We narrowed to 548 results for: PA-GFP
-
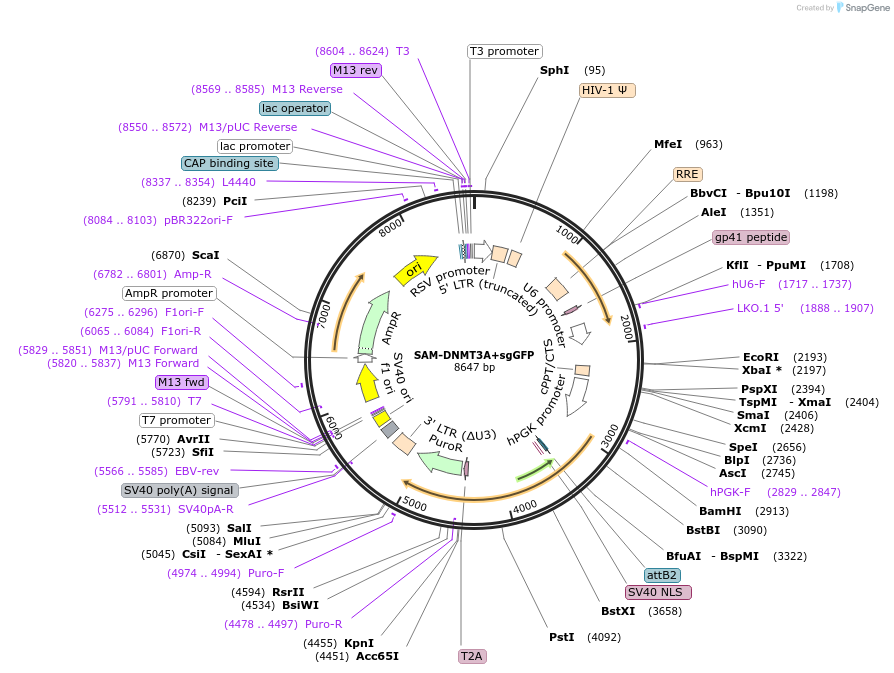
Plasmid#213165PurposeVector with sgGFP for induction of global DNA methylation.DepositorInsertsgGFP
UseCRISPR, Lentiviral, and Synthetic BiologyExpressionMammalianPromoterE1Fa and U6 and PGKAvailable SinceDec. 2, 2024AvailabilityAcademic Institutions and Nonprofits only -
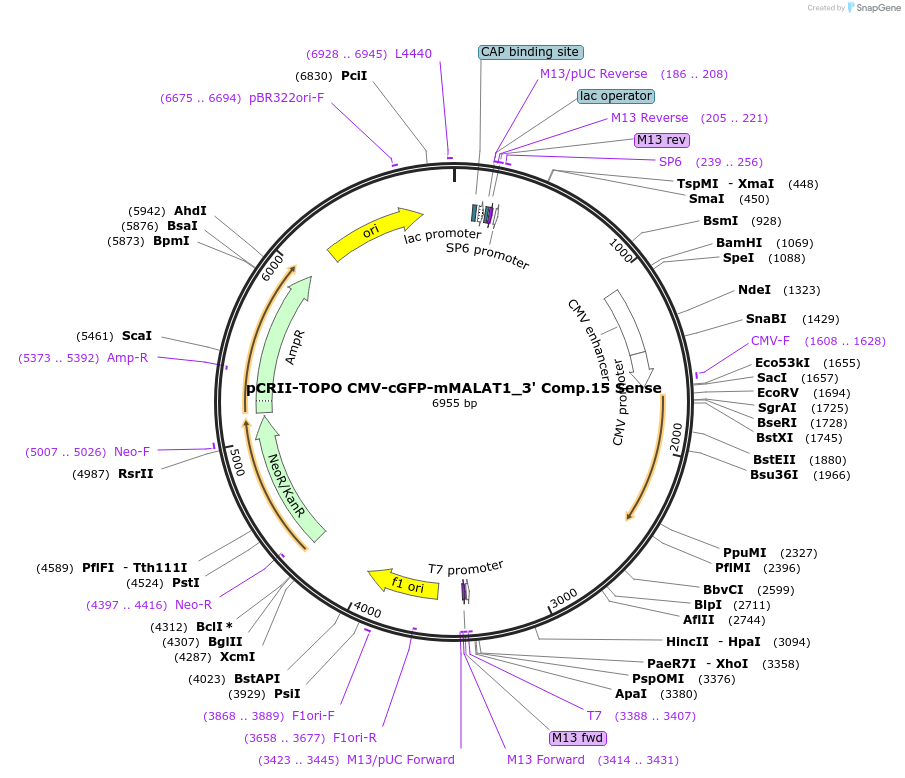
pCRII-TOPO CMV-cGFP-mMALAT1_3' Comp.15 Sense
Plasmid#46839PurposeExpress cGFP transcript ending in a mutant MALAT1 triple helix that causes the transcript to be stable but poorly translatedDepositorInsertCoral Green Fluorescent Protein (cGFP)
ExpressionMammalianPromoterCMVAvailable SinceSept. 25, 2013AvailabilityAcademic Institutions and Nonprofits only -
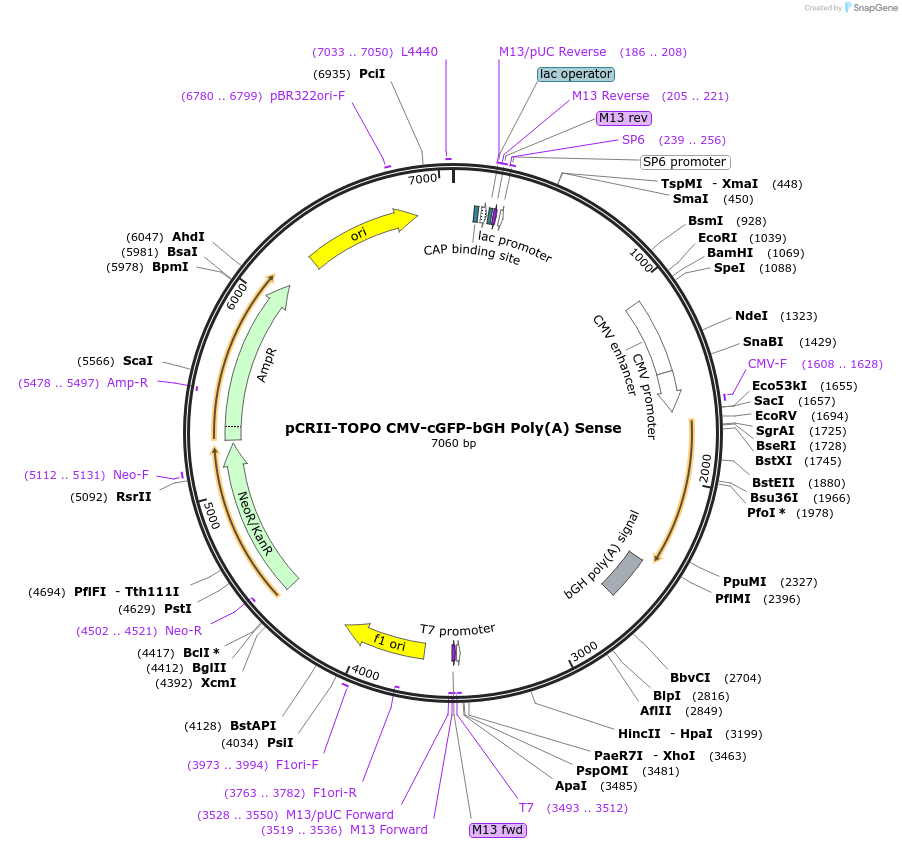
pCRII-TOPO CMV-cGFP-bGH Poly(A) Sense
