We narrowed to 11,638 results for: nar
-
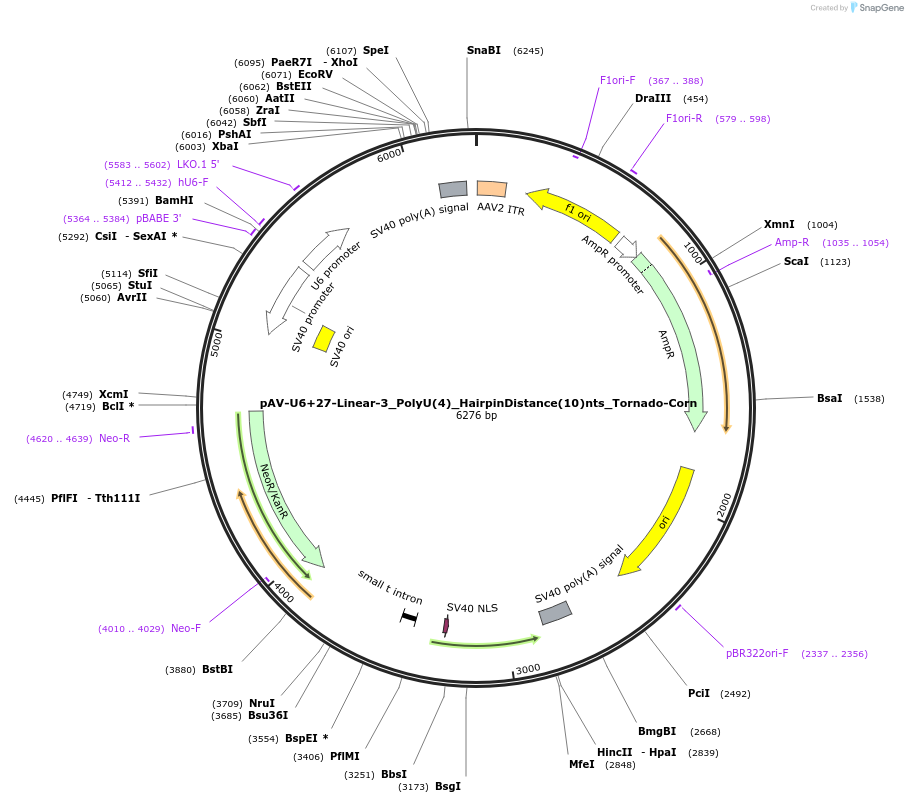
Plasmid#159506PurposeTests for the impacts 4 uracils in a poly-uracil tract in conjunctiuon with a hairpin 10 nts upstream have on human polymerase III transcription elongation from a U6 promoter.DepositorInsertTermination Module:Linear-3_PolyU(4)_HairpinDistance(10)nts
ExpressionMammalianPromoterU6-27Available SinceOct. 13, 2020AvailabilityAcademic Institutions and Nonprofits only -
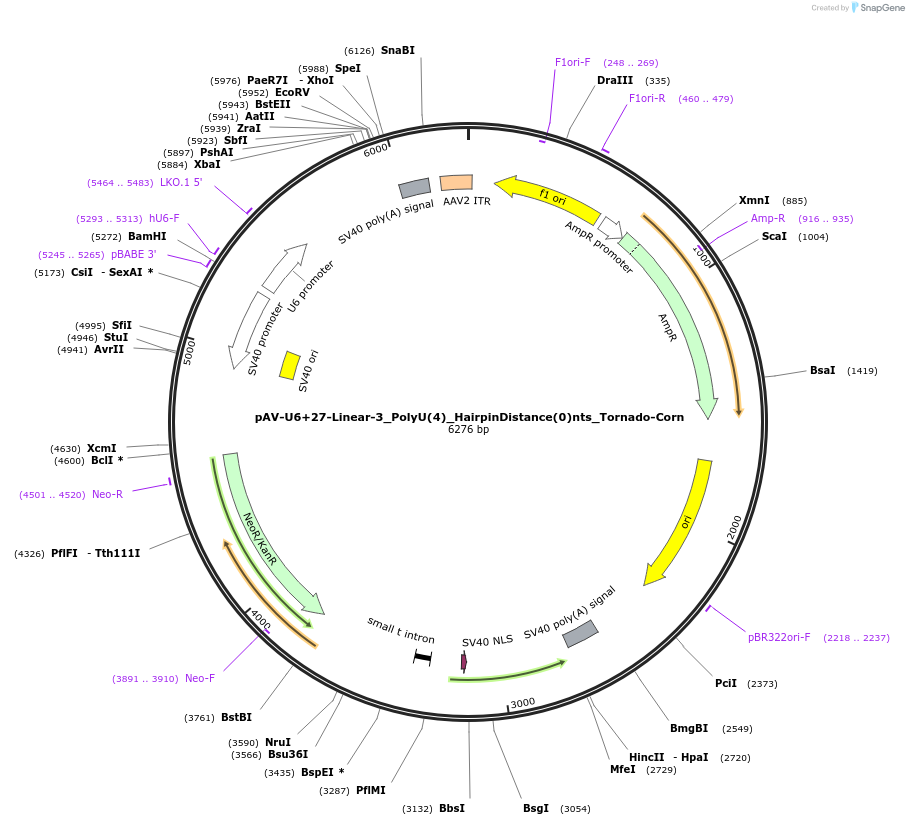
pAV-U6+27-Linear-3_PolyU(4)_HairpinDistance(0)nts_Tornado-Corn
Plasmid#159508PurposeTests for the impacts 4 uracils in a poly-uracil tract in conjunctiuon with an immediately upstream hairpin have on human polymerase III transcription elongation from a U6 promoter.DepositorInsertTermination Module:Linear-3_PolyU(4)_HairpinDistance(0)nts
ExpressionMammalianPromoterU6-27Available SinceOct. 13, 2020AvailabilityAcademic Institutions and Nonprofits only -
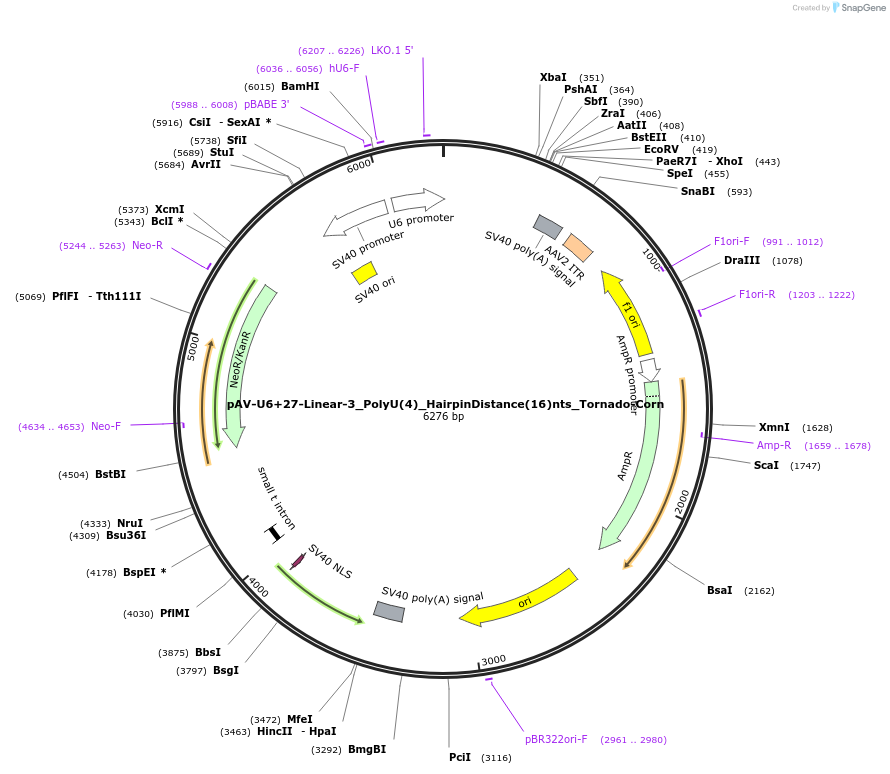
pAV-U6+27-Linear-3_PolyU(4)_HairpinDistance(16)nts_Tornado-Corn
Plasmid#159511PurposeTests for the impacts 4 uracils in a poly-uracil tract in conjunctiuon with a hairpin 16 nts upstream have on human polymerase III transcription elongation from a U6 promoter.DepositorInsertTermination Module:Linear-3_PolyU(4)_HairpinDistance(16)nts
ExpressionMammalianPromoterU6-27Available SinceOct. 13, 2020AvailabilityAcademic Institutions and Nonprofits only -
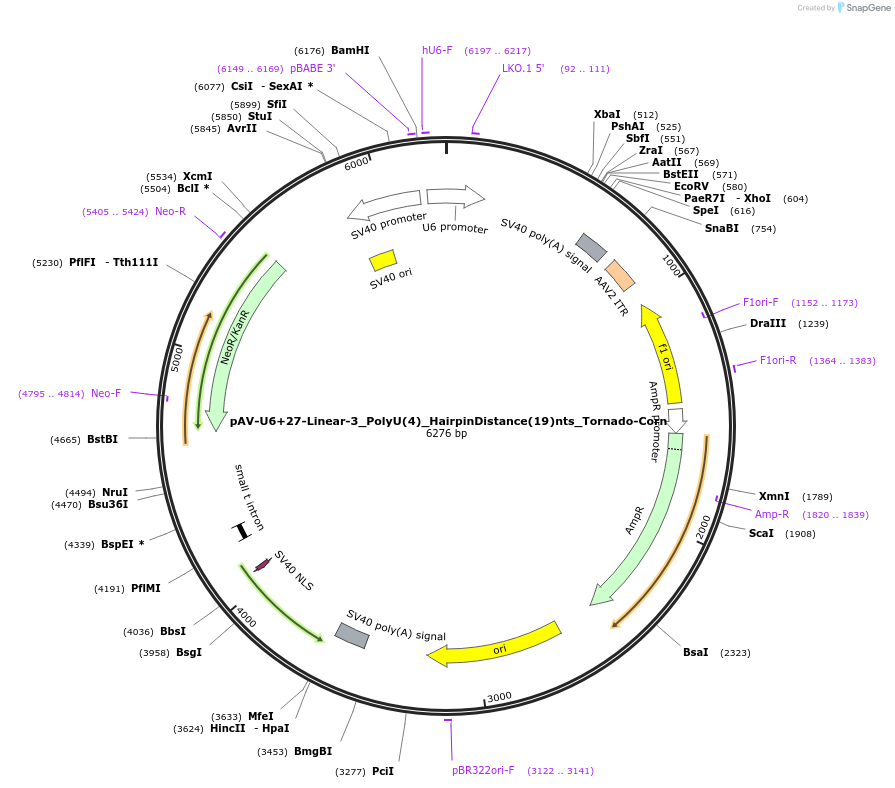
pAV-U6+27-Linear-3_PolyU(4)_HairpinDistance(19)nts_Tornado-Corn
Plasmid#159514PurposeTests for the impacts 4 uracils in a poly-uracil tract in conjunctiuon with a hairpin 19 nts upstream have on human polymerase III transcription elongation from a U6 promoter.DepositorInsertTermination Module:Linear-3_PolyU(4)_HairpinDistance(19)nts
ExpressionMammalianPromoterU6-27Available SinceOct. 13, 2020AvailabilityAcademic Institutions and Nonprofits only -
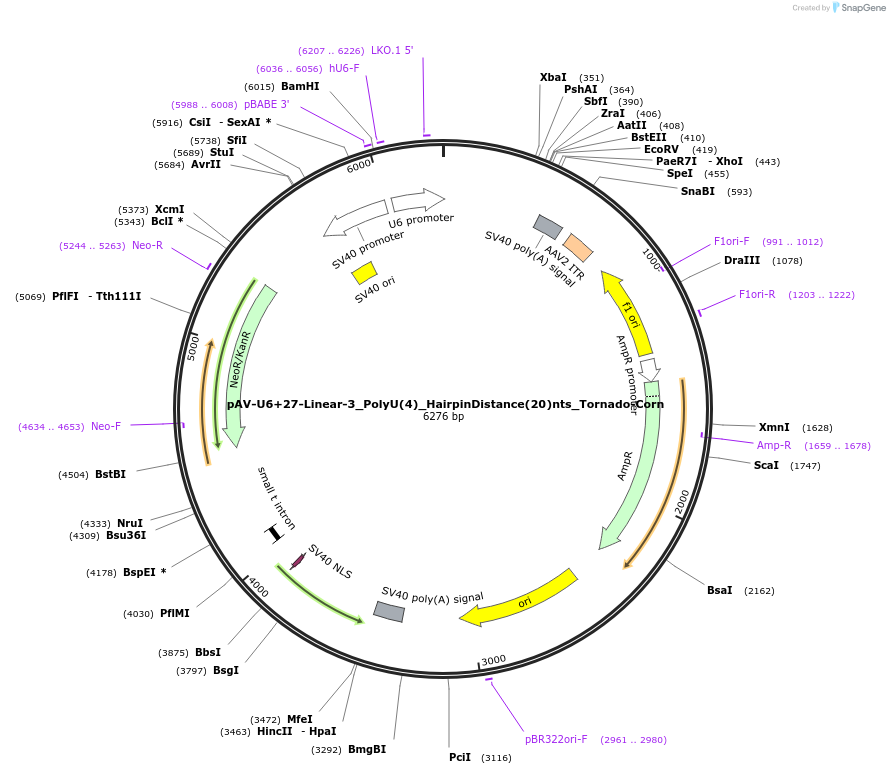
pAV-U6+27-Linear-3_PolyU(4)_HairpinDistance(20)nts_Tornado-Corn
