We narrowed to 29,147 results for: Tat
-
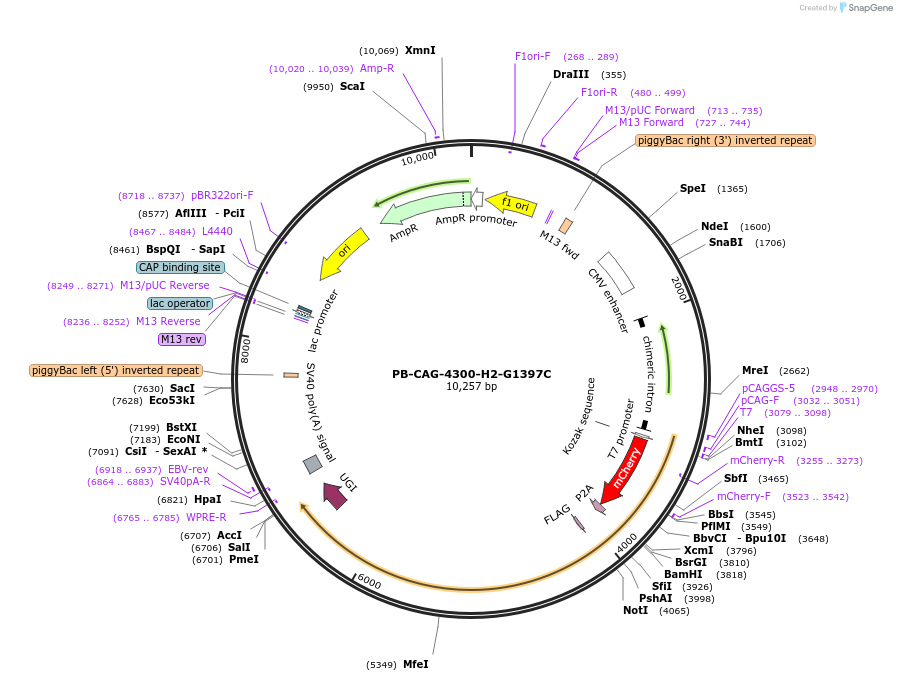
Plasmid#210395PurposeTo correct m.A4300G mutation on mtDNADepositorInsert4300-H2-G1397C
ExpressionMammalianAvailable SinceDec. 12, 2023AvailabilityAcademic Institutions and Nonprofits only -
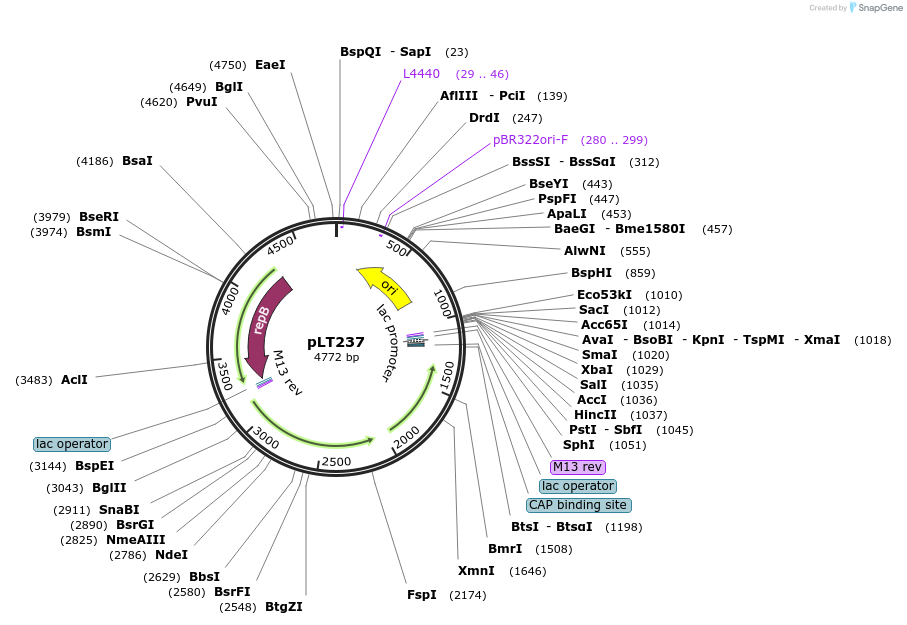
pLT237
Plasmid#174300PurposeReplicating plasmid with homology region for introducing PolCC669Y mutation. Confers thiamphenicol resistance.DepositorInsertPolC homology arm
ExpressionBacterialMutationchanged Cysteine 669 to TyrosineAvailable SinceMay 15, 2023AvailabilityAcademic Institutions and Nonprofits only -
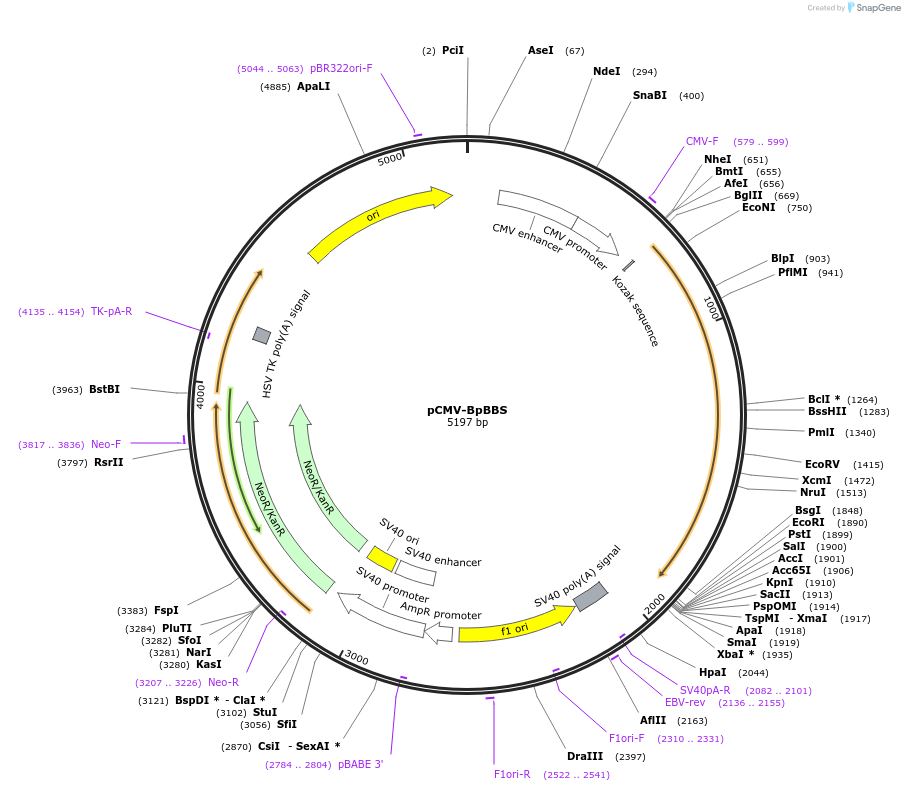
pCMV-BpBBS
Plasmid#199468PurposeExpression of the biliverdin-binding serpin of Boana punctata treefrog (BpBBS) in mammalian cells.DepositorInsertBpBBS
ExpressionMammalianAvailable SinceMay 11, 2023AvailabilityAcademic Institutions and Nonprofits only -