Plasmid#46835PurposeExpress cGFP transcript ending in a poly(A) tail [using the bGH poly(A) signal]DepositorInsertCoral Green Fluorescent Protein (cGFP)
ExpressionMammalianPromoterCMVAvailable SinceSept. 25, 2013AvailabilityAcademic Institutions and Nonprofits only -
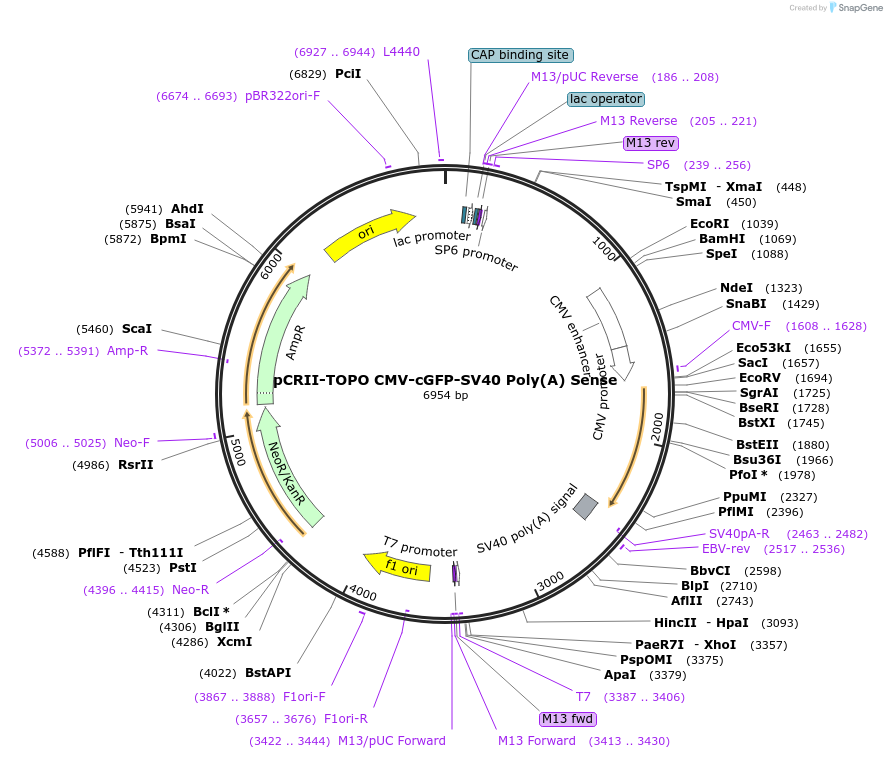
pCRII-TOPO CMV-cGFP-SV40 Poly(A) Sense
Plasmid#46836PurposeExpress cGFP transcript ending in a poly(A) tail [using the SV40 poly(A) signal]DepositorInsertCoral Green Fluorescent Protein (cGFP)
ExpressionMammalianPromoterCMVAvailable SinceSept. 25, 2013AvailabilityAcademic Institutions and Nonprofits only -
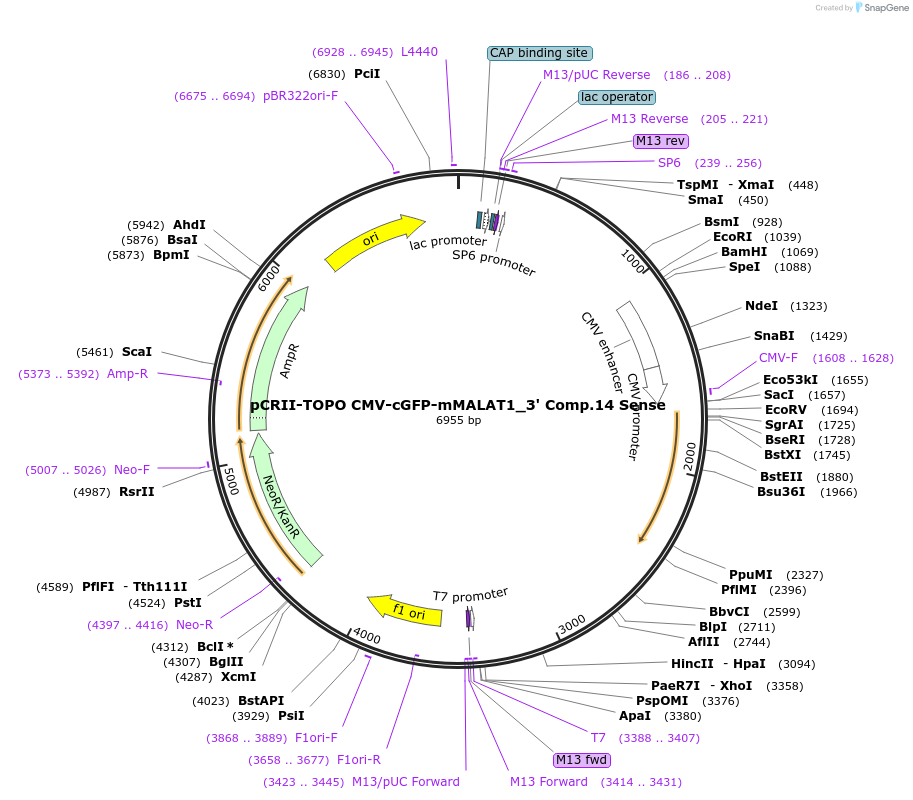
pCRII-TOPO CMV-cGFP-mMALAT1_3' Comp.14 Sense
Plasmid#46838PurposeExpress cGFP transcript ending in the minimal functional MALAT1 triple helixDepositorInsertCoral Green Fluorescent Protein (cGFP)