Plasmid#159515PurposeTests for the impacts 4 uracils in a poly-uracil tract in conjunctiuon with a hairpin 20 nts upstream have on human polymerase III transcription elongation from a U6 promoter.DepositorInsertTermination Module:Linear-3_PolyU(4)_HairpinDistance(20)nts
ExpressionMammalianPromoterU6-27Available SinceOct. 13, 2020AvailabilityAcademic Institutions and Nonprofits only -
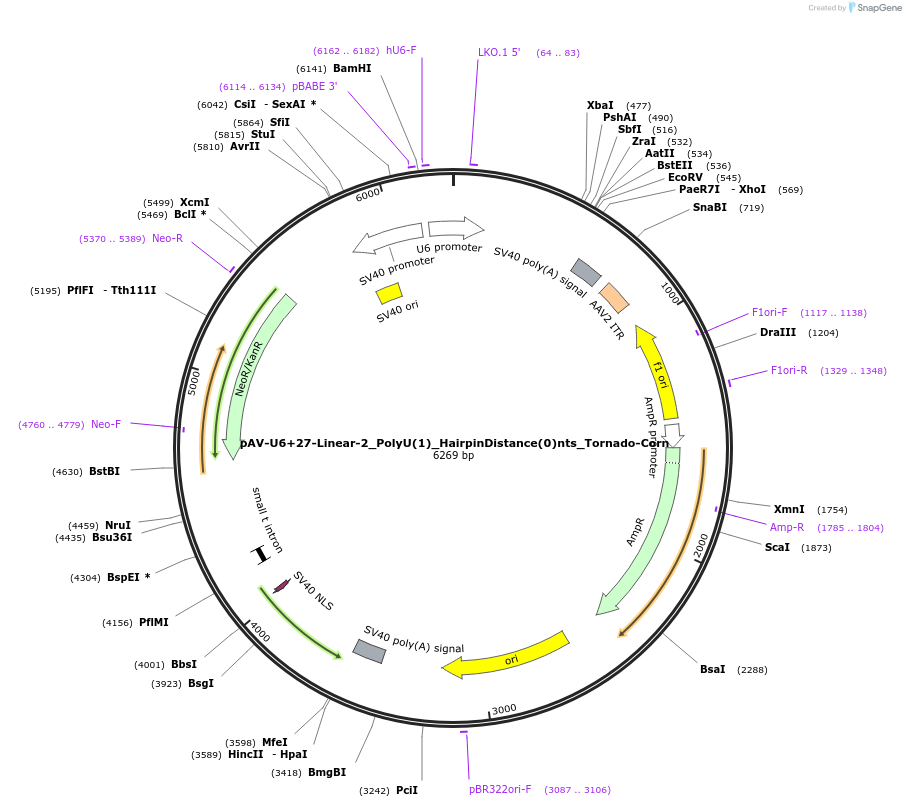
pAV-U6+27-Linear-2_PolyU(1)_HairpinDistance(0)nts_Tornado-Corn
Plasmid#159499PurposeTests for the impacts 1 uracil in a poly-uracil tract in conjunctiuon with an immediately upstream hairpin have on human polymerase III transcription elongation from a U6 promoter.DepositorInsertTermination Module:Linear-2_PolyU(1)_HairpinDistance(0)nts
ExpressionMammalianPromoterU6-27Available SinceOct. 13, 2020AvailabilityAcademic Institutions and Nonprofits only -
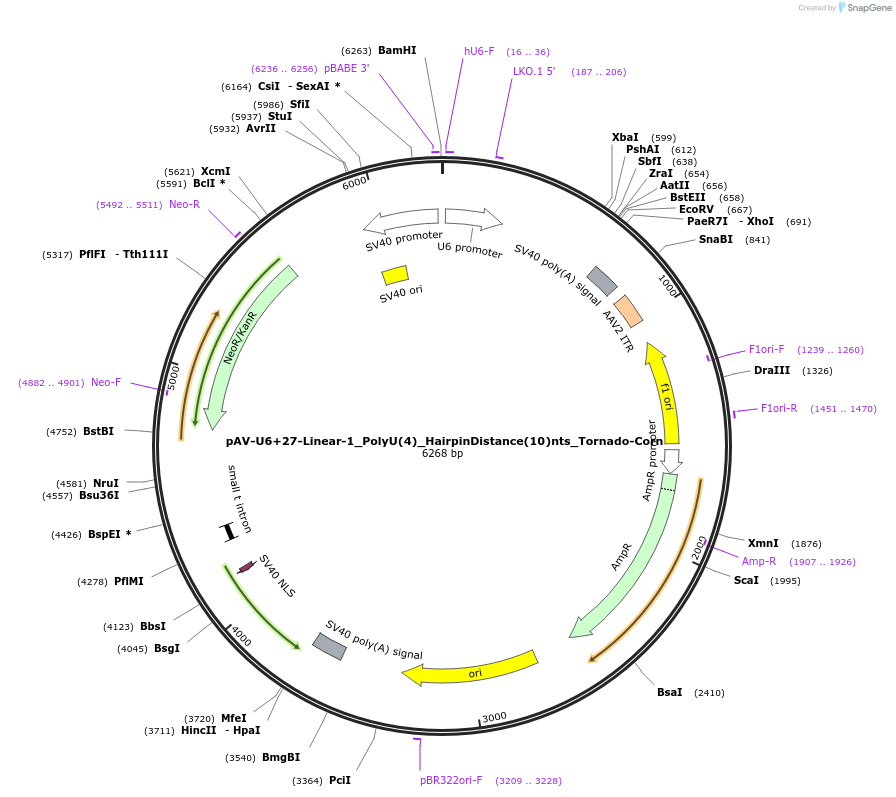
pAV-U6+27-Linear-1_PolyU(4)_HairpinDistance(10)nts_Tornado-Corn
Plasmid#159498PurposeTests for the impacts 4 uracils in a poly-uracil tract in conjunctiuon with a hairpin 10 nts upstream have on human polymerase III transcription elongation from a U6 promoter.DepositorInsertTermination Module:Linear-1_PolyU(4)_HairpinDistance(10)nts
ExpressionMammalianPromoterU6-27Available SinceOct. 13, 2020AvailabilityAcademic Institutions and Nonprofits only -
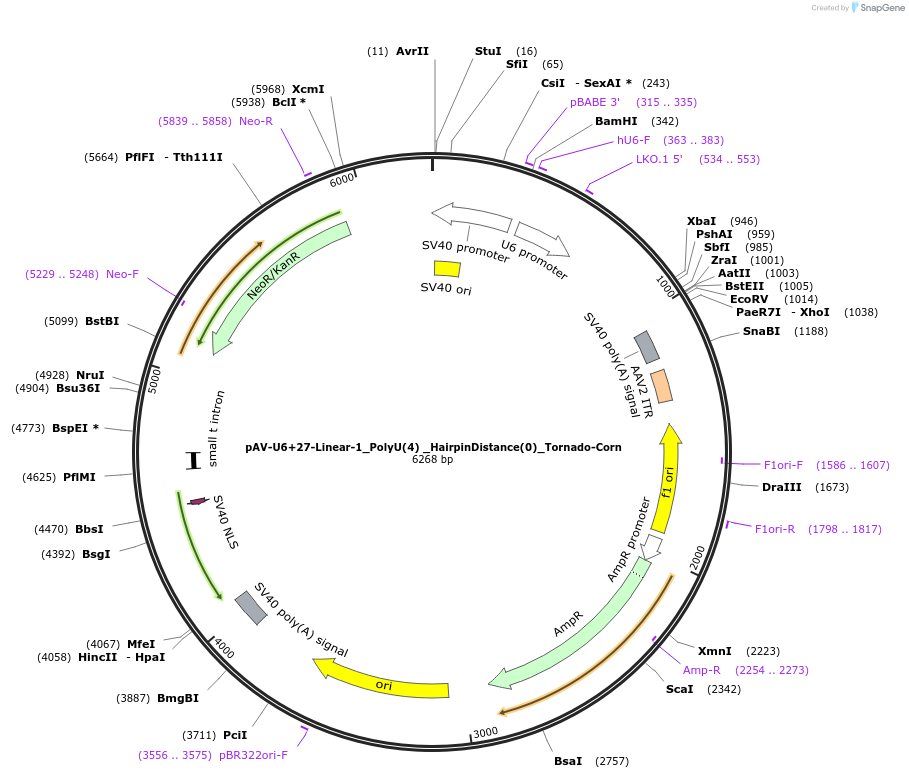
pAV-U6+27-Linear-1_PolyU(4) _HairpinDistance(0)_Tornado-Corn
Plasmid#159496PurposeTests for the impacts 4 uracils in a poly-uracil tract in conjunctiuon with an immediately upstream hairpin have on human polymerase III transcription elongation from a U6 promoter.DepositorInsertTermination Module:Linear-1_PolyU(4) _HairpinDistance(0)
ExpressionMammalianPromoterU6-27Available SinceOct. 13, 2020AvailabilityAcademic Institutions and Nonprofits only -
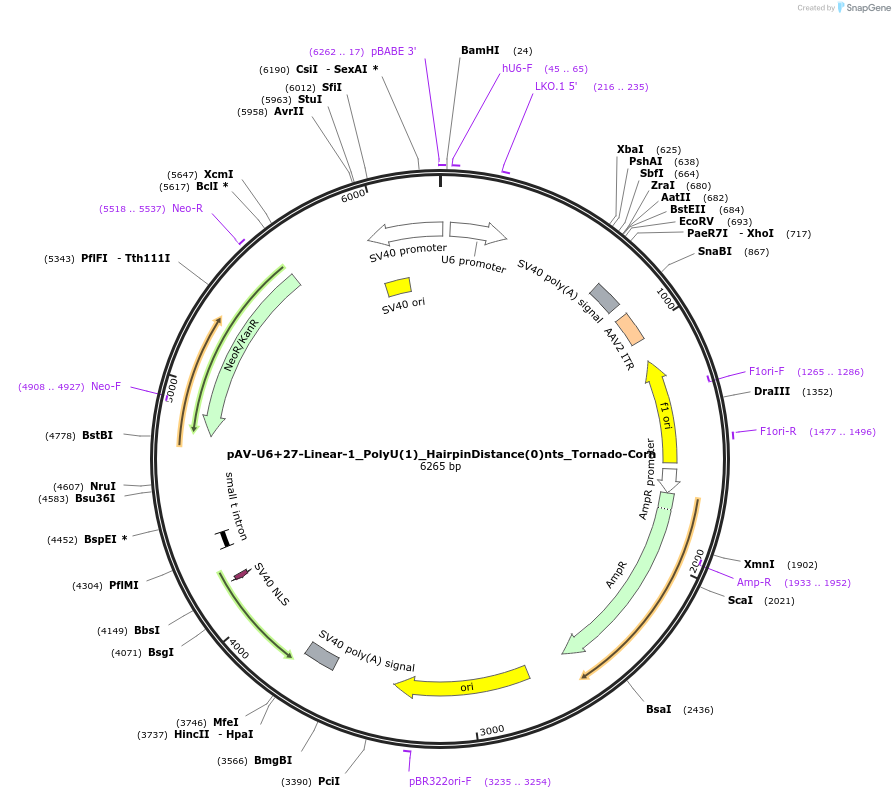
pAV-U6+27-Linear-1_PolyU(1)_HairpinDistance(0)nts_Tornado-Corn
