We narrowed to 705 results for: bacillus
-
Plasmid#79215PurposeDirected Evolution of Protein-Protein InteractionsDepositorInsertPlacZ-opt (OR1+2) gIII, luxAB; Ppro1 434cI-TnCAD-F3
ExpressionBacterialPromoterPlacZ-opt (OR1+2); Ppro1Available SinceDec. 17, 2018AvailabilityAcademic Institutions and Nonprofits only -
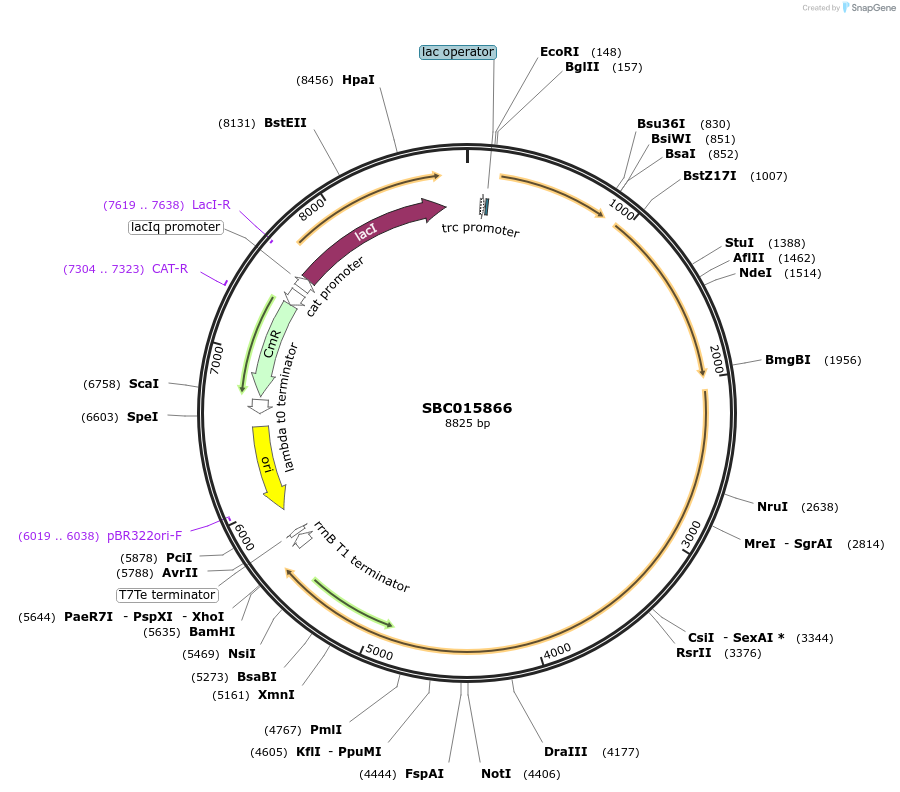
SBC015866
Plasmid#226280PurposeExpresses BsSfp, MsCAD, and SrCAR from trc promoter. Biosynthesis of (hydroxy)cinnamyl alcohols from (hydroxy)cinnamic acids.DepositorInserts4'-phosphopantetheinyl transferase Sfp
Cinnamyl alcohol dehydrogenase
Carboxylic acid reductase
ExpressionBacterialAvailable SinceNov. 25, 2024AvailabilityIndustry, Academic Institutions, and Nonprofits -
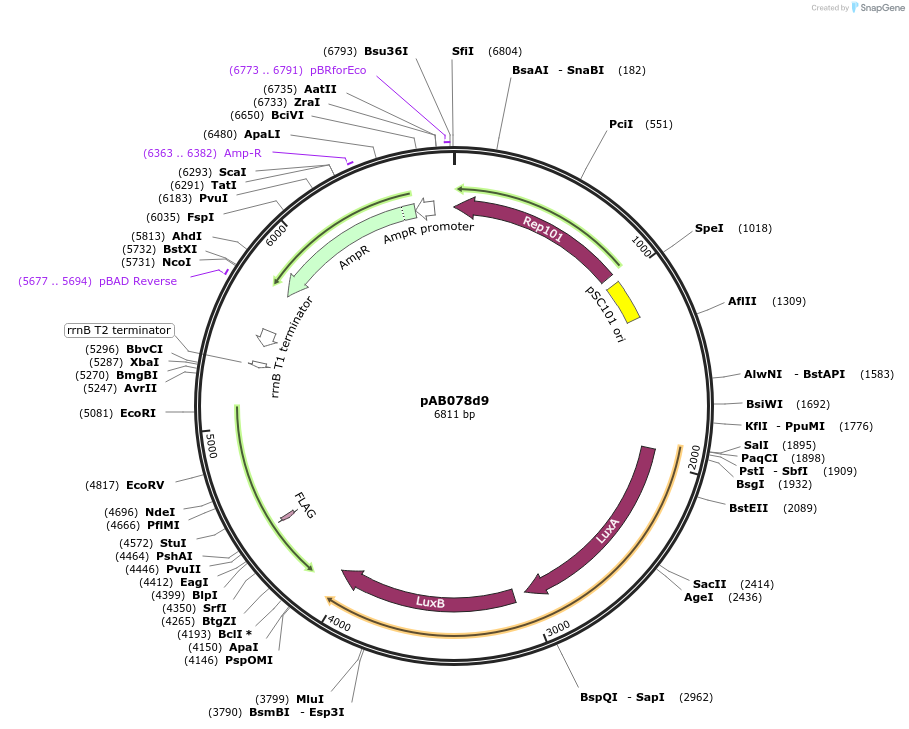
pAB078d9
Plasmid#79207PurposeDirected Evolution of Protein-Protein InteractionsDepositorInsertPlacZ-opt (OR1+2+3) luxAB; Ppro1 434cI-SH2ABL1
ExpressionBacterialPromoterPlacZ-opt (OR1+2+3); Ppro1Available SinceAug. 21, 2018AvailabilityAcademic Institutions and Nonprofits only -
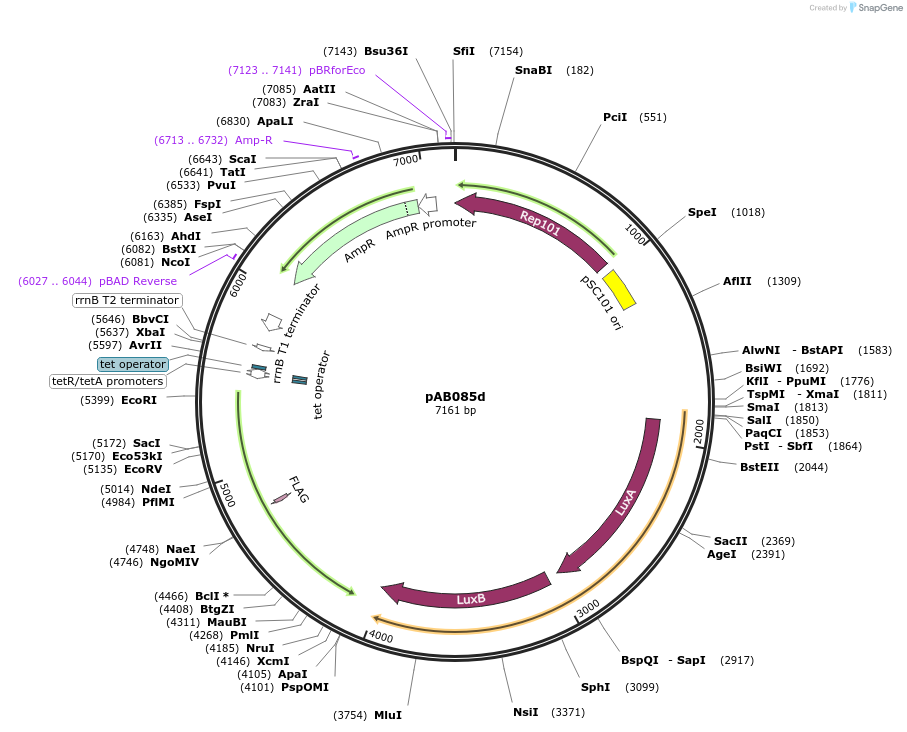
pAB085d
Plasmid#79209PurposeDirected Evolution of Protein-Protein InteractionsDepositorInsertPlacZ-opt (OR1) luxAB; Ptet 434cI-TnTBR3-F3
ExpressionBacterialPromoterPlacZ-opt (OR1); PtetAvailable SinceOct. 3, 2024AvailabilityAcademic Institutions and Nonprofits only -
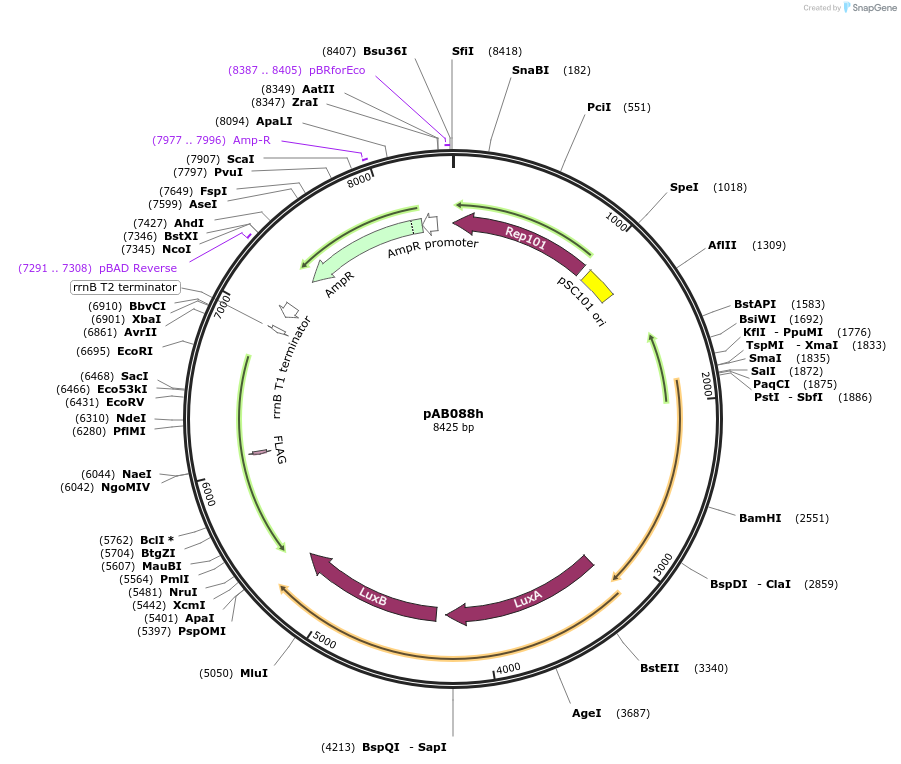
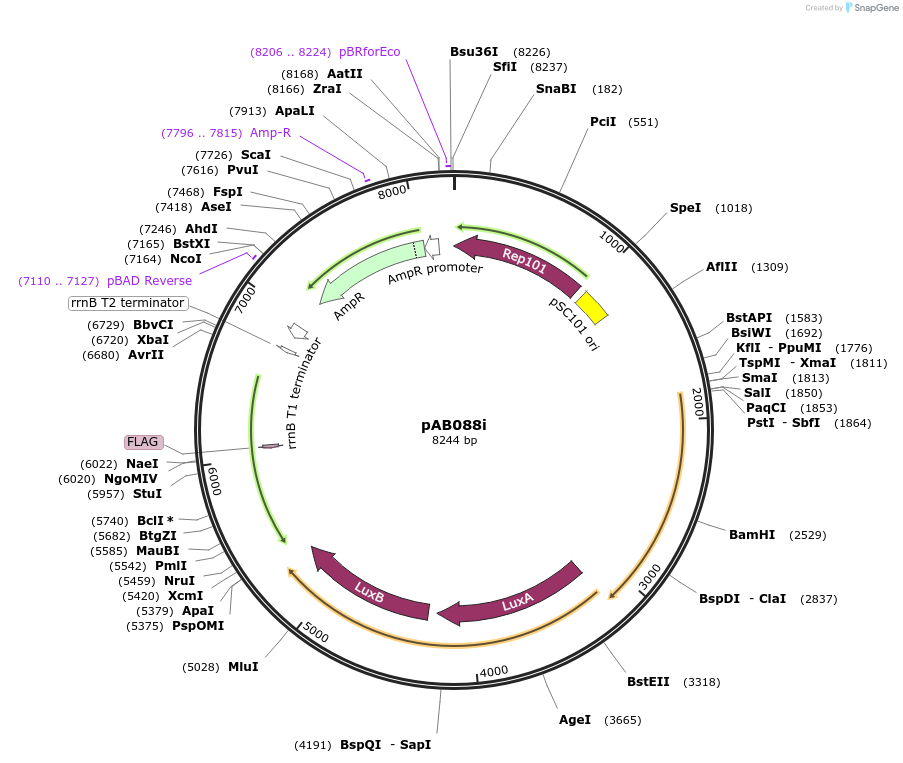
pAB088i
Plasmid#79216PurposeDirected Evolution of Protein-Protein InteractionsDepositorInsertPlacZ-opt (OR1) gIII, luxAB; Ppro1 434cI(RR69)-TnCAD-F3