ExpressionMammalianPromoterCMVAvailable SinceSept. 25, 2013AvailabilityAcademic Institutions and Nonprofits only -
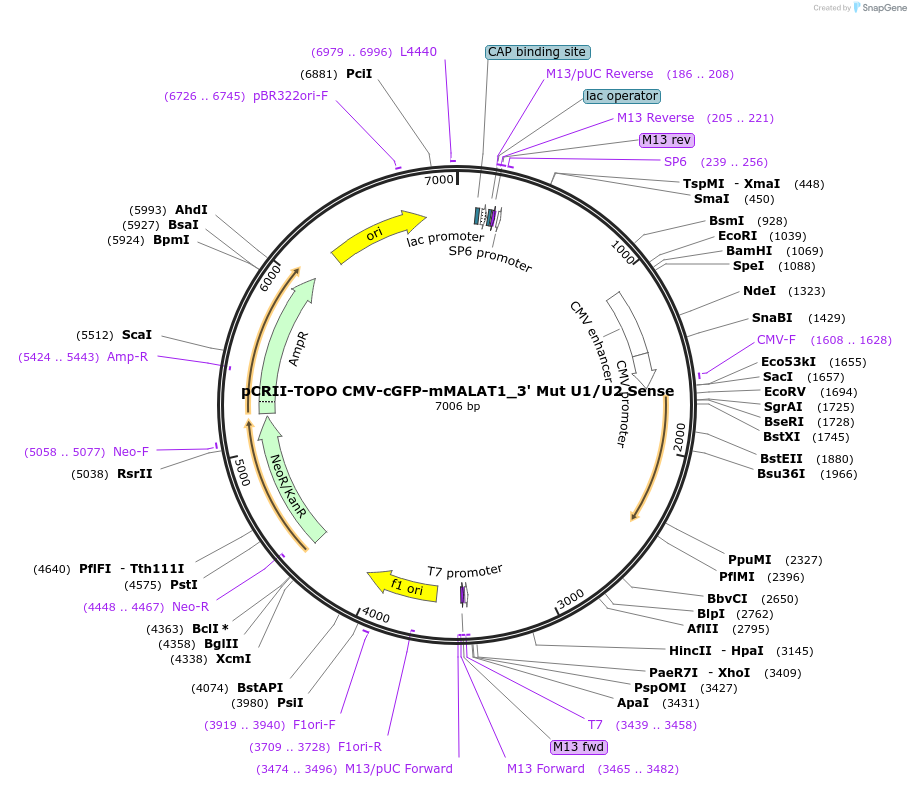
pCRII-TOPO CMV-cGFP-mMALAT1_3' Mut U1/U2 Sense
Plasmid#46837PurposeExpress cGFP transcript ending in a mutant MALAT1 triple helix which causes the transcript to be degradedDepositorInsertCoral Green Fluorescent Protein (cGFP)
ExpressionMammalianPromoterCMVAvailable SinceSept. 25, 2013AvailabilityAcademic Institutions and Nonprofits only -
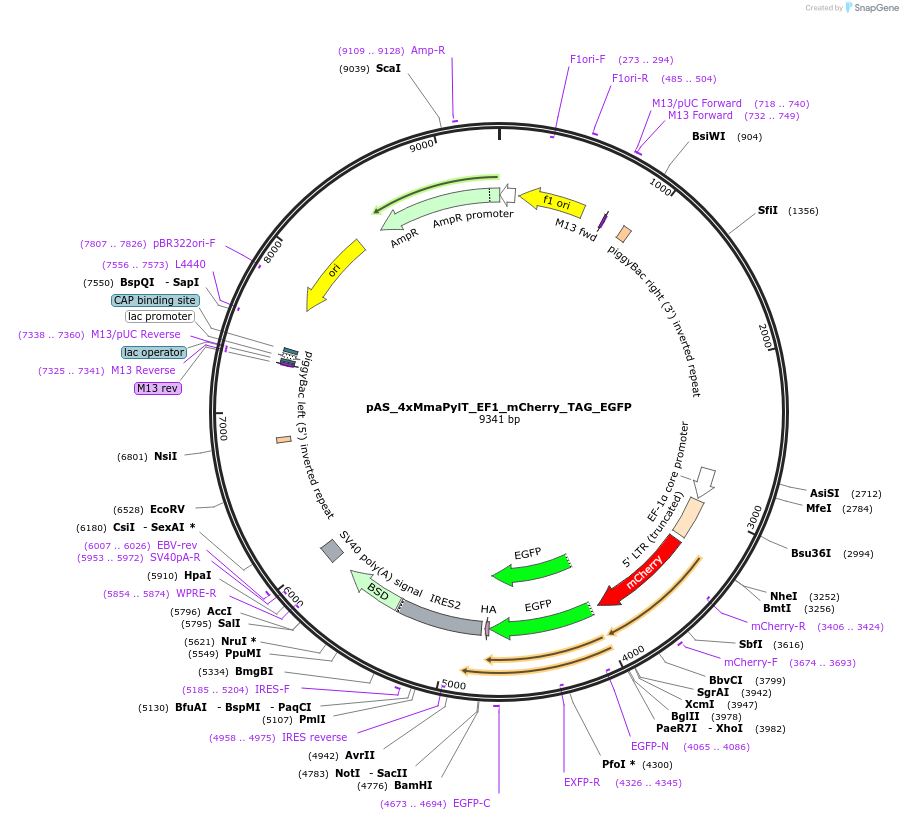
pAS_4xMmaPylT_EF1_mCherry_TAG_EGFP
Plasmid#174893Purposeamber suppression reporter mCherry-TAG-EGFP expression, with MmaPylT amber suppressor tRNA cassetteDepositorInsertmCherry and EGFP
ExpressionMammalianPromoterEF1Available SinceNov. 8, 2023AvailabilityAcademic Institutions and Nonprofits only -