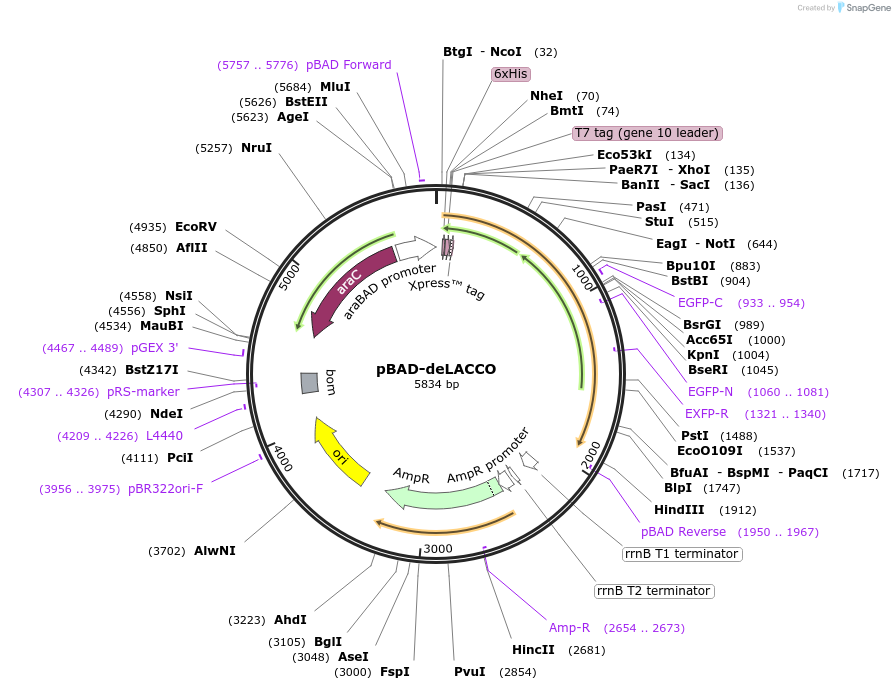
pBAD-deLACCO
Plasmid#167945PurposeFluorescent biosensor control for extracellular L-lactateDepositorInsertdeLACCO
ExpressionBacterialPromoteraraBADAvailable SinceApril 22, 2021AvailabilityAcademic Institutions and Nonprofits only -
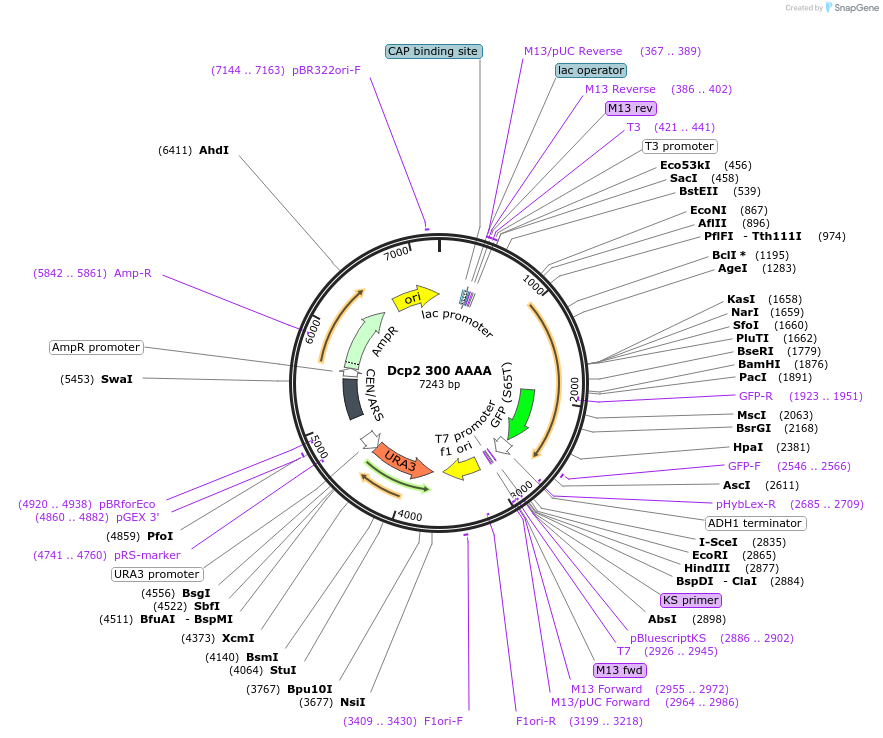
Dcp2 300 AAAA
Plasmid#163579Purposeexpress Dcp2 300 AAAA at endogenous level in S.cerevisiaeDepositorInsertDcp2 (DCP2 Budding Yeast)
TagsGFPExpressionYeastMutationTruncation of C-terminal domain (301-970); mutati…PromoterDCP2 promoterAvailable SinceApril 21, 2021AvailabilityAcademic Institutions and Nonprofits only -
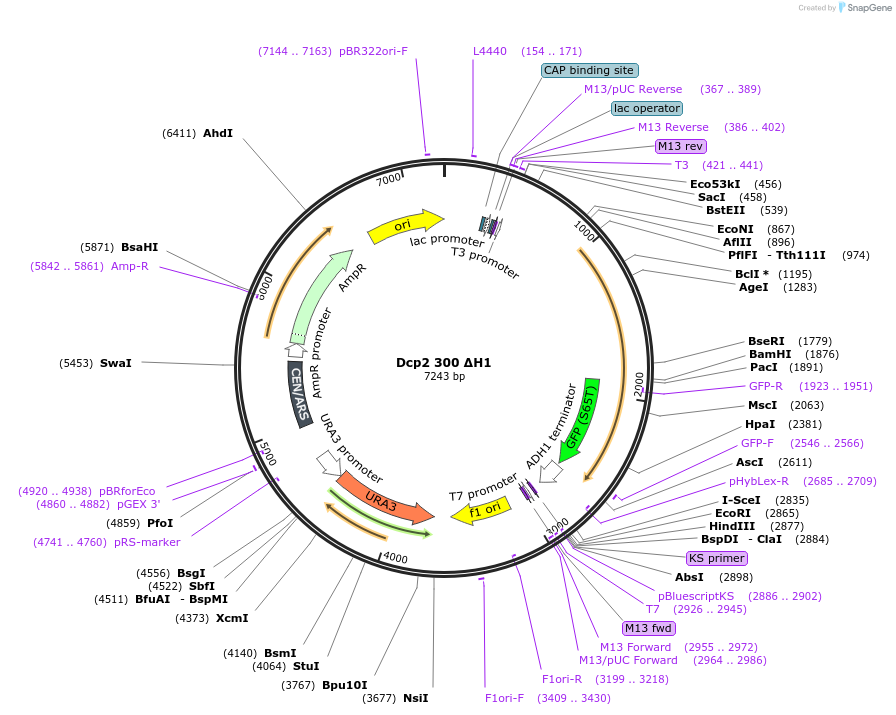
Dcp2 300 ΔH1
Plasmid#163578Purposeexpress Dcp2 300 ΔH1 at endogenous level in S.cerevisiaeDepositorInsertDcp2 (DCP2 Budding Yeast)
TagsGFPExpressionYeastMutationTruncation of C-terminal domain (301-970); Lysine…PromoterDCP2 promoterAvailable SinceApril 16, 2021AvailabilityAcademic Institutions and Nonprofits only -
