We narrowed to 4,769 results for: PC;
-
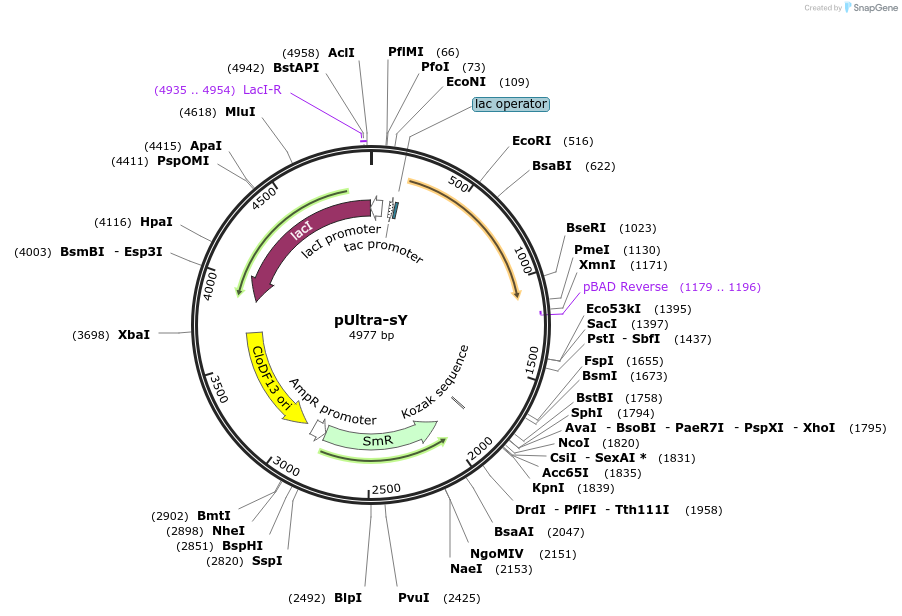
Plasmid#82417PurposeExpresses Mj aaRS for sulfotyrosine and Mj tRNA for decoding UAG codonsDepositorInsertMethanococcus jannaschii aaRS for sulfotyrosine
ExpressionBacterialMutationTyr32Leu, Leu65Pro, Asp158Gly, Ile159Cys, Leu162L…PromotertacIAvailable SinceDec. 6, 2016AvailabilityAcademic Institutions and Nonprofits only -
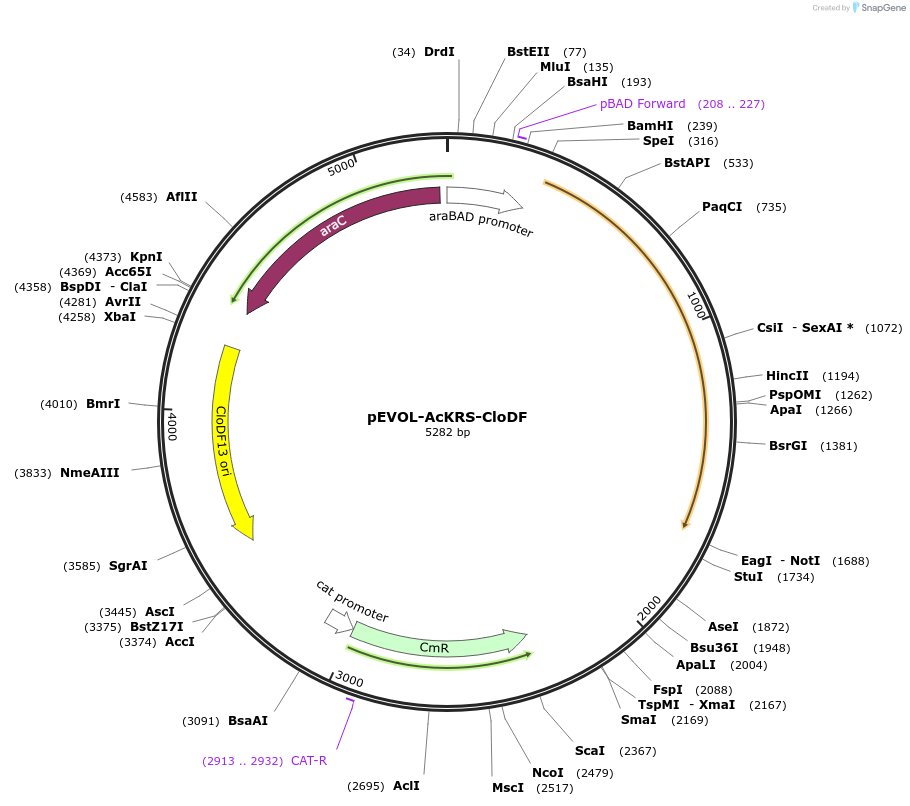
pEVOL-AcKRS-CloDF
Plasmid#127415PurposeMutant of mmPylRS designed for incorporation of non-canonical amino acids (acyl-lysine derivatives) in to M13 bacteriophageDepositorInsertPyrrolysyl-tRNA synthetase/pyrrolysyl -tRNA pair
ExpressionBacterialPromoteraraBADAvailable SinceApril 15, 2022AvailabilityAcademic Institutions and Nonprofits only -
pcDNA3/HIP1/ΔC