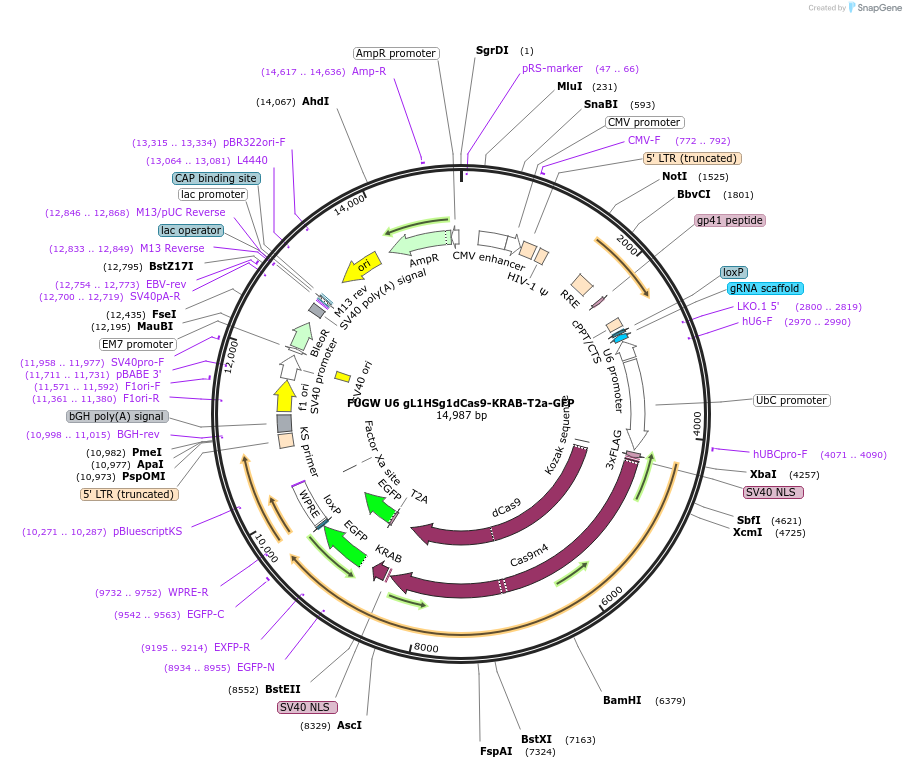
FUGW U6 gL1HSg1dCas9-KRAB-T2a-GFP
Plasmid#234882PurposeL1HS-silencing plasmid (CRISPRi gRNA1)DepositorInsertL1HS-silencing plasmid (CRISPRi gRNA1)
UseCRISPR and LentiviralTagsGFPExpressionMammalianAvailable SinceApril 16, 2025AvailabilityAcademic Institutions and Nonprofits only -
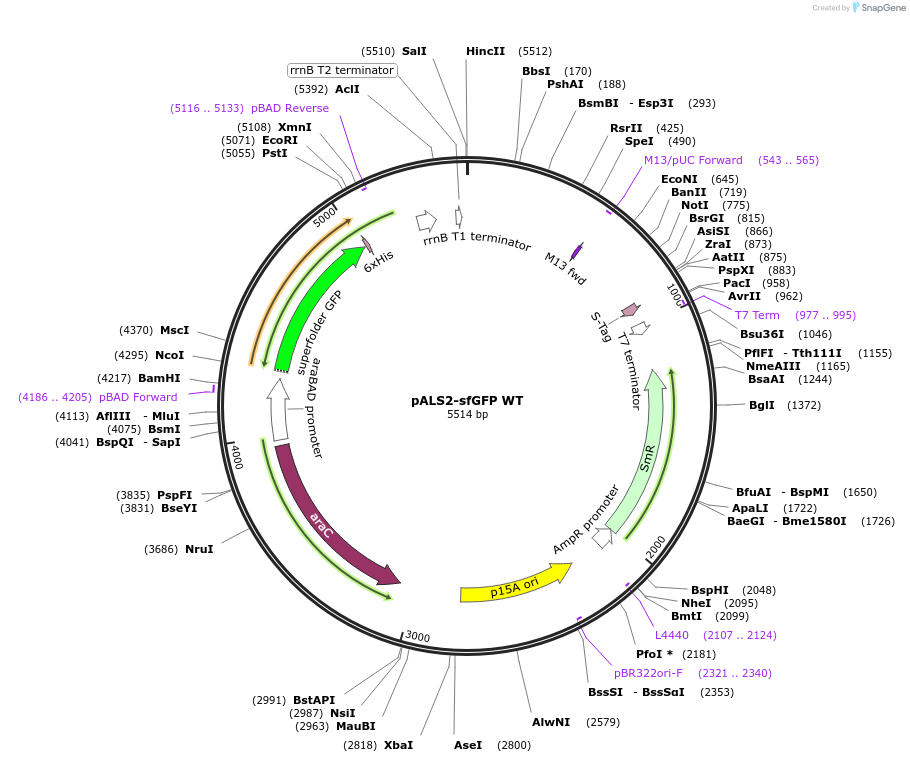
pALS2-sfGFP WT
Plasmid#197575PurposeFluorescence control plasmid for selecting ncAA-encoding Methanomethylophilus alvus Pyl-RS mutants. Expresses sfGFP-WT and M. alvus Pyl-tRNA(6). p15a origin of replicationDepositorInsertssfGFP WT
M. alvus Pyl-tRNA (6)
TagsHis6ExpressionBacterialPromoteraraC and lppAvailable SinceJune 14, 2023AvailabilityAcademic Institutions and Nonprofits only -
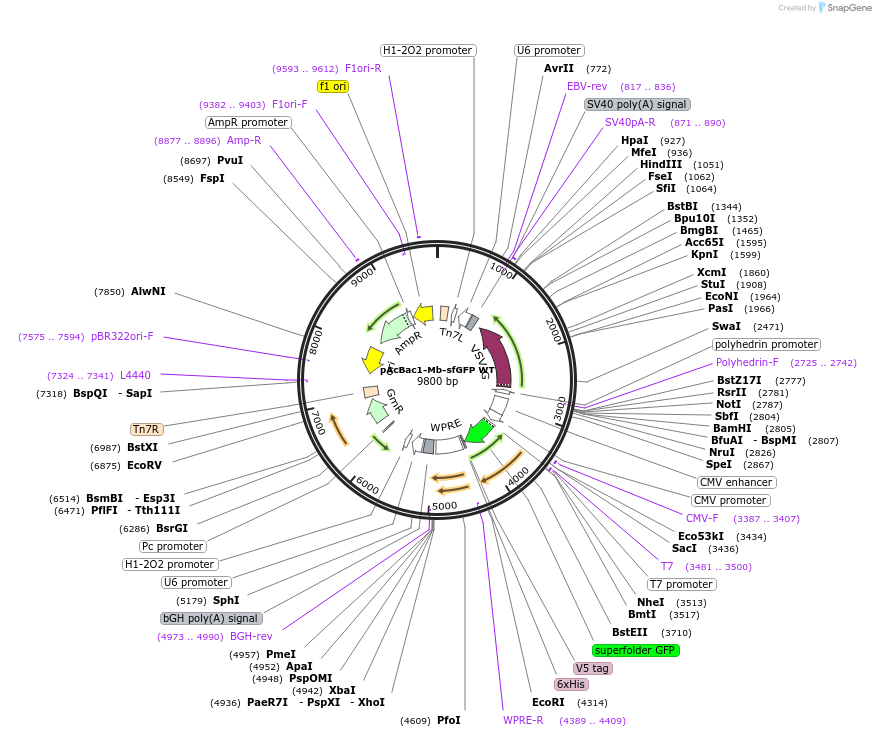
pAcBac1-Mb-sfGFP WT
