We narrowed to 171,784 results for: HTT
-
Plasmid#1692DepositorAvailable SinceApril 13, 2006AvailabilityAcademic Institutions and Nonprofits only
-
mRuby-Endo-14
Plasmid#55859PurposeLocalization: Endosomes, Excitation: 558, Emission: 605DepositorAvailable SinceMarch 12, 2015AvailabilityIndustry, Academic Institutions, and Nonprofits -
-
tdEos-Occludin-N-10
Plasmid#57651PurposeLocalization: Tight Junctions, Excitation: 505 / 569, Emission: 516 / 581DepositorAvailable SinceJan. 13, 2015AvailabilityIndustry, Academic Institutions, and Nonprofits -
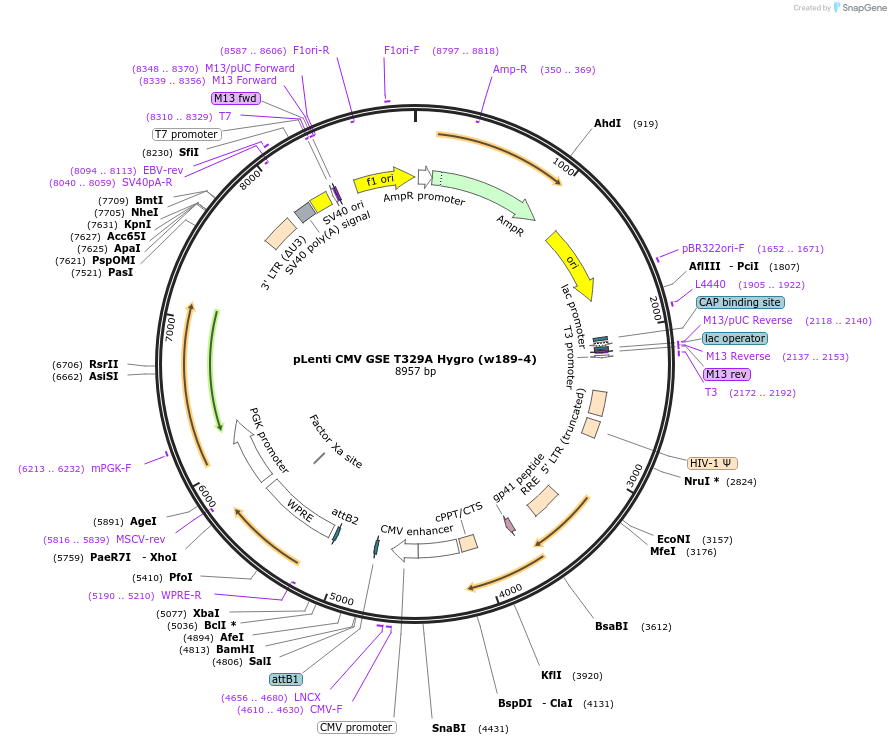
pLenti CMV GSE T329A Hygro (w189-4)
Plasmid#22255DepositorInsertGenetic suppressor element #22, T329A
