We narrowed to 5,852 results for: SPI
-
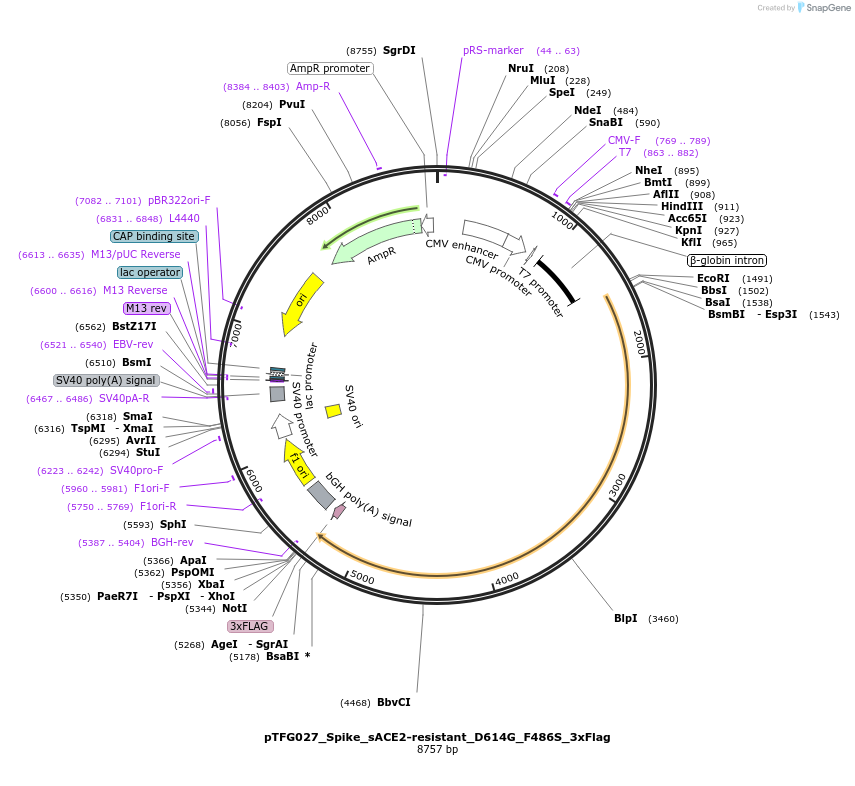
Plasmid#191572PurposeMutated SARS-CoV-2 Spike under CMV with D614G, C-term 19 amino acid truncation, and 3x FLAG. Encodes F486S mutation. For expression and pseudotyping lentiviral particles.DepositorInsertSARS-CoV-2 Spike Protein (S Synthetic)
Tags3x FLAGExpressionMammalianMutationD614G and F486S (confers reduced binding affinity…PromoterCMVAvailable SinceAug. 9, 2024AvailabilityAcademic Institutions and Nonprofits only -
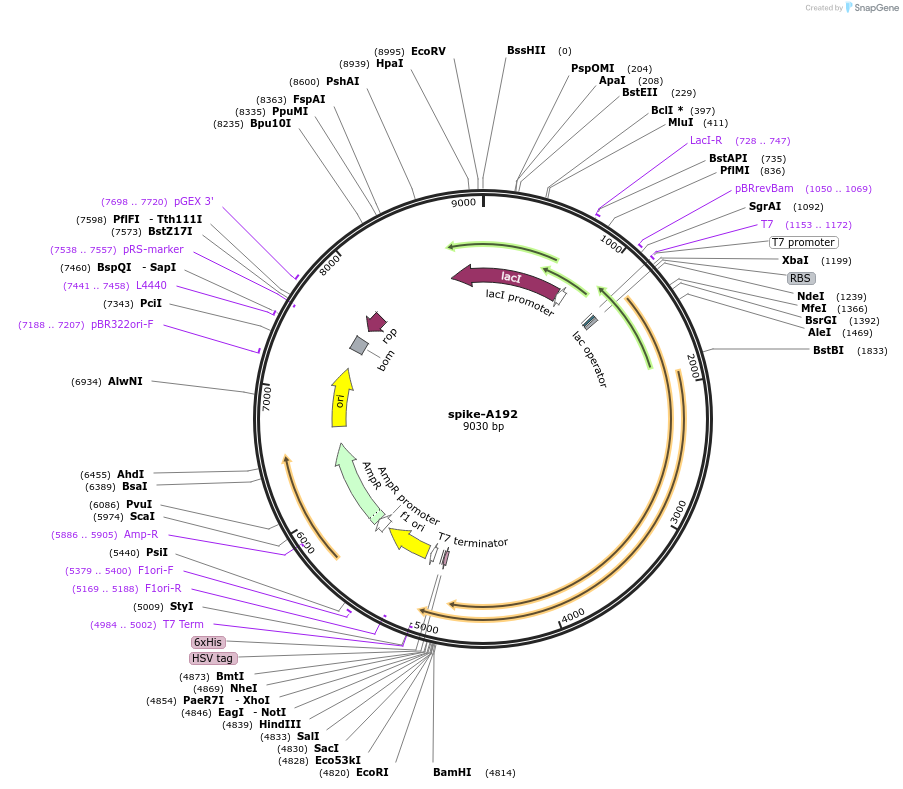
spike-A192
Plasmid#200887PurposeExpresses spike-A192 in E. coli bacterial cellsDepositorAvailable SinceAug. 10, 2023AvailabilityAcademic Institutions and Nonprofits only -