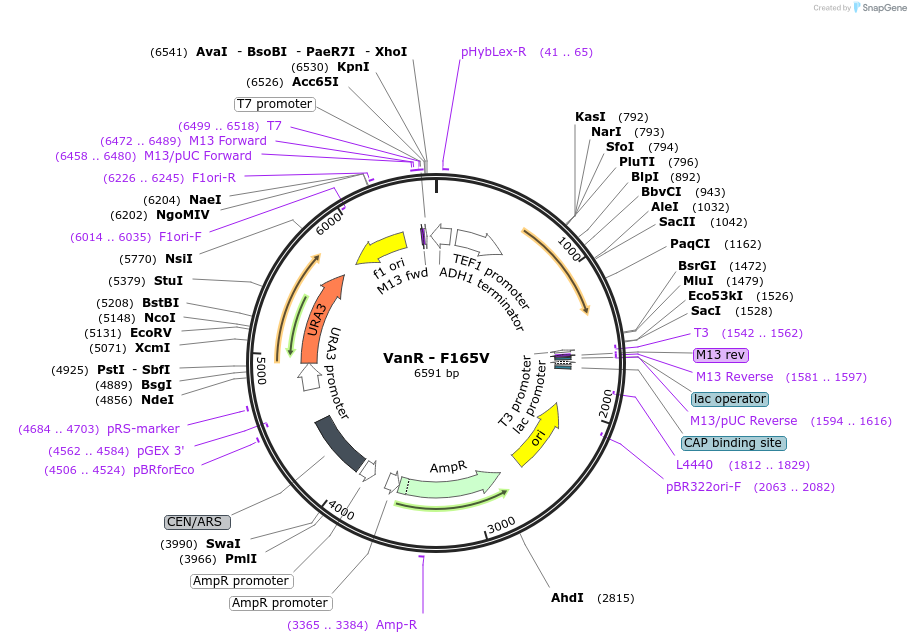
VanR - F165V
Plasmid#158965PurposeWild type VanR where the residue in position 165 was mutated from Phenylalanine to Valine. This mutation abolished the response to vanillic acid. The gene is under the control of the TEF1 promoter.DepositorInsertVanR - F165V
ExpressionYeastMutationWild type VanR from Caulobacter crescentus with m…PromoterTEF1Available SinceJan. 8, 2021AvailabilityAcademic Institutions and Nonprofits only -
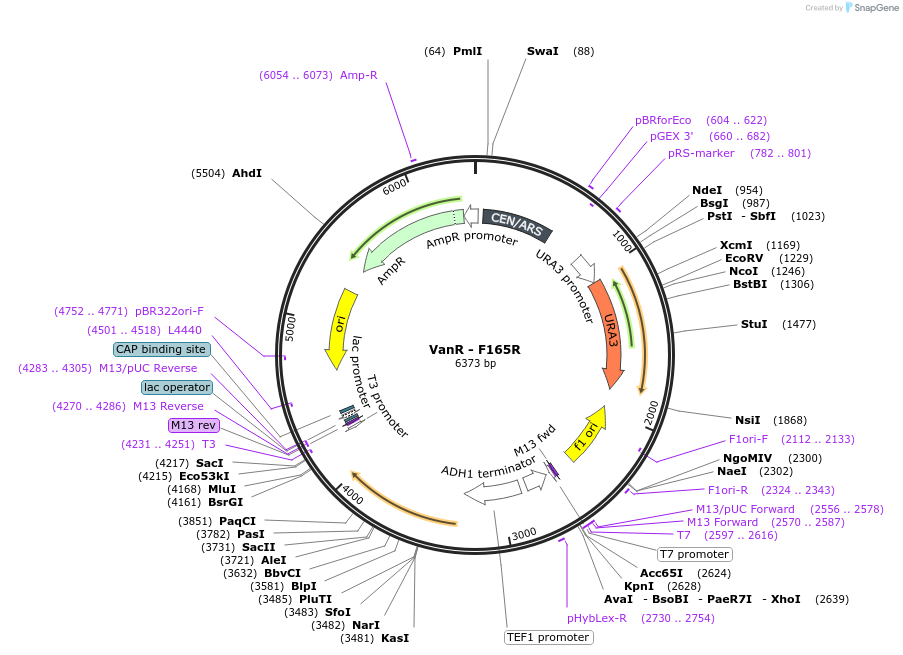
VanR - F165R
Plasmid#158966PurposeWild type VanR where the residue in position 165 was mutated from Phenylalanine to Arginine. This mutation abolished the response to vanillic acid. The gene is under the control of the TEF1 promoter.DepositorInsertVanR - F165R
ExpressionYeastMutationWild type VanR from Caulobacter crescentus with m…PromoterTEF1Available SinceJan. 8, 2021AvailabilityAcademic Institutions and Nonprofits only -
