We narrowed to 1,801 results for: nif
-
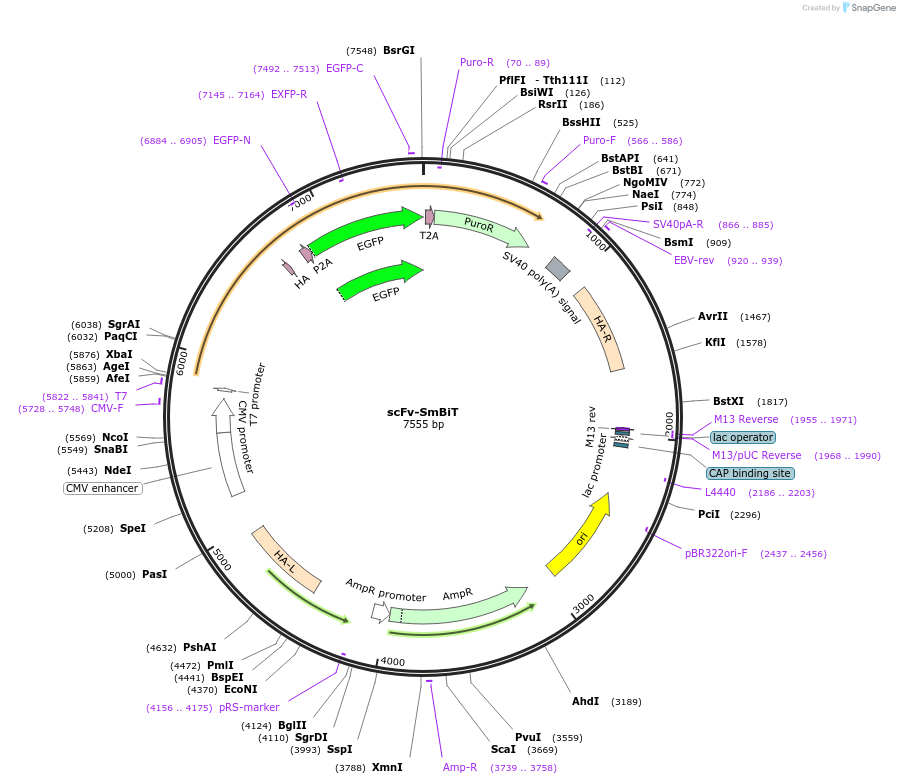
Plasmid#192541PurposeExpresses sender cell components of LOTIISDepositorInsertscFv - SmBiT86
ExpressionMammalianPromoterCMVAvailable SinceDec. 11, 2024AvailabilityAcademic Institutions and Nonprofits only -
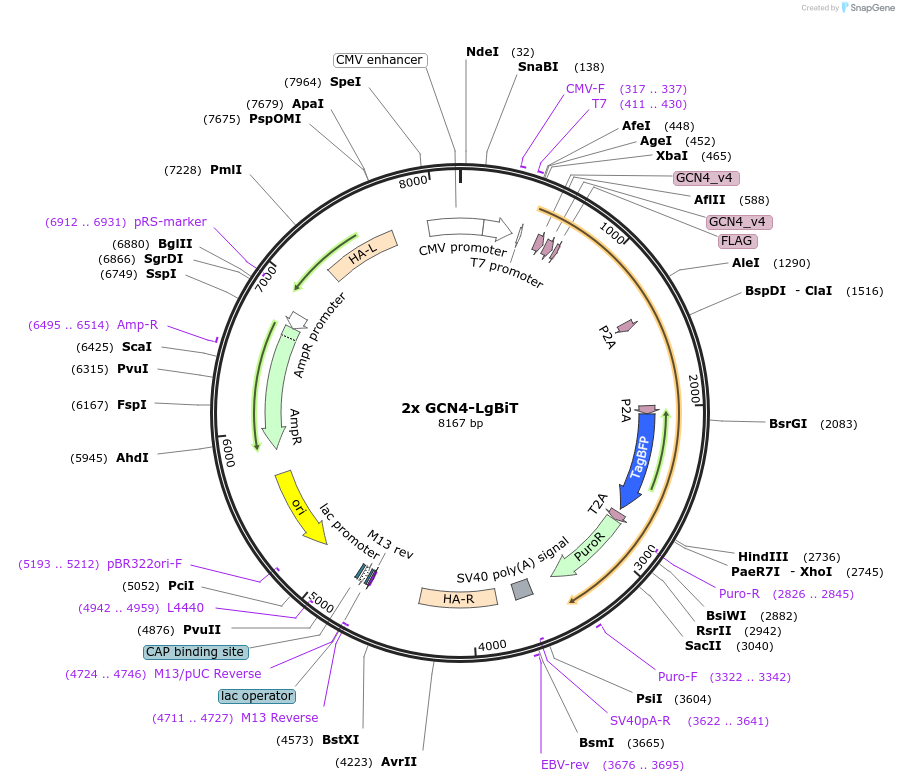
2x GCN4-LgBiT
Plasmid#192542PurposeExpresses receiver cell components of LOTIISDepositorInsert2x GCN4-LgBiT
ExpressionMammalianPromoterCMVAvailable SinceDec. 11, 2024AvailabilityAcademic Institutions and Nonprofits only -
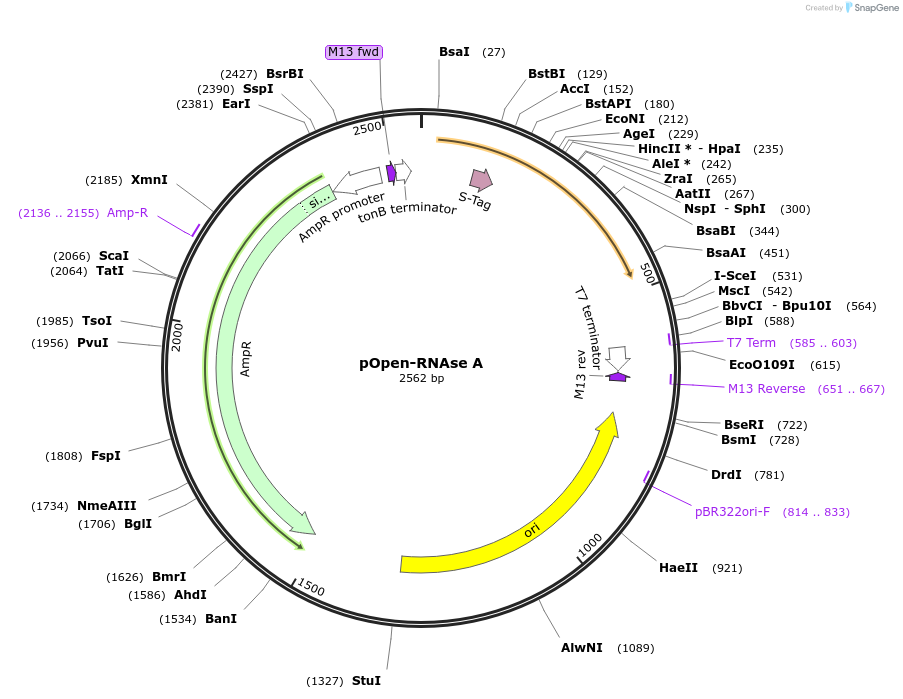
pOpen-RNAse A
Plasmid#165565PurposeRibonuclease A or RNase A; endoribonuclease purified from bovine pancreas. Important enzyme for the removal of RNA for RNA free DNA purification reactions such as plasmid DNA purification and genomic DNA purification, RNA removal from