We narrowed to 5,577 results for: dre
-
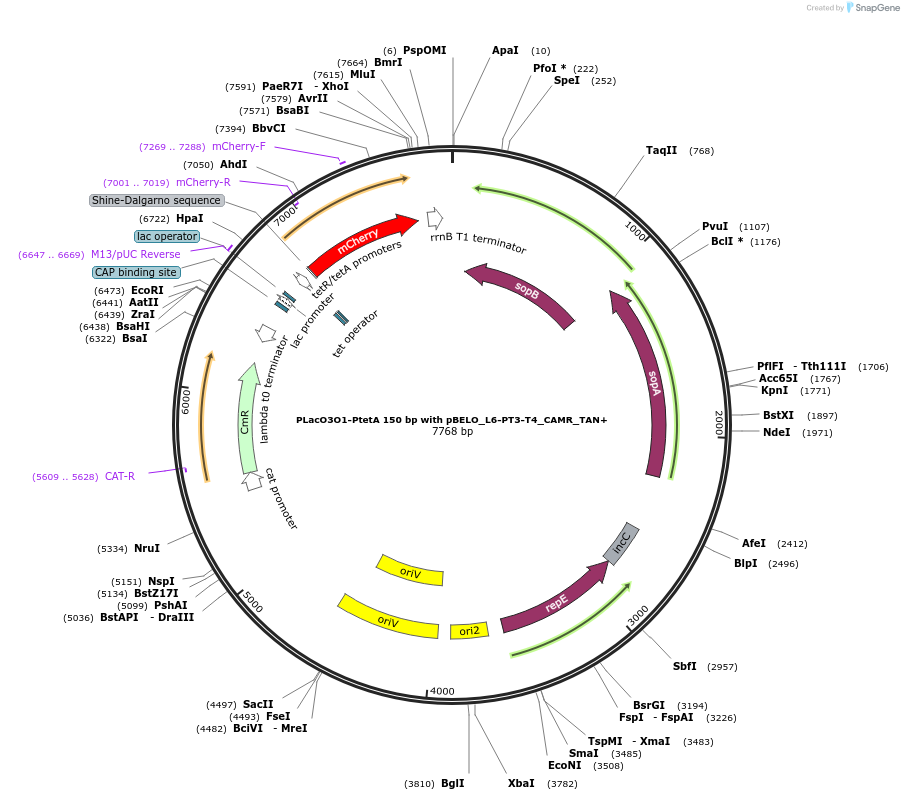
Plasmid#225647PurposeLacO3O1 and tetA promoters in tandem conformation with 150bp distance between them, expressing mCherry with pBELO_L6-PT3-T4 backbone and chloramphenicol resistanceDepositorInsertLacO3O1 and tetR/tetA in tandem conformation
TagsmCherryExpressionBacterialPromoterLacO3O1 and tetA promotersAvailable SinceOct. 11, 2024AvailabilityAcademic Institutions and Nonprofits only -
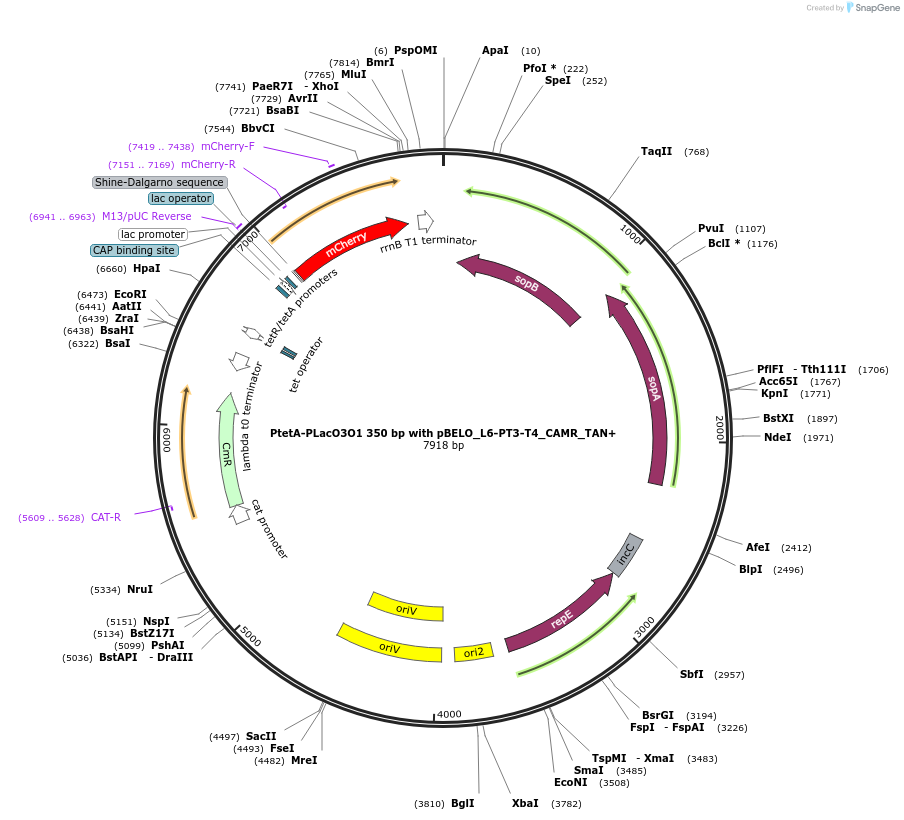
PtetA-PLacO3O1 350 bp with pBELO_L6-PT3-T4_CAMR_TAN+
Plasmid#225646PurposetetA and LacO3O1 promoters in tandem conformation with 350bp distance between them, expressing mCherry with pBELO_L6-PT3-T4 backbone and chloramphenicol resistanceDepositorInserttetR/tetA and LacO3O1 in tandem conformation
TagsmCherryExpressionBacterialPromotertetA and LacO3O1 promotersAvailable SinceOct. 11, 2024AvailabilityAcademic Institutions and Nonprofits only -