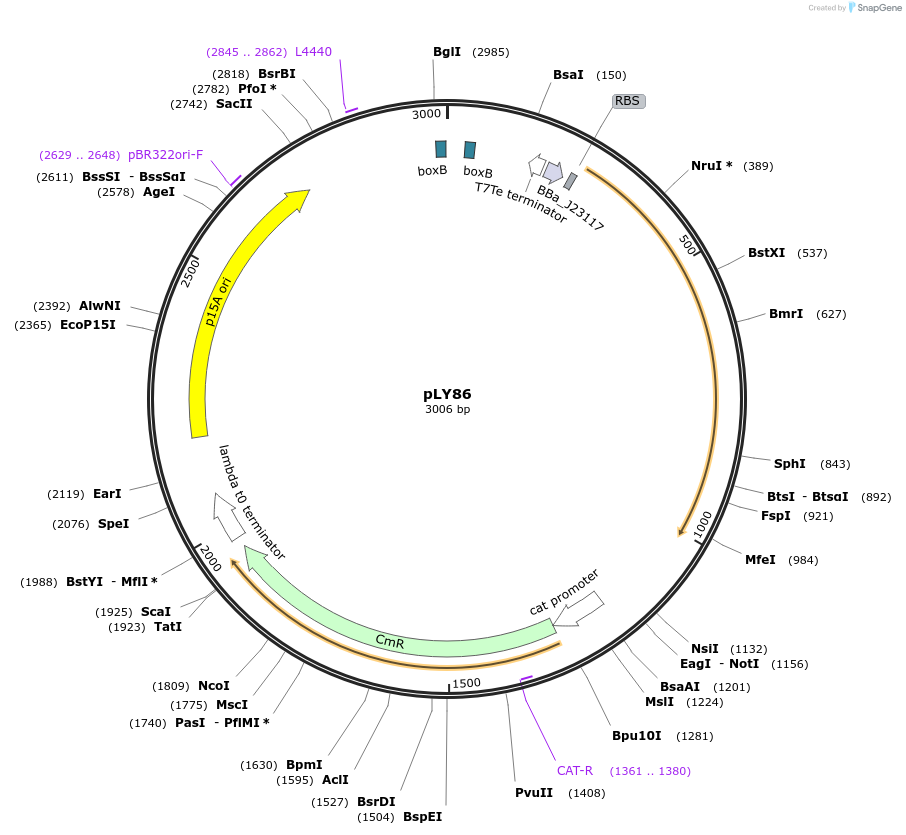
pLY86
Plasmid#130949PurposeEngineered sgRNA-LEA2-WTmis2 generator circuit including promoter Plux2, 2 BoxB aptamers and 2 mis-matches in sgRNA scaffoldDepositorInsertsgRNA-LEA2-WTmis2
UseSynthetic BiologyExpressionBacterialPromoterPlux2Available SinceOct. 8, 2019AvailabilityAcademic Institutions and Nonprofits only -
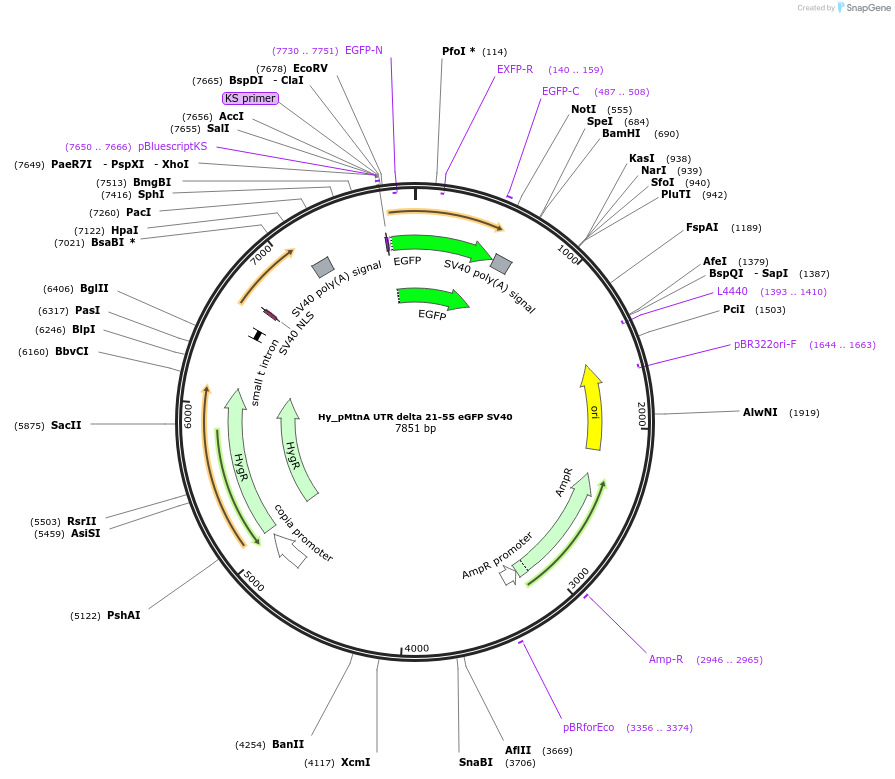
Hy_pMtnA UTR delta 21-55 eGFP SV40
Plasmid#132653PurposeExpresses an eGFP mRNA from the Drosophila Metallothionein A promoter that has 20 nt of the MtnA 5' UTRDepositorInserteGFP
ExpressionInsectPromoterMtnAAvailable SinceOct. 2, 2019AvailabilityAcademic Institutions and Nonprofits only -
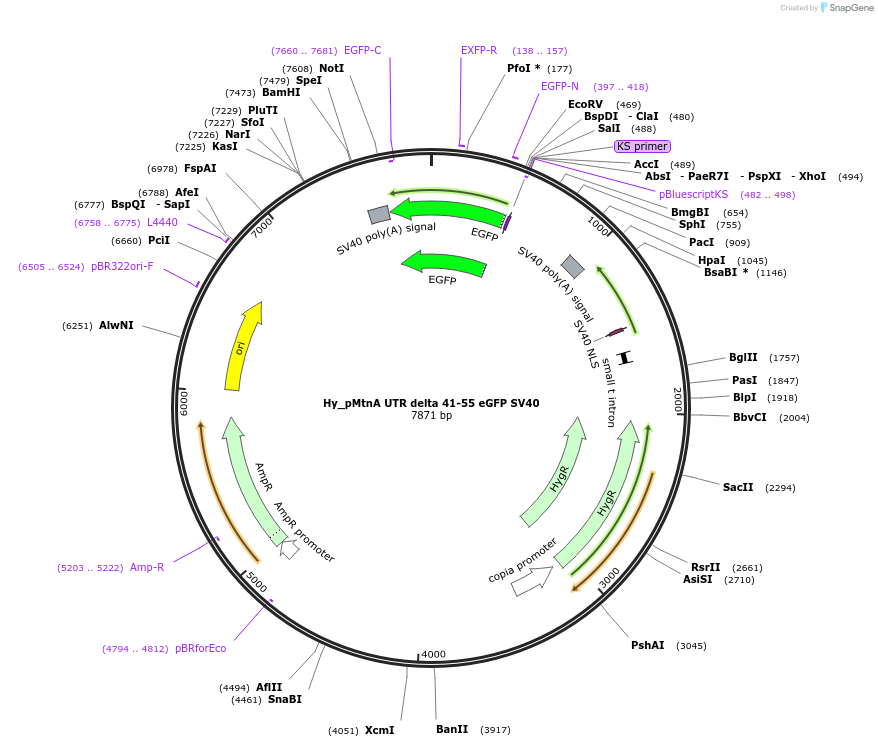
Hy_pMtnA UTR delta 41-55 eGFP SV40
Plasmid#132652PurposeExpresses an eGFP mRNA from the Drosophila Metallothionein A promoter that has 40 nt of the MtnA 5' UTRDepositorInserteGFP
