We narrowed to 20,805 results for: ATO
-
Plasmid#214332PurposeProtein overexpression in bacteriaDepositorInsertIL18
TagsHis-TEV and MycExpressionBacterialPromoterT7Available SinceFeb. 21, 2024AvailabilityAcademic Institutions and Nonprofits only -
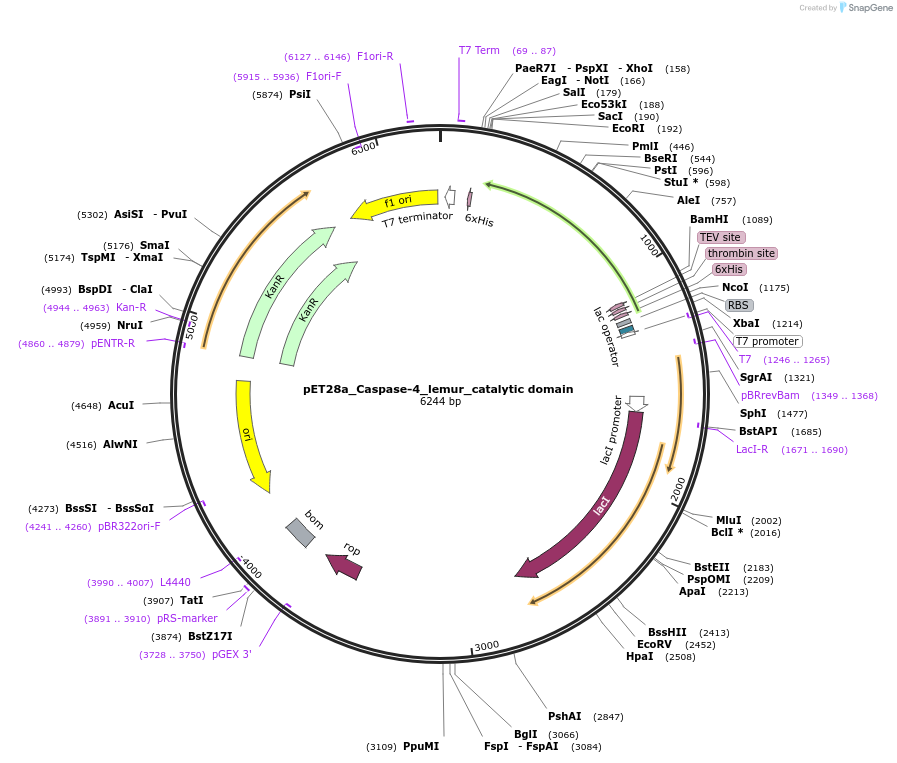
pET28a_Caspase-4_lemur_catalytic domain
Plasmid#214328PurposeProtein overexpression in bacteriaDepositorInsertCASP4
TagsHis-TEVExpressionBacterialPromoterT7Available SinceFeb. 21, 2024AvailabilityAcademic Institutions and Nonprofits only -
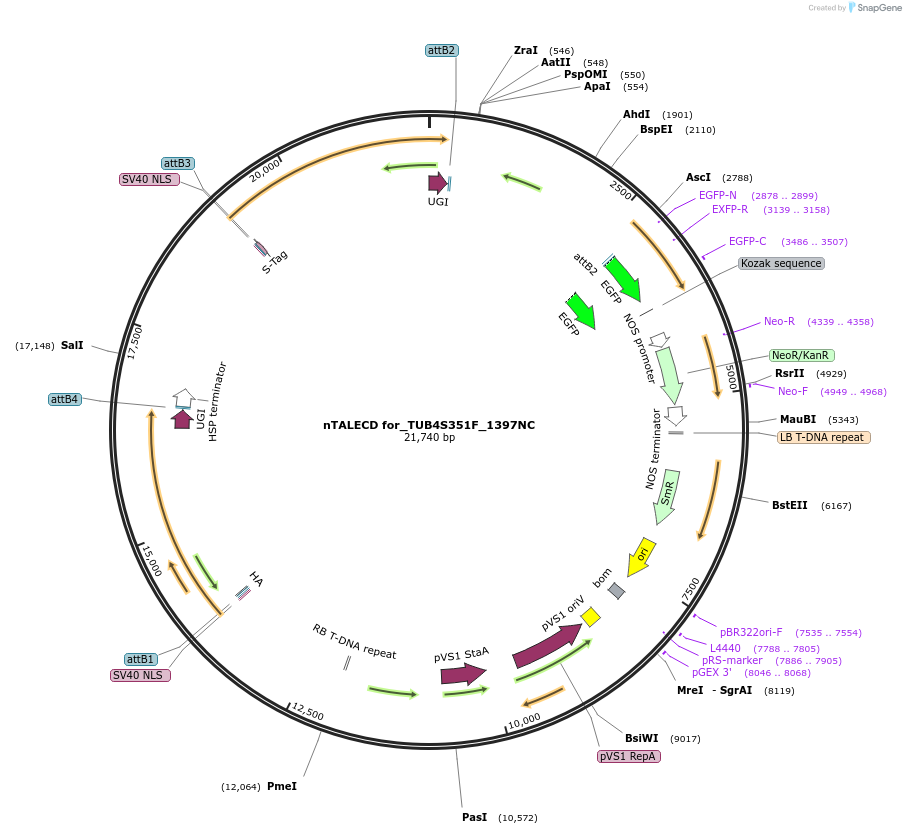
nTALECD for_TUB4S351F_1397NC
Plasmid#193382PurposenTALECD (1397NC) expression vectorDepositorInsertFor specific C-to-T base editing to introduce an amino acid mutation in the coding sequence of AtTUB4.
ExpressionPlantAvailable SinceFeb. 9, 2024AvailabilityAcademic Institutions and Nonprofits only -
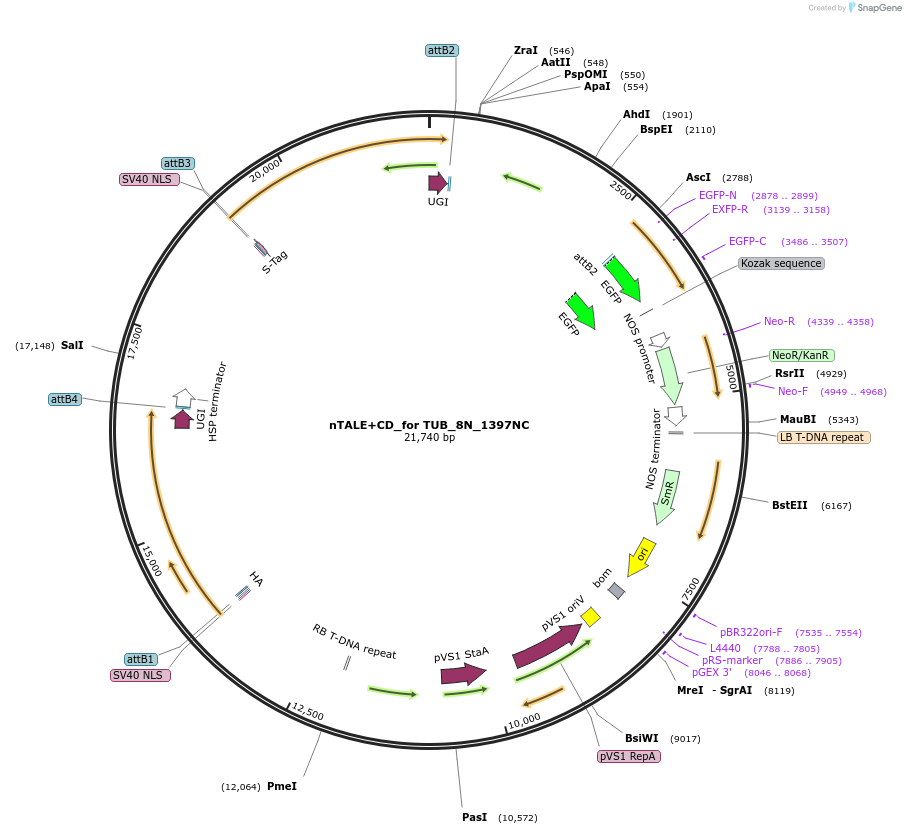
nTALE+CD_for TUB_8N_1397NC
Plasmid#193383PurposenTALECD (1397NC) expression vectorDepositorInsertFor multiplex C-to-T base editing to introduce amino acid mutations in the coding sequence of AtTUBs.
ExpressionPlantAvailable SinceFeb. 9, 2024AvailabilityAcademic Institutions and Nonprofits only -
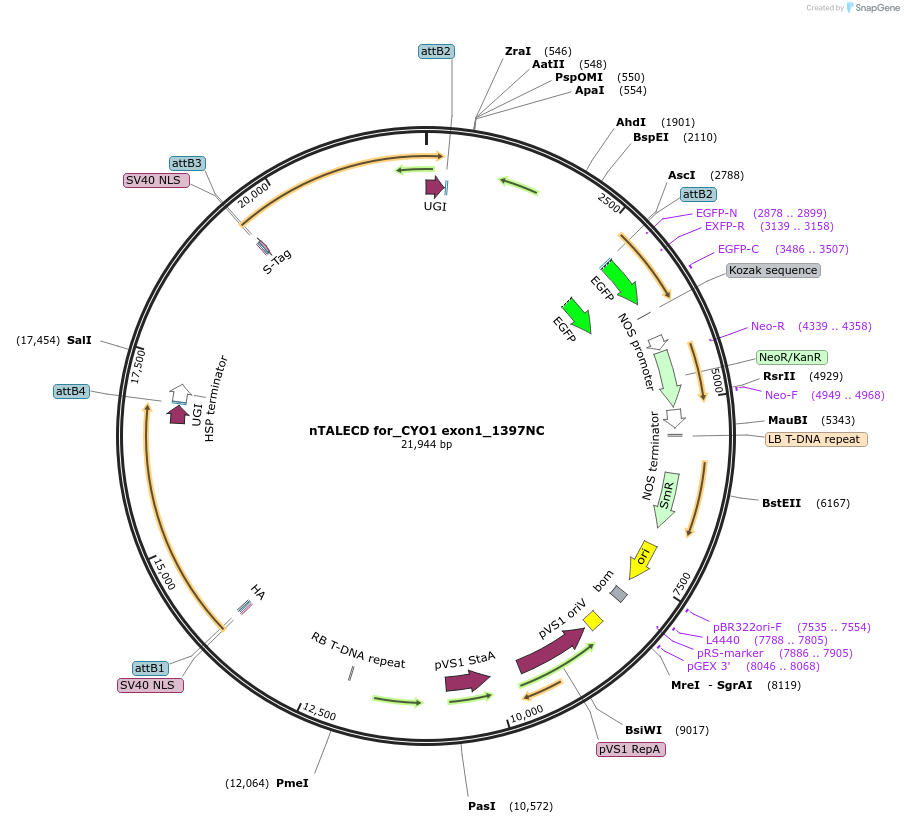
nTALECD for_CYO1 exon1_1397NC
Plasmid#193376PurposenTALECD (1397NC) expression vectorDepositorInsertFor specific C-to-T base editing to introduce a nonsense mutation in the coding sequence of AtCYO1.
ExpressionPlantAvailable SinceFeb. 9, 2024AvailabilityAcademic Institutions and Nonprofits only -
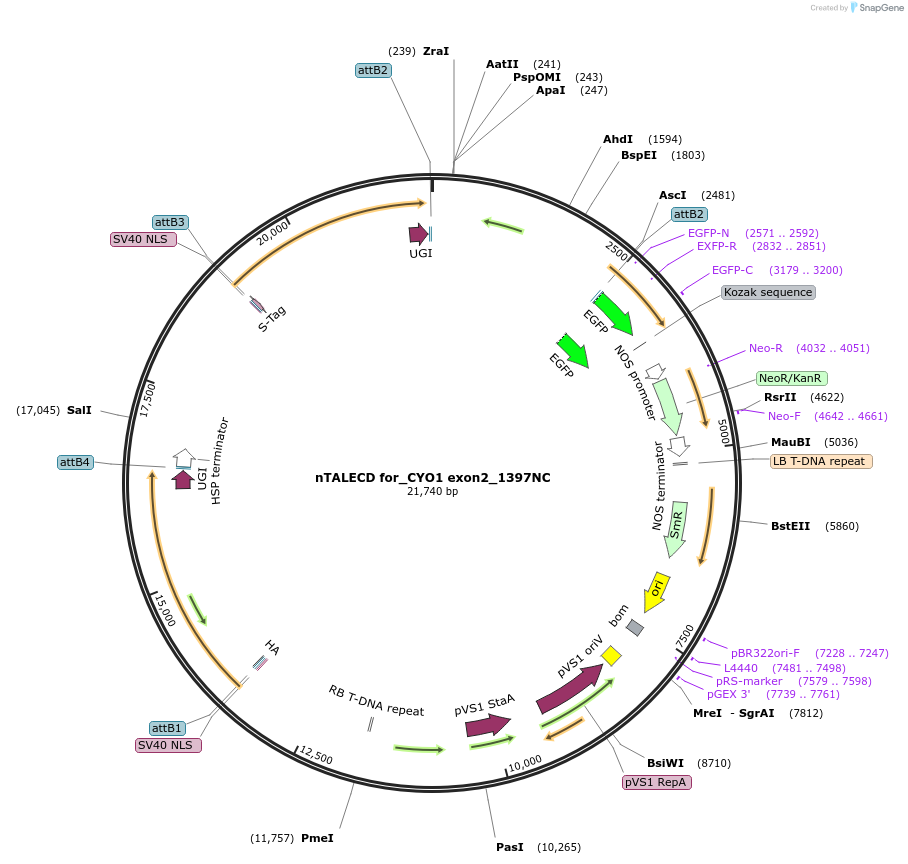
nTALECD for_CYO1 exon2_1397NC
Plasmid#193377PurposenTALECD (1397NC) expression vectorDepositorInsertFor specific C-to-T base editing to introduce a nonsense mutation in the coding sequence of AtCYO1.
ExpressionPlantAvailable SinceFeb. 9, 2024AvailabilityAcademic Institutions and Nonprofits only -
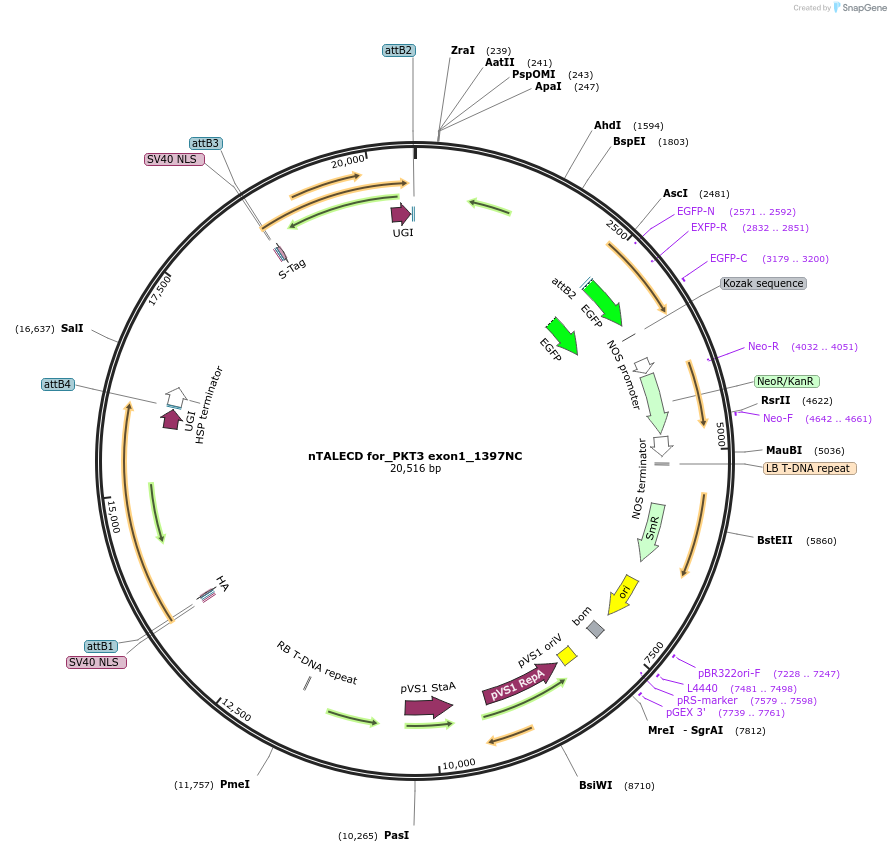
nTALECD for_PKT3 exon1_1397NC
Plasmid#193378PurposenTALECD (1397NC) expression vectorDepositorInsertFor specific C-to-T base editing to introduce a nonsense mutation in the coding sequence of AtPKT3.