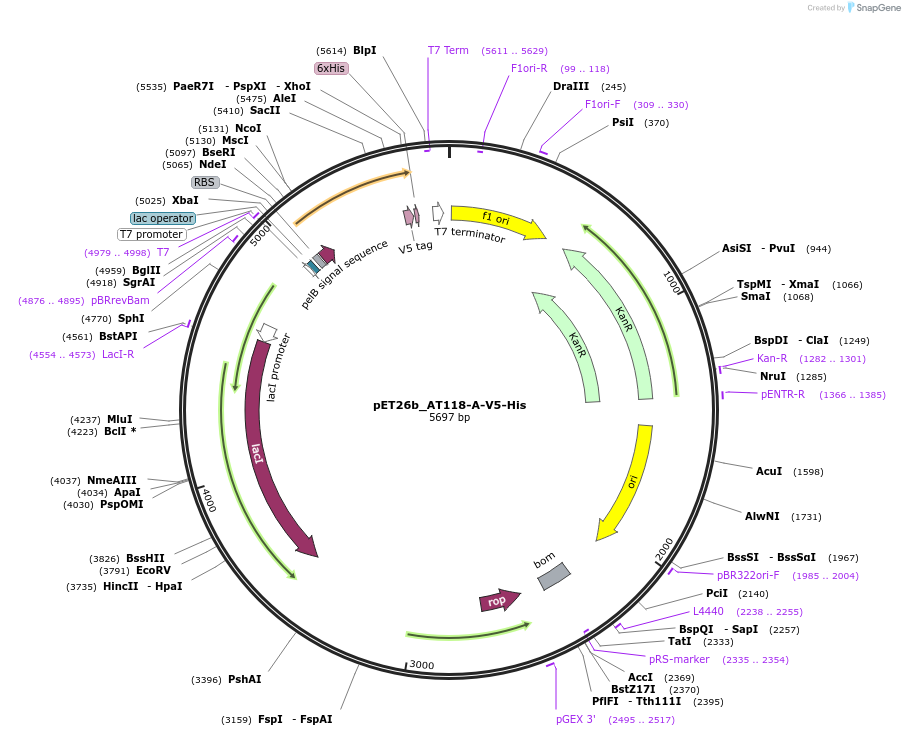
pET26b_AT118-A-V5-His
Plasmid#222849PurposeE. coli periplasmic expression plasmid encoding Nb. AT118-A, a nanobody antagonist of the human Angiotensin II type I receptor (AT1R)DepositorInsertAT118-A
TagsV5-HisExpressionBacterialAvailable SinceSept. 13, 2024AvailabilityAcademic Institutions and Nonprofits only -
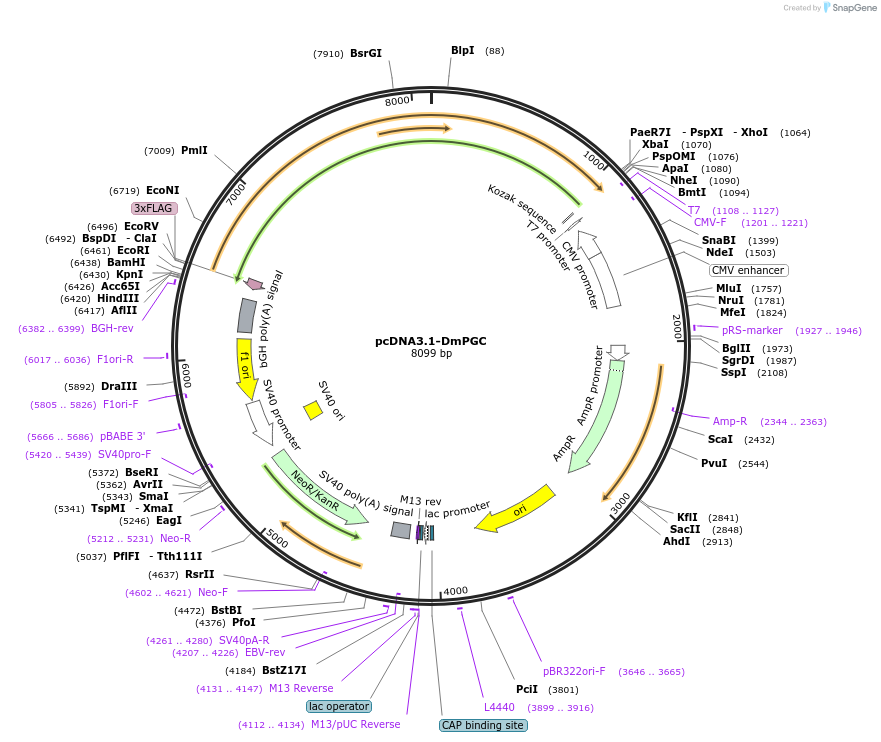
pcDNA3.1-DmPGC
Plasmid#220309Purposered-light-activated guanylyl cyclase DmPGCDepositorInsertDmPGC
ExpressionMammalianAvailable SinceJune 28, 2024AvailabilityAcademic Institutions and Nonprofits only -
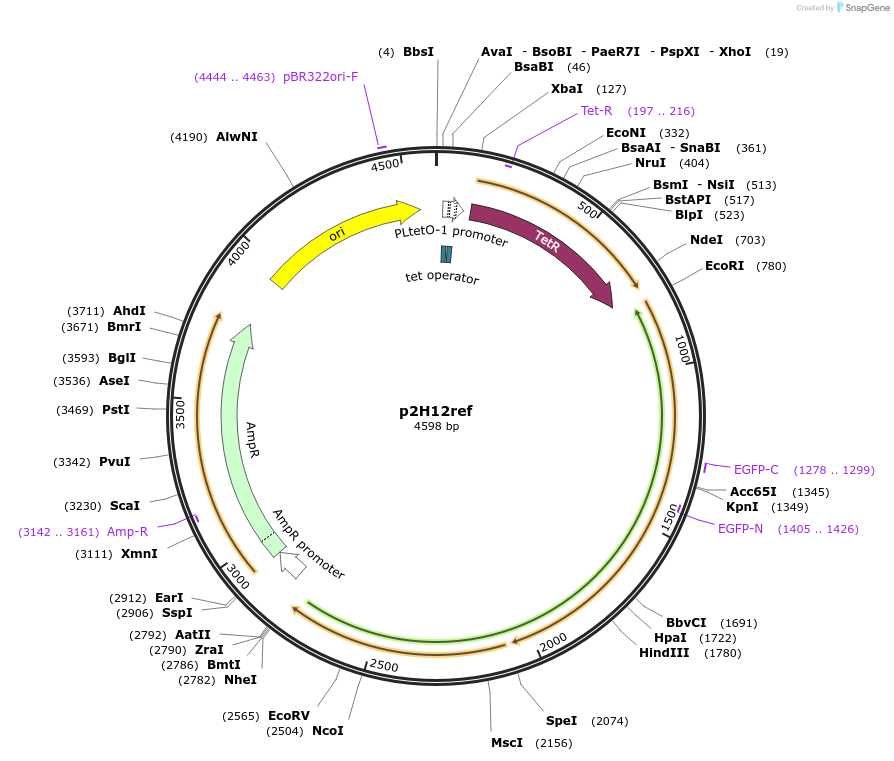
p2H12ref
Plasmid#221188PurposecdGreen expressed in operon with mScarlet-I from an aTc-inducible promoter (Internal Lab ID: UJ11206)DepositorInsertscdGreen
mScarlet-I
ExpressionBacterialMutationN-terminus changed: starts with MSKKYGEAVIKE ...PromoterNone (same as cdGreen) and PtetAvailable SinceJune 25, 2024AvailabilityAcademic Institutions and Nonprofits only -