Plasmid#31250DepositorInsertHuntingtin-interacting protein 1 (HIP1 Human)
ExpressionBacterialMutationDeltion of nucleotides 1336-2513 1386bp C to T…Available SinceSept. 29, 2011AvailabilityAcademic Institutions and Nonprofits only -
pET15b-Fab1 (Francisella tularensis)
Plasmid#61690PurposeInducible expression of the FabI enzyme (enoyl reductase) from Francisella tularensisDepositorInsertenoyl-[acyl-carrier-protein] reductase (NADH)
TagsHisExpressionBacterialPromoterT7Available SinceApril 10, 2015AvailabilityAcademic Institutions and Nonprofits only -
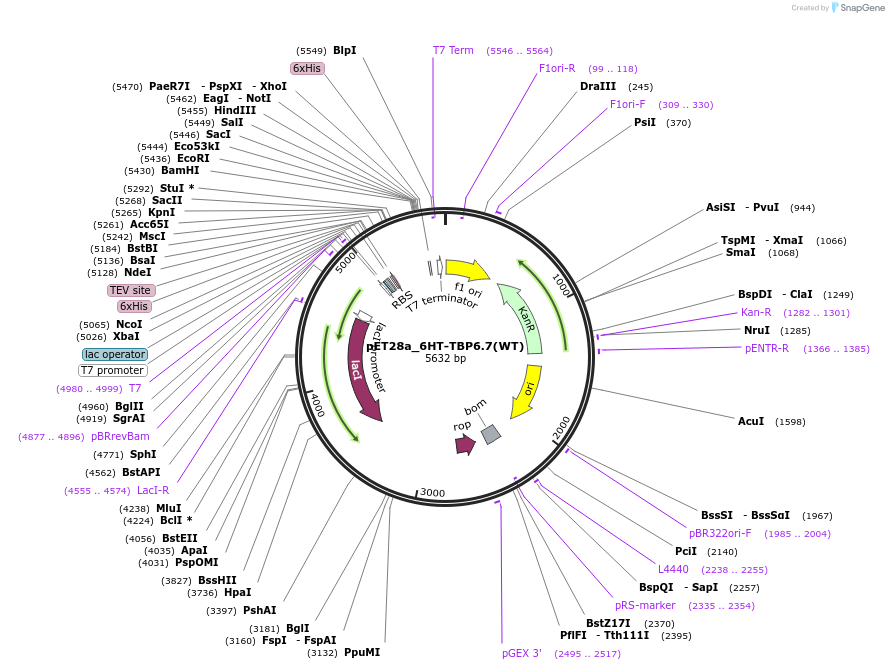
pET28a_6HT-TBP6.7(WT)
Plasmid#112720PurposeWild-type TBP6.7 was one of the first evolved TAR-binding proteins to bind with exceptional affinity. It was this protein that was used to obtain co-crystals with WT TAR RNA.DepositorInsert6His-TEV-TBP6.7(WT)