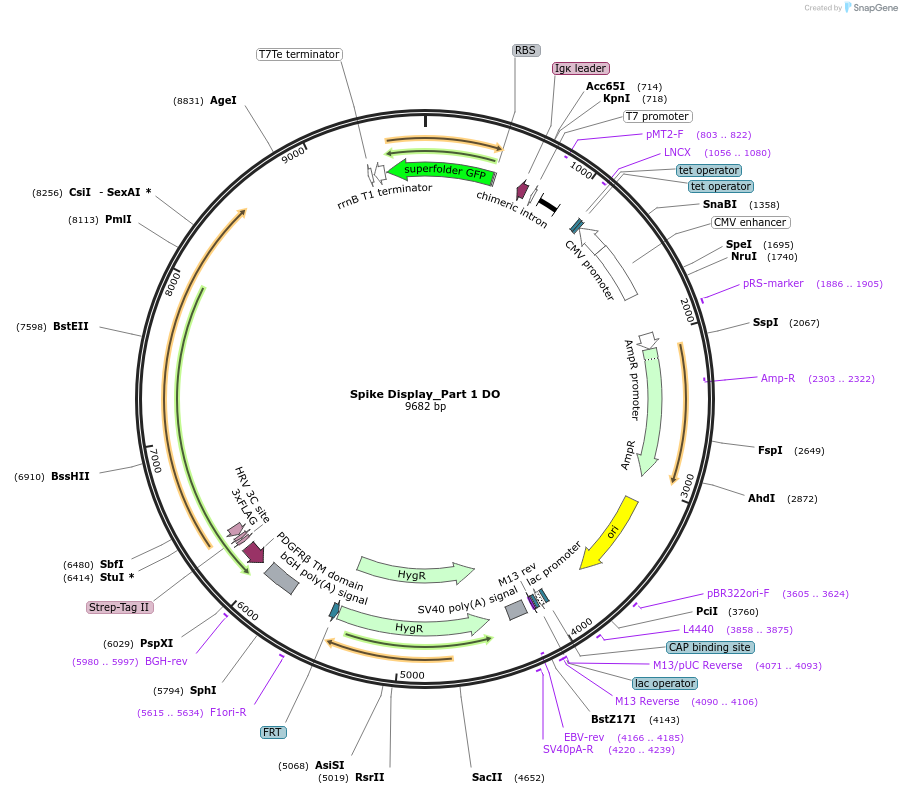
Spike Display_Part 1 DO
Plasmid#172721PurposeEncodes a sfGFP dropout expression cassette in place of Part 1 of HexaPro-D614G spike to be used in the Spike Display systemDepositorInsertSARS-CoV-2 S HexaPro-D614G (S Severe acute respiratory syndrome coronavirus 2)
Tags3X-FLAG; Strep-Tag II; HRV 3C cut site; PDGFR-B T…ExpressionMammalianMutationEctodomain only (AAs 1-1208); 682-685 (furin site…PromoterCMVAvailable SinceOct. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
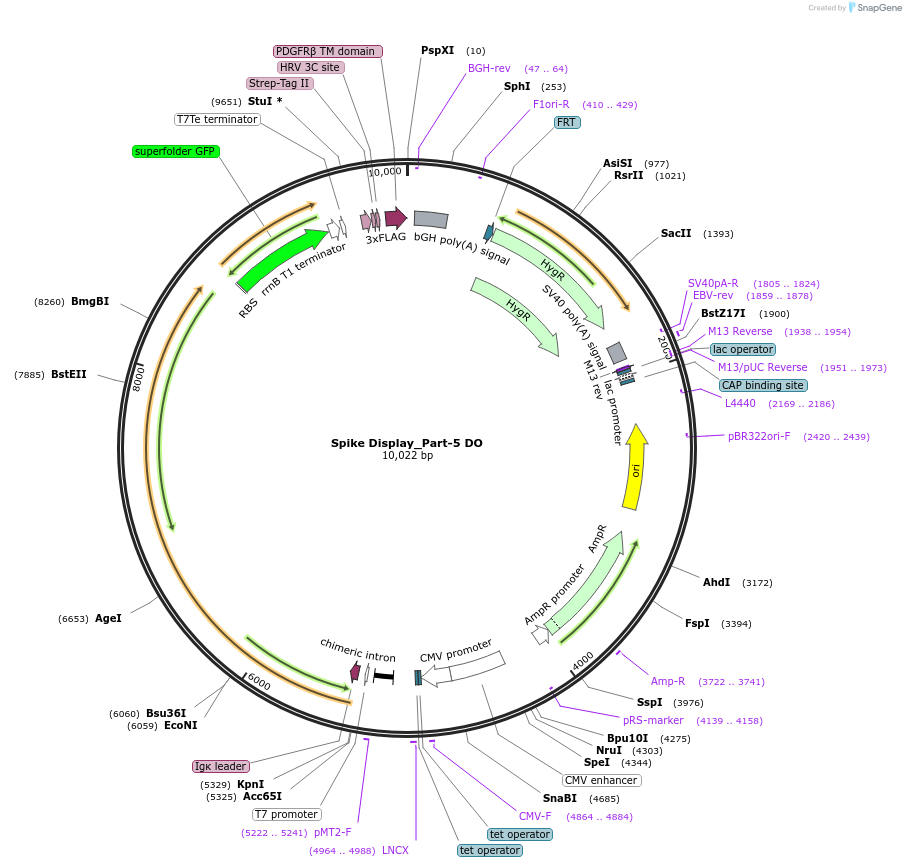
Spike Display_Part-5 DO