ExpressionBacterialPromoterPlacZ-opt (OR1); Ppro1Available SinceNov. 8, 2016AvailabilityAcademic Institutions and Nonprofits only -
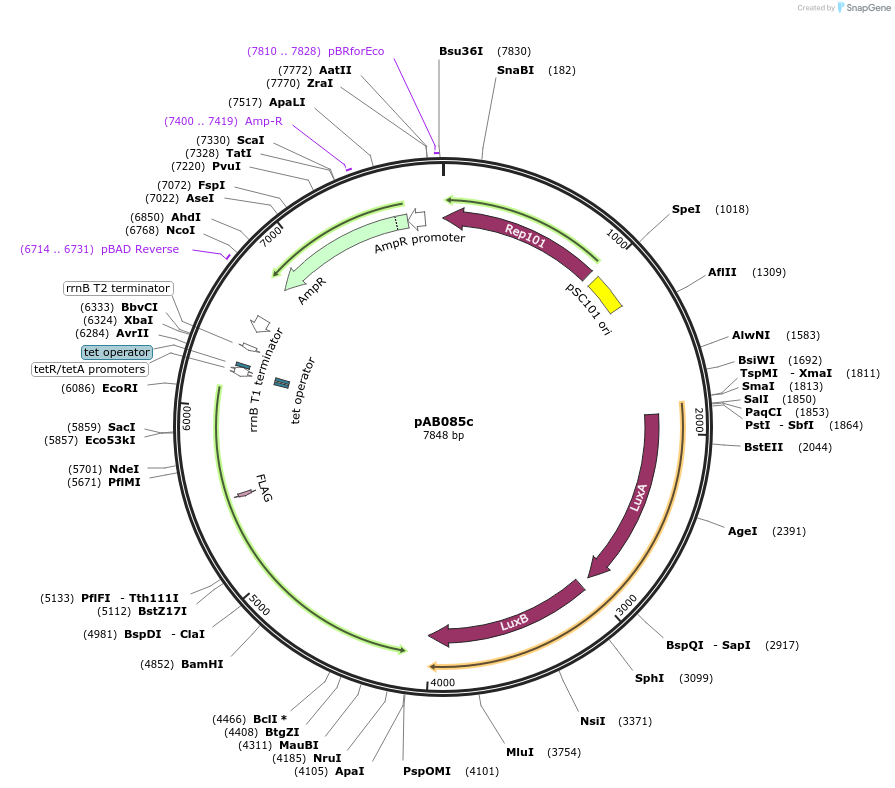
pAB085c
Plasmid#79208PurposeDirected Evolution of Protein-Protein InteractionsDepositorInsertPlacZ-opt (OR1) luxAB; Ptet 434cI-TnTBR3-FL
ExpressionBacterialPromoterPlacZ-opt (OR1); PtetAvailable SinceNov. 8, 2016AvailabilityAcademic Institutions and Nonprofits only -
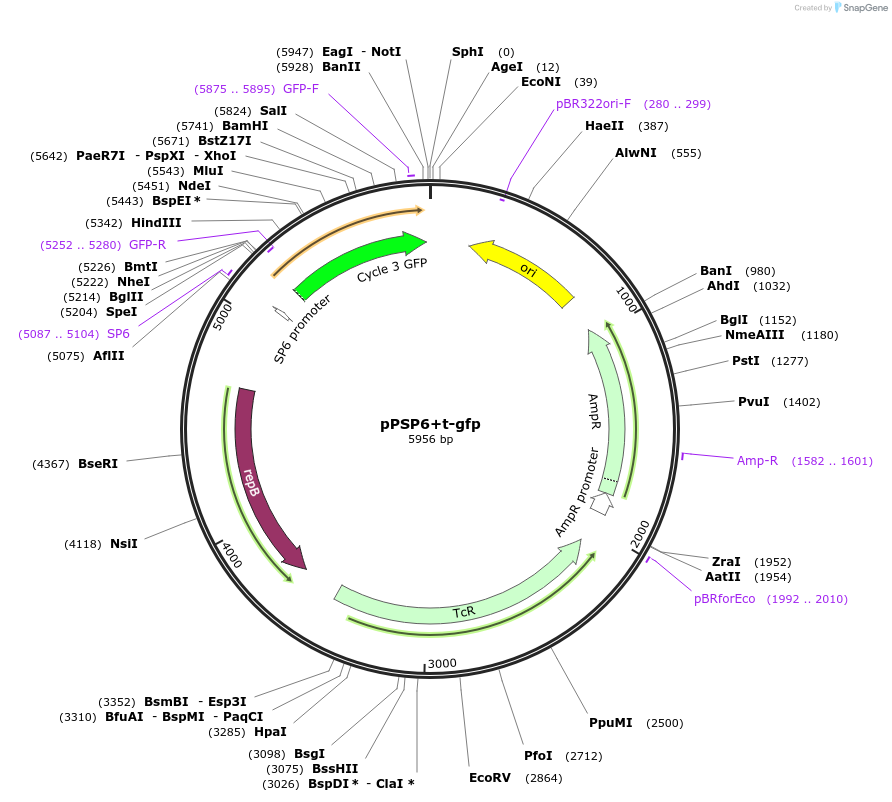
pPSP6+t-gfp
Plasmid#48130PurposeShuttle vector E. coli/B.meg.; gfp-expression under control of RNAP-SP6-inducible promoterDepositorInsertgfp
UseSynthetic BiologyExpressionBacterialAvailable SinceOct. 21, 2013AvailabilityAcademic Institutions and Nonprofits only -
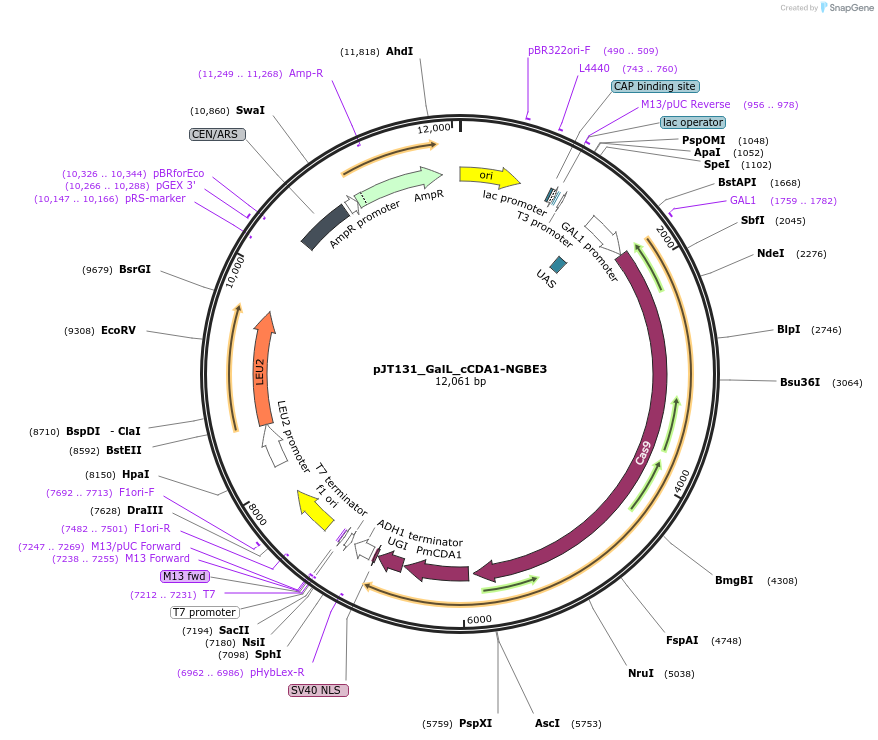
pJT131_GalL_cCDA1-NGBE3
Plasmid#145095PurposeExpressing base editor cCDA1-NGBE3 in yeast cellsDepositorInsertcCDA1-NGBE3
UseCRISPRExpressionYeastMutationspCas9(D10A/L1111R/D1135V/G1218R/E1219F/A1322R/R1…PromoterGalLAvailable SinceJuly 15, 2020AvailabilityAcademic Institutions and Nonprofits only -
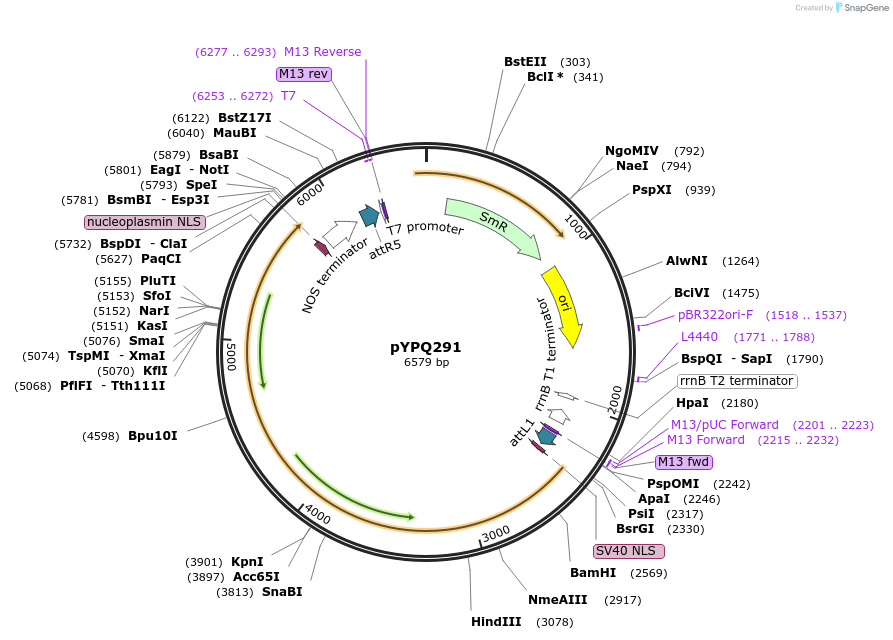
pYPQ291
Plasmid#129671PurposeGateway entry clone for CRISPR-Cas12b systemsDepositorInsertBthCas12b
UseCRISPR; Gateway compatible bthcas12b entry cloneExpressionPlantMutationBthCas12b is rice codon optimizedAvailable SinceMarch 18, 2020AvailabilityAcademic Institutions and Nonprofits only -
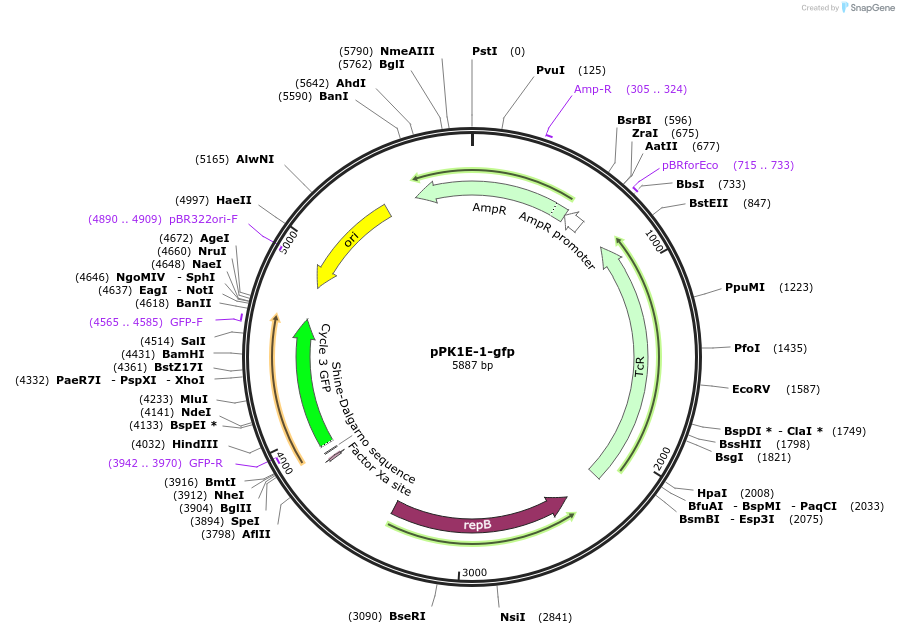
pPK1E-1-gfp
Plasmid#48133PurposeShuttle vector E. coli/B.meg.; gfp-expression under control of RNAP-K1E-inducible promoterDepositorInsertgfp
UseSynthetic BiologyExpressionBacterialAvailable SinceOct. 21, 2013AvailabilityAcademic Institutions and Nonprofits only -
