We narrowed to 124,568 results for: CIL
-
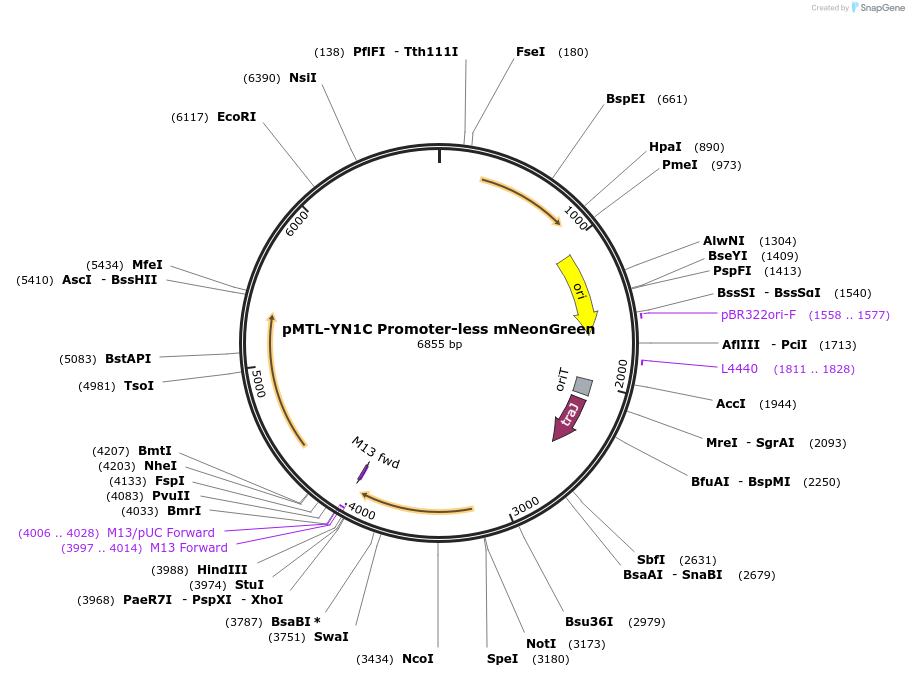
Plasmid#251223PurposepMTL-YN1C encoding a promoter-less mNeonGreen.DepositorInsertLeader peptide with mNeonGreen codon optimized for C. difficile
ExpressionBacterialPromoterPromoter-lessAvailable SinceApril 21, 2026AvailabilityAcademic Institutions and Nonprofits only -
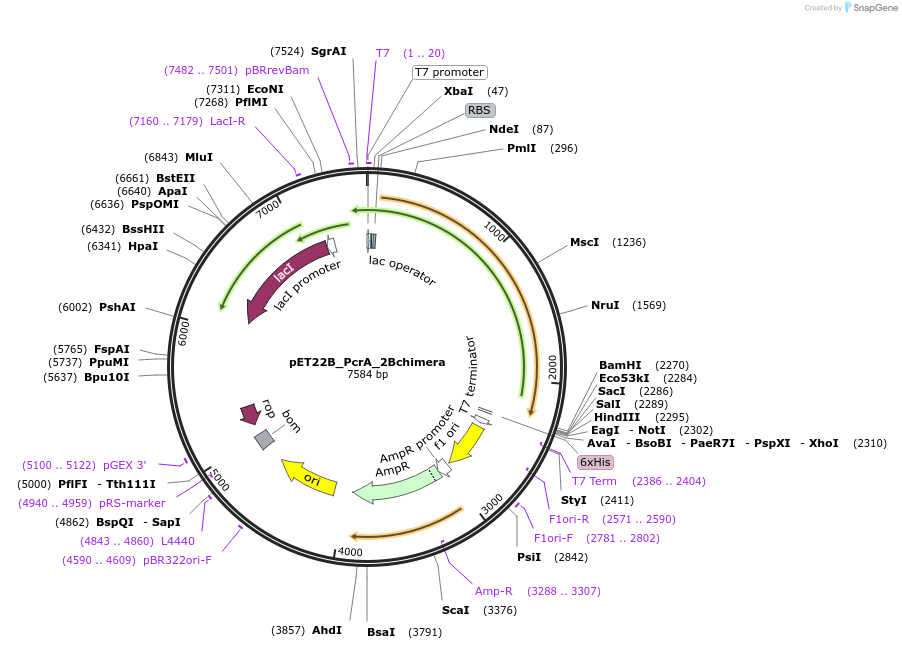
pET22B_PcrA_2Bchimera
Plasmid#102998PurposeExpress a DNA helicase, a chimeric construct of Bacilllus stearothermophilus PcrA and 2B domain of Staphylococcus aureus PcrADepositorInsertPcrA_2Bchimera
ExpressionBacterialMutationBacilllus stearothermophilus PcrA in which the 2B…PromoterT7Available SinceMay 24, 2018AvailabilityAcademic Institutions and Nonprofits only -