recombinant protein preparations, ribonuclease protection assays, mapping single-base mutations in DNA/RNA.DepositorInsertRNAse A
UseSynthetic BiologyAvailable SinceAug. 17, 2021AvailabilityIndustry, Academic Institutions, and Nonprofits -
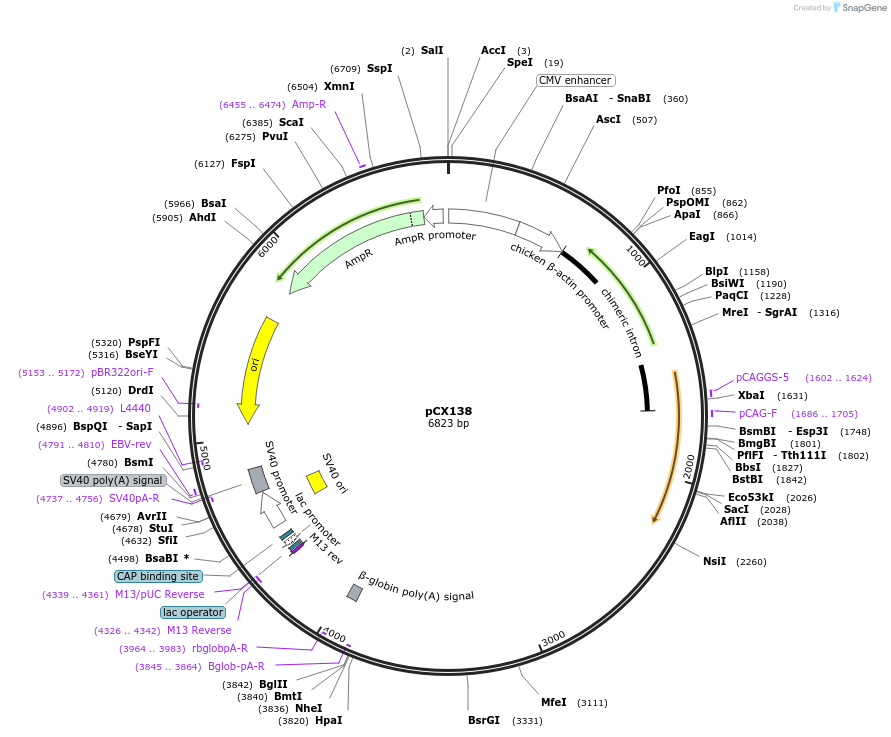
pCX138
Plasmid#229213PurposeCRISPR array plasmid for NOTCH2 (Spacers 25-48)DepositorInsertCRISPR array plasmid for NOTCH2 (Spacers 25-48)
ExpressionMammalianAvailable SinceFeb. 18, 2025AvailabilityAcademic Institutions and Nonprofits only -
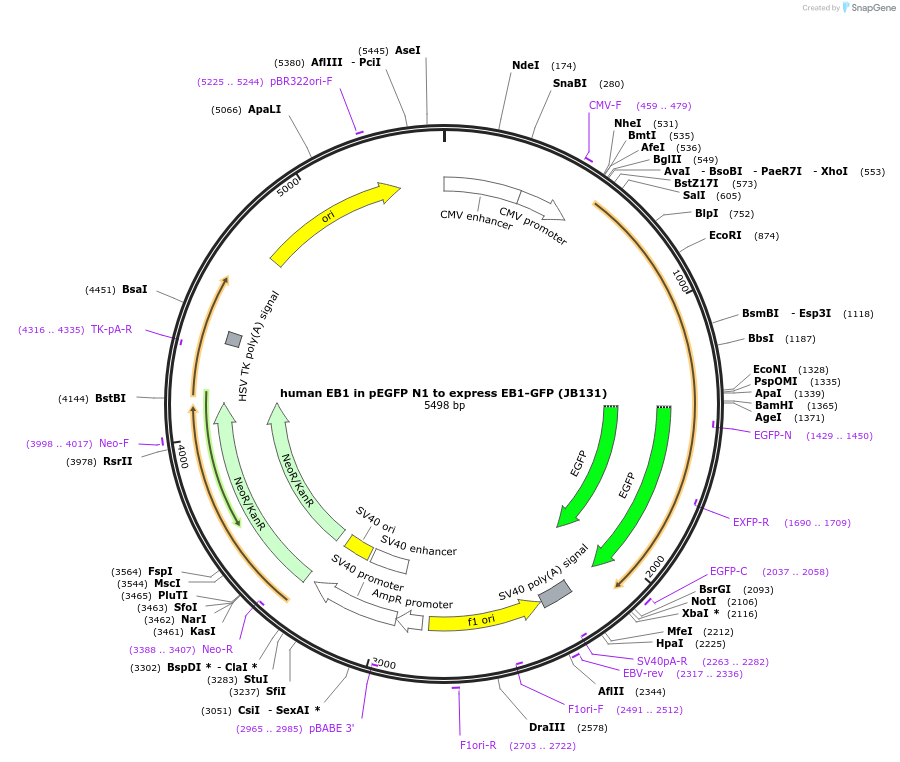
human EB1 in pEGFP N1 to express EB1-GFP (JB131)
Plasmid#39299DepositorAvailable SinceOct. 30, 2012AvailabilityAcademic Institutions and Nonprofits only -
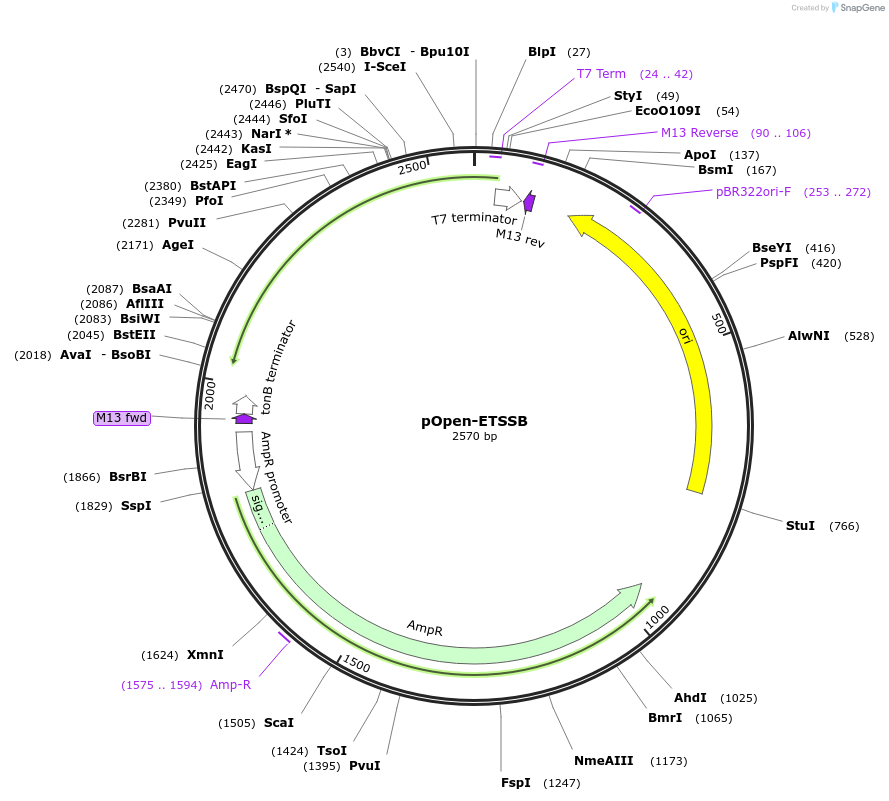
pOpen-ETSSB