Plasmid#159495PurposeTests for the impacts 1 uracil in a poly-uracil tract in conjunctiuon with an immediately upstream hairpin have on human polymerase III transcription elongation from a U6 promoter.DepositorInsertTermination Module:Linear-1_PolyU(1)_HairpinDistance(0)nts
ExpressionMammalianPromoterU6-27Available SinceOct. 13, 2020AvailabilityAcademic Institutions and Nonprofits only -
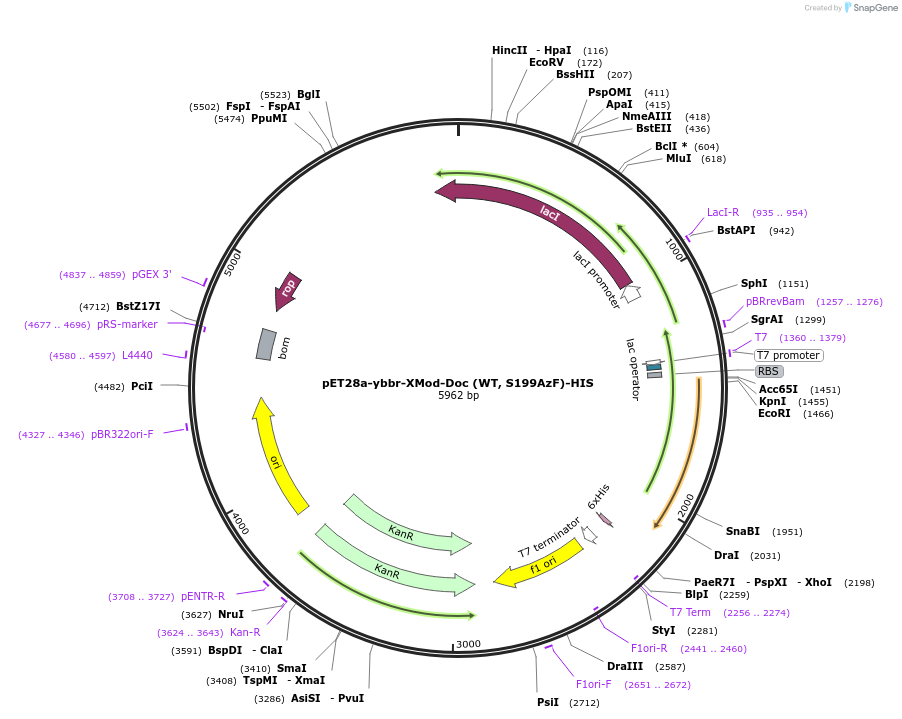
pET28a-ybbr-XMod-Doc (WT, S199AzF)-HIS
Plasmid#153443PurposeE. coli expression (amber suppression) of Rc. XDocB with serine at position 199 replaced with amber codon.DepositorInsertRc.XDocB
Tags6xHis and ybbr tagExpressionBacterialMutationSerine 199 was mutated to amber codon (TAG)PromoterT7 promoterAvailable SinceJuly 23, 2020AvailabilityAcademic Institutions and Nonprofits only -
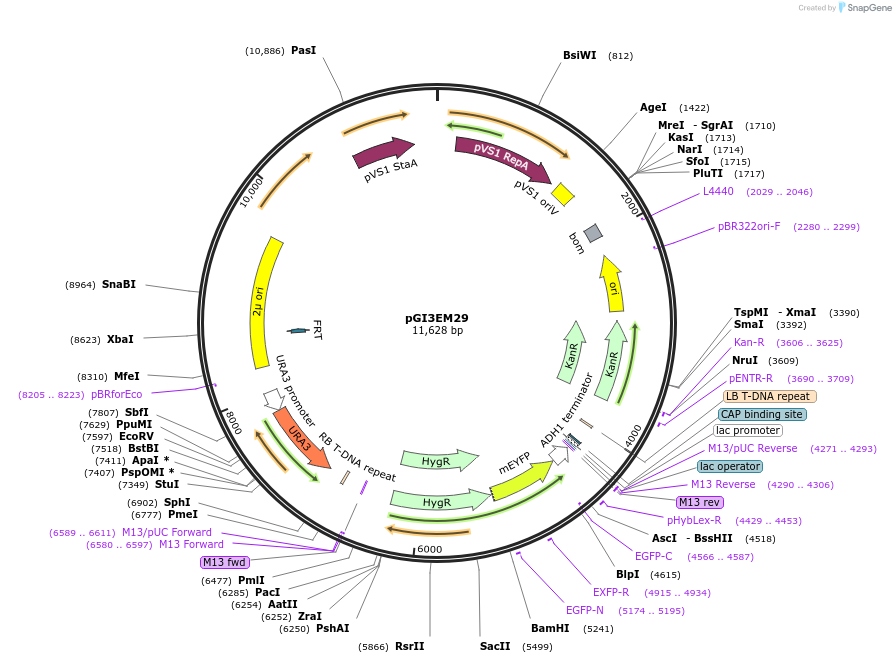
pGI3EM29
Plasmid#135489PurposeAgrobacterium T-DNA with Spun H2Bpr driving hph-mCitrineDepositorInserthph SpunH2A/Bpr hph-mCitrine
TagsmCitrineExpressionBacterial and YeastPromoterSpunH2A/Bpr (divergent)Available SinceMay 19, 2020AvailabilityAcademic Institutions and Nonprofits only -
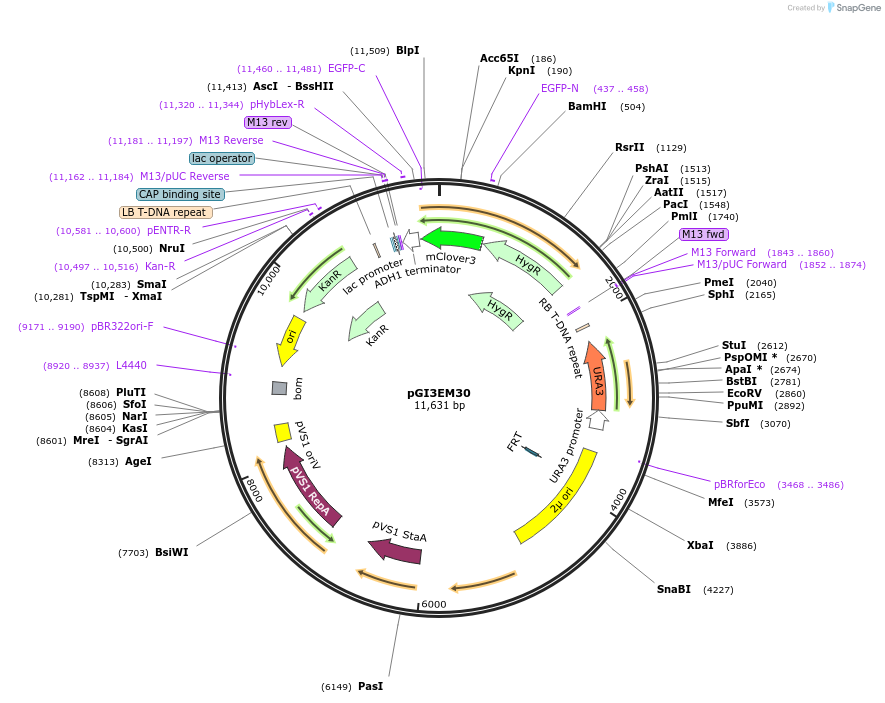
pGI3EM30
Plasmid#135490PurposeAgrobacterium T-DNA with Spun H2Bpr driving hph-mClover3DepositorInserthph SpunH2A/Bpr hph-mClover3
TagsmClover3ExpressionBacterial and YeastPromoterSpunH2A/Bpr (divergent)Available SinceMay 19, 2020AvailabilityAcademic Institutions and Nonprofits only -
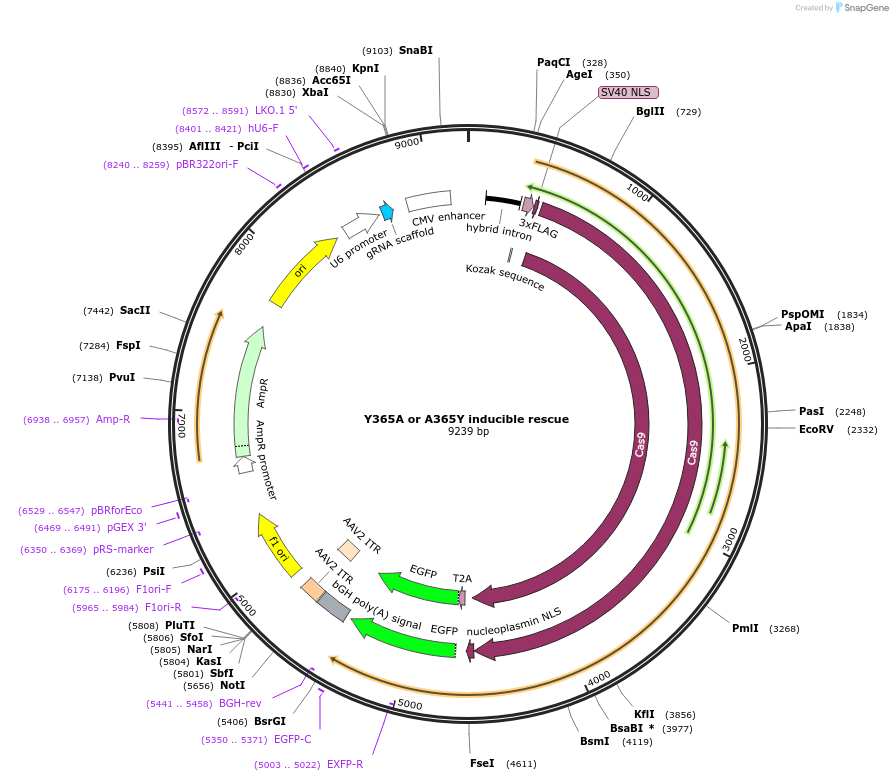
Y365A or A365Y inducible rescue
Plasmid#131323PurposegRNA to generate i-WT-r or i-MT-r systemsDepositorInsertY365A or A365Y inducible rescue
UseCRISPRExpressionMammalianAvailable SinceMarch 23, 2020AvailabilityAcademic Institutions and Nonprofits only -
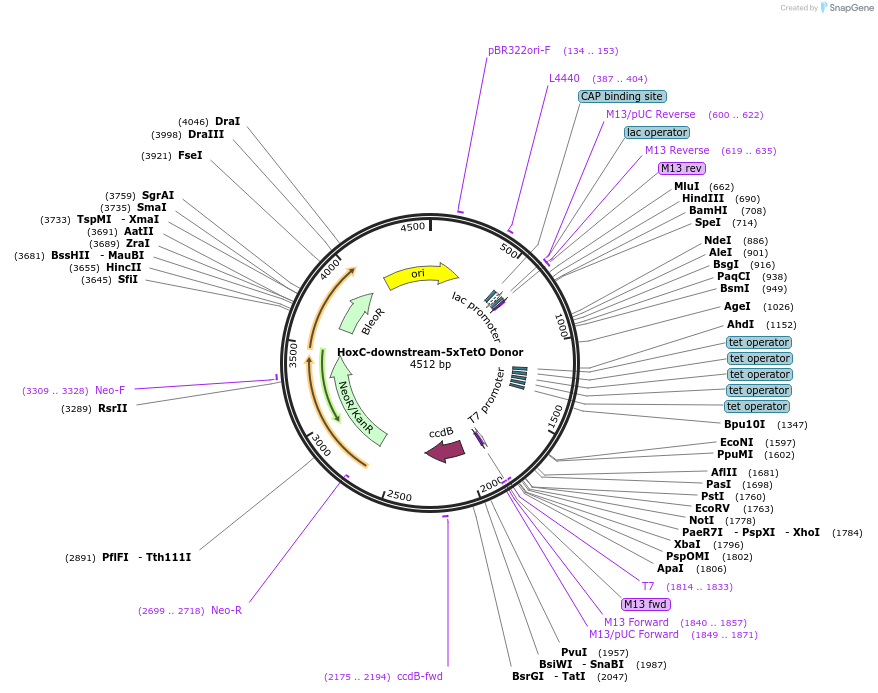
HoxC-downstream-5xTetO Donor
Plasmid#131341PurposeDonor template to insert 5xTet operator sequence into downstream of HoxC cluster.DepositorInsertHoxC-downstream-5xTetO Donor
UseCRISPR and Mouse TargetingAvailable SinceMarch 9, 2020AvailabilityAcademic Institutions and Nonprofits only -
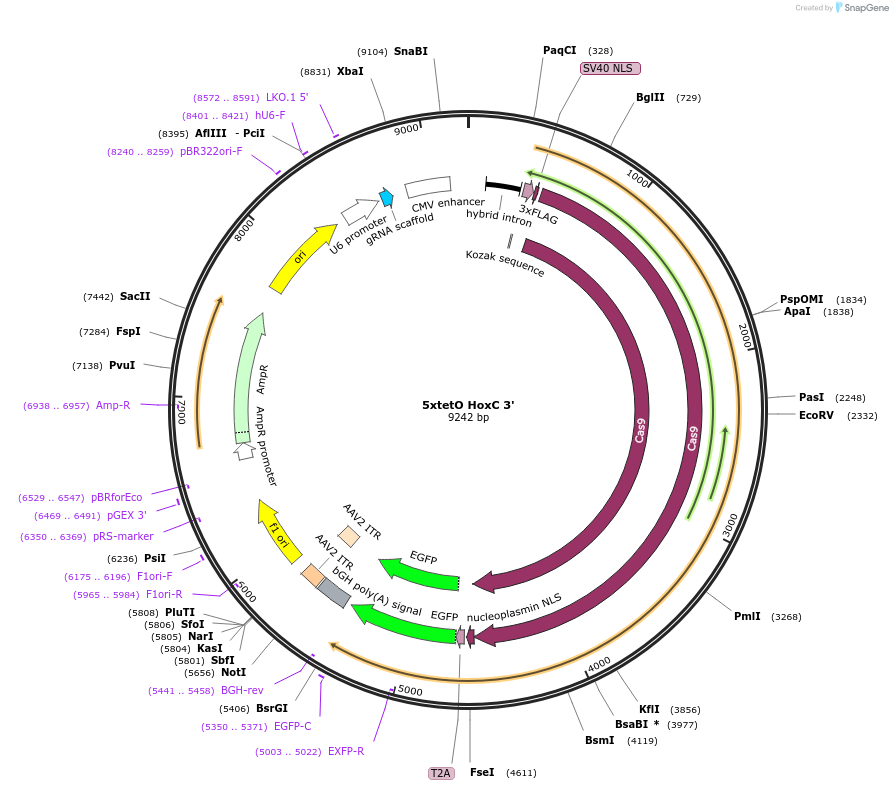
5xtetO HoxC 3'
