We narrowed to 53,132 results for: SON
-
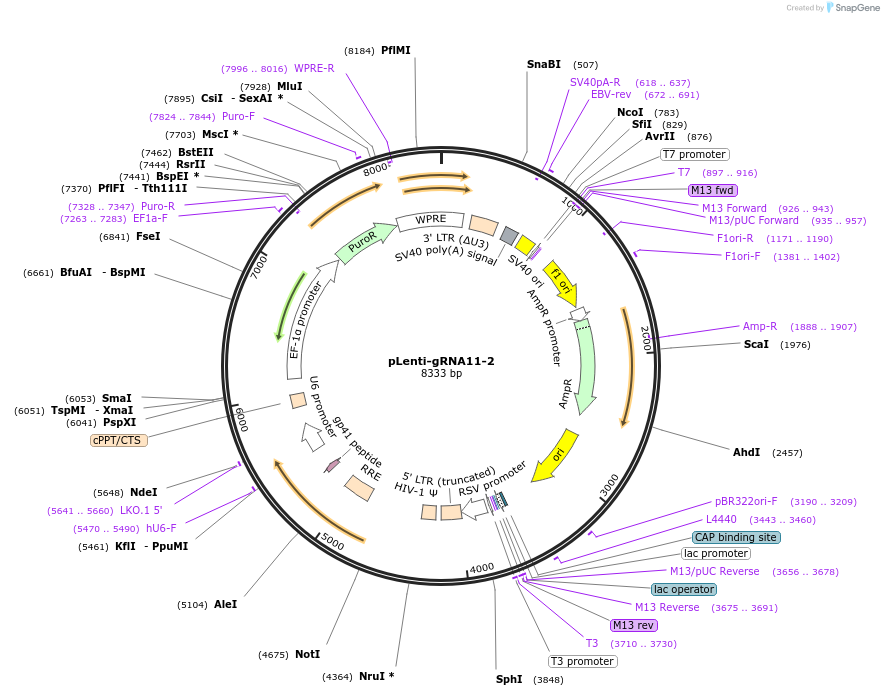
Plasmid#248538PurposeA CRISPR-based technology to study chromosome-specific aneuploidy by targeting human centromeresDepositorInsertgRNA11-2
UseCRISPRExpressionMammalianPromoterU6Available SinceNov. 18, 2025AvailabilityAcademic Institutions and Nonprofits only -
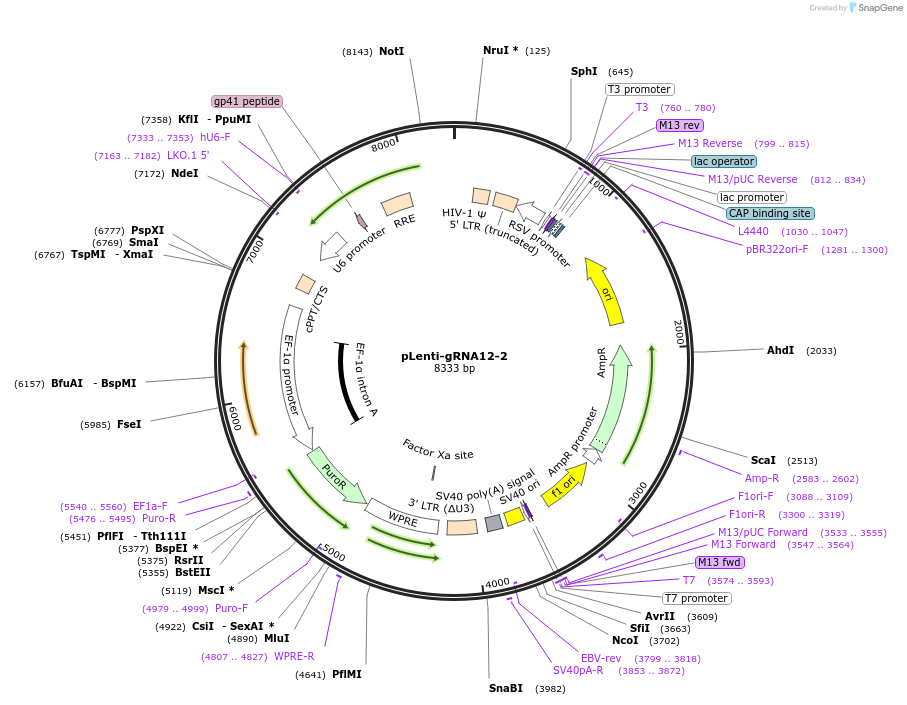
pLenti-gRNA12-2
Plasmid#248539PurposeA CRISPR-based technology to study chromosome-specific aneuploidy by targeting human centromeresDepositorInsertgRNA12-2
UseCRISPRExpressionMammalianPromoterU6Available SinceNov. 18, 2025AvailabilityAcademic Institutions and Nonprofits only -
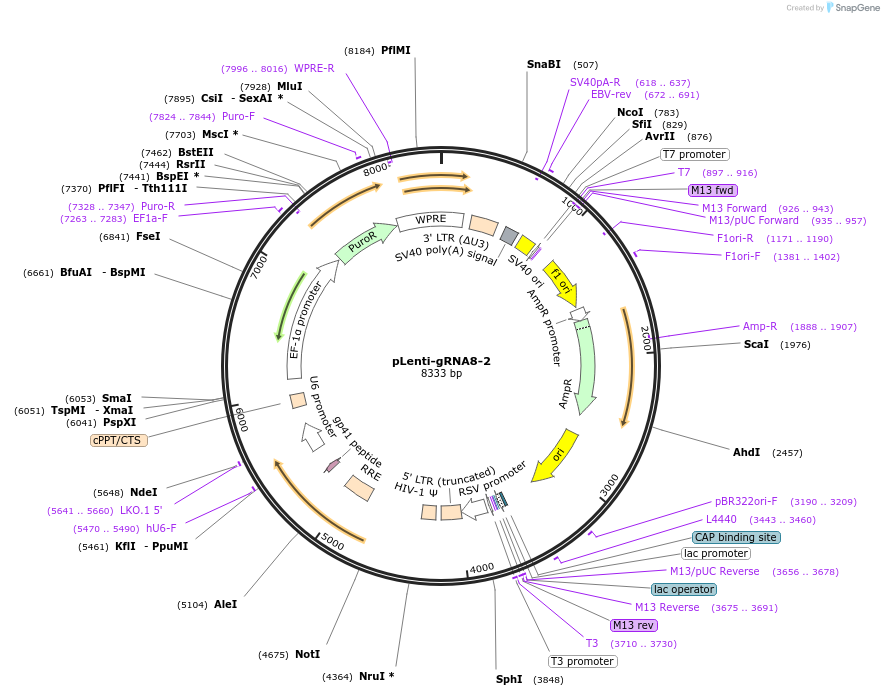
pLenti-gRNA8-2
Plasmid#248532PurposeA CRISPR-based technology to study chromosome-specific aneuploidy by targeting human centromeresDepositorInsertgRNA8-2
UseCRISPRExpressionMammalianPromoterU6Available SinceNov. 18, 2025AvailabilityAcademic Institutions and Nonprofits only -