Plasmid#165549PurposessDNA binding protein, 9kDa. Features: Improves the processivity of DNA polymerase; involved in stabilization and marking of ssDNA structure; increases the yield and specificitiy of PCR; increases the yield and processivity of RT during RT-PCR; improves DNA sequencing through regions with strong secondary structureDepositorInsertExtreme Thermostable Single-Stranded DNA Binding Protein
UseSynthetic BiologyAvailable SinceAug. 10, 2021AvailabilityIndustry, Academic Institutions, and Nonprofits -
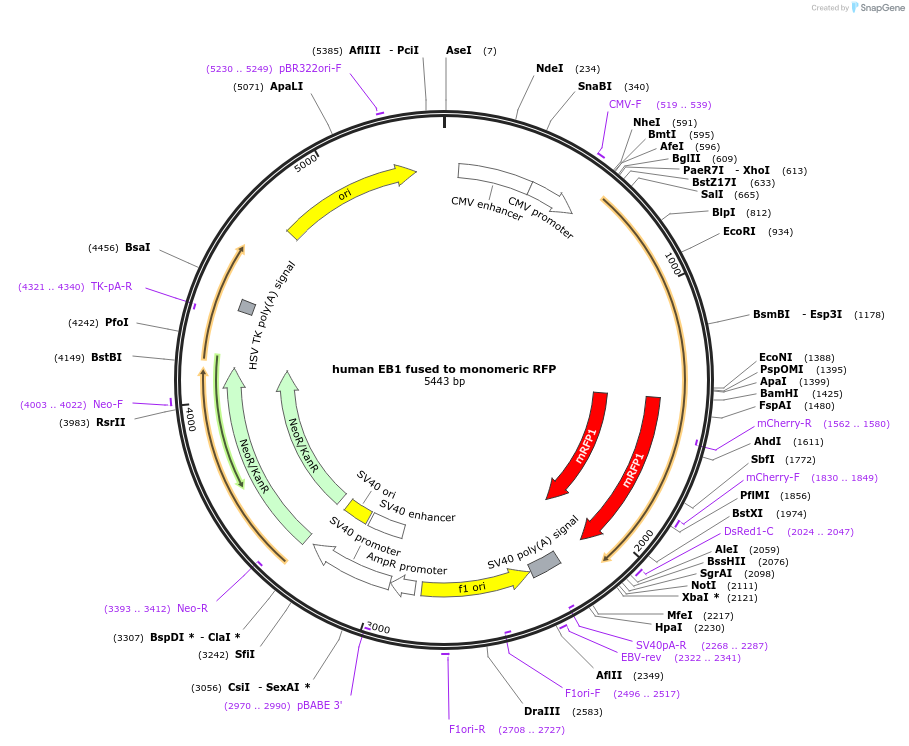
human EB1 fused to monomeric RFP
Plasmid#39323DepositorAvailable SinceOct. 30, 2012AvailabilityAcademic Institutions and Nonprofits only -
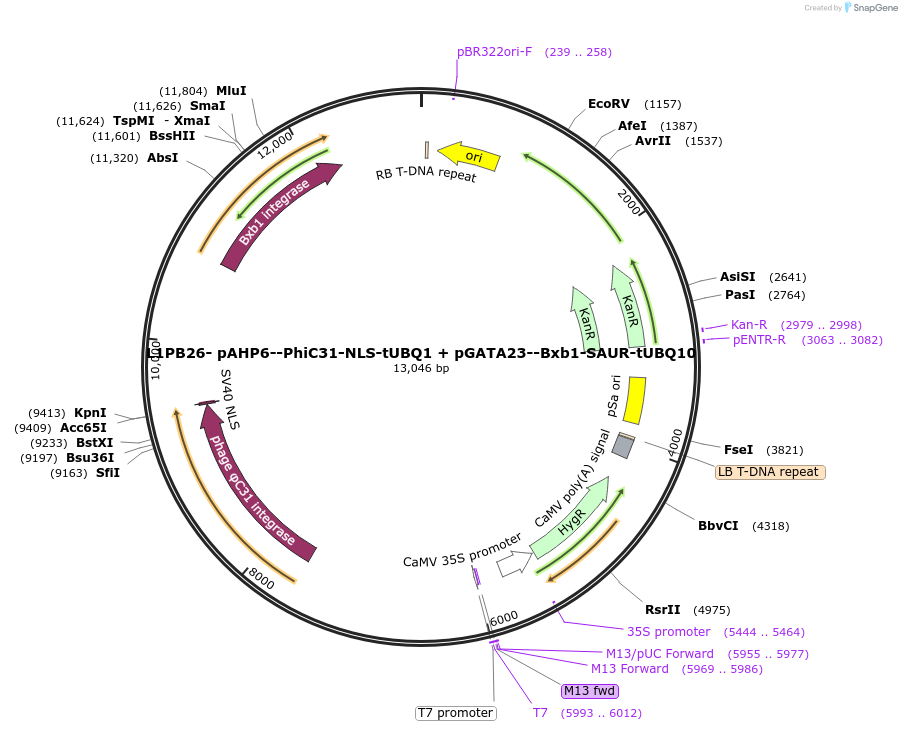
L1PB26- pAHP6--PhiC31-NLS-tUBQ1 + pGATA23--Bxb1-SAUR-tUBQ10
Plasmid#219701PurposeLevel 1 PhiC31 and Bxb1 integrase construct for tracking of lateral root development: pAHP6 promoter for PhiC31 and pGATA23 