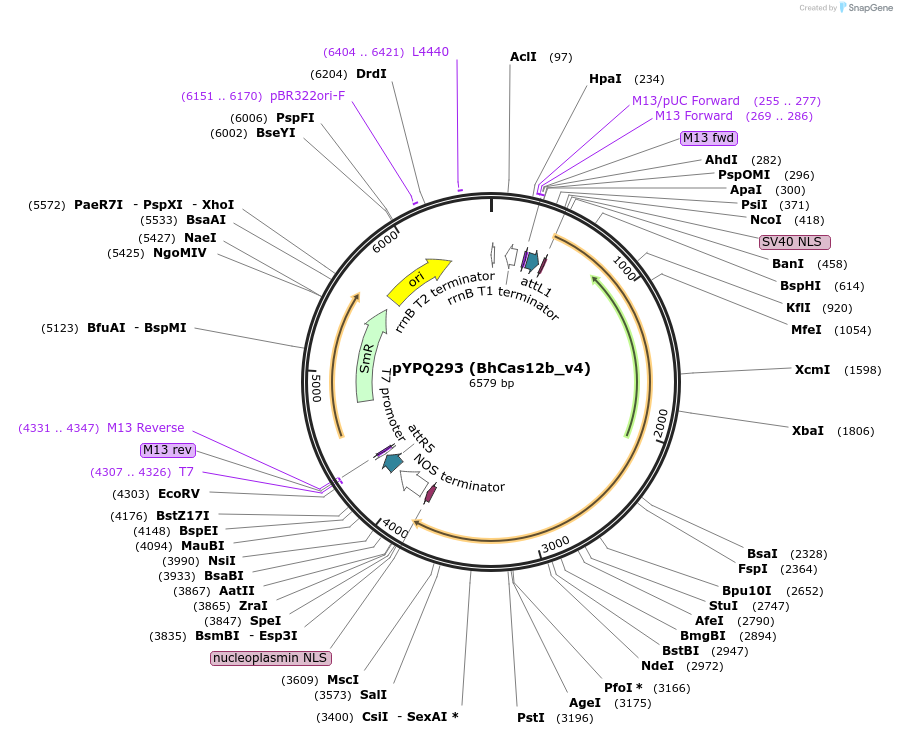
pYPQ293 (BhCas12b_v4)
Plasmid#136383PurposeGateway entry clone for BhCas12b_v4DepositorInsertBhCas12b_v4
UseCRISPRExpressionPlantMutationK846R/S893R/E837GAvailable SinceFeb. 18, 2020AvailabilityAcademic Institutions and Nonprofits only -
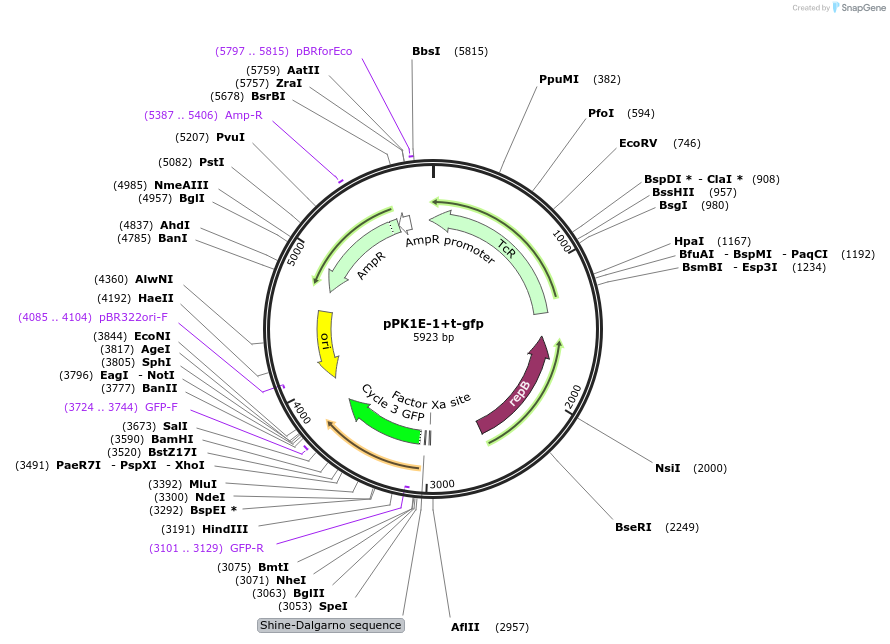
pPK1E-1+t-gfp
Plasmid#48131PurposeShuttle vector E. coli/B.meg.; gfp-expression under control of RNAP-K1E-inducible promoterDepositorInsertgfp
UseSynthetic BiologyExpressionBacterialAvailable SinceOct. 11, 2013AvailabilityAcademic Institutions and Nonprofits only -
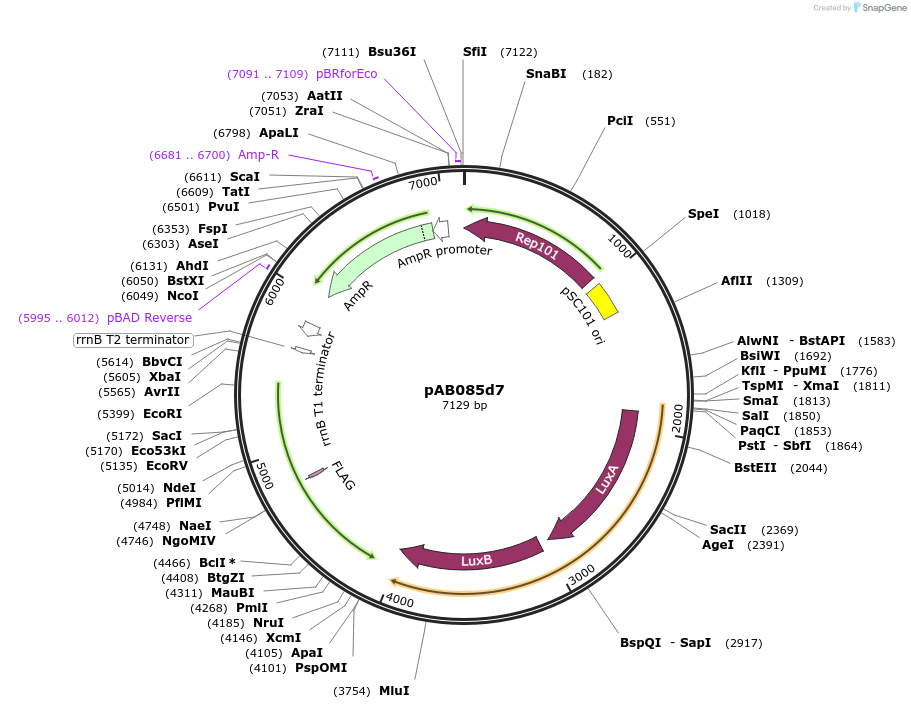
pAB085d7
Plasmid#79211PurposeDirected Evolution of Protein-Protein InteractionsDepositorInsertPlacZ-opt (OR1) luxAB; Ppro1 434cI-TnCAD-F3
ExpressionBacterialPromoterPlacZ-opt (OR1); Ppro1Available SinceDec. 17, 2018AvailabilityAcademic Institutions and Nonprofits only -
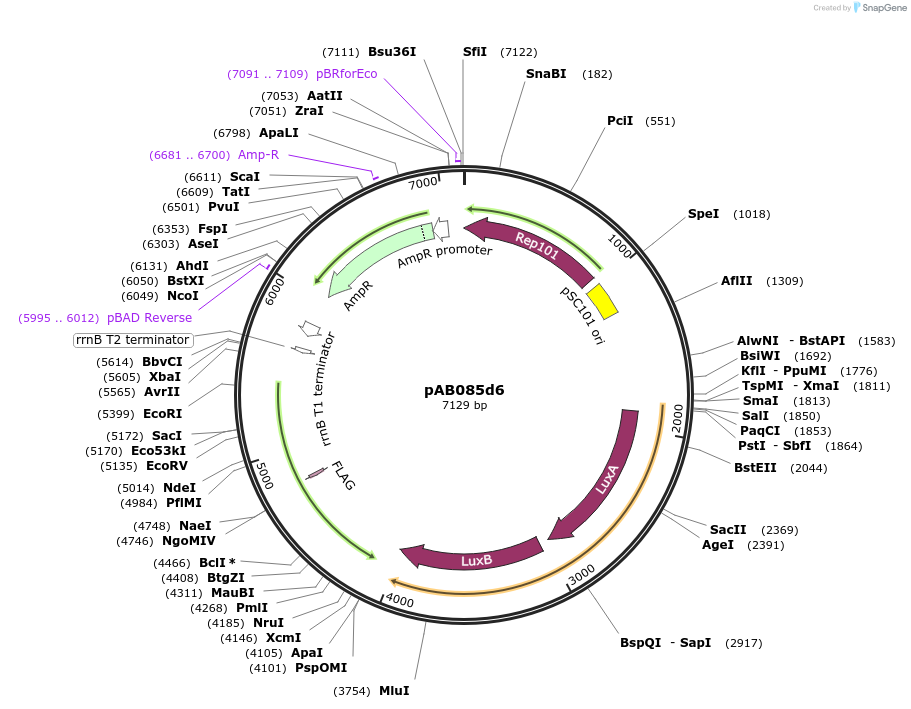
pAB085d6
Plasmid#79210PurposeDirected Evolution of Protein-Protein InteractionsDepositorInsertPlacZ-opt (OR1) luxAB; Ppro1 434cI-TnTBR3-F3
ExpressionBacterialPromoterPlacZ-opt (OR1); Ppro1Available SinceDec. 17, 2018AvailabilityAcademic Institutions and Nonprofits only -
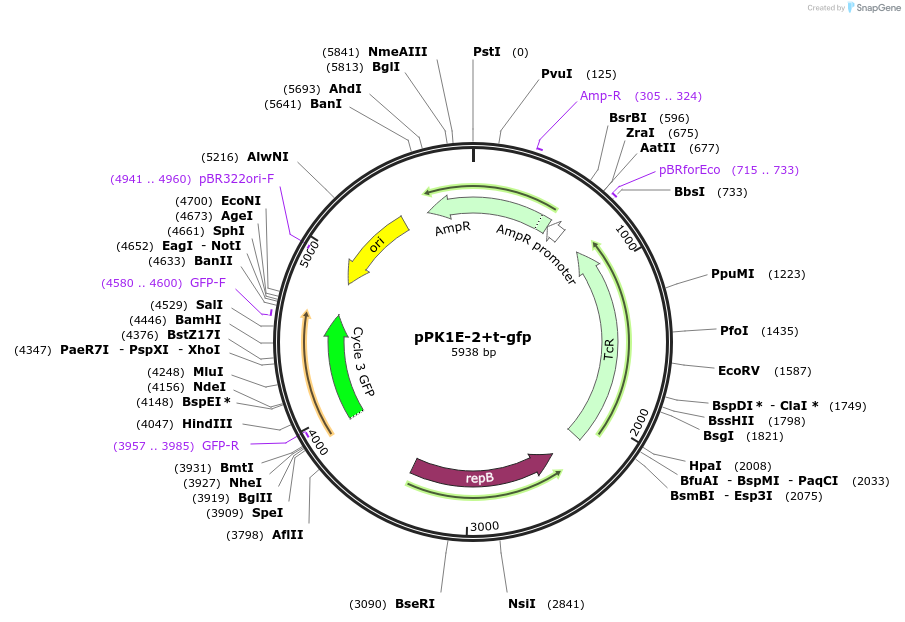
pPK1E-2+t-gfp
Plasmid#48132PurposeShuttle vector E. coli/B.meg.; gfp-expression under control of RNAP-K1E-inducible promoterDepositorInsertgfp
UseSynthetic BiologyExpressionBacterialAvailable SinceOct. 21, 2013AvailabilityAcademic Institutions and Nonprofits only -
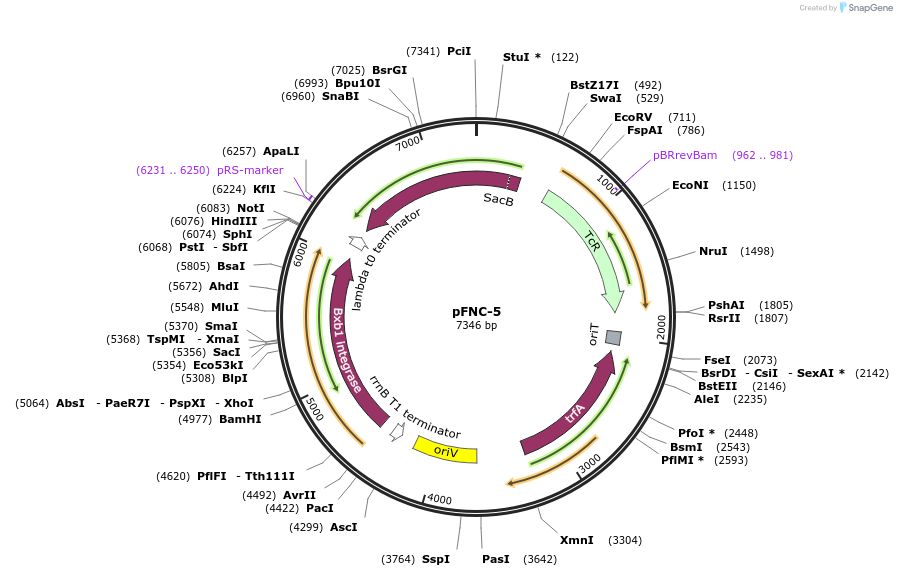
pFNC-5
