We narrowed to 6,588 results for: siae
-
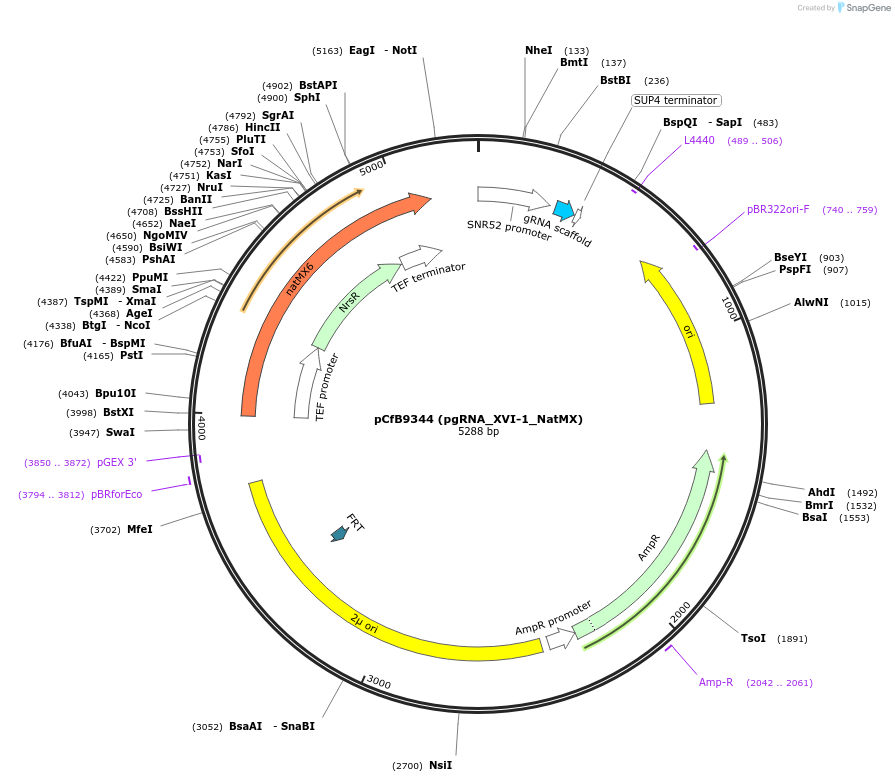
Plasmid#161596PurposeEasyClone-MarkerFree guiding RNA vector to direct Cas9 to cut at site XVI-1DepositorInsertguiding RNA
UseCRISPR and Synthetic BiologyExpressionYeastAvailable SinceAug. 9, 2022AvailabilityAcademic Institutions and Nonprofits only -
KBB438
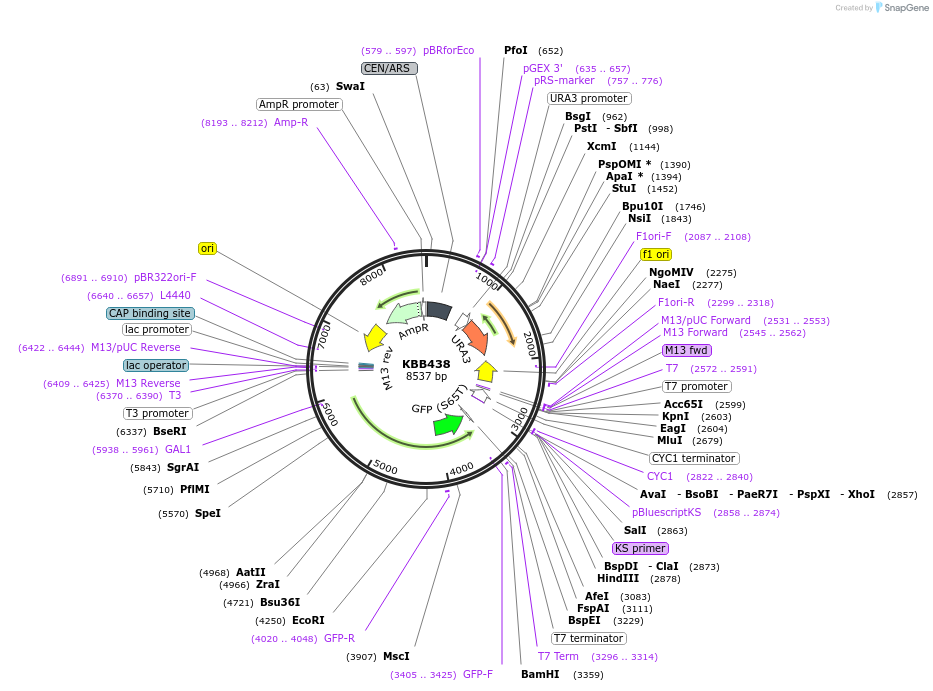
Plasmid#185092PurposeGAL1::Rfa1[1-337, 344-621]-GFP with NLS removedDepositorInsertRFA1
ExpressionYeastMutationRfa1 with amino acids 338-343 deletedPromoterGAL1Available SinceJuly 28, 2022AvailabilityAcademic Institutions and Nonprofits only -
KBB372
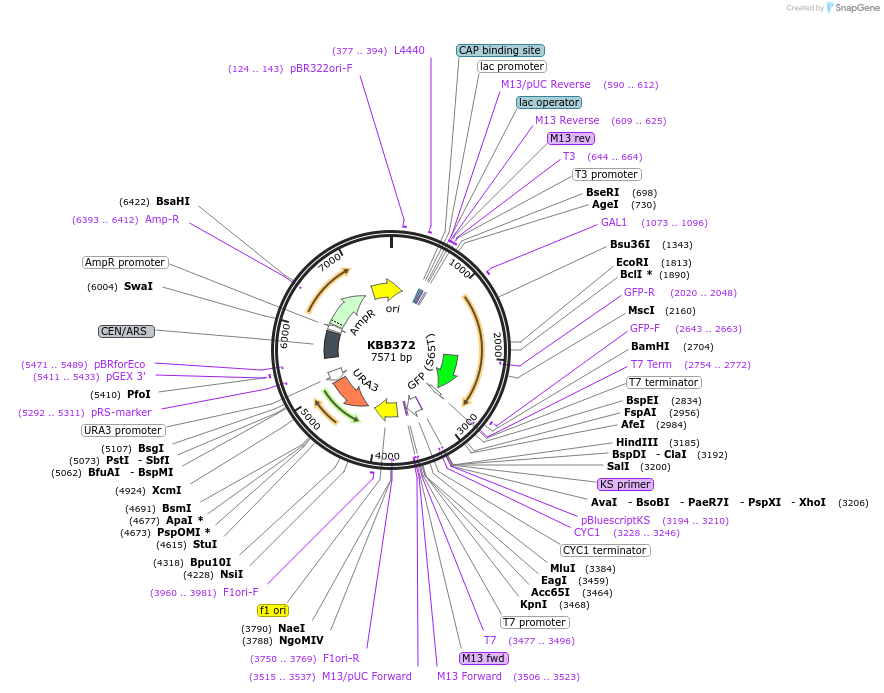