Plasmid#197569PurposeExpression of sfGFP WT with Methanosarcina barkeri (Mb) Pyl-tRNA for the encoding of non-canonical amino acids at TAG codons in HEK293 cellsDepositorInsertssfGFP WT
Pyl-tRNA (4 copies)
TagsV5-His6ExpressionMammalianPromoterCMV and U6/H1Available SinceApril 6, 2023AvailabilityAcademic Institutions and Nonprofits only -
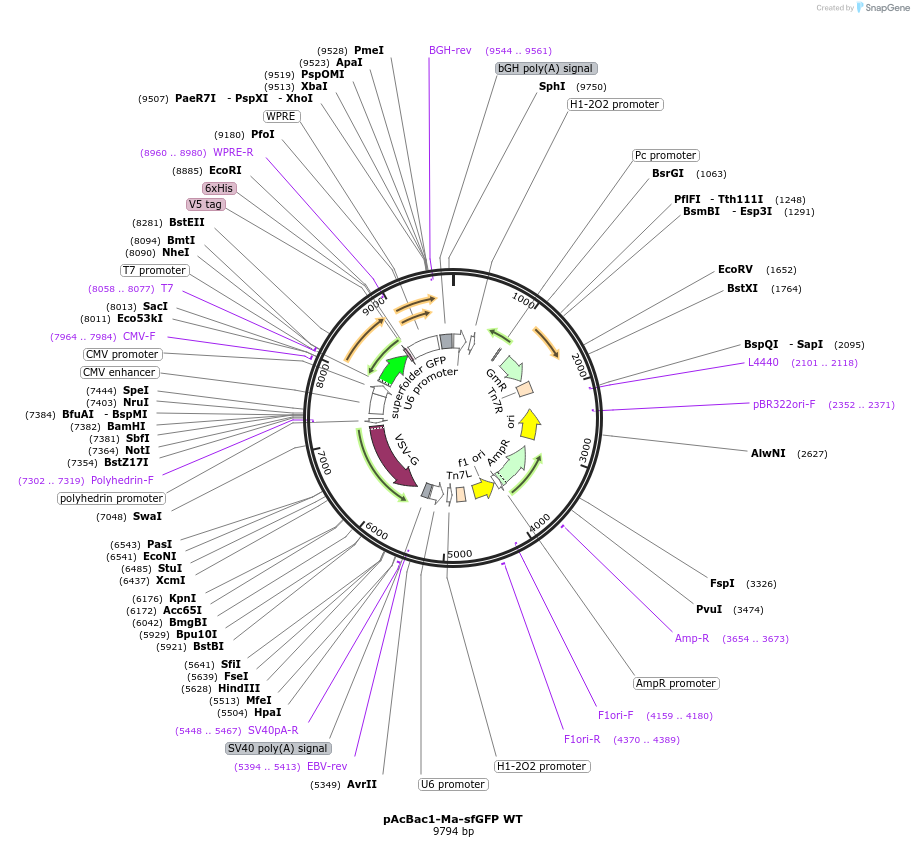
pAcBac1-Ma-sfGFP WT
Plasmid#197567PurposeExpression of sfGFP WT with Methanomethylophilus alvus (Ma) Pyl-tRNA for the encoding of non-canonical amino acids at TAG codons in HEK293 cellsDepositorInsertssfGFP WT
M. alvus Pyl-tRNA (6) (4x copies)
TagsV5-His6ExpressionMammalianPromoterCMV and U6/H1Available SinceApril 4, 2023AvailabilityAcademic Institutions and Nonprofits only -
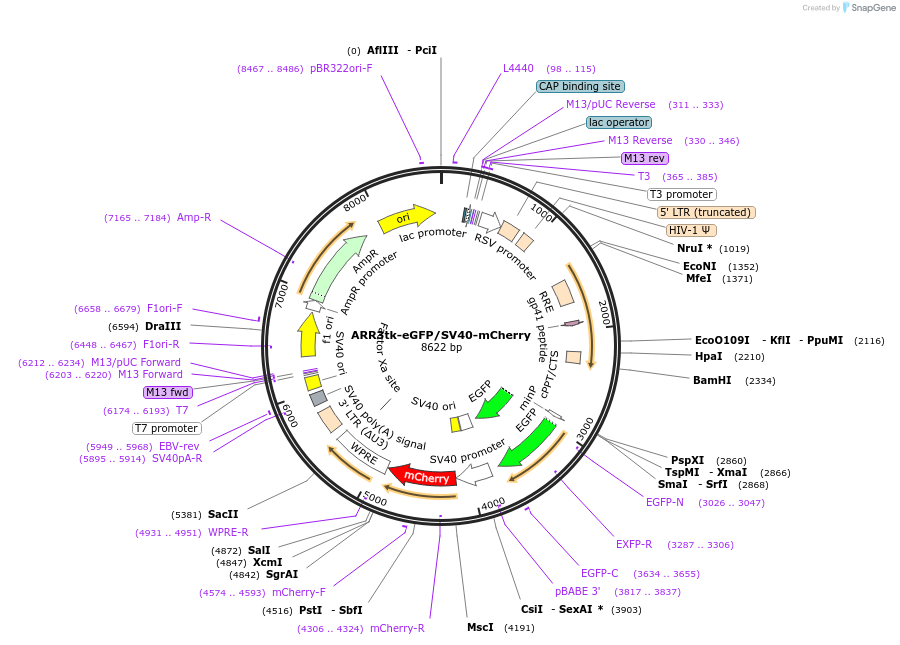
ARR3tk-eGFP/SV40-mCherry