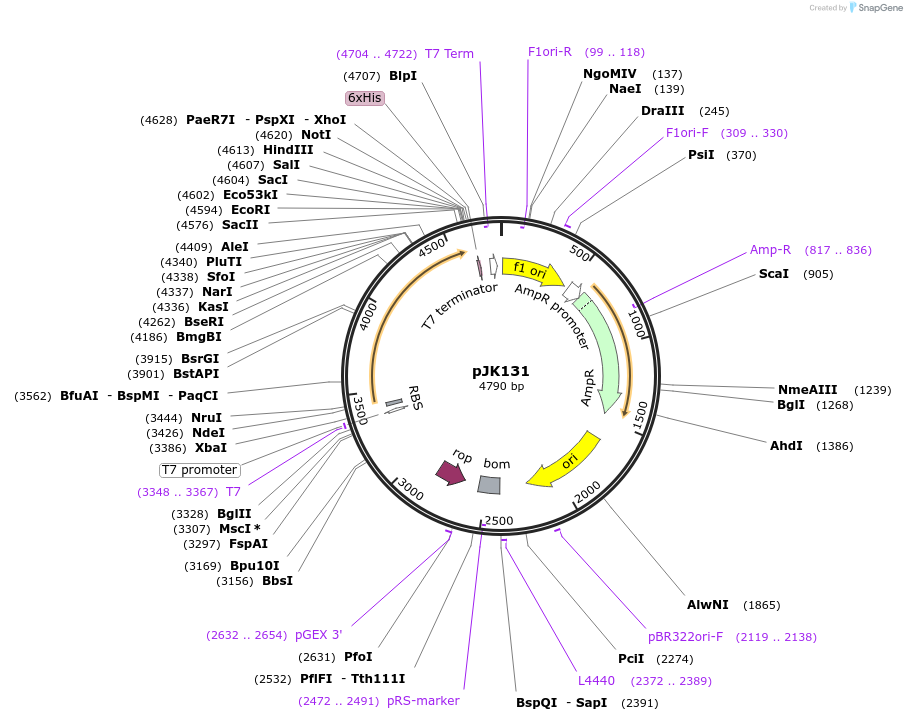
pJK131
Plasmid#71548PurposeProduces Geobacillus stearothermophilus (Bacillus stearothermophilus) alanine racemase (alr)DepositorInsertAlanine racemase
ExpressionBacterialPromoterT7Available SinceApril 1, 2016AvailabilityAcademic Institutions and Nonprofits only -
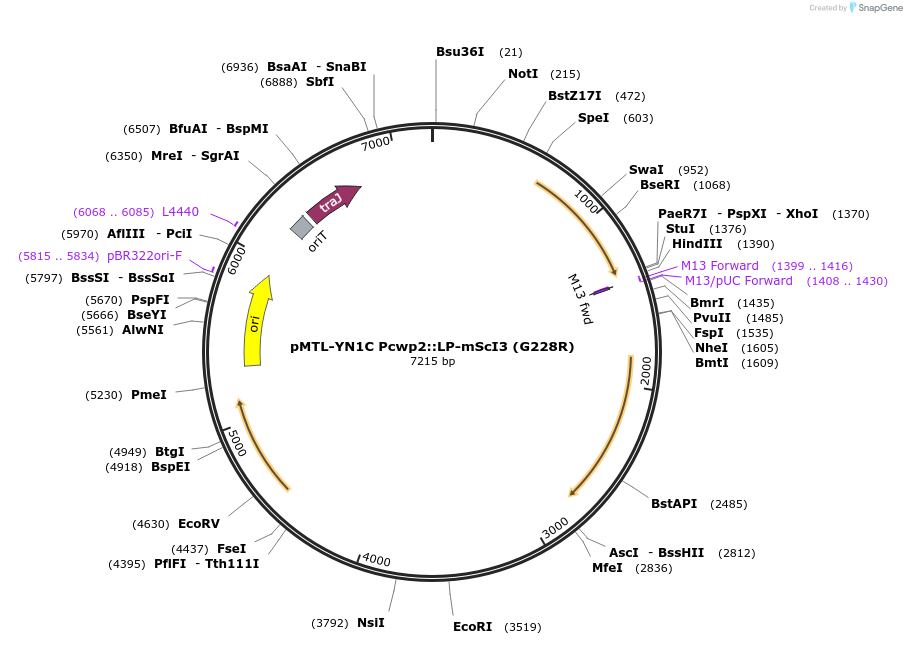
pMTL-YN1C Pcwp2::LP-mScI3 (G228R)
Plasmid#251226PurposepMTL-YN1C encoding mScarletI3 with a G228R mutation under the promoter for cwp2. This functions as a constitutive transcriptional reporter for visualizing C. difficile.DepositorInsertPcwp2::LP-mScI3 (G228R)
ExpressionBacterialMutationGlycine 228 changed to arginine.Promotercwp2Available SinceApril 21, 2026AvailabilityAcademic Institutions and Nonprofits only -