Plasmid#185084PurposeGAL::Rfa1[330-621]-GFP under control of GAL1 promoterDepositorInsertRFA1
TagsGFPExpressionYeastMutationRfa1 amino acids 330-621PromoterGAL1Available SinceJuly 28, 2022AvailabilityAcademic Institutions and Nonprofits only -
pCSN071
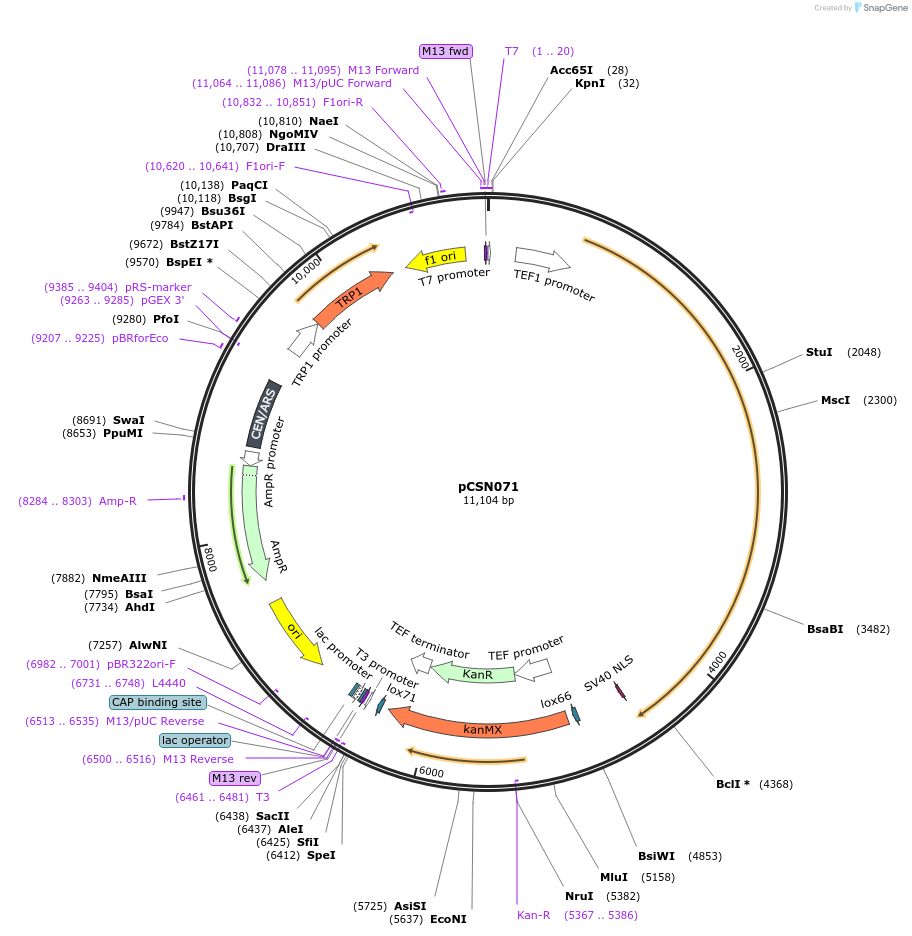
Plasmid#166729PurposeSingle copy yeast plasmid expressing DNase-dead dLbCas12a E925A fused to Mxi1 and NLS, codon optimized for Saccharomyces cerevisiaeDepositorInsertdCas12a-Mxi1-NLS
UseCRISPRTagsMxi1 and SV40 NLSExpressionYeastMutationE925APromoterS. cerevisiae TEF1Available SinceJune 10, 2021AvailabilityAcademic Institutions and Nonprofits only -
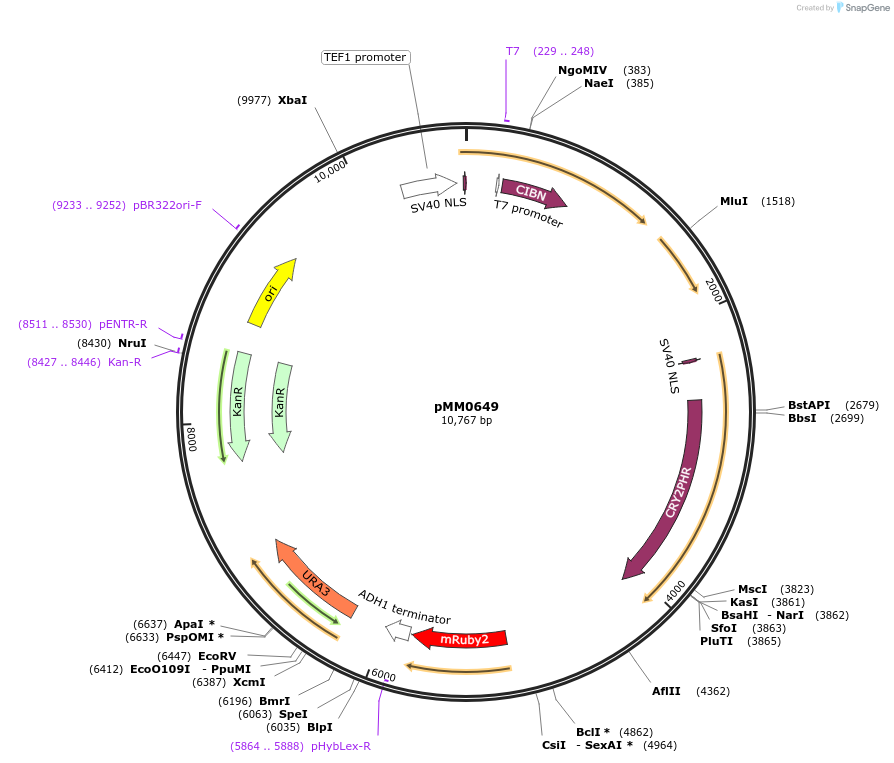