Plasmid#132360PurposeTo measure transcriptional activity of Androgen ReceptorDepositorInsertAR responsive elements (AR Human)
UseLentiviralAvailable SinceOct. 21, 2019AvailabilityAcademic Institutions and Nonprofits only -
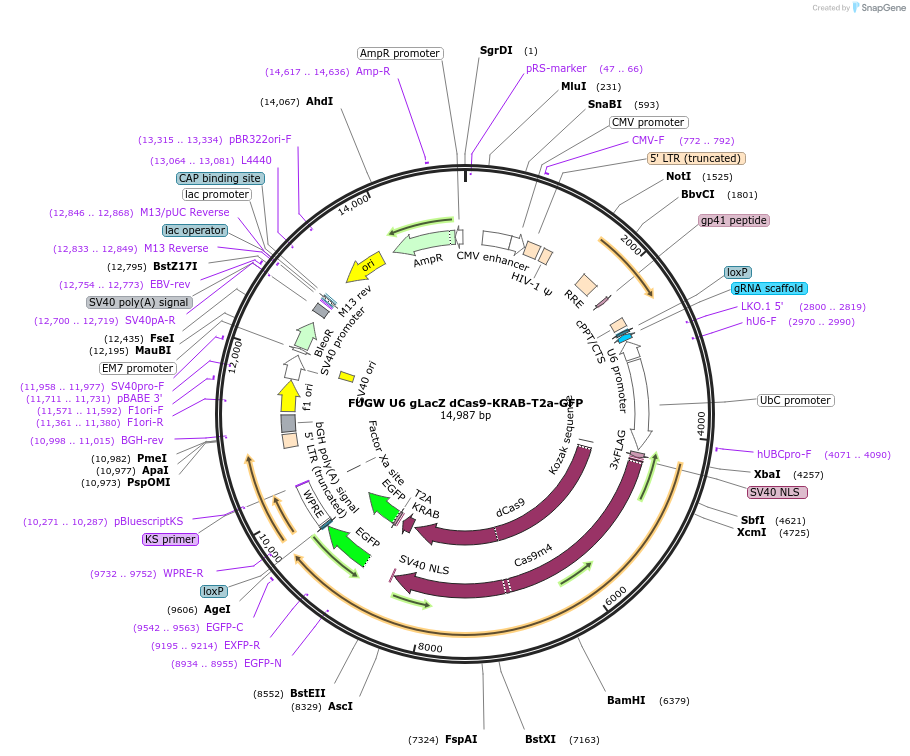
FUGW U6 gLacZ dCas9-KRAB-T2a-GFP
Plasmid#234883Purposenon-targeting CRISPRi controlDepositorInsertL1HS gRNA
UseCRISPR and LentiviralTagsGFPExpressionMammalianAvailable SinceApril 22, 2025AvailabilityAcademic Institutions and Nonprofits only -
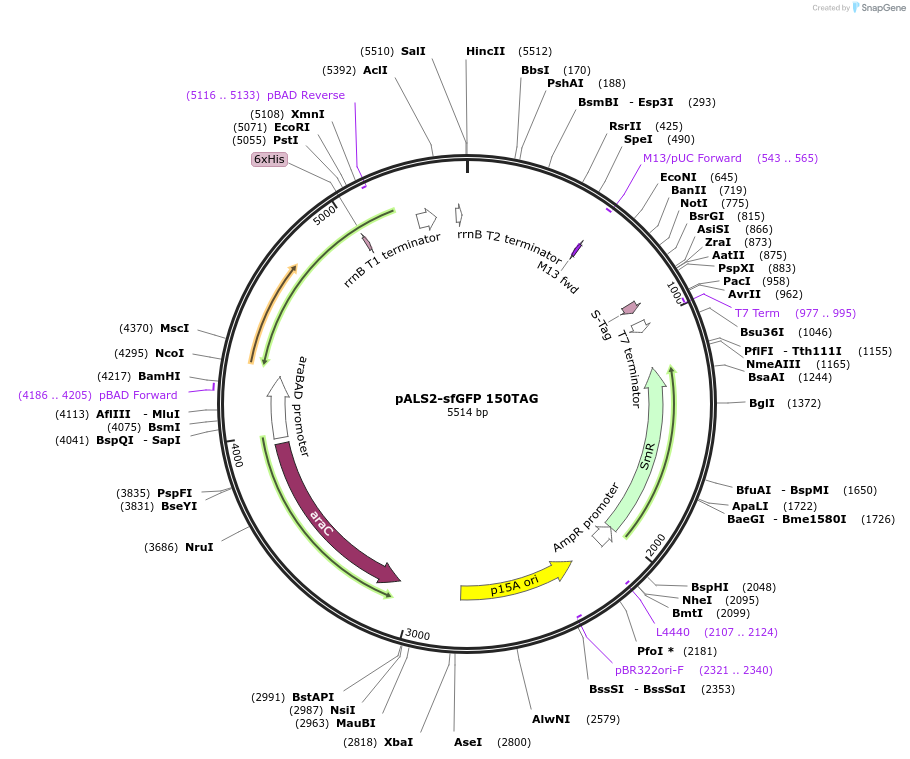
pALS2-sfGFP 150TAG
Plasmid#197574PurposeFluorescence selection plasmid for selecting ncAA-encoding Methanomethylophilus alvus Pyl-RS mutants. Expresses sfGFP-150TAG and M. alvus Pyl-tRNA(6). p15a origin of replication.DepositorInsertssfGFP 150TAG
M. alvus Pyl-tRNA (6)