promoter for Bxb1, NLS tag for PhiC31 and RNA degradation tag for Bxb1DepositorInsertpAHP6::PhiC31-NLS:tUBQ1 + pGATA23::Bxb1-SAUR:tUBQ10
ExpressionPlantAvailable SinceOct. 17, 2024AvailabilityAcademic Institutions and Nonprofits only -
pHMGWA-Pa14Csy4H29A
Plasmid#41092DepositorInsertCsy4
TagsHis6-MBPMutationHis29Ala mmutation to abolish ribonuclease activi…PromoterT7Available SinceDec. 3, 2012AvailabilityAcademic Institutions and Nonprofits only -
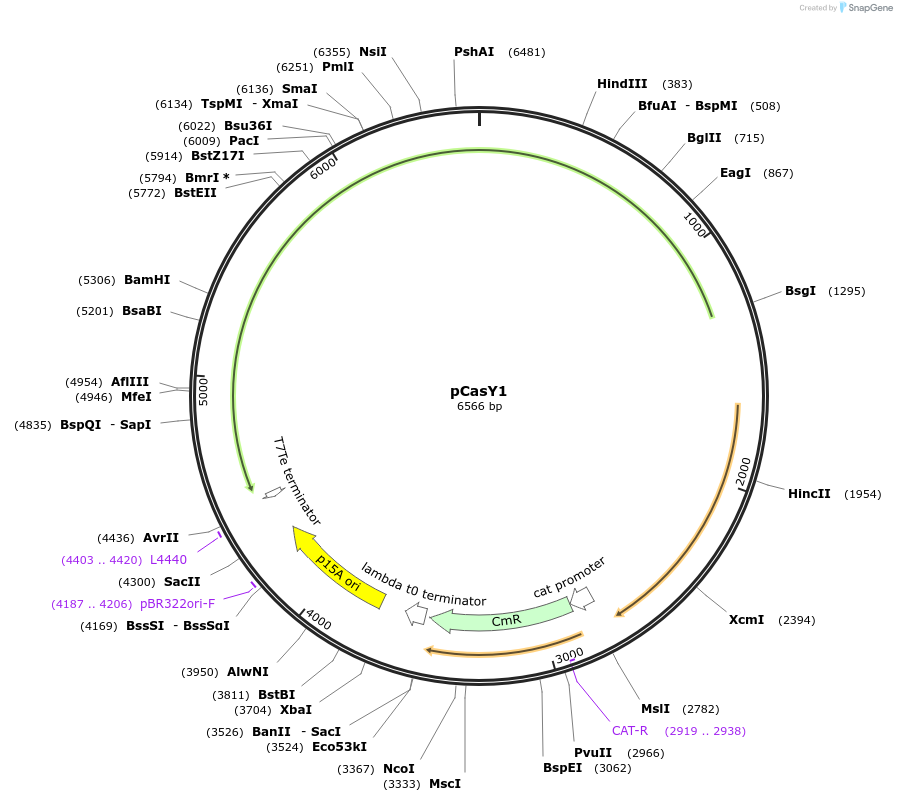
pCasY1
Plasmid#87687Purposep15A plasmid expressing CasY1DepositorInsertCasY1
ExpressionBacterialAvailable SinceApril 27, 2017AvailabilityAcademic Institutions and Nonprofits only -
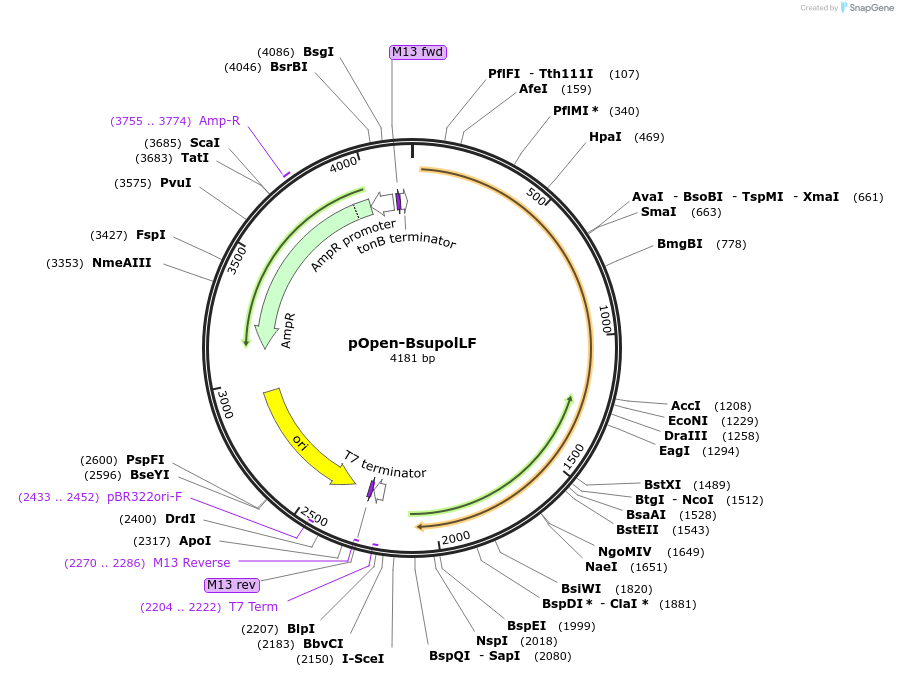
pOpen-BsupolLF
Plasmid#165563PurposeBsu DNA Polymerase I, Large Fragment retains the 5'-3' polymerase activity of the Bacillus subtilis DNA polymerase