pMM0649
Plasmid#129010PurposeControl for optogenetic machinery (pSTRONG-VP16-CIB1/pMEDIUM-ZCRY2PHR) integration with pWEAK-mRUBY2 reporterDepositorInsert5' URA3-pTEF1-SV40NLS-VP16-CIB1-tENO1-pRPL18B-SV40NLS-ZiF268DBD-CRY2PHR-tSSA1-pREV1-mRUBY2-tADH1-URA3 URA3 3'-KanR-ColE1
ExpressionYeastAvailable SinceFeb. 3, 2020AvailabilityAcademic Institutions and Nonprofits only -
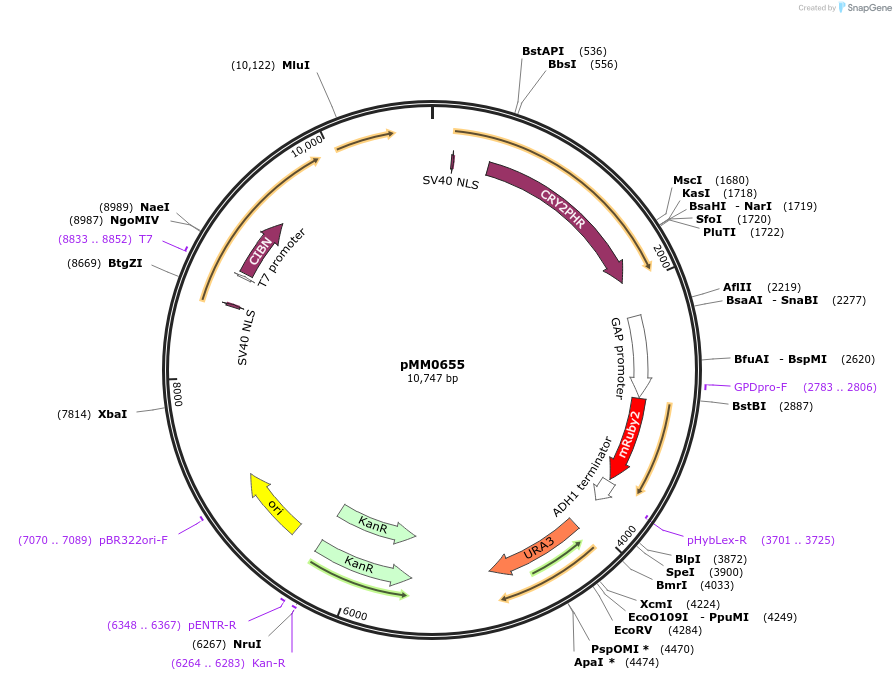
pMM0655
Plasmid#129016PurposeControl for optogenetic machinery (pMEDIUM-VP16-CIB1/pMEDIUM-ZCRY2PHR) integration with pSTRONG-mRUBY2 reporterDepositorInsert5' URA3-pRPL18B-SV40NLS-VP16-CIB1-tENO1-pRPL18B-SV40NLS-ZCRY2PHR-tSSA1-pTDH3-mRUBY2-tADH1-URA3 URA3 3'-KanR-ColE1