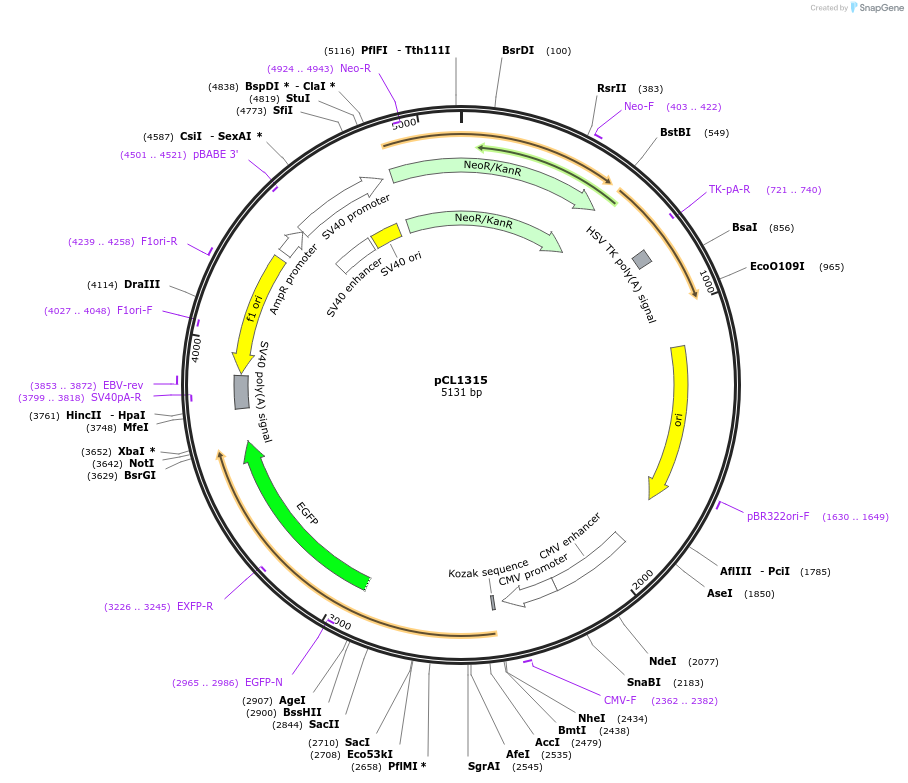
pCL1315
Plasmid#217957PurposeExpression of human H3.1 carrying a triple point mutation to disrupt folding with histone H4. Expression in mammalian cells driven by the CMV promoter.DepositorInsertHistone H3.1 (H3C1 Human)
TagseGFPExpressionMammalianMutationF84A, L92A, Y99APromoterCMVAvailable SinceApril 18, 2024AvailabilityAcademic Institutions and Nonprofits only -
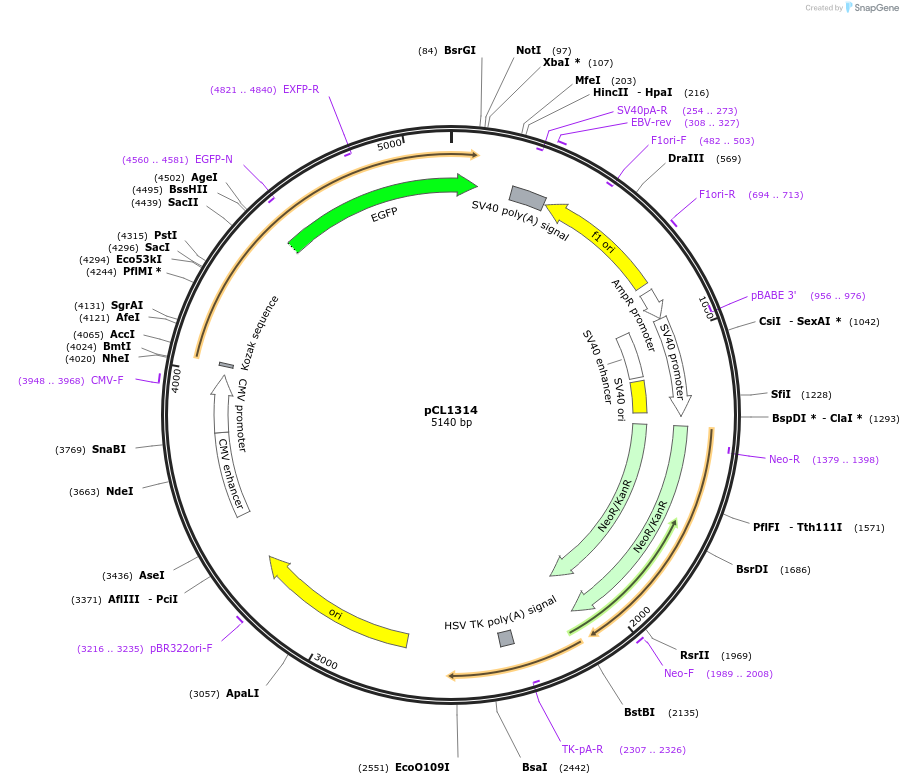
pCL1314
Plasmid#217956PurposeExpression of human H3.1 carrying a triple glycine insertion to disrupt folding with H4. Expression in mammalian cells driven by the CMV promoter.DepositorInsertHistone H3.1 (H3C1 Human)
TagseGFPExpressionMammalianMutationGGG insertion between A95 and C96PromoterCMVAvailable SinceApril 18, 2024AvailabilityAcademic Institutions and Nonprofits only -
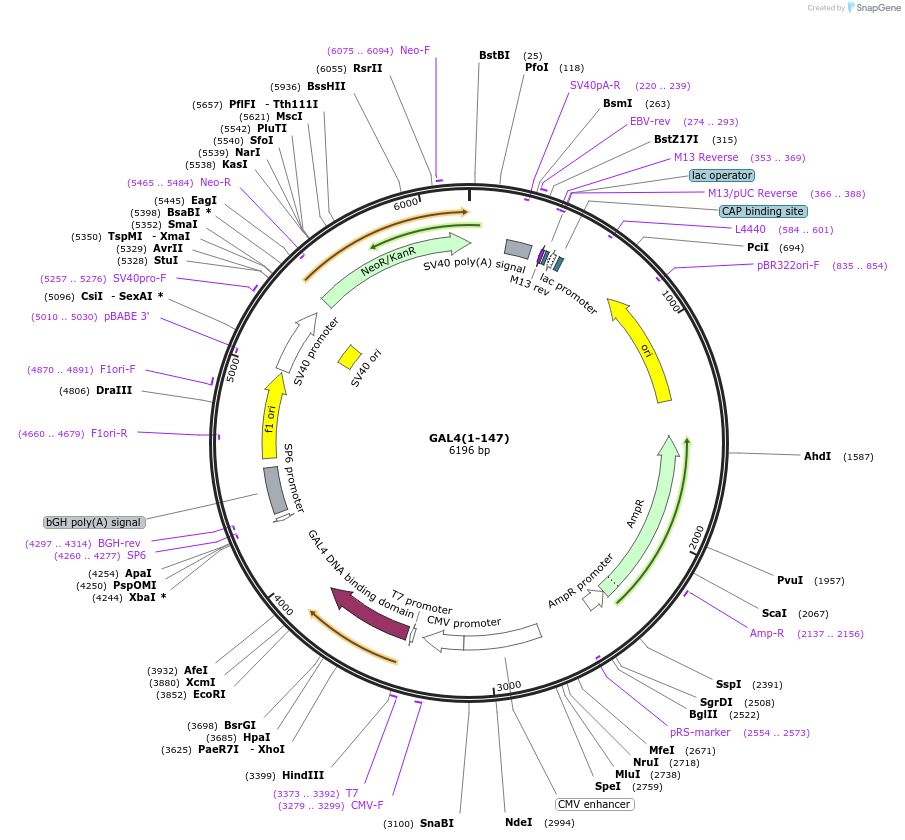
GAL4(1-147)