UseLentiviralExpressionMammalianMutationThreonine 329 is mutated to Alanine (p53 nomencat…Available SinceDec. 15, 2009AvailabilityAcademic Institutions and Nonprofits only -
pLIC
Plasmid#27989DepositorTypeEmpty backboneExpressionBacterialAvailable SinceMarch 22, 2011AvailabilityAcademic Institutions and Nonprofits only -
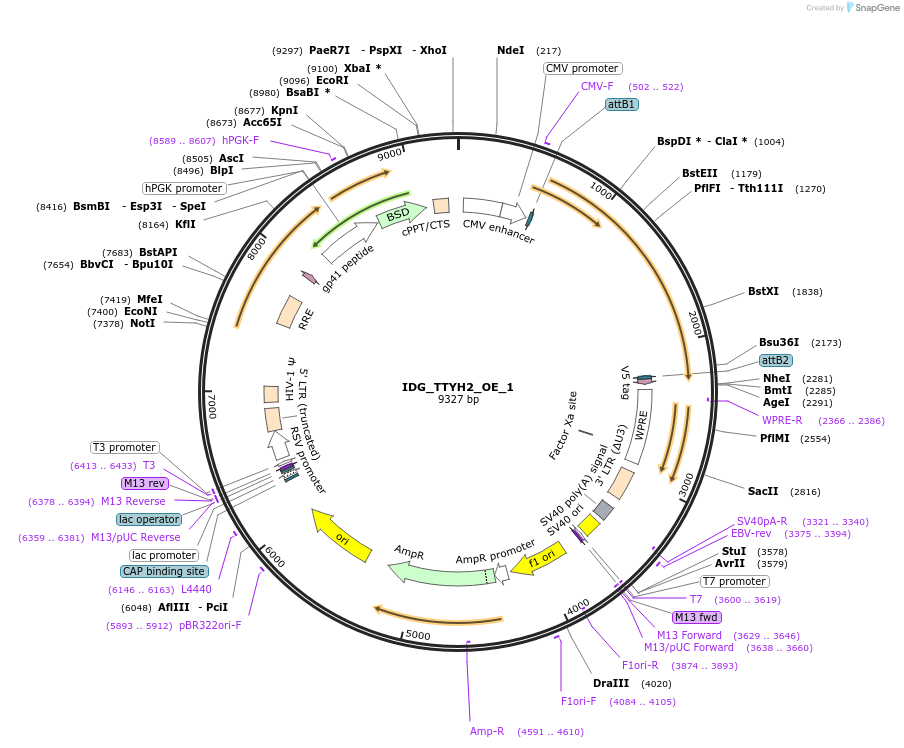
IDG_TTYH2_OE_1
Plasmid#161676PurposeOverexpresses TTYH2DepositorAvailable SinceApril 20, 2022AvailabilityAcademic Institutions and Nonprofits only -
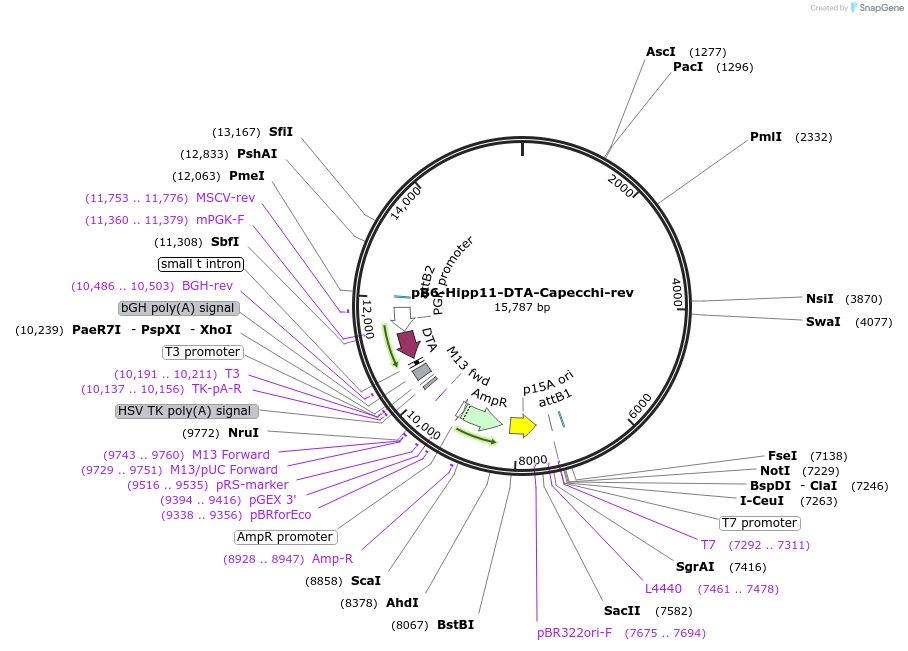
pB6-Hipp11-DTA-Capecchi-rev
Plasmid#127054PurposeB6J Hipp11 targeting vector homology arms, Capecchi configuration, DTA, I-CeuI, reverse