Plasmid#172725PurposeEncodes a sfGFP dropout expression cassette in place of Part 5 of HexaPro-D614G spike to be used in the Spike Display systemDepositorInsertSARS-CoV-2 S HexaPro-D614G (S Severe acute respiratory syndrome coronavirus 2)
Tags3X-FLAG; Strep-Tag II; HRV 3C cut site; PDGFR-B T…ExpressionMammalianMutationEctodomain only (AAs 1-1208); 682-685 (furin site…PromoterCMVAvailable SinceAug. 18, 2021AvailabilityAcademic Institutions and Nonprofits only -
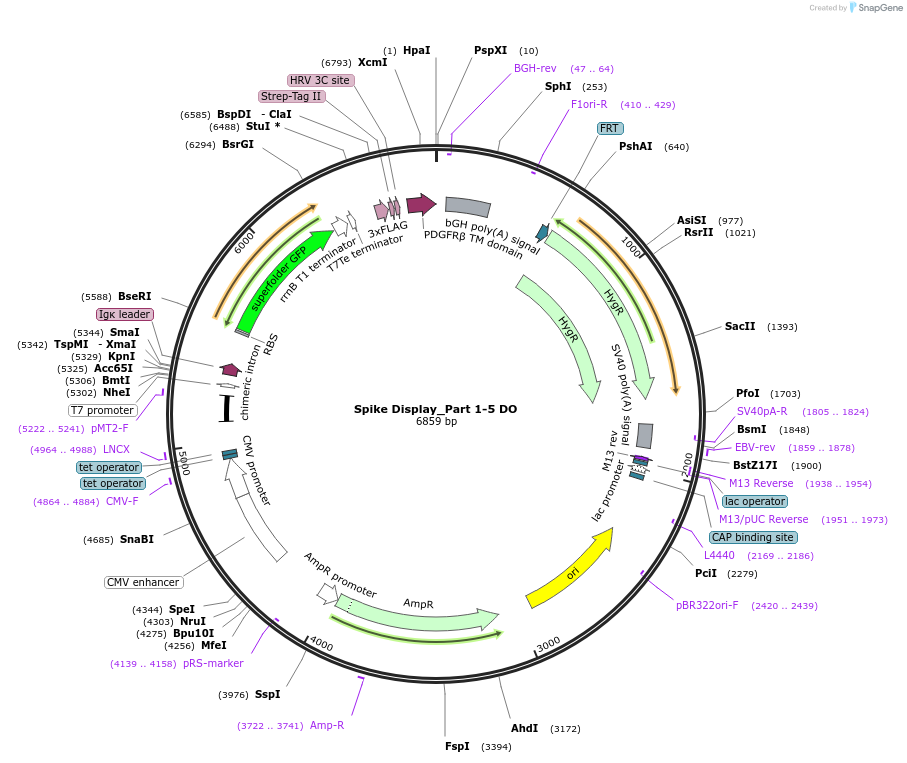
Spike Display_Part 1-5 DO
Plasmid#172726PurposeEncodes a sfGFP dropout expression cassette in place of Parts 1-5 to be used in the Spike Display