Plasmid#195029PurposeThe control vector GAL4 (1-147) was generated by introducing a stop codon immediately after the GAL4 (1-147) sequence in the GAL4-Creb5 (1-128) vectorDepositorInsertGAL4 (1-147)
ExpressionMammalianAvailable SinceFeb. 8, 2024AvailabilityAcademic Institutions and Nonprofits only -
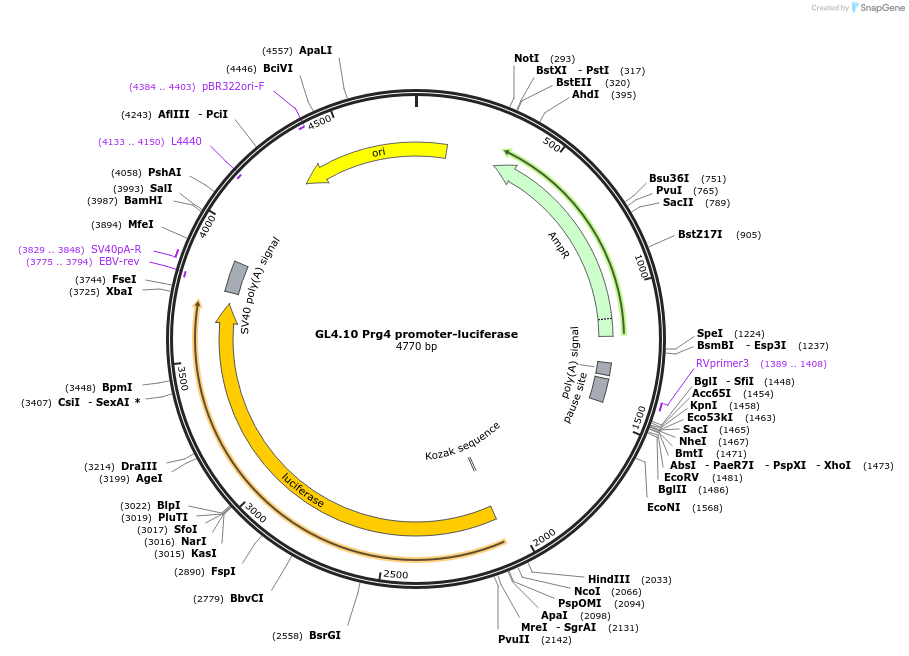
GL4.10 Prg4 promoter-luciferase
Plasmid#195034PurposePrg4 enhancer/promoterfirefly luciferase constructs were constructed by cloning various Prg4 regulatory fragments into pGL4.10[luc2] vectorDepositorInsertPrg4 promoter (PRG4 Bovine)
UseLuciferaseAvailable SinceFeb. 8, 2024AvailabilityAcademic Institutions and Nonprofits only -
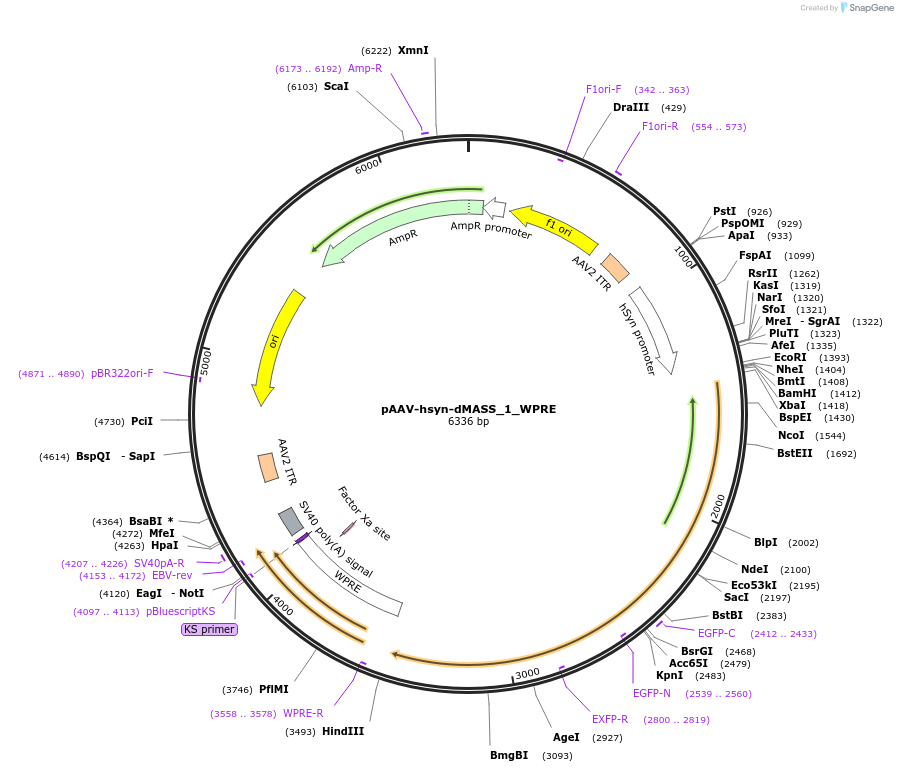
pAAV-hsyn-dMASS_1_WPRE
Plasmid#209765PurposeAAV virus production for neuronal expression of dMASS_1 under hSyn promoterDepositorInsertdMASS_1
UseAAVExpressionMammalianAvailable SinceNov. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