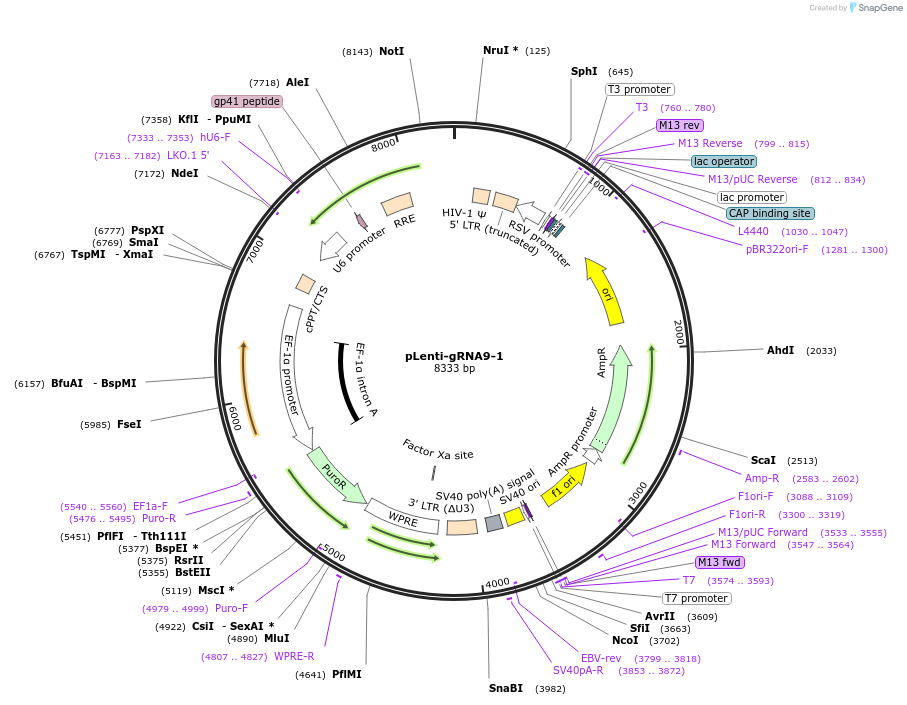
pLenti-gRNA9-1
Plasmid#248533PurposeA CRISPR-based technology to study chromosome-specific aneuploidy by targeting human centromeresDepositorInsertgRNA9-1
UseCRISPRExpressionMammalianPromoterU6Available SinceNov. 18, 2025AvailabilityAcademic Institutions and Nonprofits only -
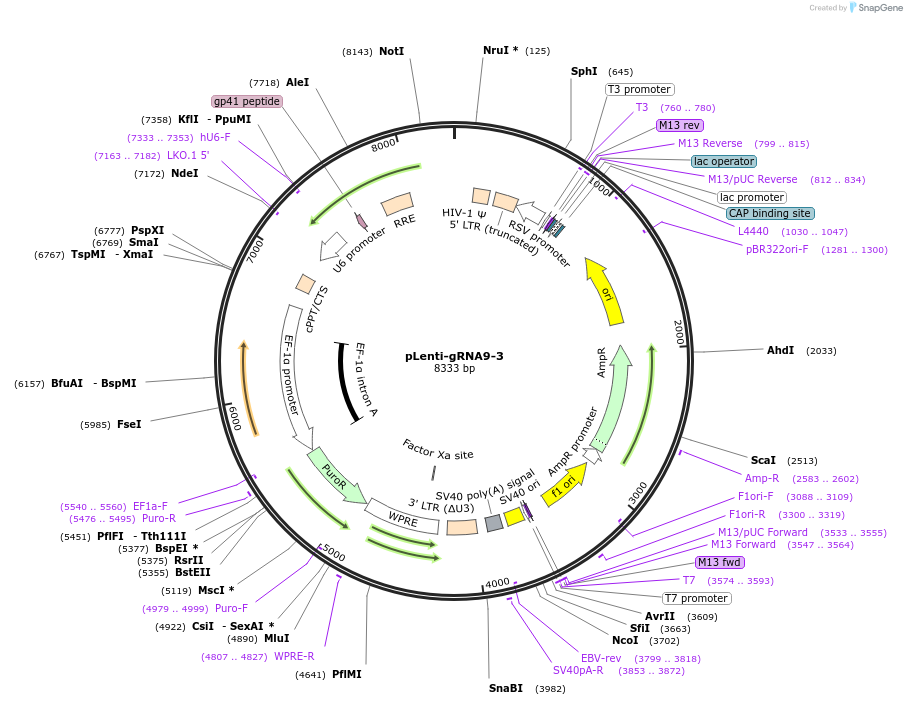
pLenti-gRNA9-3
Plasmid#248534PurposeA CRISPR-based technology to study chromosome-specific aneuploidy by targeting human centromeresDepositorInsertgRNA9-3
UseCRISPRExpressionMammalianPromoterU6Available SinceNov. 18, 2025AvailabilityAcademic Institutions and Nonprofits only -
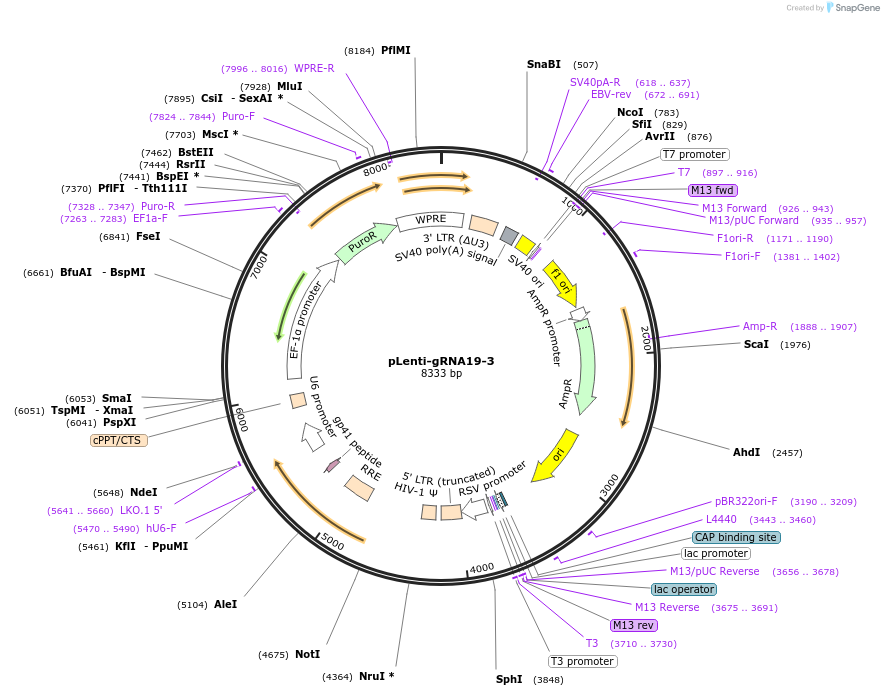
pLenti-gRNA19-3
Plasmid#248547PurposeA CRISPR-based technology to study chromosome-specific aneuploidy by targeting human centromeresDepositorInsertgRNA19-3