Plasmid#131340PurposegRNA to insert 5xTet operator sequence into downstream of HoxC cluster.DepositorInsertHoxC-downstream-gRNA
UseCRISPRExpressionMammalianAvailable SinceMarch 9, 2020AvailabilityAcademic Institutions and Nonprofits only -
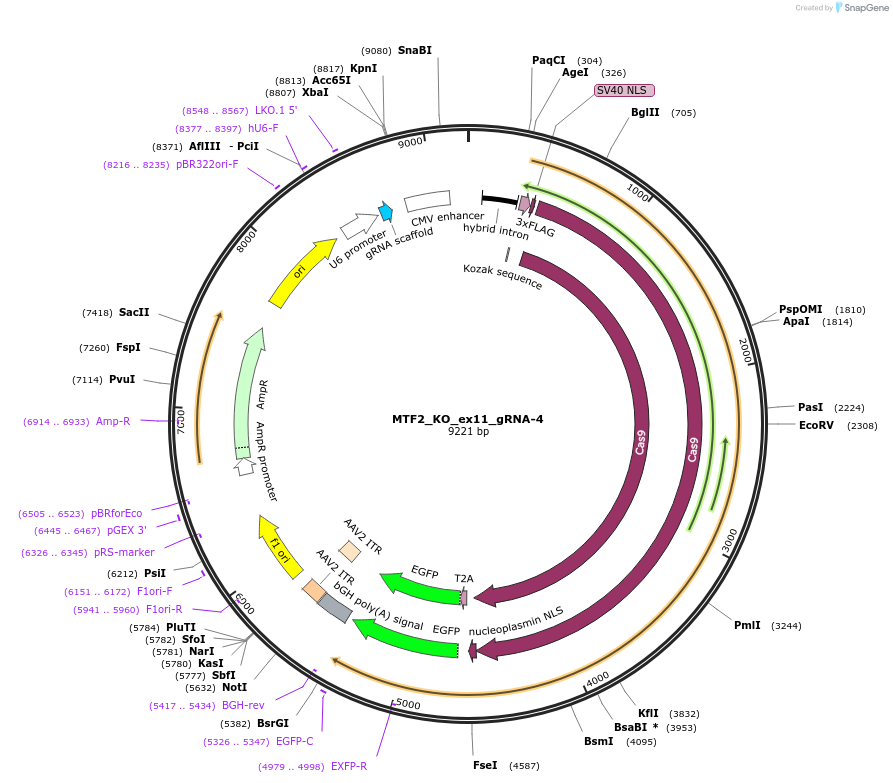
MTF2_KO_ex11_gRNA-4
Plasmid#131334PurposegRNA to knockout MTF2. This gRNA is also used in Haojie Li et al. Nature 2017. It targets exon 11 of MTF2.DepositorInsertMTF2_KO_ex11_gRNA
UseCRISPRExpressionMammalianAvailable SinceMarch 9, 2020AvailabilityAcademic Institutions and Nonprofits only -
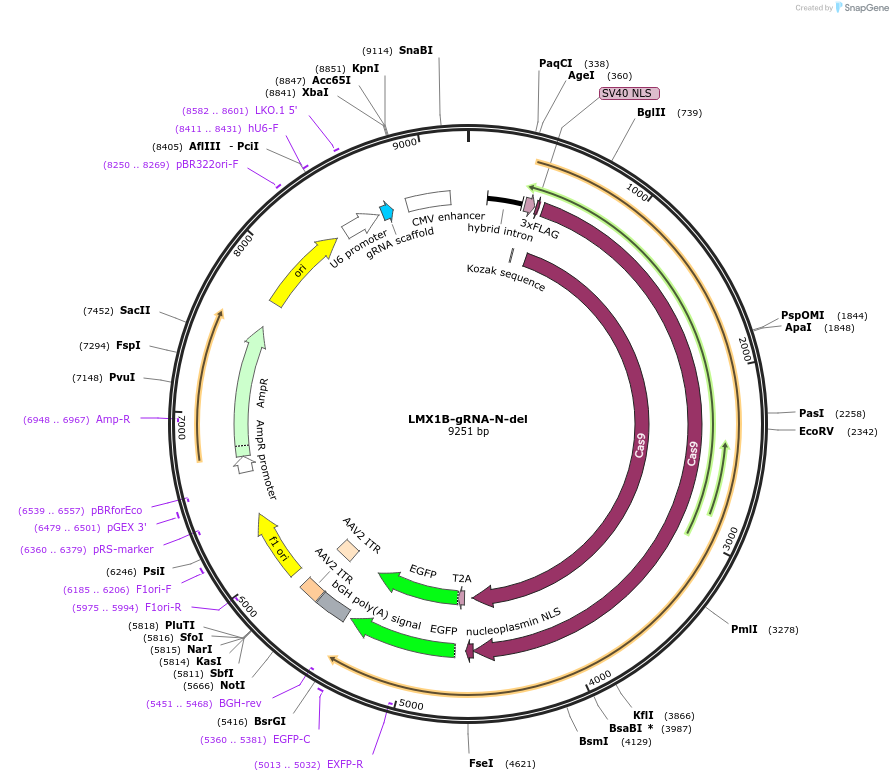
LMX1B-gRNA-N-del
Plasmid#131333PurposegRNA to delete a nucleation site near Lmx1b. Use with LMX1B-gRNA-C-delDepositorInsertLMX1B-gRNA-N-del
UseCRISPRExpressionMammalianAvailable SinceMarch 6, 2020AvailabilityAcademic Institutions and Nonprofits only -
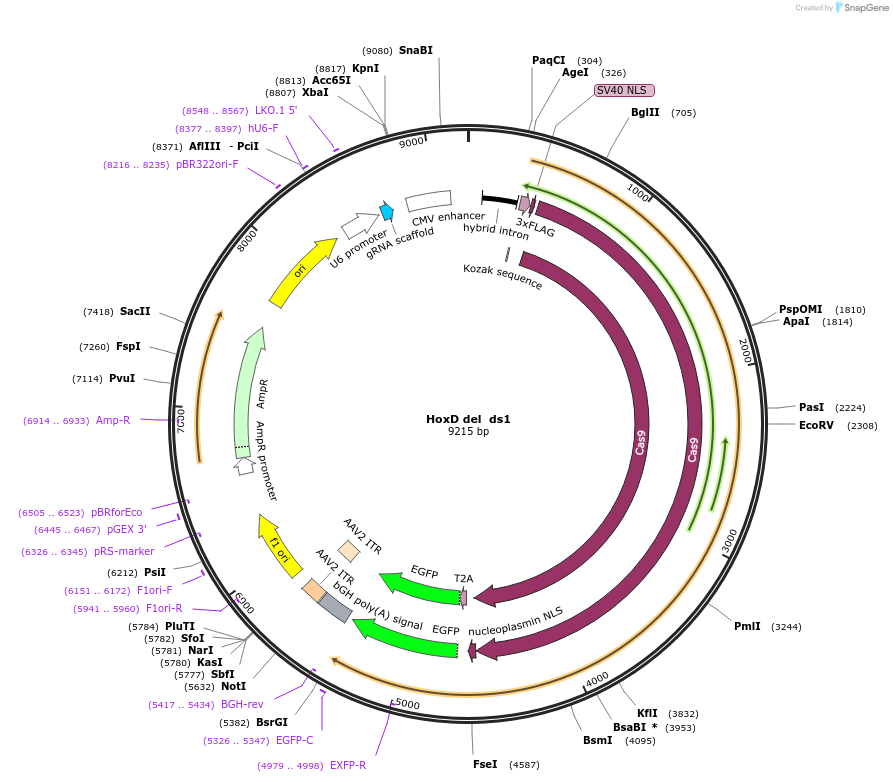
HoxD del ds1
Plasmid#131338PurposegRNA to delete a nucleation site near HoxD. Use with HoxD del us1DepositorInsertEVX2-gRNA-C-del
UseCRISPRExpressionMammalianAvailable SinceMarch 6, 2020AvailabilityAcademic Institutions and Nonprofits only -
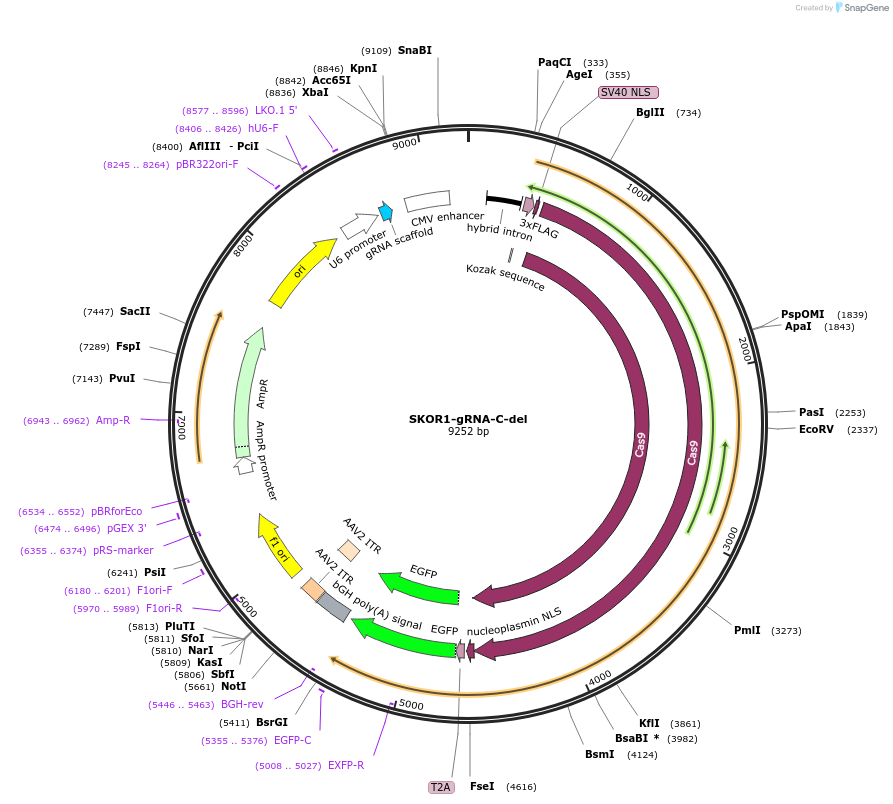
SKOR1-gRNA-C-del
Plasmid#131332PurposegRNA to delete a nucleation site near Skor1. Use with SKOR1-gRNA-N-delDepositorInsertSKOR1-gRNA-C-del
UseCRISPRExpressionMammalianAvailable SinceMarch 6, 2020AvailabilityAcademic Institutions and Nonprofits only -
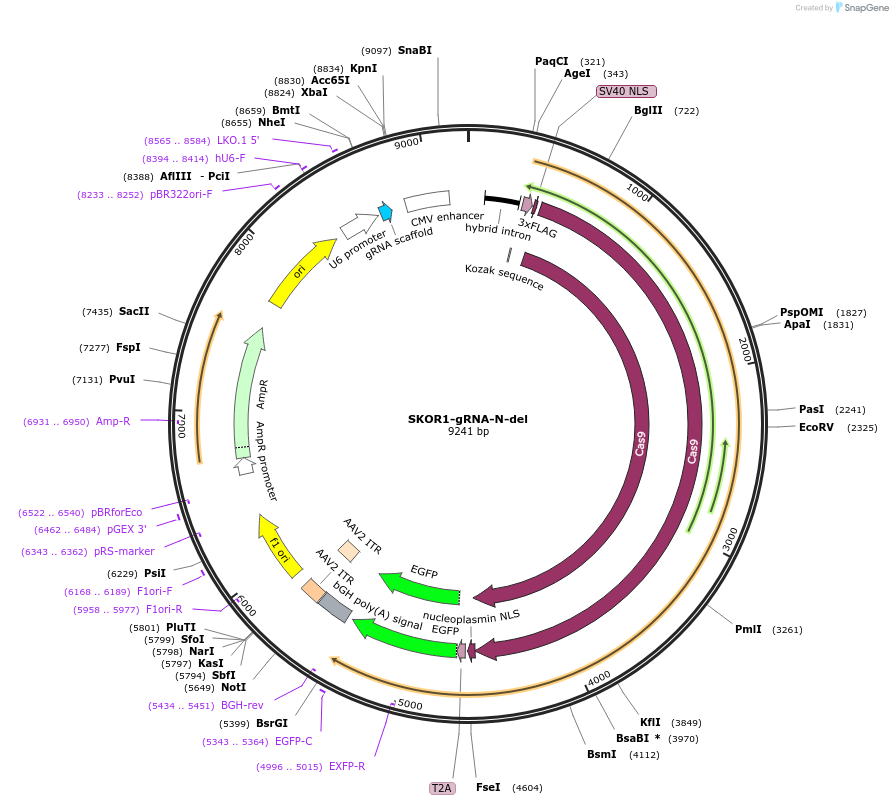
SKOR1-gRNA-N-del
Plasmid#131331PurposegRNA to delete a nucleation site near Skor1. Use with SKOR1-gRNA-C-delDepositorInsertSKOR1-gRNA-N-del
UseCRISPRExpressionMammalianAvailable SinceMarch 6, 2020AvailabilityAcademic Institutions and Nonprofits only -
HoxD del control ds
Plasmid#131336PurposegRNA to delete control region near HoxD. Use with HoxD del control usDepositorInsertCTRL-gRNA-C-del
UseCRISPRExpressionMammalianAvailable SinceMarch 6, 2020AvailabilityAcademic Institutions and Nonprofits only -
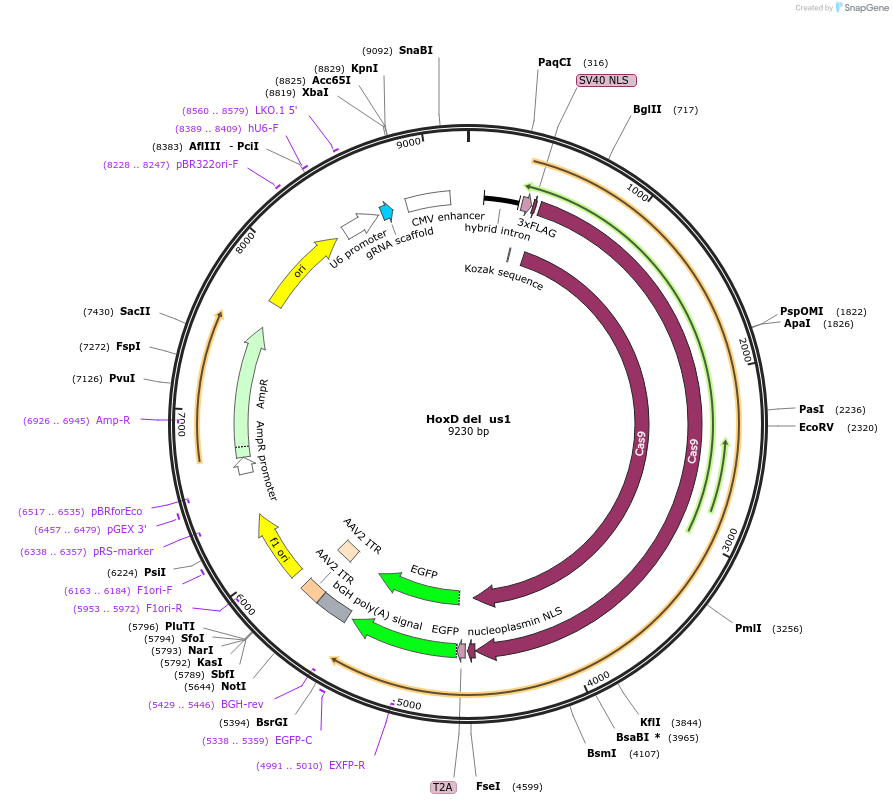
HoxD del us1
Plasmid#131337PurposegRNA to delete a nucleation site near HoxD. Use with HoxD del ds1DepositorInsertEVX2-gRNA-N-del