ExpressionYeastAvailable SinceOct. 29, 2019AvailabilityAcademic Institutions and Nonprofits only -
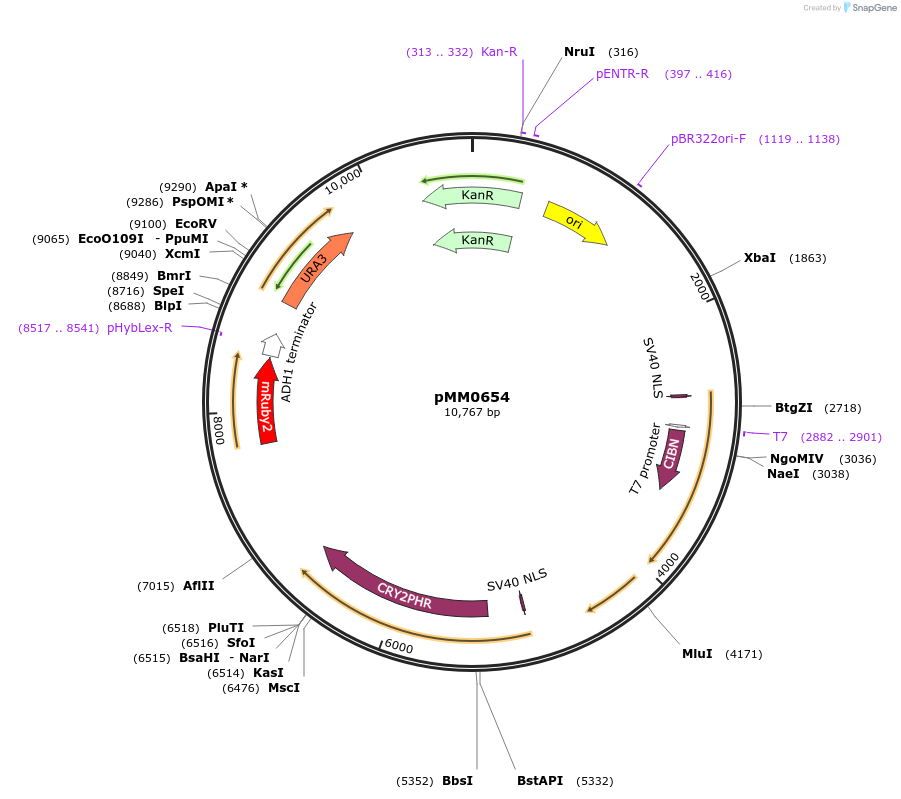
pMM0654
Plasmid#129015PurposeControl for optogenetic machinery (pMEDIUM-VP16-CIB1/pMEDIUM-ZCRY2PHR) integration with pMEDIUM-mRUBY2 reporterDepositorInsert5' URA3-pRPL18B-SV40NLS-VP16-CIB1-tENO1-pRPL18B-SV40NLS-ZCRY2PHR-tSSA1-pRPL18B-mRUBY2-tADH1-URA3 URA3 3'-KanR-ColE1
ExpressionYeastAvailable SinceOct. 16, 2019AvailabilityAcademic Institutions and Nonprofits only -
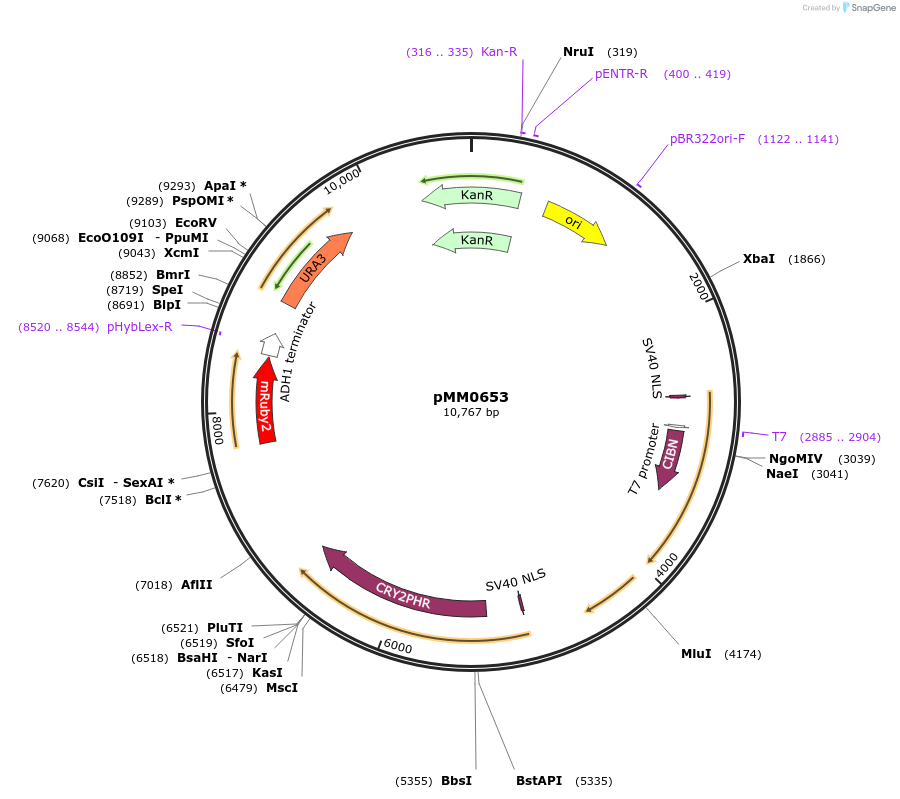
pMM0653