ExpressionInsectPromoterMtnAAvailable SinceOct. 2, 2019AvailabilityAcademic Institutions and Nonprofits only -
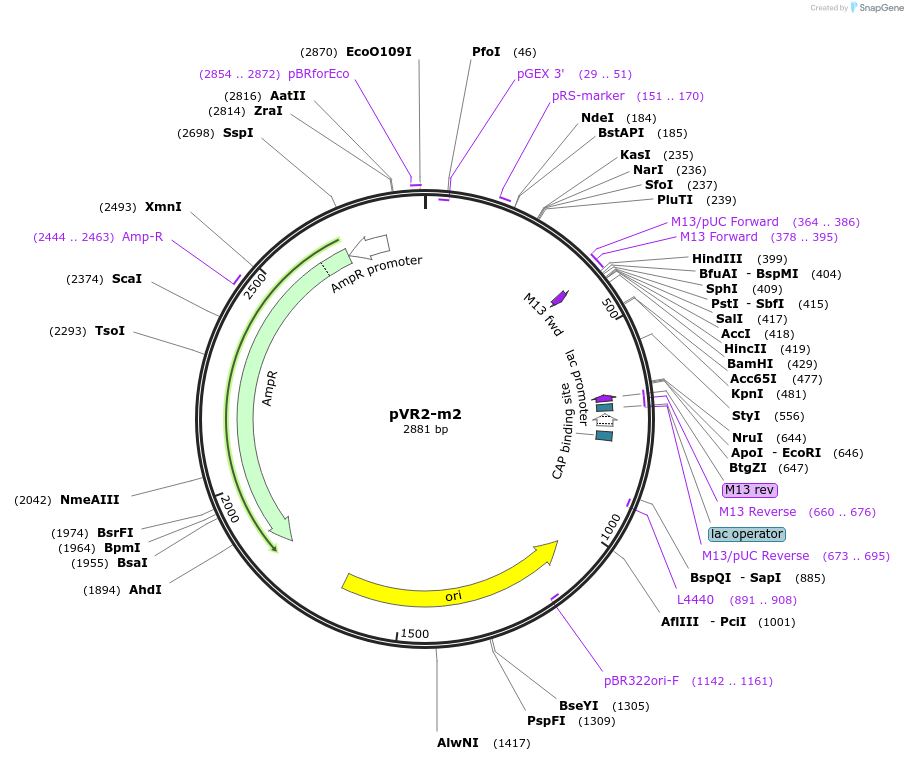
pVR2-m2
Plasmid#84721PurposepVR2 but with a G to U mutation in the riboswitch aptamerDepositorInsertsE. coli RecA promoter
stretch lacking A's followed by 5' UTR of queC gene
ExpressionBacterialMutationG to U mutation at position 16 of annotated "…Available SinceMarch 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
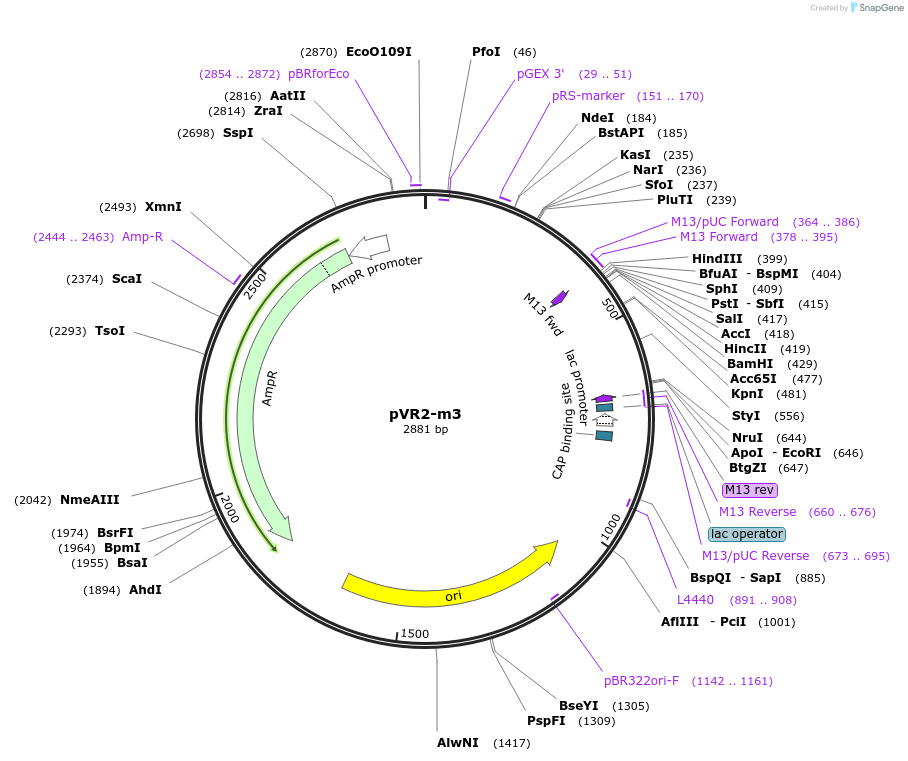
pVR2-m3
Plasmid#84722PurposepVR2 but with a C to G mutation in the riboswitch aptamerDepositorInsertsE. coli RecA promoter
stretch lacking A's followed by 5' UTR of queC gene
ExpressionBacterialMutationC to G mutation at position 22 of annotated "…Available SinceMarch 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
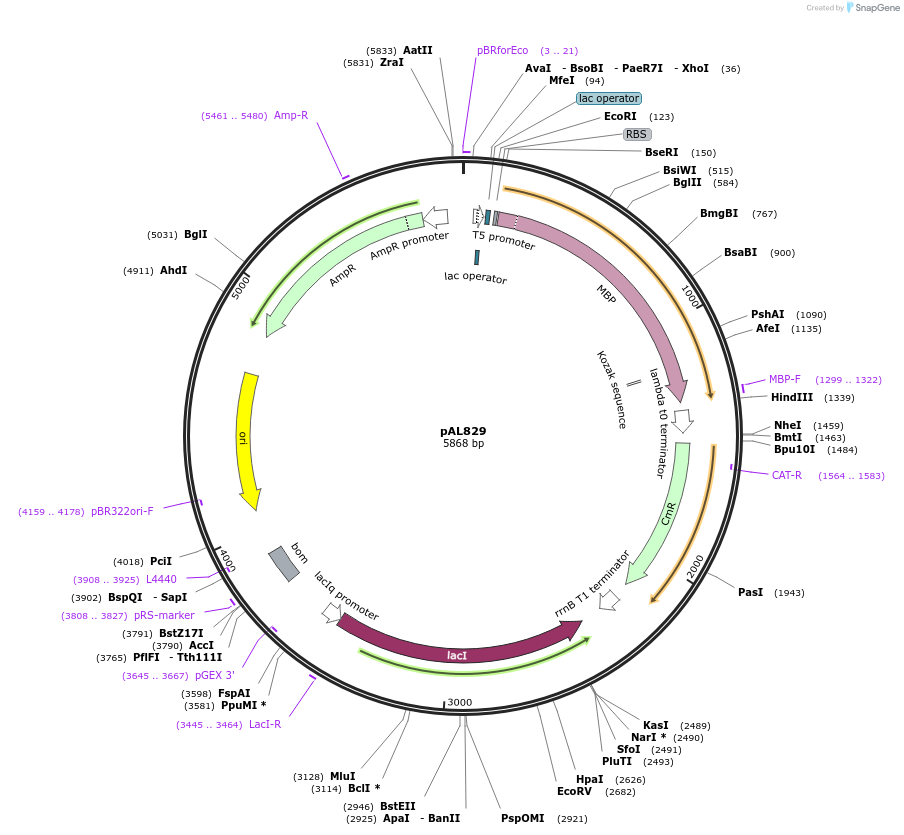
pAL829