UseCRISPRExpressionMammalianPromoterU6Available SinceNov. 18, 2025AvailabilityAcademic Institutions and Nonprofits only -
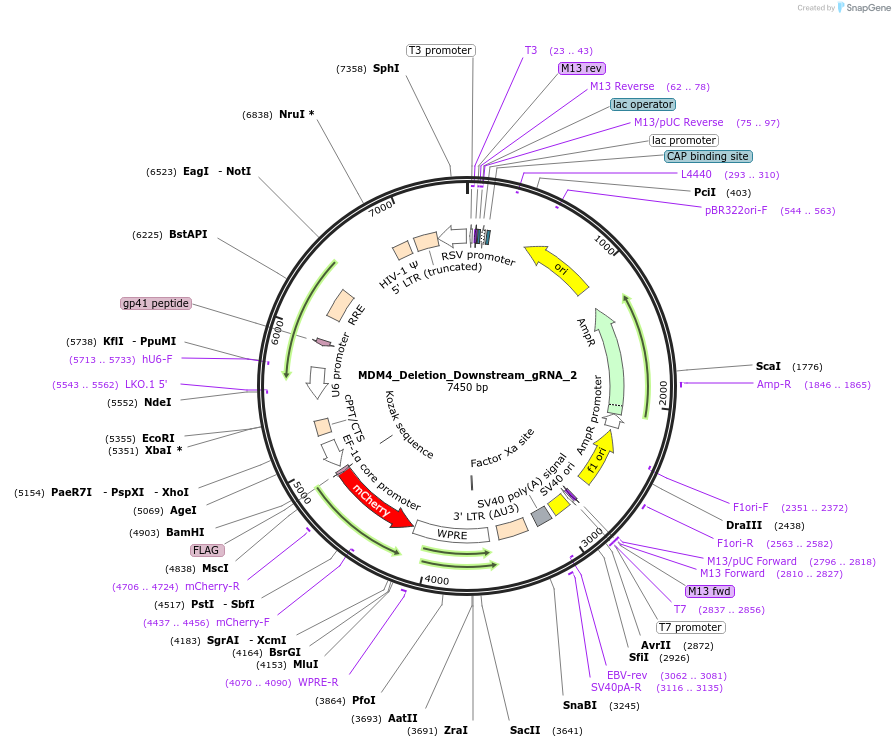
MDM4_Deletion_Downstream_gRNA_2
Plasmid#195137PurposegRNA in a third generation gRNA backbone with mCherry, targeting region immediately downstream of MDM4, to be used with MDM4_Deletion_Upstream_gRNA_1/2 for MDM4 deletionDepositorInsertMDM4 Deletion Downstream gRNA 2 (MDM4 Human)
ExpressionMammalianAvailable SinceFeb. 6, 2023AvailabilityAcademic Institutions and Nonprofits only -
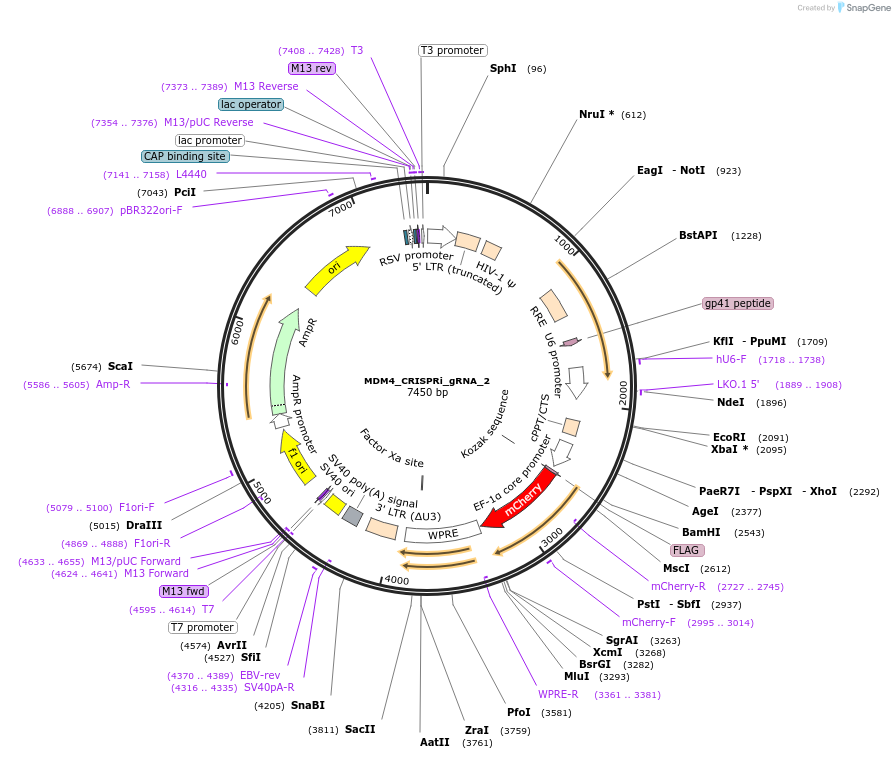
MDM4_CRISPRi_gRNA_2
Plasmid#195133PurposegRNA from the Weissman CRISPRi library targeting MDM4, cloned into a third generation gRNA backbone with mCherryDepositorInsertMDM4 CRISPRi gRNA 2 (MDM4 Human)
ExpressionMammalianAvailable SinceFeb. 3, 2023AvailabilityAcademic Institutions and Nonprofits only -
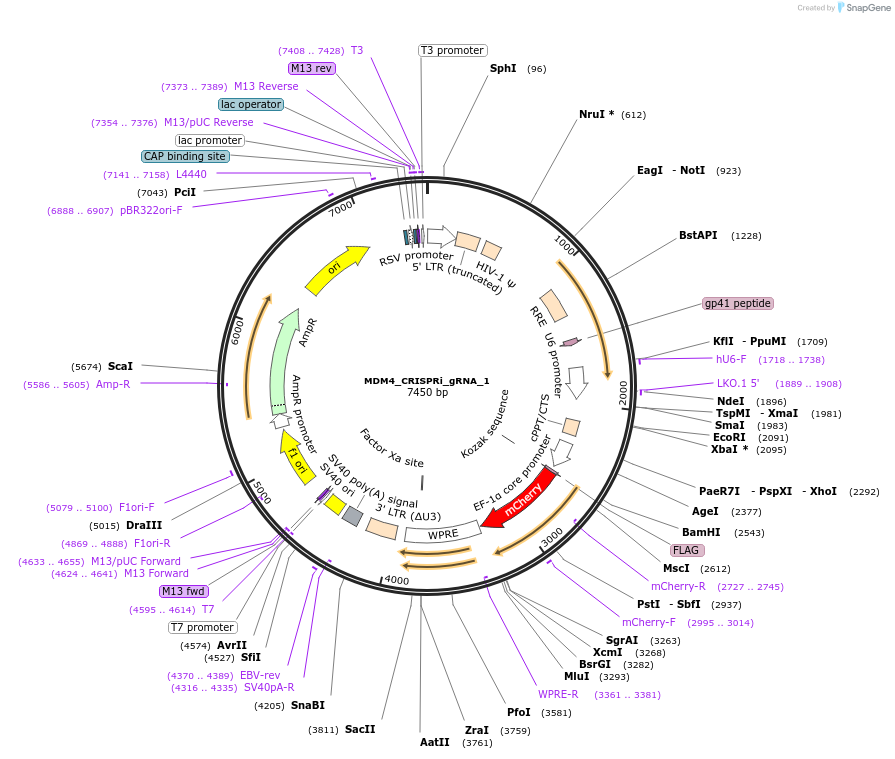
MDM4_CRISPRi_gRNA_1