Plasmid#233297PurposePlasmid encoding the BxbI phage integrase. It can be used to remove the antibiotic resistance cassette integrated with pCIFR. TcR, easily curable via sucrose counterselection.DepositorInsertsBxbI phage integrase
sacB
ExpressionBacterialPromoterPEM7 and unknownAvailable SinceApril 9, 2025AvailabilityAcademic Institutions and Nonprofits only -
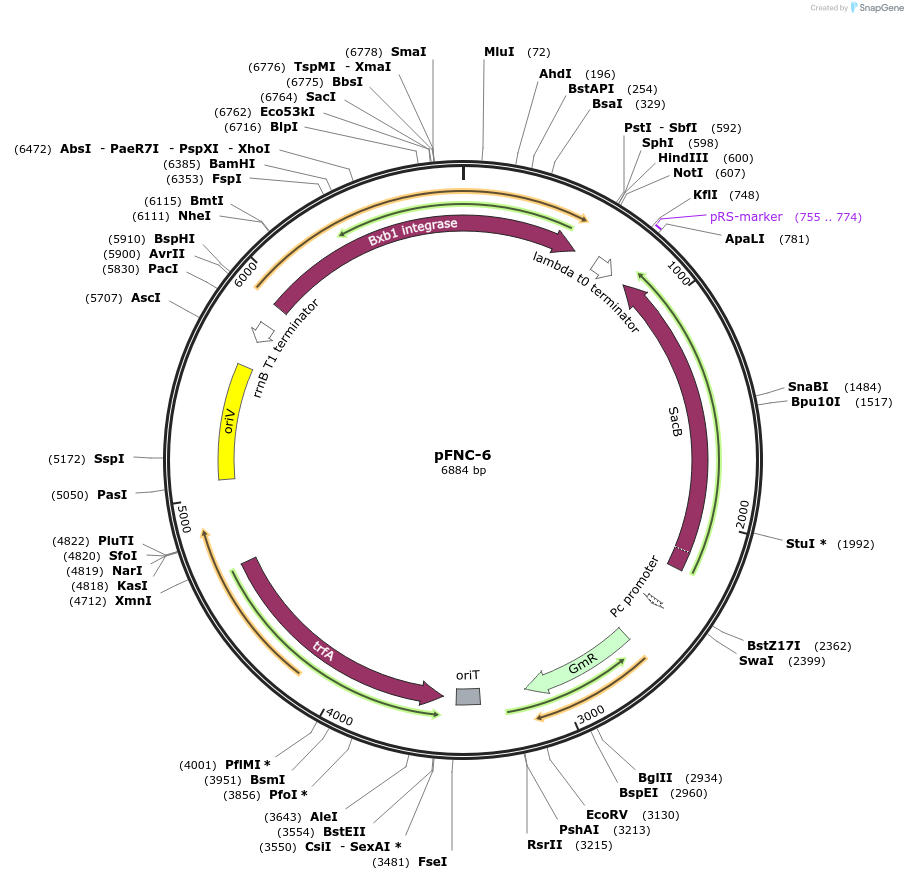
pFNC-6
Plasmid#227669PurposePlasmid encoding the BxbI phage integrase. It can be used to remove the antibiotic resistance cassette integrated with pCIFR. GmR, easily curable via sucrose counterselection.DepositorInsertsBxbI phage integrase
sacB
ExpressionBacterialPromoterPEM7Available SinceJan. 22, 2025AvailabilityAcademic Institutions and Nonprofits only -
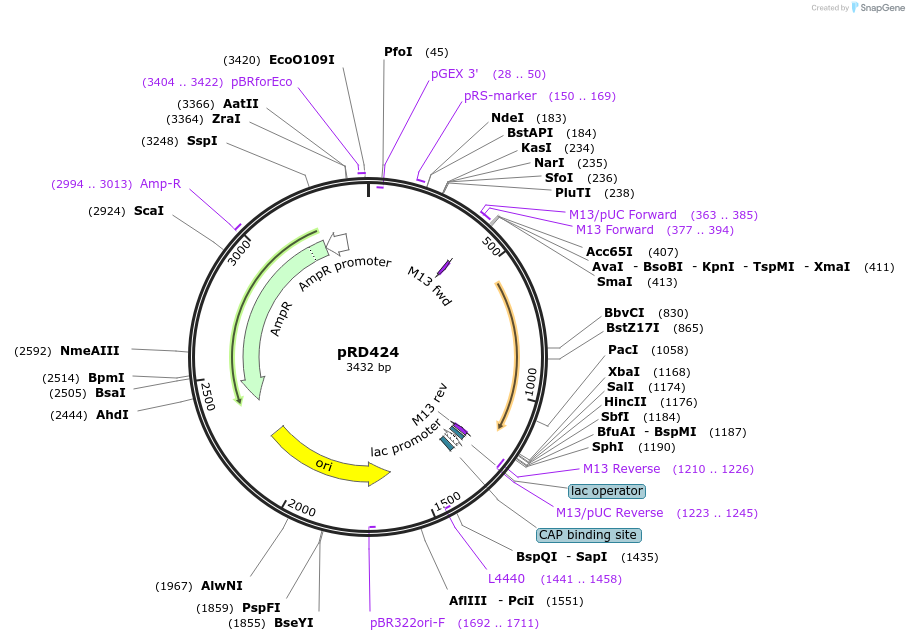
pRD424
Plasmid#178202PurposeExpression of Rok from the hns promoterDepositorInsertphns + rok
ExpressionBacterialPromoterhns promoterAvailable SinceJan. 3, 2023AvailabilityAcademic Institutions and Nonprofits only -
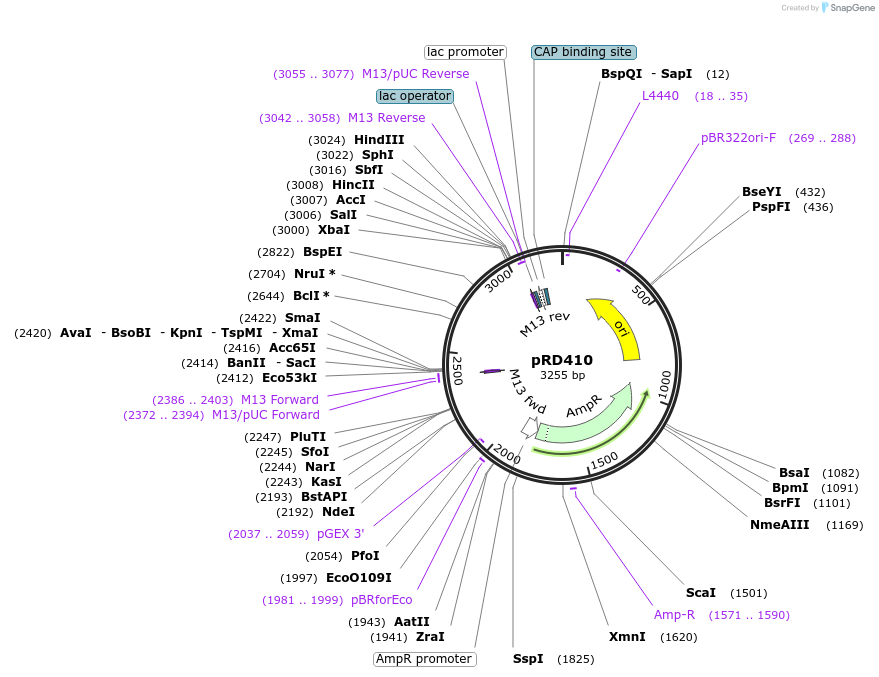
pRD410
Plasmid#178200PurposeExpression of sRok from the hns promoterDepositorInsertphns + srok
ExpressionBacterialPromoterhns promoterAvailable SinceNov. 22, 2022AvailabilityAcademic Institutions and Nonprofits only -
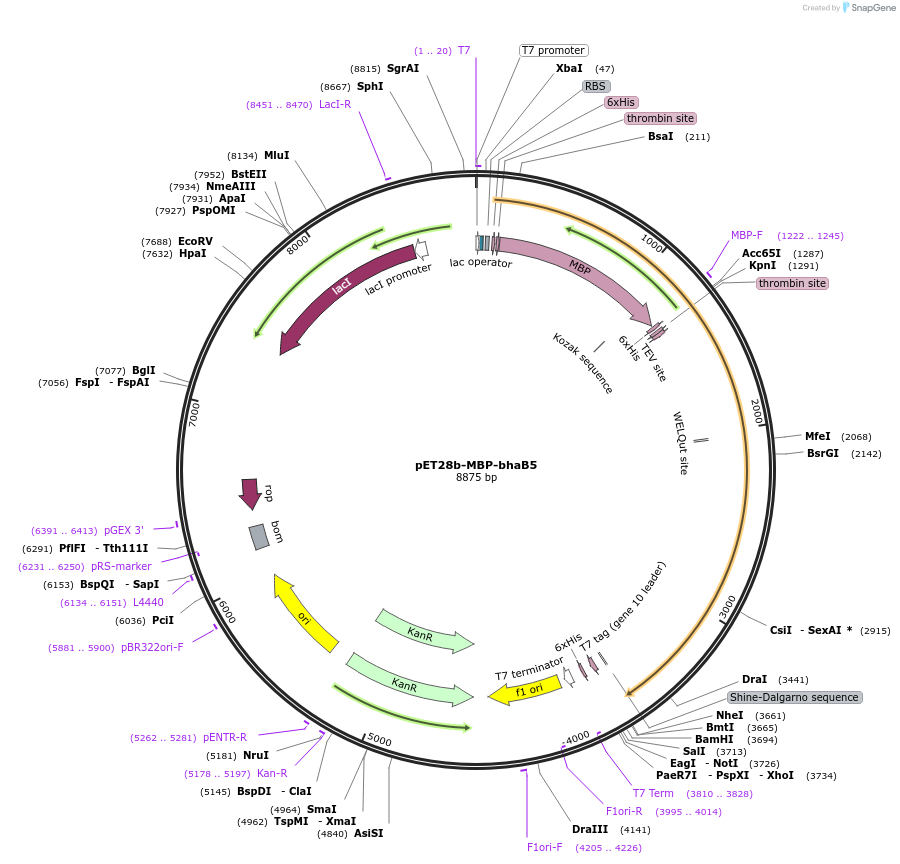
pET28b-MBP-bhaB5
Plasmid#167812PurposeExpresses His6- and MBP-tagged BhaB5 in E. coliDepositorInsertBhaB5
TagsHis6, MBPExpressionBacterialPromoterT7Available SinceAug. 11, 2022AvailabilityAcademic Institutions and Nonprofits only -
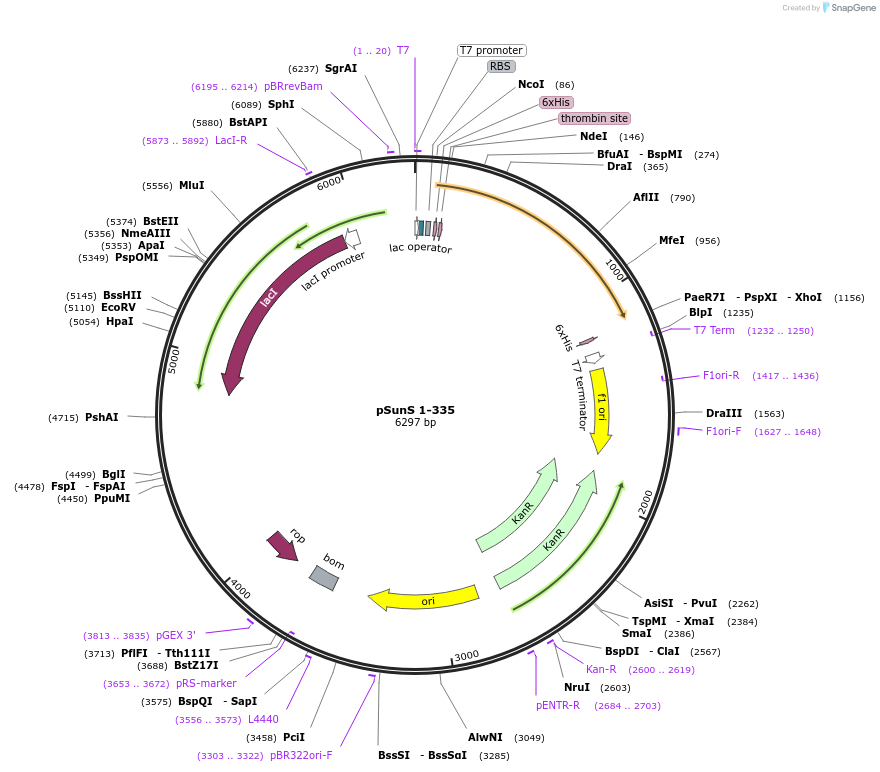
pSunS 1-335
Plasmid#171673PurposeSunS 1-335 expression in E. coliDepositorInsertGlycosyltransferase SunS (sunS Bacillus subtilis (strain 168))