systemDepositorInsertSARS-CoV-2 S HexaPro-D614G (S Severe acute respiratory syndrome coronavirus 2)
Tags3X-FLAG; Strep-Tag II; HRV 3C cut site; PDGFR-B T…ExpressionMammalianMutationEctodomain only (AAs 1-1208); 682-685 (furin site…PromoterCMVAvailable SinceAug. 11, 2021AvailabilityAcademic Institutions and Nonprofits only -
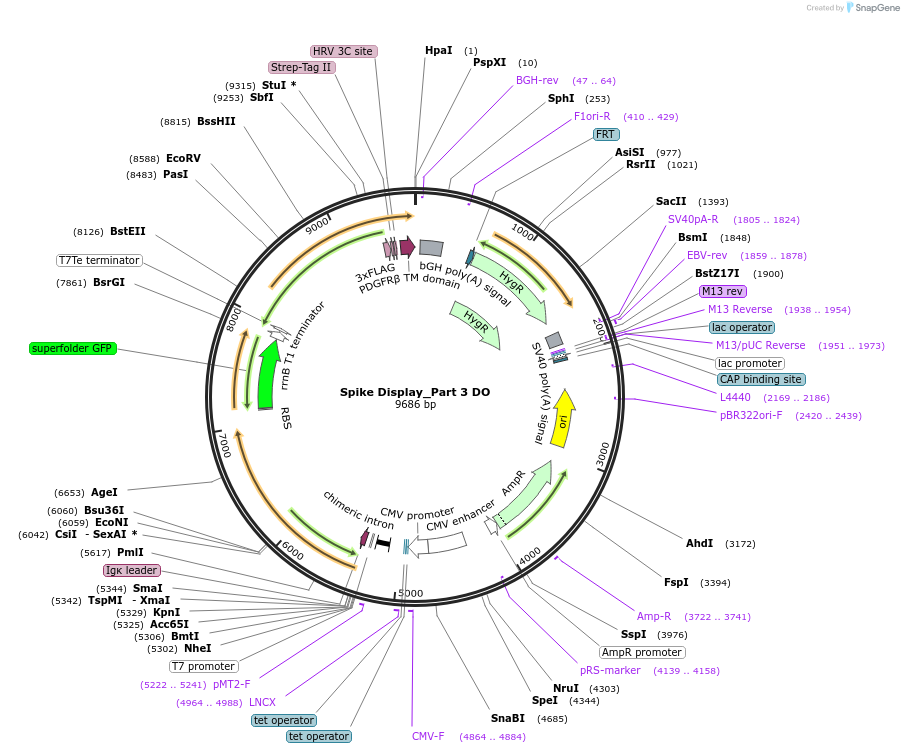
Spike Display_Part 3 DO
Plasmid#172723PurposeEncodes a sfGFP dropout expression cassette in place of Part 3 of HexaPro-D614G spike to be used in the Spike Display systemDepositorInsertSARS-CoV-2 S HexaPro-D614G (S Severe acute respiratory syndrome coronavirus 2)