Plasmid#195132PurposegRNA from the Weissman CRISPRi library targeting MDM4, cloned into a third generation gRNA backbone with mCherryDepositorInsertMDM4 CRISPRi gRNA 1 (MDM4 Human)
ExpressionMammalianAvailable SinceFeb. 3, 2023AvailabilityAcademic Institutions and Nonprofits only -
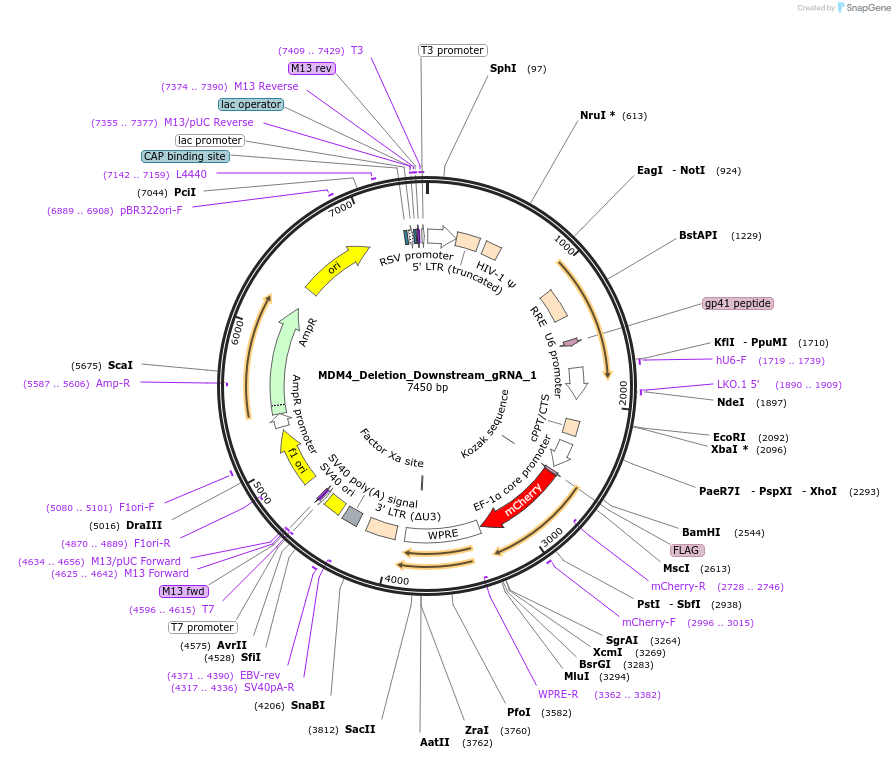
MDM4_Deletion_Downstream_gRNA_1
Plasmid#195136PurposegRNA in a third generation gRNA backbone with mCherry, targeting region immediately downstream of MDM4, to be used with MDM4_Deletion_Upstream_gRNA_1/2 for MDM4 deletionDepositorInsertMDM4 Deletion Downstream gRNA 1 (MDM4 Human)
ExpressionMammalianAvailable SinceFeb. 3, 2023AvailabilityAcademic Institutions and Nonprofits only -
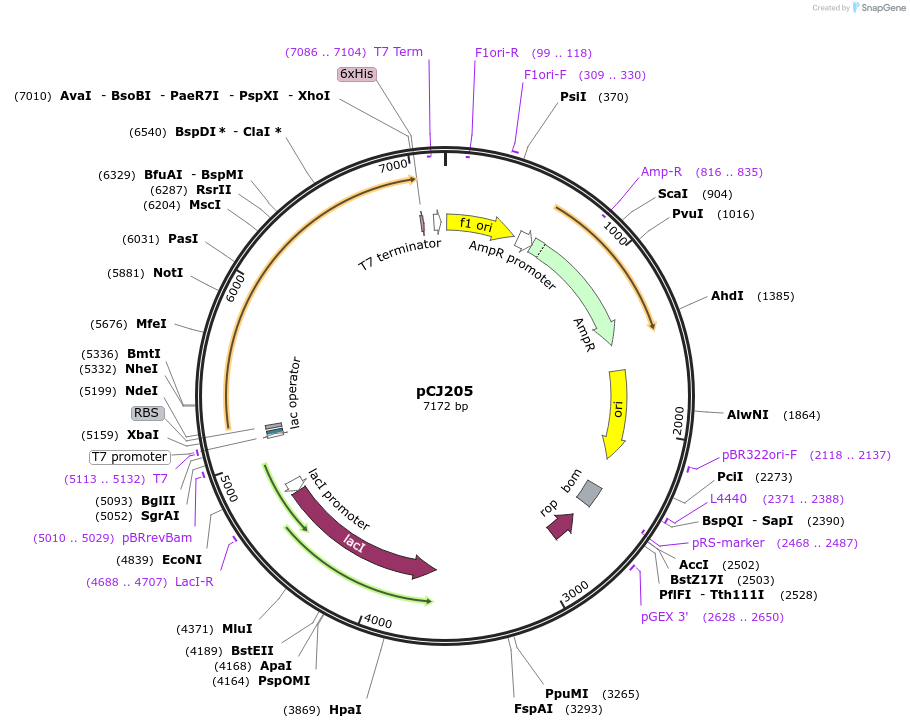
pCJ205
Plasmid#162676PurposepET-21b(+) based plasmid for expression of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating C224A anDepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1)
