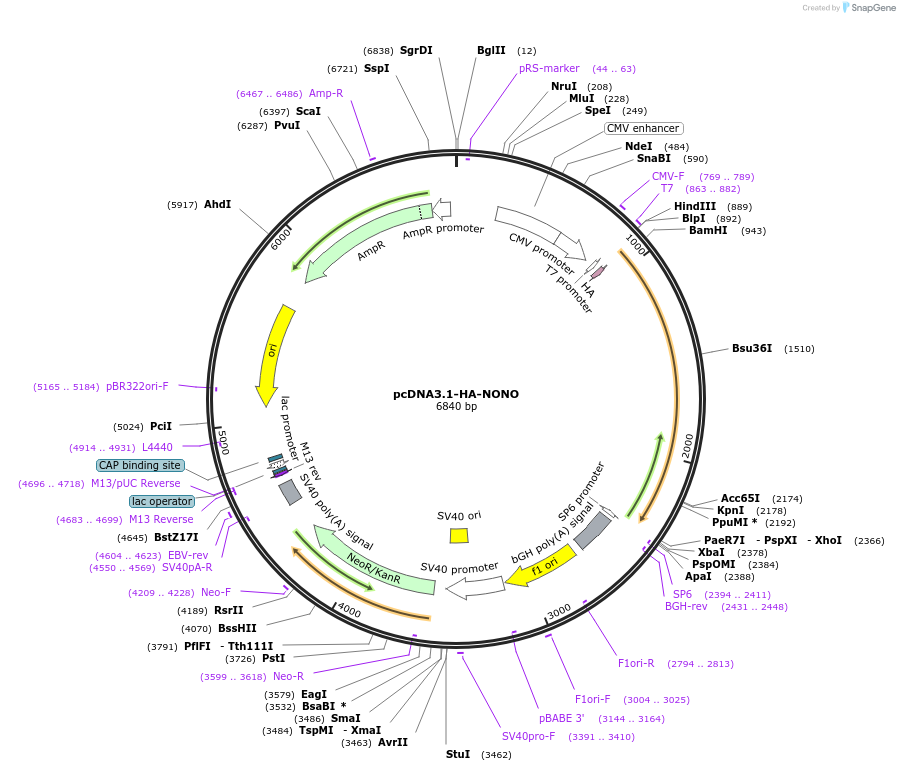
pcDNA3.1-HA-NONO
Plasmid#127655PurposeOver-express HA-tagged human NONODepositorAvailable SinceJuly 9, 2019AvailabilityAcademic Institutions and Nonprofits only -
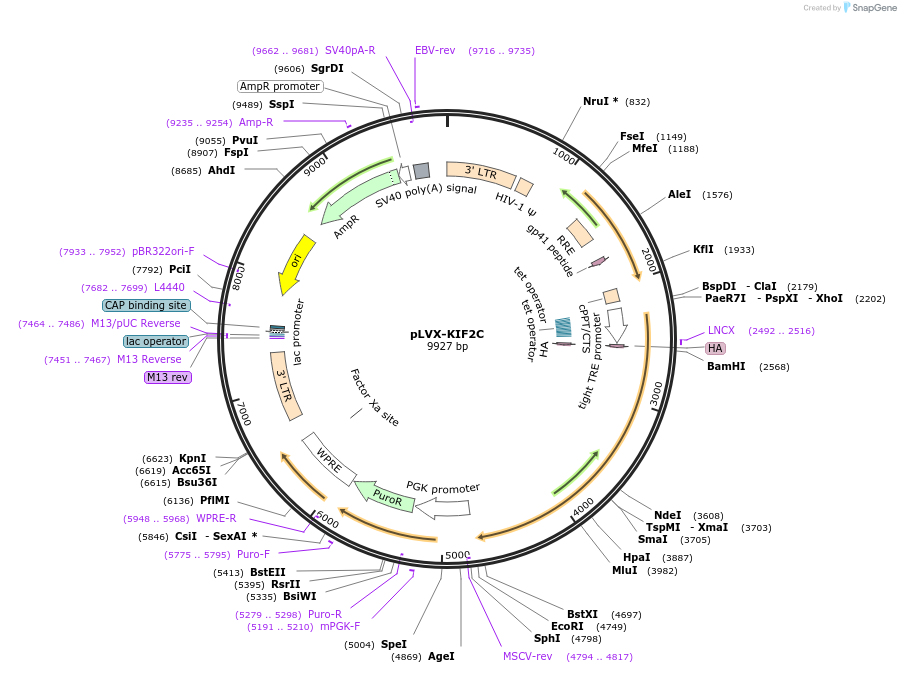
pLVX-KIF2C
Plasmid#205996Purposeinducible expression of HA-KIF2CDepositorAvailable SinceNov. 20, 2023AvailabilityAcademic Institutions and Nonprofits only