Tags3X-FLAG; Strep-Tag II; HRV 3C cut site; PDGFR-B T…ExpressionMammalianMutationEctodomain only (AAs 1-1208); 682-685 (furin site…PromoterCMVAvailable SinceAug. 11, 2021AvailabilityAcademic Institutions and Nonprofits only -
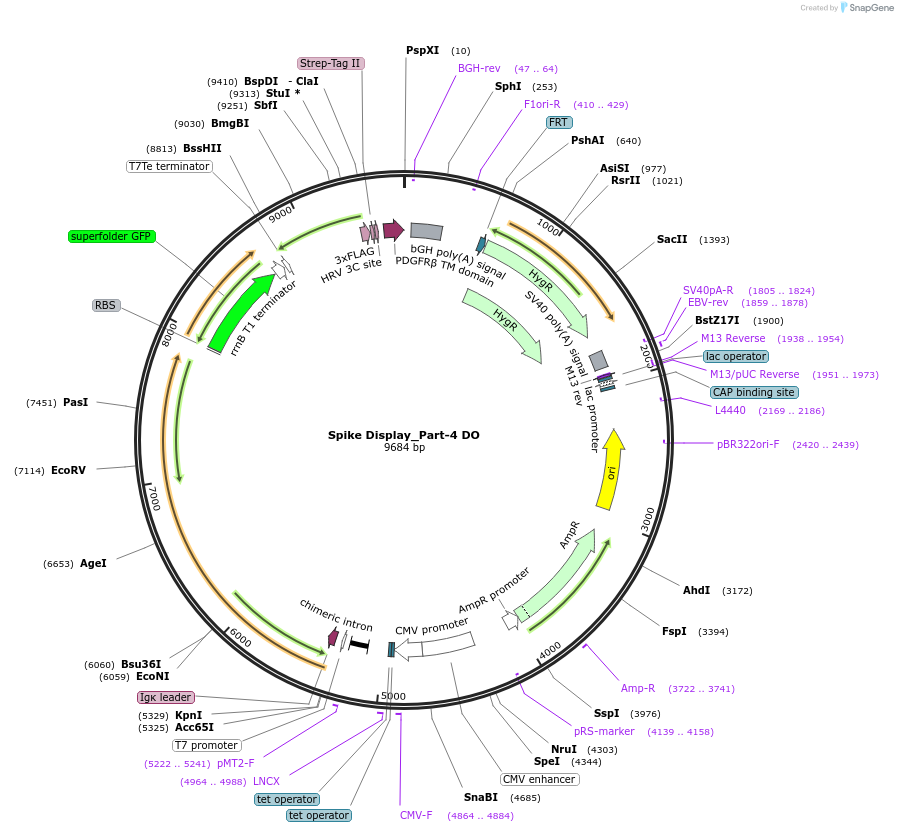
Spike Display_Part-4 DO
Plasmid#172724PurposeEncodes a sfGFP dropout expression cassette in place of Part 4 of HexaPro-D614G spike to be used in the Spike Display systemDepositorInsertSARS-CoV-2 S HexaPro-D614G (S Severe acute respiratory syndrome coronavirus 2)
Tags3X-FLAG; Strep-Tag II; HRV 3C cut site; PDGFR-B T…ExpressionMammalianMutationEctodomain only (AAs 1-1208); 682-685 (furin site…PromoterCMVAvailable SinceAug. 9, 2021AvailabilityAcademic Institutions and Nonprofits only -