TagsHis6ExpressionBacterialMutationN150TAGPromoteraraC and lppAvailable SinceMarch 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
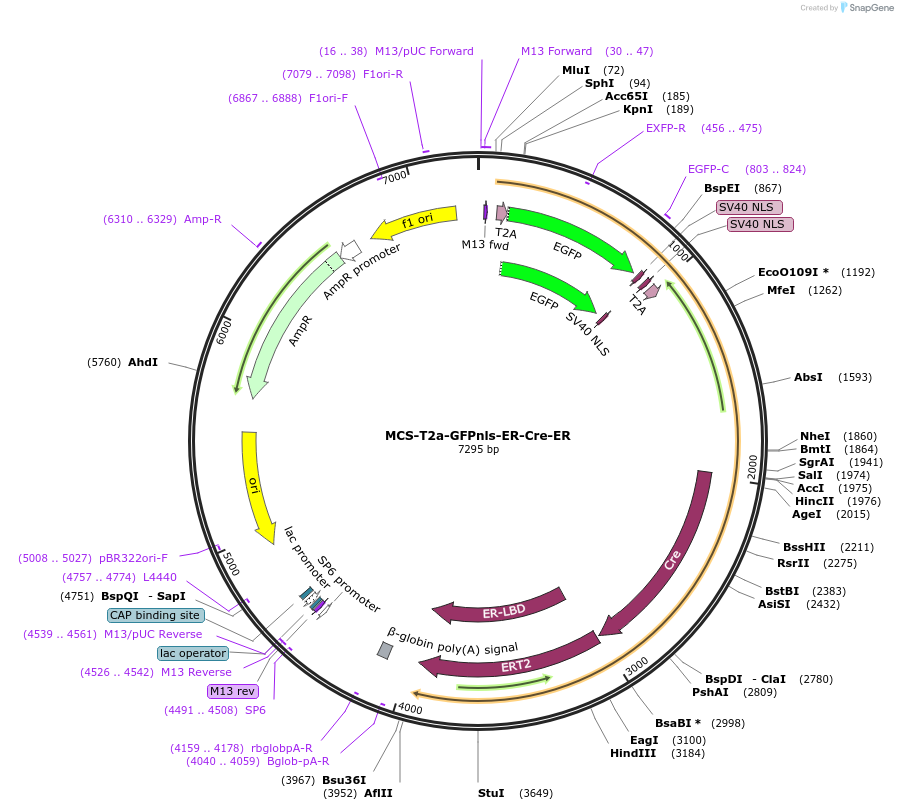
MCS-T2a-GFPnls-ER-Cre-ER
Plasmid#200104PurposeVector For TwistDepositorTypeEmpty backboneUseUnspecifiedAvailable SinceJune 9, 2023AvailabilityAcademic Institutions and Nonprofits only -
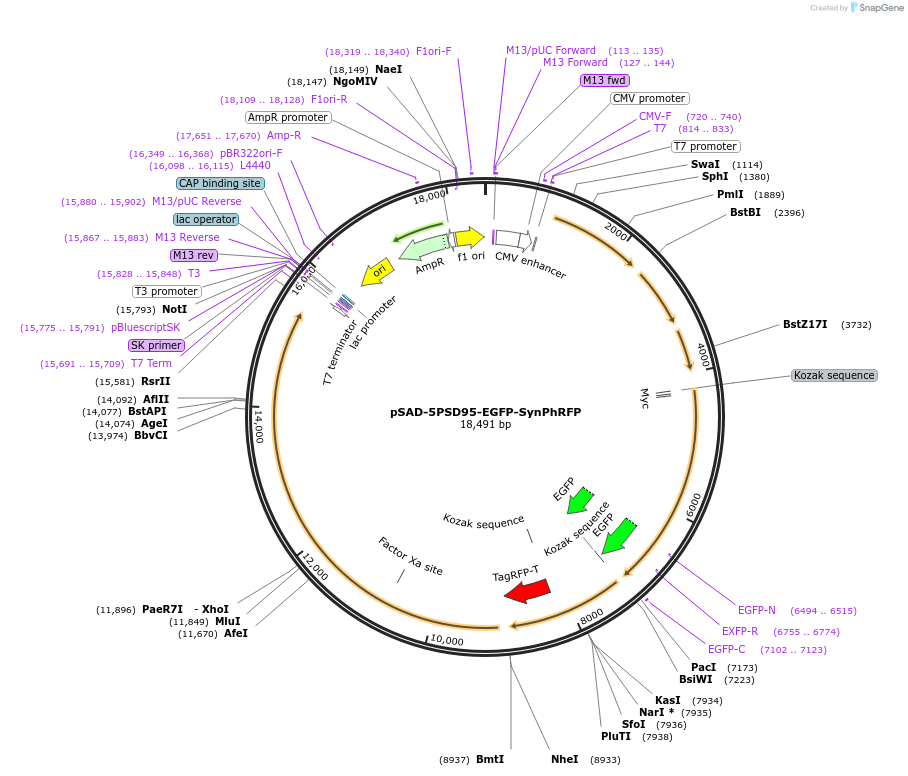
pSAD-5PSD95-EGFP-SynPhRFP
Plasmid#217979PurposeG-Deleted Rabies genomic plasmid to express 5PSD95-EGFP and SynPhRFPDepositorUseG-deleted rabiesTagsEGFP and RFPPromoterNoneAvailable SinceApril 22, 2024AvailabilityAcademic Institutions and Nonprofits only -
MSCV-dGFP-JARID1C A388P
Plasmid#74778Purposeretroviral expression of JARID1C A388PDepositorAvailable SinceMay 3, 2016AvailabilityAcademic Institutions and Nonprofits only -
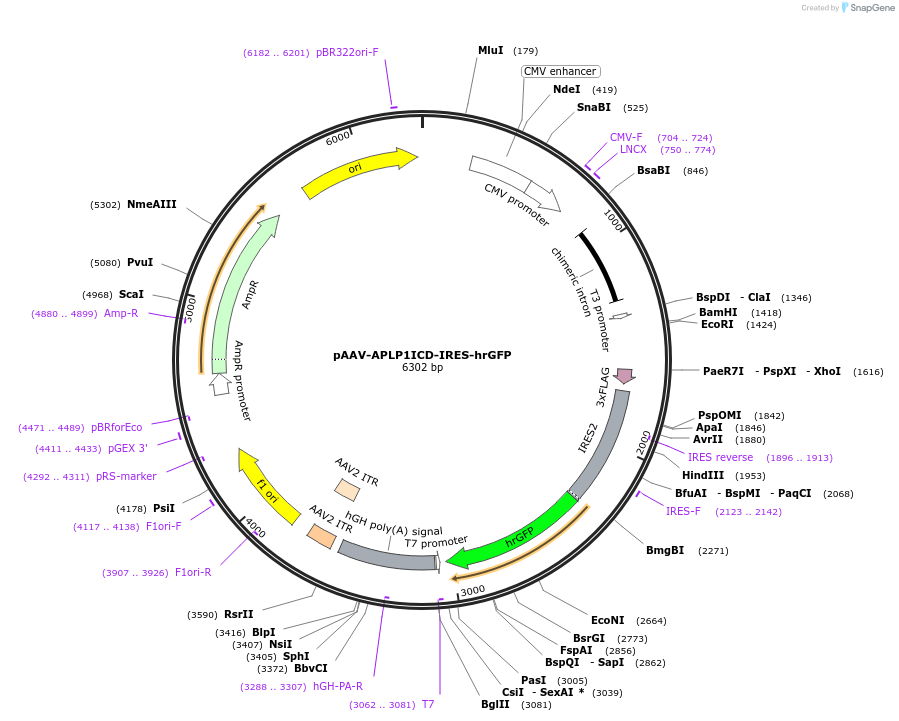