ExpressionPlantAvailable SinceFeb. 9, 2024AvailabilityAcademic Institutions and Nonprofits only -
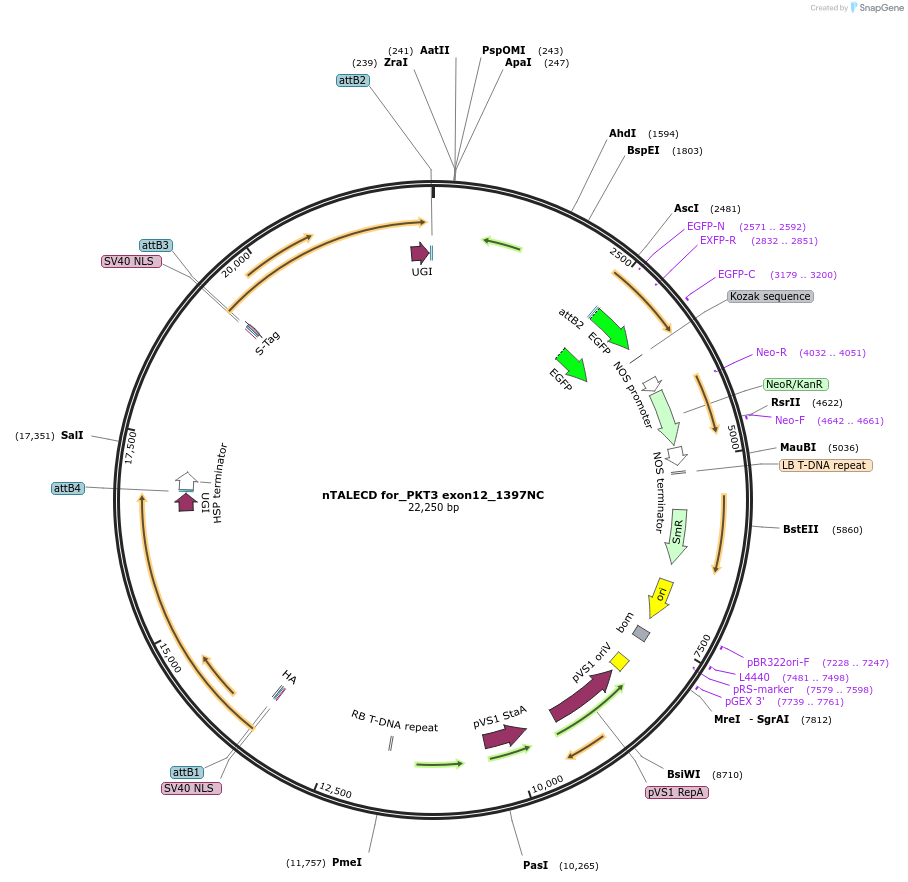
nTALECD for_PKT3 exon12_1397NC
Plasmid#193379PurposenTALECD (1397NC) expression vectorDepositorInsertFor specific C-to-T base editing to introduce an amino acid mutation in the coding sequence of AtPKT3.
ExpressionPlantAvailable SinceFeb. 9, 2024AvailabilityAcademic Institutions and Nonprofits only -
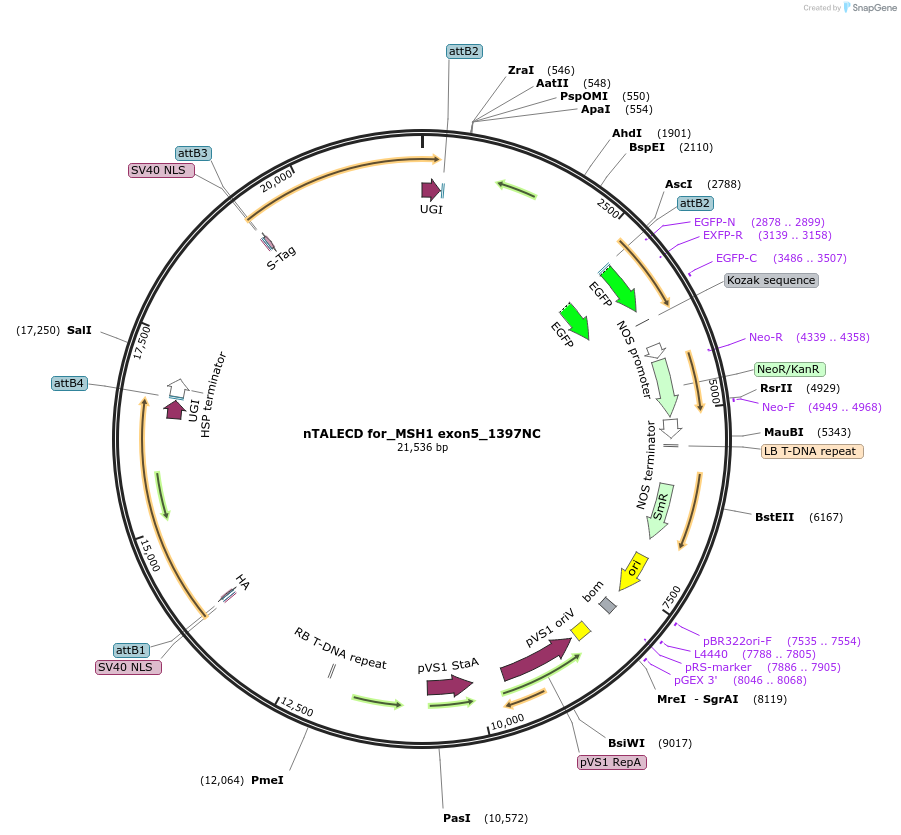
nTALECD for_MSH1 exon5_1397NC
Plasmid#193380PurposenTALECD (1397NC) expression vectorDepositorInsertFor specific C-to-T base editing to introduce an amino acid mutation in the coding sequence of AtMSH1.
ExpressionPlantAvailable SinceFeb. 9, 2024AvailabilityAcademic Institutions and Nonprofits only -
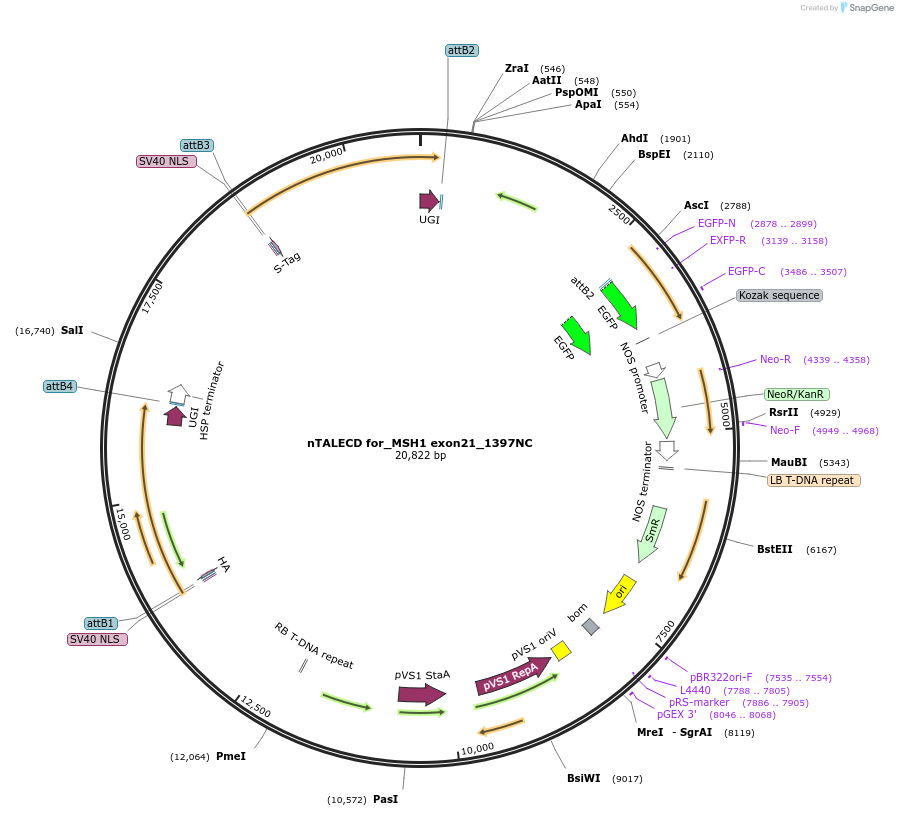
nTALECD for_MSH1 exon21_1397NC
Plasmid#193381PurposenTALECD (1397NC) expression vectorDepositorInsertFor specific C-to-T base editing to introduce an amino acid mutation in the coding sequence of AtMSH1.
ExpressionPlantAvailable SinceFeb. 9, 2024AvailabilityAcademic Institutions and Nonprofits only -
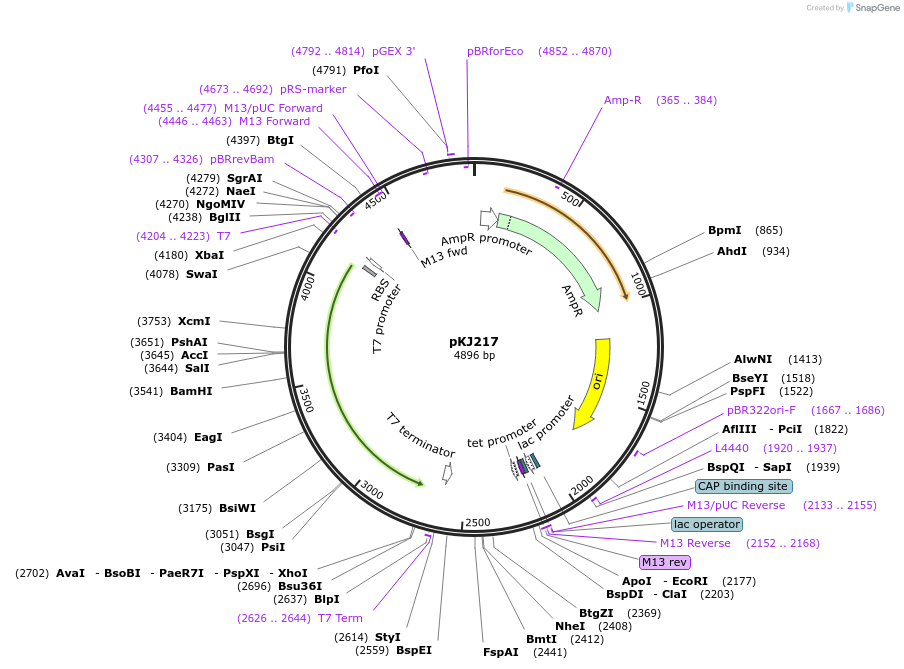
pKJ217
Plasmid#212318PurposeExpress MmFnuc with pureexpressDepositorInsertMercenaria
ExpressionBacterialAvailable SinceJan. 2, 2024AvailabilityAcademic Institutions and Nonprofits only -
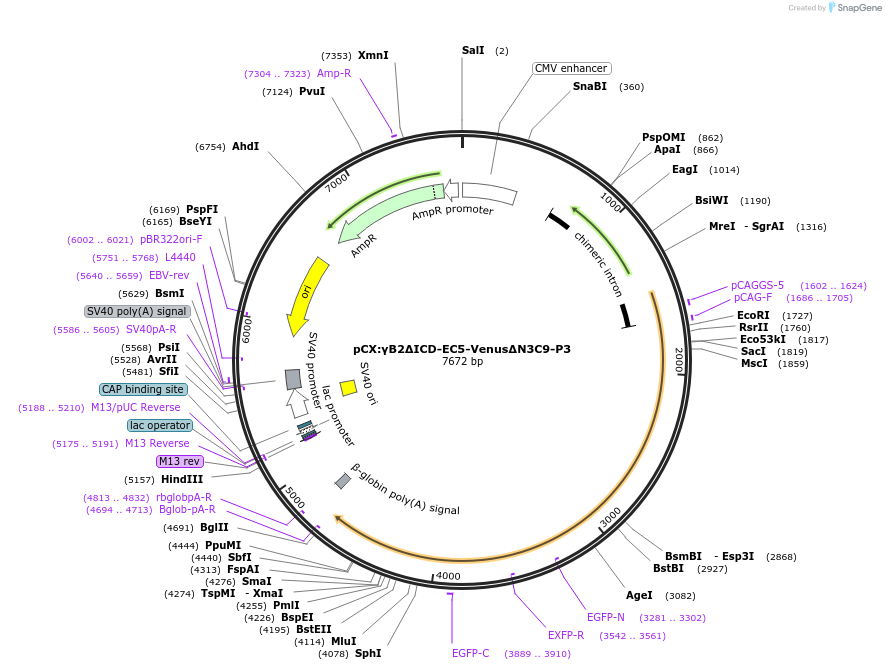
pCX:γB2ΔICD-EC5-VenusΔN3C9-P3
Plasmid#209090PurposemVenus-inserted PcdhgB2 lacking an intracellular domain (ICD)DepositorAvailable SinceDec. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
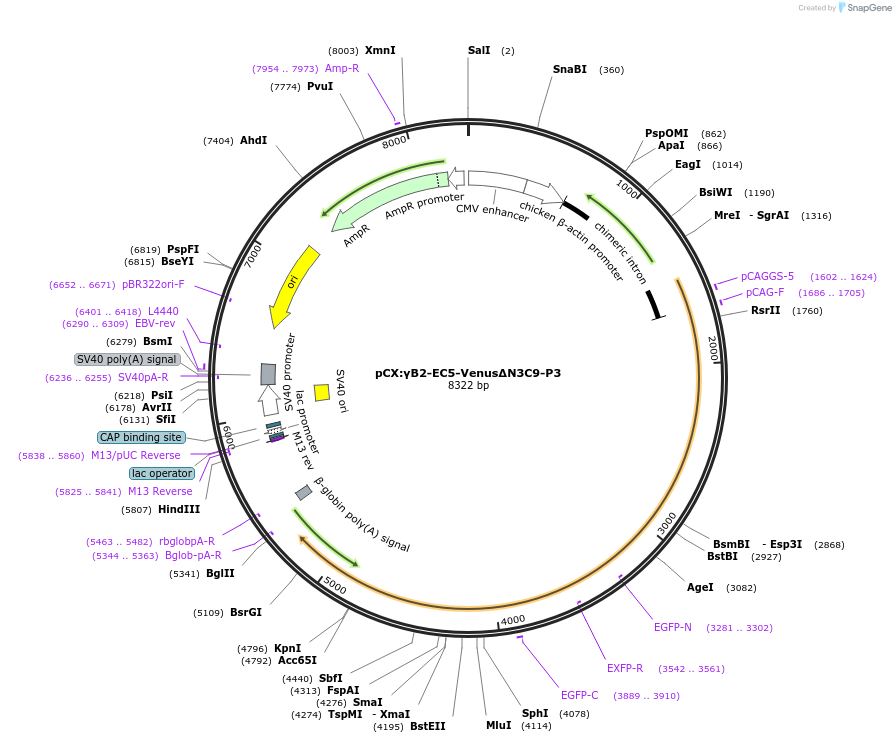
pCX:γB2-EC5-VenusΔN3C9-P3
Plasmid#209092PurposemVenus-inserted PcdhgB2DepositorAvailable SinceDec. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
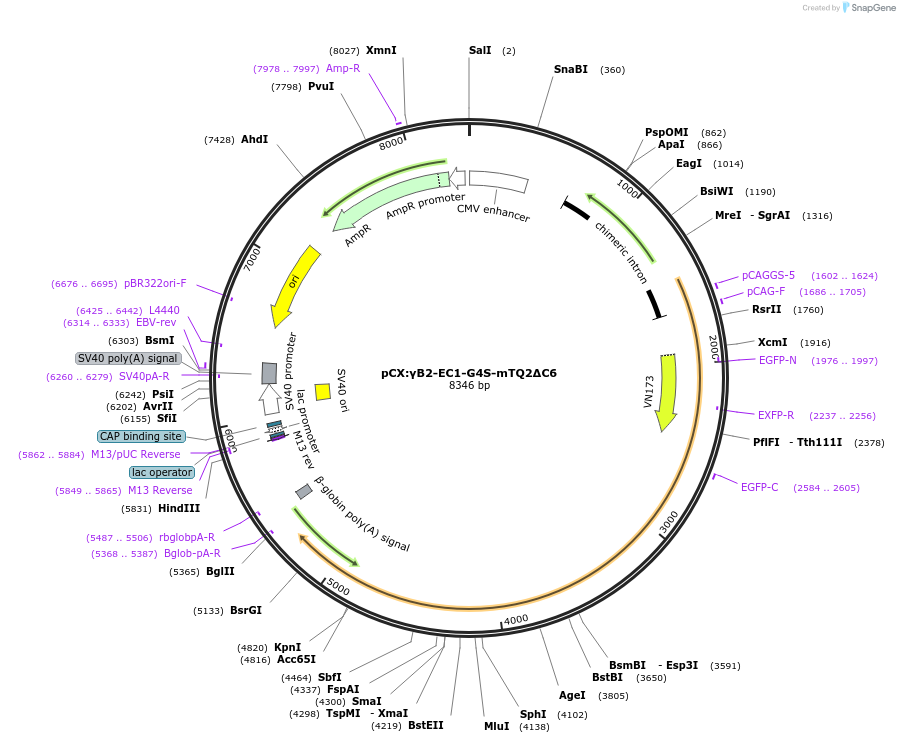
pCX:γB2-EC1-G4S-mTQ2ΔC6