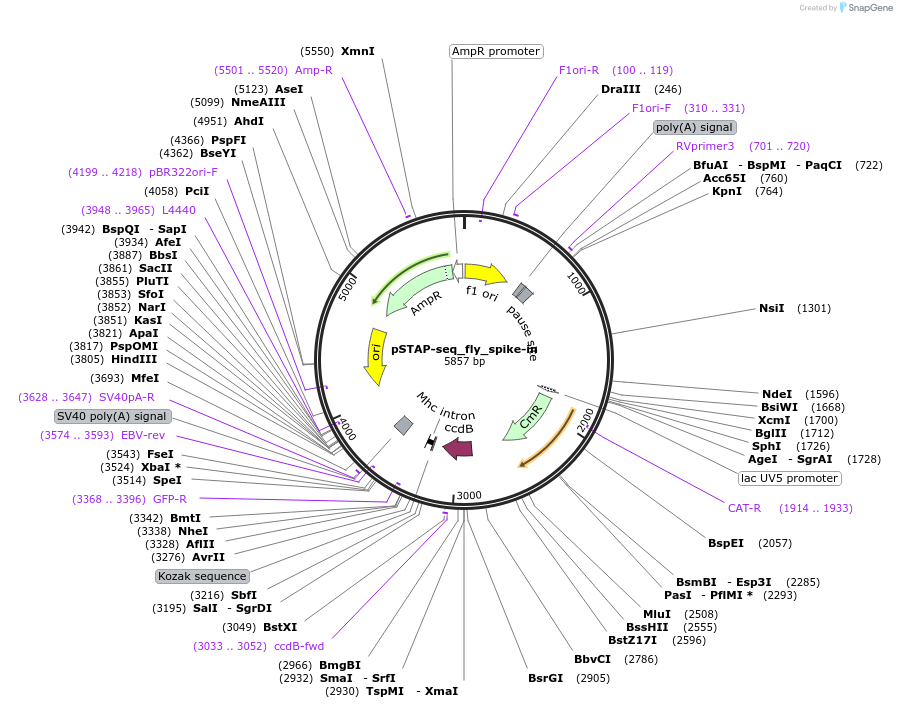
pSTAP-seq_fly_spike-in
Plasmid#125151PurposeVector to measure CP candidates (spike-in) expression by determining the abundance of transcripts originating from the candidate by a D.pse enhancer in fly cellsDepositorInsertD. pseudoobscura peak #47 enhancer
UseStap-seq screening vectorAvailable SinceAug. 1, 2019AvailabilityAcademic Institutions and Nonprofits only -
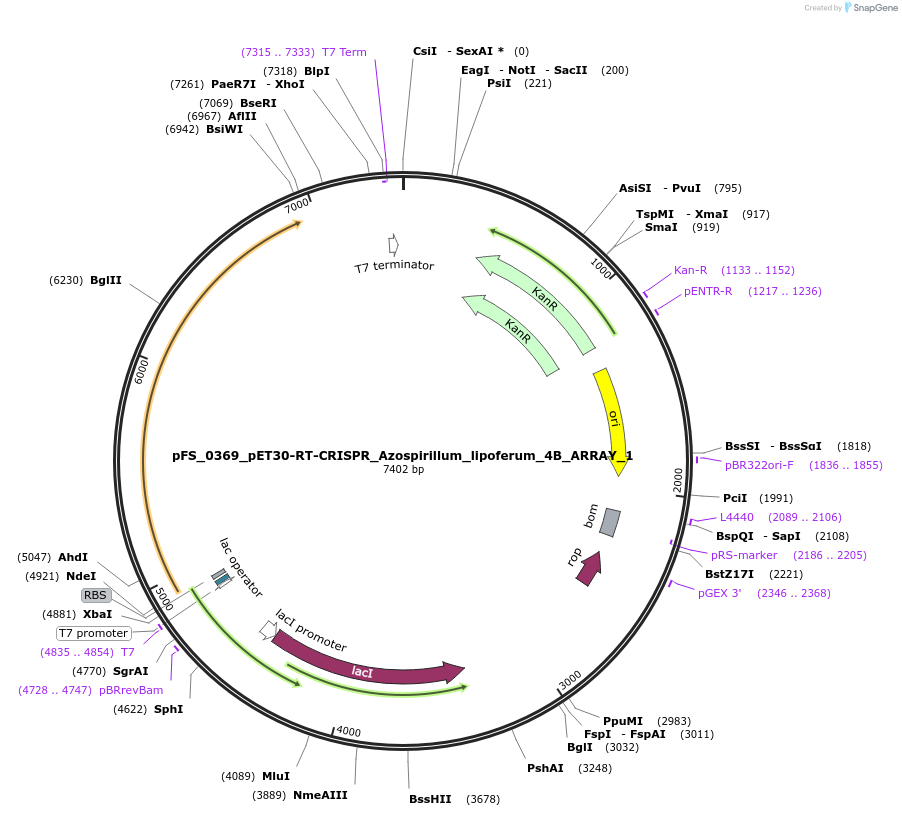
pFS_0369_pET30-RT-CRISPR_Azospirillum_lipoferum_4B_ARRAY_1