Tags6His-TEVExpressionBacterialPromoterT7Available SinceAug. 8, 2018AvailabilityAcademic Institutions and Nonprofits only -
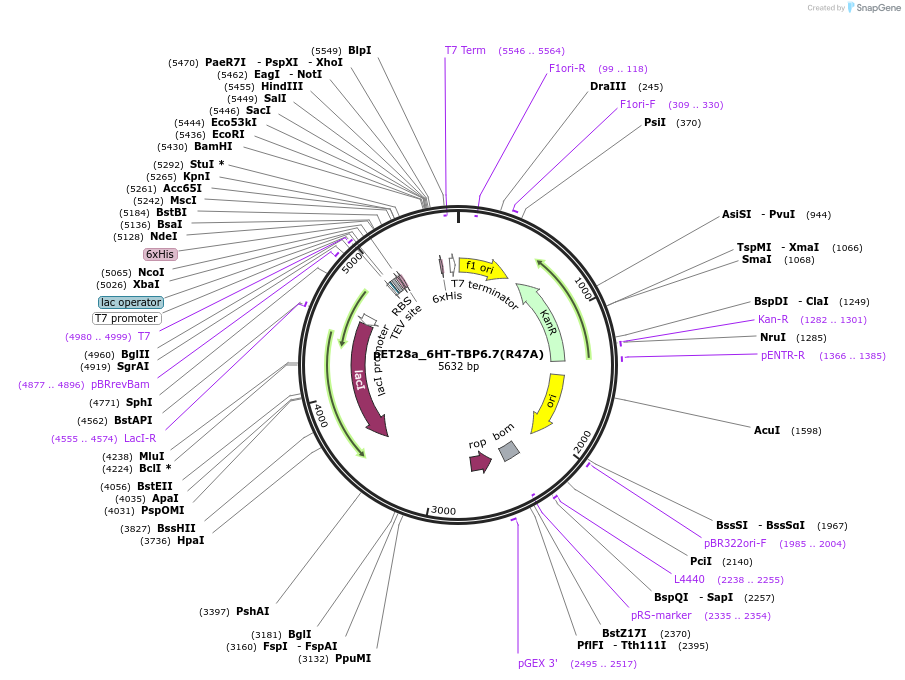
pET28a_6HT-TBP6.7(R47A)
Plasmid#112722PurposeArg47 within TBP6.7 is the most critical residue in terms of forming interactions with WT TAR. Mutating this amino acid to Ala results in ~600-fold reduction in Kd.DepositorInsert6His-TEV-TBP6.7(R47A)
Tags6His-TEVExpressionBacterialMutationR47APromoterT7Available SinceAug. 8, 2018AvailabilityAcademic Institutions and Nonprofits only -
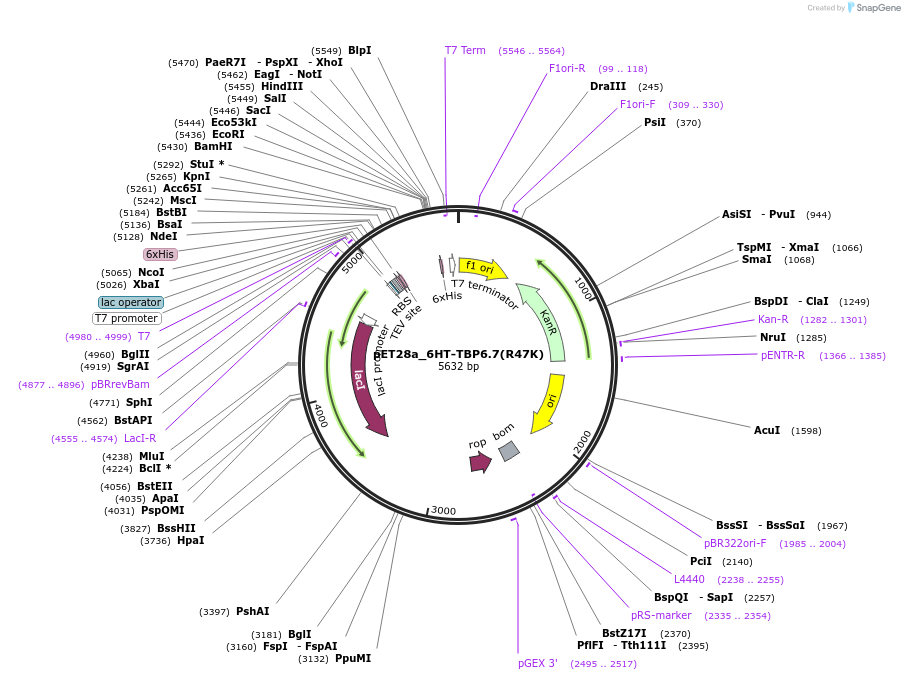
pET28a_6HT-TBP6.7(R47K)
Plasmid#112723PurposeArg47 of TBP6.7 interacts with Hoogsteen edge of TAR's Gua26. Mutating this amino acid to Lys reduces binding to the WT TAR RNA ~330-fold.DepositorInsert6His-TEV-TBP6.7(R47K)