Plasmid#129014PurposeControl for optogenetic machinery (pMEDIUM-VP16-CIB1/pMEDIUM-ZCRY2PHR) integration with pWEAK-mRUBY2 reporterDepositorInsert5' URA3-pRPL18B-SV40NLS-VP16-CIB1-tENO1-pRPL18B-SV40NLS-ZCRY2PHR-tSSA1-pREV1-mRUBY2-tADH1-URA3 URA3 3'-KanR-ColE1
ExpressionYeastAvailable SinceOct. 16, 2019AvailabilityAcademic Institutions and Nonprofits only -
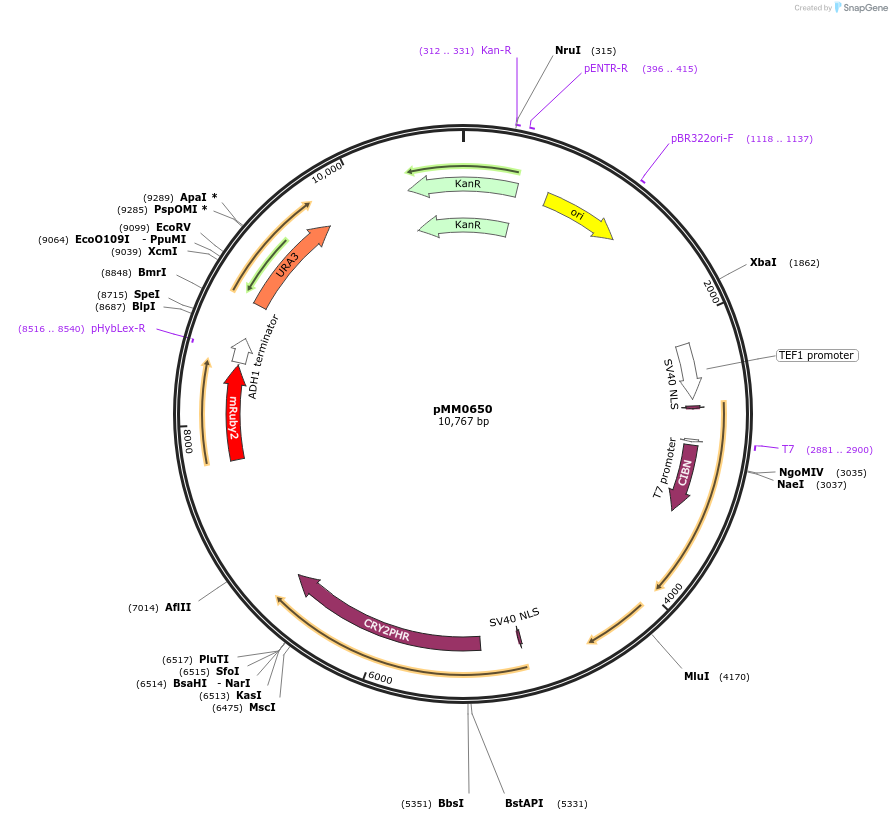
pMM0650
Plasmid#129011PurposeControl for optogenetic machinery (pSTRONG-VP16-CIB1/pMEDIUM-ZCRY2PHR) integration with pMEDIUM-mRUBY2 reporterDepositorInsert5' URA3-pTEF1-SV40NLS-VP16-CIB1-tENO1-pRPL18B-SV40NLS-ZCRY2PHR-tSSA1-pRPL18B-mRUBY2-tADH1-URA3 URA3 3'-KanR-ColE1
ExpressionYeastAvailable SinceOct. 16, 2019AvailabilityAcademic Institutions and Nonprofits only -