UseCRISPRExpressionMammalianAvailable SinceMarch 6, 2020AvailabilityAcademic Institutions and Nonprofits only -
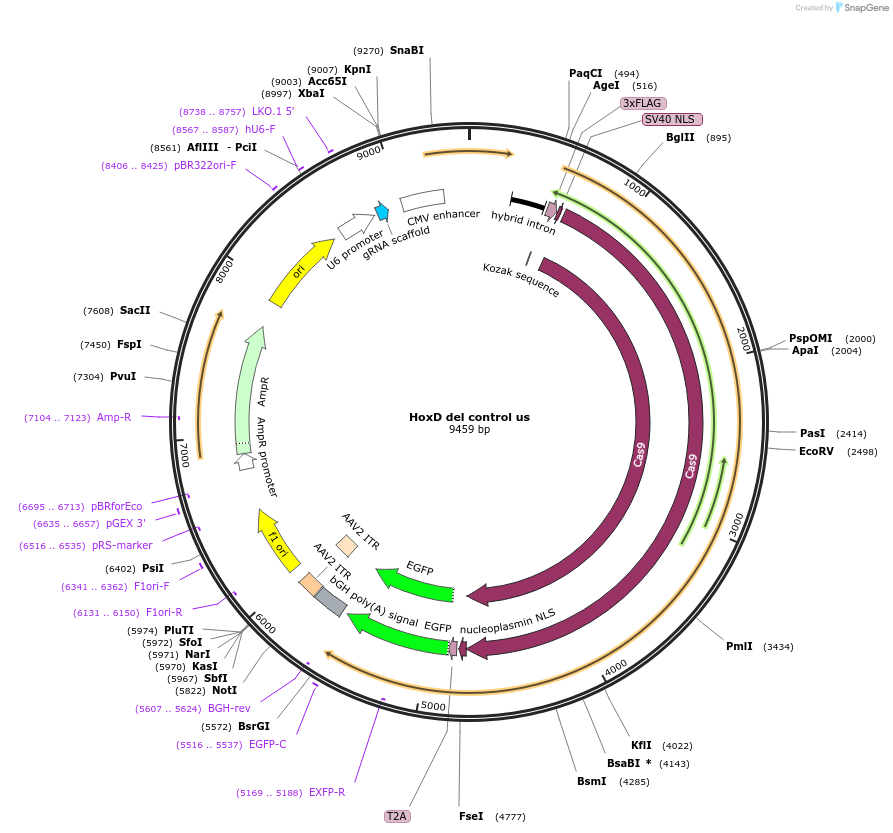
HoxD del control us
Plasmid#131335PurposegRNA to delete control region near HoxD. Use with HoxD del control dsDepositorInsertCTRL-gRNA-N-del
UseCRISPRExpressionMammalianAvailable SinceMarch 6, 2020AvailabilityAcademic Institutions and Nonprofits only -
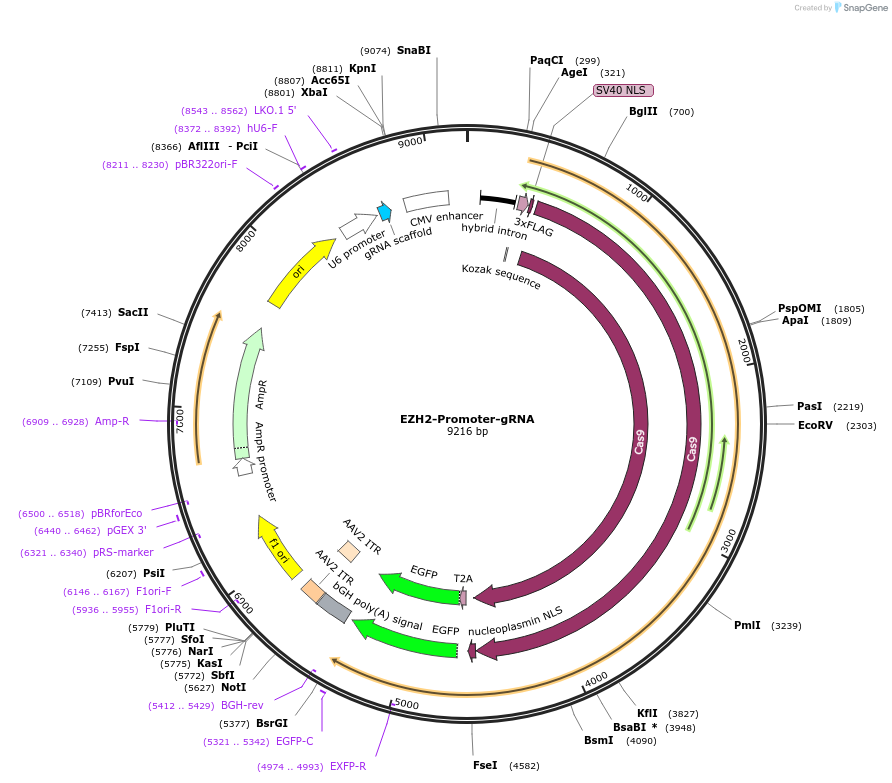
EZH2-Promoter-gRNA
Plasmid#131329PurposegRNA to generate endogenously tagged -tetR-EZH2DepositorInsertEZH2-Promoter-gRNA
UseCRISPRExpressionMammalianAvailable SinceMarch 6, 2020AvailabilityAcademic Institutions and Nonprofits only -
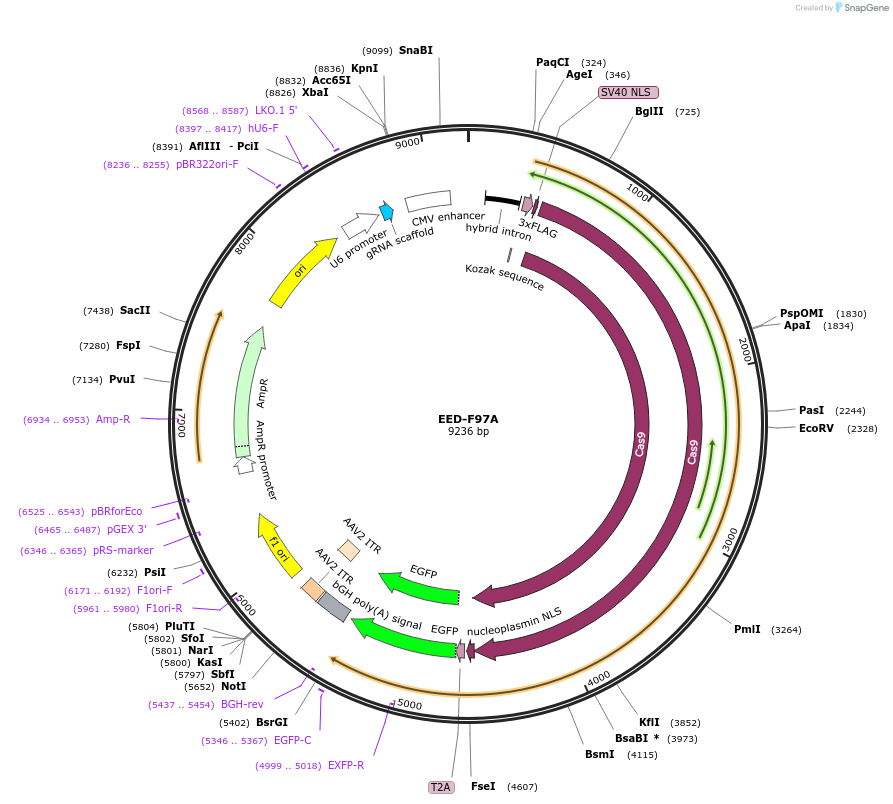
EED-F97A
Plasmid#131326PurposegRNA to generate F97A cage-mutant EEDDepositorInsertEED-KO-gRNA-2
UseCRISPRExpressionMammalianAvailable SinceMarch 6, 2020AvailabilityAcademic Institutions and Nonprofits only -
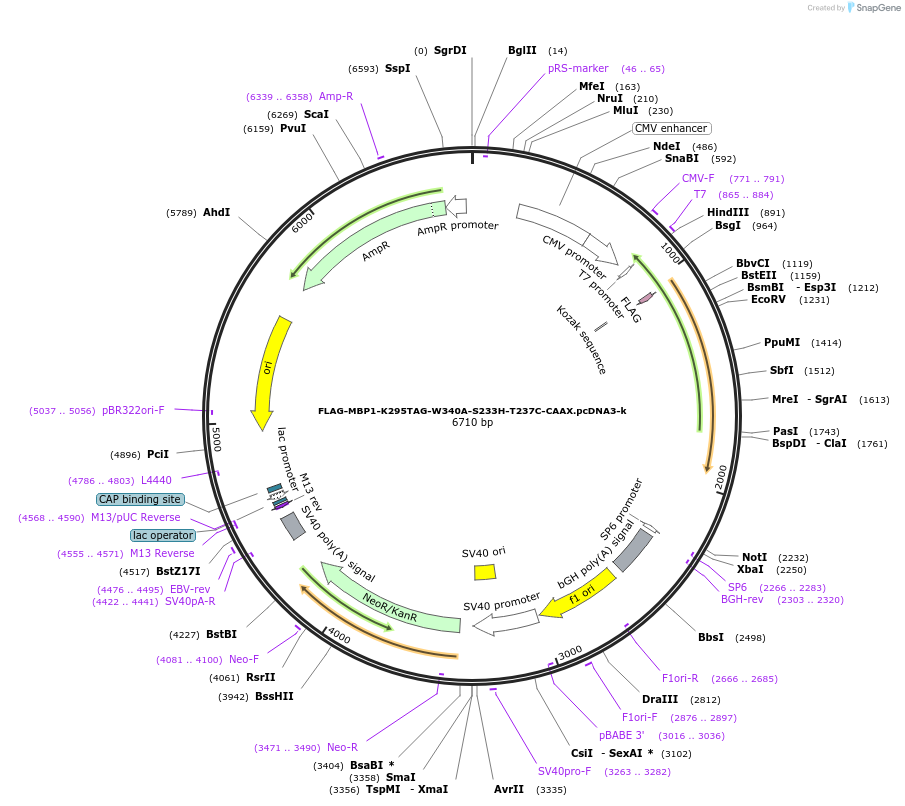
FLAG-MBP1-K295TAG-W340A-S233H-T237C-CAAX.pcDNA3-k
Plasmid#127403PurposeExpresses MBP in mammalian cells. Amber codon at AA# 295 as a noncanonical amino acid incorporation siteDepositorInsertMaltose Binding Protein
TagsCAAX and FLAGExpressionMammalianMutationAmino acid mutations: K295 to Amber codon, W340A,…PromoterCMVAvailable SinceJuly 3, 2019AvailabilityAcademic Institutions and Nonprofits only -
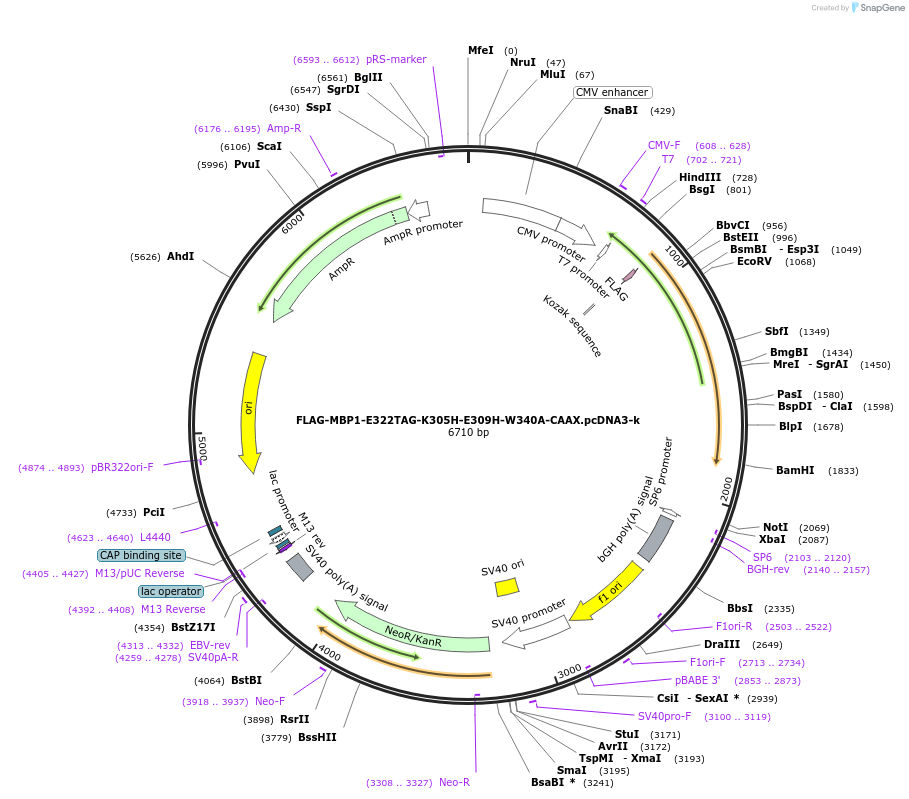
FLAG-MBP1-E322TAG-K305H-E309H-W340A-CAAX.pcDNA3-k
Plasmid#127407PurposeExpresses MBP in mammalian cells. Amber codon at AA# 322 as a noncanonical amino acid incorporation siteDepositorInsertMaltose Binding Protein
TagsCAAX and FLAGExpressionMammalianMutationAmino acid mutations: E322 to Amber codon, K305H,…PromoterCMVAvailable SinceJune 27, 2019AvailabilityAcademic Institutions and Nonprofits only -
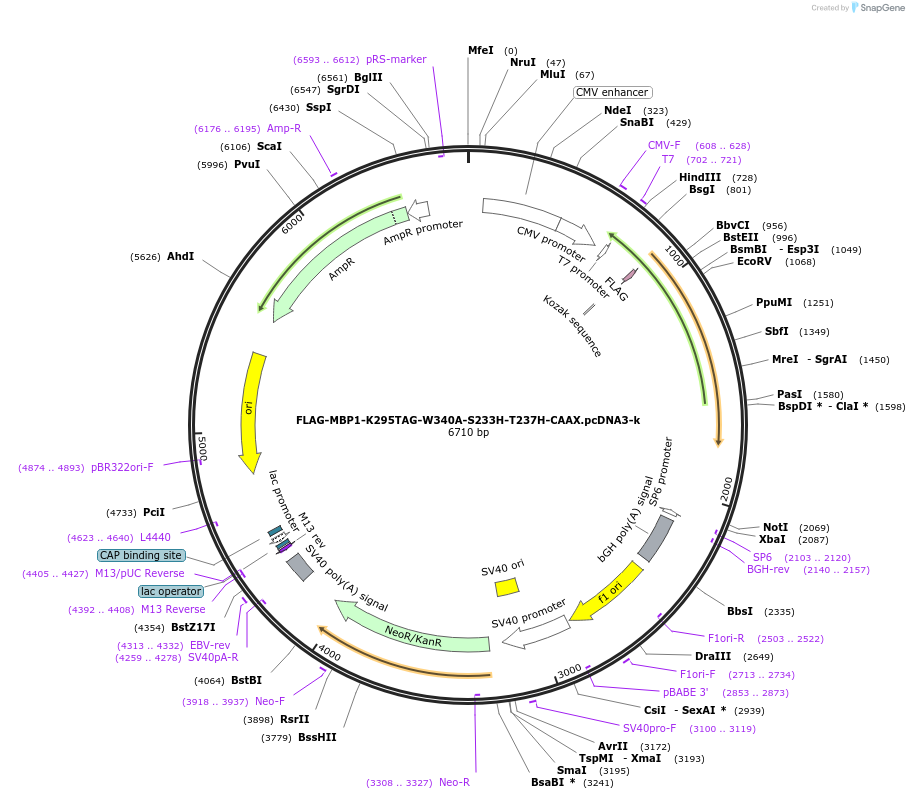
FLAG-MBP1-K295TAG-W340A-S233H-T237H-CAAX.pcDNA3-k
Plasmid#127401PurposeExpresses MBP in mammalian cells. Amber codon at AA# 295 as a noncanonical amino acid incorporation siteDepositorInsertMaltose Binding Protein