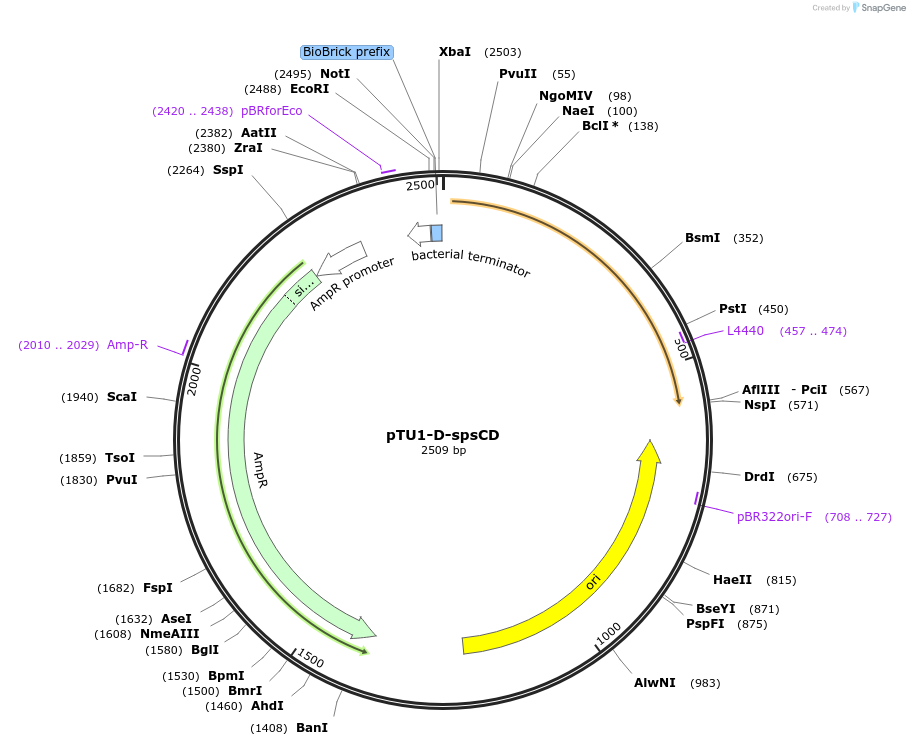
pTU1-D-spsCD
Plasmid#112788PurposeDownstream homology arm, SpsC is involved in spore coat polysaccharide synthesisDepositorArticleInsertspsCD
ExpressionBacterialAvailable SinceAug. 15, 2018AvailabilityAcademic Institutions and Nonprofits only -
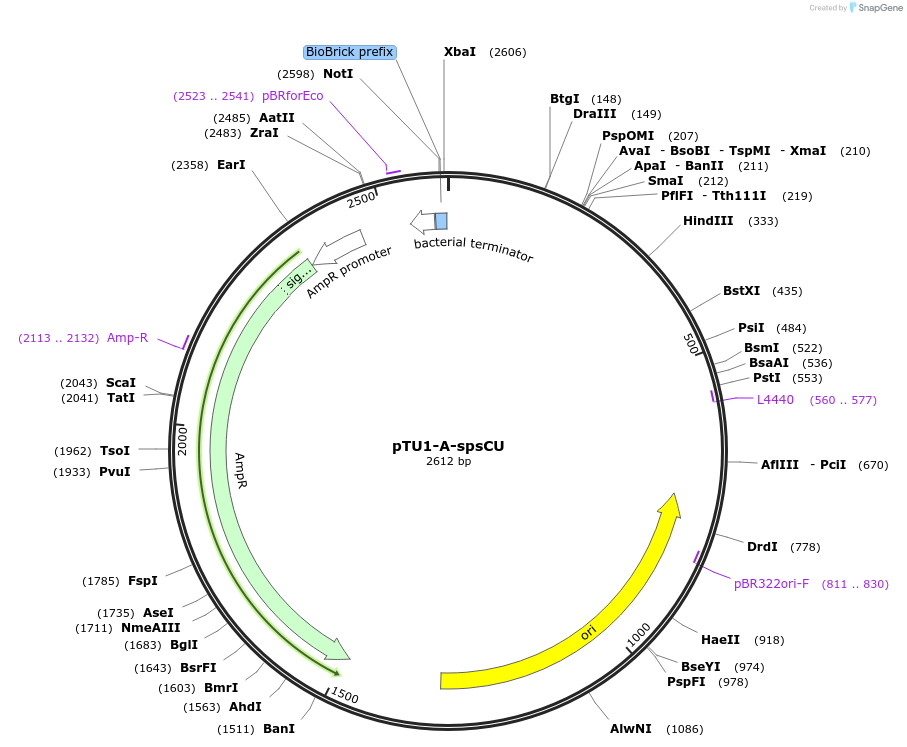
pTU1-A-spsCU
Plasmid#112787PurposeUpstream homology arm, SpsC is involved in spore coat polysaccharide synthesisDepositorArticleInsertspsCU
ExpressionBacterialAvailable SinceAug. 15, 2018AvailabilityAcademic Institutions and Nonprofits only -
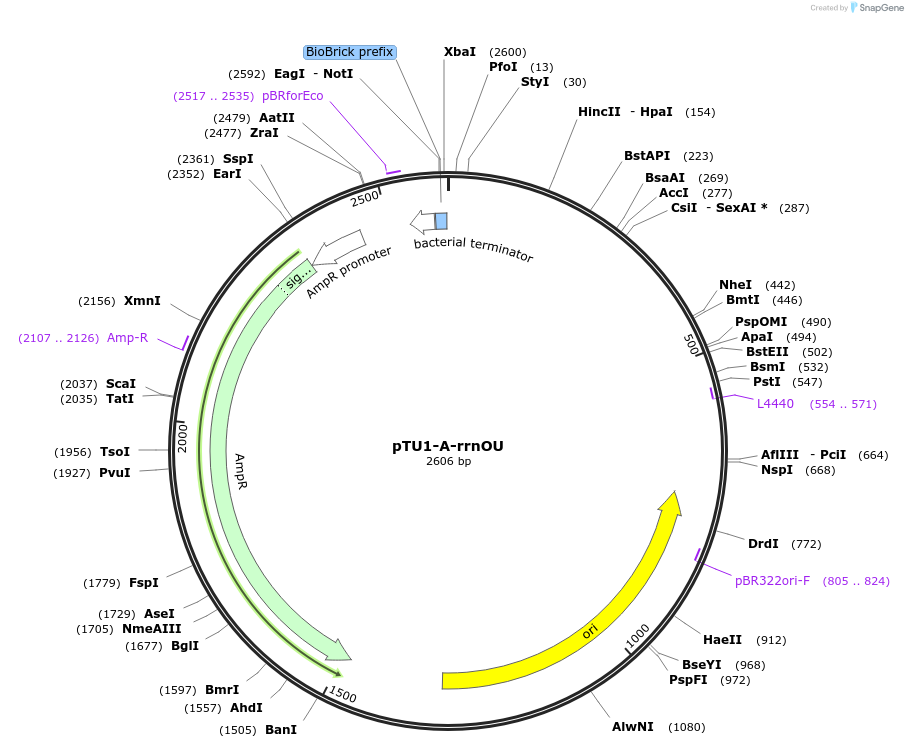
pTU1-A-rrnOU
Plasmid#112785PurposeUpstream homology arm, rrnO-23S (23S rRNA)DepositorArticleInsertrrnOU