Plasmid#209091PurposemTurquoise2-inserted PcdhgB2DepositorAvailable SinceDec. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
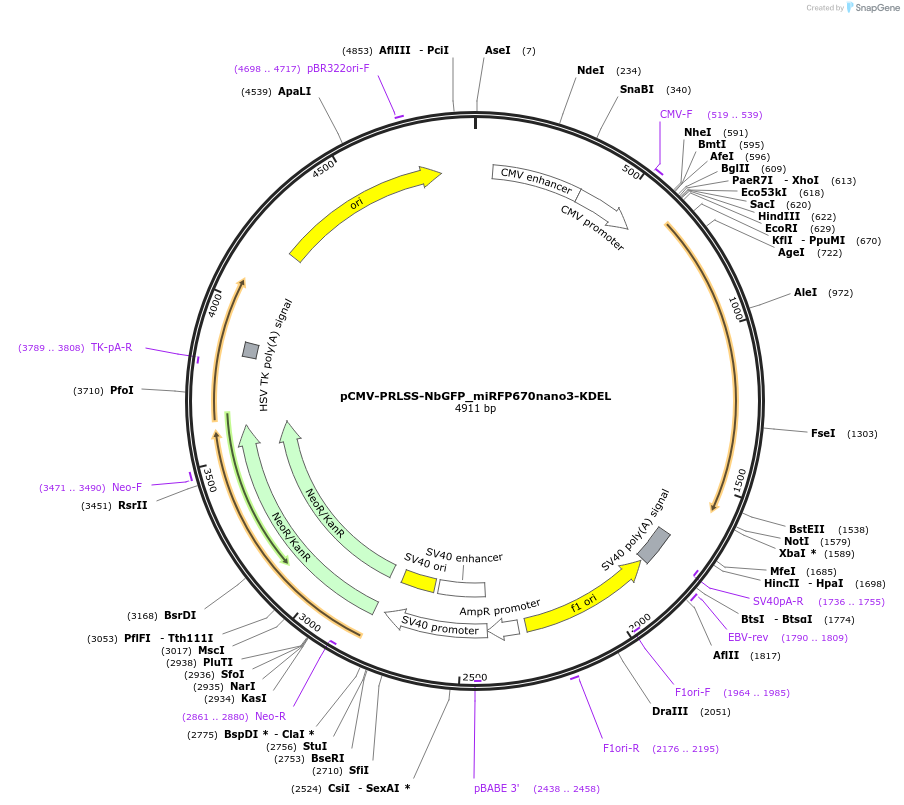
pCMV-PRLSS-NbGFP_miRFP670nano3-KDEL
Plasmid#209718PurposeMammalian expression of NbGFP-miRFP670nano3 localized to endoplasmic reticulumDepositorInsertPRLSS-NbGFP_miRFP670nano3-KDEL
TagsPRLSSExpressionMammalianAvailable SinceNov. 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
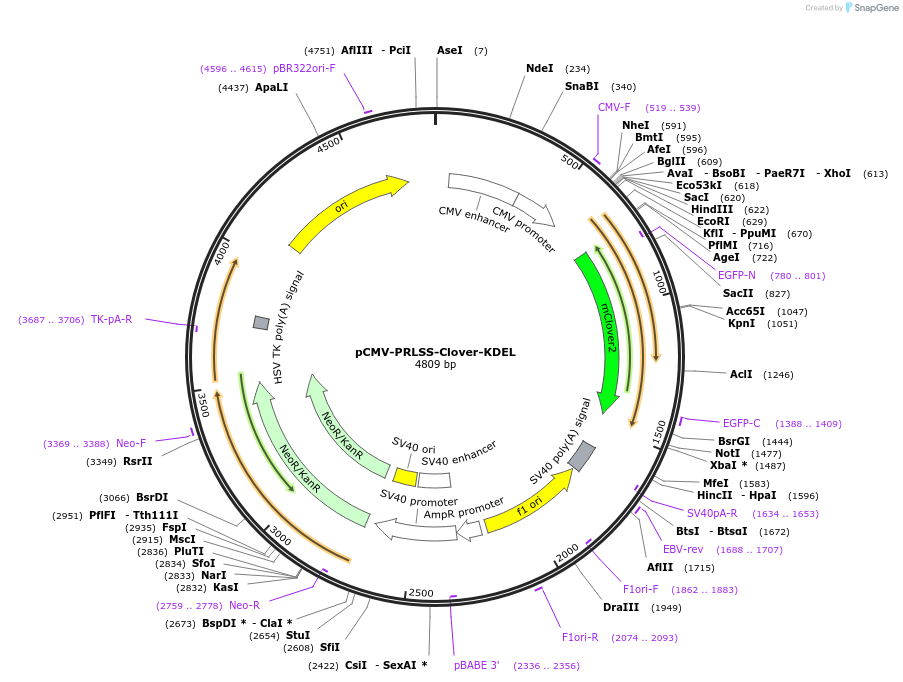
pCMV-PRLSS-Clover-KDEL
Plasmid#209720PurposeMammalian expression of clover localized to endoplasmic reticulumDepositorInsertPRLSS-Clover-KDEL
TagsPRLSSExpressionMammalianAvailable SinceNov. 2, 2023AvailabilityAcademic Institutions and Nonprofits only -
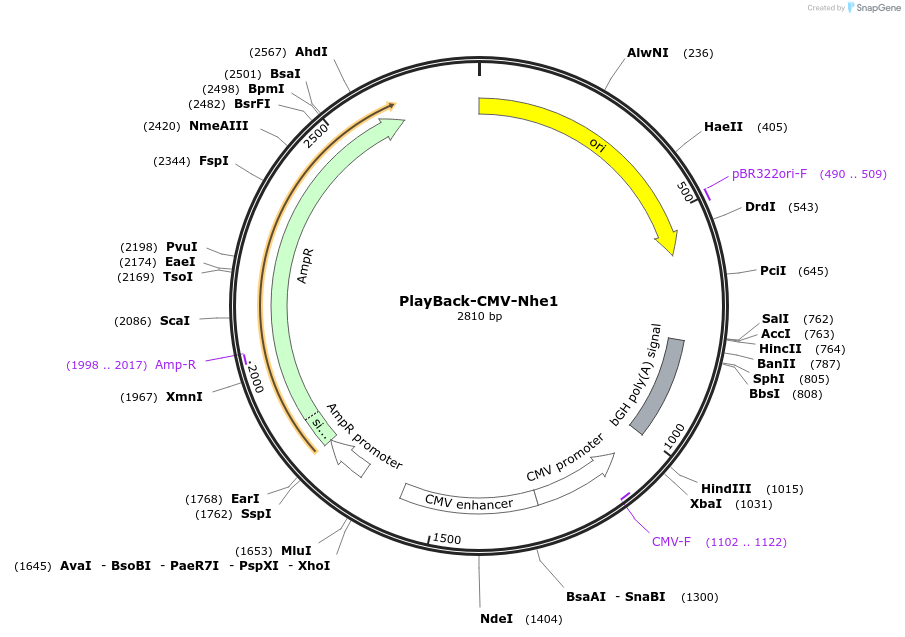
PlayBack-CMV-Nhe1
Plasmid#203304PurposeFor expression of eukaryotic genes of interest with CMV promoter with NheI as a PlayBack Restriction EnzymeDepositorTypeEmpty backboneExpressionMammalianAvailable SinceOct. 16, 2023AvailabilityAcademic Institutions and Nonprofits only -
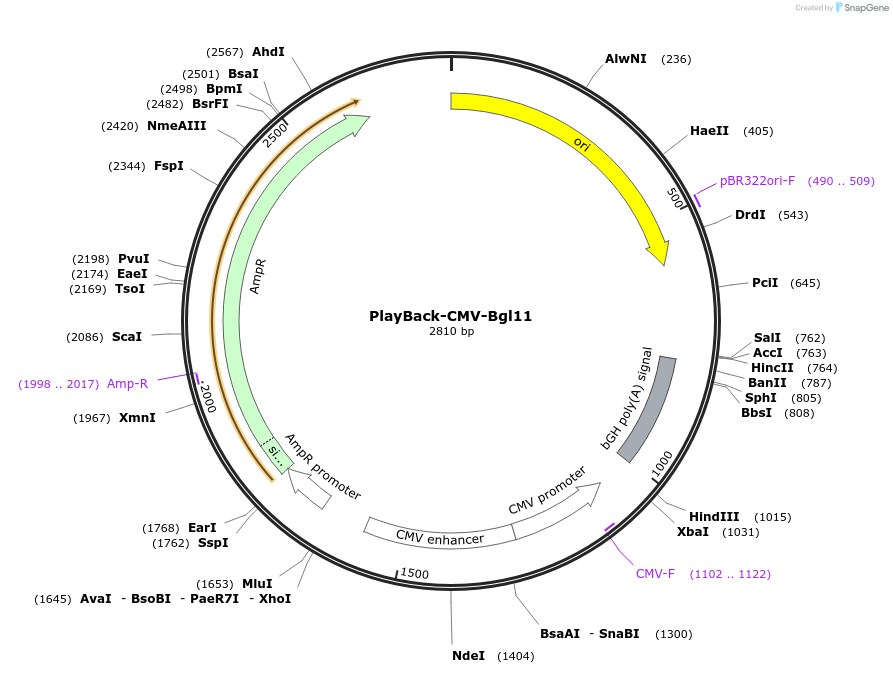
PlayBack-CMV-Bgl11
Plasmid#203306PurposeFor expression of eukaryotic genes of interest with CMV promoter with BglII as a PlayBack Restriction EnzymeDepositorTypeEmpty backboneExpressionMammalianAvailable SinceOct. 16, 2023AvailabilityAcademic Institutions and Nonprofits only -
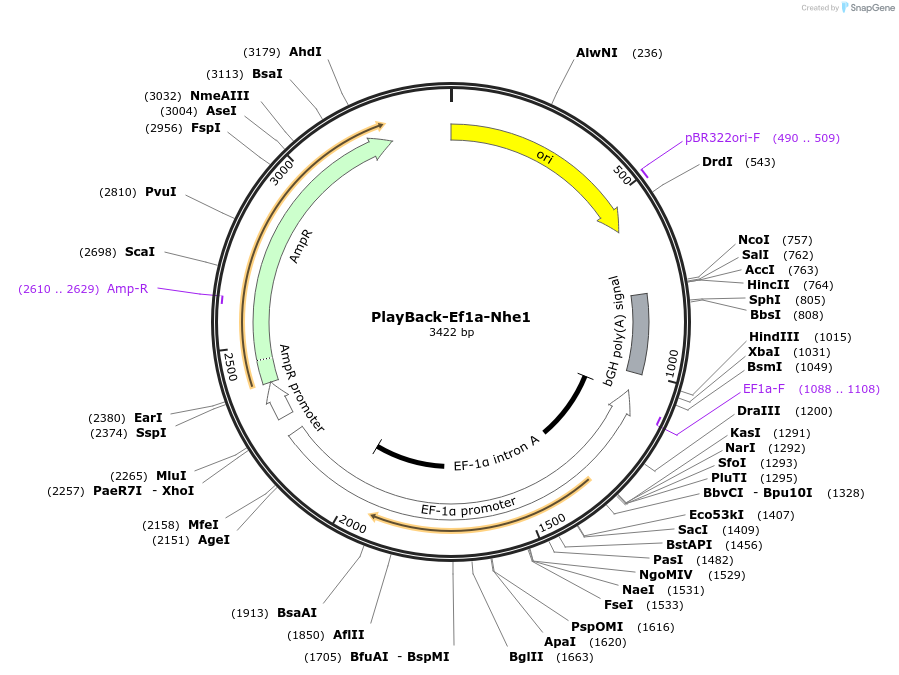
PlayBack-Ef1a-Nhe1
Plasmid#203310PurposeFor expression of eukaryotic genes of interest with Ef1a promoter with NheI as a PlayBack Restriction EnzymeDepositorTypeEmpty backboneExpressionMammalianAvailable SinceOct. 16, 2023AvailabilityAcademic Institutions and Nonprofits only -
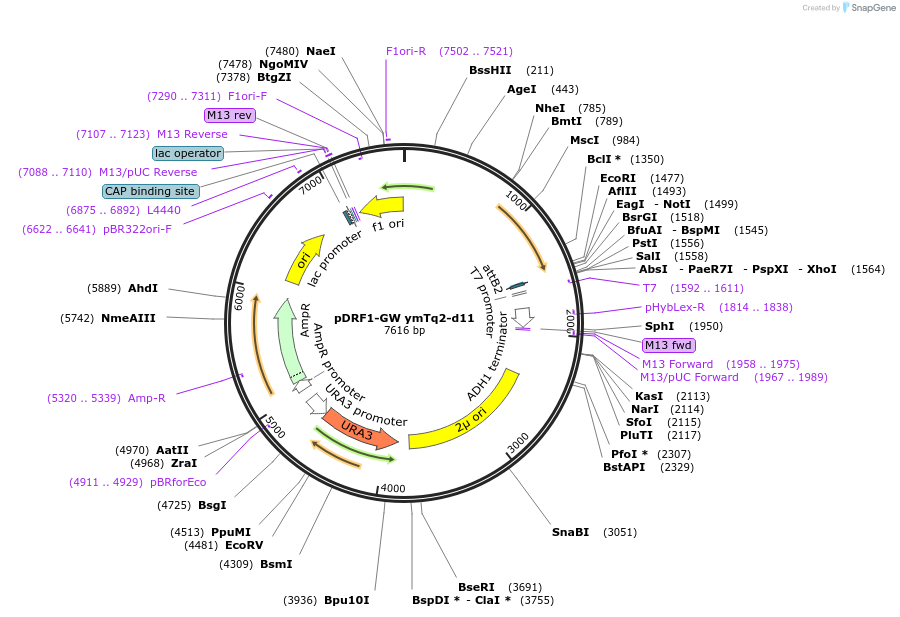
pDRF1-GW ymTq2-d11
Plasmid#205490Purposeexpress ymTq2-d11 as donor control for yAT1.03DepositorInsertymTq2-d11
ExpressionYeastAvailable SinceSept. 14, 2023AvailabilityAcademic Institutions and Nonprofits only -