TagsHisExpressionBacterialMutationC224A, C529A; codon optimized for expression in E…PromoterT7/lacAvailable SinceFeb. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
pCJ198
Plasmid#162671PurposepET-21b(+) based plasmid for expression of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating F495I mutation.DepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1)
TagsHisExpressionBacterialMutationF495I; codon optimized for expression in E. coli …PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
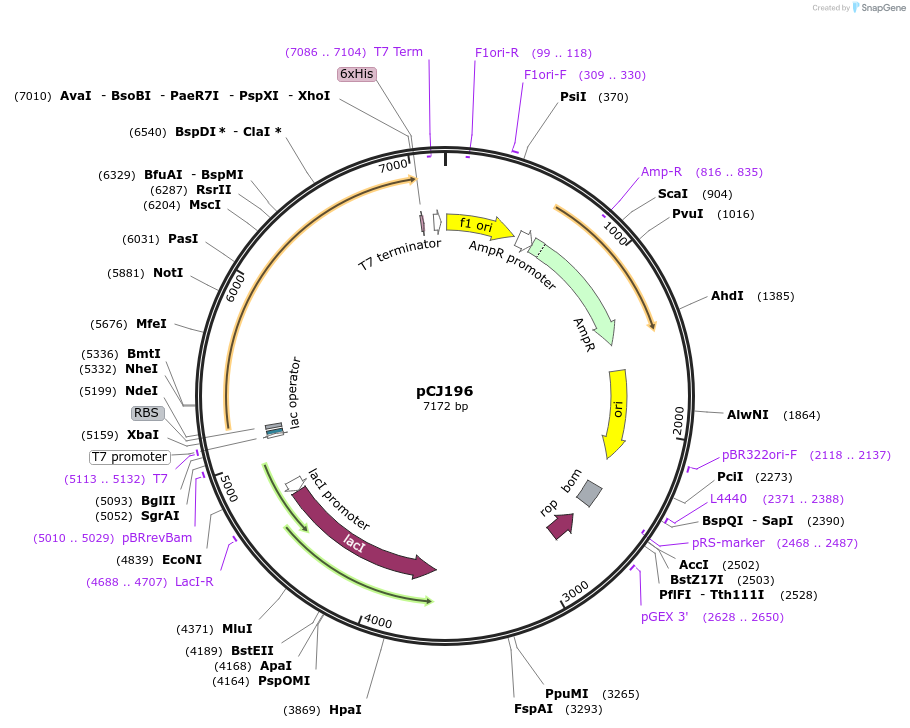
pCJ196
Plasmid#162669PurposepET-21b(+) based plasmid for expression of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating catalytic mutation, S225ADepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1)
TagsHisExpressionBacterialMutationS225A; codon optimized for expression in E. coli …PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