Plasmid#116981PurposeExpresses AlRT-Cas1-Cas2 under pT7lac promoter, encodes AlCRISPR array 1, compatible with SENECA acquisition readoutDepositorInsertAzospirillum lipoferum 4B RT-Cas1-Cas2
ExpressionBacterialPromoterT7Available SinceNov. 29, 2018AvailabilityAcademic Institutions and Nonprofits only -
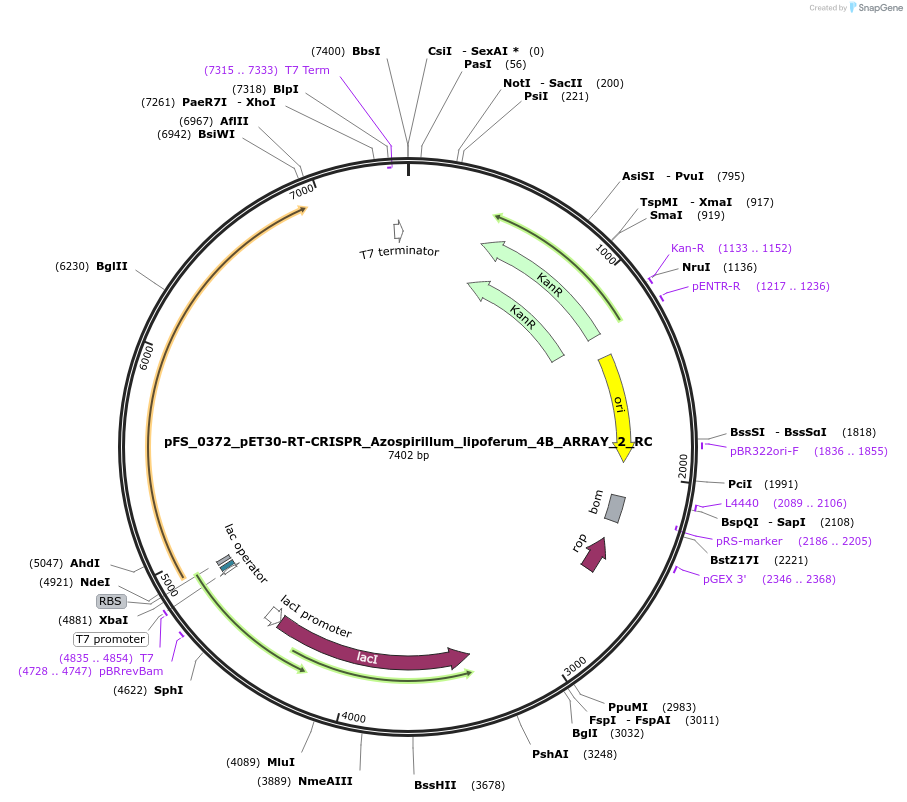
pFS_0372_pET30-RT-CRISPR_Azospirillum_lipoferum_4B_ARRAY_2_RC
Plasmid#116984PurposeExpresses AlRT-Cas1-Cas2 under pT7lac promoter, encodes AlCRISPR array 2 RC, compatible with SENECA acquisition readoutDepositorInsertAzospirillum lipoferum 4B RT-Cas1-Cas2
ExpressionBacterialPromoterT7Available SinceNov. 2, 2018AvailabilityAcademic Institutions and Nonprofits only