TagsCAAX and FLAGExpressionMammalianMutationAmino acid mutations: K295 to Amber codon, W340A,…PromoterCMVAvailable SinceJune 27, 2019AvailabilityAcademic Institutions and Nonprofits only -
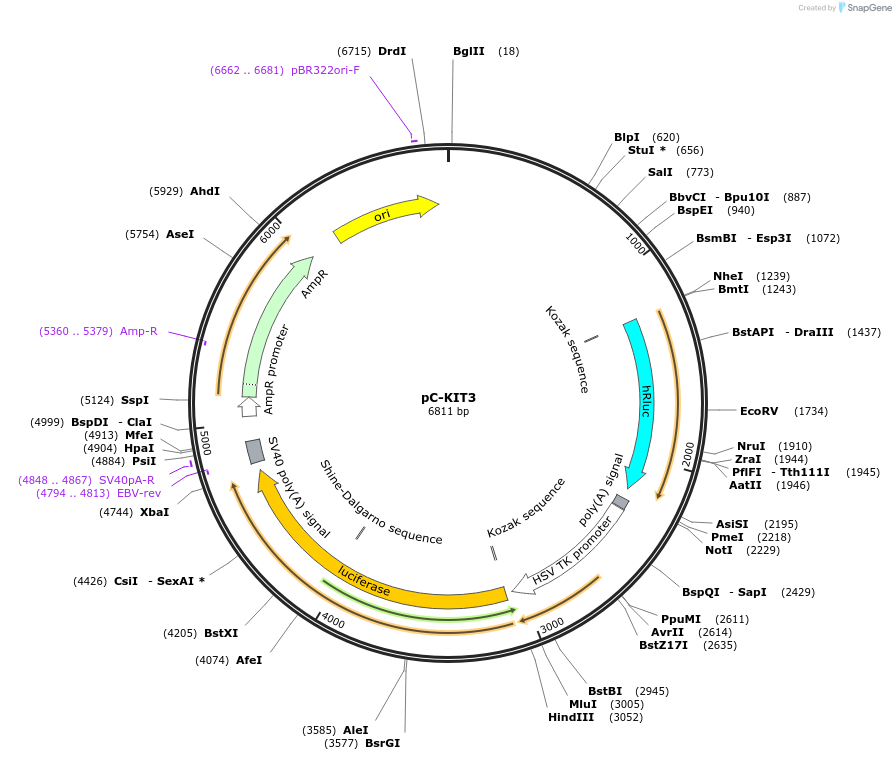
pC-KIT3
Plasmid#118985PurposeMutational dissection of the G rich region of the c-kit promoterDepositorAvailable SinceJune 10, 2019AvailabilityAcademic Institutions and Nonprofits only -
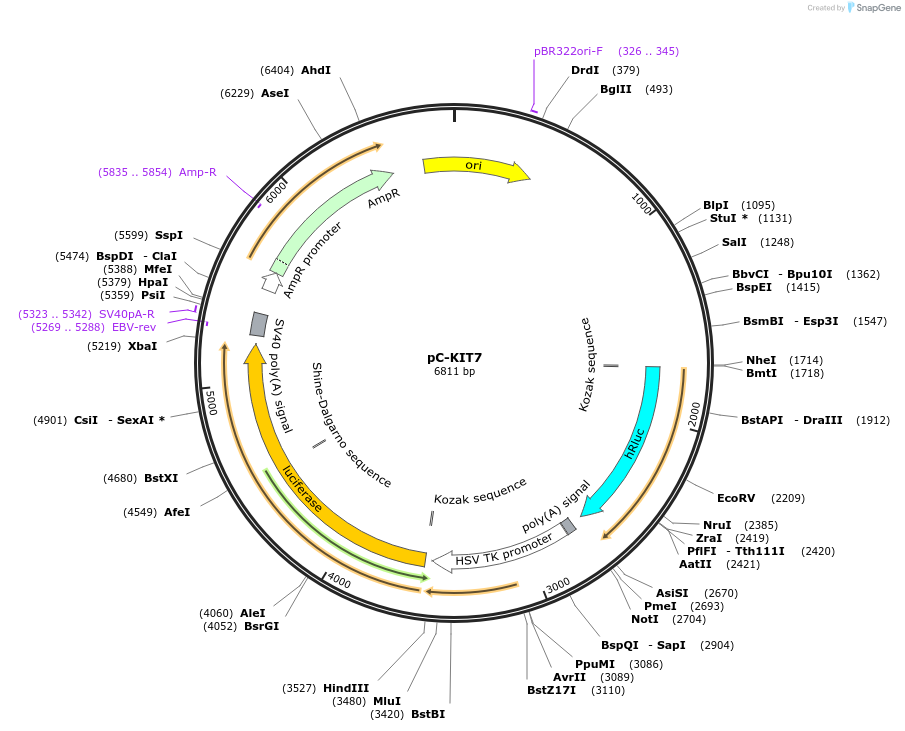
pC-KIT7
Plasmid#118989PurposeMutational dissection of the G rich region of the c-kit promoterDepositorAvailable SinceJune 10, 2019AvailabilityAcademic Institutions and Nonprofits only -
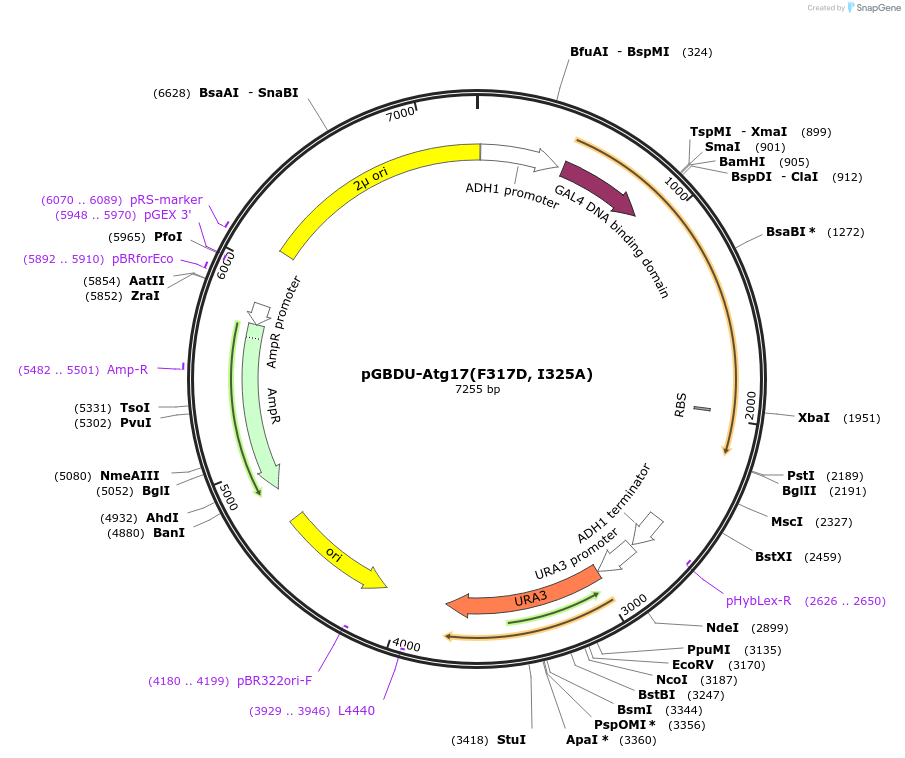
pGBDU-Atg17(F317D, I325A)
Plasmid#122275PurposeFor Yeast-two-hybrid analysisDepositorInsertATG17
ExpressionYeastMutationPhenylalanine 317 to Aspartic acid; Isoleucine 32…Available SinceApril 2, 2019AvailabilityAcademic Institutions and Nonprofits only -
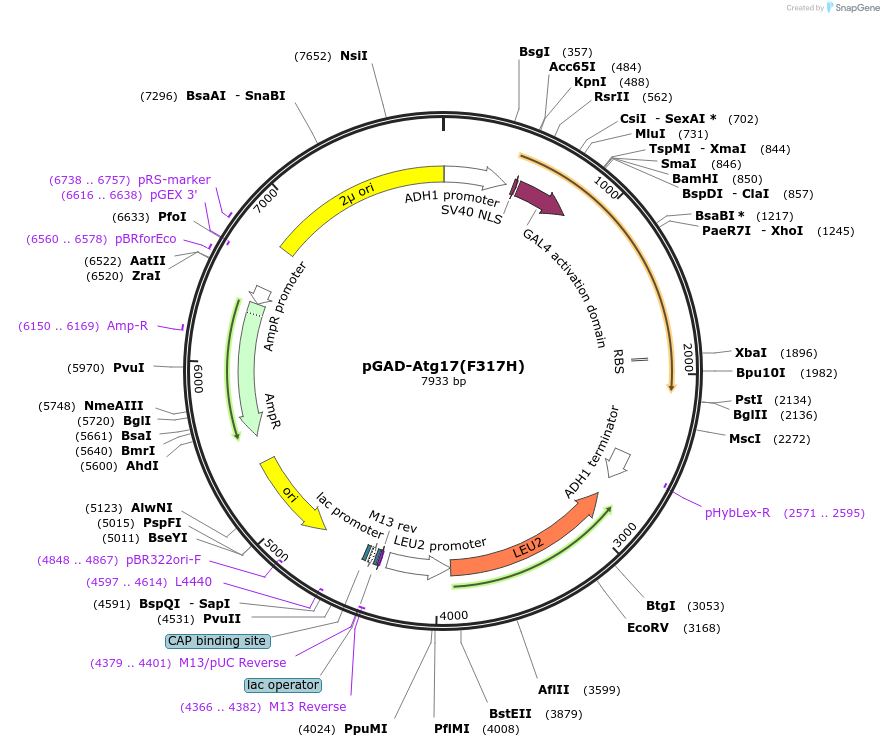
pGAD-Atg17(F317H)
Plasmid#122269PurposeFor Yeast-two-hybrid analysisDepositorInsertATG17
ExpressionYeastMutationPhenylalanine 317 to HistidineAvailable SinceApril 1, 2019AvailabilityAcademic Institutions and Nonprofits only -
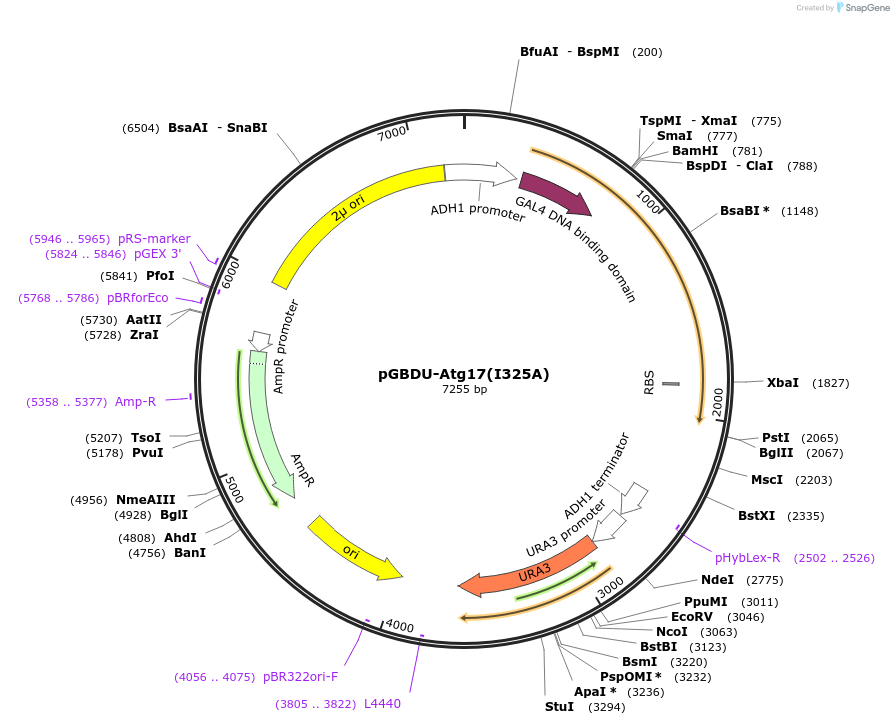
pGBDU-Atg17(I325A)
Plasmid#122274PurposeFor Yeast-two-hybrid analysisDepositorInsertATG17
ExpressionYeastMutationIsoleucine 325 to alanineAvailable SinceApril 1, 2019AvailabilityAcademic Institutions and Nonprofits only -
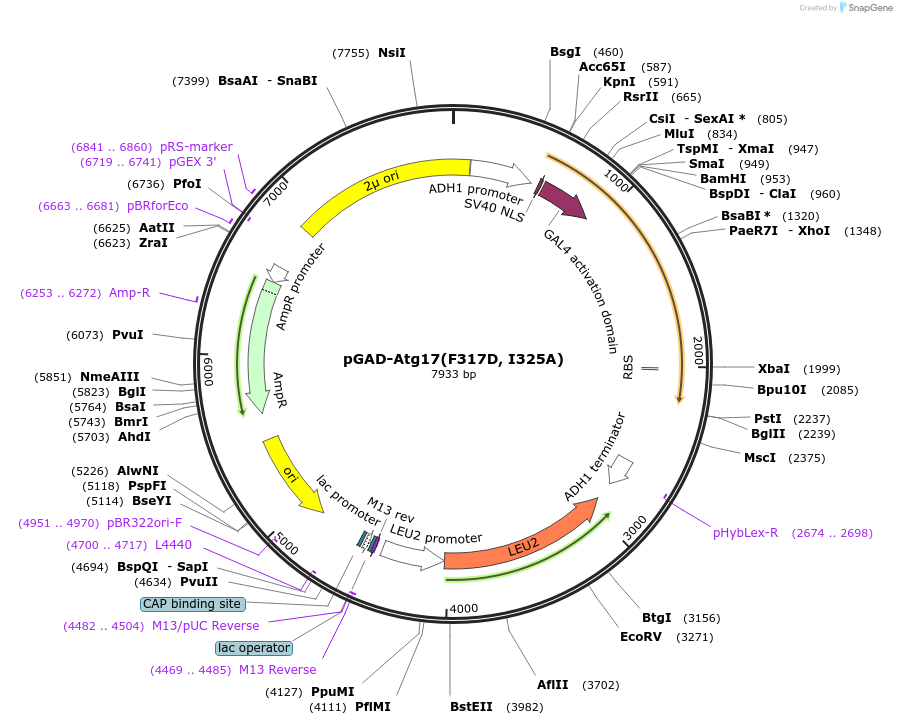
pGAD-Atg17(F317D, I325A)
Plasmid#122273PurposeFor Yeast-two-hybrid analysisDepositorInsertATG17
ExpressionYeastMutationPhenylalanine 317 to Aspartic acid; Isoleucine 32…Available SinceMarch 29, 2019AvailabilityAcademic Institutions and Nonprofits only -
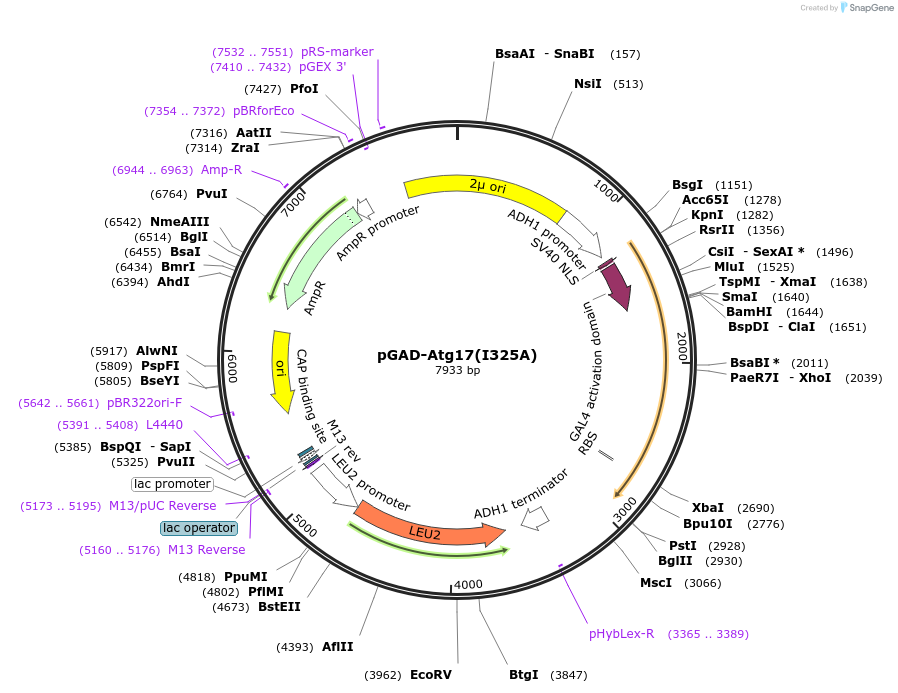
pGAD-Atg17(I325A)
Plasmid#122272PurposeFor Yeast-two-hybrid analysisDepositorInsertATG17
ExpressionYeastMutationIsoleucine 325 to alanineAvailable SinceMarch 26, 2019AvailabilityAcademic Institutions and Nonprofits only -
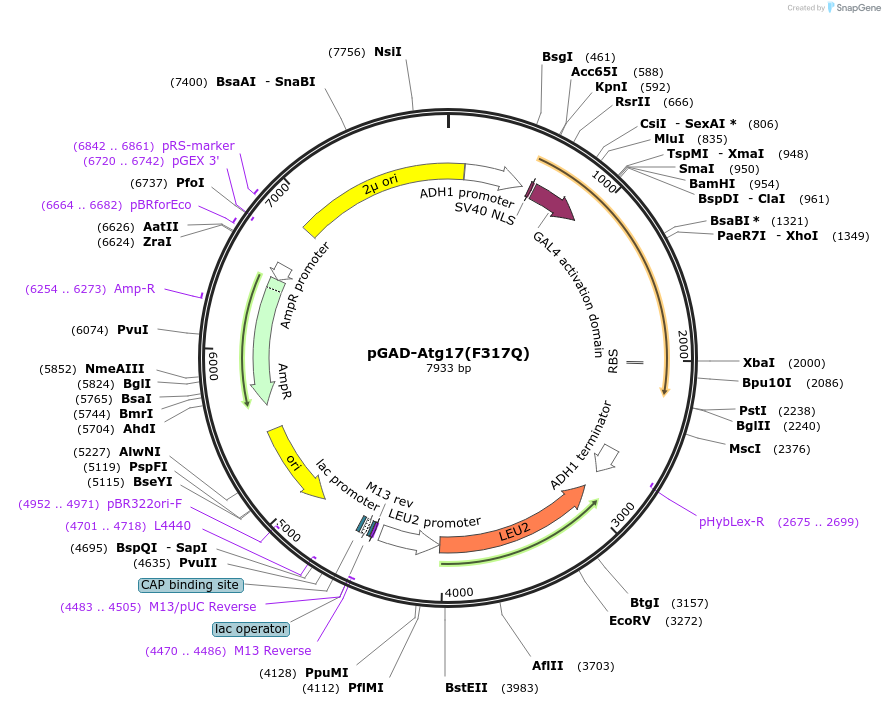