TagsHis-tagExpressionBacterialPromoterT7Available SinceAug. 12, 2021AvailabilityAcademic Institutions and Nonprofits only -
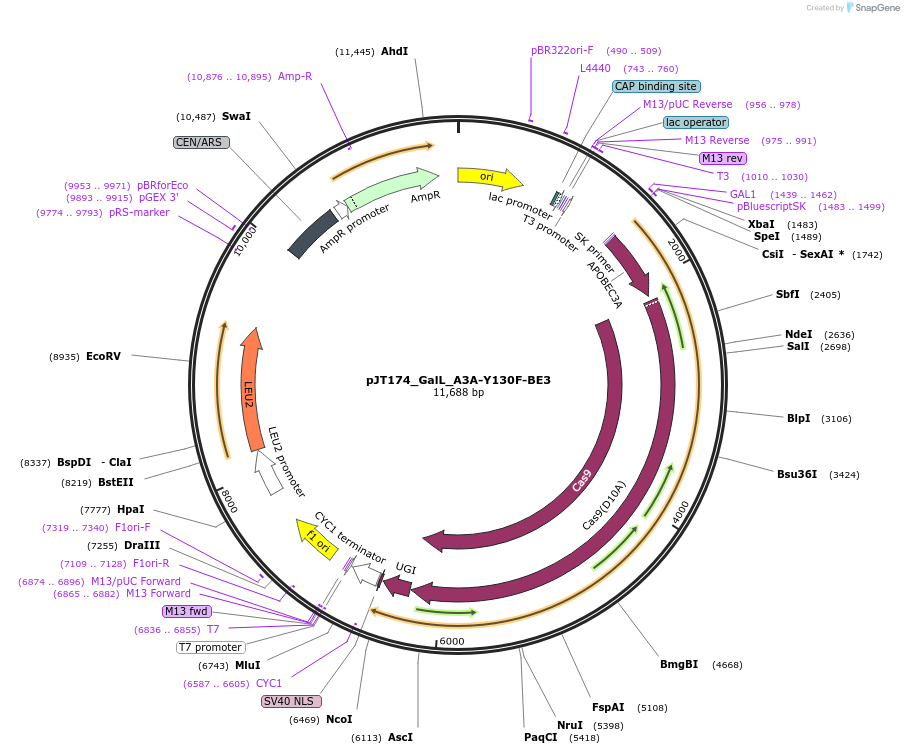
pJT174_GalL_A3A-Y130F-BE3
Plasmid#145123PurposeExpressing base editor A3A-Y130F-BE3 in yeast cellsDepositorInsertA3A(Y130F)-BE3
UseCRISPRExpressionYeastMutationA3A(Y130F); spCas9(D10A)PromoterGalLAvailable SinceJuly 22, 2020AvailabilityAcademic Institutions and Nonprofits only -
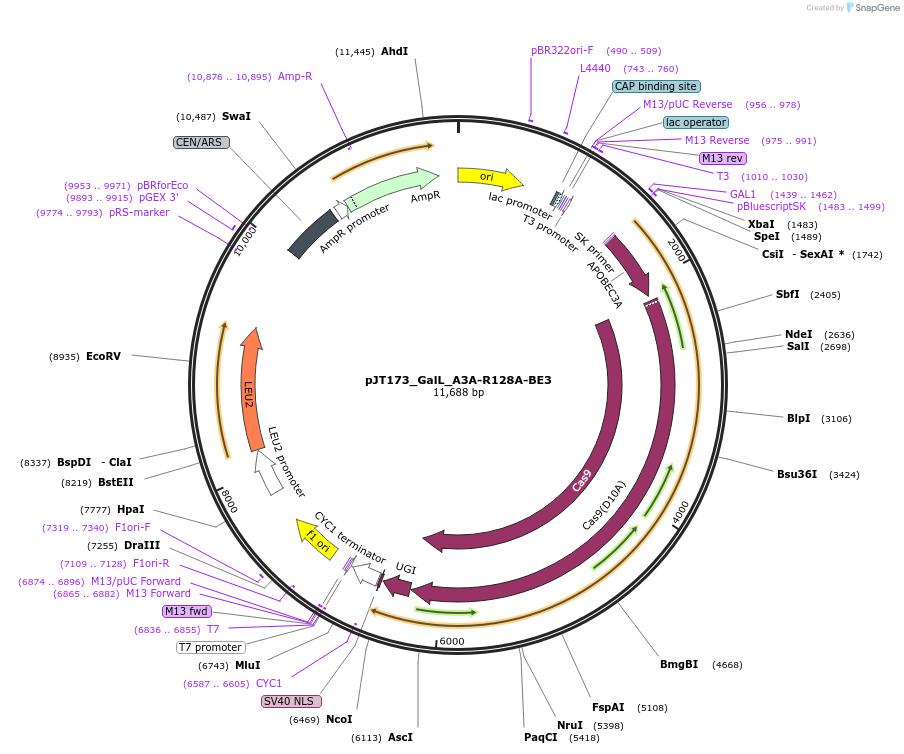
pJT173_GalL_A3A-R128A-BE3
Plasmid#145122PurposeExpressing base editor A3A-R128A-BE3 in yeast cellsDepositorInsertA3A(R128A)-BE3
UseCRISPRExpressionYeastMutationA3A(R128A); spCas9(D10A)PromoterGalLAvailable SinceJuly 22, 2020AvailabilityAcademic Institutions and Nonprofits only -
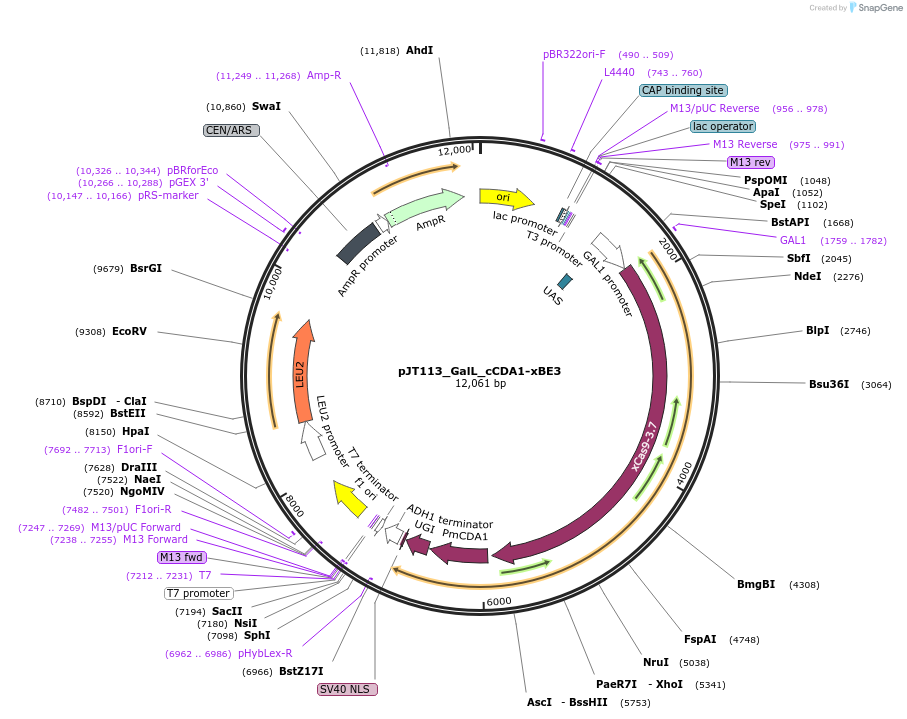
pJT113_GalL_cCDA1-xBE3
Plasmid#145079PurposeExpressing base editor cCDA1-xBE3 in yeast cellsDepositorInsertcCDA1-xBE3
UseCRISPRExpressionYeastMutationspCas9(D10A, A262T, R324L, S409I, E480K, E543D, M…PromoterGalLAvailable SinceJuly 15, 2020AvailabilityAcademic Institutions and Nonprofits only -
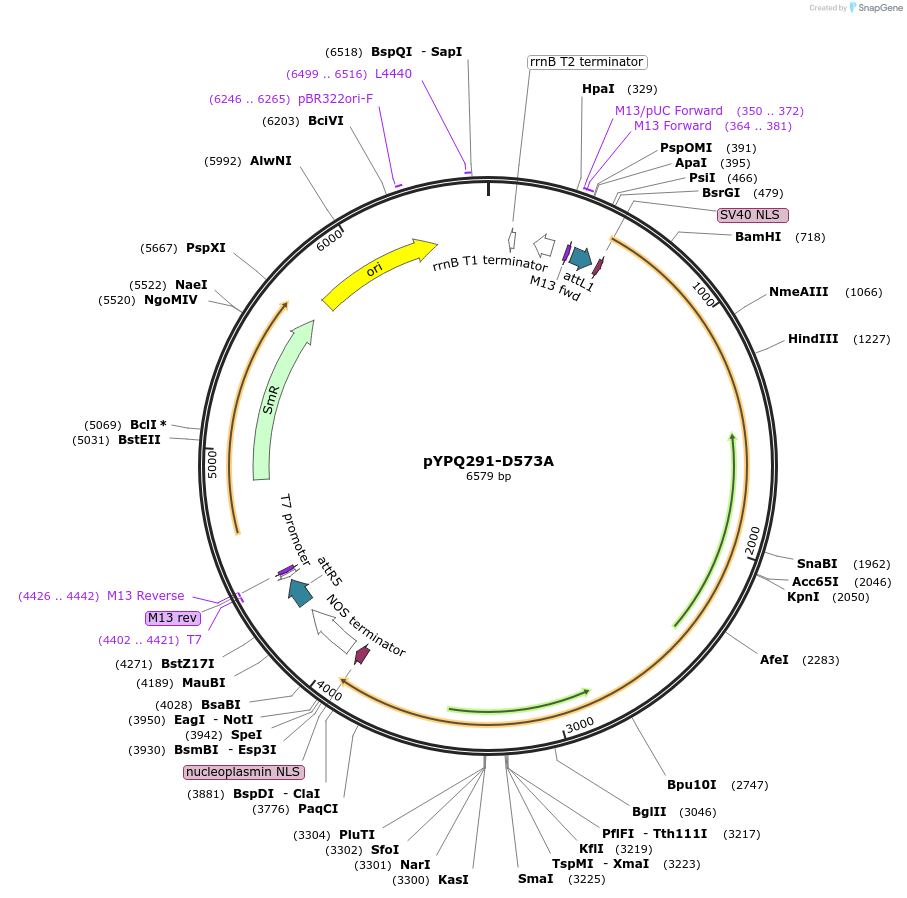
pYPQ291-D573A
Plasmid#129676PurposeGateway entry clone for CRISPR-Cas12b systemsDepositorInsertBthCas12b
UseCRISPR; Gateway compatible bthcas12b entry cloneExpressionPlantMutationD573A; BthCas12b is rice codon optimizedAvailable SinceMarch 18, 2020AvailabilityAcademic Institutions and Nonprofits only -
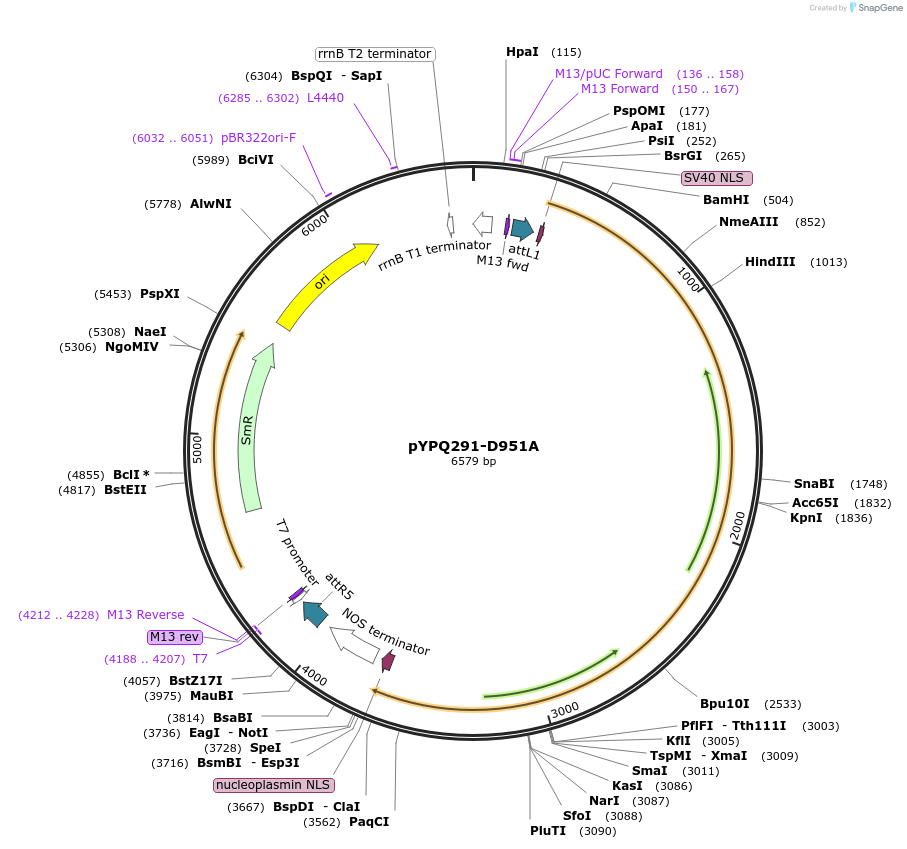
pYPQ291-D951A
Plasmid#129677PurposeGateway entry clone for CRISPR-Cas12b systemsDepositorInsertBthCas12b