cloning sites, low copy numberDepositorTypeEmpty backboneUseMouse TargetingAvailable SinceJune 26, 2019AvailabilityAcademic Institutions and Nonprofits only -
mKate-H4-23
Plasmid#56061PurposeLocalization: Nucleus/Histones, Excitation: 488, Emission: 635DepositorAvailable SinceApril 28, 2015AvailabilityIndustry, Academic Institutions, and Nonprofits -
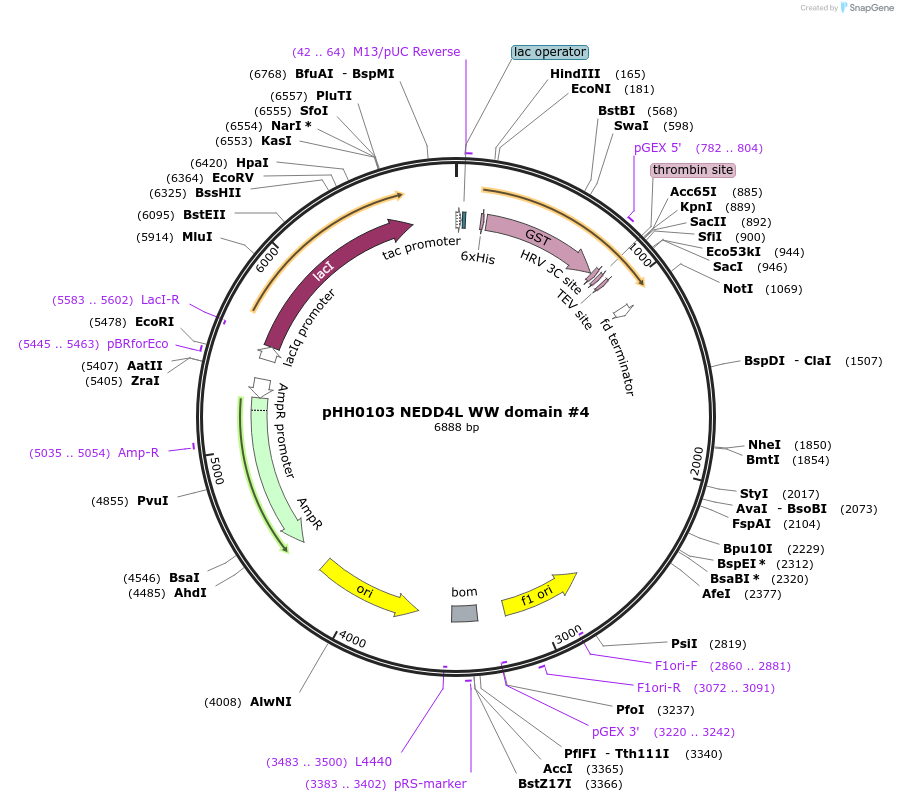
pHH0103 NEDD4L WW domain #4
Plasmid#104207PurposeBacterial expression of WW domain #4 from NEDD4LDepositorInsertNEDD4L WW domain #4 (NEDD4L Human)
TagsGSTExpressionBacterialMutationcodon optimized for expression in bacteriaPromotertacAvailable SinceAug. 28, 2018AvailabilityAcademic Institutions and Nonprofits only -