I, but lacks the 5'-3' exonuclease domain. This large fragment naturally lacks 3'-5' exonuclease activity. Applications include random primer labeling, second strand cDNA synthesis, single dA tailing, and strand displacement DNA synthesis.DepositorInsertBsu DNA Polymerase I, Large Fragment
UseSynthetic BiologyAvailable SinceAug. 17, 2021AvailabilityIndustry, Academic Institutions, and Nonprofits -
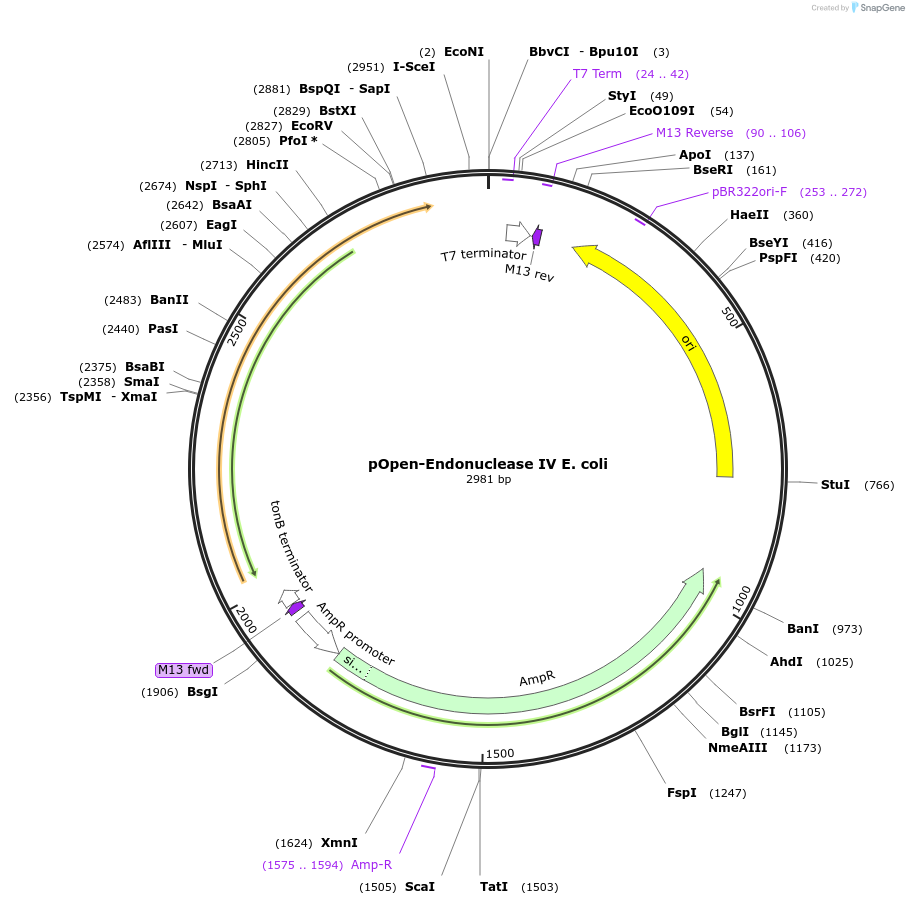
pOpen-Endonuclease IV E. coli
Plasmid#165551PurposeDNA AP endonuclease; Catalyzes the cleavage of DNA phosphodiester backbone at AP sites via hydrolysis leaving a 1 nucleotide gap with 3'-hydroxyl and 5' deoxyribose phosphate (dRP) termini; Also has 3'-diesterase activity which can remove 3' phosphate, 3'-alpha, beta-unsaturated aldehyde, phosphoglycoaldehyde, and other 3' blocking groups.DepositorInsertEndonuclease IV E. coli