UseCRISPR; Gateway compatible bthcas12b entry cloneExpressionPlantMutationD951A; BthCas12b is rice codon optimizedAvailable SinceMarch 18, 2020AvailabilityAcademic Institutions and Nonprofits only -
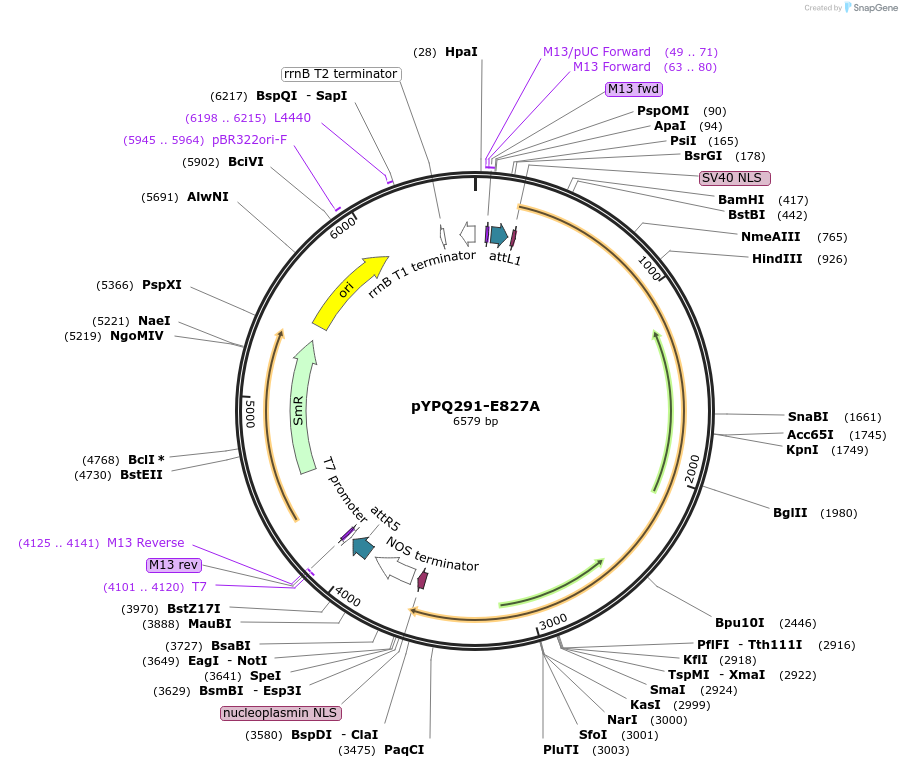
pYPQ291-E827A
Plasmid#129678PurposeGateway entry clone for CRISPR-Cas12b systemsDepositorInsertBthCas12b
UseCRISPR; Gateway compatible bthcas12b entry cloneExpressionPlantMutationE827A; BthCas12b is rice codon optimizedAvailable SinceMarch 18, 2020AvailabilityAcademic Institutions and Nonprofits only -
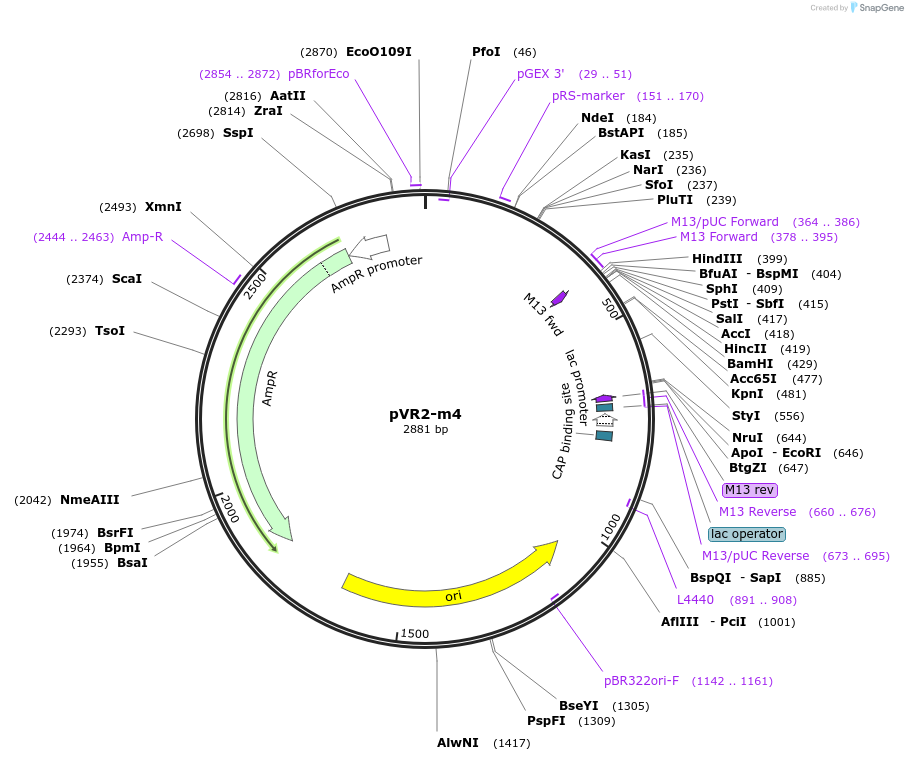
pVR2-m4
Plasmid#84723PurposepVR2 but with G to U and A to C mutations following the riboswitch aptamerDepositorInsertsE. coli RecA promoter
stretch lacking A's followed by 5' UTR of queC gene
ExpressionBacterialMutationG to U mutation at the first nucleotide after the…Available SinceMarch 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
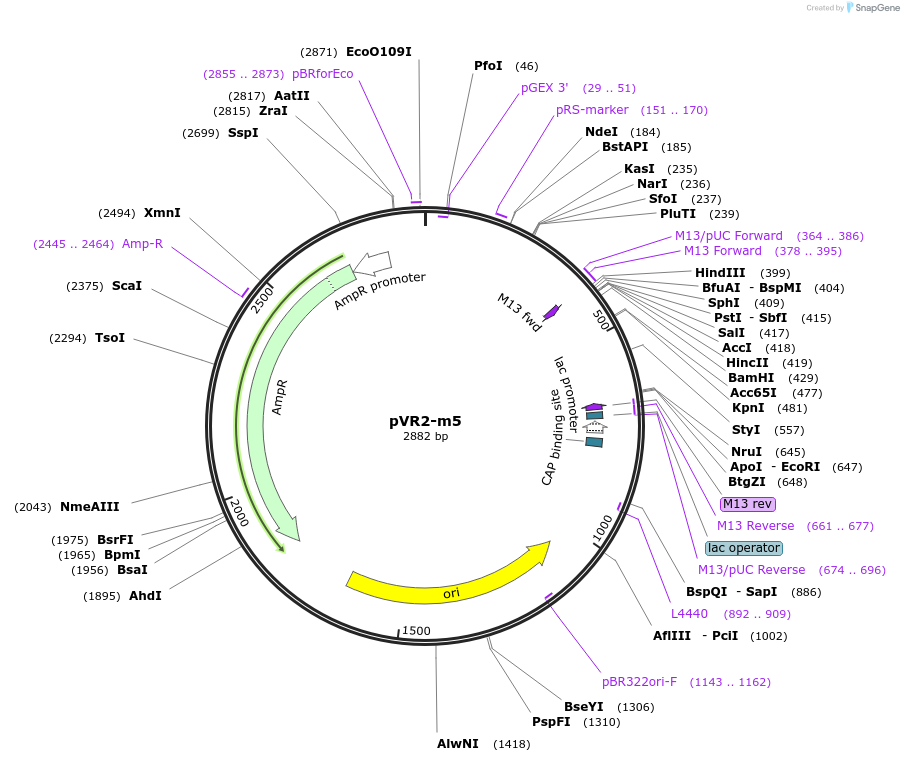
pVR2-m5
Plasmid#84724PurposepVR2 but with an insertion of 1 U immediately following the riboswitch aptamerDepositorInsertsE. coli RecA promoter
stretch lacking A's followed by 5' UTR of queC gene
ExpressionBacterialMutationinsertion of 1 U between positions 39 and 40 of a…Available SinceMarch 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
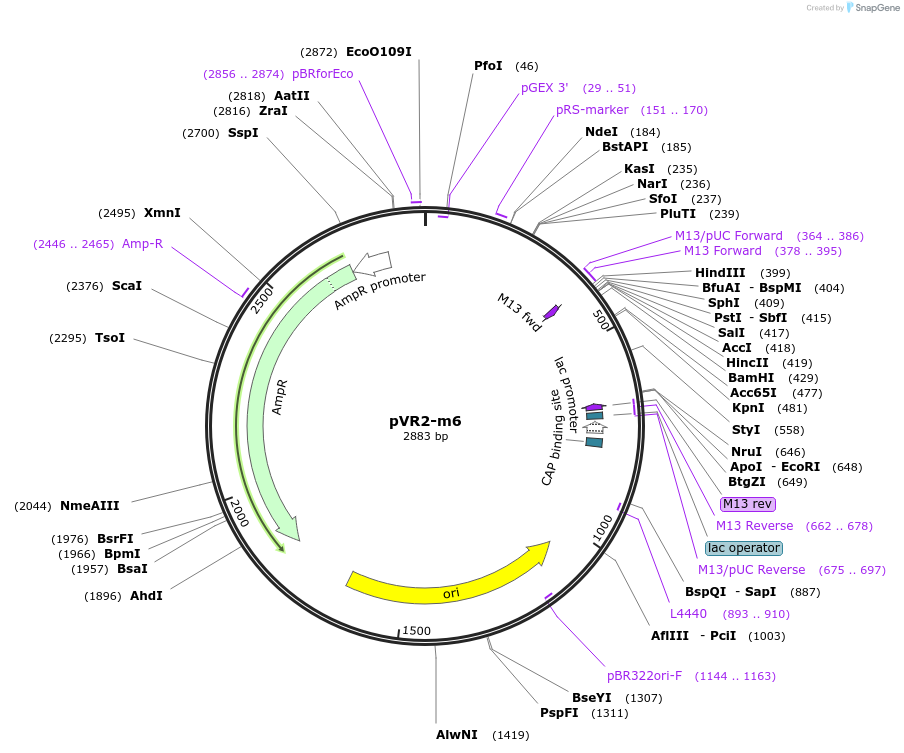
pVR2-m6
Plasmid#84725PurposepVR2 but with an insertion of 2 U's immediately following the riboswitch aptamerDepositorInsertsE. coli RecA promoter
stretch lacking A's followed by 5' UTR of queC gene
ExpressionBacterialMutationinsertion of 2 Us between positions 39 and 40 of …Available SinceMarch 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