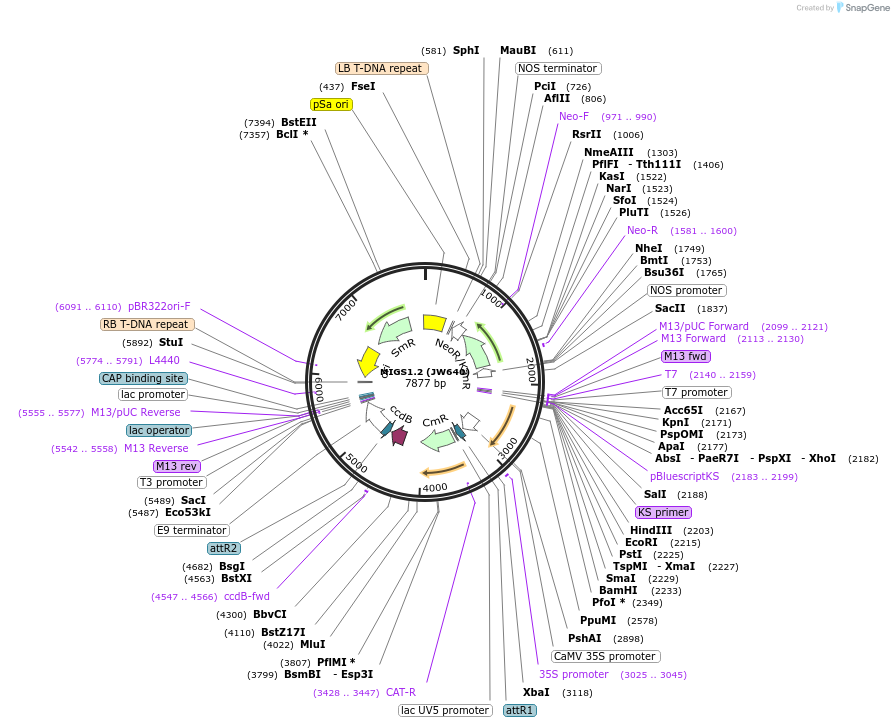
MIGS1.2 (JW640)
Plasmid#35246DepositorTypeEmpty backboneExpressionPlantPromoterCaMV 35SAvailable SinceMarch 23, 2012AvailabilityAcademic Institutions and Nonprofits only -
mApple-PDHA1-N-10
Plasmid#54938PurposeLocalization: Mitochondria, Excitation: 568, Emission: 592DepositorAvailable SinceAug. 29, 2014AvailabilityAcademic Institutions and Nonprofits only -
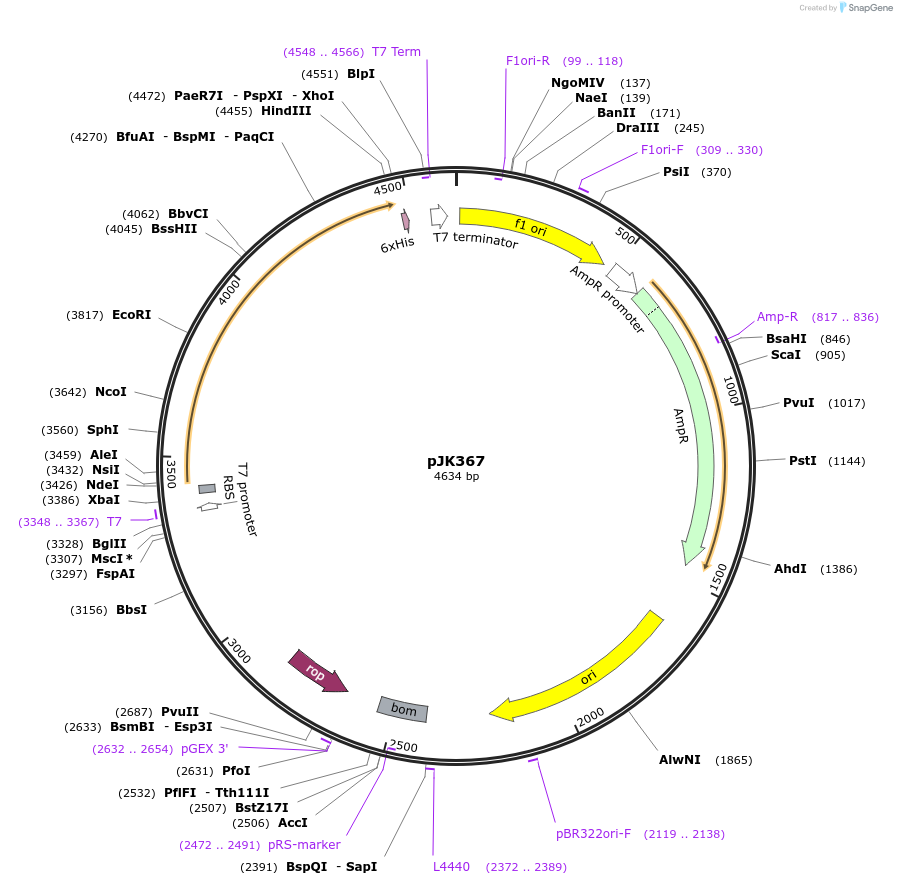
pJK367
Plasmid#72406PurposeProduces Acetobacter aceti 1023 Est06DepositorInsertlipase
ExpressionBacterialMutationlacks Met1 and Thr2 in annotated (v1) sequencePromoterT7Available SinceFeb. 25, 2016AvailabilityAcademic Institutions and Nonprofits only -
