We narrowed to 29,080 results for: Tat
-
Plasmid#63214PurposeConstitutively active NFAT1, mutated to interfere selectively with the NFAT:AP-1 interactio. Contains Puromycin selection markerDepositorInsertCA-RIT-NFAT1 (Nfatc2 Mouse)
UseRetroviralTagsHAExpressionMammalianMutationAmino acids 1-3 removed; CA: PASSGSSASF mutated t…Available SinceJuly 22, 2015AvailabilityAcademic Institutions and Nonprofits only -
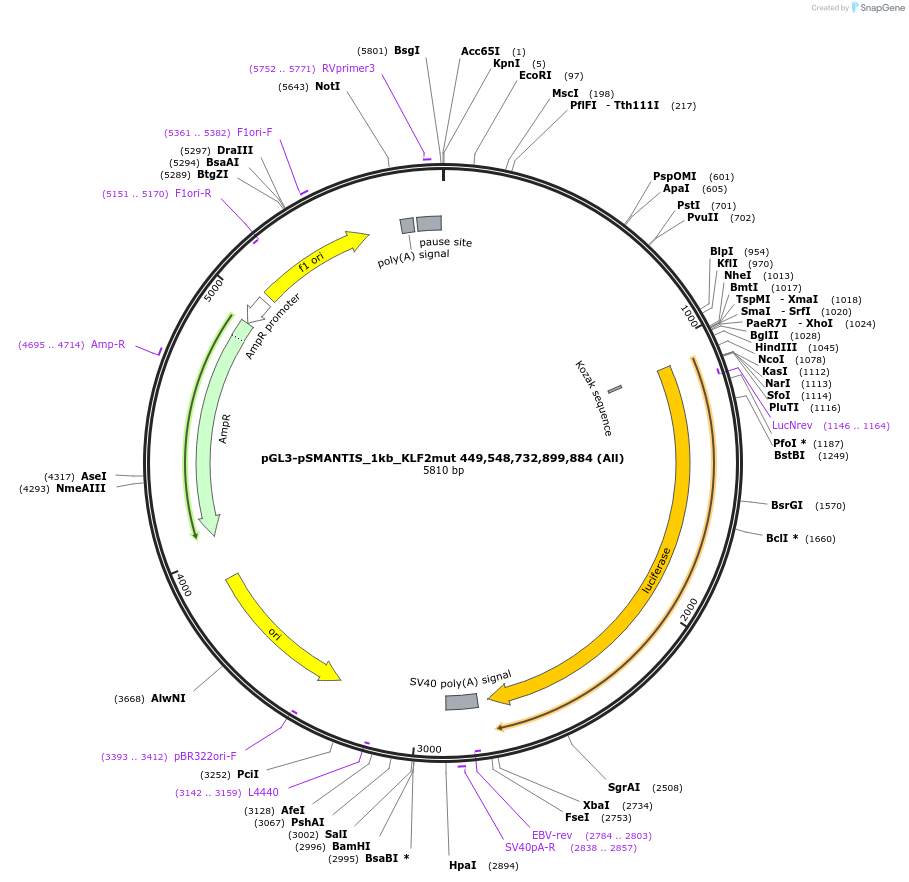
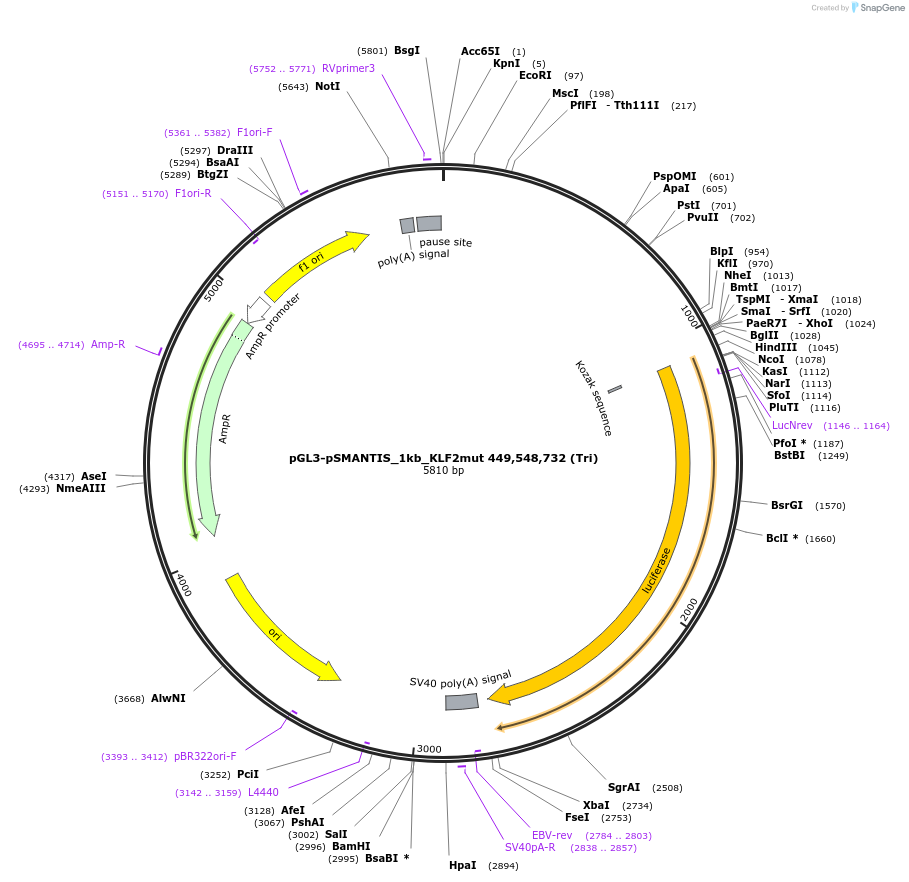
pGL3-pSMANTIS_1kb_KLF2mut 449,548,732,899,884 (All)
Plasmid#194190PurposeIncludes the promoter (1kb) of SMTS with 5 mutated KLF binding sites (positions 449,548,732,899,884)DepositorInsertSMANTIS promoter (1kb) (SMANTIS Human)
UseLuciferaseMutation5 mutated KLF binding sites (positions 449,548,73…PromoterSMANTIS promoter (1kb), mutatedAvailable SinceJan. 24, 2023AvailabilityAcademic Institutions and Nonprofits only -
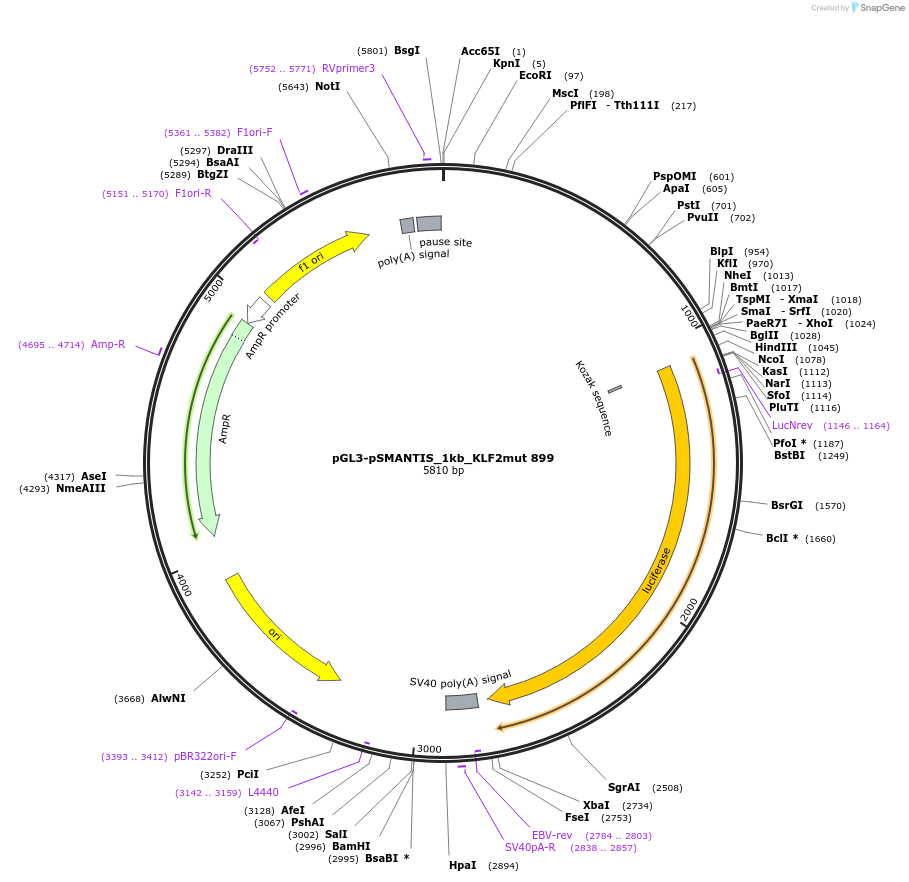
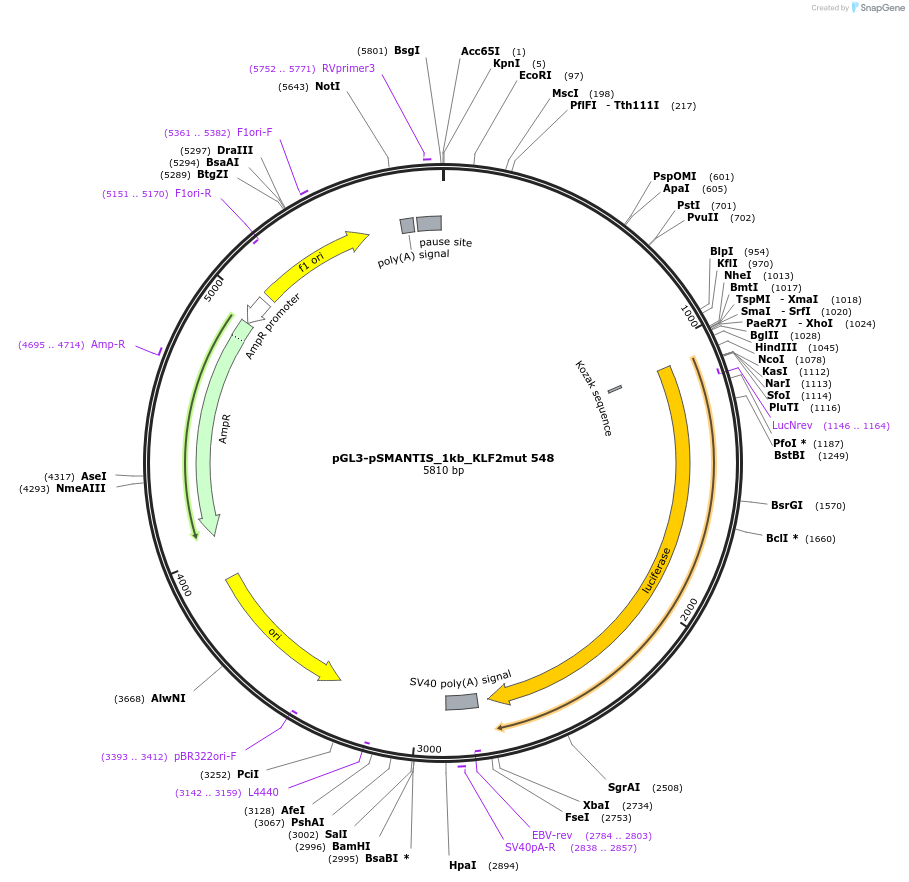
pGL3-pSMANTIS_1kb_KLF2mut 899
Plasmid#194193PurposeIncludes the promoter (1kb) of SMTS with mutated KLF binding sites (position 899)DepositorInsertSMANTIS promoter (1kb) (SMANTIS Human)
UseLuciferaseMutationmutated KLF binding sites (position 899)PromoterSMANTIS promoter (1kb), mutatedAvailable SinceJan. 24, 2023AvailabilityAcademic Institutions and Nonprofits only -
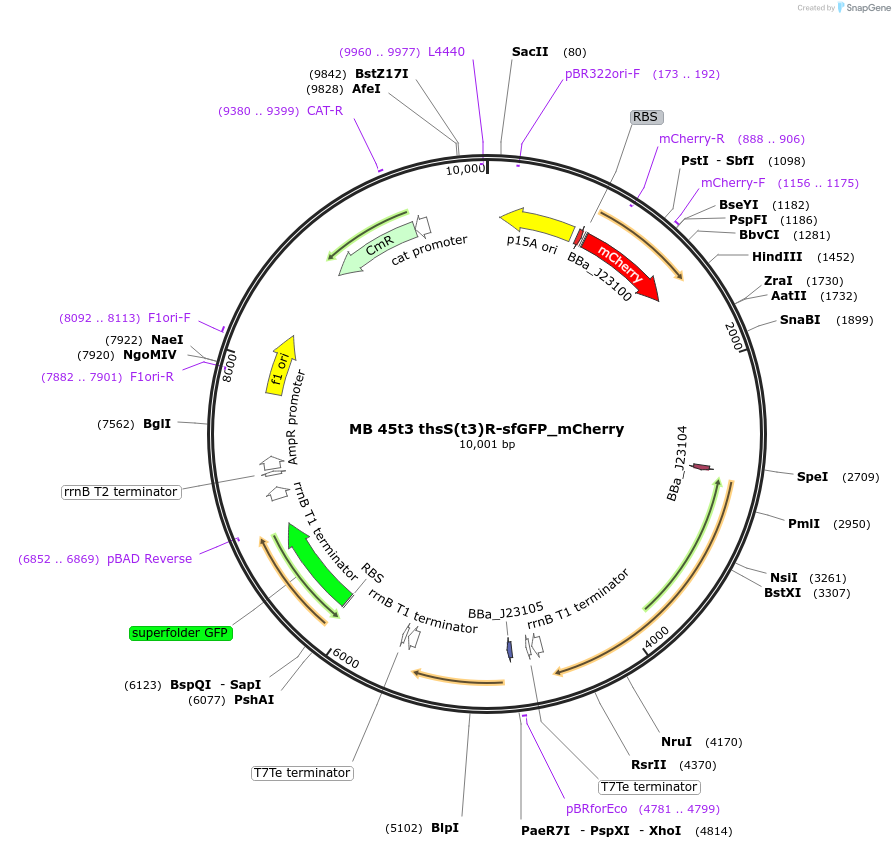
MB 45t3 thsS(t3)R-sfGFP_mCherry
Plasmid#232470PurposeOptimized thiosulfate sensor with fluorescent reporter (GFP), constitutive mCherryDepositorInsertsthsS(t3)
thsR
PphsA342
sfGFP
AxeTxe
mCherry
ExpressionBacterialMutationContains the following mutations K286Q, Q350H, I…PromoterJ23100 (TTGACGGCTAGCTCAGTCCTAGGTACAGTGCTAGC), J23…Available SinceSept. 17, 2025AvailabilityAcademic Institutions and Nonprofits only -