ExpressionBacterialAvailable SinceAug. 15, 2018AvailabilityAcademic Institutions and Nonprofits only -
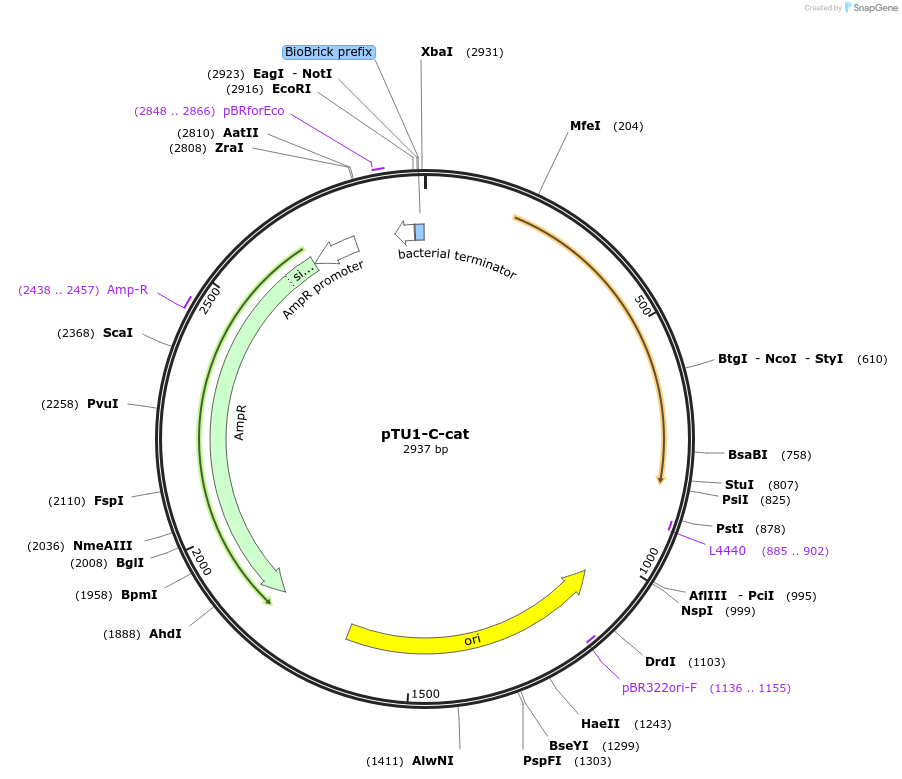
pTU1-C-cat
Plasmid#112791PurposeChloramphenicol resistance cassette (expressing chloramphenicol acetyltransferase)DepositorArticleInsertcat
ExpressionBacterialAvailable SinceAug. 14, 2018AvailabilityAcademic Institutions and Nonprofits only -
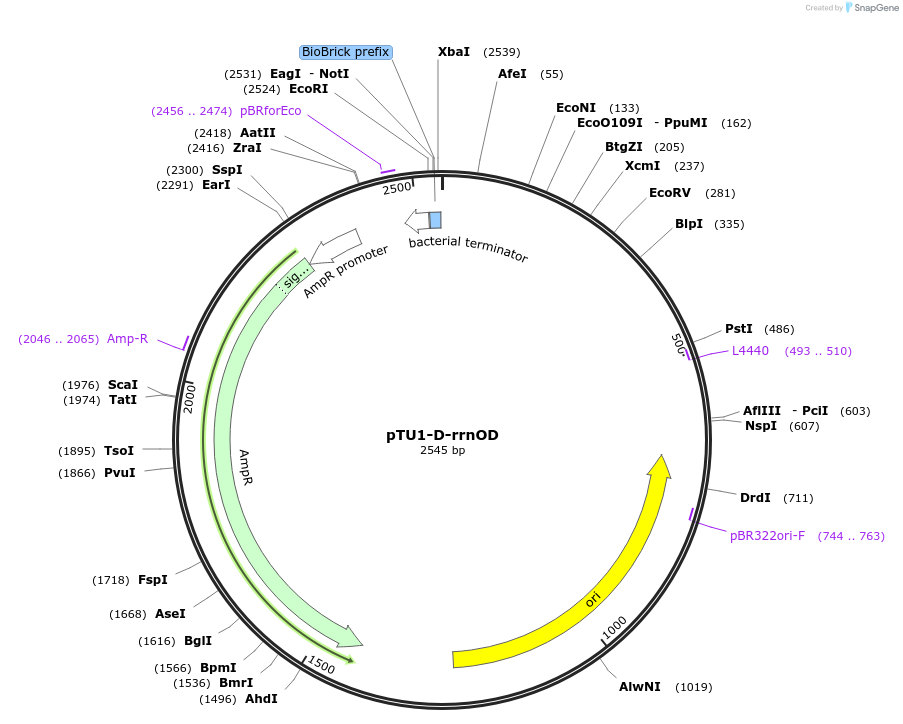
pTU1-D-rrnOD
Plasmid#112786PurposeDownstream homology arm, rrnO-23S (23S rRNA)DepositorArticleInsertrrnOD
ExpressionBacterialAvailable SinceAug. 14, 2018AvailabilityAcademic Institutions and Nonprofits only -
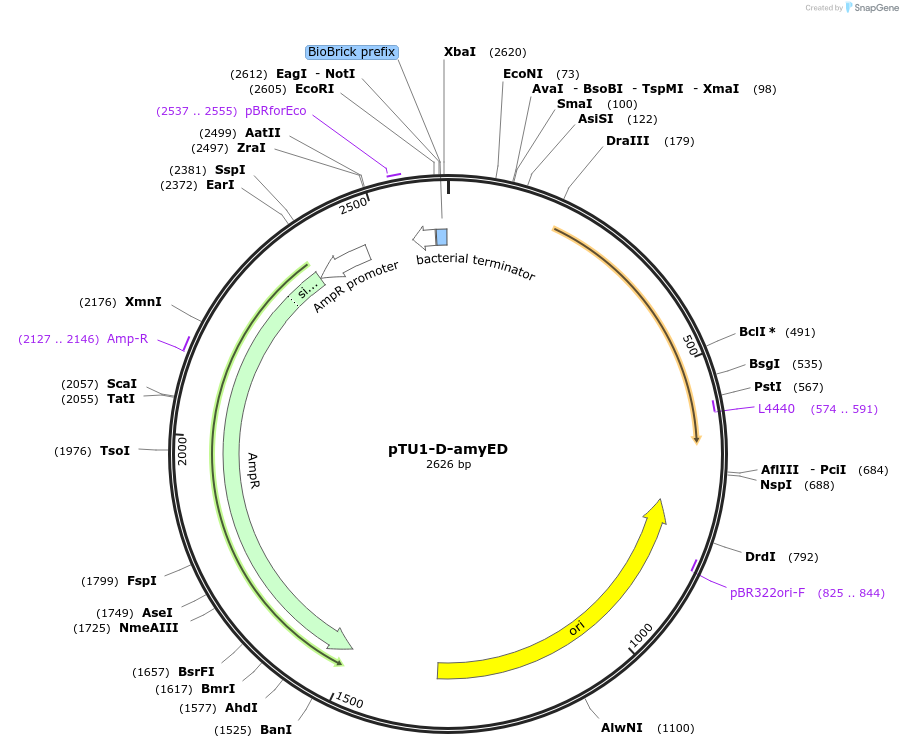
pTU1-D-amyED