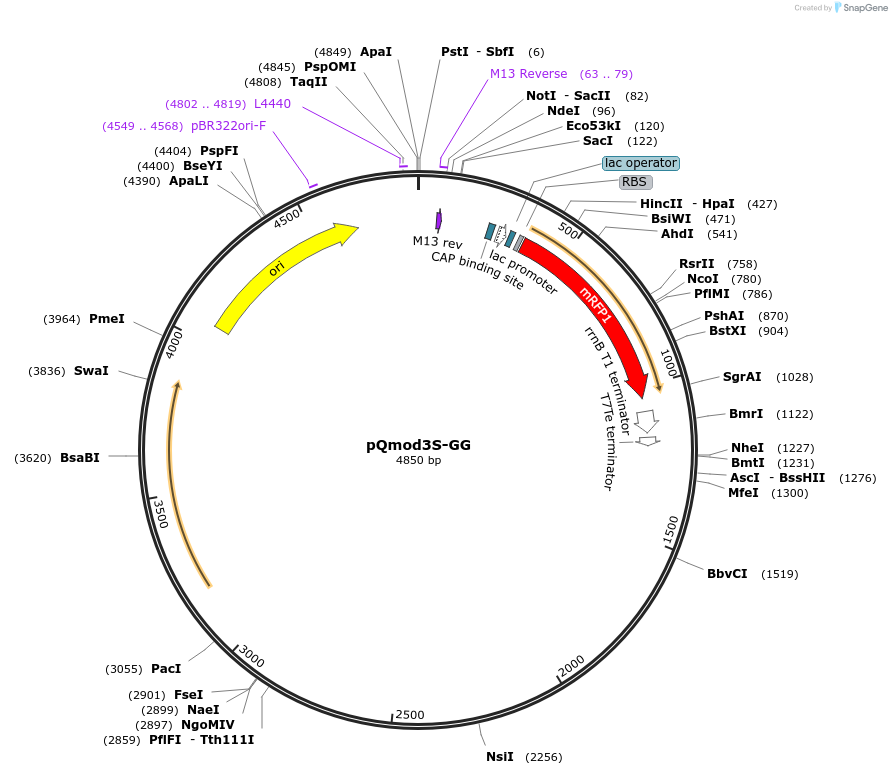
pQmod3S-GG
Plasmid#191349PurposeClostridium expression vector (pCB102 origin, specR) with drop-out Plac-RFP cassette flanked with BsaI sitesDepositorInsertPlac-RFP (IPTG-inducible red fluorescent protein)
ExpressionBacterialPromoterPlacAvailable SinceNov. 29, 2022AvailabilityAcademic Institutions and Nonprofits only -
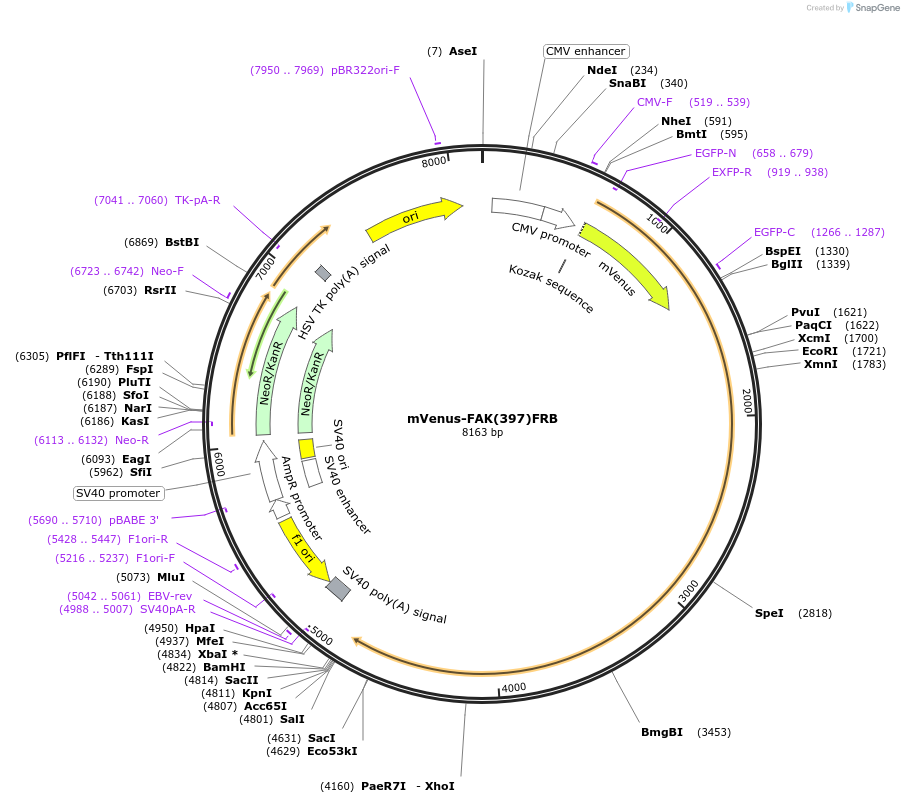
mVenus-FAK(397)FRB
Plasmid#188660PurposemVenus tagged Focal Adhesion Kinase with FRB inserted at Y397DepositorInsertFocal Adhesion Kinase
TagsmVenusExpressionMammalianMutationInsertion of FRB at Tyrosine 397PromoterCMVAvailable SinceAug. 19, 2022AvailabilityAcademic Institutions and Nonprofits only -
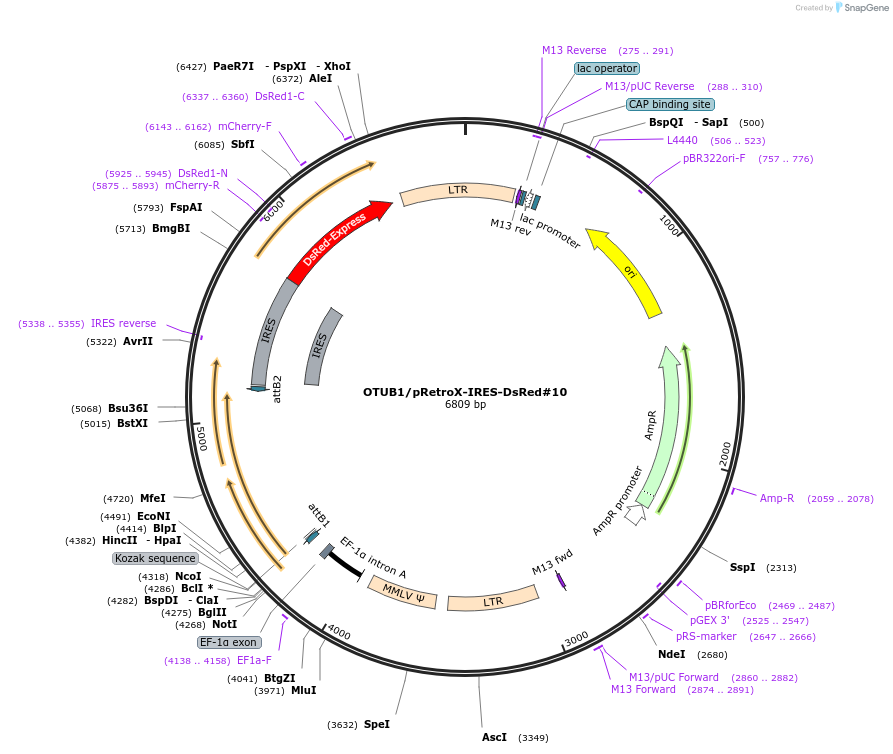
OTUB1/pRetroX-IRES-DsRed#10
Plasmid#180279PurposePlasmid encoding the human protease OTUB1 without a tag. Polycistronic red fluorescent retroviral vector (unmodified from TAKARA BIO).DepositorAvailable SinceMarch 22, 2022AvailabilityAcademic Institutions and Nonprofits only -