Plasmid#119697Purposeprecursor slow-folding maltodextrin binding protein, MBP or malE, with mutations Y283D, G289C and V347CDepositorInsertslow-folding maltose ABC transporter periplasmic binding protein, malE, with mutations Y283D, G289C and V347C
TagsnoneExpressionBacterialMutationmutations Y283D, G289C and V347CPromoterT5Available SinceJan. 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
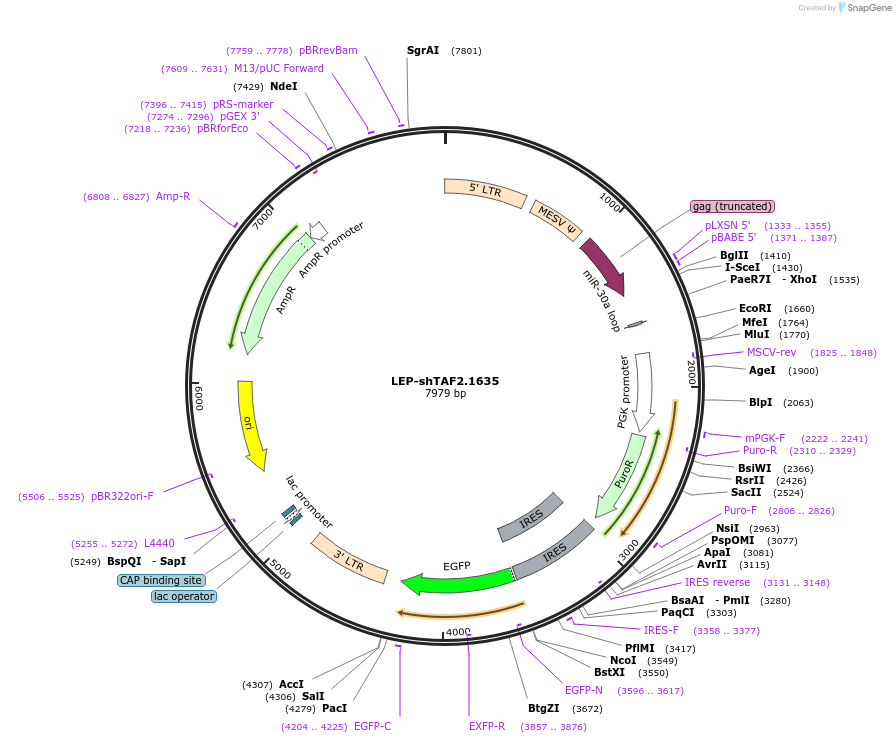
LEP-shTAF2.1635
Plasmid#105561Purposeretrovirally express TAF2 shRNA with puro resistance and GFP markerDepositorInsertTAF2 shRNA
UseRNAi and RetroviralExpressionMammalianPromoterMSCV-LTRAvailable SinceFeb. 13, 2018AvailabilityAcademic Institutions and Nonprofits only -
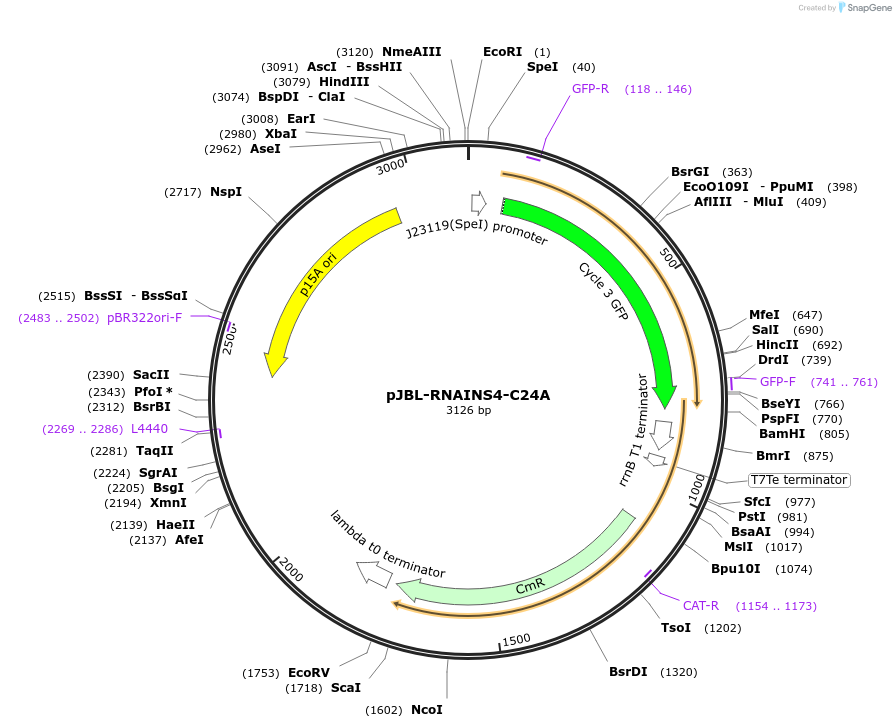
pJBL-RNAINS4-C24A
Plasmid#69462PurposeRNA-IN S4 C24A variant 5' UTR sequence controlling expression of superfolder GFP (UTR sequence from Mutalik et al., Nat. Chem. Biol., 2012)DepositorInsertRNA-IN S4 with C24A mutation