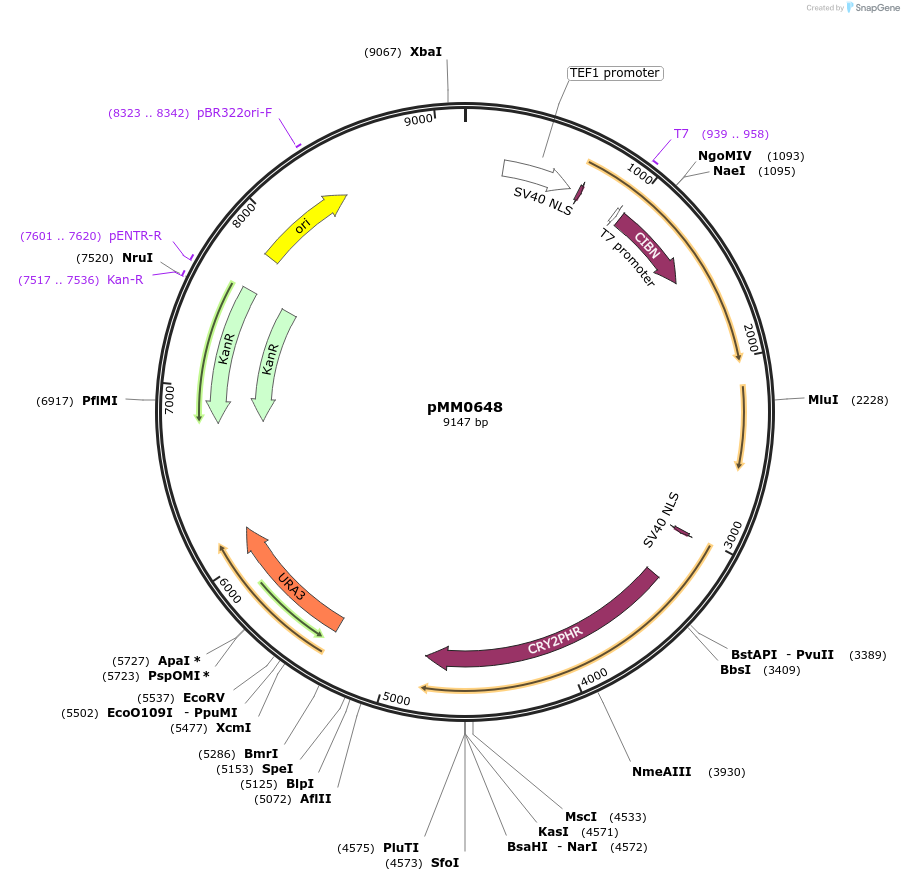
pMM0648
Plasmid#129009PurposeControl for optogenetic machinery (pSTRONG-VP16-CIB1/pMEDIUM-ZCRY2PHR) integration effectsDepositorInsert5' URA3 homology-pTEF1-SV40NLS-VP16-CIB1-tENO1-pRPL18B-SV40NLS-ZCRY2PHR-tSSA1-Spacer-URA3 URA3 3'-homology KanR-ColE1
ExpressionYeastAvailable SinceOct. 16, 2019AvailabilityAcademic Institutions and Nonprofits only -
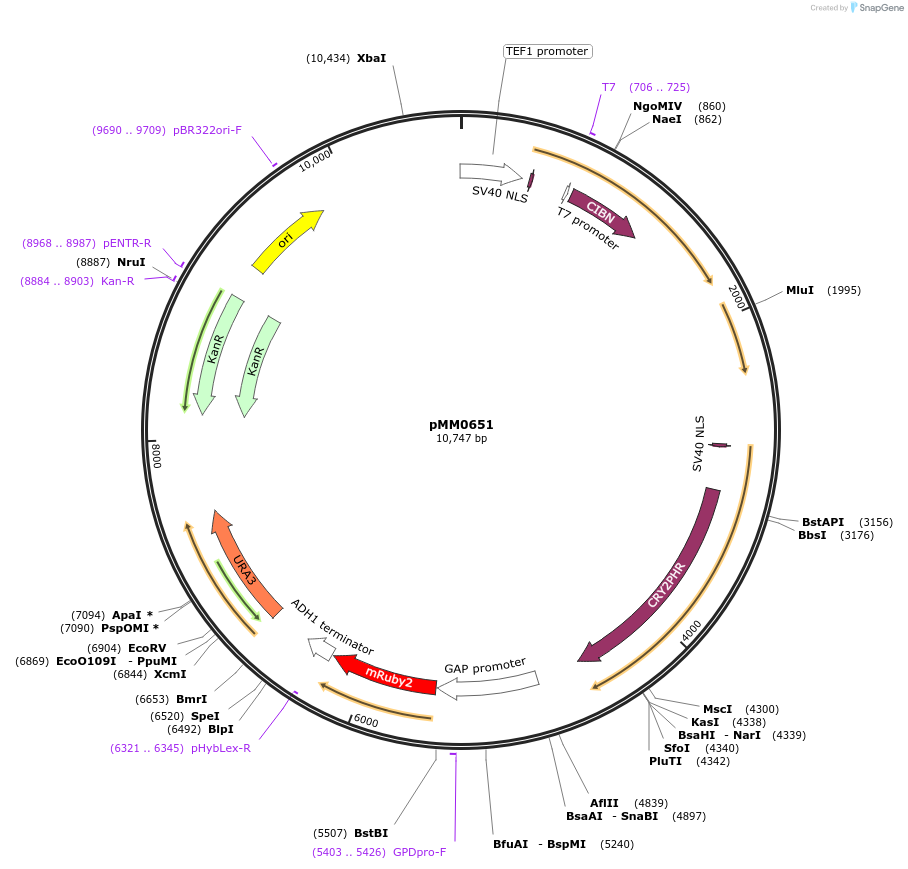
pMM0651
Plasmid#129012PurposeControl for optogenetic machinery (pSTRONG-VP16-CIB1/pMEDIUM-ZCRY2PHR) integration with pSTRONG-mRUBY2 reporterDepositorInsert5' URA3-pTEF1-SV40NLS-VP16-CIB1-tENO1-pRPL18B-SV40NLS-ZCRY2PHR-tSSA1-pTDH3-mRUBY2-tADH1-URA3 URA3 3'-KanR-ColE1