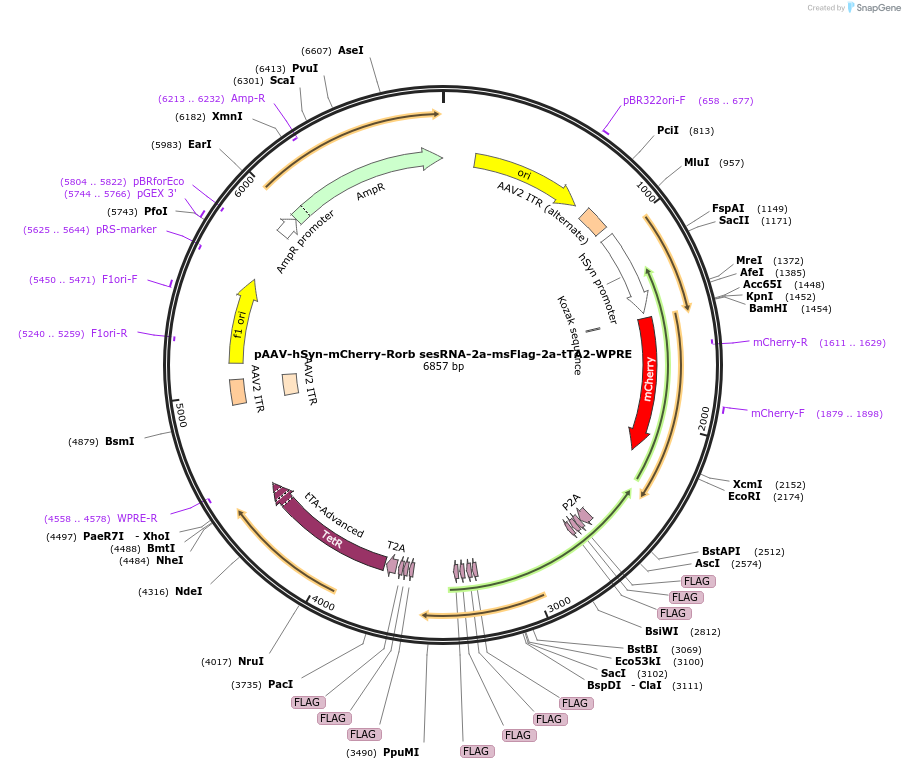
pAAV-hSyn-mCherry-Rorb sesRNA-2a-msFlag-2a-tTA2-WPRE
Plasmid#239029PurposeExpression of mCherry, sesRNA in mammalian cells, with msFlag and tTA2 as efRNA.DepositorAvailable SinceMay 14, 2025AvailabilityAcademic Institutions and Nonprofits only -
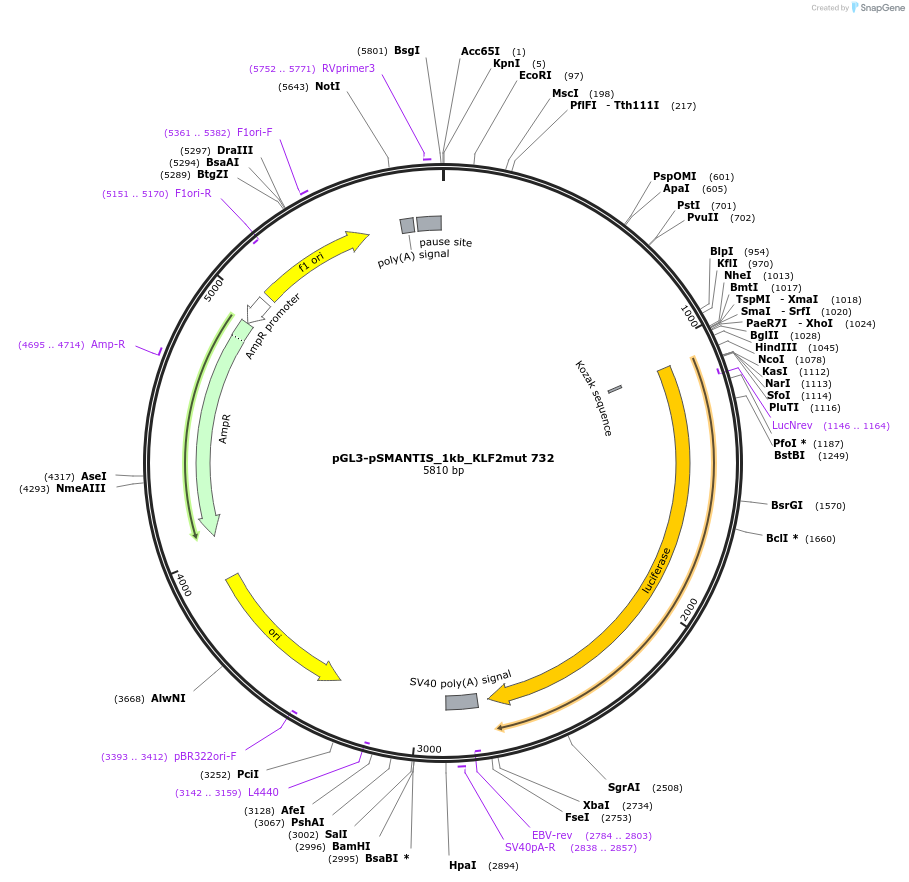
pGL3-pSMANTIS_1kb_KLF2mut 732
Plasmid#194195PurposeIncludes the promoter (1kb) of SMTS with mutated KLF binding sites (position 732)DepositorInsertSMANTIS promoter (1kb) (SMANTIS Human)
UseLuciferaseMutationmutated KLF binding sites (position 732)PromoterSMANTIS promoter (1kb), mutatedAvailable SinceJan. 24, 2023AvailabilityAcademic Institutions and Nonprofits only -
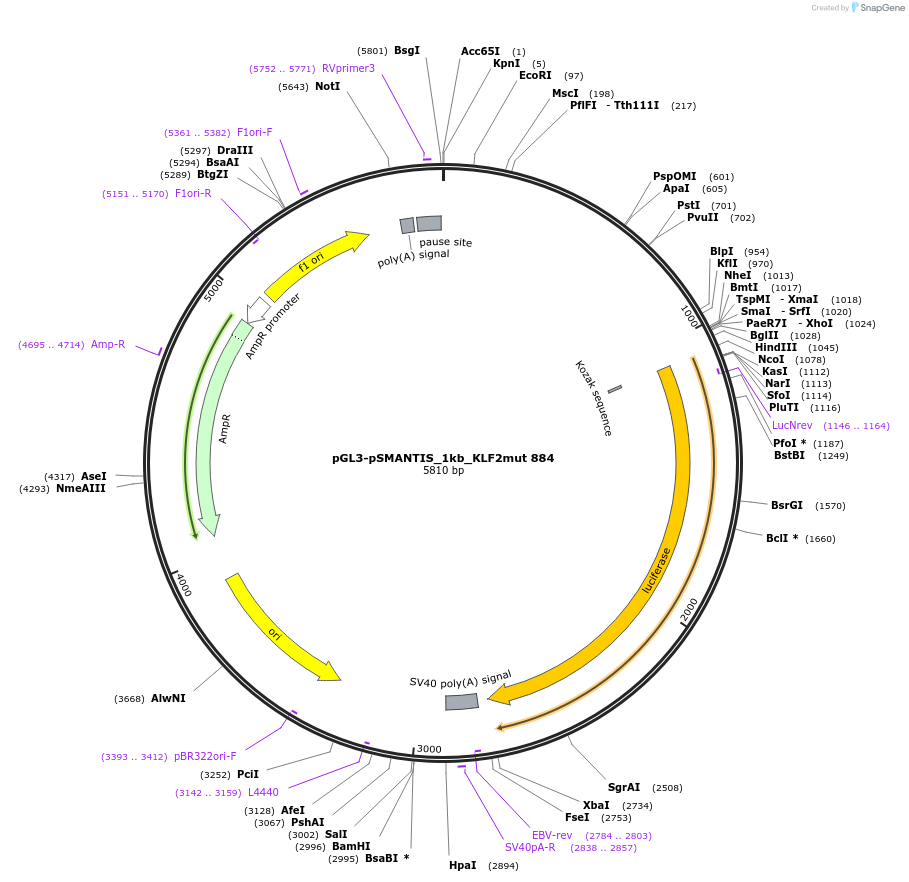
pGL3-pSMANTIS_1kb_KLF2mut 884
Plasmid#194196PurposeIncludes the promoter (1kb) of SMTS with mutated KLF binding sites (position 884)DepositorInsertSMANTIS promoter (1kb) (SMANTIS Human)
UseLuciferaseMutationmutated KLF binding sites (position 884)PromoterSMANTIS promoter (1kb), mutatedAvailable SinceJan. 24, 2023AvailabilityAcademic Institutions and Nonprofits only -
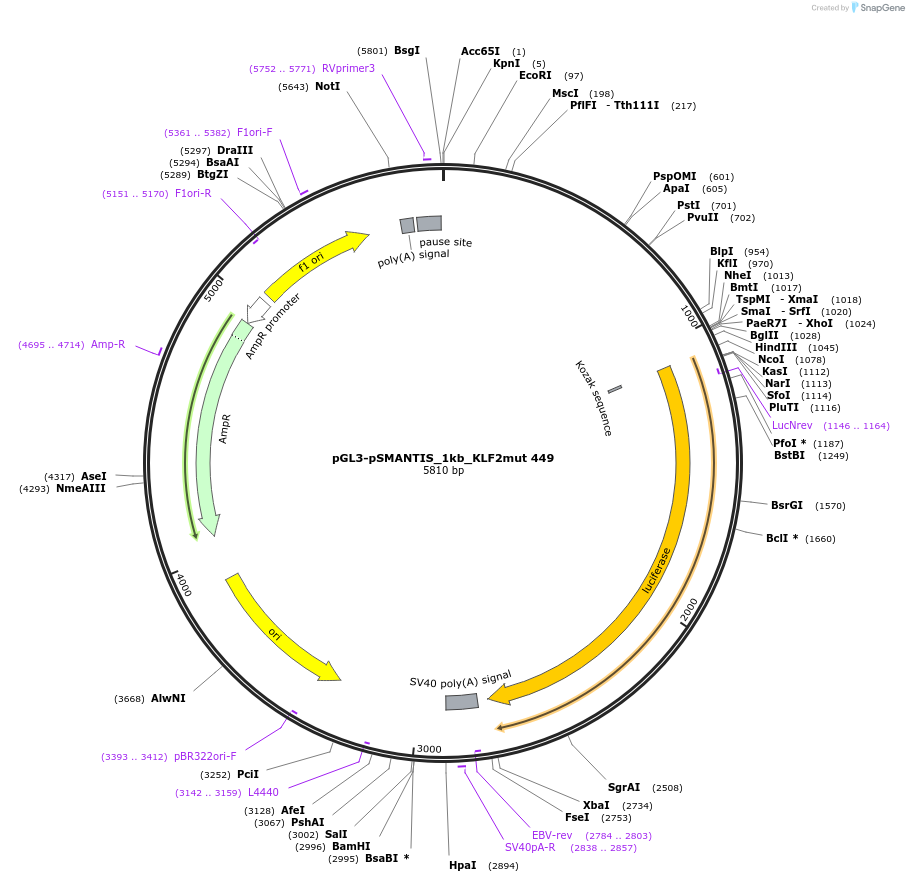
pGL3-pSMANTIS_1kb_KLF2mut 449
Plasmid#194197PurposeIncludes the promoter (1kb) of SMTS with mutated KLF binding sites (position 449)DepositorInsertSMANTIS promoter (1kb) (SMANTIS Human)
UseLuciferaseMutationmutated KLF binding sites (position 449)PromoterSMANTIS promoter (1kb), mutatedAvailable SinceJan. 24, 2023AvailabilityAcademic Institutions and Nonprofits only -
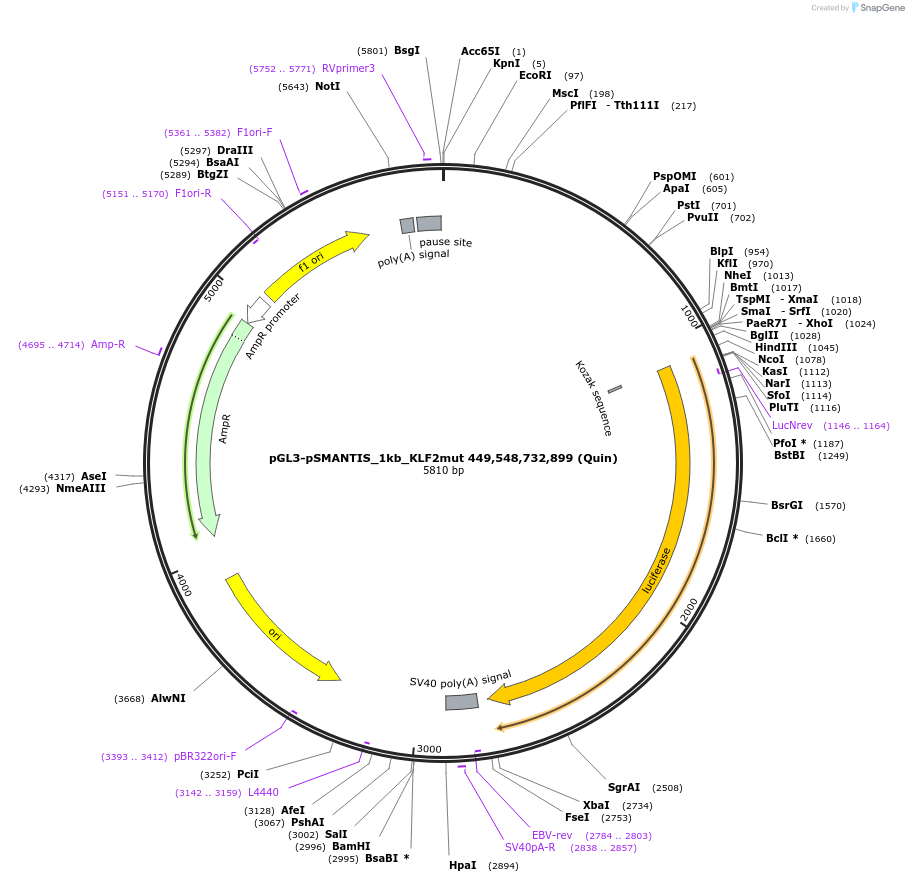
pGL3-pSMANTIS_1kb_KLF2mut 449,548,732,899 (Quin)
Plasmid#194191PurposeIncludes the promoter (1kb) of SMTS with 4 mutated KLF binding sites (positions 449,548,732,899)DepositorInsertSMANTIS promoter (1kb) (SMANTIS Human)