Plasmid#112782PurposeDownstream homology arm, amyE encodes _-amylaseDepositorArticleInsertamyED
ExpressionBacterialAvailable SinceAug. 14, 2018AvailabilityAcademic Institutions and Nonprofits only -
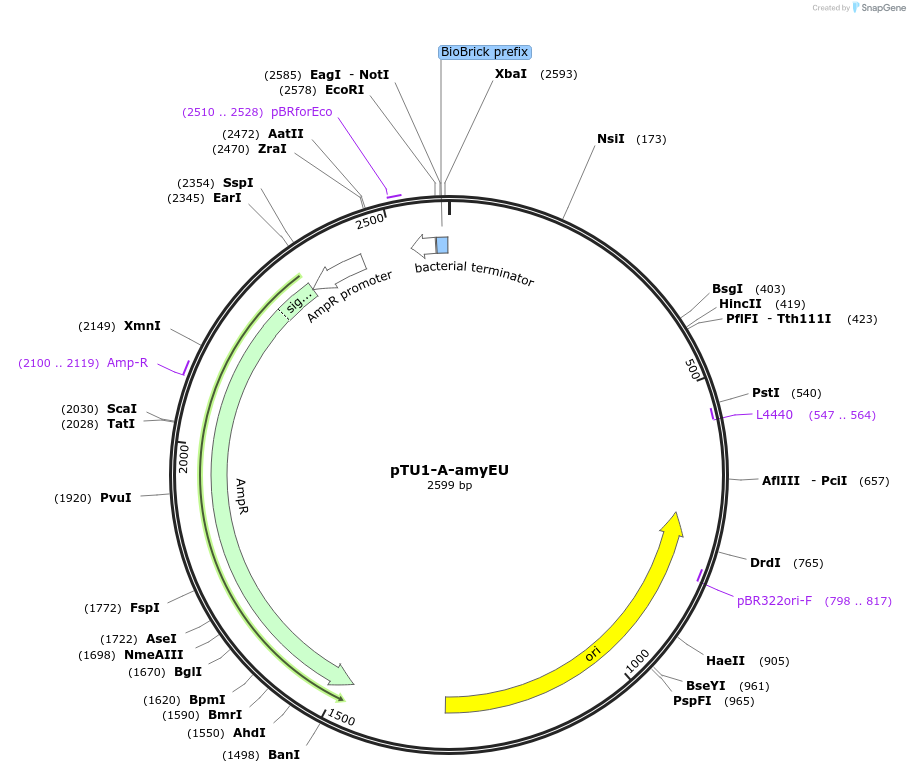
pTU1-A-amyEU
Plasmid#112781PurposeUpstream homology arm, amyE encodes _-amylaseDepositorArticleInsertamyEU
ExpressionBacterialAvailable SinceAug. 14, 2018AvailabilityAcademic Institutions and Nonprofits only -
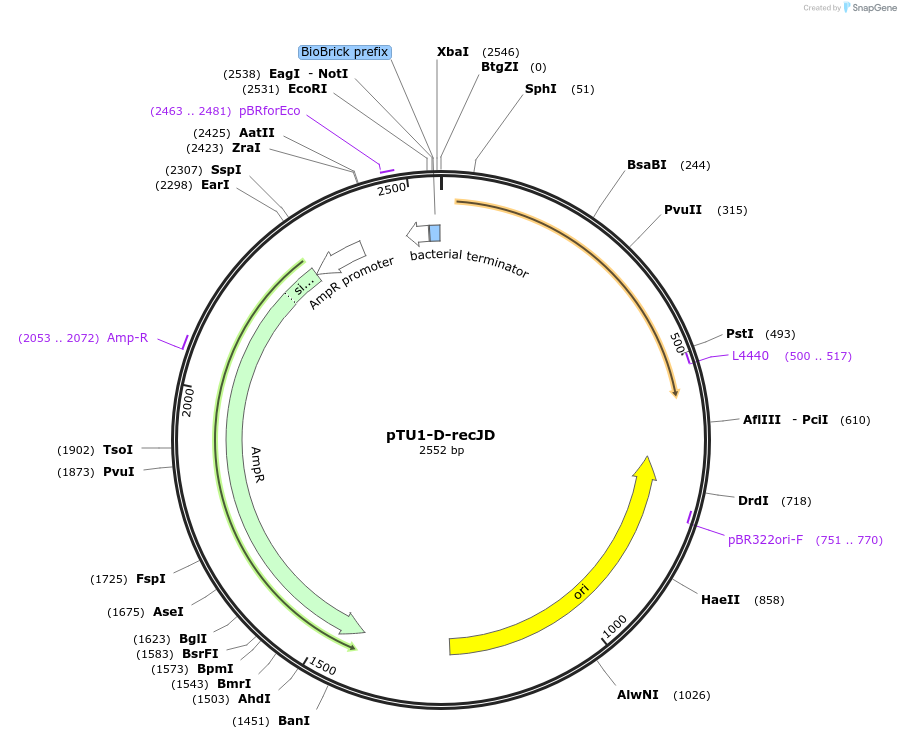
pTU1-D-recJD
Plasmid#112784PurposeDownstream homology arm, RecJ is a ssDNA specific exonuclease, involved in genome repair and maintenanceDepositorArticleInsertrecJD
ExpressionBacterialAvailable SinceAug. 14, 2018AvailabilityAcademic Institutions and Nonprofits only -