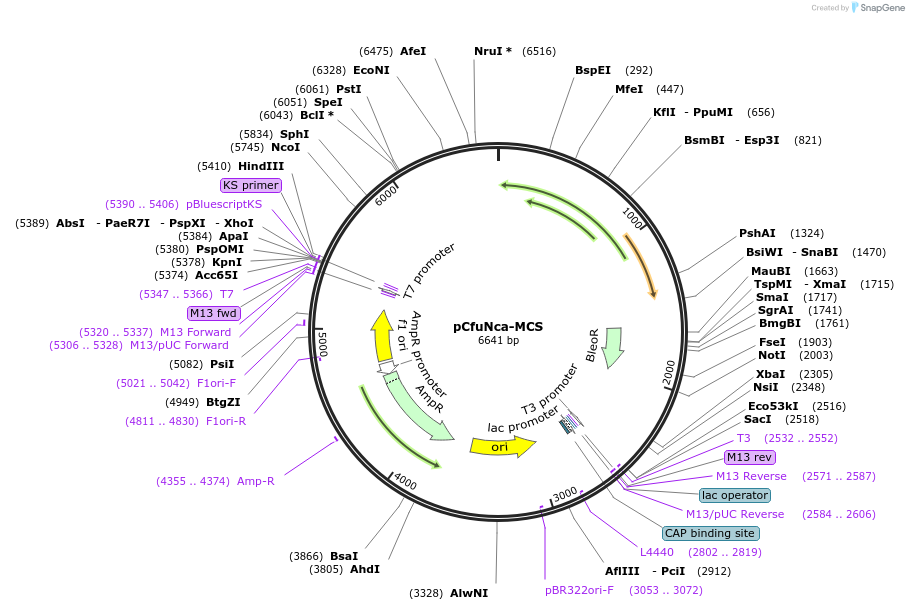
pCfuNca-MCS
Plasmid#190820PurposeMultiple cloning site between Nitzschia captiva promoter and termination regionsDepositorInsertsMultiple cloning site with SpeI and PstI
Bleomycin resistance gene
ExpressionBacterialAvailable SinceAug. 14, 2023AvailabilityAcademic Institutions and Nonprofits only -
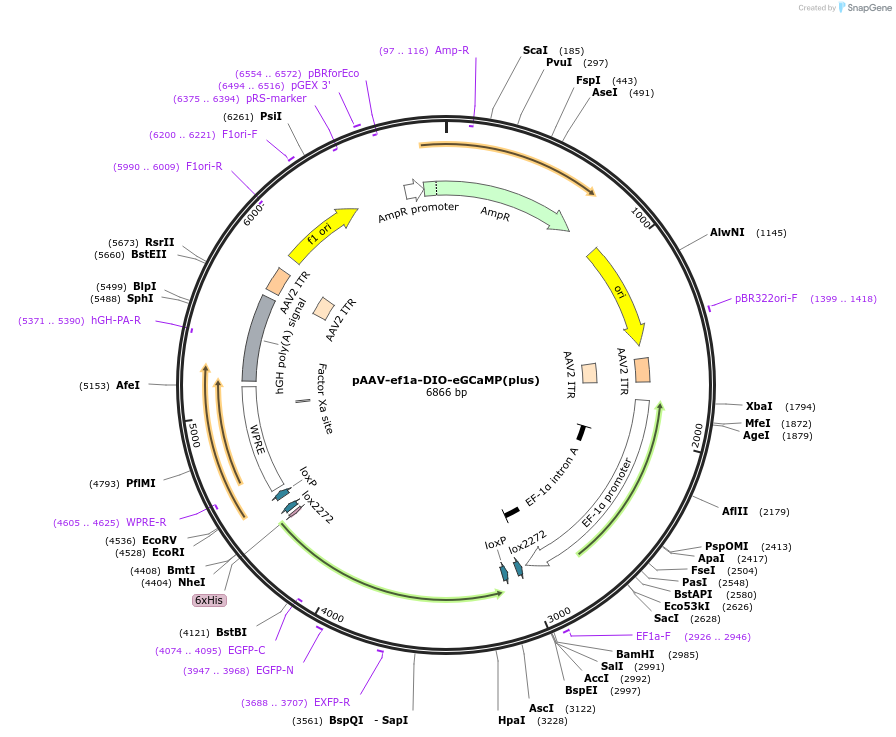
pAAV-ef1a-DIO-eGCaMP(plus)
Plasmid#201149PurposeAAV virus production for Cre recombinase dependent expression of eGCaMP(plus)DepositorInserteGCaMP+
UseAAV, Cre/Lox, and Mouse TargetingExpressionMammalianAvailable SinceJune 27, 2023AvailabilityAcademic Institutions and Nonprofits only -
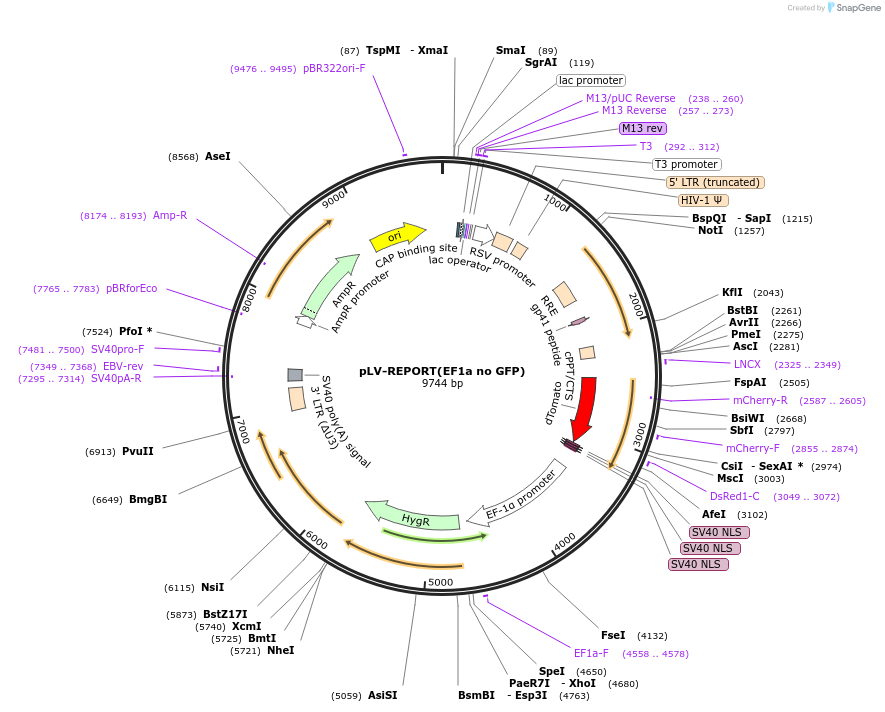
pLV-REPORT(EF1a no GFP)
Plasmid#172339PurposemnucTomatoDepositorInsertsmonomeric nuclear Tomato
HygR
UseLentiviral and Synthetic BiologyTags3x SV40 NLSExpressionMammalianPromoterEF1alphaAvailable SinceJune 26, 2023AvailabilityAcademic Institutions and Nonprofits only -
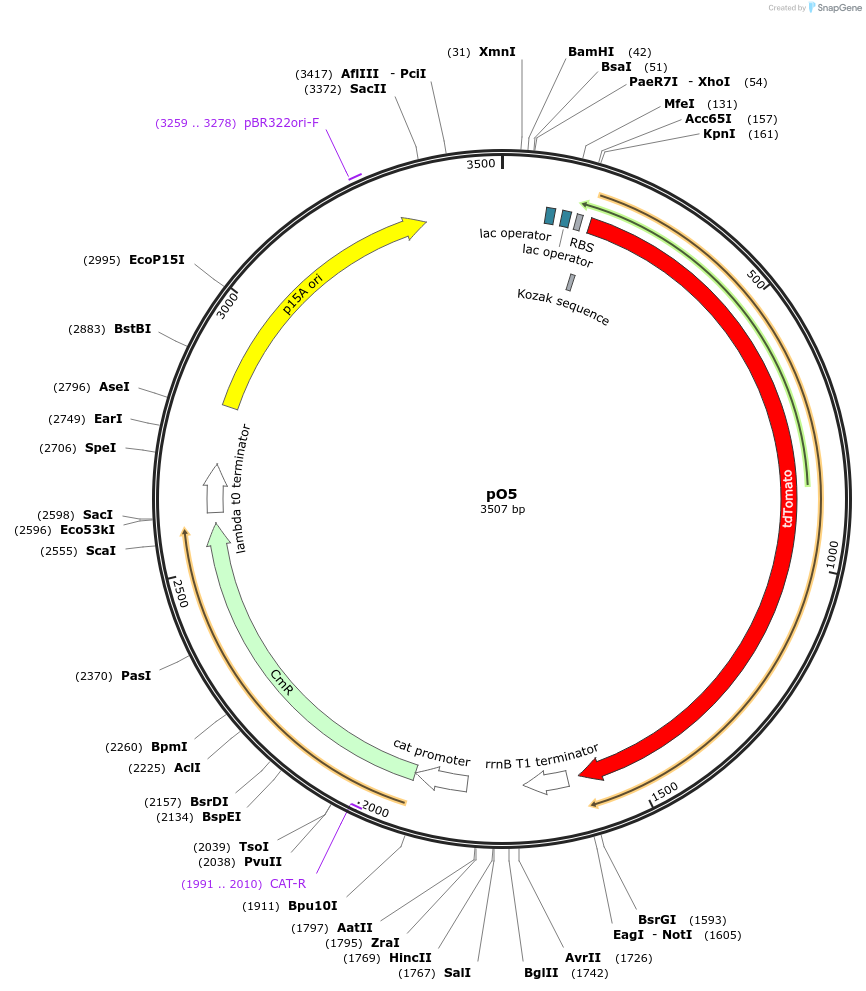
pO5
Plasmid#193749PurposeExpresses tdTomato under the control of PIAH synthetic promoter regulated by AHL and IPTG, p15A origin of replication, Chloramphenicol selectionDepositorInserttdTomato
ExpressionBacterialPromoterPIAHAvailable SinceMay 22, 2023AvailabilityAcademic Institutions and Nonprofits only -
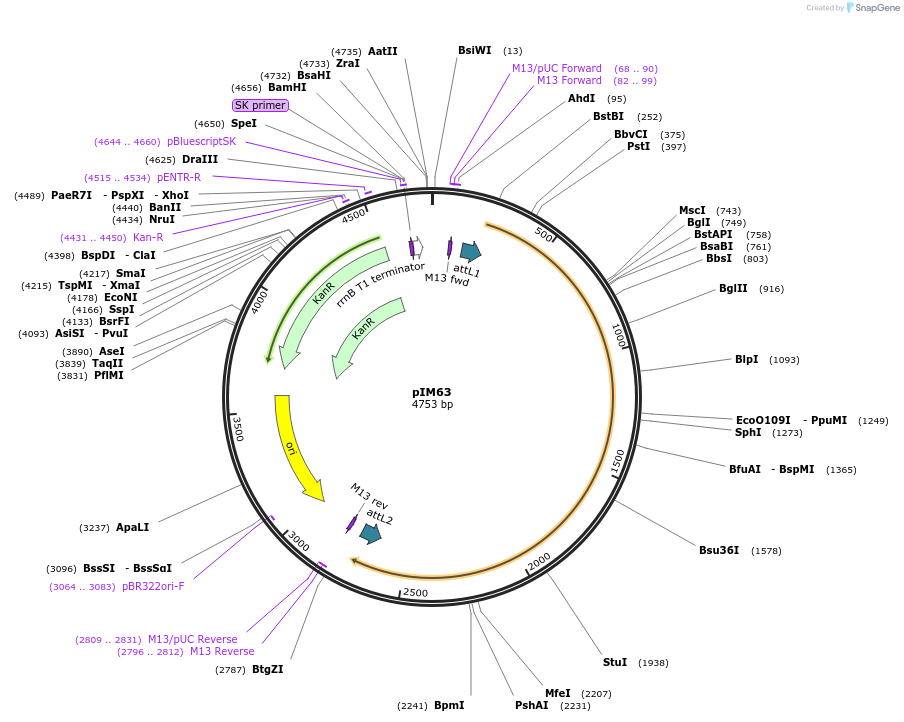
pIM63
Plasmid#193545PurposepENTRY clone for NLR Integrated Domain ID#19DepositorInsertID#19
UsePentry cloneAvailable SinceFeb. 14, 2023AvailabilityAcademic Institutions and Nonprofits only -
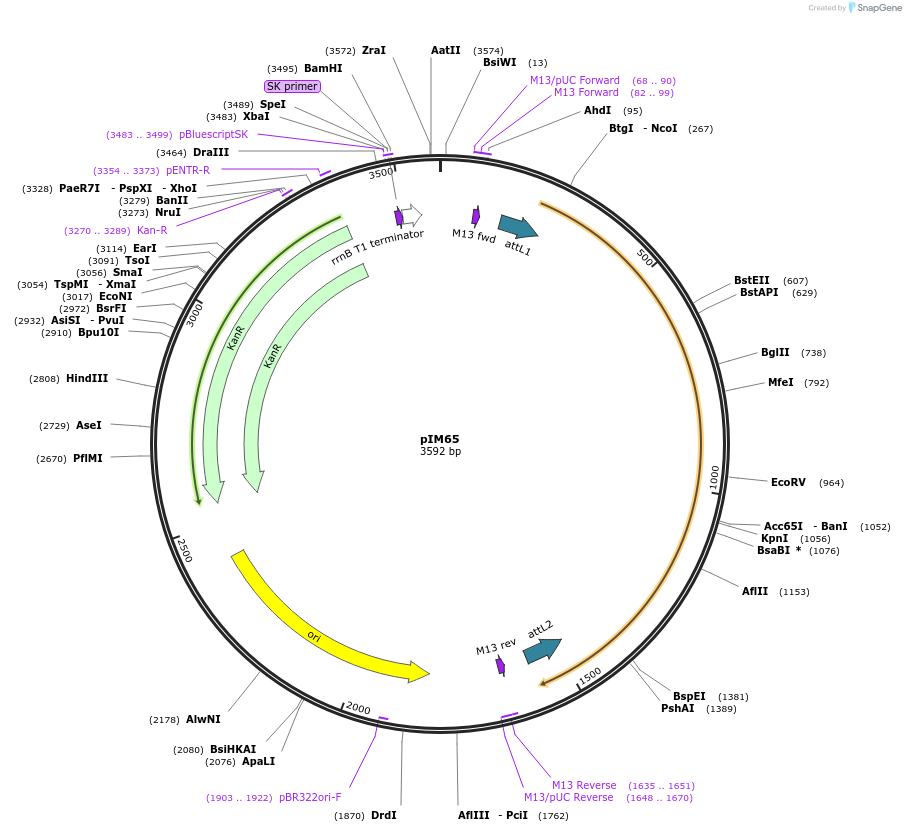
pIM65
Plasmid#193547PurposepENTRY clone for NLR Integrated Domain ID#14DepositorInsertID#14
UsePentry cloneAvailable SinceJan. 25, 2023AvailabilityAcademic Institutions and Nonprofits only -
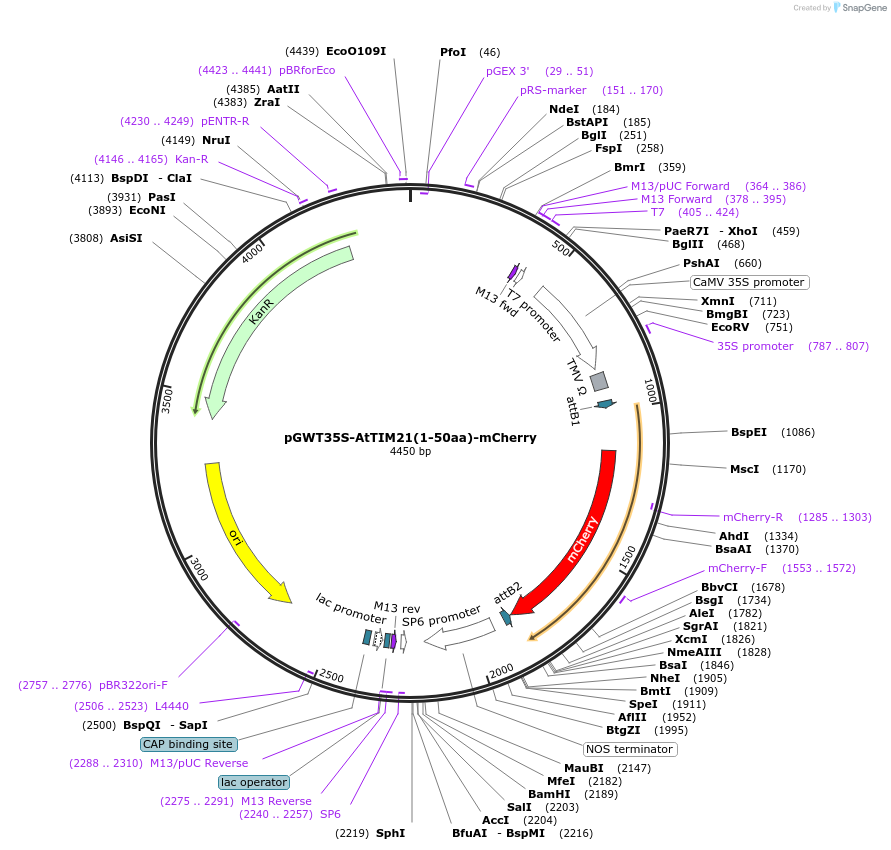
pGWT35S-AtTIM21(1-50aa)-mCherry