UseLuciferaseMutation4 mutated KLF binding sites (positions 449,548,73…PromoterSMANTIS promoter (1kb), mutatedAvailable SinceJan. 24, 2023AvailabilityAcademic Institutions and Nonprofits only -
pGL3-pSMANTIS_1kb_KLF2mut 449,548,732 (Tri)
Plasmid#194192PurposeIncludes the promoter (1kb) of SMTS with 3 mutated KLF binding sites (positions 449,548,732)DepositorInsertSMANTIS promoter (1kb) (SMANTIS Human)
UseLuciferaseMutation3 mutated KLF binding sites (positions 449,548,73…PromoterSMANTIS promoter (1kb), mutatedAvailable SinceJan. 24, 2023AvailabilityAcademic Institutions and Nonprofits only -
pGL3-pSMANTIS_1kb_KLF2mut 548
Plasmid#194194PurposeIncludes the promoter (1kb) of SMTS with mutated KLF binding sites (position 548)DepositorInsertSMANTIS promoter (1kb) (SMANTIS Human)
UseLuciferaseMutationmutated KLF binding sites (position 548)PromoterSMANTIS promoter (1kb), mutatedAvailable SinceJan. 24, 2023AvailabilityAcademic Institutions and Nonprofits only -
pCMV-hAR-H874Y
Plasmid#89087Purposemammalian expression of human androgen receptor with prostate cancer somatic mutation H874Y from CWR22 xenograftDepositorInserthAR-H874Y prostate cancer mutant (AR Human)
ExpressionMammalianMutationhAR-H874YPromoterCMVAvailable SinceApril 11, 2017AvailabilityAcademic Institutions and Nonprofits only -
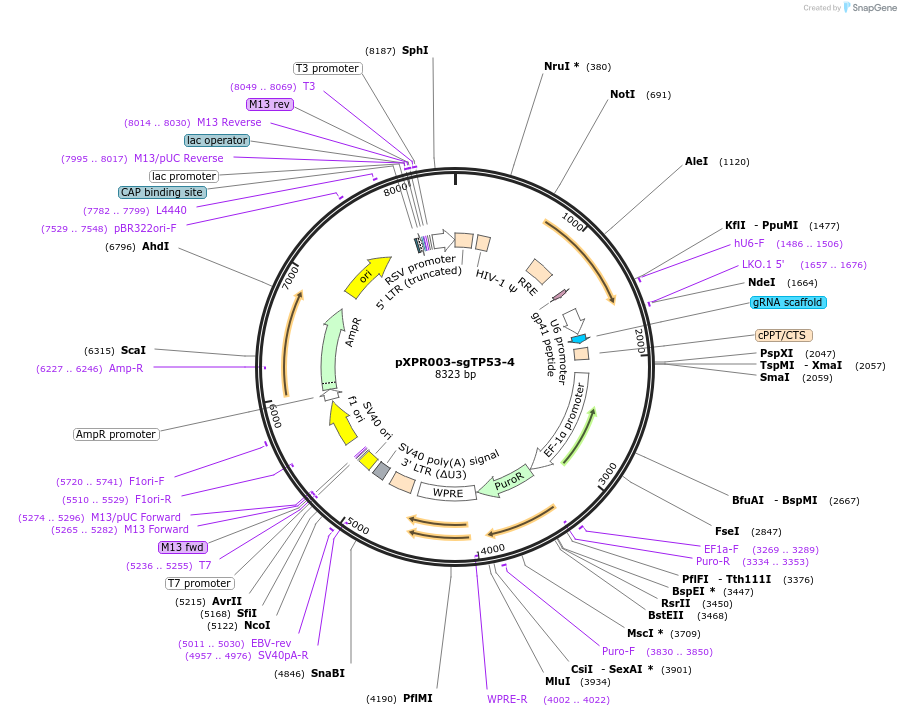
pXPR003-sgTP53-4
Plasmid#118022PurposeConstitutive expression of sgRNA targeting human TP53DepositorAvailable SinceOct. 29, 2018AvailabilityAcademic Institutions and Nonprofits only -
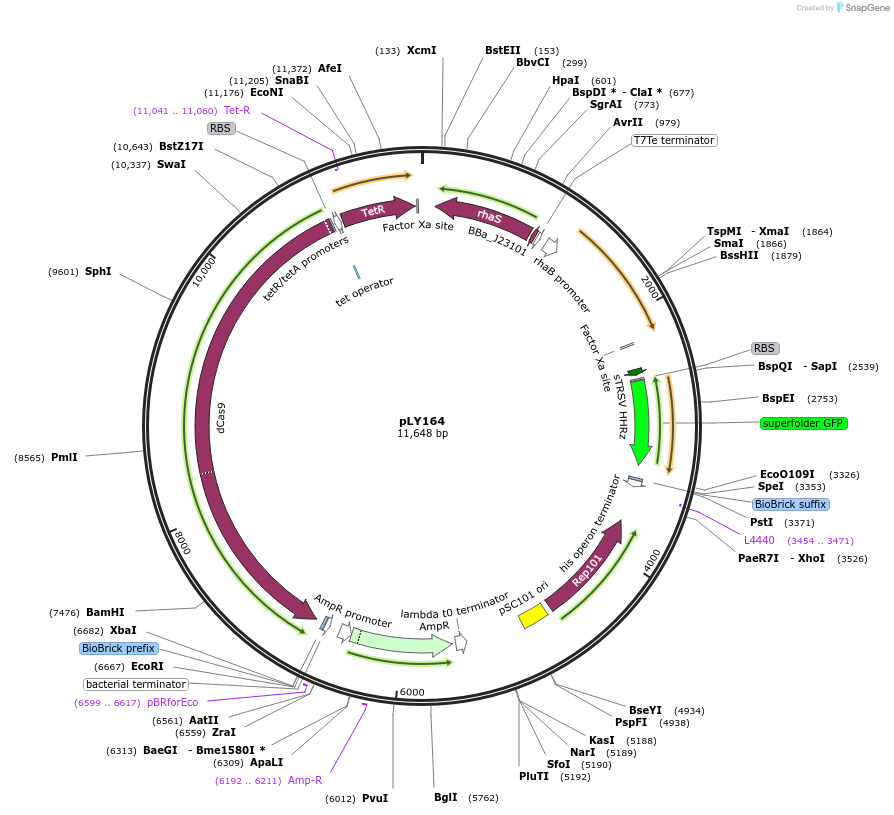
pLY164
Plasmid#184140PurposeReporter circuit for hijacking RFP mRNA as CRISPR RNA (target site 3)DepositorInsertsdCas9
tetR
pspFΔHTH::λN22plus
sfgfp
rhaS