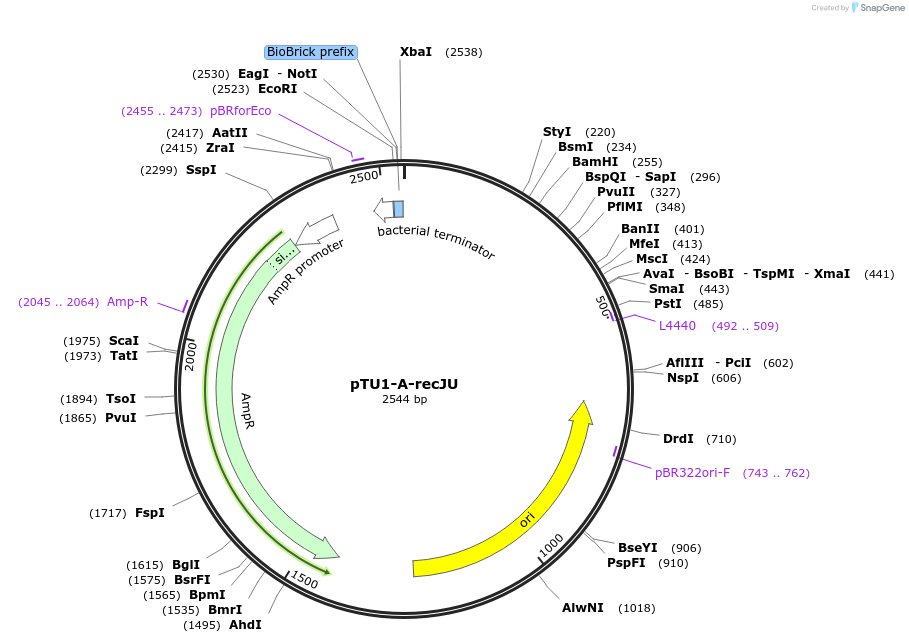
pTU1-A-recJU
Plasmid#112783PurposeUpstream homology arm, RecJ is a ssDNA specifiC exonuclease, involved in genome repair and maintenanceDepositorArticleInsertrecJU
ExpressionBacterialAvailable SinceAug. 15, 2018AvailabilityAcademic Institutions and Nonprofits only -
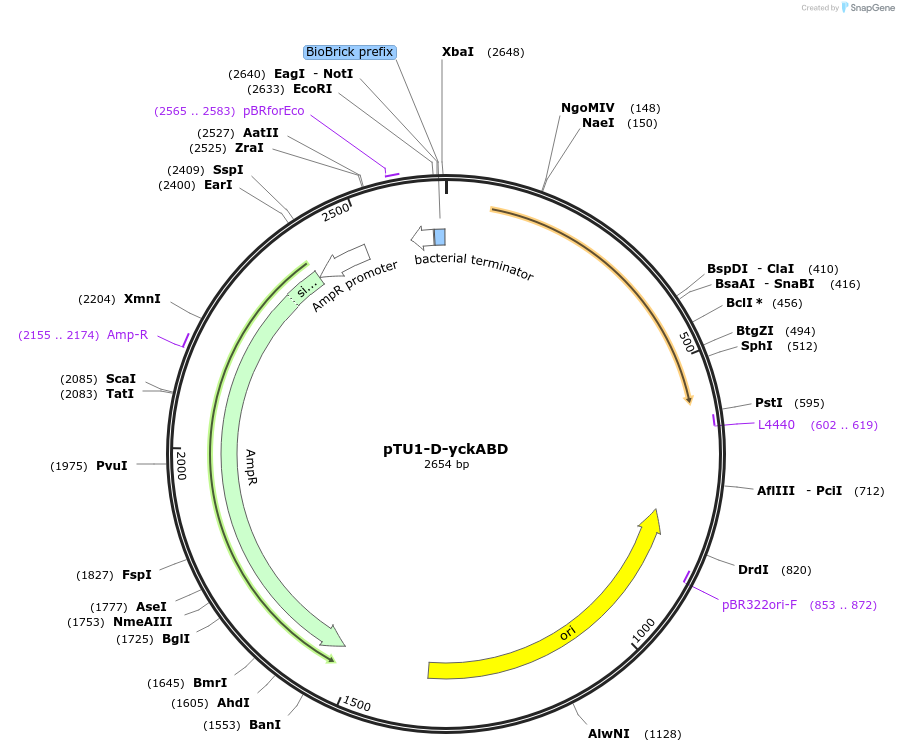
pTU1-D-yckABD
Plasmid#112790PurposeDownstream homology arm, Pair of HAs spans yckA and yckB, which share homology with ABC transporters in B. subtilisDepositorArticleInsertyckABD
ExpressionBacterialAvailable SinceAug. 15, 2018AvailabilityAcademic Institutions and Nonprofits only -
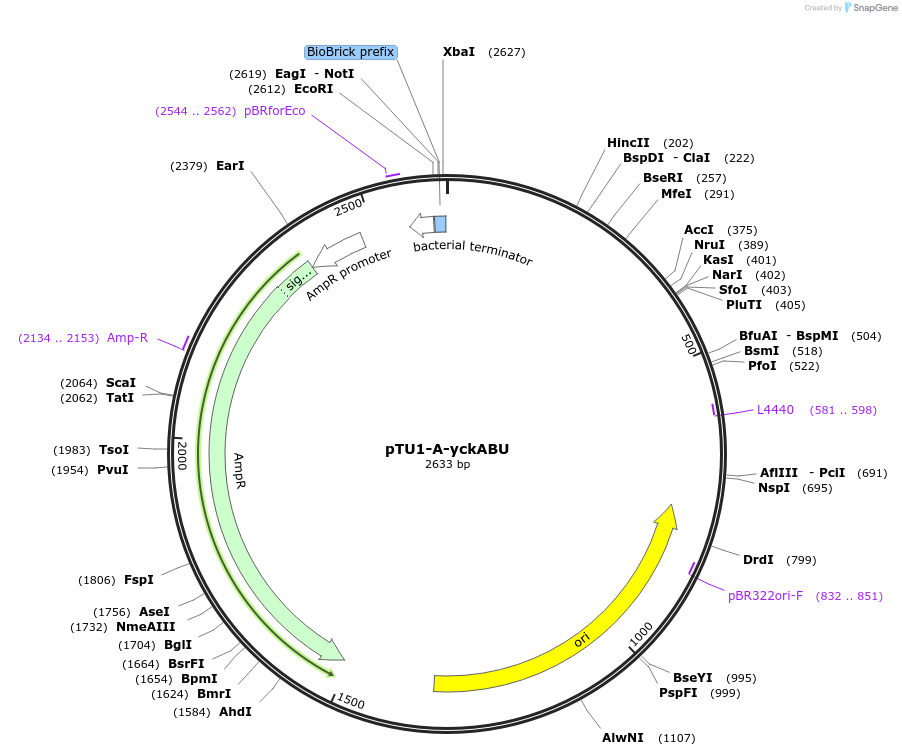
pTU1-A-yckABU