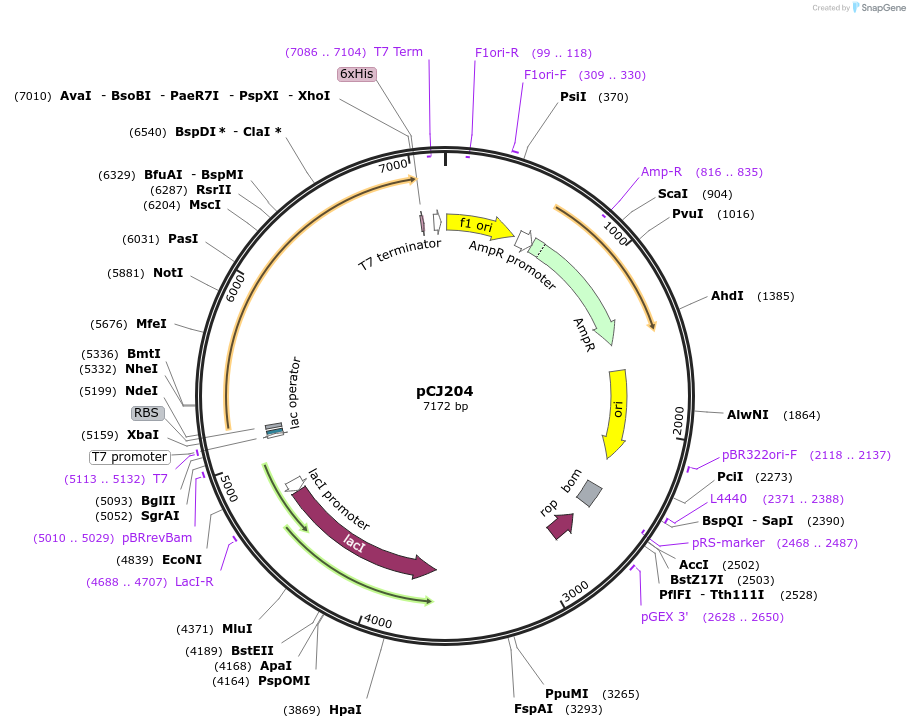
pCJ204
Plasmid#162677PurposepET-21b(+) based plasmid for expression of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating C224H and C529F mutations.DepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1)
TagsHisExpressionBacterialMutationC224H, C529F; codon optimized for expression in E…PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
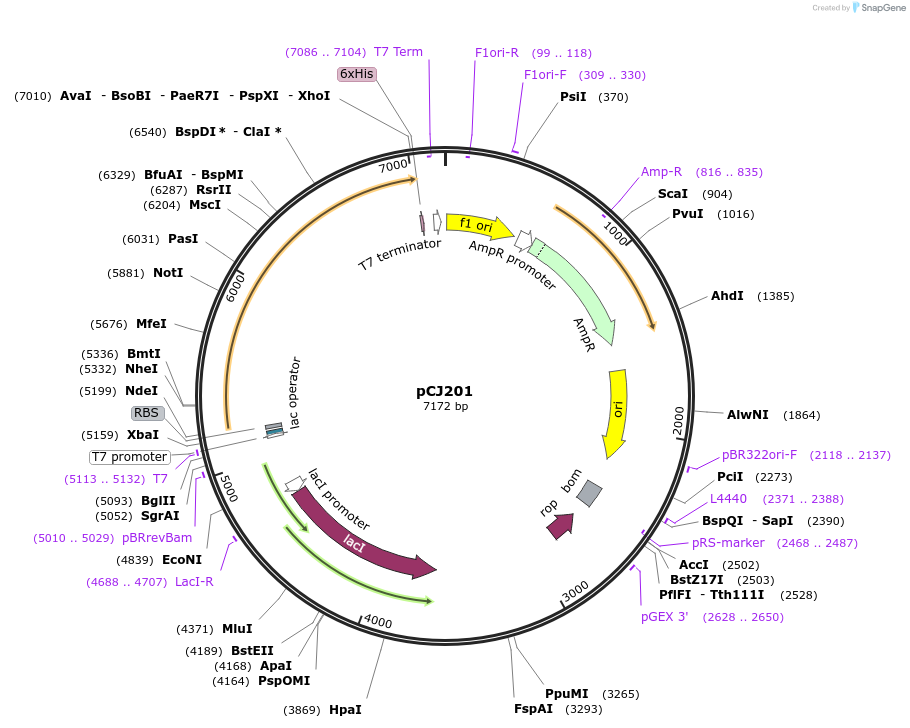
pCJ201
Plasmid#162674PurposepET-21b(+) based plasmid for expression of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating C224W and C529SS mutations.DepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1)
TagsHisExpressionBacterialMutationC224W, C529SS; codon optimized for expression in …PromoterT7/lacAvailable SinceJan. 6, 2021AvailabilityAcademic Institutions and Nonprofits only -
mKusabira Orange-N1