UseSynthetic BiologyTagsASV tag and λN22plusExpressionBacterialMutationMutations D10A and H840A on WT cas9. Synonymous m…PromoterPcon, PpspA-R3, PrhaB, and PtetAvailable SinceNov. 29, 2022AvailabilityAcademic Institutions and Nonprofits only -
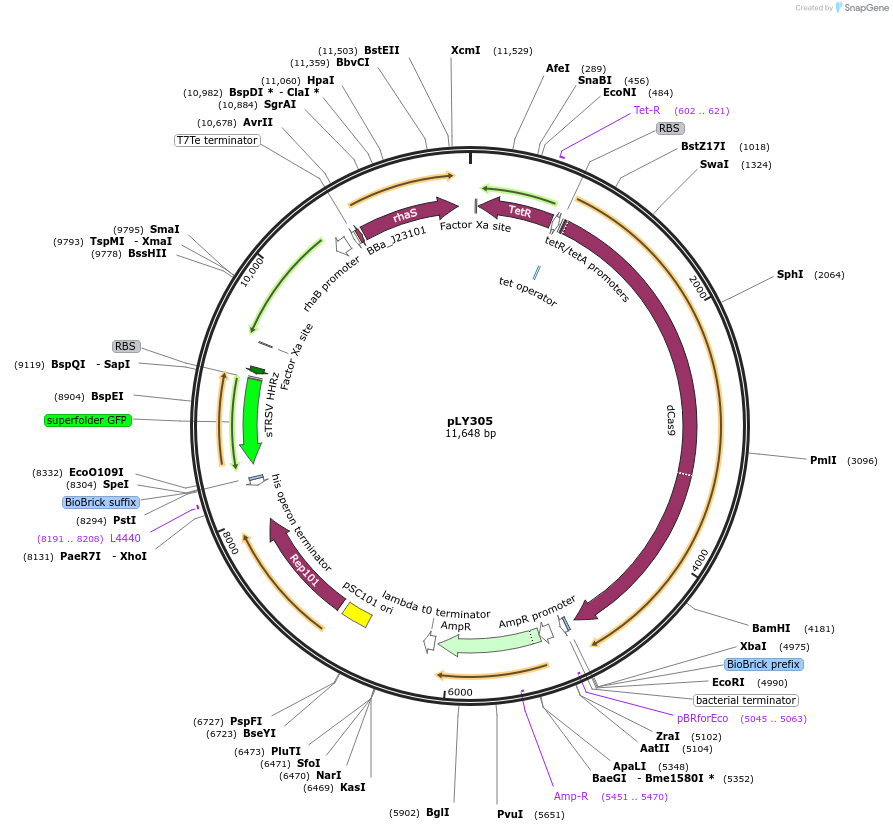
pLY305
Plasmid#184159PurposeReporter circuit for hijacking mRNA of zraP as CRISPR RNADepositorInsertsdCas9
tetR
pspFΔHTH::λN22plus
mutsfgfp
rhaS
UseSynthetic BiologyTagsASV tag and λN22plusExpressionBacterialMutationK140N, Mutations D10A and H840A on WT cas9. Synon…PromoterPcon, PpspA-zraP-22, PrhaB, and PtetAvailable SinceOct. 31, 2022AvailabilityAcademic Institutions and Nonprofits only -
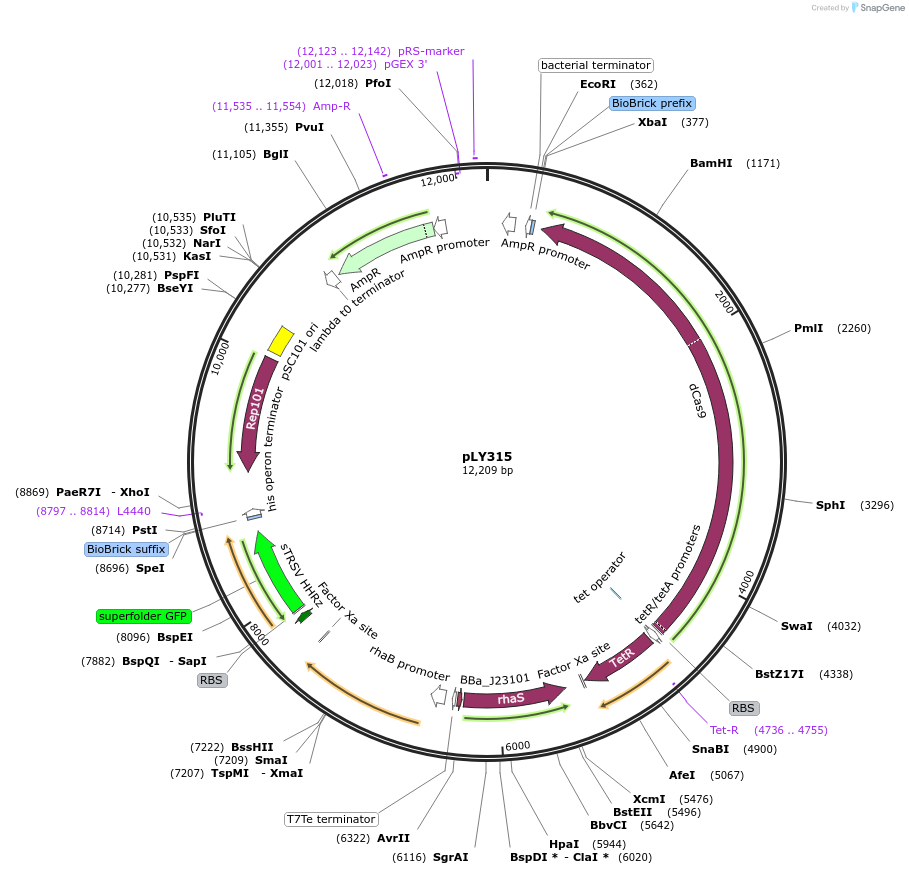
pLY315
Plasmid#184160PurposeReporter circuit for hijacking mRNA of zntA as CRISPR RNADepositorInsertsdCas9
tetR
pspFΔHTH::λN22plus
sfgfp
rhaS
UseSynthetic BiologyTagsASV tag and λN22plusExpressionBacterialMutationMutations D10A and H840A on WT cas9. Synonymous m…PromoterPcon, PpspA-zntA-1034, PrhaB, and PtetAvailable SinceDec. 12, 2022AvailabilityAcademic Institutions and Nonprofits only -
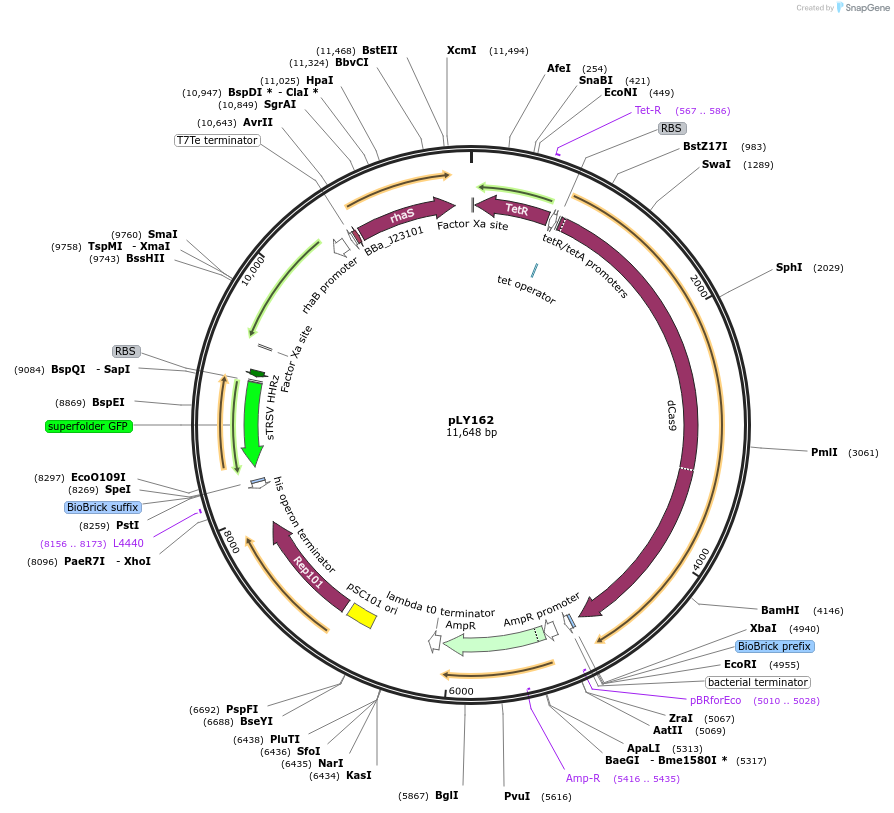
pLY162
Plasmid#184138PurposeReporter circuit for hijacking RFP mRNA as CRISPR RNA (target site 1)DepositorInsertsdCas9
tetR
pspFΔHTH::λN22plus
sfgfp
rhaS
UseSynthetic BiologyTagsASV tag and λN22plusExpressionBacterialMutationMutations D10A and H840A on WT cas9. Synonymous m…PromoterPcon, PpspA-R1, PrhaB, and PtetAvailable SinceOct. 31, 2022AvailabilityAcademic Institutions and Nonprofits only -
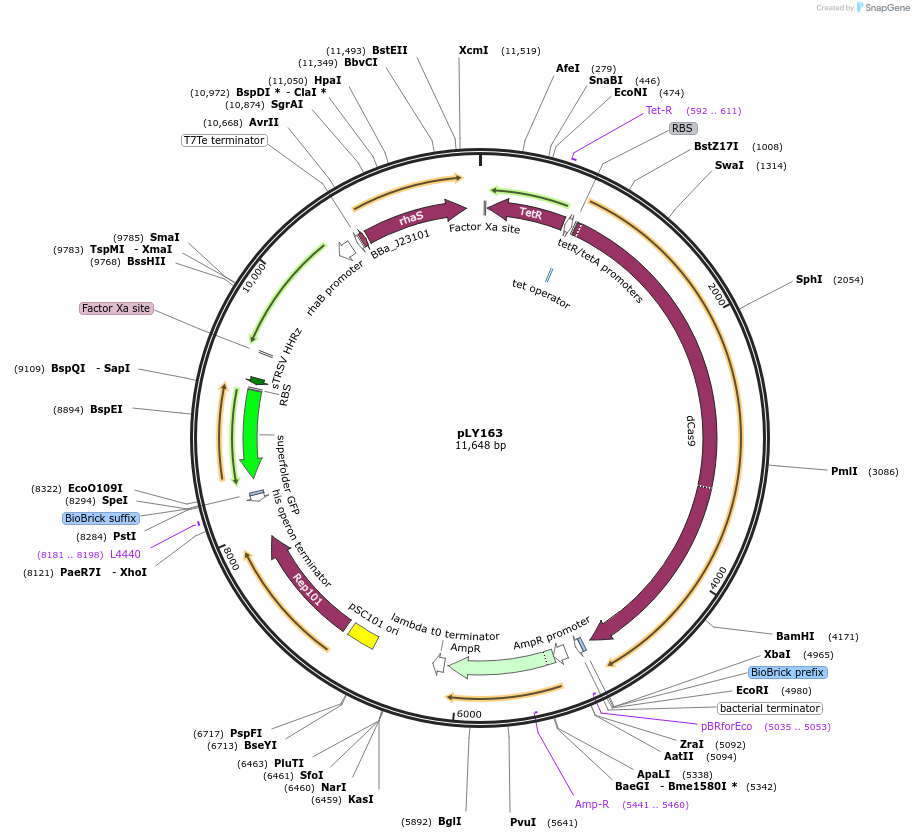
pLY163
Plasmid#184141PurposeReporter circuit for hijacking RFP mRNA as CRISPR RNA (target site 2)DepositorInsertsdCas9
tetR
pspFΔHTH::λN22plus
sfgfp
rhaS