UseSynthetic BiologyAvailable SinceAug. 16, 2021AvailabilityIndustry, Academic Institutions, and Nonprofits -
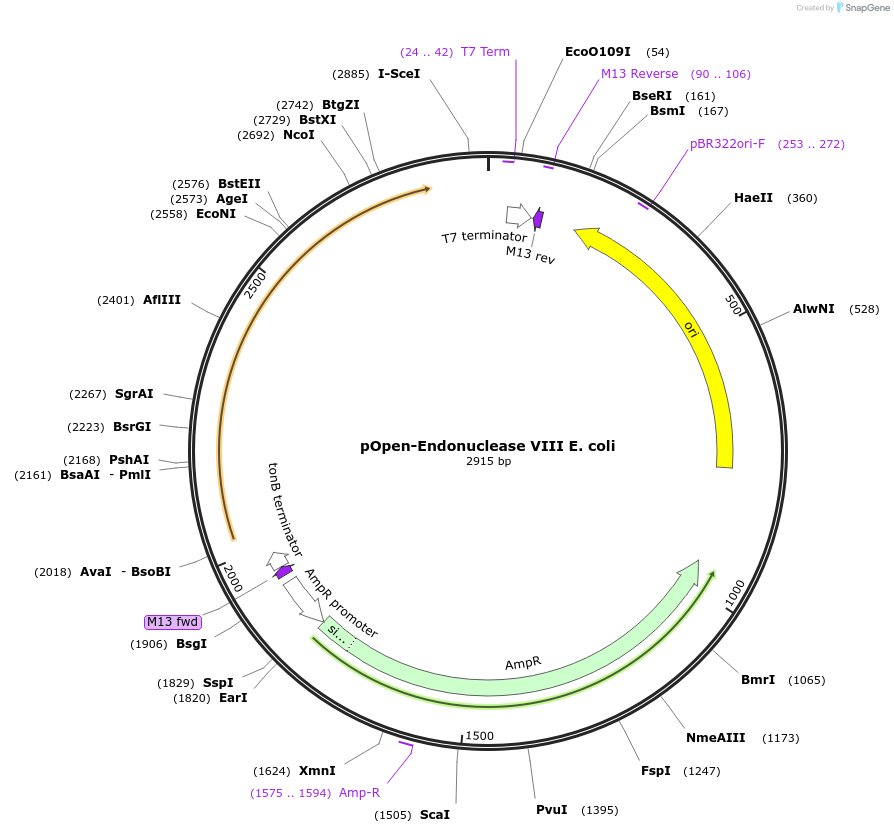
pOpen-Endonuclease VIII E. coli
Plasmid#165561PurposeBifunctional DNA glycosylase with DNA N-glycosylase and AP lyase activities; The N-glycosylase activity releases damaged pyrimidines, including thymine glycol and uracil glycol. The AP lyase activity cleaves DNA phosphodiester backbone at AP sites via beta and delta-elimination, creating a 1 nucleotide DNA gap with 5' and 3' phosphate termini.DepositorInsertEndonuclease VIII E. coli
UseSynthetic BiologyAvailable SinceAug. 16, 2021AvailabilityIndustry, Academic Institutions, and Nonprofits -
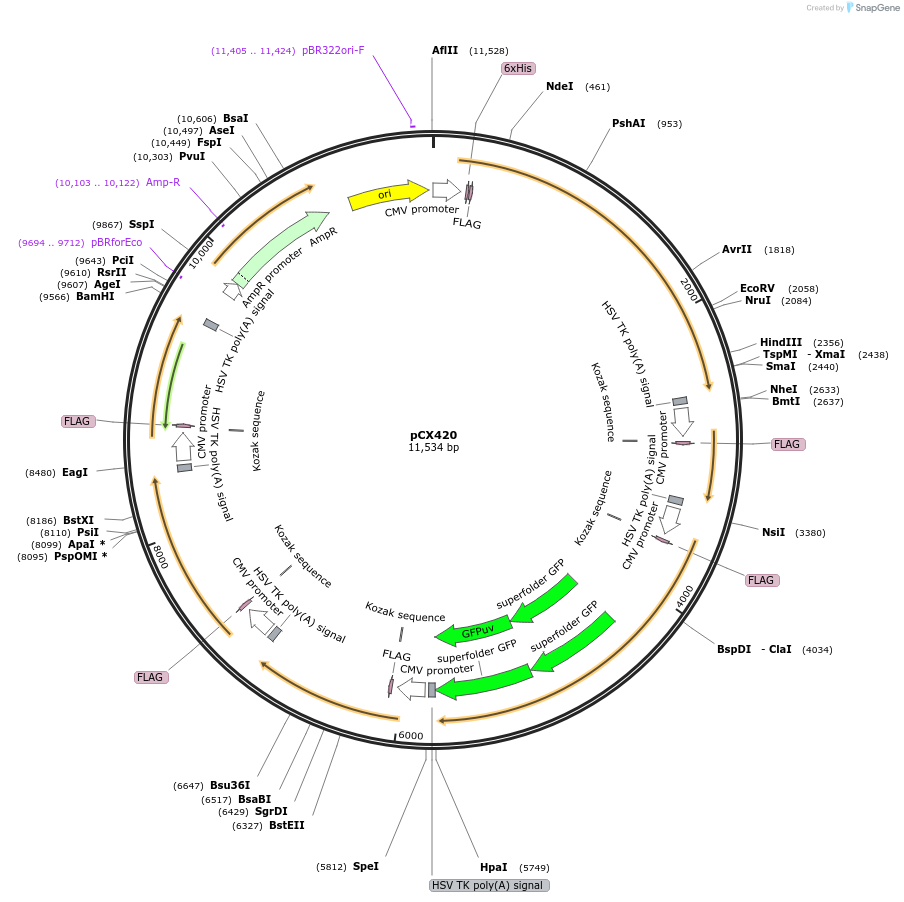
pCX420
Plasmid#229211PurposeCytoplasmic-targeting Csm complex plasmidDepositorInsertCsm1; Csm2; Csm3(RNase mut)-2xsfGFP; Csm4; Csm5; Cas6