Tags6His-TEVExpressionBacterialMutationR47KPromoterT7Available SinceAug. 8, 2018AvailabilityAcademic Institutions and Nonprofits only -
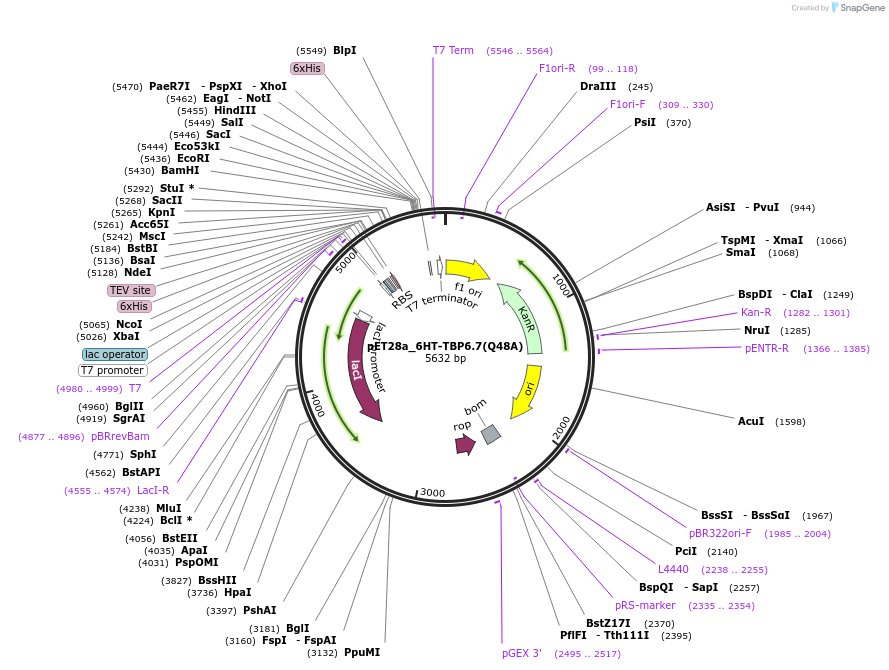
pET28a_6HT-TBP6.7(Q48A)
Plasmid#112724PurposeGln48 contacts the backbone of TAR RNA as well as mediates intramoecular interaction to stabilize the conformation of beta2-beta3 loop within TBP6.7.DepositorInsert6His-TEV-TBP6.7(Q48A)
Tags6His-TEVExpressionBacterialMutationQ48APromoterT7Available SinceAug. 8, 2018AvailabilityAcademic Institutions and Nonprofits only -
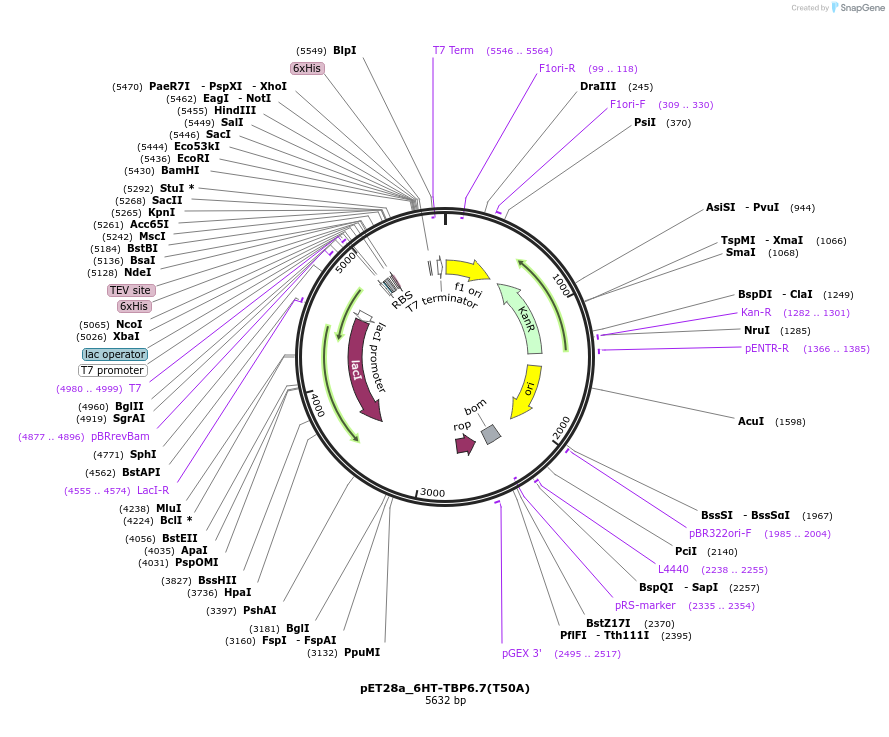
pET28a_6HT-TBP6.7(T50A)
Plasmid#112727PurposeLav-evolved amino acid T50 within TBPs is important in engaging intramolecularly to steer also lab-evolved R49 into conformation compatible binding to RNA.DepositorInsert6His-TEV-TBP6.7(T50A)