UseSynthetic BiologyExpressionBacterialMutationC24A and mutations for S4 variant (from wt)PromoterJ23119 and J23119 (sigma 70)Available SinceSept. 22, 2015AvailabilityAcademic Institutions and Nonprofits only -
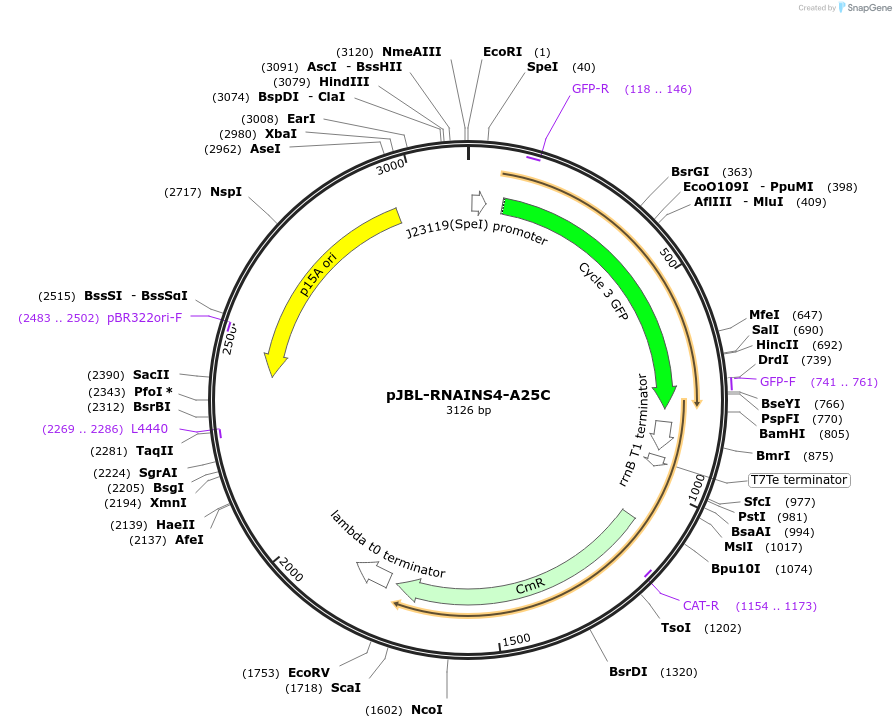
pJBL-RNAINS4-A25C
Plasmid#69461PurposeRNA-IN S4 A25C variant 5' UTR sequence controlling expression of superfolder GFP (UTR sequence from Mutalik et al., Nat. Chem. Biol., 2012)DepositorInsertRNA-IN S4 with A25C mutation
UseSynthetic BiologyExpressionBacterialMutationA25C and mutations for S4 variant (from wt)PromoterJ23119 and J23119 (sigma 70)Available SinceSept. 22, 2015AvailabilityAcademic Institutions and Nonprofits only -
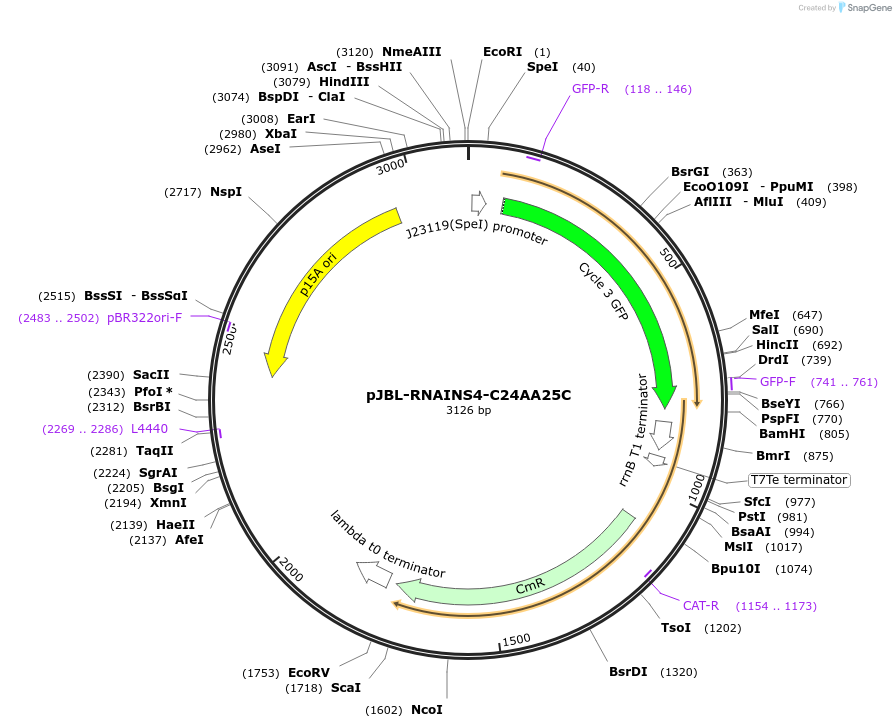
pJBL-RNAINS4-C24AA25C
Plasmid#69463PurposeRNA-IN S4 C24AA25C variant 5' UTR sequence controlling expression of superfolder GFP (UTR sequence from Mutalik et al., Nat. Chem. Biol., 2012)DepositorInsertRNA-IN S4 with C24A, A25C mutations
UseSynthetic BiologyExpressionBacterialMutationC24A, A25C, and mutations for S4 variant (from wt)PromoterJ23119 and J23119 (sigma 70)Available SinceSept. 22, 2015AvailabilityAcademic Institutions and Nonprofits only -
pJexpress414_OmpA171_WTnoGirdles
Plasmid#62653PurposeWild-type OmpA with aromatic girdle residues replaced by leucineDepositorInsertOmpA_noGirdles
UseSynthetic BiologyTags6His and FlAsHExpressionBacterialMutationAromatic girdle residues mutated to Leu (W7, W15,…PromoterT7Available SinceMay 27, 2015AvailabilityAcademic Institutions and Nonprofits only -
pMiR-HPV16-E7/E1-miR-375-m1-5
Plasmid#53704PurposeLuciferase reporter assay for HPV16 E7/E1 with point mutation on miR-375 binding siteDepositorInsertHPV16-E7/E1
UseLuciferaseMutationMutation in miR-375 binding sites m1-5 (refer to …Available SinceJuly 22, 2014AvailabilityAcademic Institutions and Nonprofits only