Plasmid#194047PurposeTransient expression of AtTIM21-mCherry in plant cell (Mitochondria)DepositorInsertmCherry
TagsAtTIM21 1–50aaExpressionPlantPromoterCaMV 35S promoterAvailable SinceJan. 18, 2023AvailabilityAcademic Institutions and Nonprofits only -
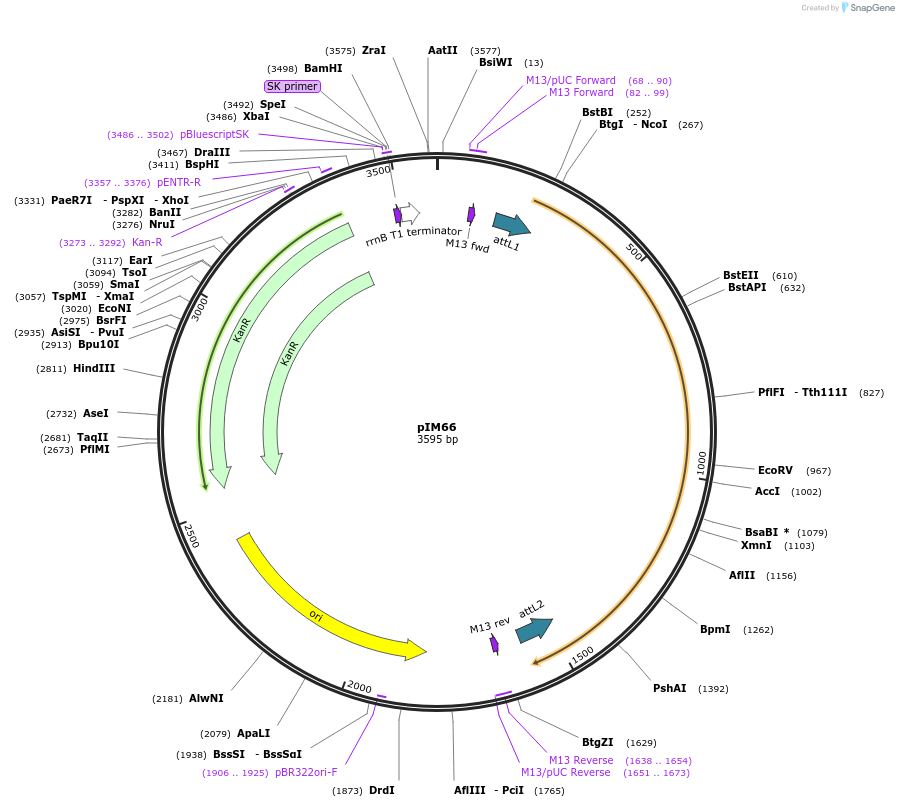
pIM66
Plasmid#193548PurposepENTRY clone for NLR Integrated Domain ID#12DepositorInsertID#12
UsePentry cloneAvailable SinceDec. 2, 2022AvailabilityAcademic Institutions and Nonprofits only -
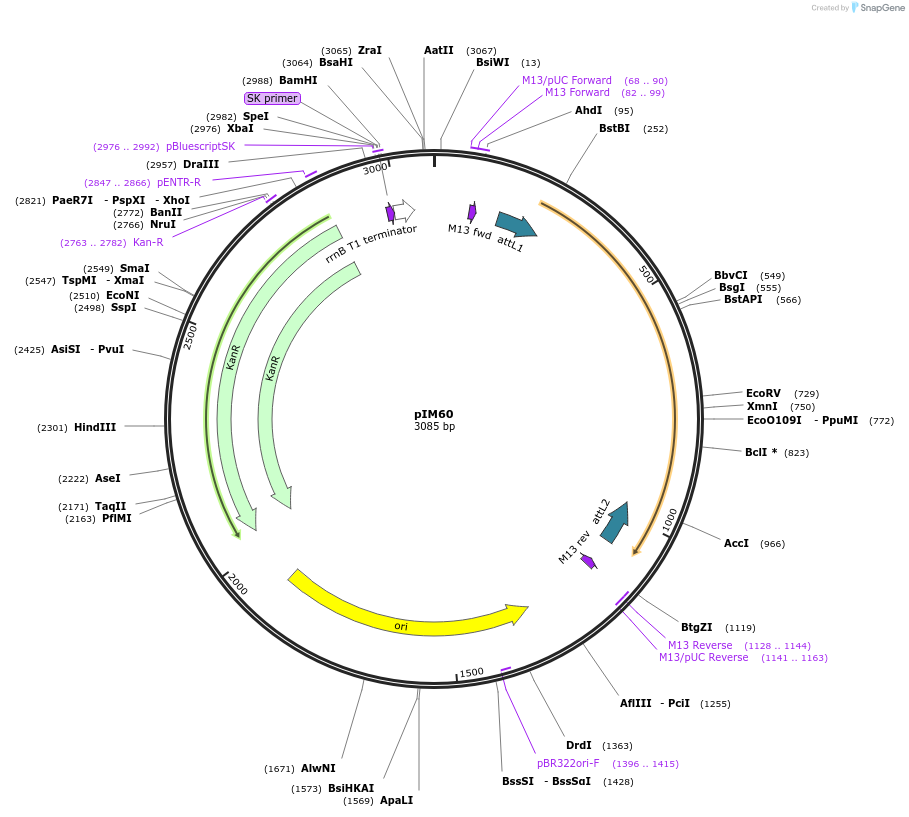
pIM60
Plasmid#193542PurposepENTRY clone for NLR Integrated Domain ID#22DepositorInsertID#22
UsePentry cloneAvailable SinceDec. 2, 2022AvailabilityAcademic Institutions and Nonprofits only -
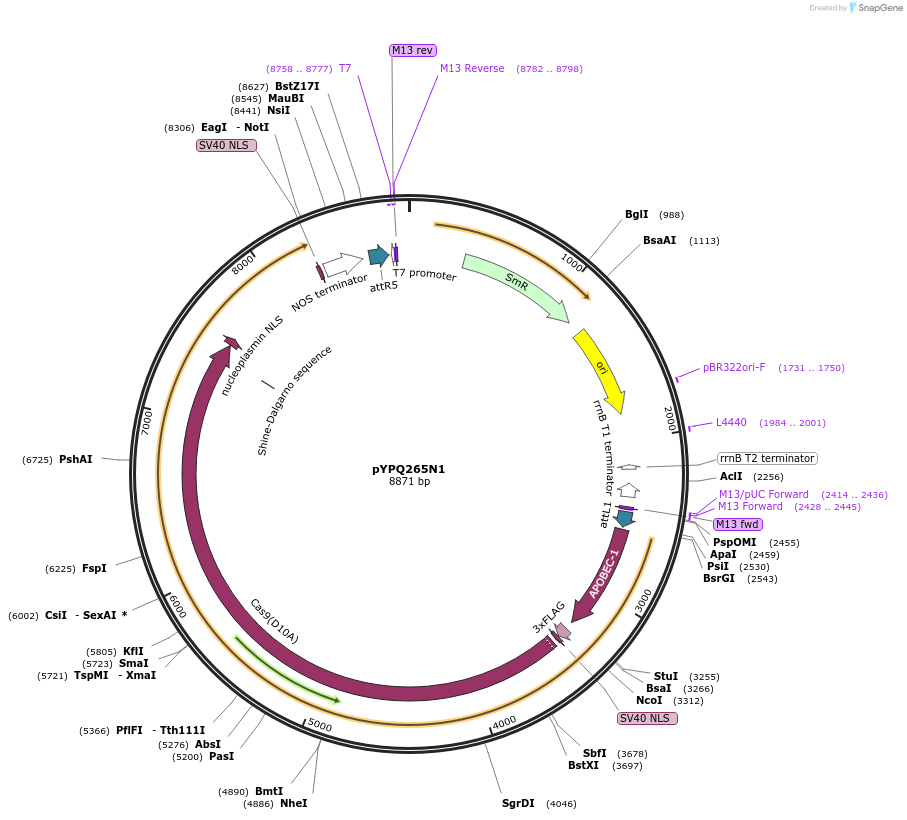
pYPQ265N1
Plasmid#173998PurposeGateway entry clone (attL1 & attR5) rAPOBEC1-zCas9(D10A)-UNG for C-G base editingDepositorInsertrAPOBEC1-zCas9(D10A)-UNG
UseCRISPRTags3X FLAG, NLSExpressionPlantMutationD10AAvailable SinceOct. 17, 2022AvailabilityAcademic Institutions and Nonprofits only -
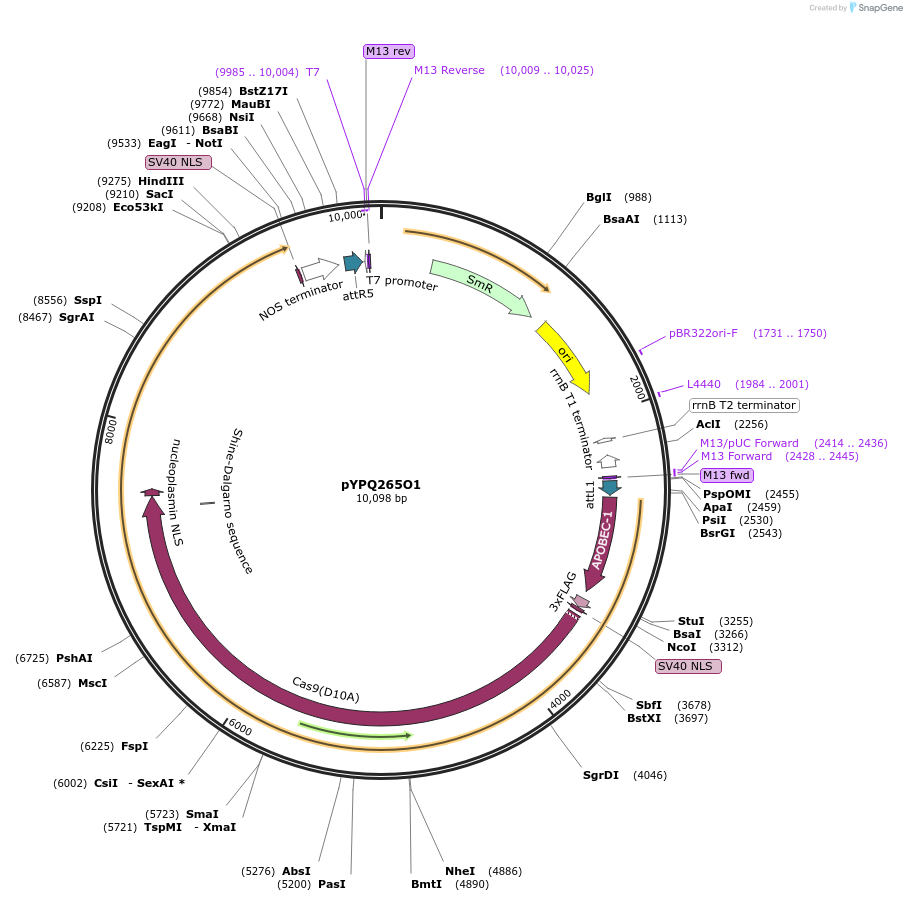
pYPQ265O1
Plasmid#173999PurposeGateway entry clone (attL1 & attR5) rAPOBEC1-zCas9(D10A)-rXRCC1 for C-G base editingDepositorInsertrAPOBEC1-zCas9(D10A)-rXRCC1
UseCRISPRTags3X FLAG, NLSExpressionPlantMutationD10AAvailable SinceOct. 17, 2022AvailabilityAcademic Institutions and Nonprofits only -
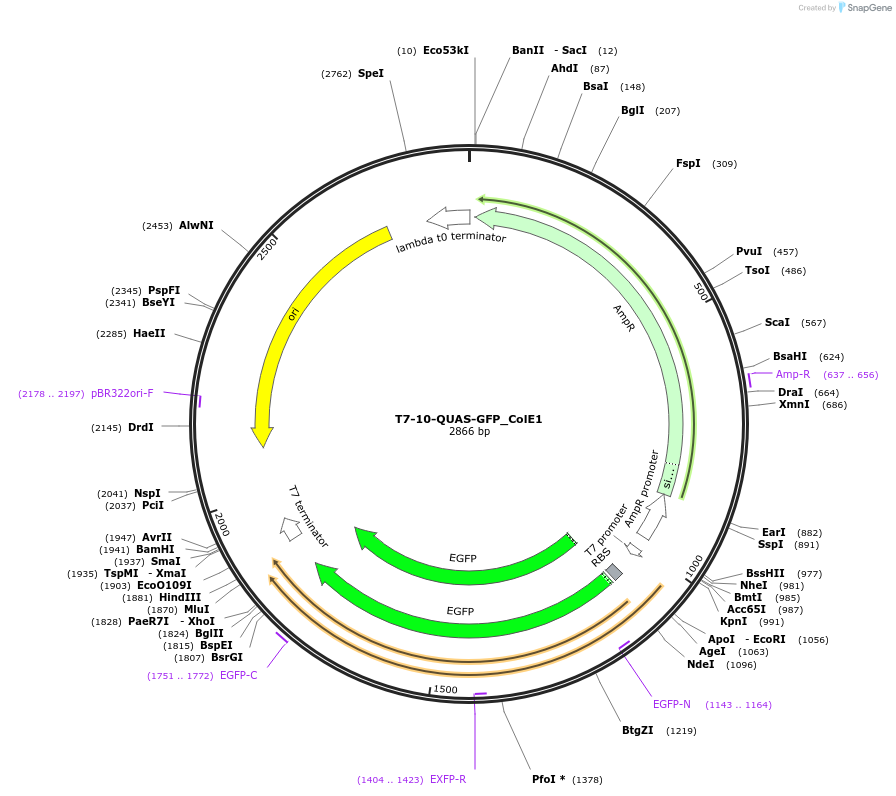
T7-10-QUAS-GFP_ColE1
Plasmid#171657PurposeQUAS ten base pairs downstream of a T7 promoter. Expresses GFP in BL21(DE3) E. coli. Contains ColE1 origin of replication.DepositorInsertT7-10-QUAS
TagsssrA degradation tag (DAS+4)ExpressionBacterialPromoterT7Available SinceOct. 14, 2022AvailabilityAcademic Institutions and Nonprofits only -
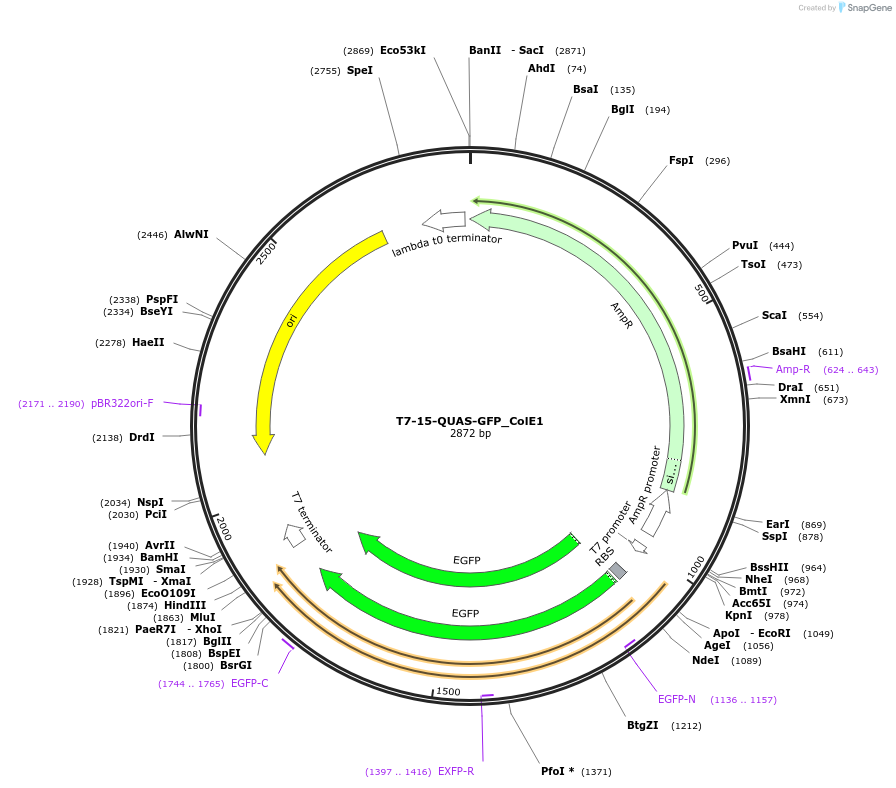
T7-15-QUAS-GFP_ColE1