ExpressionYeastAvailable SinceOct. 16, 2019AvailabilityAcademic Institutions and Nonprofits only -
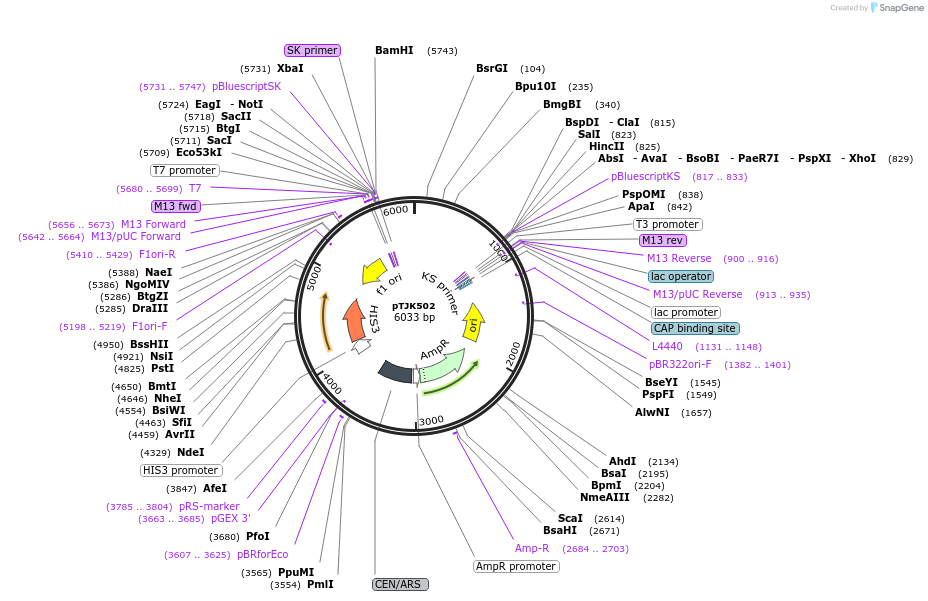
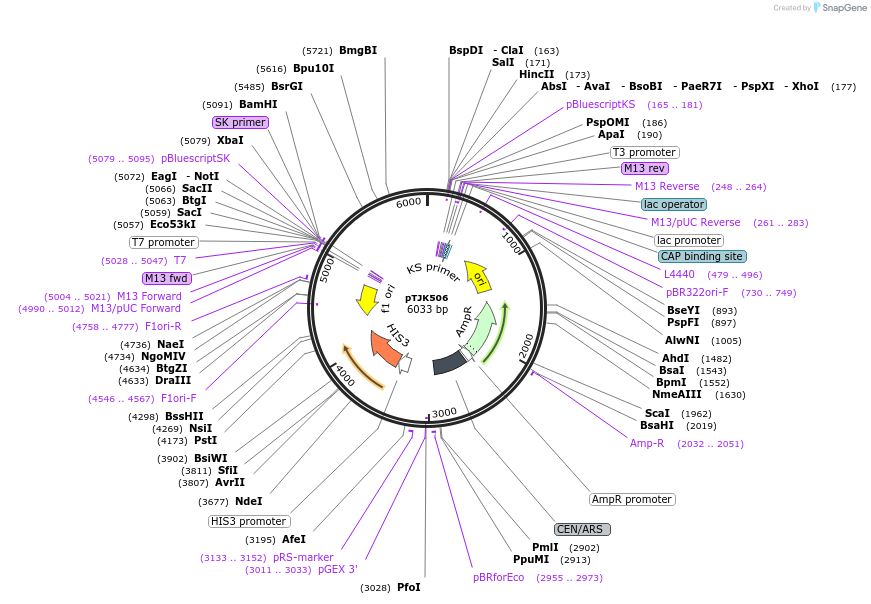
pTJK502
Plasmid#121457PurposeCEN HIS3 CNB1DepositorAvailable SinceMay 2, 2019AvailabilityAcademic Institutions and Nonprofits only -
-
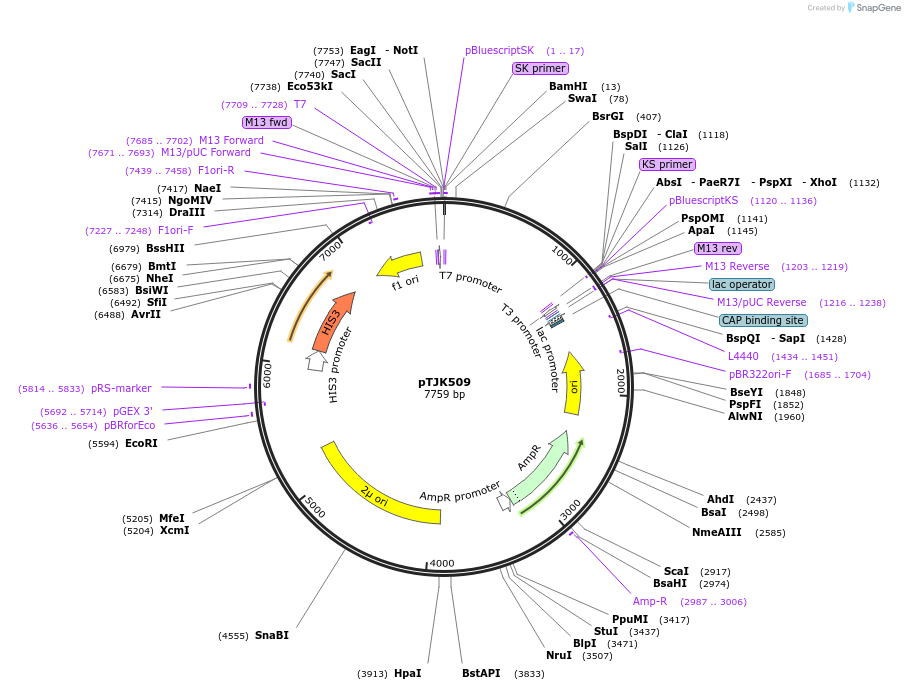
pTJK509
Plasmid#121464Purpose2µm HIS3 CNB1mutEF2DepositorInsertCNB1mutEF2 (CNB1 Budding Yeast)
UseYeast high copy 2 micron vectorMutationChanged aspartic acid 66 to alaninePromoterEndogenous genomic promoter regionAvailable SinceApril 23, 2019AvailabilityAcademic Institutions and Nonprofits only -