pAAV-APLP1ICD-IRES-hrGFP
Plasmid#107546PurposeAAV-mediated expression of last 50 amino acids of APLP1 protein (APLP1ICD) and IRES-mediated co-expression of hrGFP to recognize labelled cellsDepositorAvailable SinceMay 1, 2018AvailabilityAcademic Institutions and Nonprofits only -
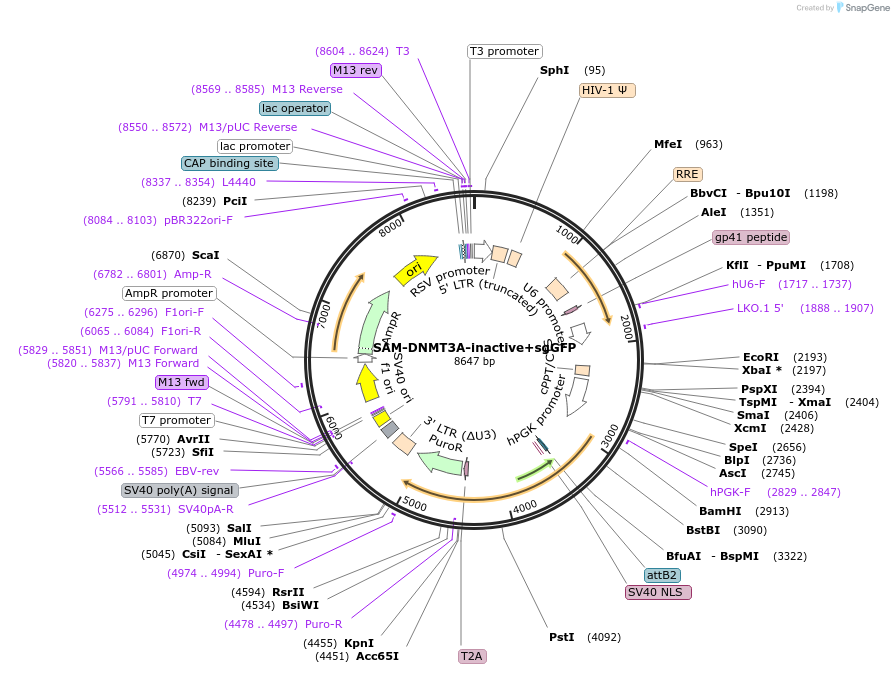
SAM-DNMT3A-inactive+sgGFP
Plasmid#213169PurposeVector with sgGFP used as a negative control (inactive DNMT3A) for induction of global DNA methylation.DepositorInsertsgGFP
UseCRISPR, Lentiviral, and Synthetic BiologyExpressionMammalianPromoterE1Fa and U6Available SinceDec. 2, 2024AvailabilityAcademic Institutions and Nonprofits only -
sgGFP_3
Plasmid#78165PurposesgGFPDepositorInsertGFP
UseCRISPR and LentiviralExpressionMammalianPromoterhU6Available SinceJune 22, 2016AvailabilityAcademic Institutions and Nonprofits only