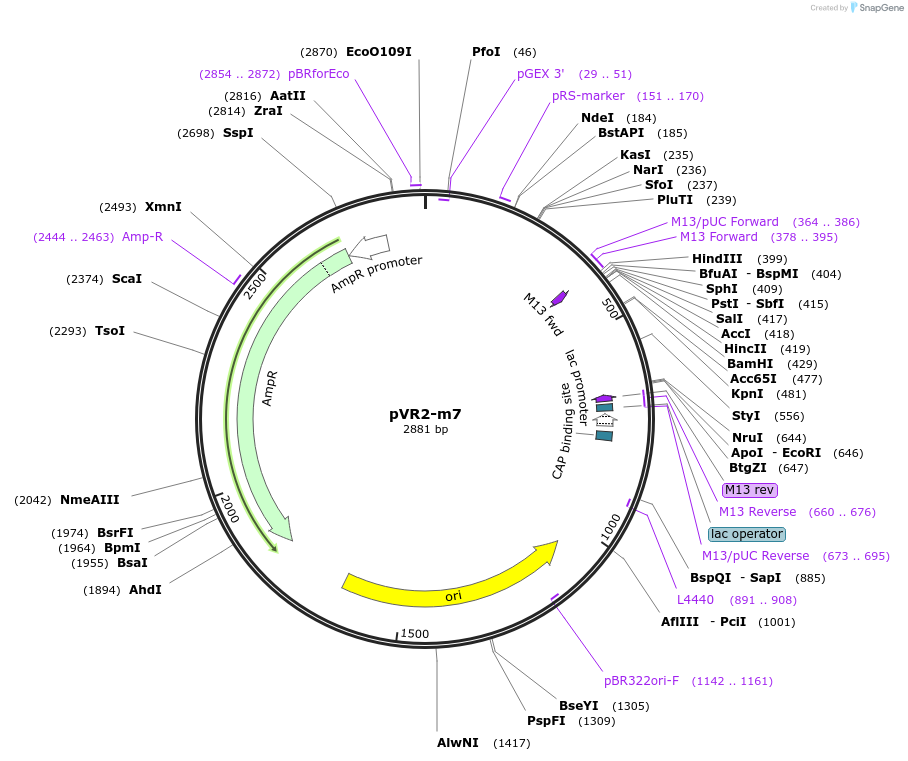
pVR2-m7
Plasmid#84726PurposepVR2 but with a G to U mutation immediately following the riboswitch aptamerDepositorInsertsE. coli RecA promoter
stretch lacking A's followed by 5' UTR of queC gene
ExpressionBacterialMutationG to U mutation at position 41 of annotated "…Available SinceMarch 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
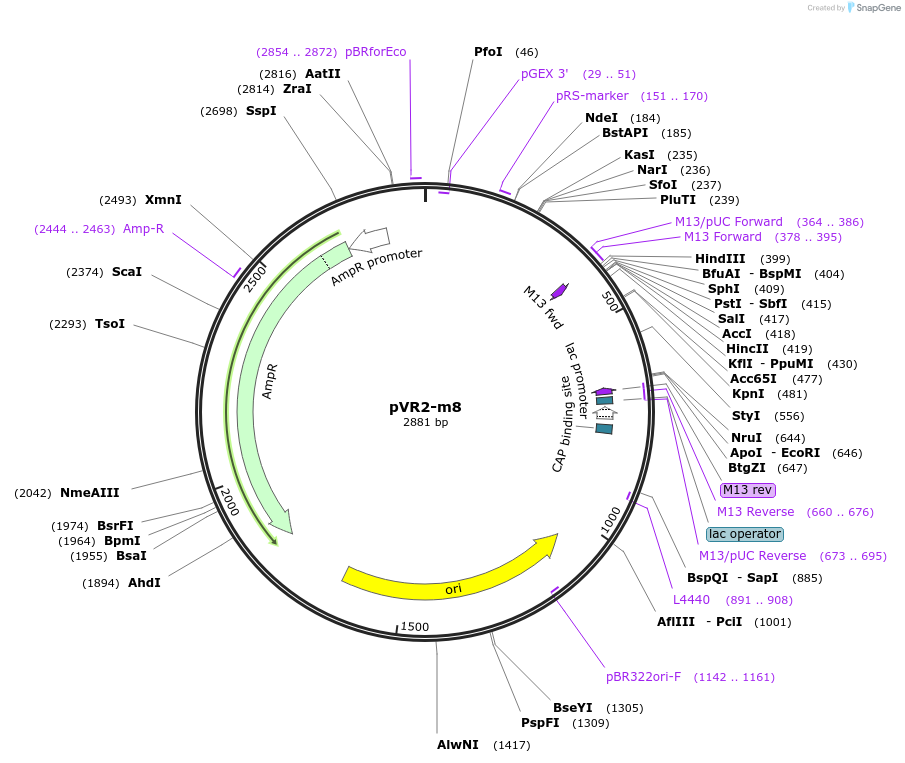
pVR2-m8
Plasmid#84727PurposepVR2 but with two G to U mutations immediately following the riboswitch aptamerDepositorInsertsE. coli RecA promoter
stretch lacking A's followed by 5' UTR of queC gene
ExpressionBacterialMutationG to U mutations at positions 40 and 41 of annota…Available SinceMarch 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
pVR2
Plasmid#84719PurposeE. coli RecA promoter followed by stretch with no A's followed by riboswitch for single-round transcriptionDepositorInsertsE. coli RecA promoter
stretch lacking A's followed by 5' UTR of queC gene
ExpressionBacterialMutation12 nt stretch lacking A's inserted before 5&…Available SinceMarch 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
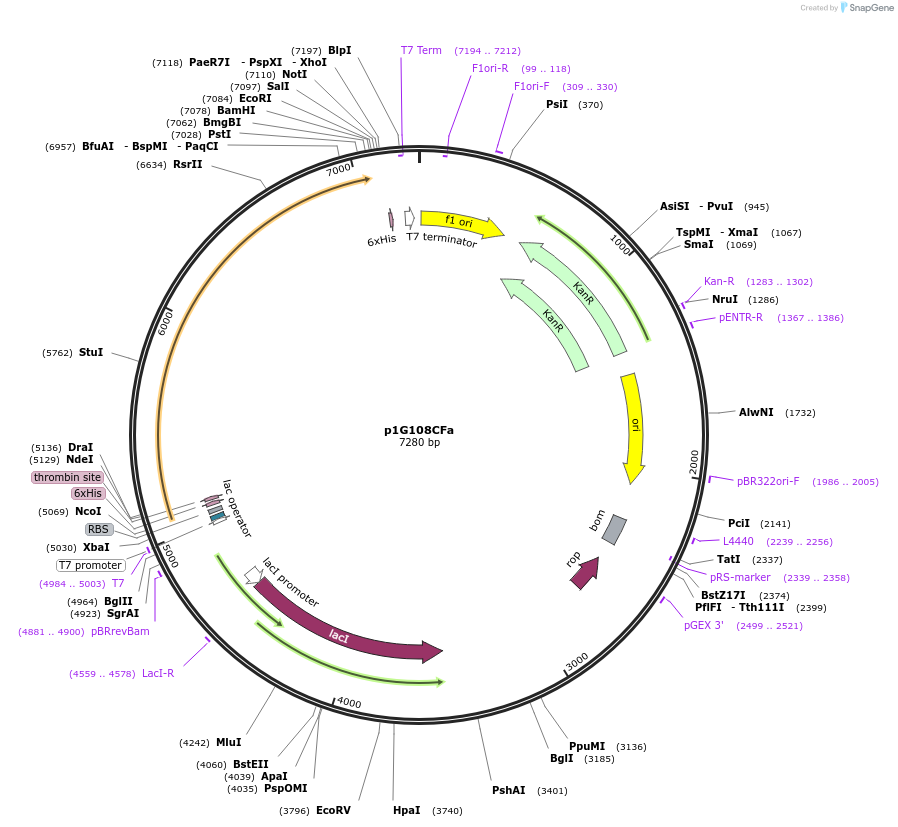
p1G108CFa
Plasmid#85089PurposeExpresses PCNA1 variant (G108C) fused to an FAD domain of P450 BM3 (A74G/C773S/C810S) in E. coliDepositorInsertThe G108C variant of PCNA1 fused to an FAD domain of P450 BM3
TagsHis6 tagExpressionBacterialMutationG108C in PCNA1, A74G/C773S/C810S in FAD domain of…PromoterT7 promoterAvailable SinceNov. 29, 2016AvailabilityAcademic Institutions and Nonprofits only -
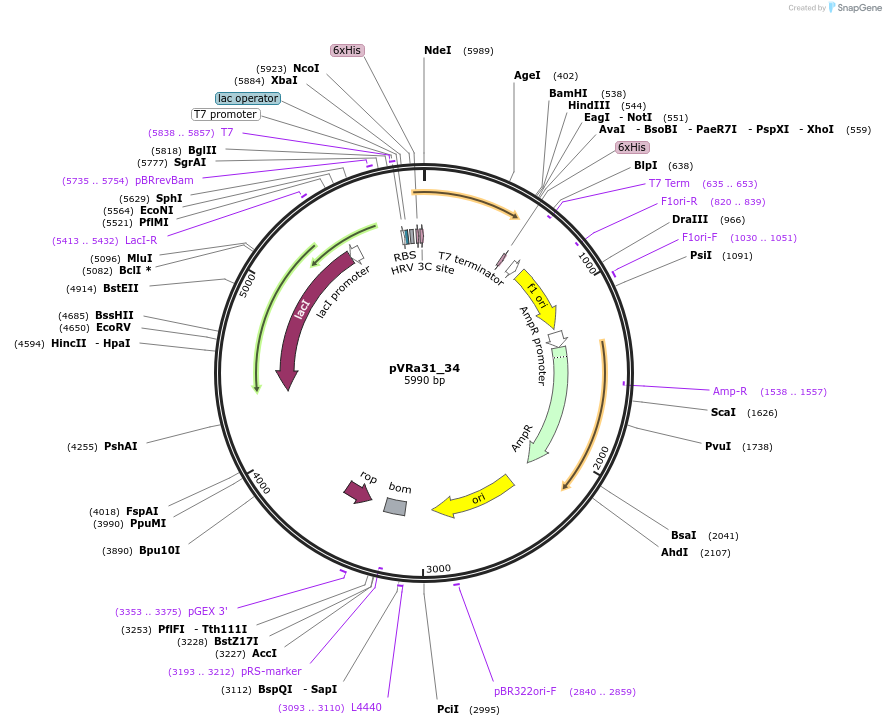