Plasmid#112789PurposeUpstream homology arm, Pair of HAs spans yckA and yckB, which share homology with ABC transporters in B. subtilisDepositorArticleInsertyckABU
ExpressionBacterialAvailable SinceAug. 15, 2018AvailabilityAcademic Institutions and Nonprofits only -
BS.YxiN.Twin1.a1-479.E152Q
Plasmid#17518DepositorInsertB. subtilis DEAD-box helicase YxiN
Tagsintein-chitin binding domainExpressionBacterialMutationE152Q mutationAvailable SinceMarch 21, 2008AvailabilityAcademic Institutions and Nonprofits only -
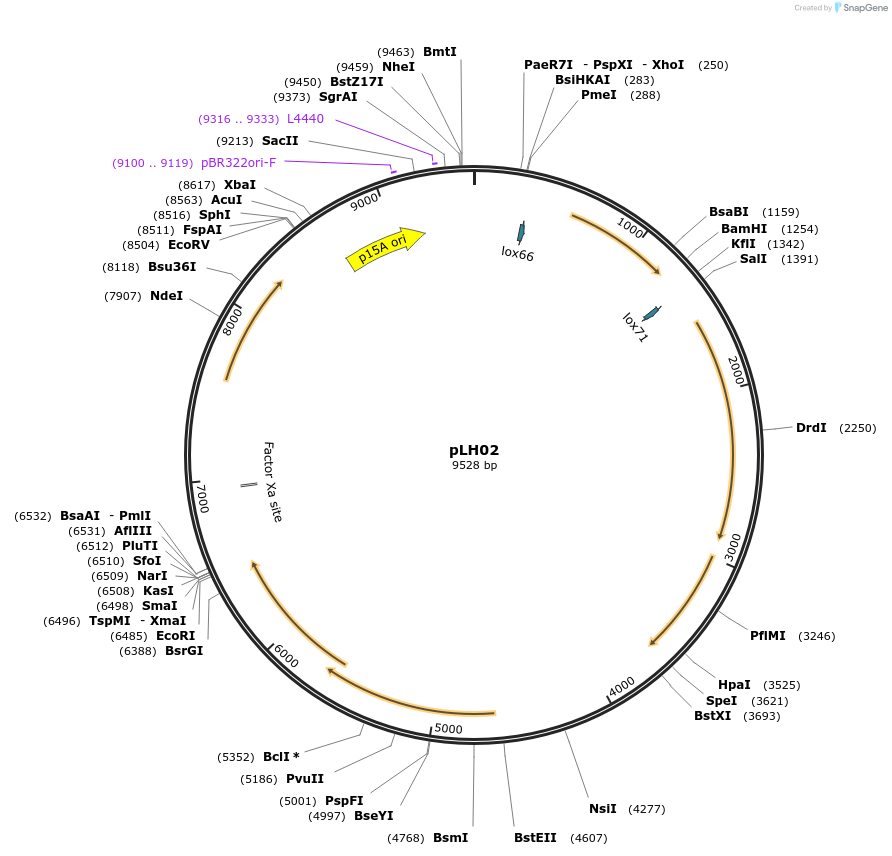
pLH02
Plasmid#117262PurposeGenome editing for Lactobacillus brevis ATCC367DepositorInsertPorfX-LVISKB_1732-30 , sapK-sappR
UseOtherAvailable SinceDec. 9, 2019AvailabilityAcademic Institutions and Nonprofits only -