Plasmid#55046PurposeEmpty backbone for fusing gene of interest to N-term of mKODepositorTypeEmpty backboneTagsmKOExpressionMammalianPromoterCMVAvailable SinceSept. 17, 2014AvailabilityAcademic Institutions and Nonprofits only -
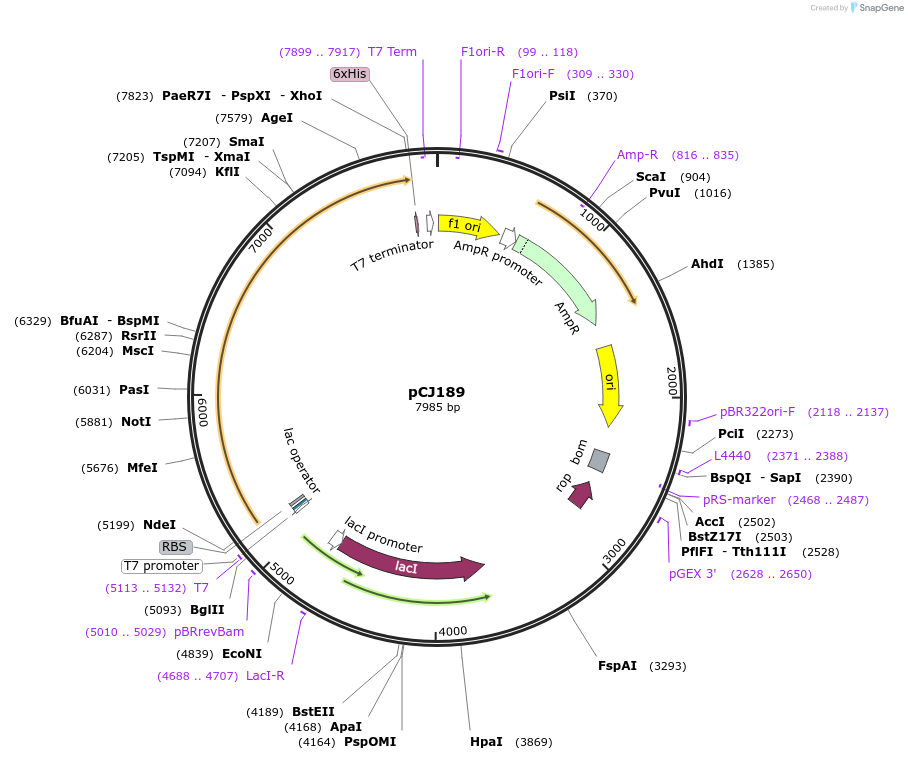
pCJ189
Plasmid#162666PurposepET-21b(+) based plasmid for expresion of MHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by an 8 residue Gly/Ser linker; codon optimized for expression in E. coli K12, with C-terminal His tag.DepositorInsertMHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by an 8 residue Gly/Ser linker
TagsHisExpressionBacterialMutationcodon optimized for expression in E. coli K12PromoterT7/lacAvailable SinceFeb. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
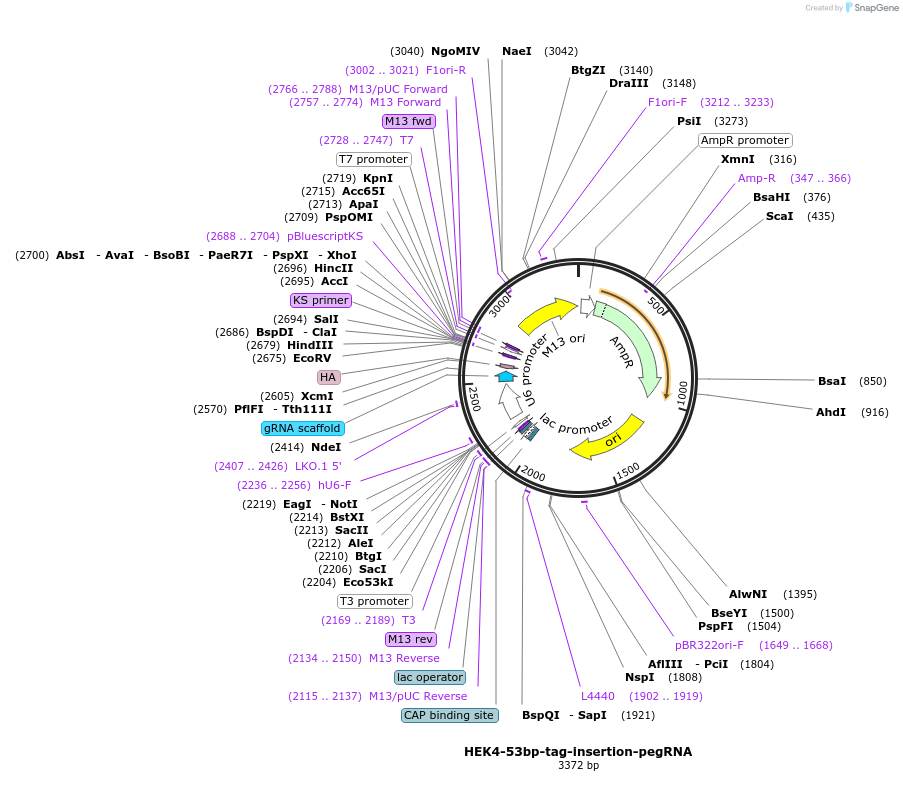
HEK4-53bp-tag-insertion-pegRNA
Plasmid#227673PurposePegRNA for 53 bp insertionDepositorAvailable SinceOct. 22, 2024AvailabilityAcademic Institutions and Nonprofits only -