pVRa31_34
Plasmid#49687DepositorInsertECF31_34
UseSynthetic BiologyTagsHis6-PreScissionExpressionBacterialMutationCodon optimized for E.coliPromoterpT7-lacOAvailable SinceJan. 6, 2014AvailabilityAcademic Institutions and Nonprofits only -
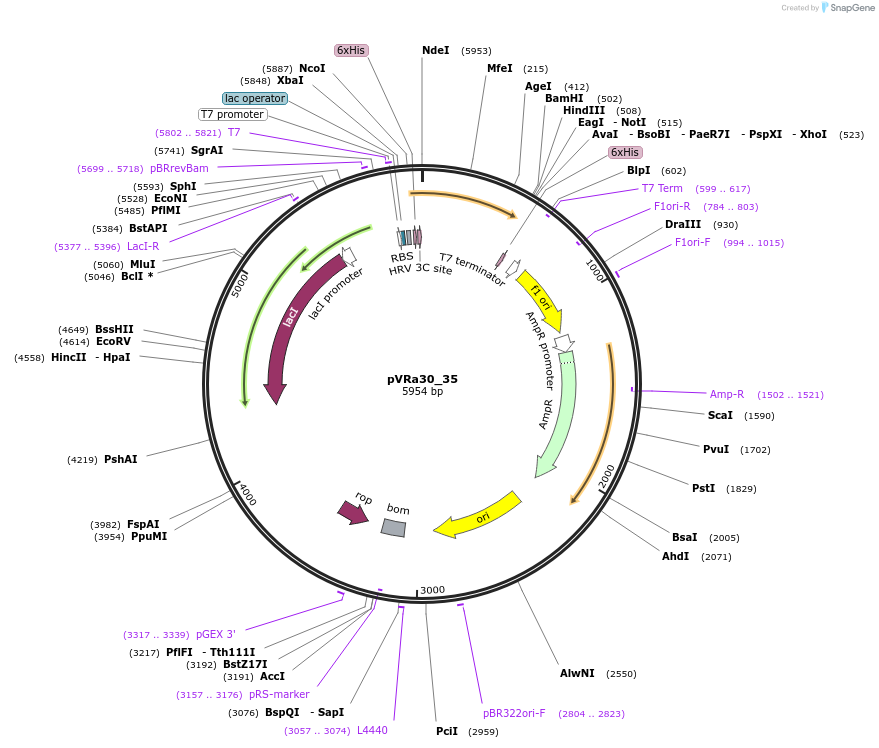
pVRa30_35
Plasmid#49685DepositorInsertECF30_35
UseSynthetic BiologyTagsHis6-PreScissionExpressionBacterialMutationCodon optimized for E.coliPromoterpT7-lacOAvailable SinceJan. 6, 2014AvailabilityAcademic Institutions and Nonprofits only -
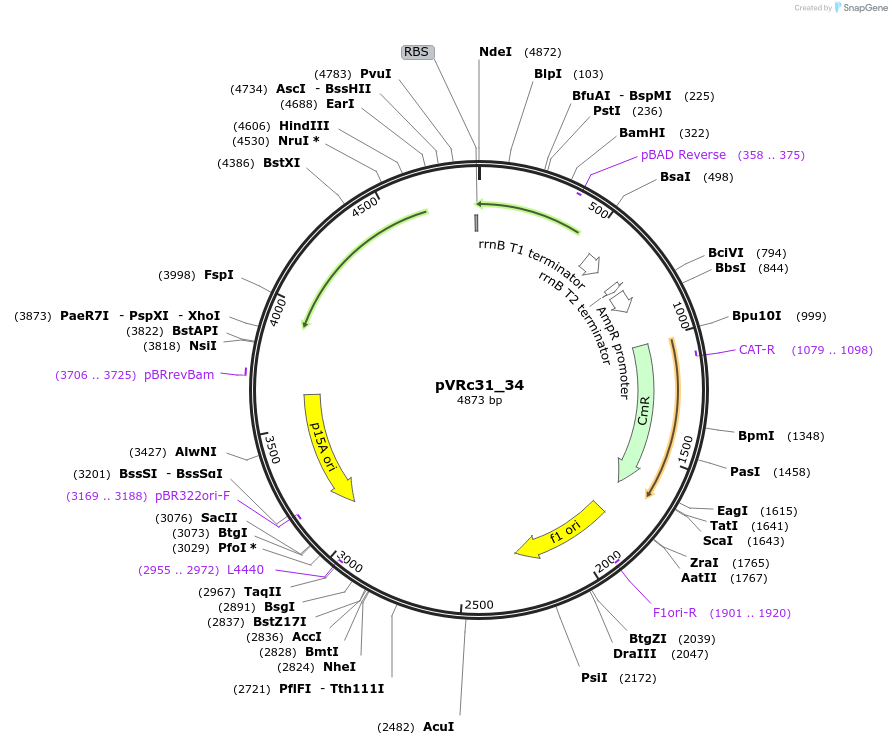
pVRc31_34
Plasmid#49747DepositorInsertAS31_34
UseSynthetic BiologyExpressionBacterialMutationCodon optimized for E.coliPromoterpLuxAvailable SinceJan. 6, 2014AvailabilityAcademic Institutions and Nonprofits only -
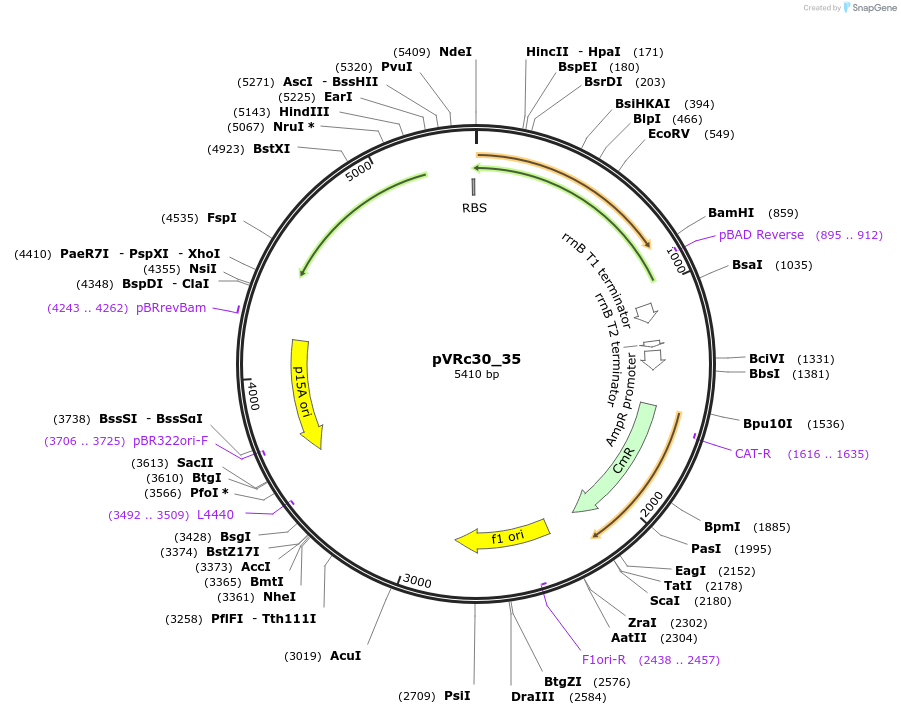
pVRc30_35
Plasmid#49746DepositorInsertAS30_35
UseSynthetic BiologyExpressionBacterialMutationCodon optimized for E.coliPromoterpLuxAvailable SinceJan. 6, 2014AvailabilityAcademic Institutions and Nonprofits only -
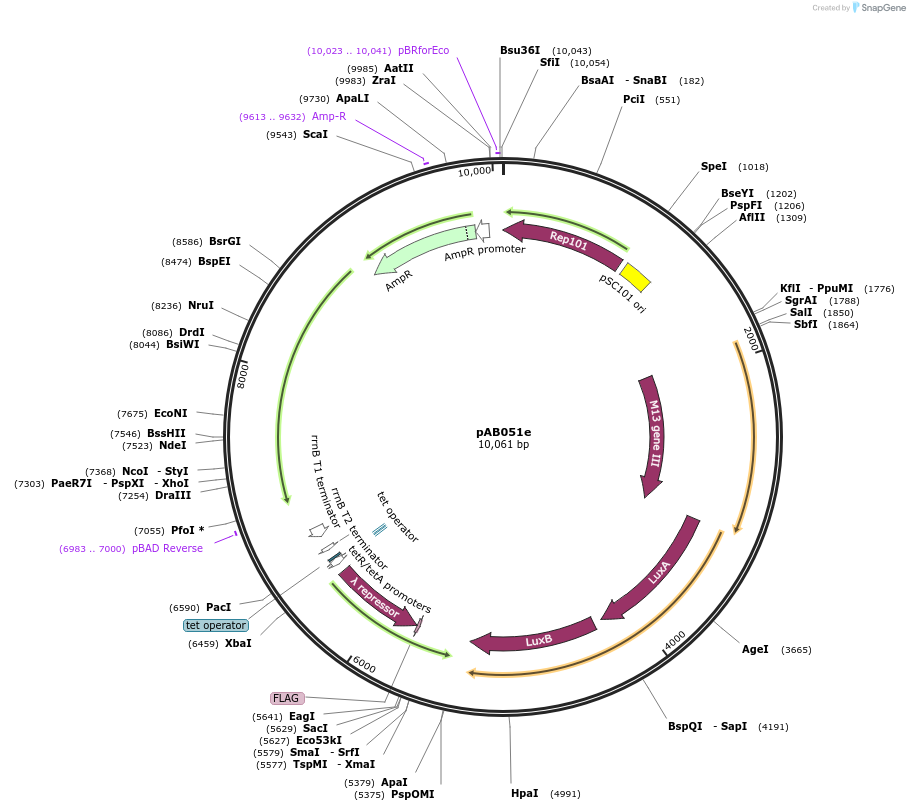
pAB051e
Plasmid#79195PurposeDirected Evolution of Protein-Protein InteractionsDepositorInsertPExt-10cons lambda op-62 gIII, luxAB; Ptet lambda cI-SH2ABL1
ExpressionBacterialPromoterPExt-10cons lambda op-62; PtetAvailabilityAcademic Institutions and Nonprofits only -
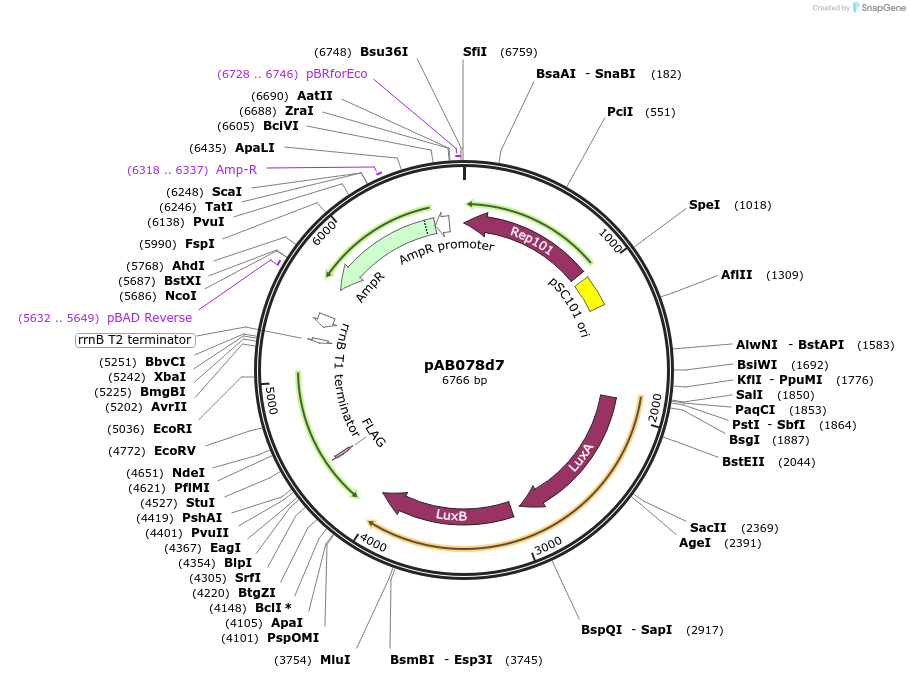
pAB078d7
Plasmid#79205PurposeDirected Evolution of Protein-Protein InteractionsDepositorInsertPlacZ-opt (off-target) luxAB; Ppro1 434cI-SH2ABL1
ExpressionBacterialPromoterPlacZ-opt (off-target); Ppro1AvailabilityAcademic Institutions and Nonprofits only