Plasmid#171658PurposeQUAS fifteen base pairs downstream of a T7 promoter. Expresses GFP in BL21(DE3) E. coli. Contains ColE1 origin of replication.DepositorInsertT7-15-QUAS
TagsssrA degradation tag (DAS+4)ExpressionBacterialPromoterT7Available SinceOct. 14, 2022AvailabilityAcademic Institutions and Nonprofits only -
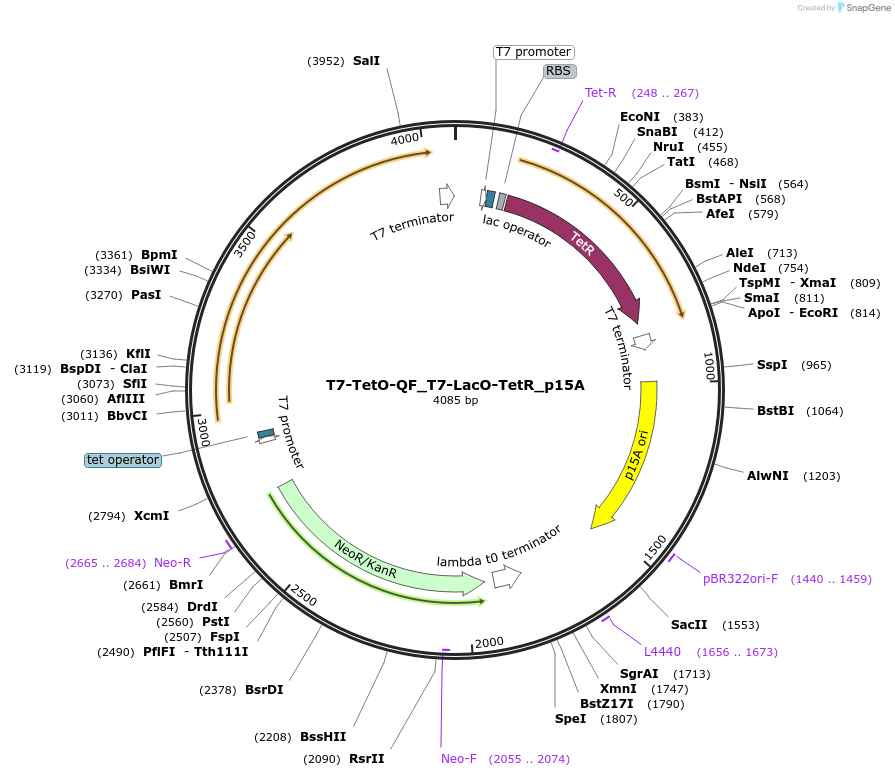
T7-TetO-QF_T7-LacO-TetR_p15A
Plasmid#171659PurposeExpresses QF and/or TetR in BL21(DE3) E. coli according to the presence of aTc. Contains a p15A origin of replication.DepositorInsertT7-LacO-TetR-T7 Stop
ExpressionBacterialPromoterT7Available SinceOct. 14, 2022AvailabilityAcademic Institutions and Nonprofits only -
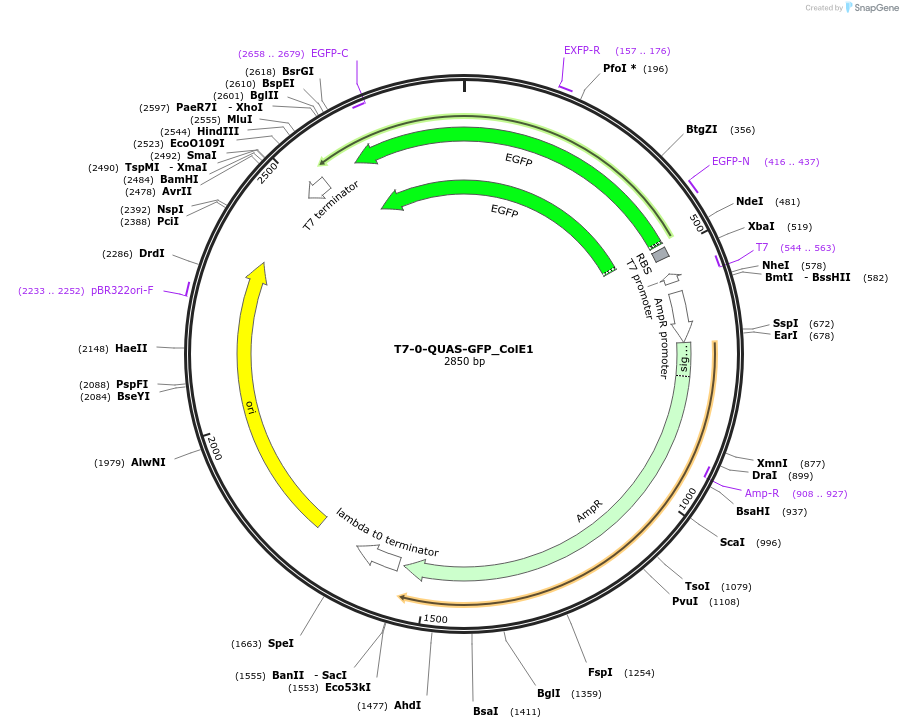
T7-0-QUAS-GFP_ColE1
Plasmid#171650PurposeQUAS directly downstream of a T7 promoter. Expresses GFP in BL21(DE3) E. coli. Contains ColE1 origin of replication.DepositorInsertT7-0-QUAS
TagsssrA degradation tag (DAS+4)ExpressionBacterialPromoterT7Available SinceOct. 14, 2022AvailabilityAcademic Institutions and Nonprofits only -
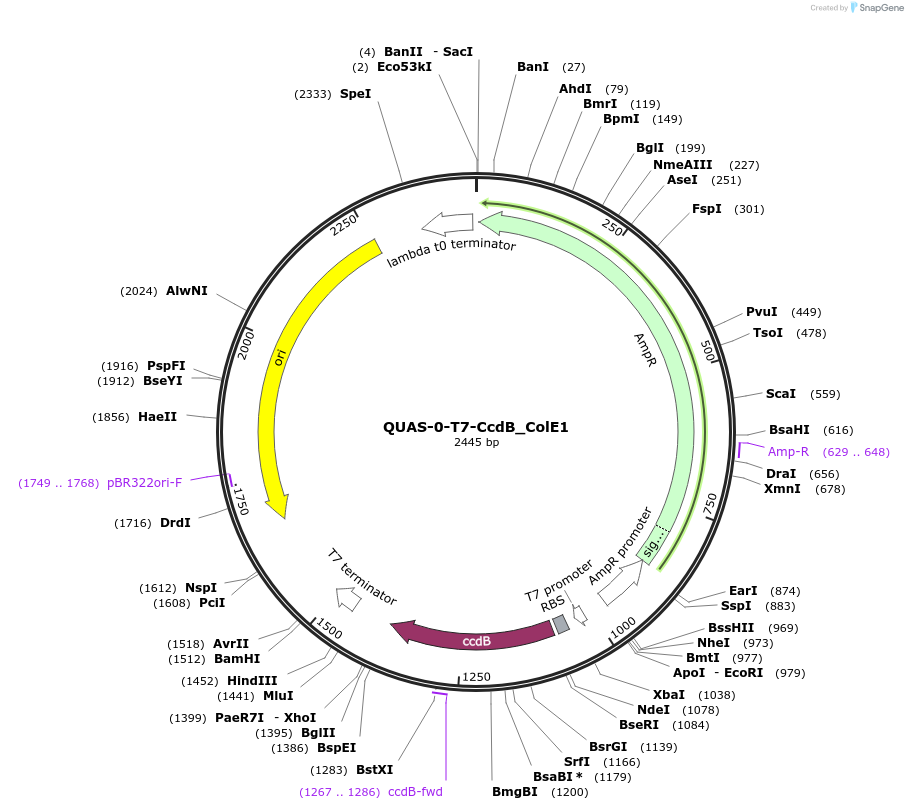
QUAS-0-T7-CcdB_ColE1
Plasmid#171652PurposeQUAS directly upstream of a T7 promoter. Expresses CcdB in BL21(DE3) E. coli. Contains ColE1 origin of replication.DepositorInsertCcdB-ssrA
TagsssrA degradation tag (DAS+4)ExpressionBacterialPromoterT7Available SinceOct. 14, 2022AvailabilityAcademic Institutions and Nonprofits only -
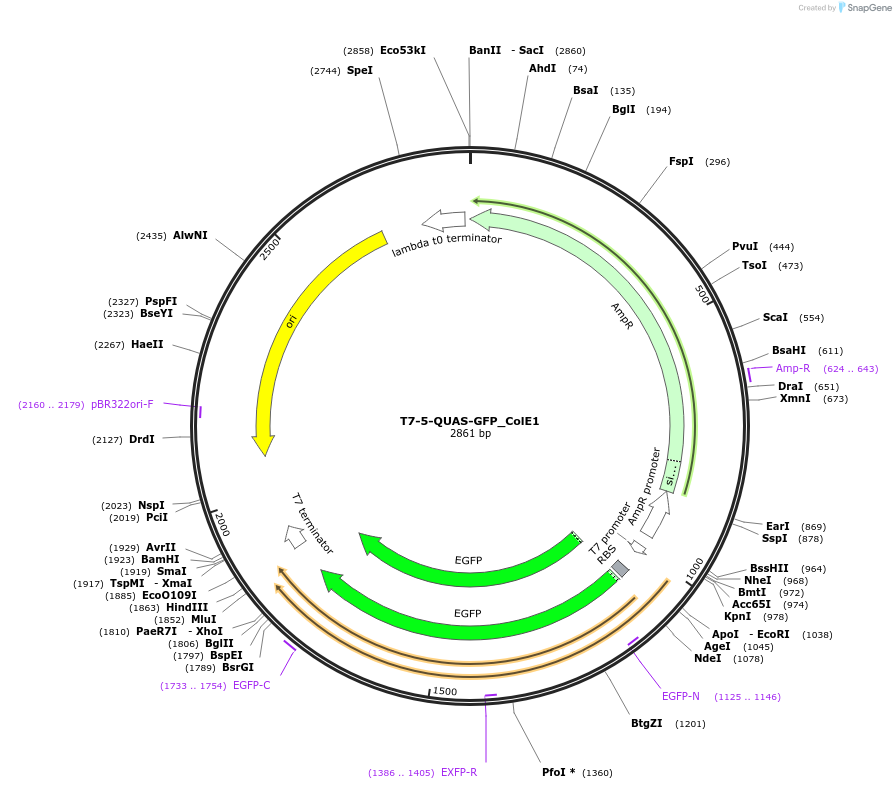
T7-5-QUAS-GFP_ColE1
Plasmid#171656PurposeQUAS five base pairs downstream of a T7 promoter. Expresses GFP in BL21(DE3) E. coli. Contains ColE1 origin of replication.DepositorInsertT7-5-QUAS
TagsssrA degradation tag (DAS+4)ExpressionBacterialPromoterT7Available SinceOct. 14, 2022AvailabilityAcademic Institutions and Nonprofits only -
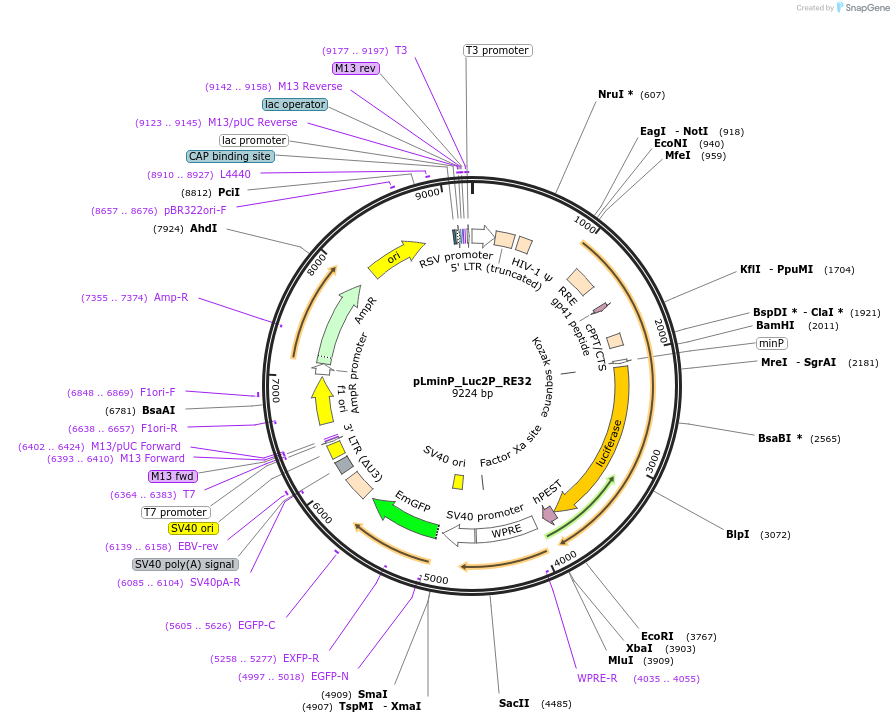
pLminP_Luc2P_RE32
Plasmid#90374PurposeSP1 - constitutive enhancer gene reporterDepositorInsertFirefly luciferase