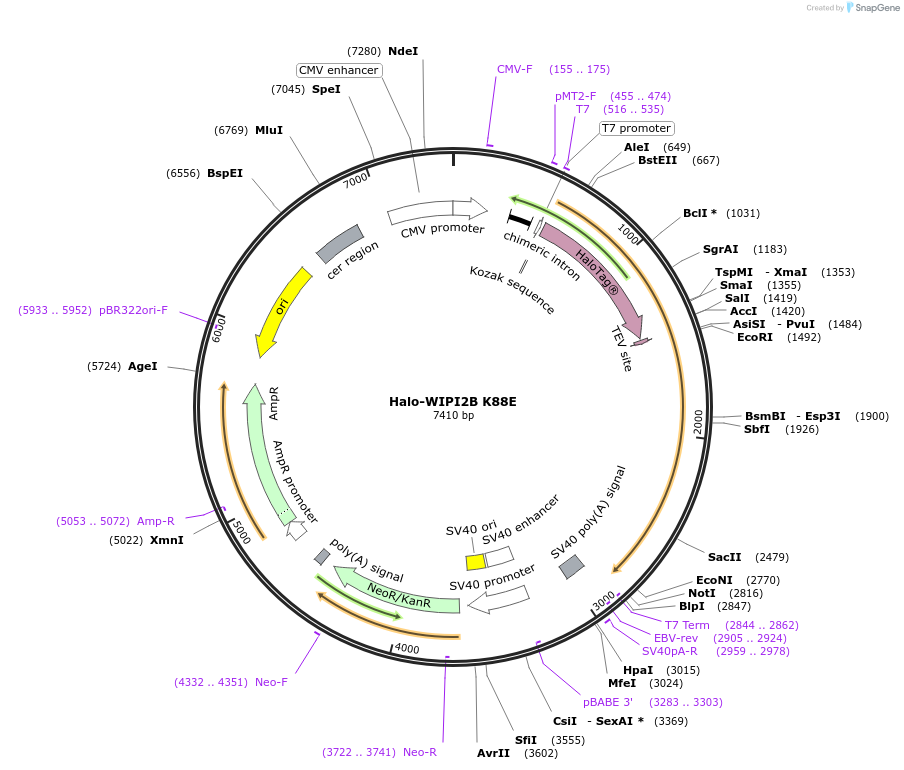
Halo-WIPI2B K88E
Plasmid#175028PurposeExpresses WIPI2B (K88E) fused to Halo in mammalian cells to abrogate interaction with ATG16L1DepositorInsertWIPI2B (K88E) (WIPI2 Human)
TagsHaloTagExpressionMammalianMutationLysine 88 to Glutamate (AAG to GAG)PromoterCMVAvailable SinceDec. 15, 2021AvailabilityAcademic Institutions and Nonprofits only -
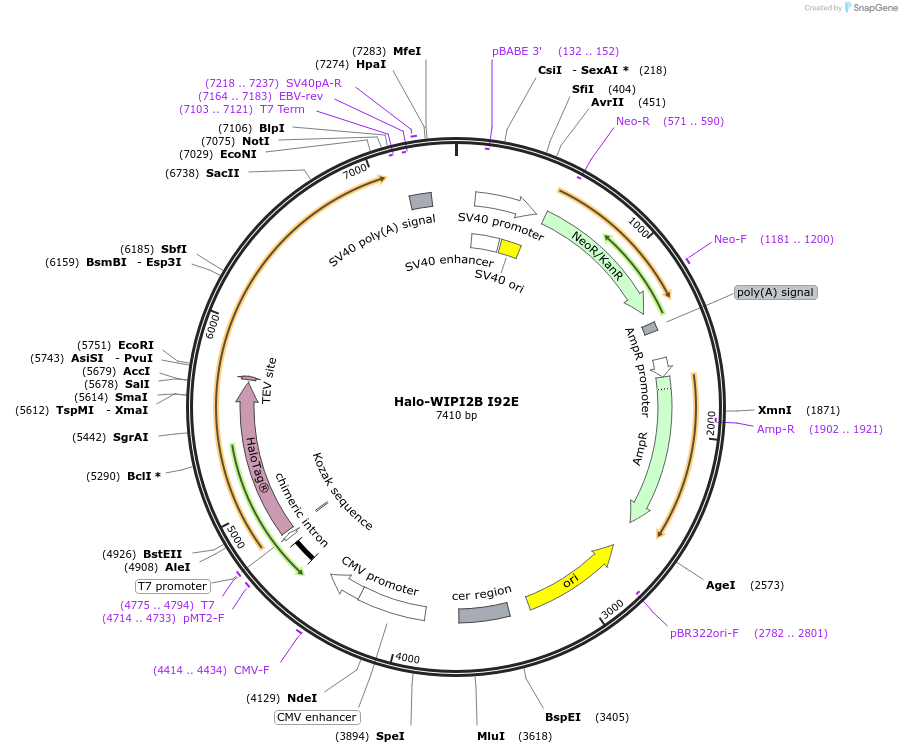
Halo-WIPI2B I92E
Plasmid#175029PurposeExpresses WIPI2B (I92E) fused to Halo in mammalian cells to abrogate interaction with ATG16L1DepositorInsertWIPI2B (I92E) (WIPI2 Human)
TagsHaloTagExpressionMammalianMutationIsoleucine 92 to Glutamate (ATC to GAA)PromoterCMVAvailable SinceDec. 15, 2021AvailabilityAcademic Institutions and Nonprofits only -
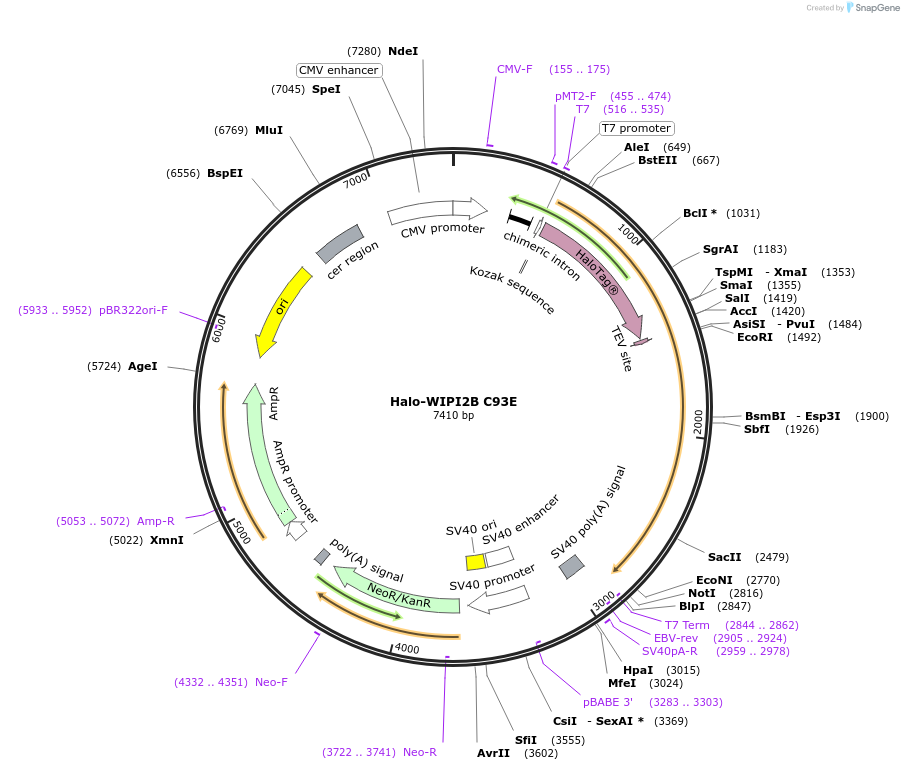
Halo-WIPI2B C93E