pGAD-Atg17(F317Q)
Plasmid#122271PurposeFor Yeast-two-hybrid analysisDepositorInsertATG17
ExpressionYeastMutationPhenylalanine 317 to glutamineAvailable SinceMarch 26, 2019AvailabilityAcademic Institutions and Nonprofits only -
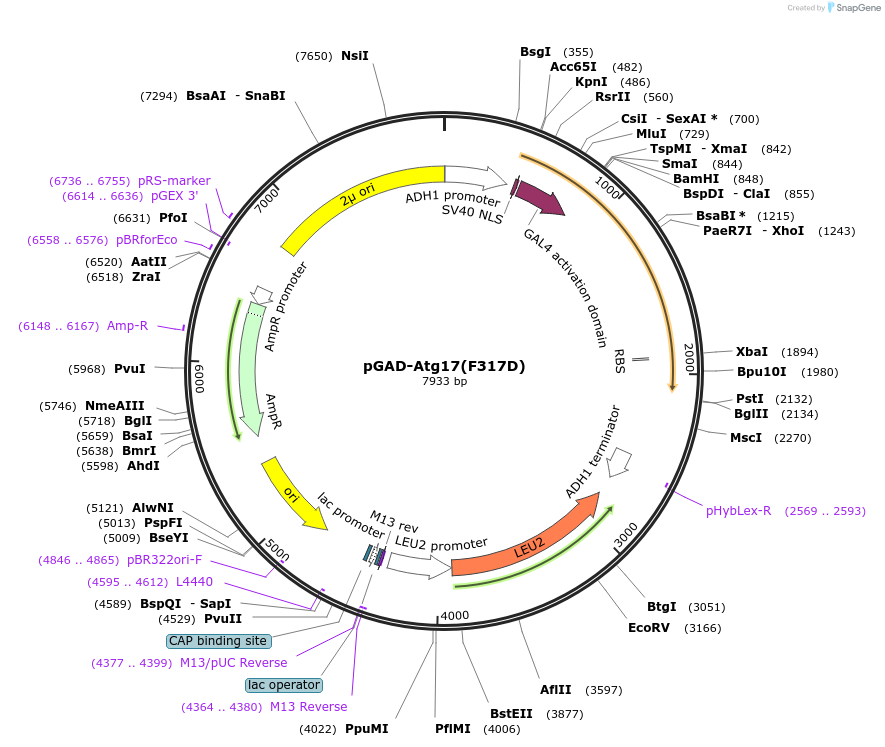
pGAD-Atg17(F317D)
Plasmid#121391PurposeFor Yeast-two-hybrid analysisDepositorInsertATG17
ExpressionYeastMutationPhenylalanine 317 to Aspartic acidPromoterADH1 promoterAvailable SinceMarch 26, 2019AvailabilityAcademic Institutions and Nonprofits only -
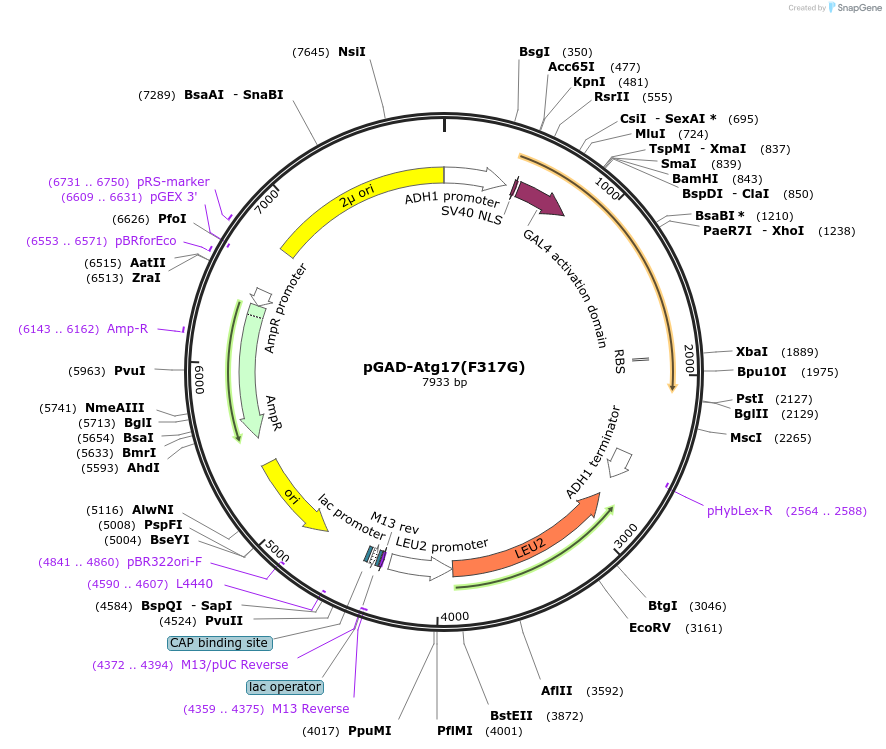
pGAD-Atg17(F317G)
Plasmid#122268PurposeFor Yeast-two-hybrid analysisDepositorInsertATG17
ExpressionYeastMutationPhenylalanine 317 to glycineAvailable SinceMarch 25, 2019AvailabilityAcademic Institutions and Nonprofits only -
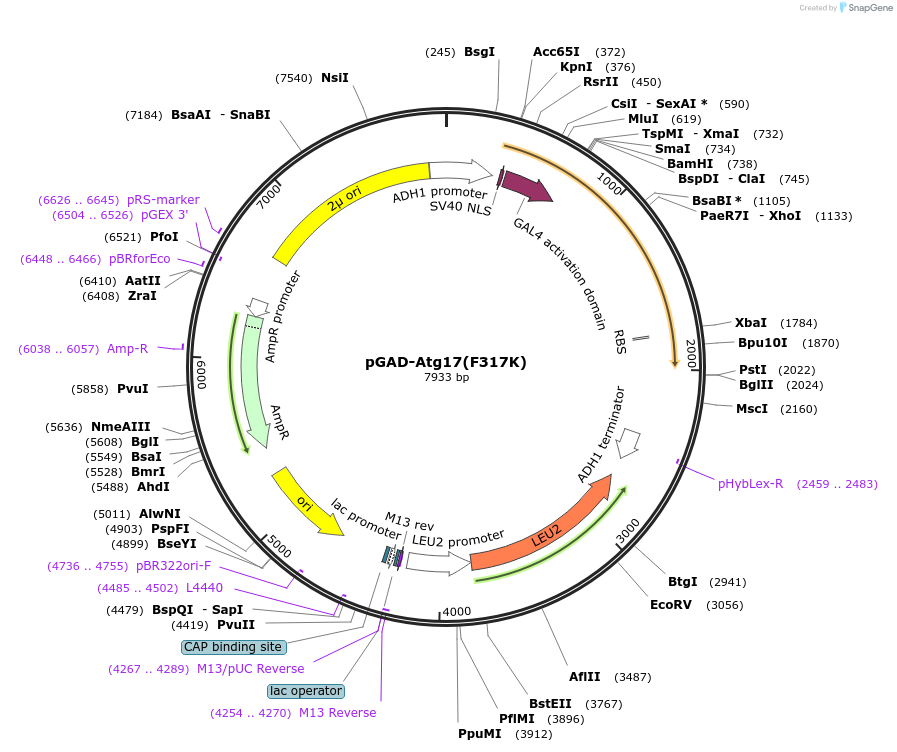
pGAD-Atg17(F317K)
Plasmid#122270PurposeFor Yeast-two-hybrid analysisDepositorInsertATG17
ExpressionYeastMutationPhenylalanine 317 to lysineAvailable SinceMarch 25, 2019AvailabilityAcademic Institutions and Nonprofits only -
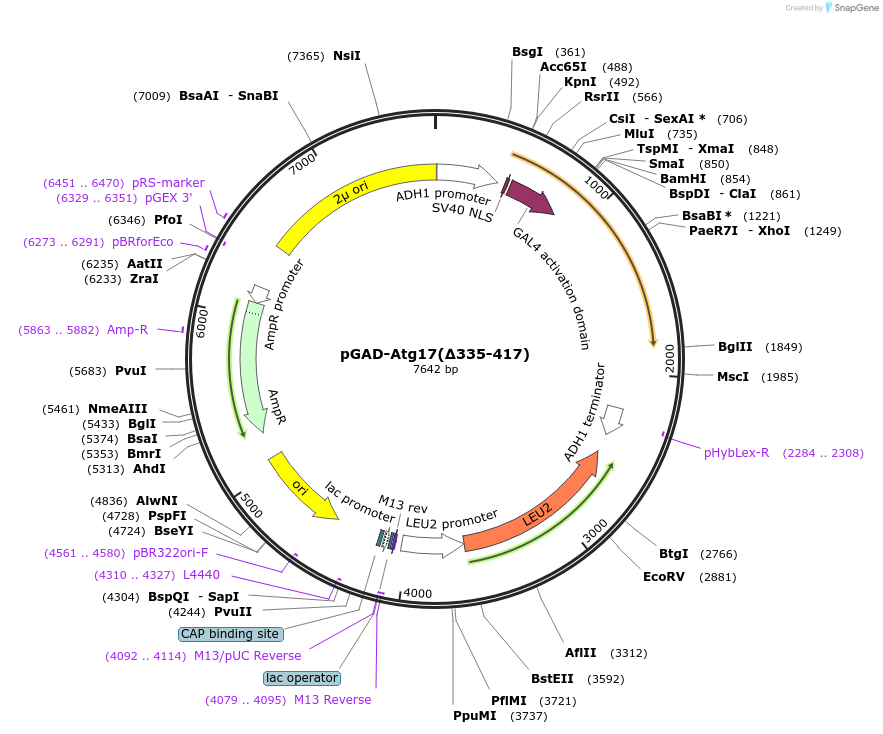
pGAD-Atg17(∆335-417)
Plasmid#121525PurposeFor Yeast-two-hybrid analysisDepositorInsertATG17
ExpressionYeastMutationdeleted amino acids 335-417*Available SinceMarch 22, 2019AvailabilityAcademic Institutions and Nonprofits only -
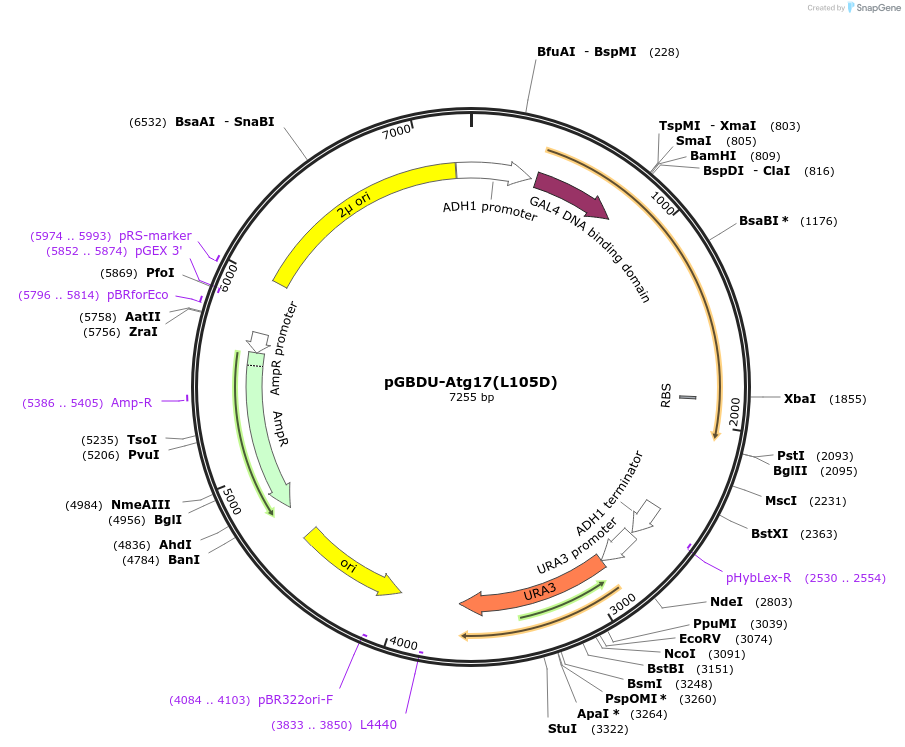
pGBDU-Atg17(L105D)
Plasmid#121395PurposeFor Yeast-two-hybrid analysisDepositorInsertATG17
ExpressionYeastMutationLeucine 105 to Aspartic acidPromoterADH1 promoterAvailable SinceMarch 14, 2019AvailabilityAcademic Institutions and Nonprofits only -
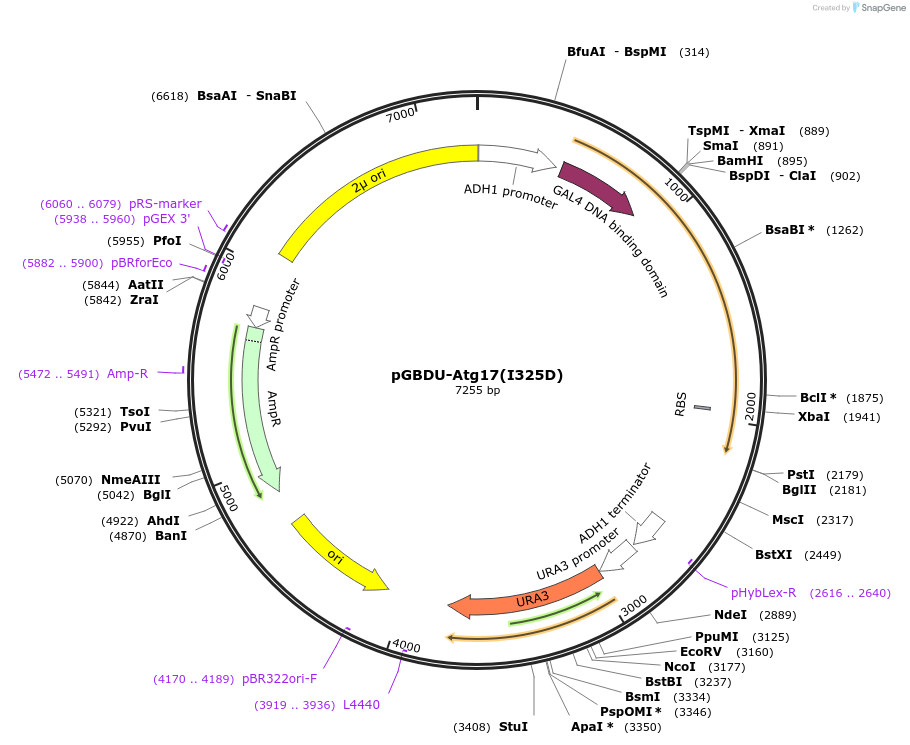
pGBDU-Atg17(I325D)
Plasmid#121397PurposeFor Yeast-two-hybrid analysisDepositorInsertATG17
ExpressionYeastMutationIsoleucine 325 to Aspartic acidPromoterADH1 promoterAvailable SinceMarch 14, 2019AvailabilityAcademic Institutions and Nonprofits only -
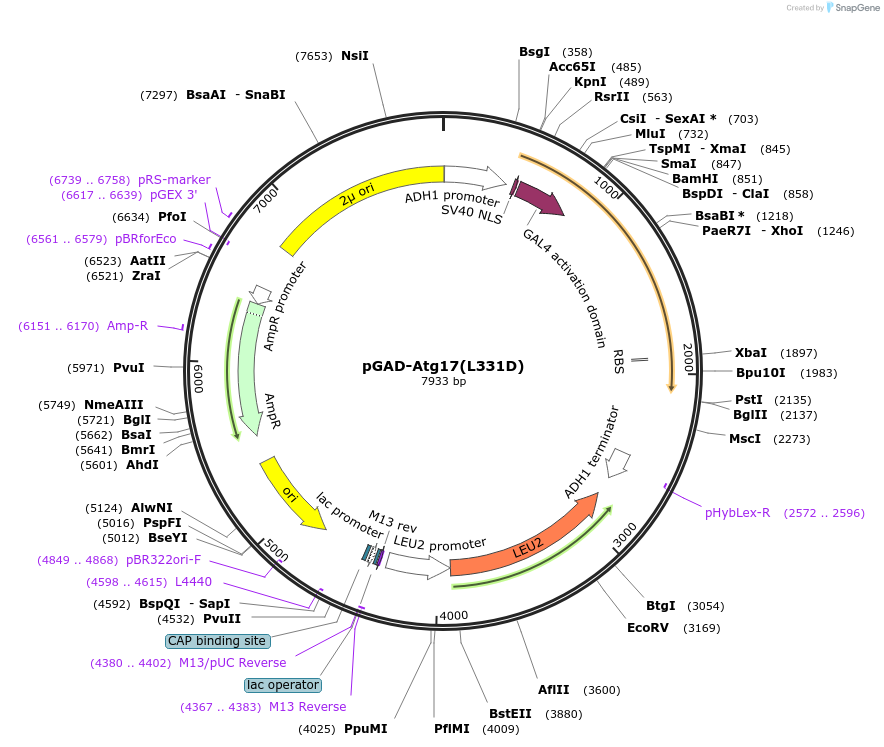
pGAD-Atg17(L331D)
Plasmid#121393PurposeFor Yeast-two-hybrid analysisDepositorInsertATG17