UseLentiviralPromoterSP1Available SinceAug. 24, 2022AvailabilityAcademic Institutions and Nonprofits only -
pLminP_Luc2P_RE11
Plasmid#90345PurposeSRF - Serum response factor gene reporterDepositorInsertFirefly luciferase
UseLentiviralPromoterSRFAvailable SinceAug. 22, 2022AvailabilityAcademic Institutions and Nonprofits only -
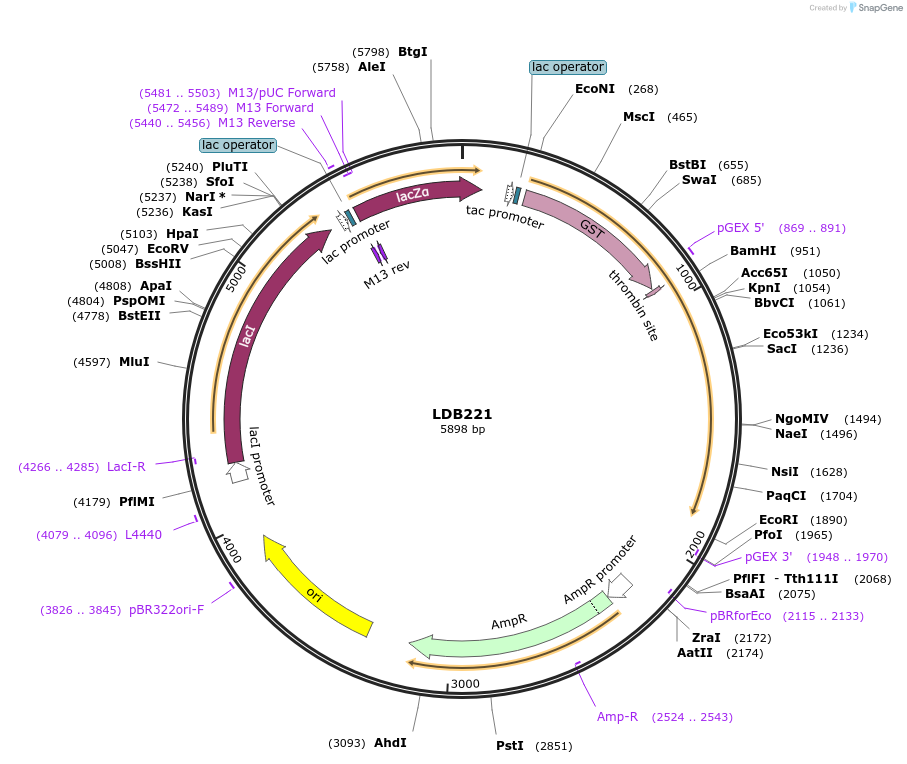
LDB221
Plasmid#185427PurposeBacterial expression of Nup1 C-terminal residues 777-1076 as GST fusionDepositorInsertNUP1
TagsGSTExpressionBacterialMutationNup1 C-terminal residues 777-1076Available SinceJuly 25, 2022AvailabilityAcademic Institutions and Nonprofits only -
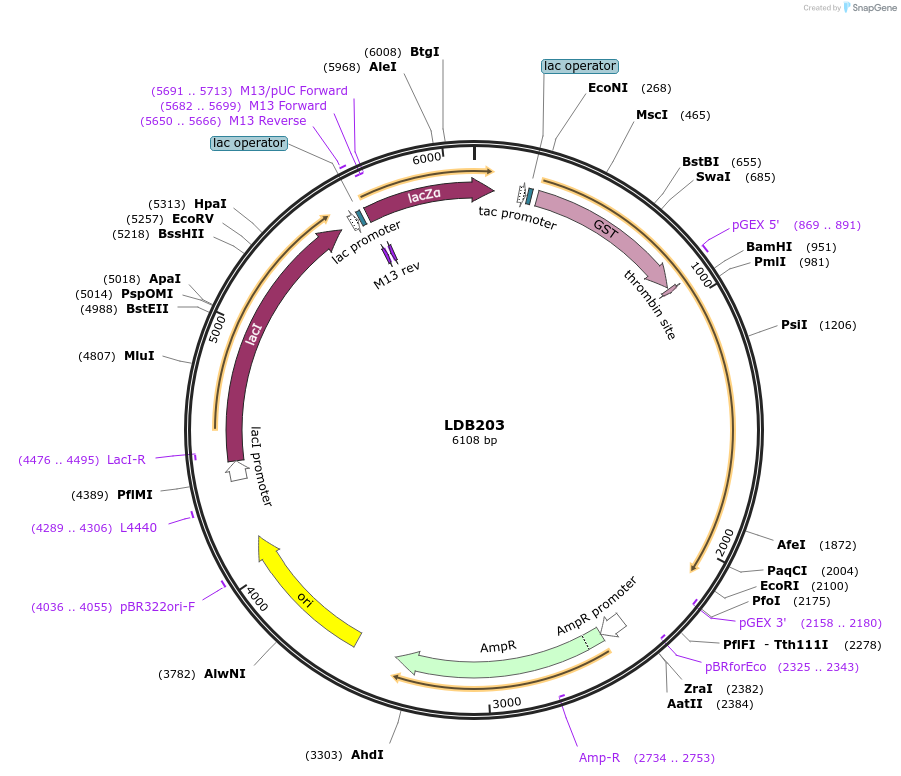
LDB203
Plasmid#185424PurposeBacterial expression of N-terminus of NUP1 nucleoporin as GST-Nup1-N fusionDepositorInsertNUP1
TagsGSTExpressionBacterialMutationNup1 N-terminal domain onlyAvailable SinceJuly 25, 2022AvailabilityAcademic Institutions and Nonprofits only -
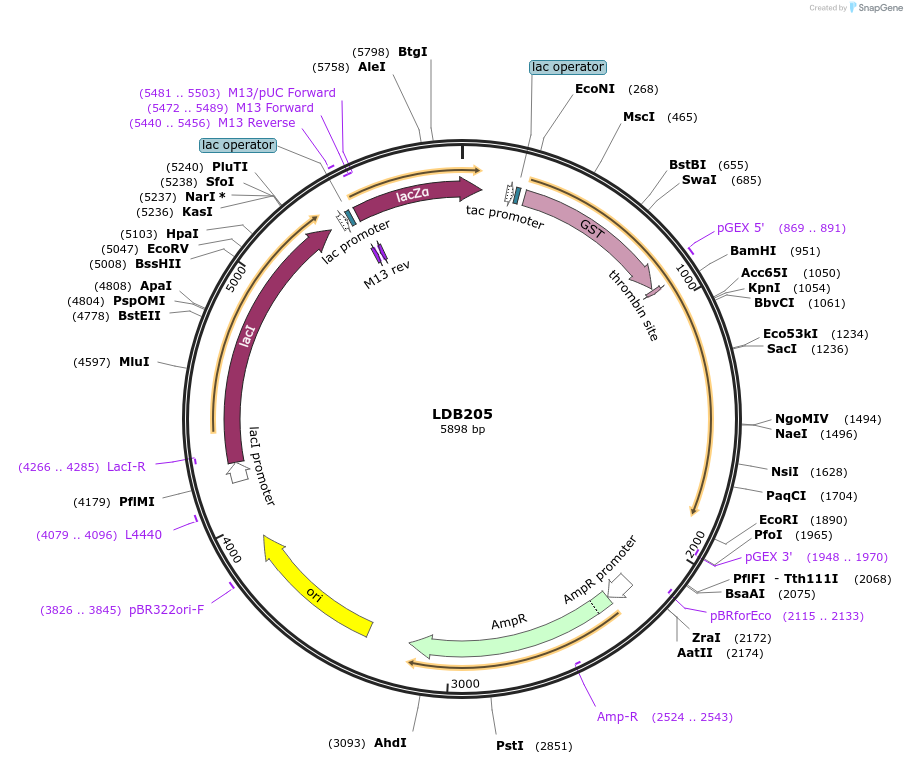
LDB205
Plasmid#185425PurposeBacterial expression of C-terminus of NUP1 nucleoporin as GST-Nup1-C fusionDepositorInsertNUP1
TagsGSTExpressionBacterialMutationNup1 C-terminal domain onlyAvailable SinceJuly 22, 2022AvailabilityAcademic Institutions and Nonprofits only -
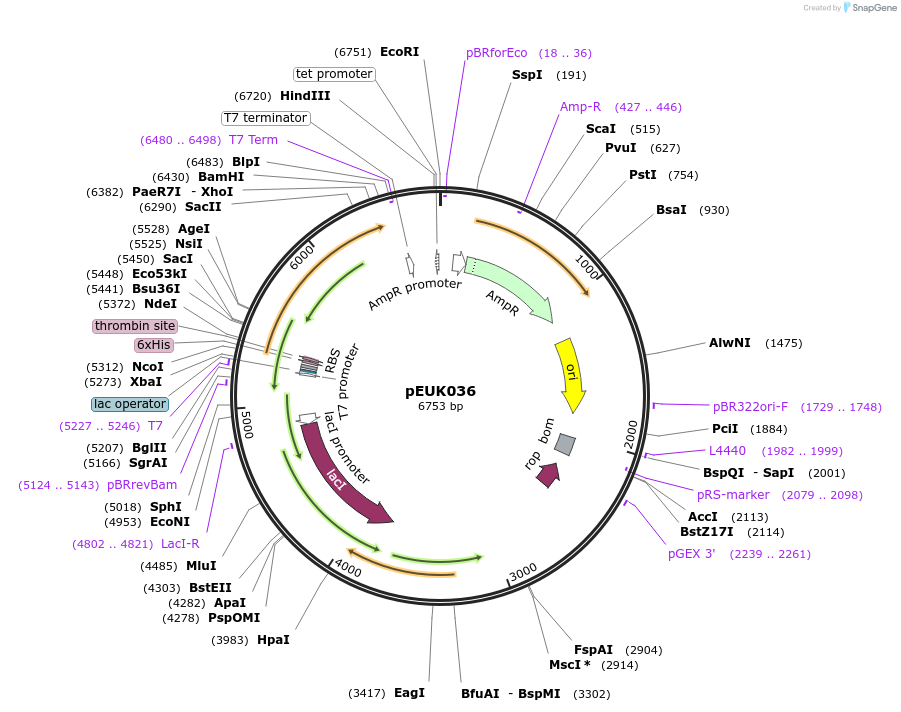
pEUK036
Plasmid#180828PurposeT7-inducible expression construct to produce CnGdmA (CNE_RS35785) in E. coliDepositorArticleInsertCnGdmA
Tags6X HisExpressionBacterialPromoterT7Available SinceJune 28, 2022AvailabilityAcademic Institutions and Nonprofits only -
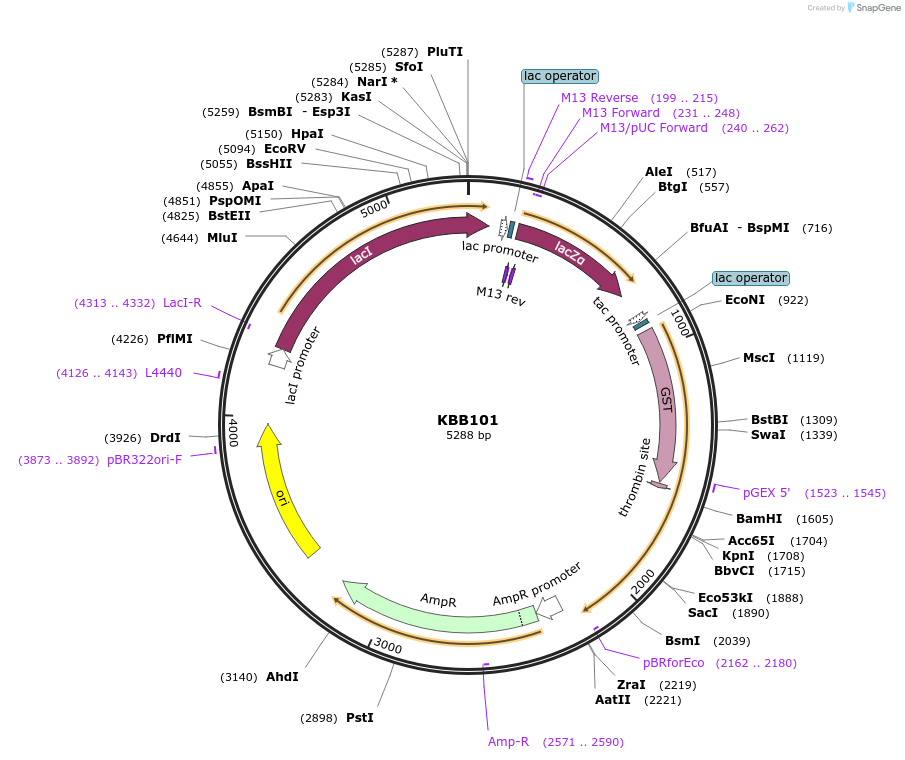
KBB101
Plasmid#185064PurposeBacterial expression of C-terminus of NUP1 nucleoporin as GST-Nup1-C fusion truncated after the last FXFG repeatDepositorInsertNUP1
TagsGSTExpressionBacterialMutationNup1 C-terminal region, truncatedAvailable SinceJune 13, 2022AvailabilityAcademic Institutions and Nonprofits only -