Plasmid#175030PurposeExpresses WIPI2B (C93E) fused to Halo in mammalian cells to abrogate interaction with ATG16L1DepositorInsertWIPI2B (C93E) (WIPI2 Human)
TagsHaloTagExpressionMammalianMutationCysteine 93 to Glutamate (TGC to GAG)PromoterCMVAvailable SinceDec. 15, 2021AvailabilityAcademic Institutions and Nonprofits only -
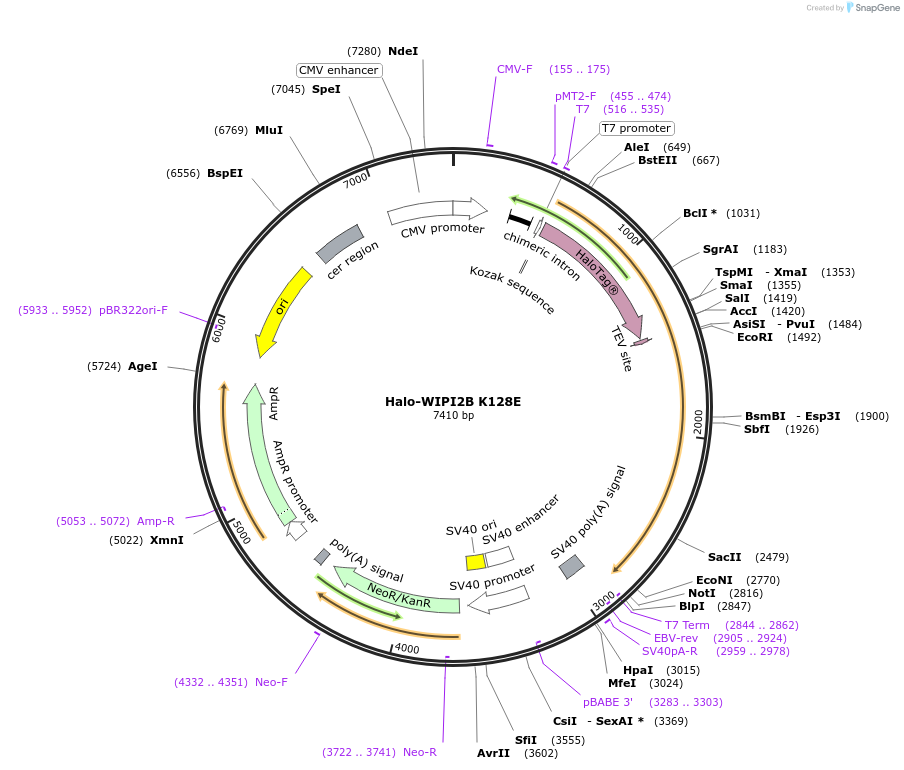
Halo-WIPI2B K128E
Plasmid#175031PurposeExpresses WIPI2B (K128E) fused to Halo in mammalian cells to abrogate interaction with ATG16L1DepositorInsertWIPI2B (K128E) (WIPI2 Human)
TagsHaloTagExpressionMammalianMutationLysine 128 to Glutamate (AAG to GAG)PromoterCMVAvailable SinceDec. 15, 2021AvailabilityAcademic Institutions and Nonprofits only -
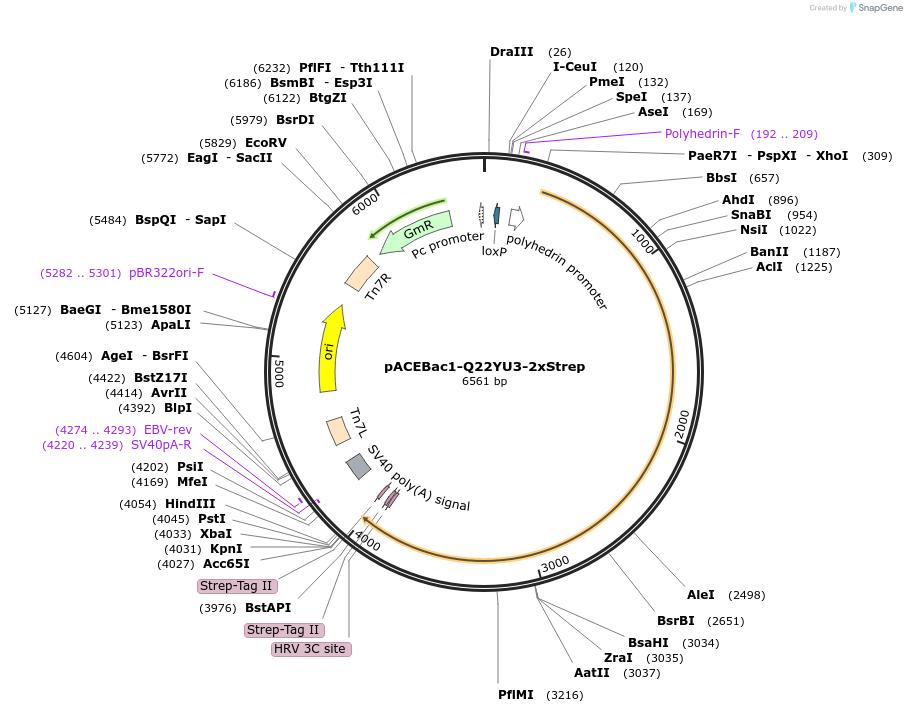
pACEBac1-Q22YU3-2xStrep
Plasmid#170315PurposePlasmid for insect cell expression of Tetrahymena thermophila Q22YU3 C-terminally tagged with 2x Strep-tag IIDepositorInsertQ22YU3 (TTHERM_00122270 Tetrahymena thermophila)