ExpressionYeastMutationLeucine 331 to Aspartic acidPromoterADH1 promoterAvailable SinceMarch 11, 2019AvailabilityAcademic Institutions and Nonprofits only -
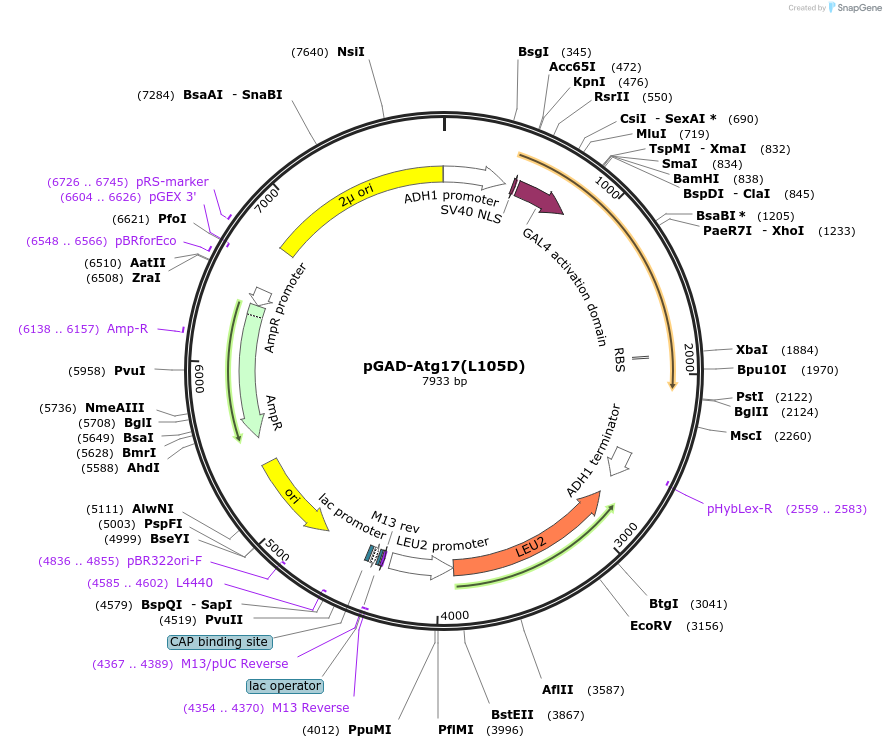
pGAD-Atg17(L105D)
Plasmid#121388PurposeFor Yeast-two-hybrid analysisDepositorInsertATG17
ExpressionYeastMutationLeucine 105 to Aspartic acidPromoterADH1 promoterAvailable SinceMarch 8, 2019AvailabilityAcademic Institutions and Nonprofits only -
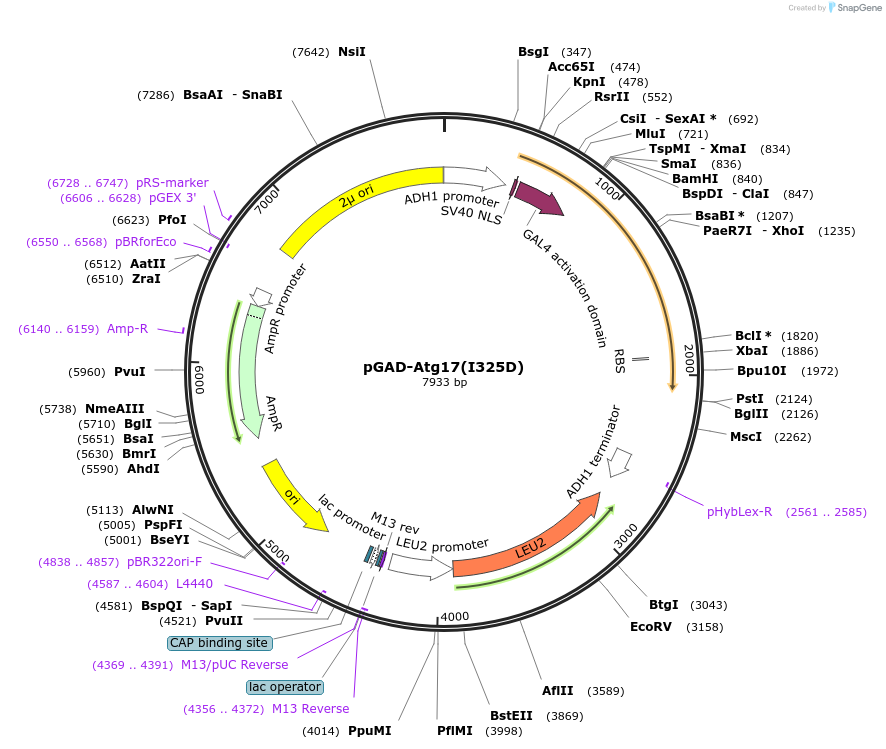
pGAD-Atg17(I325D)
Plasmid#121392PurposeFor Yeast-two-hybrid analysisDepositorInsertATG17
ExpressionYeastMutationIsoleucine 325 to Aspartic acidPromoterADH1 promoterAvailable SinceMarch 8, 2019AvailabilityAcademic Institutions and Nonprofits only -
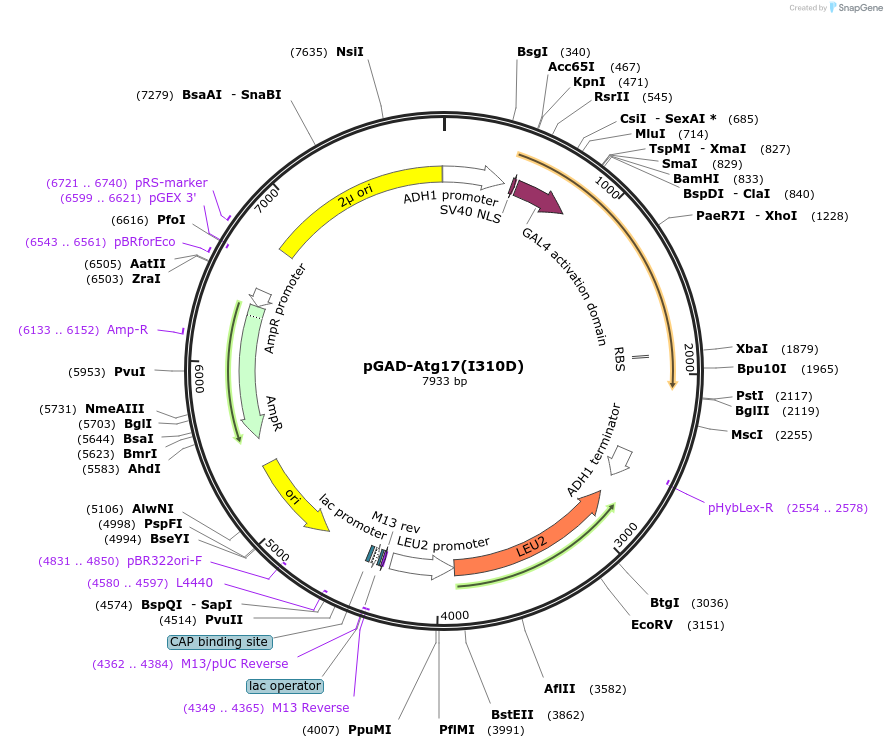
pGAD-Atg17(I310D)
Plasmid#121389PurposeFor Yeast-two-hybrid analysisDepositorInsertATG17
ExpressionYeastMutationIsoleucine 310 to Aspartic acidPromoterADH1 promoterAvailable SinceMarch 8, 2019AvailabilityAcademic Institutions and Nonprofits only -
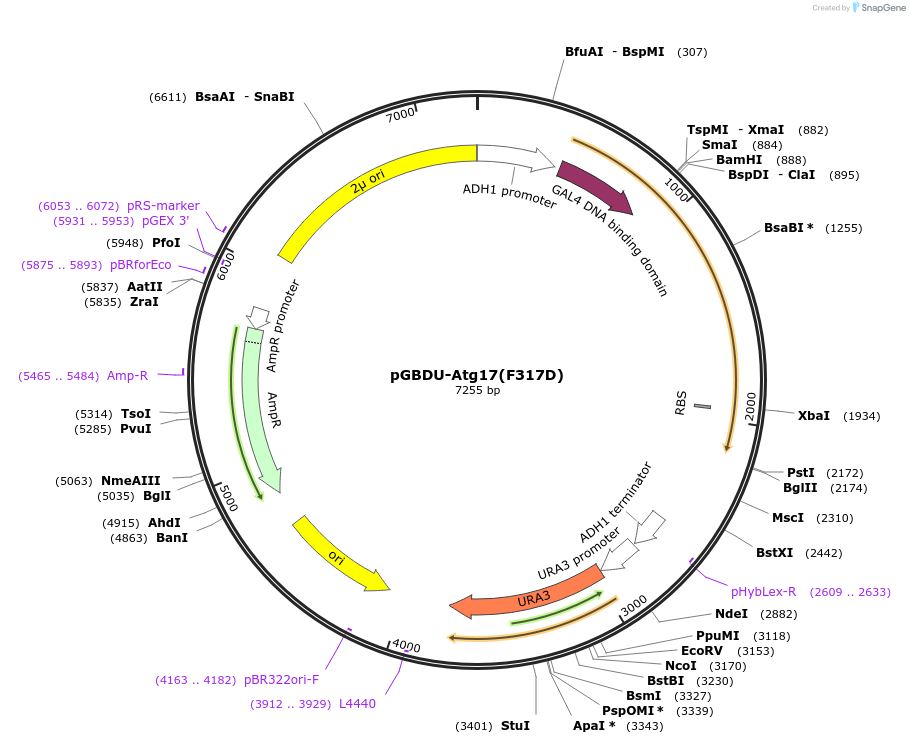
pGBDU-Atg17(F317D)
Plasmid#121396PurposeFor Yeast-two-hybrid analysisDepositorInsertATG17
ExpressionYeastMutationPhenylalanine 317 to Aspartic acidPromoterADH1 promoterAvailable SinceMarch 5, 2019AvailabilityAcademic Institutions and Nonprofits only -
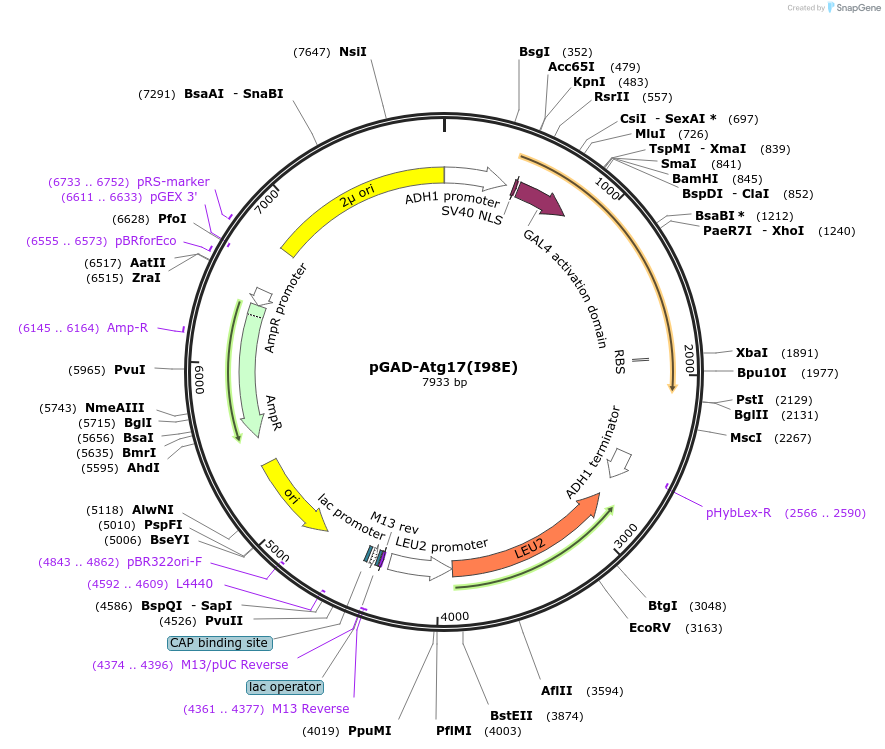
pGAD-Atg17(I98E)
Plasmid#121387PurposeFor Yeast-two-hybrid analysisDepositorInsertATG17
ExpressionYeastMutationIsoleucine 98 to Glutamic acidPromoterADH1 promoterAvailable SinceFeb. 25, 2019AvailabilityAcademic Institutions and Nonprofits only -
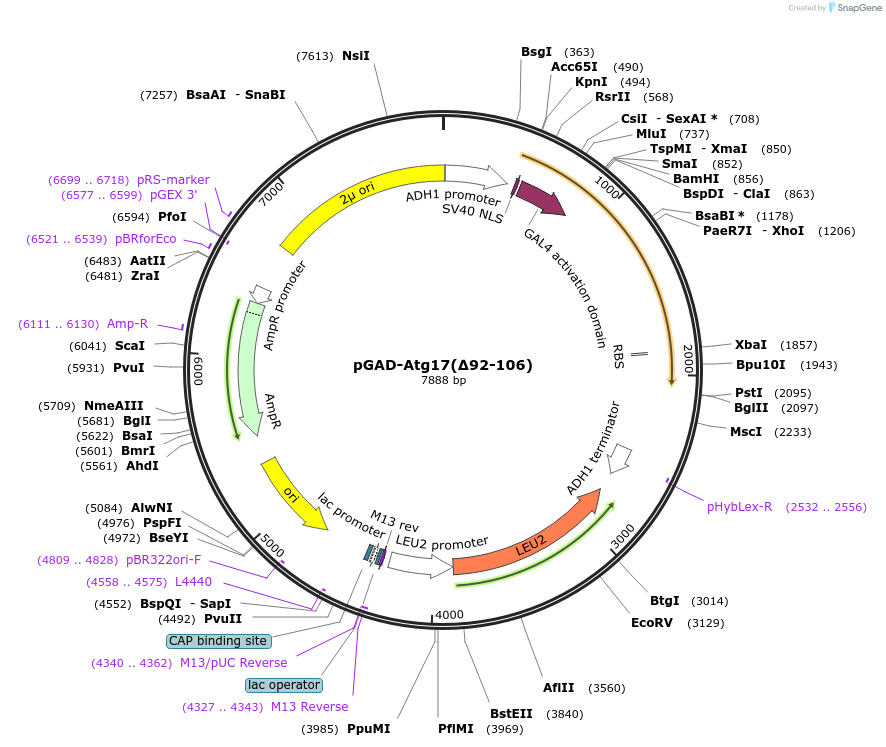
pGAD-Atg17(∆92-106)
Plasmid#121517PurposeFor Yeast-two-hybrid analysisDepositorInsertATG17
ExpressionYeastMutationdeleted amino acids 92-106Available SinceFeb. 25, 2019AvailabilityAcademic Institutions and Nonprofits only -
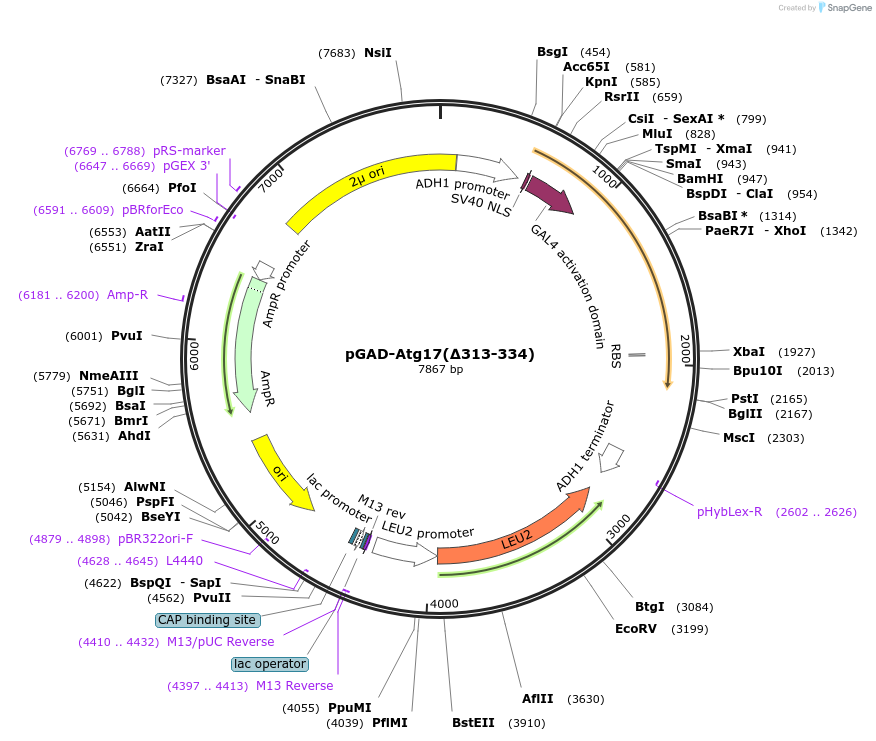
pGAD-Atg17(∆313-334)
Plasmid#121524PurposeFor Yeast-two-hybrid analysisDepositorInsertATG17
ExpressionYeastMutationdeleted amino acids 313-334Available SinceFeb. 25, 2019AvailabilityAcademic Institutions and Nonprofits only