UseSynthetic BiologyTagsASV tag and λN22plusExpressionBacterialMutationMutations D10A and H840A on WT cas9. Synonymous m…PromoterPcon, PpspA-R2, PrhaB, and PtetAvailable SinceOct. 31, 2022AvailabilityAcademic Institutions and Nonprofits only -
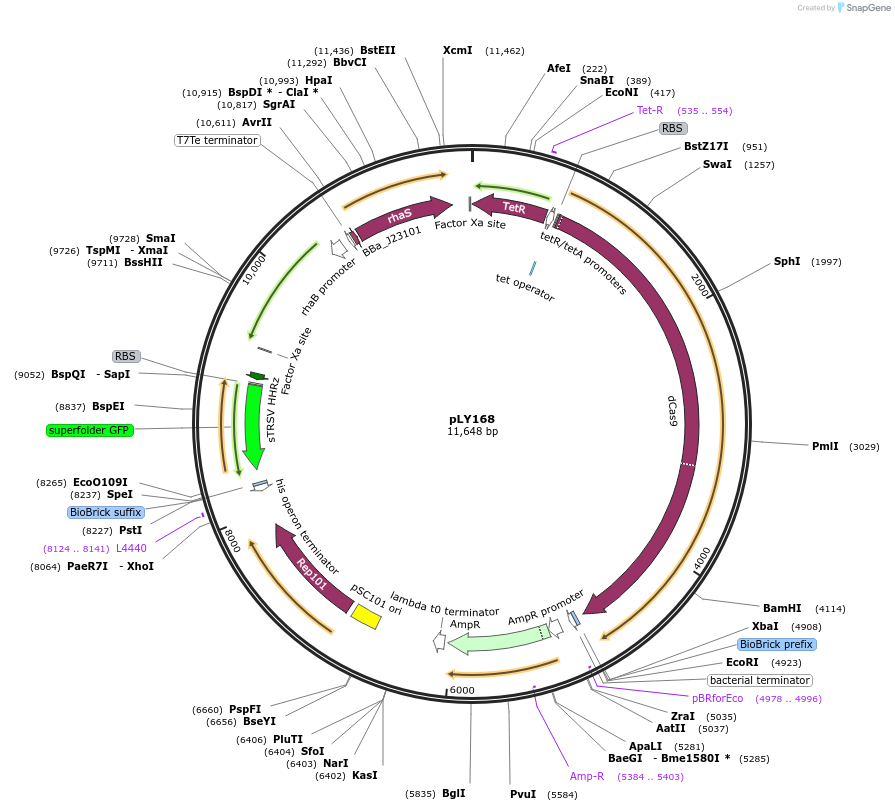
pLY168
Plasmid#184142PurposeReporter circuit for hijacking mRNA of arsenic responsive gene cluster as CRISPR RNA (site 2)DepositorInsertsdCas9
tetR
pspFΔHTH::λN22plus
sfgfp
rhaS
UseSynthetic BiologyTagsASV tag and λN22plusExpressionBacterialMutationMutations D10A and H840A on WT cas9. Synonymous m…PromoterPcon, PpspA-Ar2, PrhaB, and PtetAvailable SinceOct. 31, 2022AvailabilityAcademic Institutions and Nonprofits only -
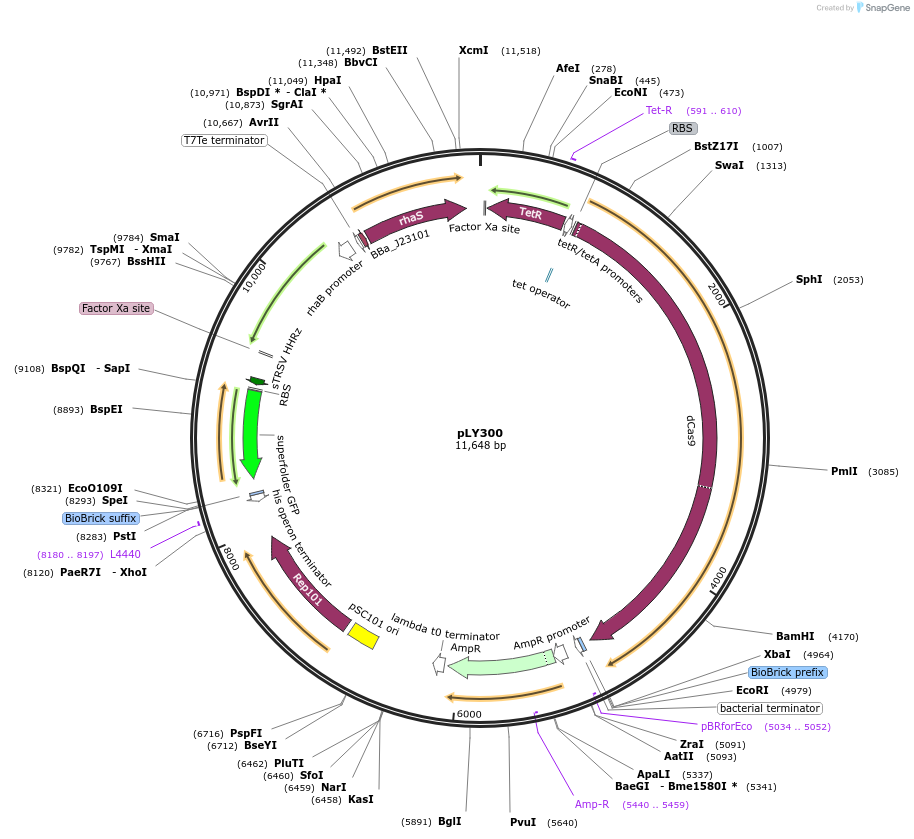
pLY300
Plasmid#184158PurposeReporter circuit for hijacking small RNA oxyS as CRISPR RNADepositorInsertsdCas9
tetR
pspFΔHTH::λN22plus
sfgfp
rhaS
UseSynthetic BiologyTagsASV tag and λN22plusExpressionBacterialMutationMutations D10A and H840A on WT cas9. Synonymous m…PromoterPcon, PpspA-oxyS-86, PrhaB, and PtetAvailable SinceOct. 31, 2022AvailabilityAcademic Institutions and Nonprofits only -
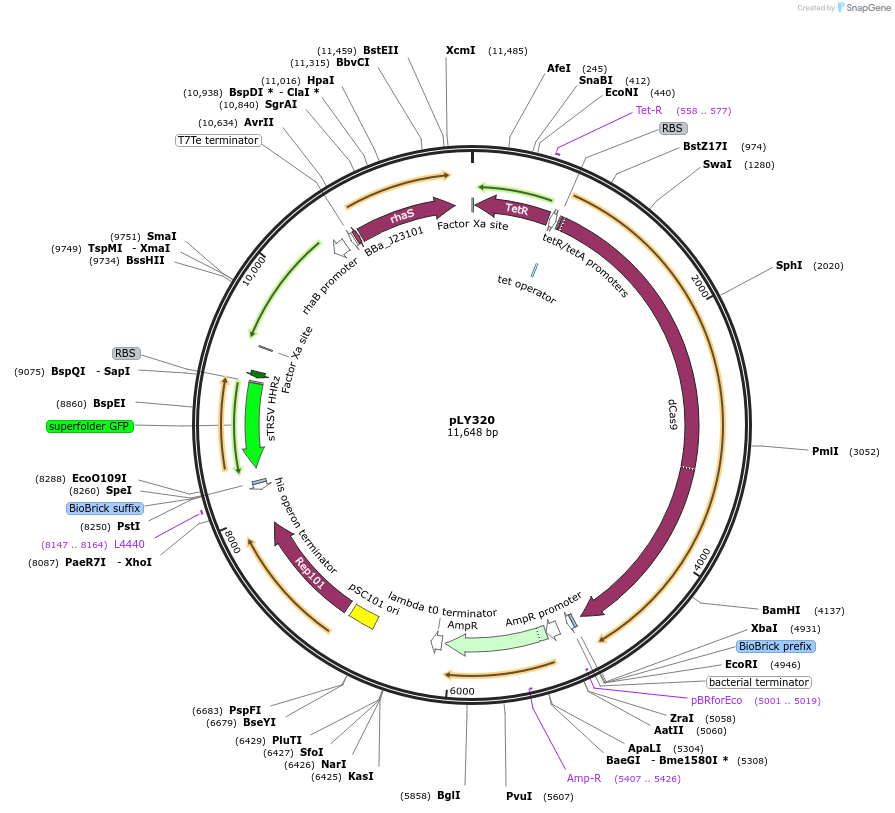
pLY320
Plasmid#184161PurposeReporter circuit for hijacking mRNA of cusCFAB as CRISPR RNADepositorInsertsdCas9
tetR
pspFΔHTH::λN22plus
sfgfp
rhaS
UseSynthetic BiologyTagsASV tag and λN22plusExpressionBacterialMutationMutations D10A and H840A on WT cas9. Synonymous m…PromoterPcon, PpspA-cusCFAB-656, PrhaB, and PtetAvailable SinceOct. 31, 2022AvailabilityAcademic Institutions and Nonprofits only -
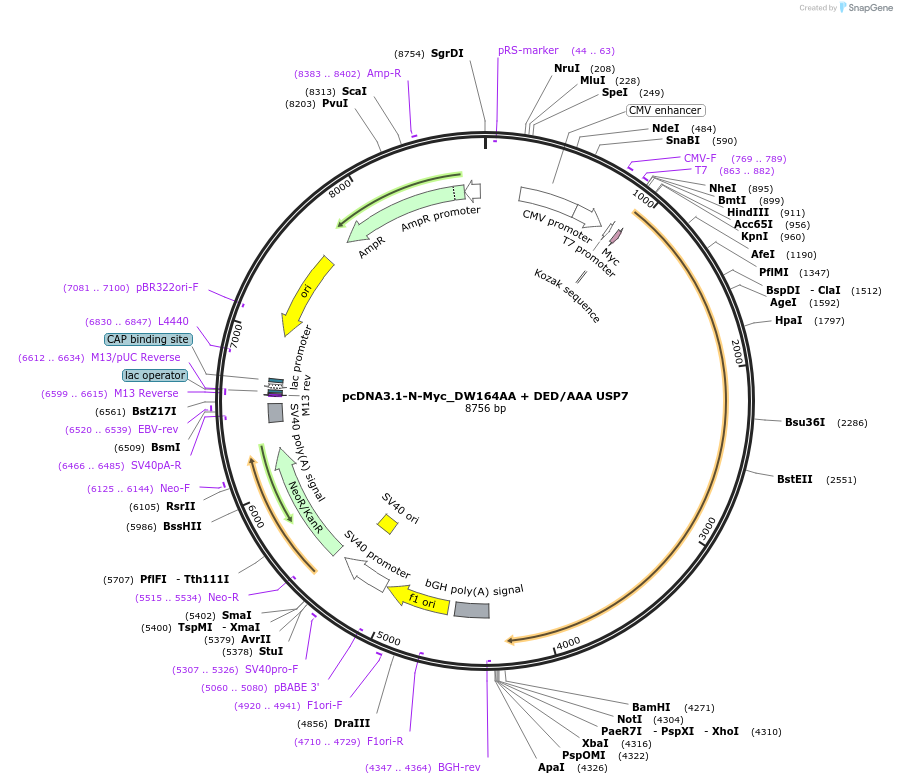
pcDNA3.1-N-Myc_DW164AA + DED/AAA USP7
Plasmid#131256PurposeMammalian expression of N-terminally Myc-tagged USP7, with mutations in the TRAF-like domain (DW164AA) and UBL2 (DE758AA_D764A)DepositorInsertUbiquitin carboxyl-terminal hydrolase 7 (USP7 Human)