TagsStrep-Tag IIExpressionInsectPromoterPolyhedrinAvailable SinceJuly 23, 2021AvailabilityAcademic Institutions and Nonprofits only -
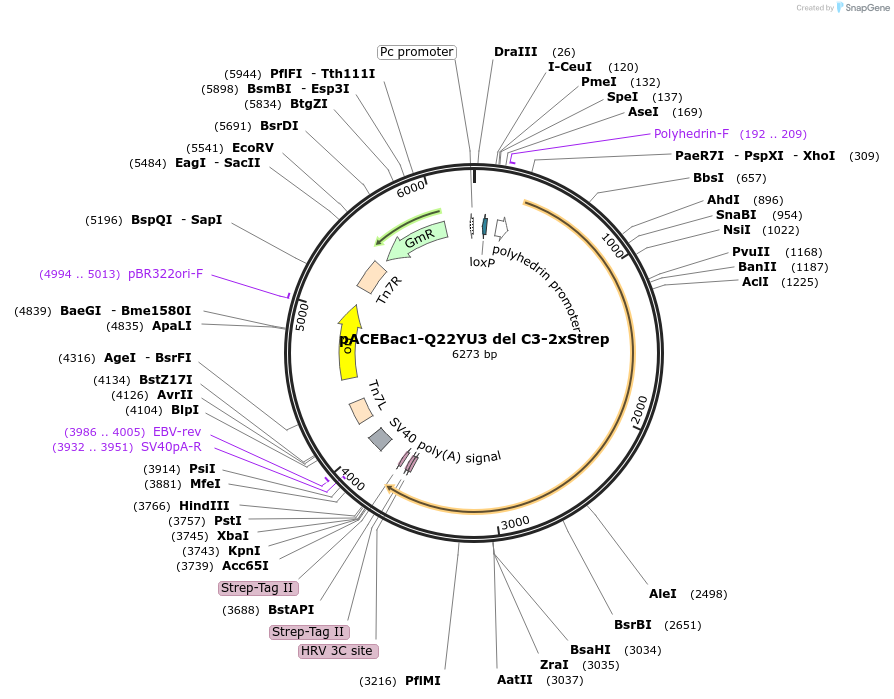
pACEBac1-Q22YU3 del C3-2xStrep
Plasmid#170316PurposePlasmid for insect cell expression of Tetrahymena thermophila Q22YU3 with a C-terminal truncation (C3-finger). Tagged with 2x Strep-tag II at the C-terminus.DepositorInsertQ22YU3 C-terminal truncation (Delta-C3)
TagsStrep-Tag IIExpressionInsectMutationDeletion of C-terminal C3-fingerPromoterpolyhedrinAvailable SinceMay 28, 2021AvailabilityAcademic Institutions and Nonprofits only -
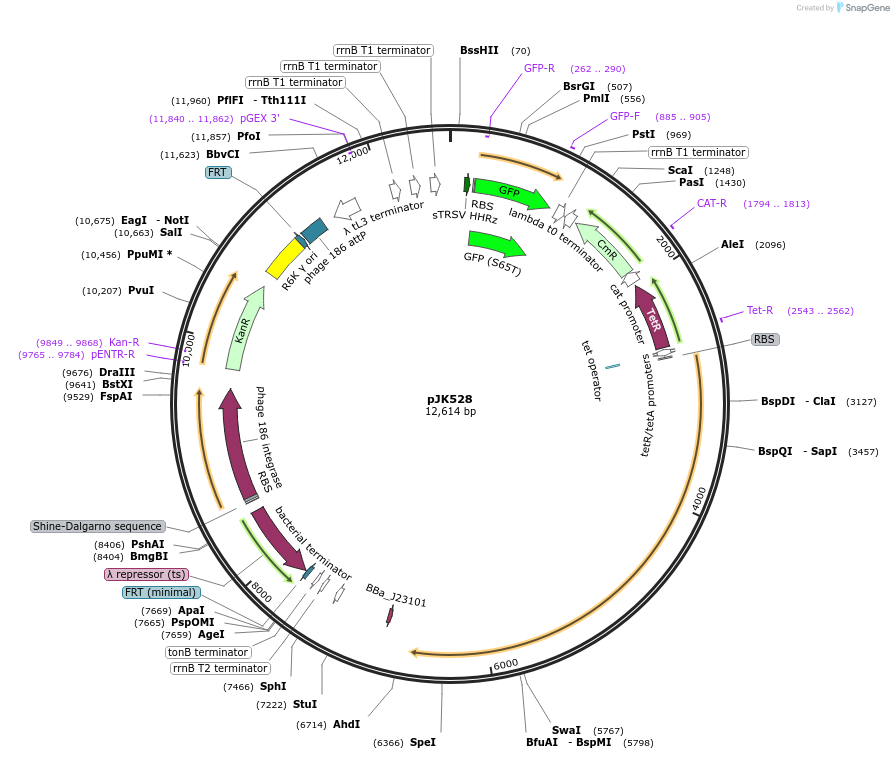
pJK528
Plasmid#155388PurposedCas12a oscillator, reverse crRNAs + PA4-mVenus. (Integration of 511; pOSIP)DepositorInsertsdCas12a (F. novicida)
PA4-mVenus
dCas12a oscillator crRNAs
UseCRISPRExpressionBacterialMutationD917A (nuclease-deactivating)Available SinceFeb. 12, 2021AvailabilityAcademic Institutions and Nonprofits only