Tags6His-TEVExpressionBacterialMutationT50APromoterT7Available SinceAug. 8, 2018AvailabilityAcademic Institutions and Nonprofits only -
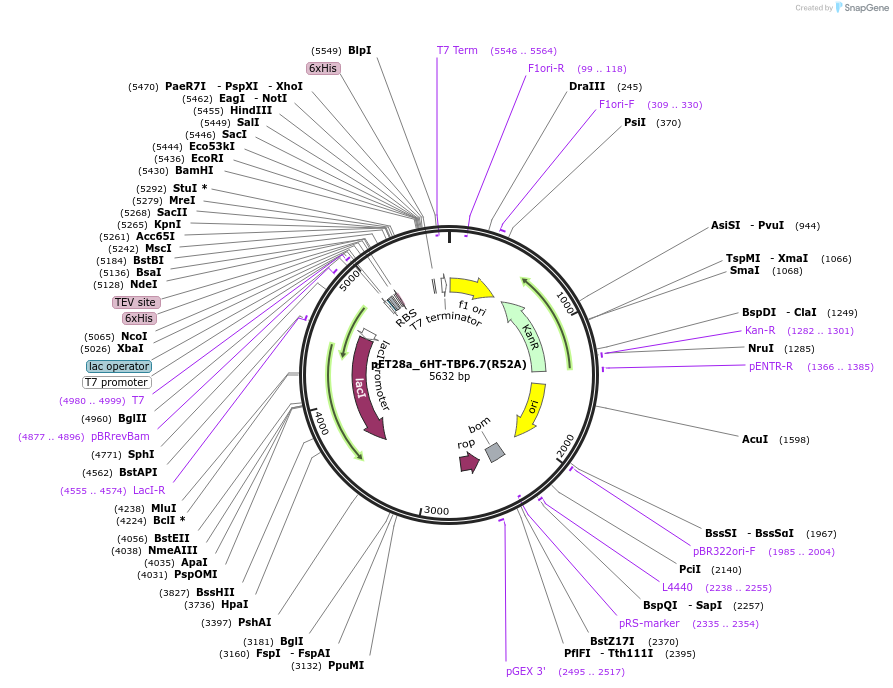
pET28a_6HT-TBP6.7(R52A)
Plasmid#112729Purpose52nd position is Arg in both wild-type U1A and TBPs, but the RNA recognition by this residue in both types of proteins is fundamentally different.DepositorInsert6His-TEV-TBP6.7(R52A)
Tags6His-TEVExpressionBacterialMutationR52APromoterT7Available SinceAug. 8, 2018AvailabilityAcademic Institutions and Nonprofits only -
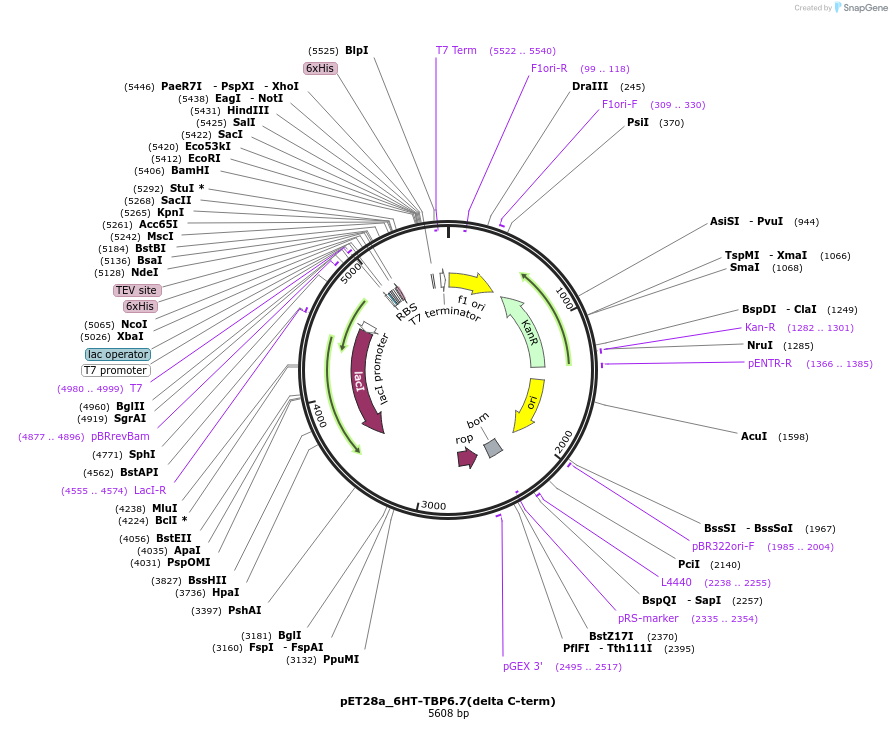
pET28a_6HT-TBP6.7(delta C-term)
Plasmid#112731PurposeThis mutant of TBP6.7 is lacking residues 91-98 at C-terminus, and was used to test the involvement of those residues in TAR recognition.DepositorInsert6His-TEV-TBP6.7(delta C-term)