TagsMycExpressionMammalianMutationContains point mutations that convert USP7 aspart…Available SinceDec. 17, 2020AvailabilityAcademic Institutions and Nonprofits only -
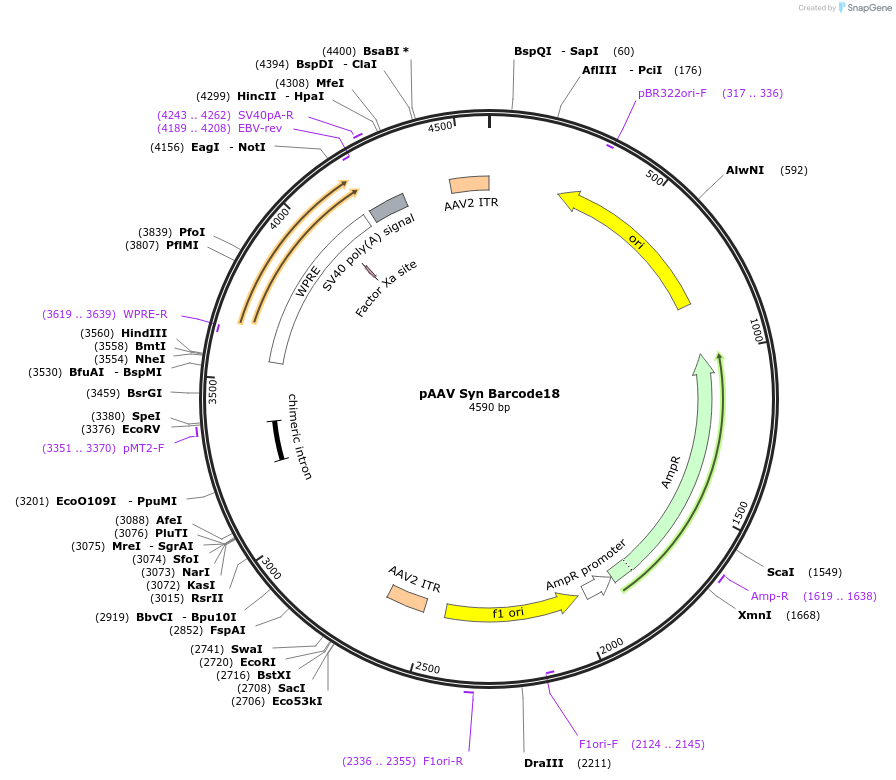
pAAV Syn Barcode18
Plasmid#226191PurposeExpression mappingDepositorInsertSyn Barcode18
UseAAVAvailable SinceOct. 30, 2024AvailabilityAcademic Institutions and Nonprofits only -
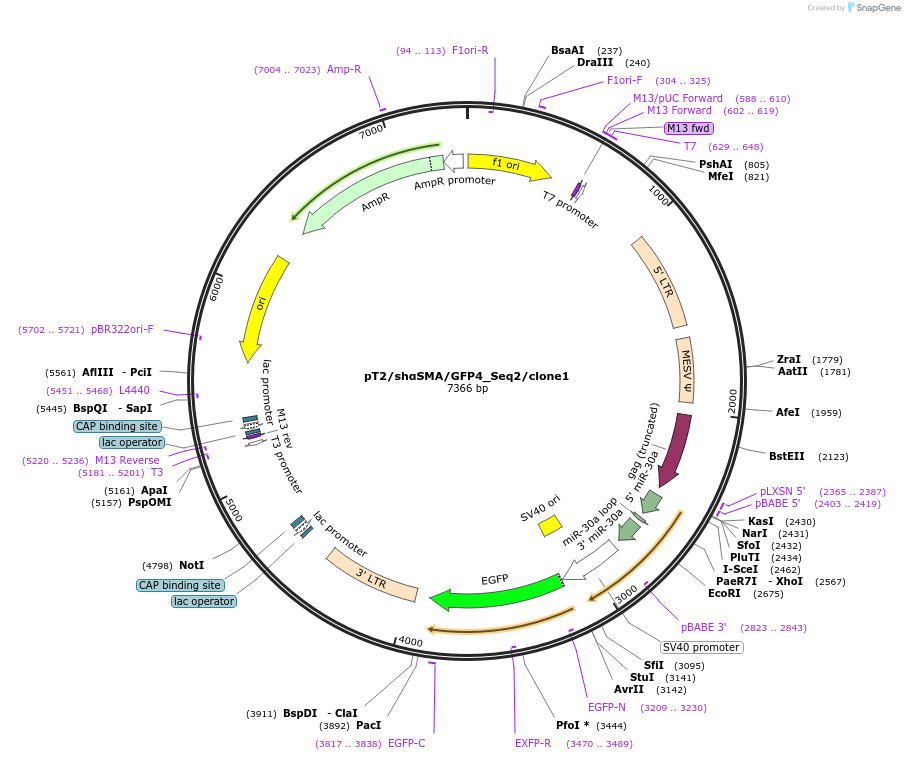
pT2/shαSMA/GFP4_Seq2/clone1
Plasmid#201402PurposeKnockdown of alpha-SMA. Construct has inverted repeats to be used in Sleeping Beauty transposon system.DepositorInsertalpha-SMA
ExpressionMammalianAvailable SinceMay 24, 2023AvailabilityAcademic Institutions and Nonprofits only -
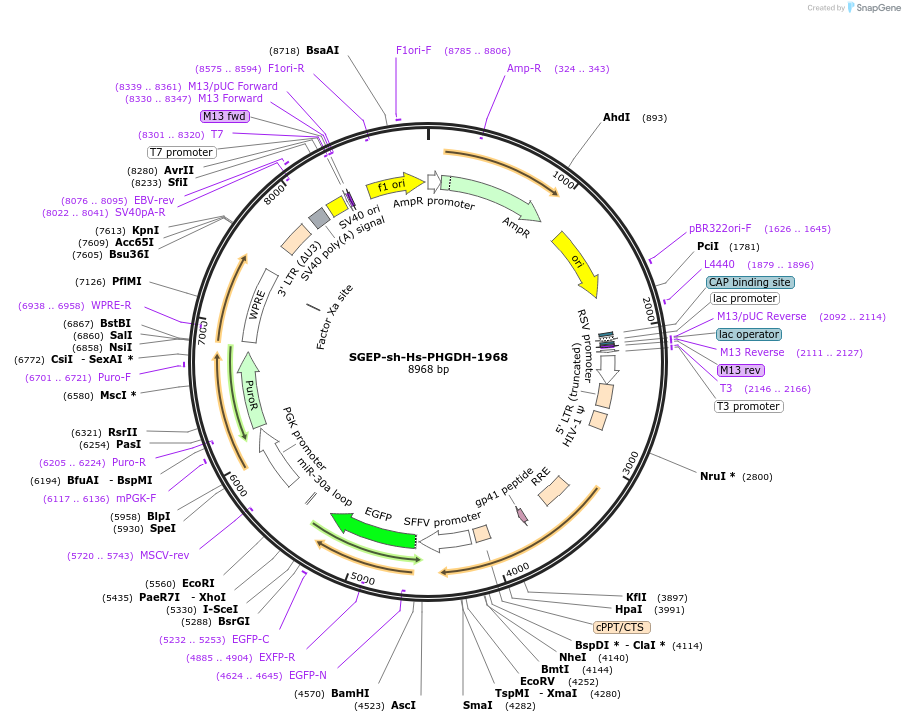
SGEP-sh-Hs-PHGDH-1968
Plasmid#188673PurposeshRNADepositorAvailable SinceOct. 21, 2022AvailabilityAcademic Institutions and Nonprofits only -
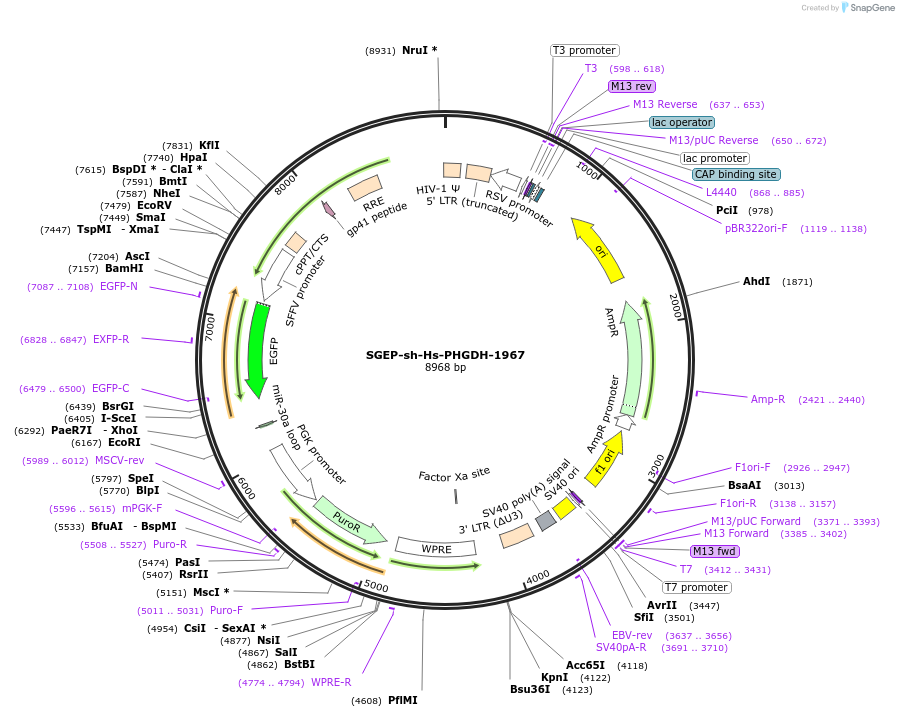
SGEP-sh-Hs-PHGDH-1967
Plasmid#188672PurposeshRNADepositorAvailable SinceOct. 20, 2022AvailabilityAcademic Institutions and Nonprofits only -
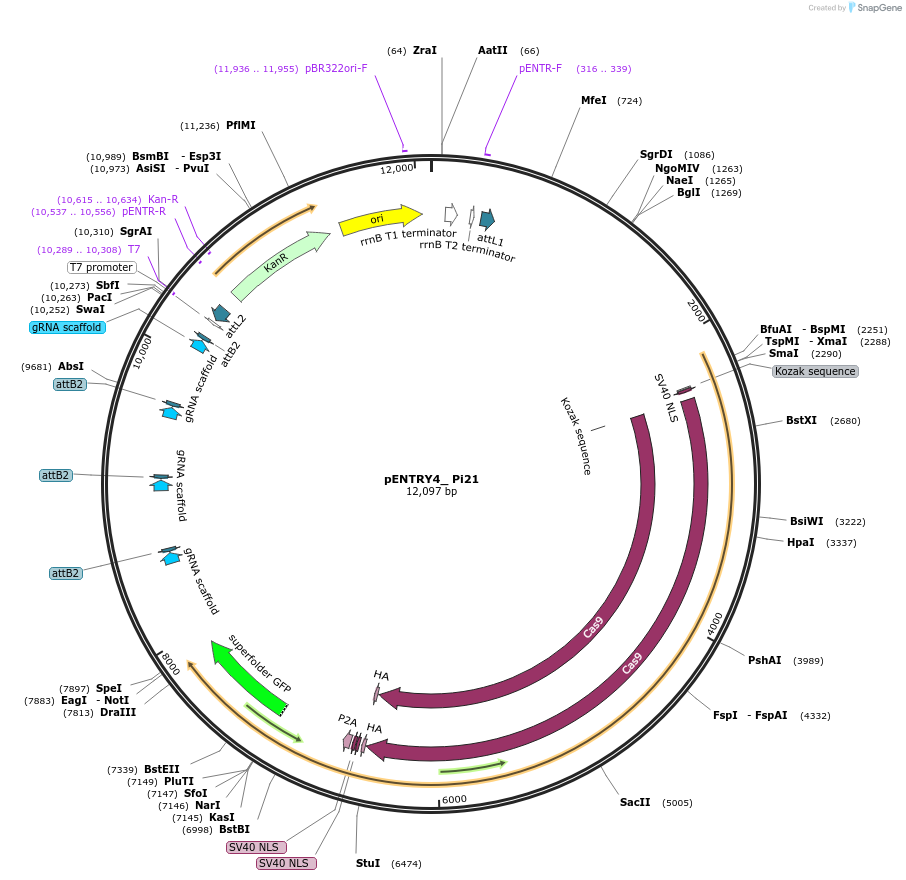
pENTRY4_ Pi21
Plasmid#126904PurposeGateway pENTR-plasmid to generate a binary construct for genome editing of the wheat homolog of a susceptibility gene of O. sativa. Encodes wheat optimized Cas9 and gene specific sgRNA modules.DepositorInsertWheat_live_Cas9
UseCRISPRAvailable SinceJuly 12, 2019AvailabilityAcademic Institutions and Nonprofits only -
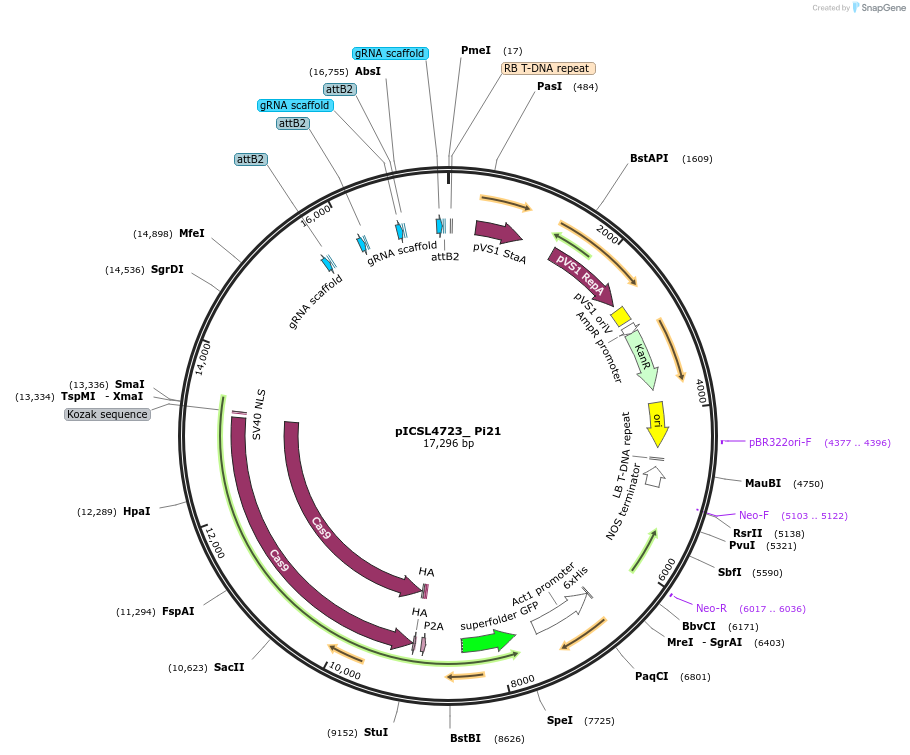
pICSL4723_ Pi21
Plasmid#126892PurposeGenome editing of the wheat homolog of a susceptibility gene of O. sativa. Encodes wheat optimized Cas9 module, nptII resistance gene, and specific sgRNA modules on the binary plasmid pICSL4723.DepositorInsertWheat_live_Cas9
UseCRISPRAvailable SinceJuly 24, 2019AvailabilityAcademic Institutions and Nonprofits only -