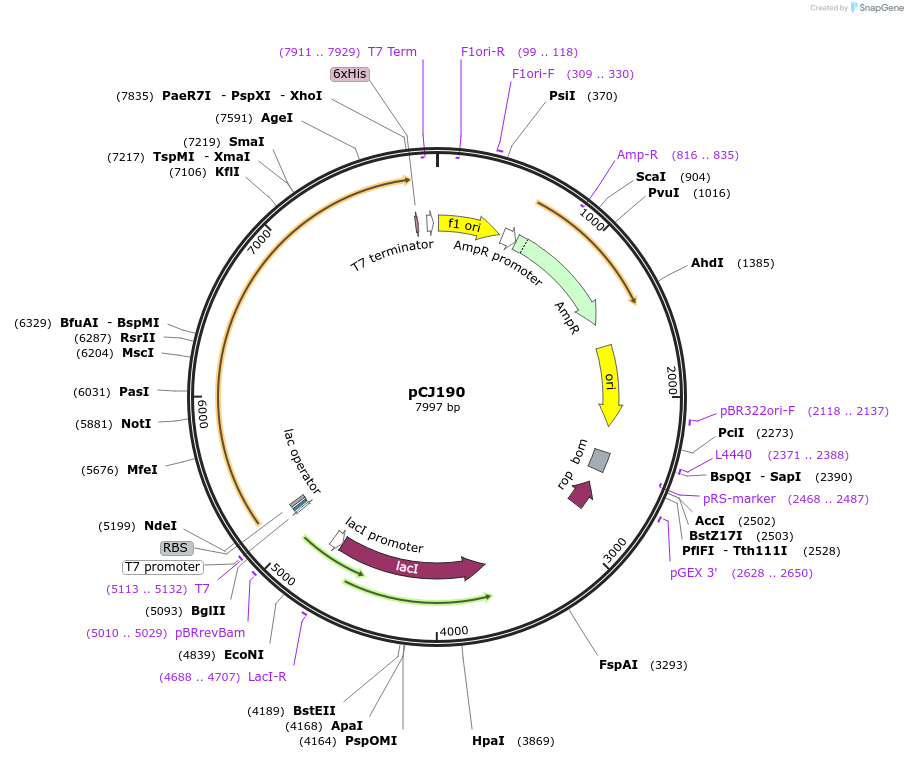
pCJ190
Plasmid#162667PurposepET-21b(+) based plasmid for expresion of MHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by a 12 residue Gly/Ser linker; codon optimized for expression in E. coli K12, with C-terminal His tag.DepositorInsertMHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by a 12 residue Gly/Ser linker
TagsHisExpressionBacterialMutationcodon optimized for expression in E. coli K12PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
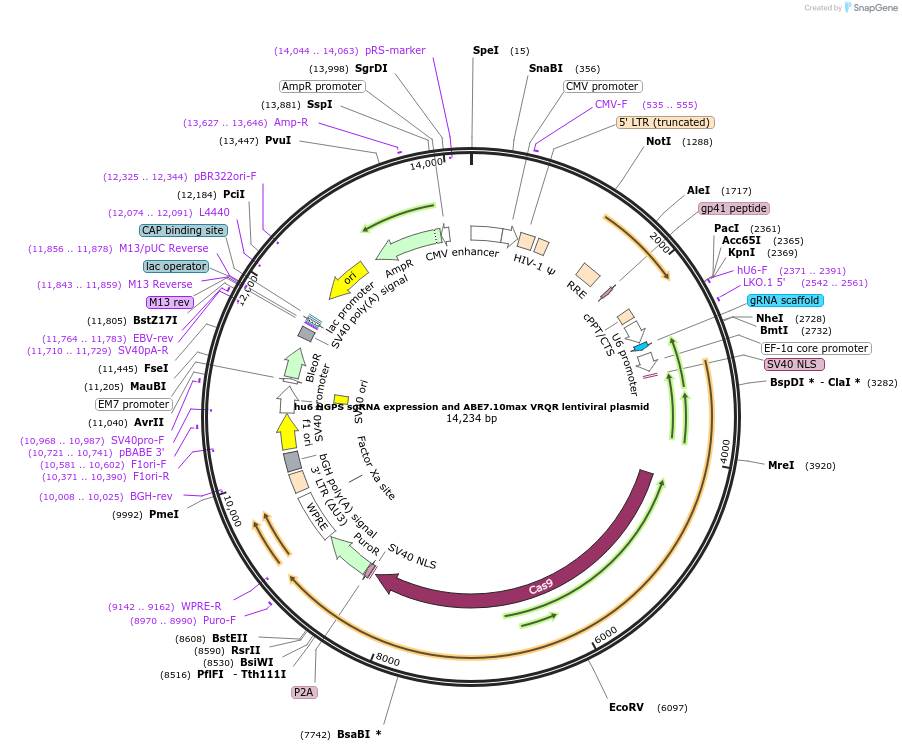
hu6 HGPS sgRNA expression and ABE7.10max VRQR lentiviral plasmid
Plasmid#154429Purposehu6 HGPS sgRNA expression and ABE7.10max VRQR lentiviral plasmidDepositorInserthu6 HGPS sgRNA expression and ABE7.10max VRQR lentiviral plasmid
UseCRISPR and LentiviralExpressionMammalianMutationVRQR point mutations in SpCas9Available SinceFeb. 19, 2021AvailabilityAcademic Institutions and Nonprofits only