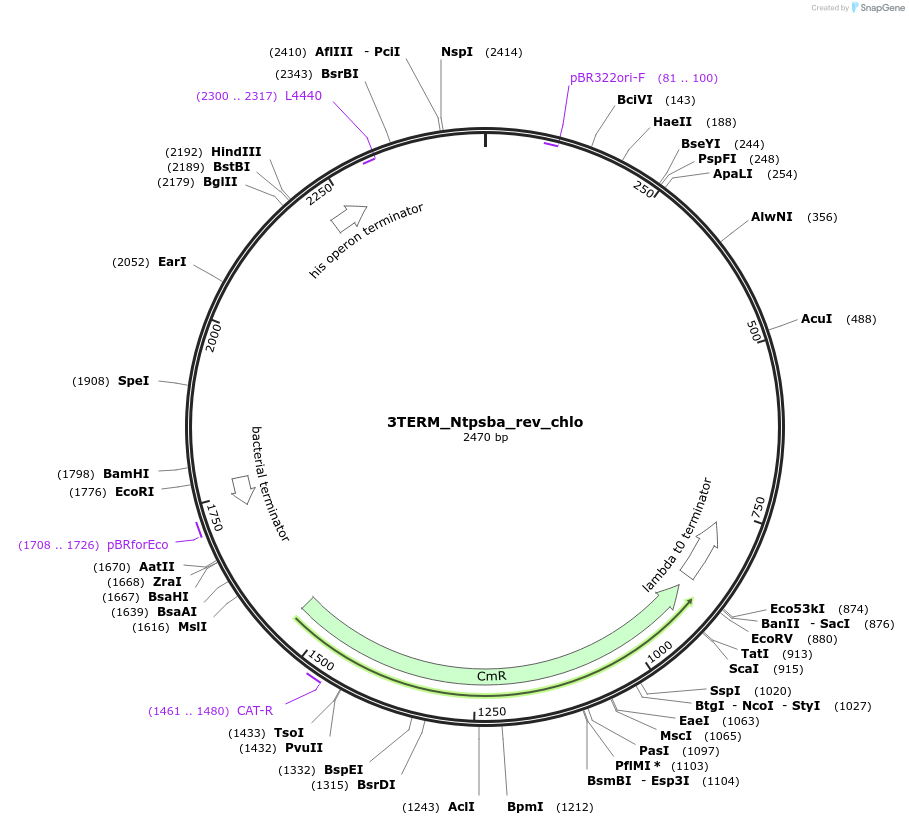
3TERM_Ntpsba_rev_chlo
Plasmid#163932PurposeL0 part - terminatorDepositorInsertNicotiana tabacum psba terminator
ExpressionBacterialAvailable SinceNov. 30, 2021AvailabilityIndustry, Academic Institutions, and Nonprofits -
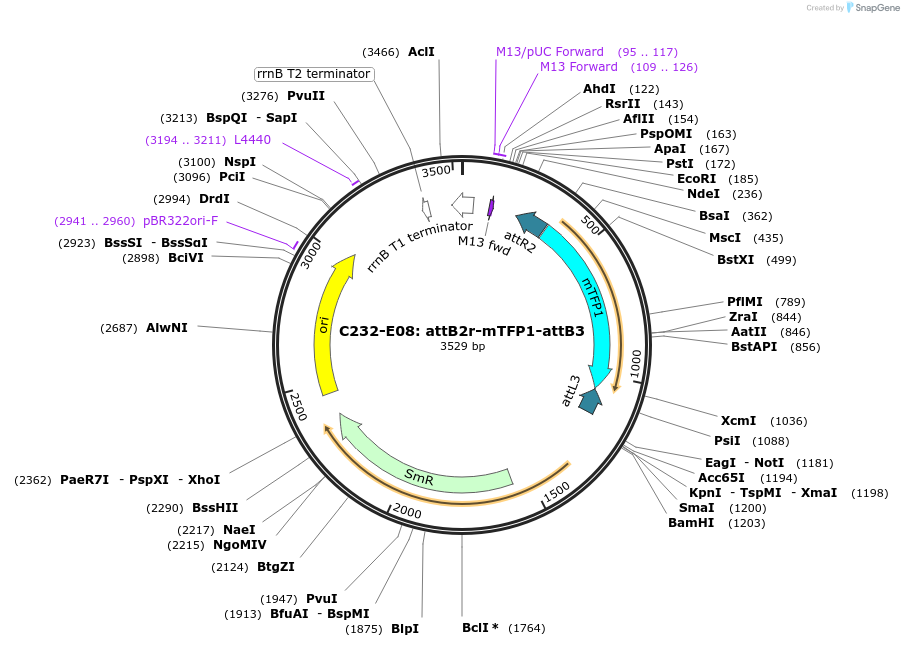
C232-E08: attB2r-mTFP1-attB3
Plasmid#162904PurposeGateway attB2r/attB3 entry clone for construction of C-terminal mTFP1 fusion proteinsDepositorInsertmonomeric Teal fluorescent protein
UseSynthetic BiologyAvailable SinceNov. 11, 2021AvailabilityAcademic Institutions and Nonprofits only -
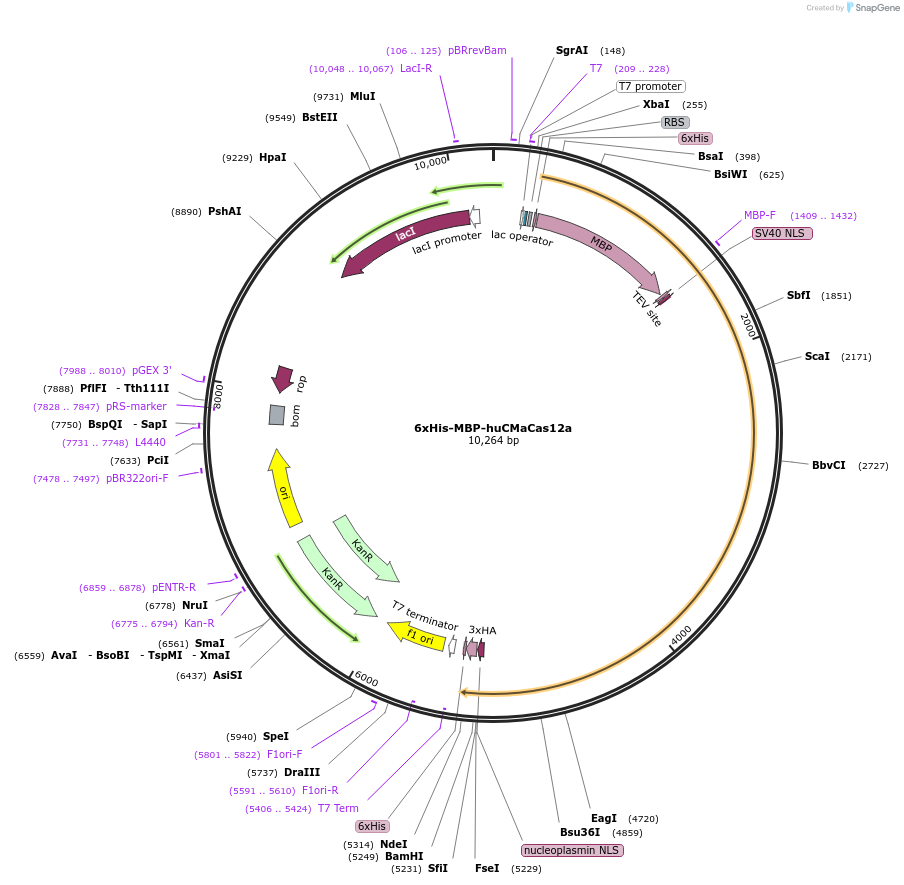
6xHis-MBP-huCMaCas12a
Plasmid#174679PurposeExpresses 6xHis-MBP tagged CMaCas12aDepositorArticleInserthuCMaCas12a
UseCRISPR and Synthetic BiologyTags6xHis, MBP, TEV and NLS, 6xHis, 3xHAExpressionBacterialPromoterT7Available SinceOct. 25, 2021AvailabilityAcademic Institutions and Nonprofits only -
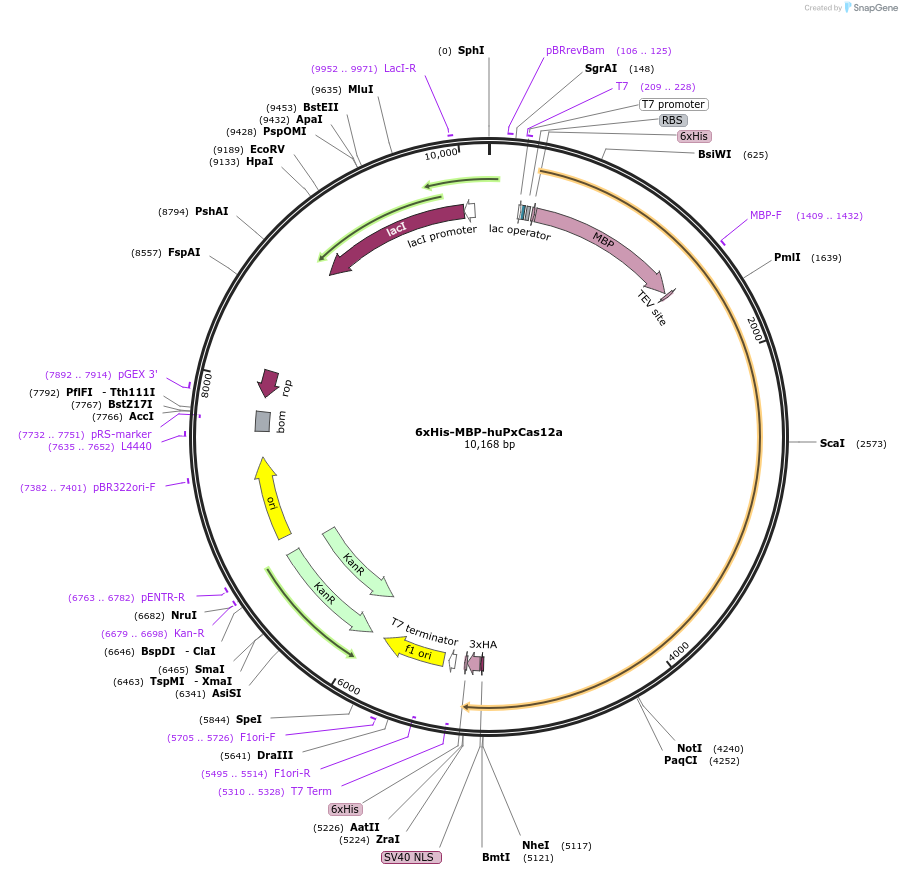
6xHis-MBP-huPxCas12a
Plasmid#174675PurposeExpresses 6xHis-MBP tagged PxCas12aDepositorArticleInserthuPxCas12a
UseCRISPR and Synthetic BiologyTags6xHis, MBP, TEV and NLS, 6xHis, 3xHAExpressionBacterialPromoterT7Available SinceOct. 22, 2021AvailabilityAcademic Institutions and Nonprofits only -
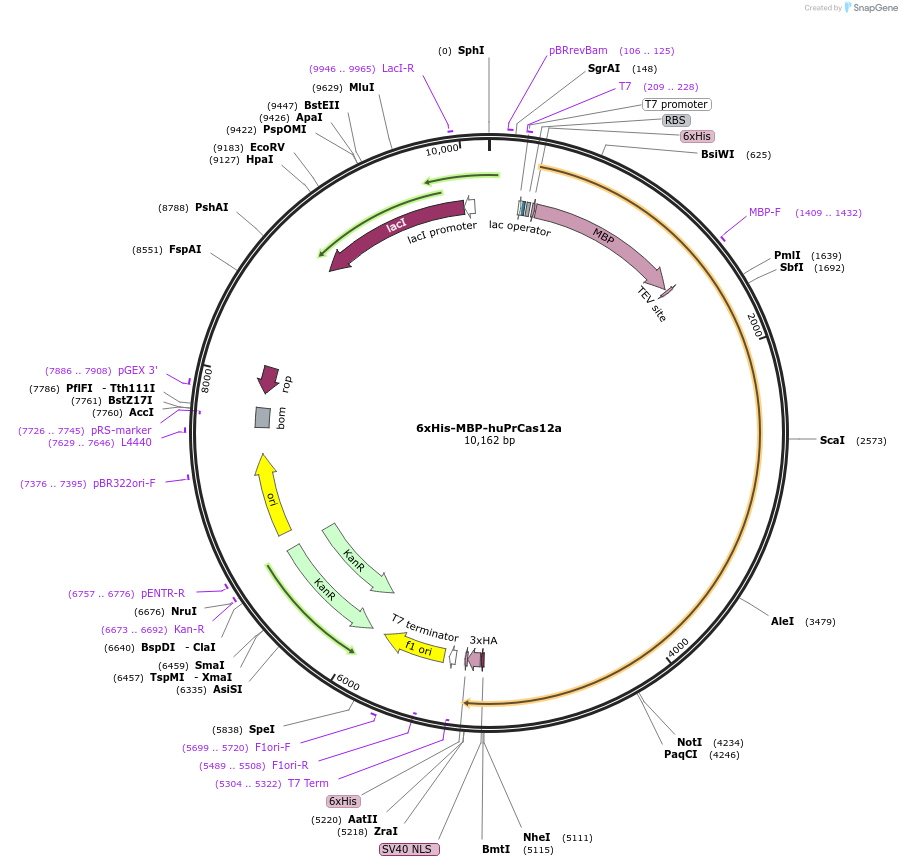
6xHis-MBP-huPrCas12a
Plasmid#174674PurposeExpresses 6xHis-MBP tagged PrCas12aDepositorArticleInserthuPrCas12a
UseCRISPR and Synthetic BiologyTags6xHis, MBP, TEV and NLS, 6xHis, 3xHAExpressionBacterialPromoterT7Available SinceOct. 22, 2021AvailabilityAcademic Institutions and Nonprofits only -
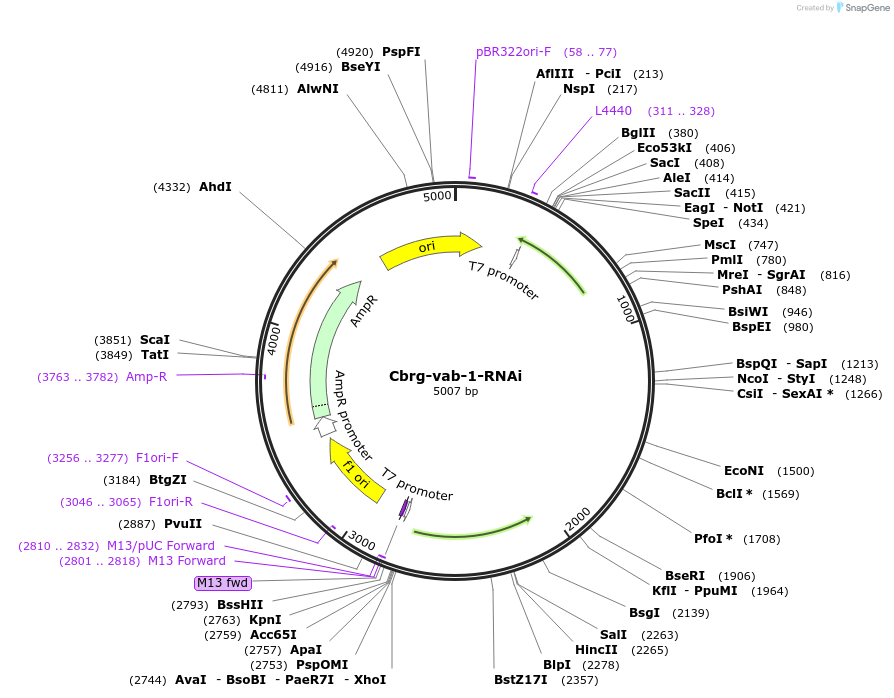
Cbrg-vab-1-RNAi
Plasmid#175249PurposeRNAi feeding bacteria/plasmid to knock-down vab-1 in C. briggsaeDepositorInsertvab-1
UseRNAiPromoterT7Available SinceSept. 20, 2021AvailabilityAcademic Institutions and Nonprofits only