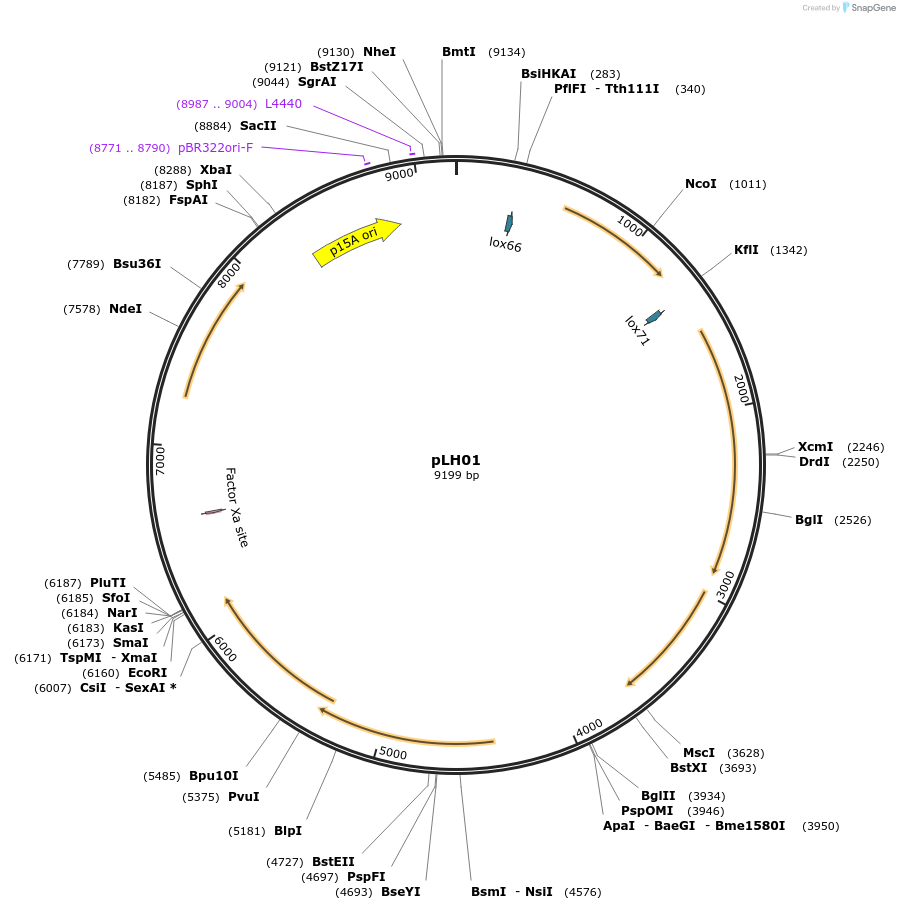
pLH01
Plasmid#117261PurposeGenome editing for Lactobacillus plantarum WCSF1DepositorInsertPorfX-Lp_0640-42, sapK-sappR
UseOtherAvailable SinceSept. 12, 2019AvailabilityAcademic Institutions and Nonprofits only -
pLenti-EF1α-IRES-EGFP-hMCIDAS
Plasmid#149696PurposeExpresses untagged human MCIDAS in mammalian cellsDepositorAvailable SinceJuly 24, 2020AvailabilityAcademic Institutions and Nonprofits only -
BS.YxiN.Twin1.a207-479.wt
Plasmid#17516DepositorInsertB. subtilis DEAD-box helicase YxiN second domain
Tagsintein-chitin binding domainExpressionBacterialMutationdeleted amino acid residues 1-206Available SinceMarch 21, 2008AvailabilityAcademic Institutions and Nonprofits only