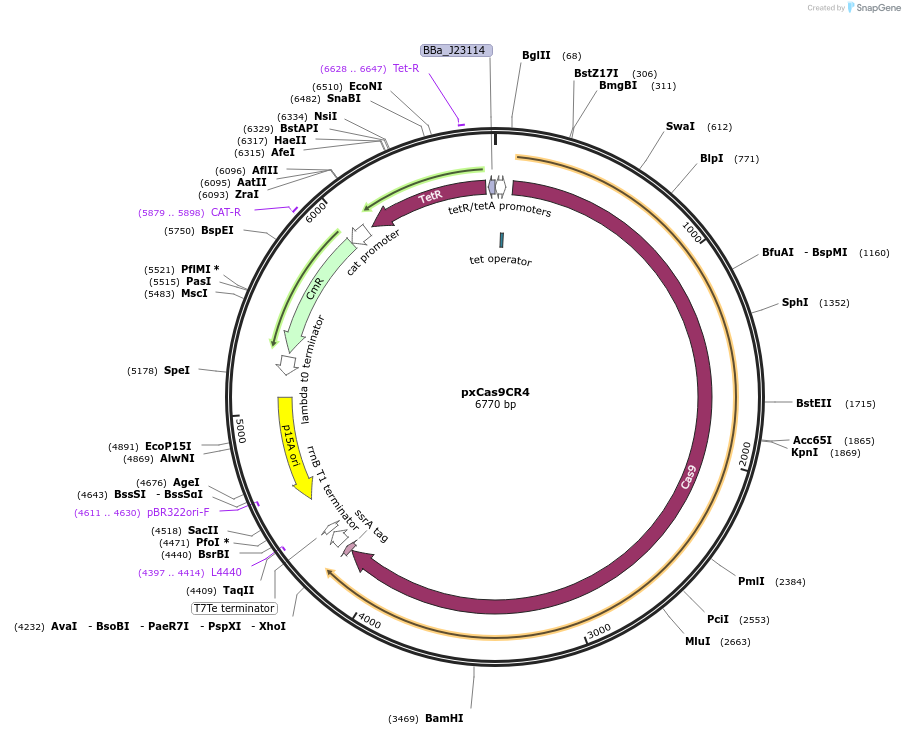
pxCas9CR4
Plasmid#111656PurposePCas9CR4 with the xCas9 mutations allowing NG PAM sitesDepositorInsertTetR and xCas9
UseSynthetic BiologyTagsssrAExpressionBacterialMutationxCas9 mutationsAvailable SinceJuly 30, 2018AvailabilityAcademic Institutions and Nonprofits only -
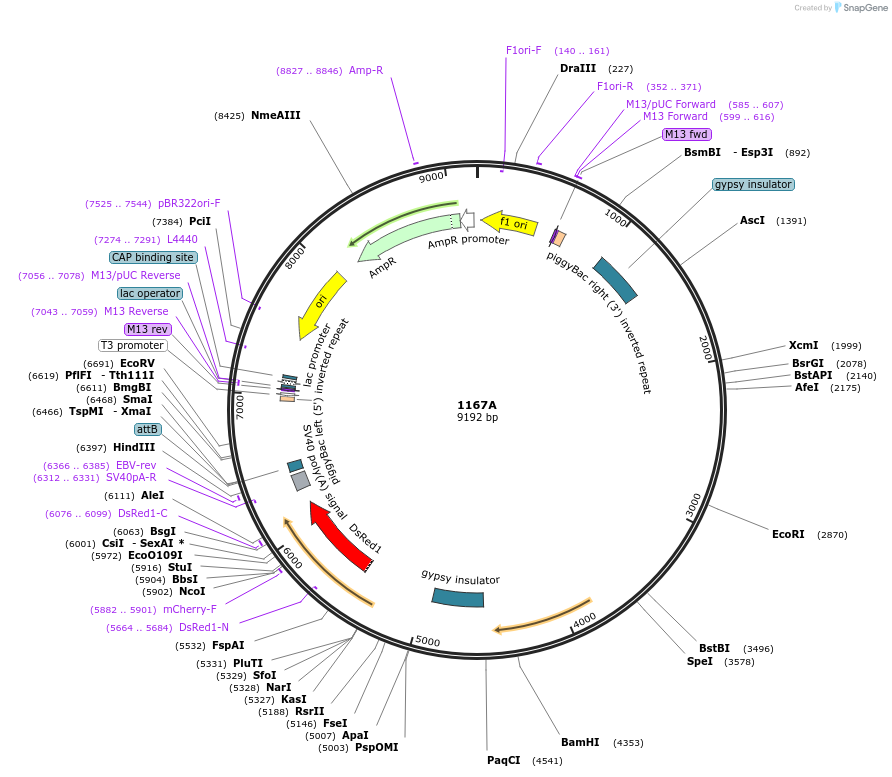
1167A
Plasmid#218233PurposeHDR plasmid for inserting Ceratitis capitata transformer female intron into the white pupae geneDepositorInsertputative metabolite transport protein
ExpressionInsectAvailable SinceMay 16, 2024AvailabilityAcademic Institutions and Nonprofits only -
pCAR-IF reporter
Plasmid#16628DepositorInsertCAR-IF
Tagsbeta-galactosidaseExpressionMammalianAvailable SinceMarch 12, 2008AvailabilityAcademic Institutions and Nonprofits only -
pCAR-OF reporter
Plasmid#16627DepositorInsertCAR-OF
Tagsbeta-galactosidaseExpressionMammalianAvailable SinceMarch 12, 2008AvailabilityAcademic Institutions and Nonprofits only -
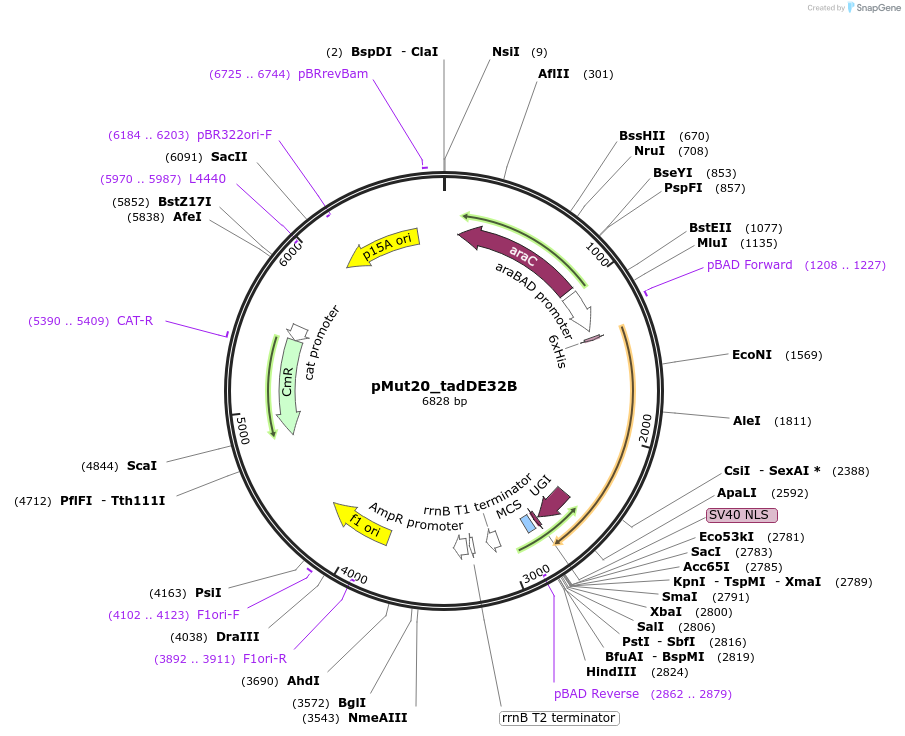
pMut20_tadDE32B
Plasmid#247074PurposeL-arabinose inducible TadDE-XTEN32-CasB(Cas11)-GSSG-UGIDepositorInsertTadDE-XTEN16-CasB-UGI
UseCRISPRExpressionBacterialAvailable SinceNov. 4, 2025AvailabilityAcademic Institutions and Nonprofits only -
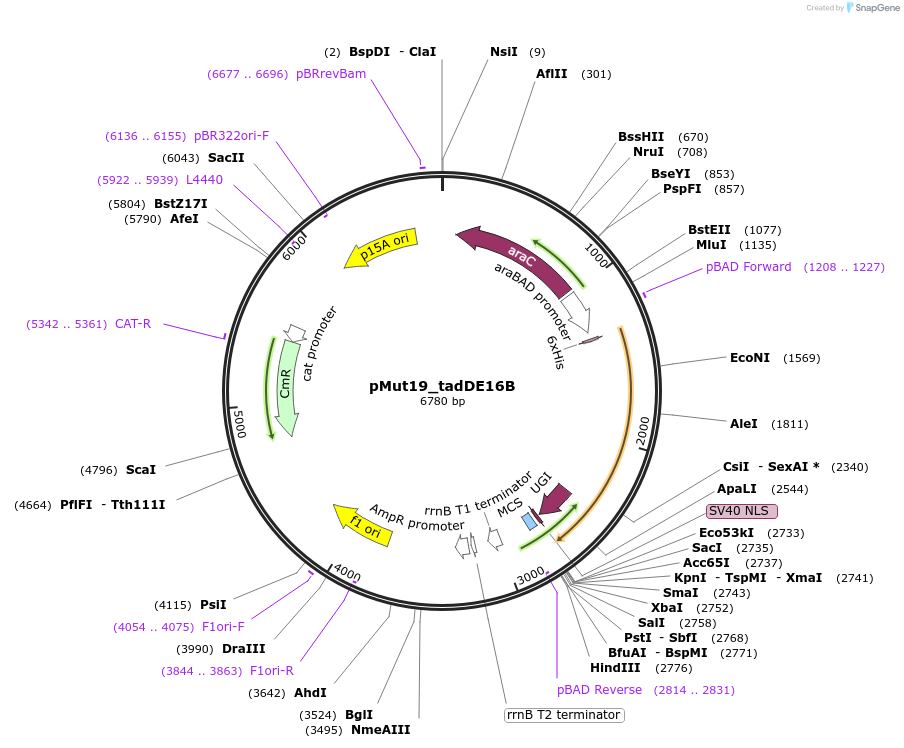
pMut19_tadDE16B
Plasmid#247073PurposeL-arabinose inducible TadDE-XTEN16-CasB(Cas11)-GSSG-UGIDepositorInsertTadDE-XTEN16-CasB-UGI
UseCRISPRExpressionBacterialAvailable SinceNov. 4, 2025AvailabilityAcademic Institutions and Nonprofits only -
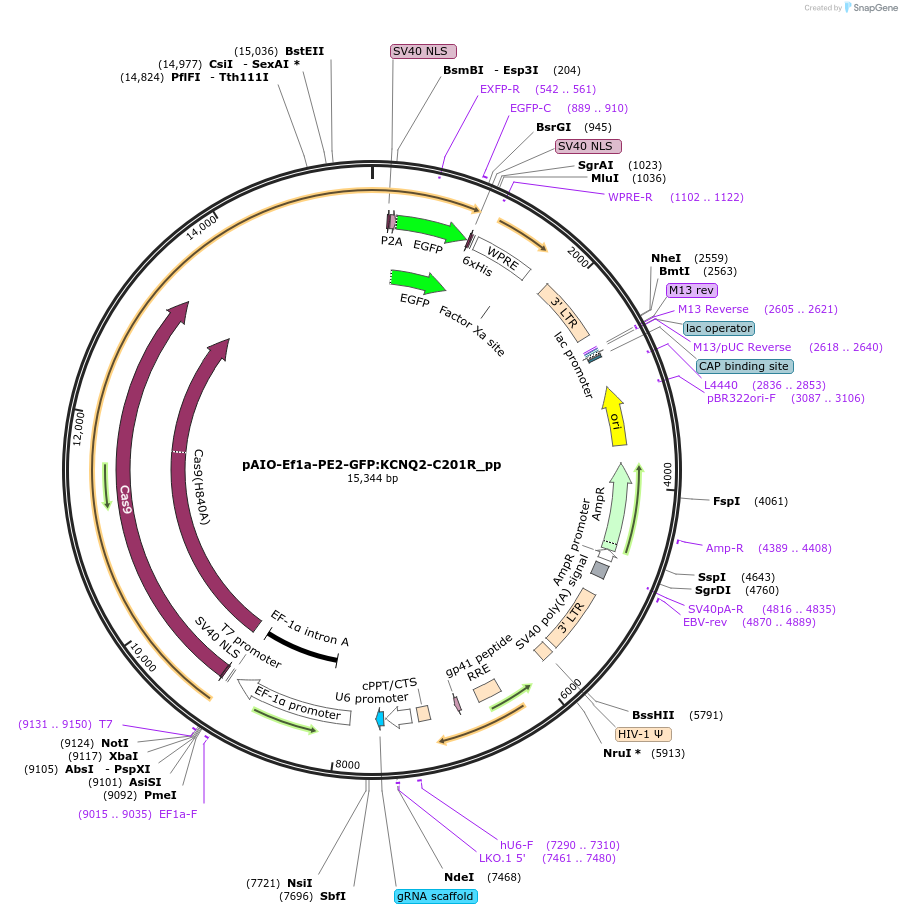
pAIO-Ef1a-PE2-GFP:KCNQ2-C201R_pp
Plasmid#185061PurposeEf1a driven PE2 with pegRNA for editing C201R mutation (including silent PAM mutation) in KCNQ2. See Addgene plasmid #184445DepositorInsertU6:pegRNA:scaffold:PBS+RT template