Tags6His-TEVExpressionBacterialMutationdelta C-termPromoterT7Available SinceAug. 8, 2018AvailabilityAcademic Institutions and Nonprofits only -
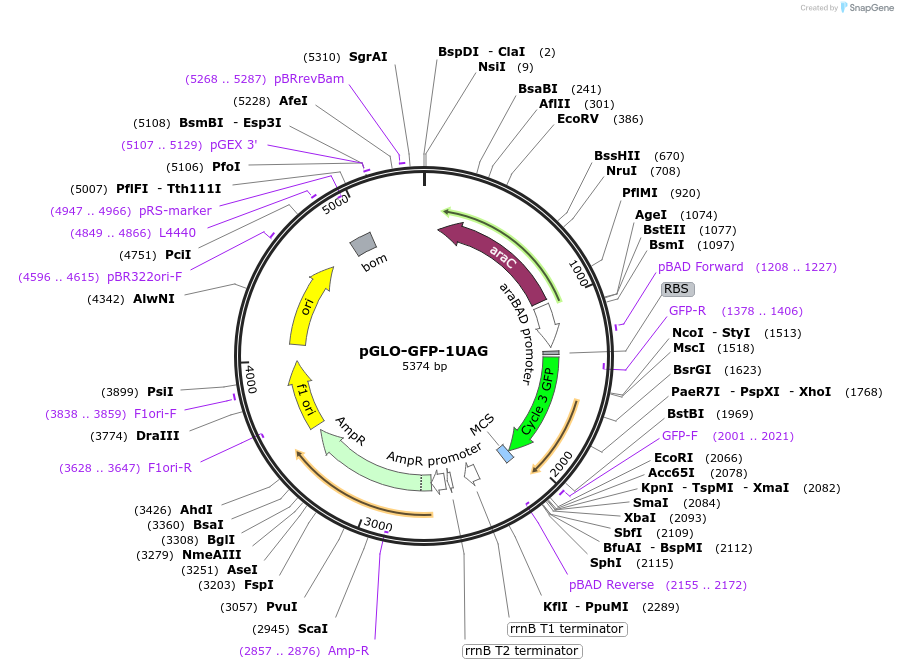
pGLO-GFP-1UAG
Plasmid#82500PurposeExpresses GFP with 1 UAG codon at the amino acid position 3DepositorInsertGreen Fluorescent Protein with 1 UAG codon
ExpressionBacterialMutationAdded an UAG codon at the amino acid position 3PromoterAraBADAvailable SinceSept. 22, 2016AvailabilityAcademic Institutions and Nonprofits only -
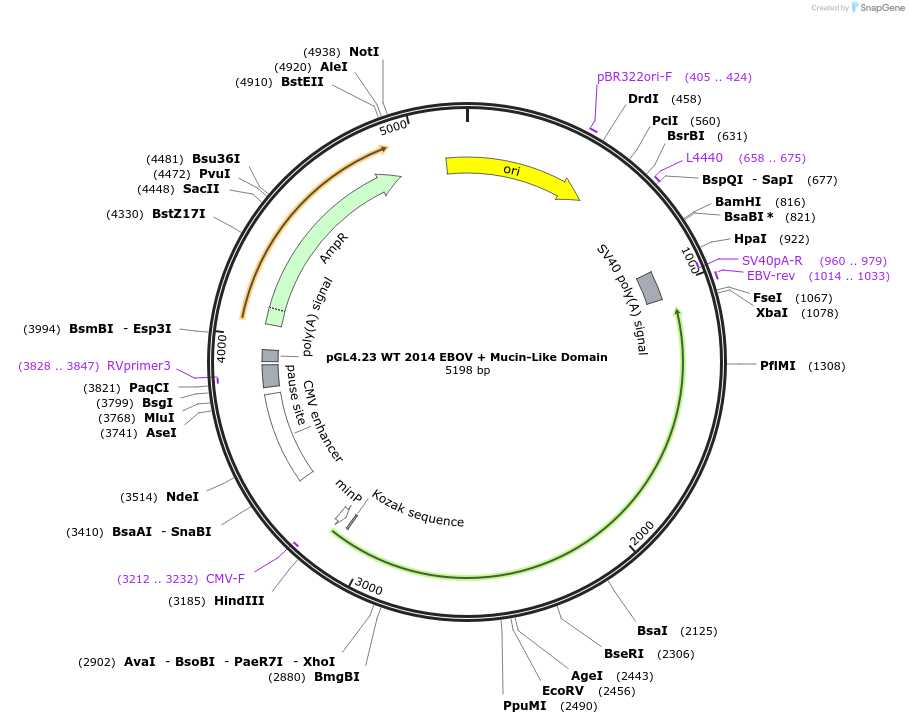
pGL4.23 WT 2014 EBOV + Mucin-Like Domain
Plasmid#85794PurposeEbola Virus (Makona) Glycoprotein in CMV driven mammalian expression vectorDepositorInsertMakona Ebola Virus Glycoprotein
ExpressionMammalianPromoterCMV IEAvailable SinceJan. 11, 2017AvailabilityAcademic Institutions and Nonprofits only -
plenti6.3/hCx43-stop