-
MPSTA
Plasmid#42482PurposeBacterial expression for structure determination; may not be full ORFDepositorInsertMPSTA
TagsHis6-TEVExpressionBacterialMutationSee CommentsAvailable SinceMarch 18, 2013AvailabilityAcademic Institutions and Nonprofits only -
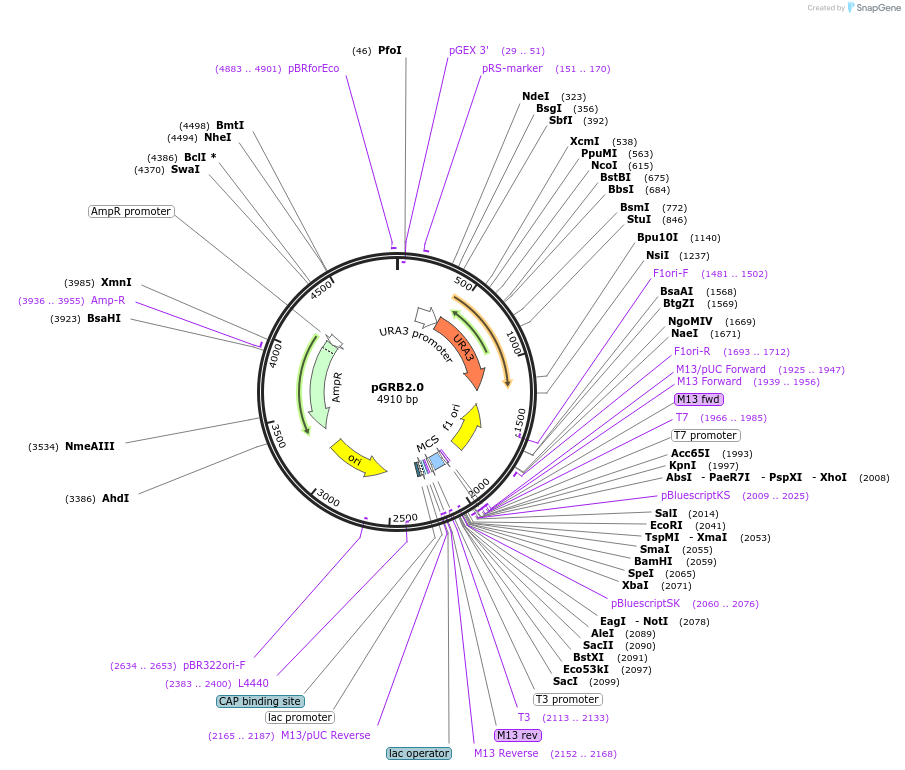
pGRB2.0
Plasmid#45340DepositorTypeEmpty backboneUseC. glabrataAvailable SinceJuly 8, 2013AvailabilityAcademic Institutions and Nonprofits only -
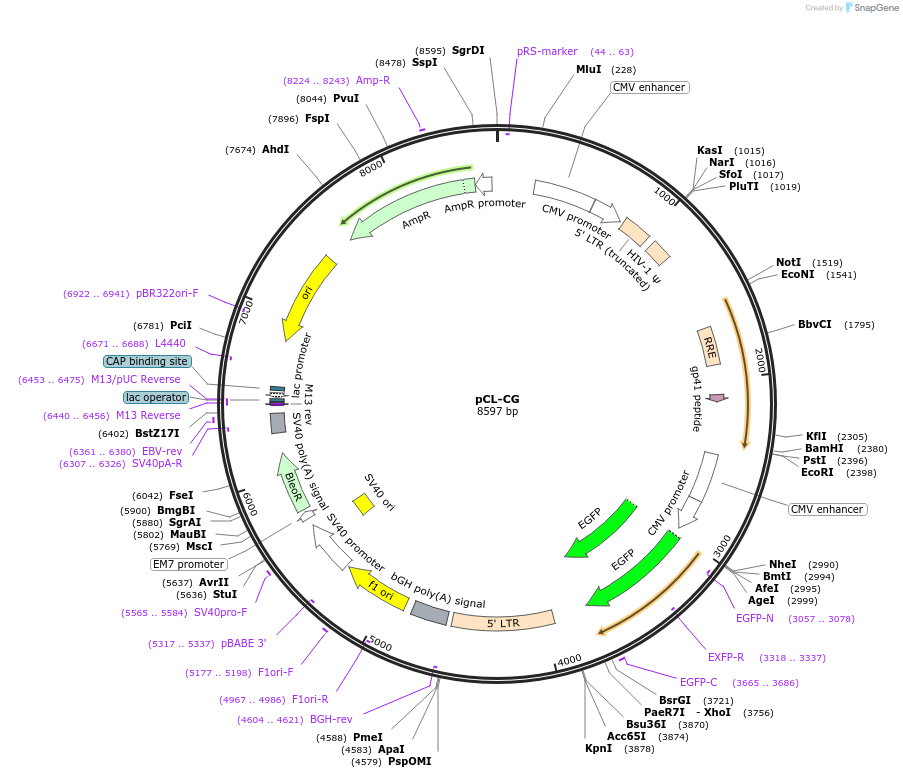
pCL-CG
Plasmid#12157DepositorInsertCMV-GFP
UseLentiviralExpressionMammalianAvailable SinceAug. 17, 2006AvailabilityAcademic Institutions and Nonprofits only -
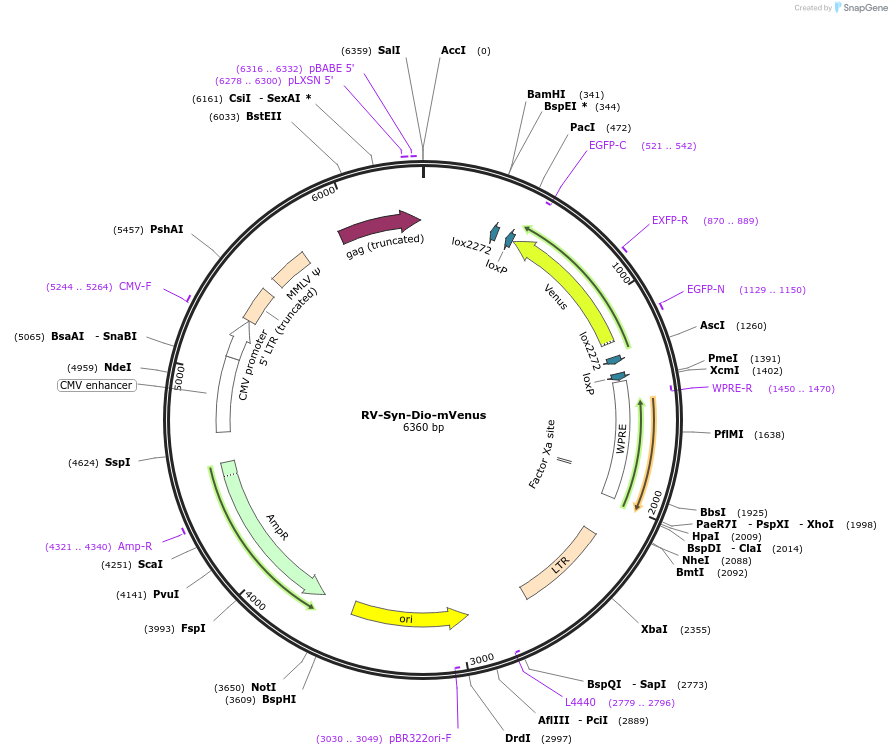
RV-Syn-Dio-mVenus
Plasmid#153204PurposeCan be used to generate retrovirus that will mark the membrane (Palmitoylated-Venus) in the presence of Cre from the rat synapsin promoterDepositorInsertmembrane-tagged-Venus
UseRetroviralPromoterrat synapsinAvailable SinceJune 4, 2020AvailabilityAcademic Institutions and Nonprofits only -
EYFP-Endosomes-13
Plasmid#56588PurposeLocalization: Endosomes, Excitation: 513, Emission: 527DepositorAvailable SinceJan. 6, 2015AvailabilityIndustry, Academic Institutions, and Nonprofits -
pSLAX13 HA (CT#233)
Plasmid#14027DepositorTypeEmpty backboneTags3x HAExpressionBacterialAvailable SinceFeb. 9, 2007AvailabilityAcademic Institutions and Nonprofits only