ExpressionMammalianAvailable SinceFeb. 18, 2025AvailabilityAcademic Institutions and Nonprofits only -
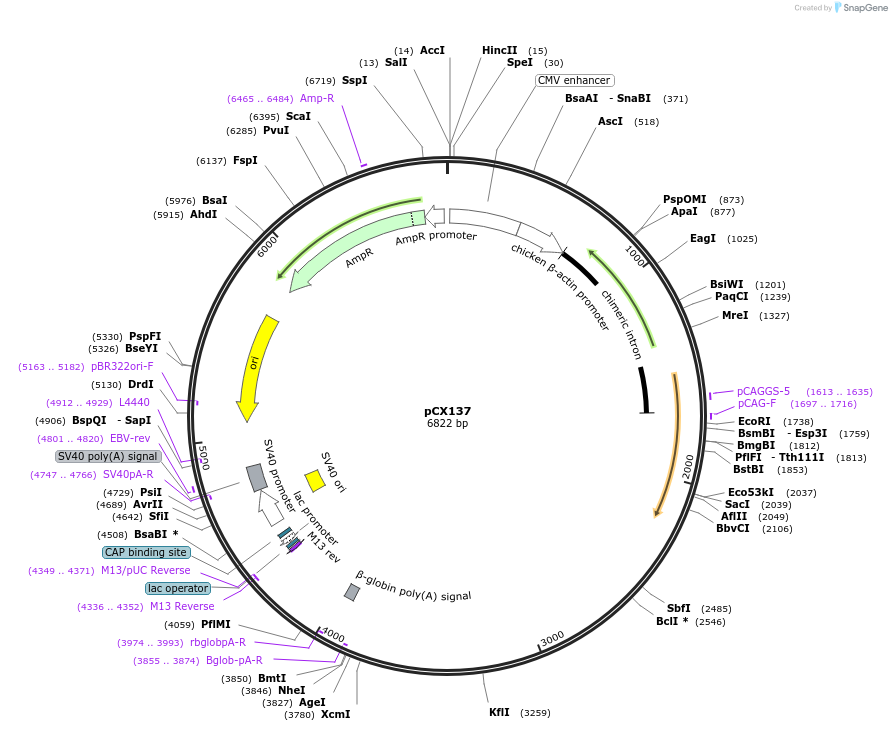
pCX137
Plasmid#229212PurposeCRISPR array plasmid for NOTCH2 (Spacers 1-24)DepositorInsertCRISPR array plasmid for NOTCH2 (Spacers 1-24)
ExpressionMammalianAvailable SinceFeb. 18, 2025AvailabilityAcademic Institutions and Nonprofits only -
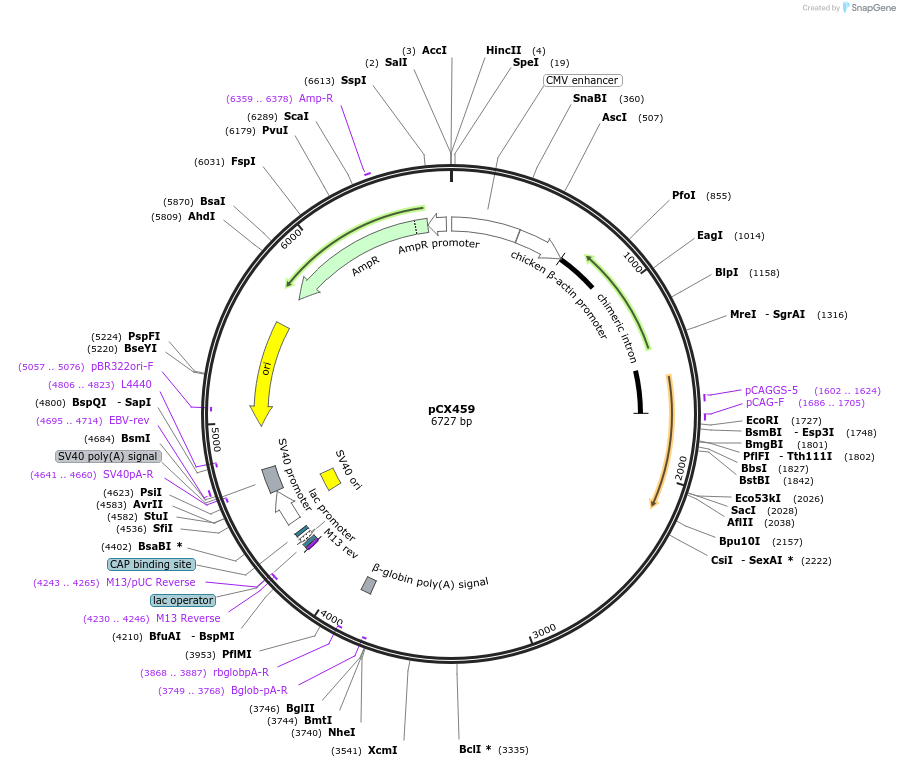
pCX459
Plasmid#229214PurposeCRISPR array plasmid for MAP1B (Spacers 1-24)DepositorInsertCRISPR array plasmid for MAP1B (Spacers 1-24)
ExpressionMammalianAvailable SinceFeb. 18, 2025AvailabilityAcademic Institutions and Nonprofits only -
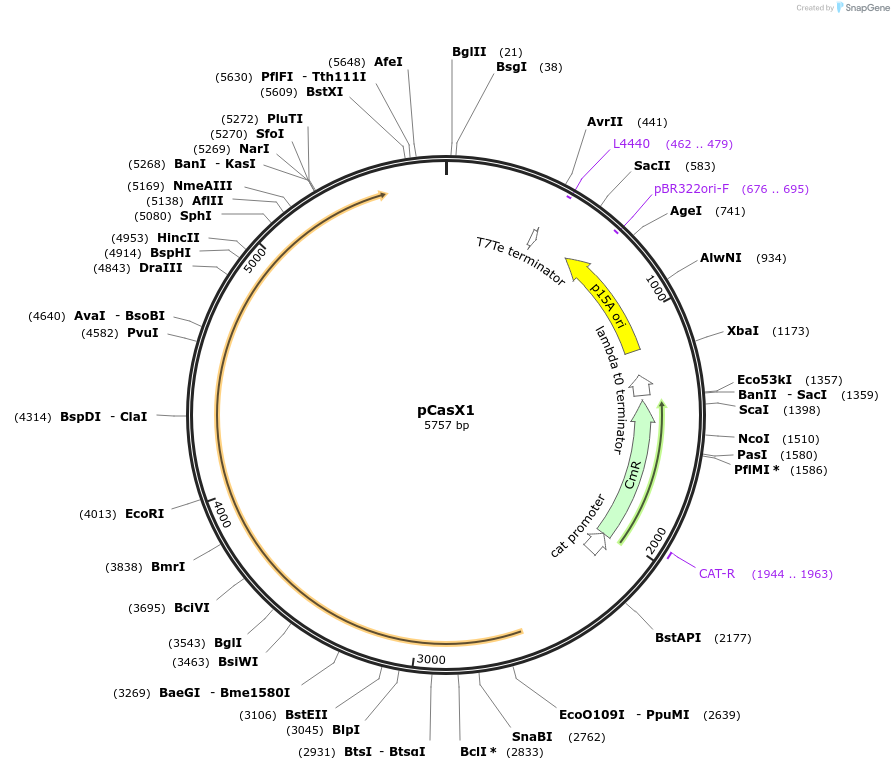
pCasX1
Plasmid#87685Purposep15A plasmid expressing CasX locus from DeltaproteobacteriaDepositorInsertCasX1