Plasmid#27383DepositorAvailable SinceJan. 20, 2011AvailabilityAcademic Institutions and Nonprofits only -
pcDNAz-mBACEflag (C474A/C478A/C482A/C485A)
Plasmid#26736DepositorInsertBACE (Bace1 Mouse)
TagsFLAGExpressionMammalianMutationCys 474, 478, 482, and 485 mutated to AlaAvailable SinceDec. 3, 2010AvailabilityAcademic Institutions and Nonprofits only -
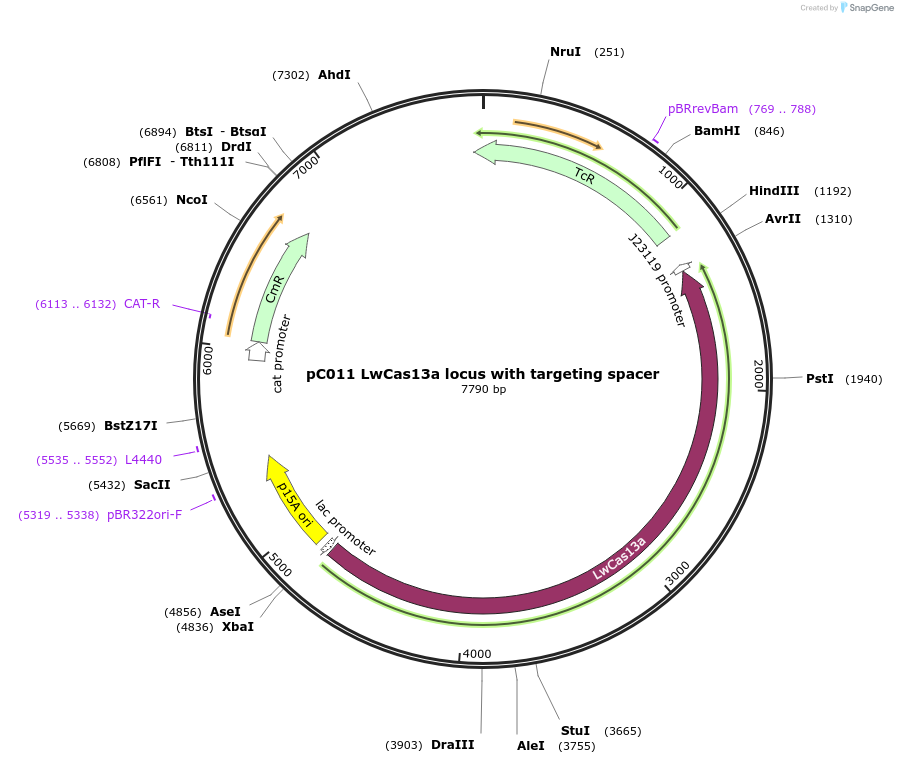
pC011 LwCas13a locus with targeting spacer
Plasmid#91903PurposeLwCas13a locus into pACYC184 with targeting spacerDepositorInsertLwCas13a with targeting spacer
UseCRISPRExpressionBacterialAvailable SinceOct. 16, 2017AvailabilityIndustry, Academic Institutions, and Nonprofits -
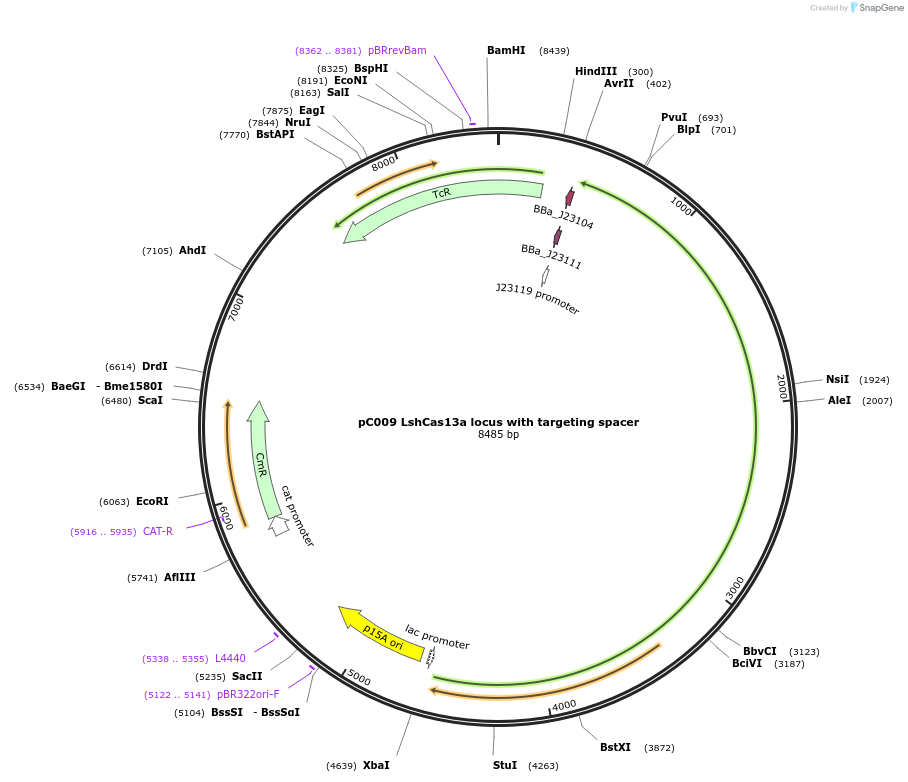
pC009 LshCas13a locus with targeting spacer
Plasmid#91900PurposeLshCas13a locus into pACYC184 with targeting spacerDepositorInsertLshCas13a locus with targeting spacer