UseCRISPR and LentiviralExpressionMammalianPromoterU6Available SinceNov. 7, 2022AvailabilityAcademic Institutions and Nonprofits only -
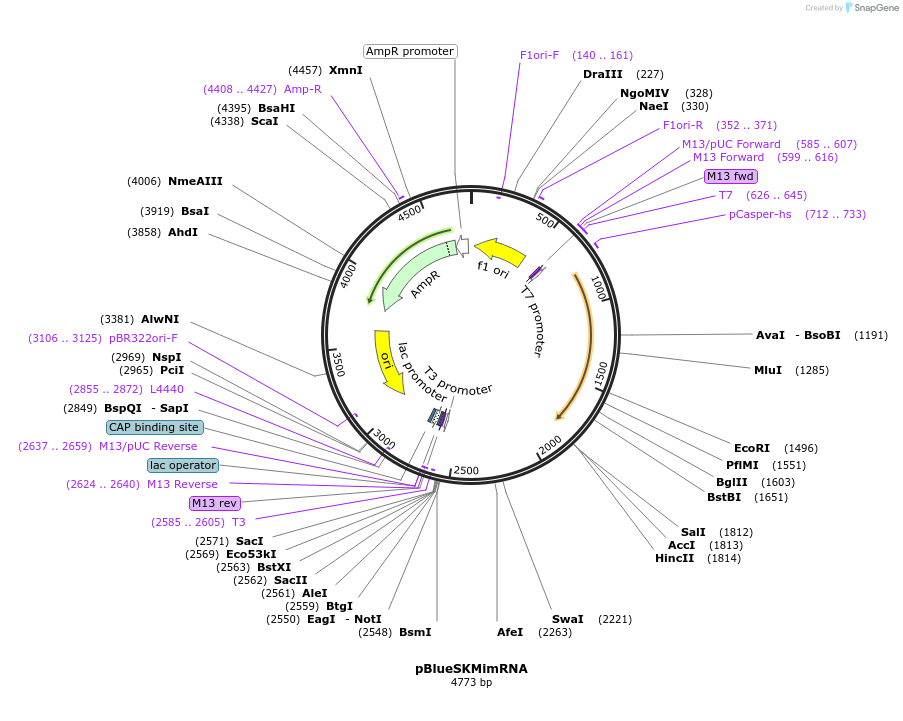
pBlueSKMimRNA
Plasmid#102535PurposeTemplate for synthesis of Minos transposase mRNADepositorInsertMinos transposase mRNA
Available SinceApril 13, 2018AvailabilityAcademic Institutions and Nonprofits only -
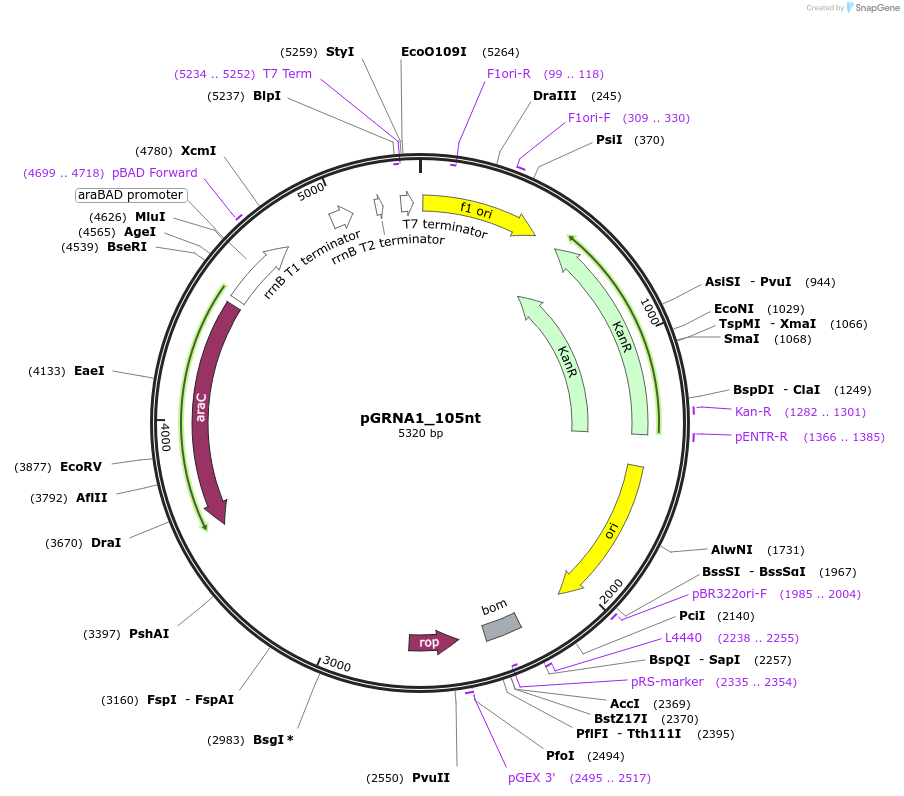
pGRNA1_105nt
Plasmid#247075PurposeL-arabinose inducible crRNA of type I-E CRISPR cas (Escherichia coli K-12). encoding 105 nt spacer which targets pheS*_codonDepositorInserttype I-E CRISPR cas crRNA
UseCRISPRExpressionBacterialAvailable SinceNov. 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
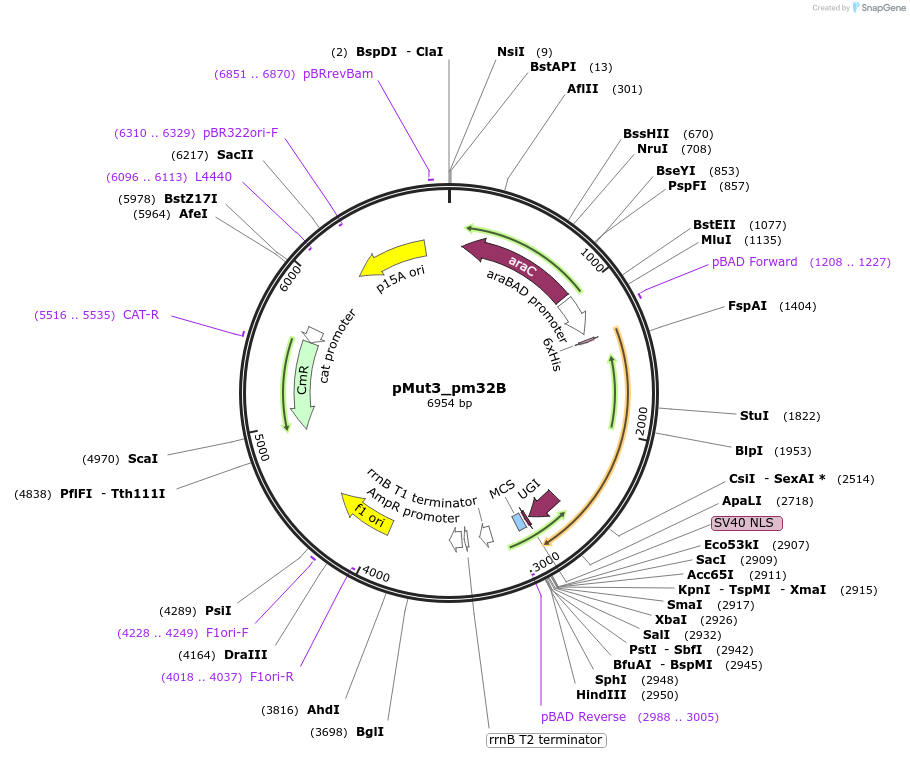
pMut3_pm32B
Plasmid#247071PurposeL-arabinose inducible pmCDA-XTEN32-CasB(Cas11)-GSSG-UGIDepositorInsertPmCDA-XTEN32-Cas11(CasB)-UGI
UseCRISPRExpressionBacterialAvailable SinceNov. 4, 2025AvailabilityAcademic Institutions and Nonprofits only -
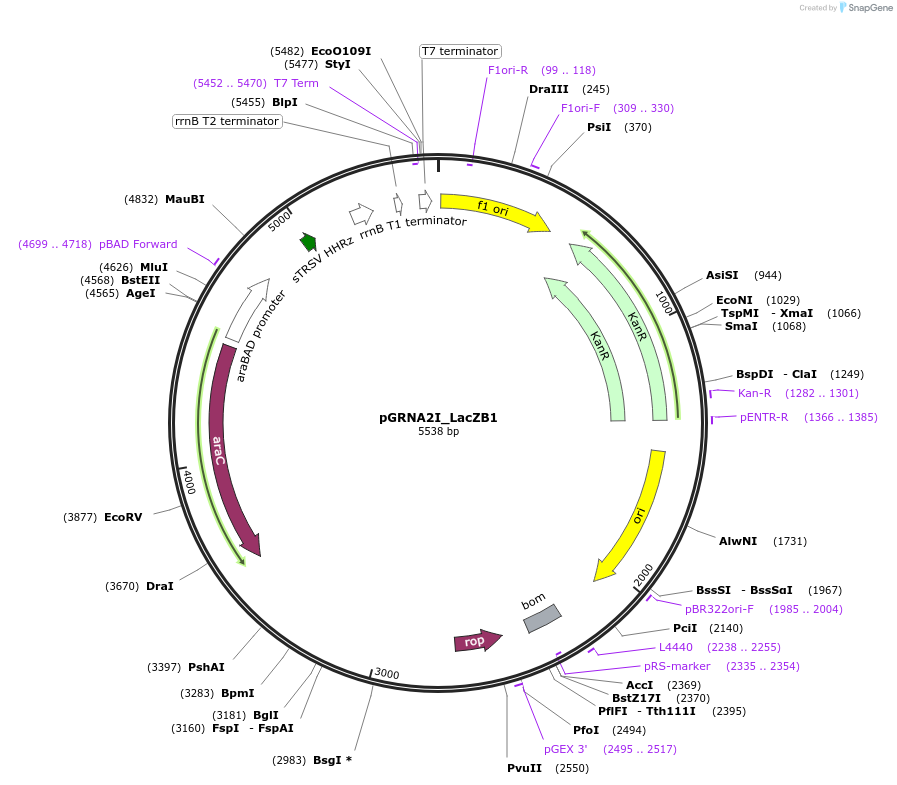
pGRNA2I_LacZB1
Plasmid#247076PurposeL-arabinose inducible crRNA of type I-E CRISPR cas (Escherichia coli K-12). encoding 105 nt spacer which targets Ecoli LacZ B1 siteDepositorInserttype I-E CRISPR cas crRNA
UseCRISPRExpressionBacterialAvailable SinceNov. 4, 2025AvailabilityAcademic Institutions and Nonprofits only -
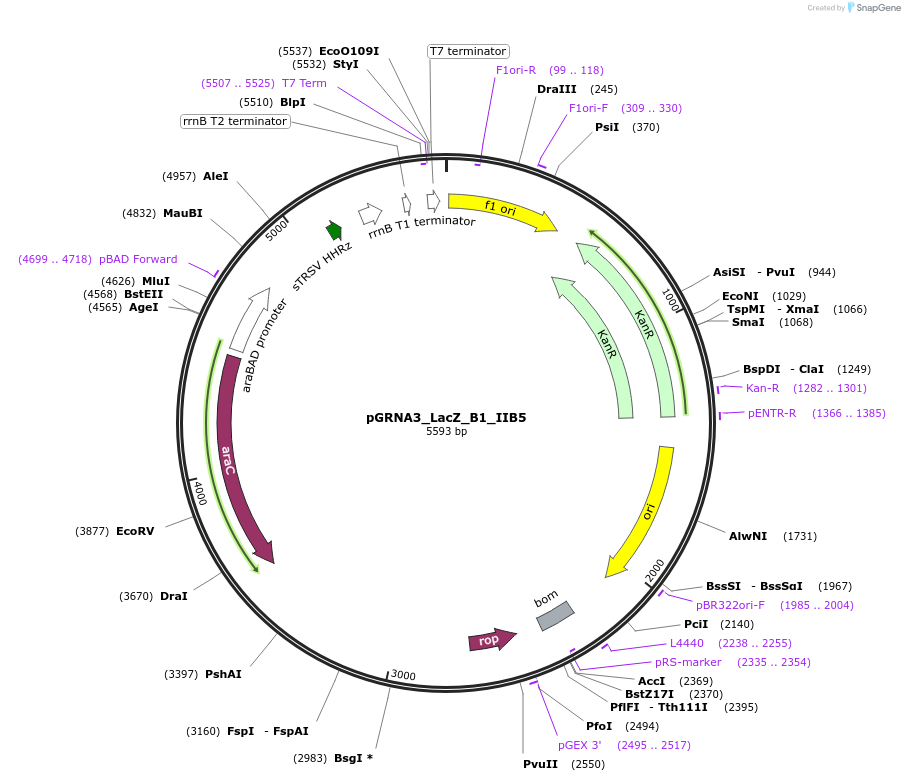
pGRNA3_LacZ_B1_IIB5
Plasmid#247081PurposeL-arabinose inducible crRNA of type I-E CRISPR cas (Escherichia coli K-12). encoding 105 nt spacer which targets both Ecoli LacZ B1 and B5 siteDepositorInserttype I-E CRISPR cas crRNA
UseCRISPRExpressionBacterialAvailable SinceNov. 4, 2025AvailabilityAcademic Institutions and Nonprofits only