ExpressionBacterialPromoterNoneAvailable SinceApril 27, 2017AvailabilityAcademic Institutions and Nonprofits only -
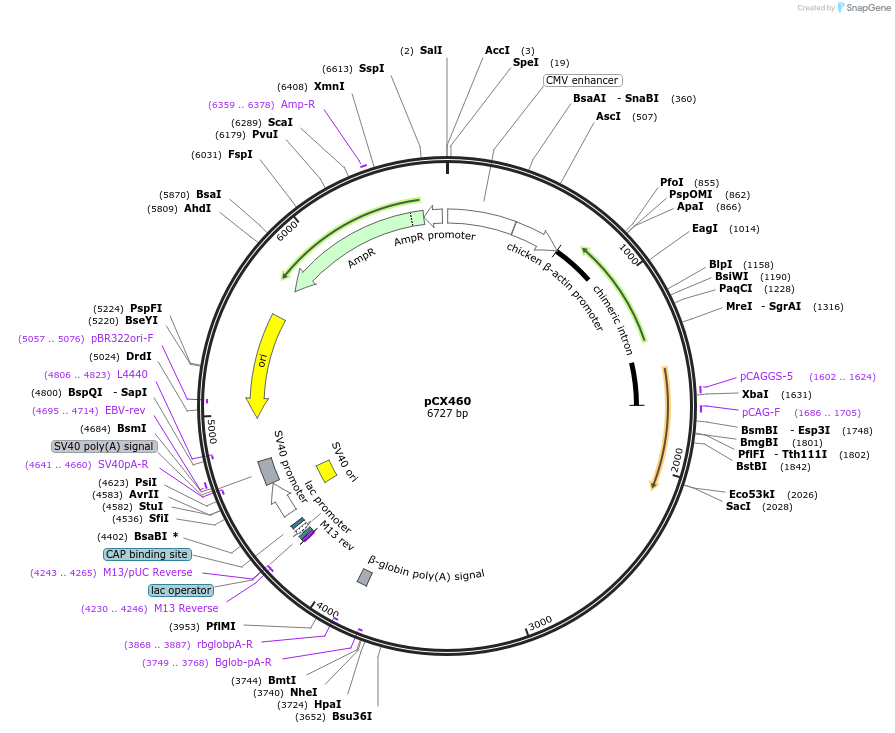
pCX460
Plasmid#229215PurposeCRISPR array plasmid for MAP1B (Spacers 25-48)DepositorInsertCRISPR array plasmid for MAP1B (Spacers 25-48)
ExpressionMammalianAvailable SinceFeb. 18, 2025AvailabilityAcademic Institutions and Nonprofits only -
p2CT-Cdi
Plasmid#83044PurposeExpresses Corynebacterium diphtheriae Cas9 (CdiCas9)DepositorInsertCdiCas9
TagsHis-MBPExpressionBacterialAvailable SinceNov. 1, 2016AvailabilityAcademic Institutions and Nonprofits only -
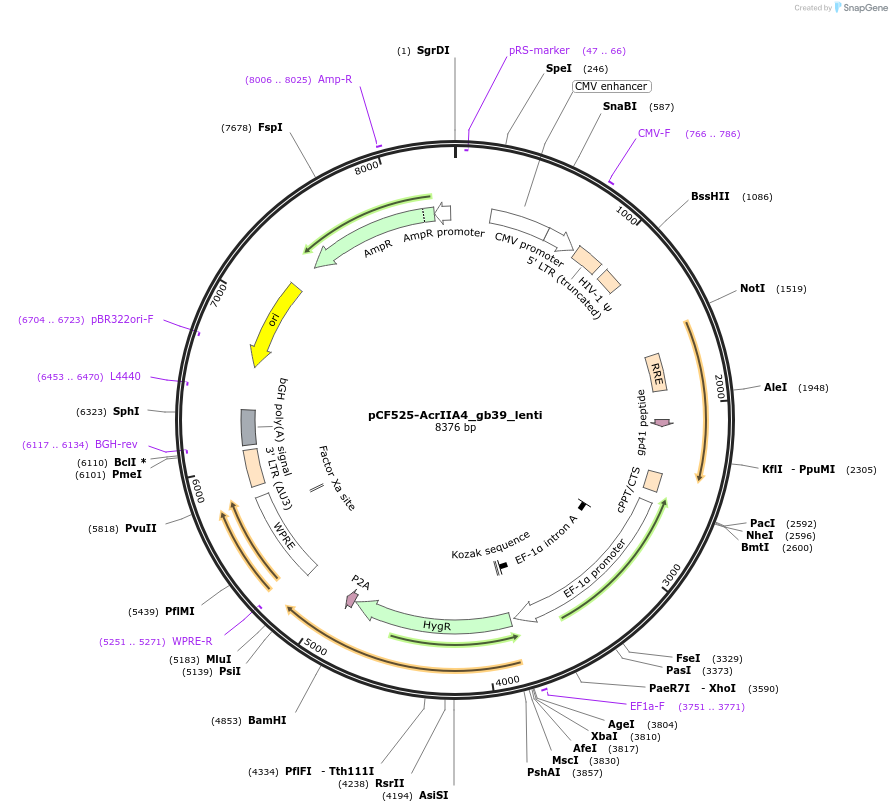
pCF525-AcrIIA4_gb39_lenti
Plasmid#138305PurposeLenti expression of AcrIIA4DepositorInsertAcrIIA4
UseCRISPR and LentiviralExpressionMammalianAvailable SinceMarch 13, 2020AvailabilityAcademic Institutions and Nonprofits only