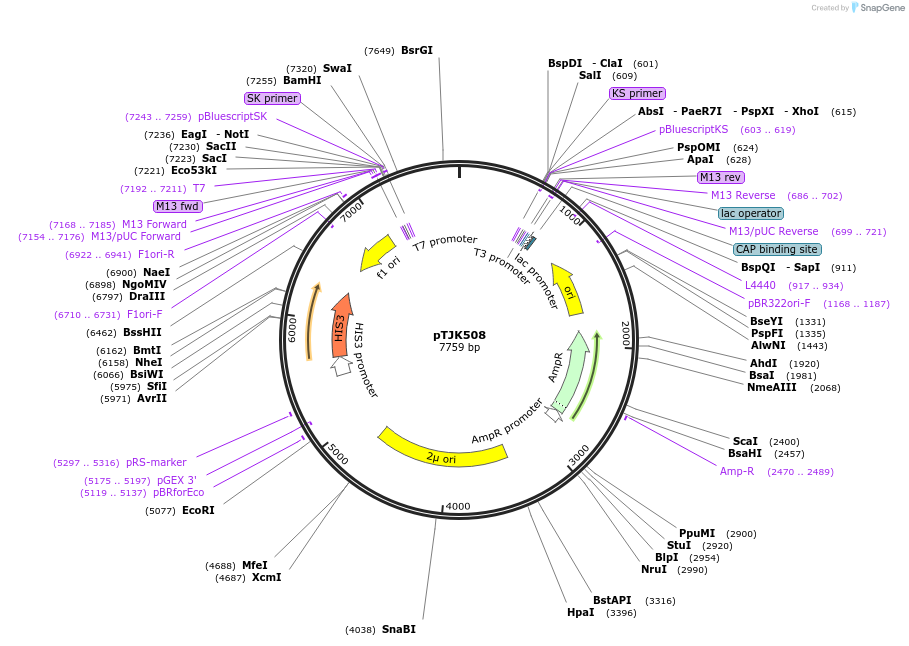
pTJK508
Plasmid#121463Purpose2µm HIS3 CNB1mutEF1DepositorInsertCNB1mutEF1 (CNB1 Budding Yeast)
UseYeast high copy 2 micron vectorMutationChanged aspartic acid 34 to alaninePromoterEndogenous genomic promoter regionAvailable SinceApril 23, 2019AvailabilityAcademic Institutions and Nonprofits only -
-
-
-
-
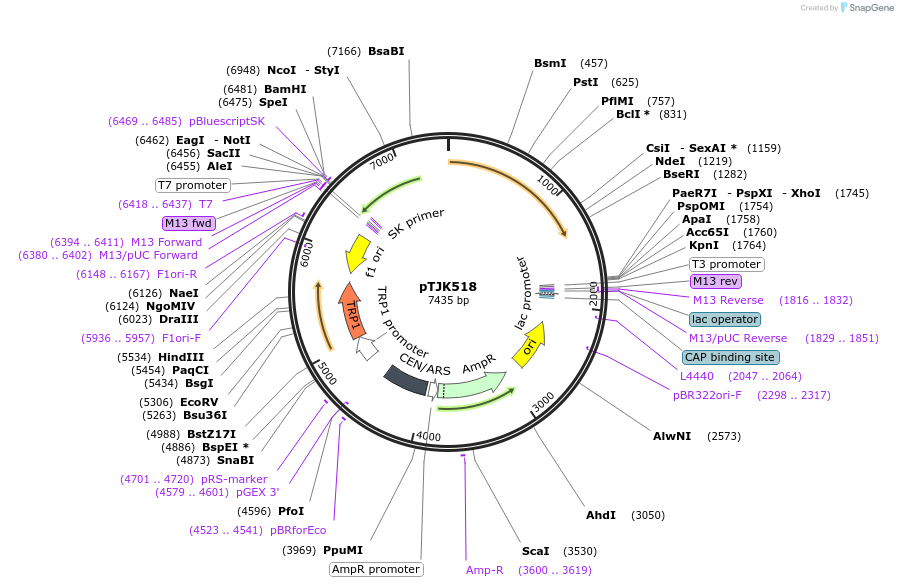
pTJK518
Plasmid#121475PurposeCEN TRP1 CNA1ΔAIDDepositorInsertCNA1deltaAID
UseYeast cen vectorMutationTruncated after amino acid 508PromoterEndogenous genomic promoter regionAvailable SinceApril 23, 2019AvailabilityAcademic Institutions and Nonprofits only