ExpressionBacterialAvailable SinceOct. 16, 2017AvailabilityIndustry, Academic Institutions, and Nonprofits -
pTRE-TIGHT-Cx43-eYFP
Plasmid#31807DepositorAvailable SinceSept. 14, 2011AvailabilityAcademic Institutions and Nonprofits only -
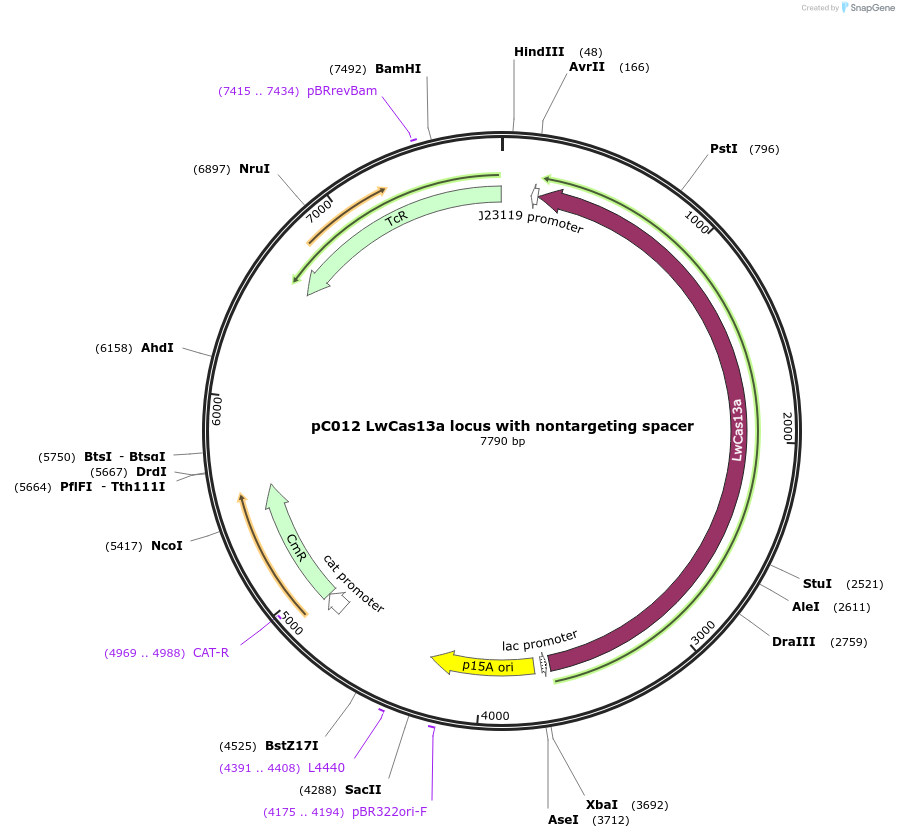
pC012 LwCas13a locus with nontargeting spacer
Plasmid#91904PurposeLwCas13a locus into pACYC184 with nontargeting spacerDepositorInsertLwCas13a with non-targeting spacer
UseCRISPRExpressionBacterialAvailable SinceOct. 16, 2017AvailabilityIndustry, Academic Institutions, and Nonprofits -
pDEST/N1-hEB1-GFP
Plasmid#27382DepositorInsertmicrotubule-associated protein, RP/EB family, member 1 (MAPRE1 Human)
TagsGFPExpressionMammalianAvailable SinceJan. 27, 2011AvailabilityAcademic Institutions and Nonprofits only