We narrowed to 10,090 results for: Coli
-
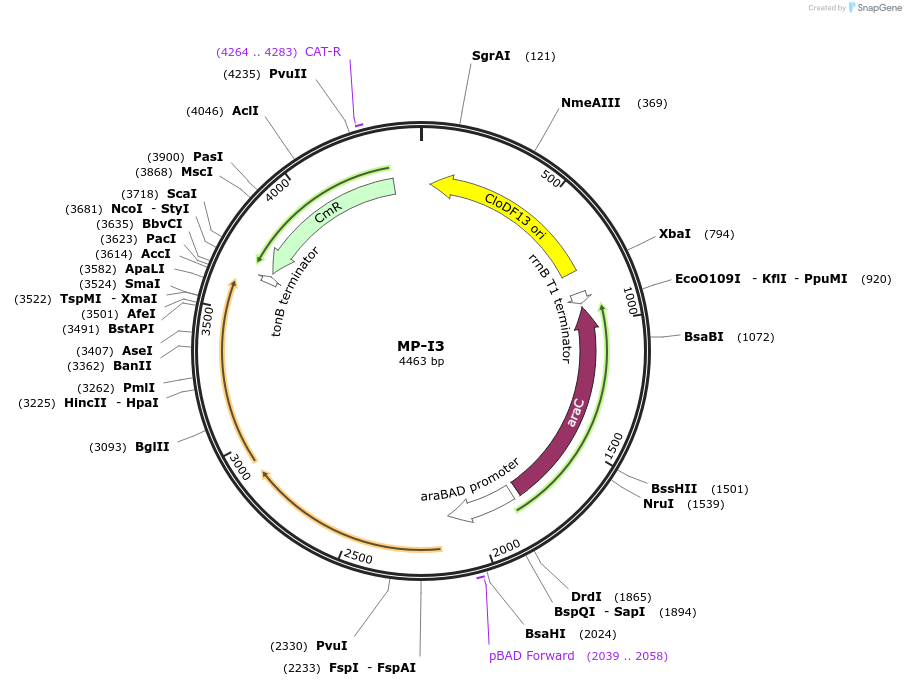
Plasmid#69646PurposeUntargetted Bacterial MutagenesisDepositorInsertdnaQ926 mutH(K116E)
ExpressionBacterialPromoterE. coli PBADAvailable SinceOct. 8, 2015AvailabilityAcademic Institutions and Nonprofits only -
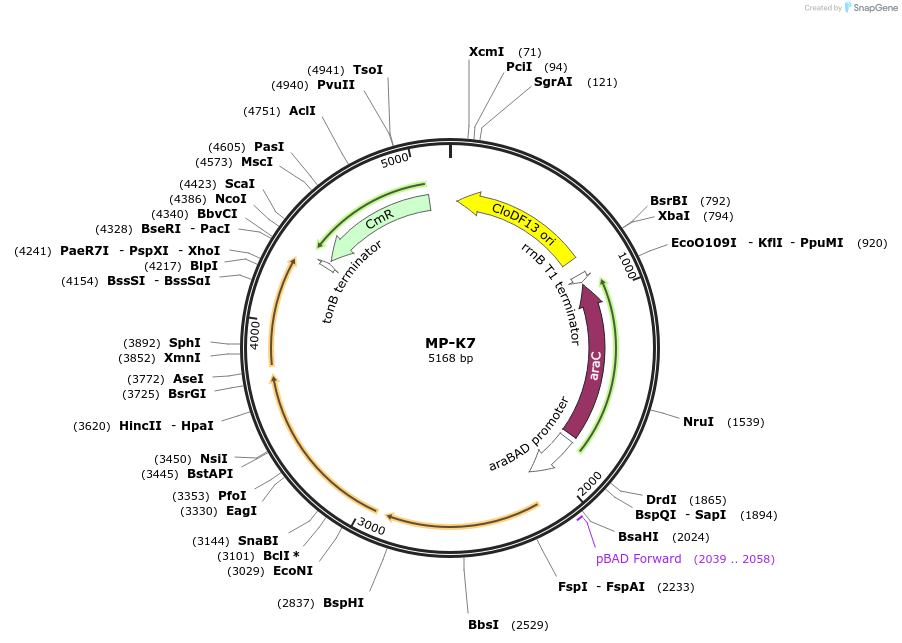
MP-K7
Plasmid#69651PurposeUntargetted Bacterial MutagenesisDepositorInsertdnaQ926 dam emrR
ExpressionBacterialPromoterE. coli PBADAvailable SinceOct. 8, 2015AvailabilityAcademic Institutions and Nonprofits only -
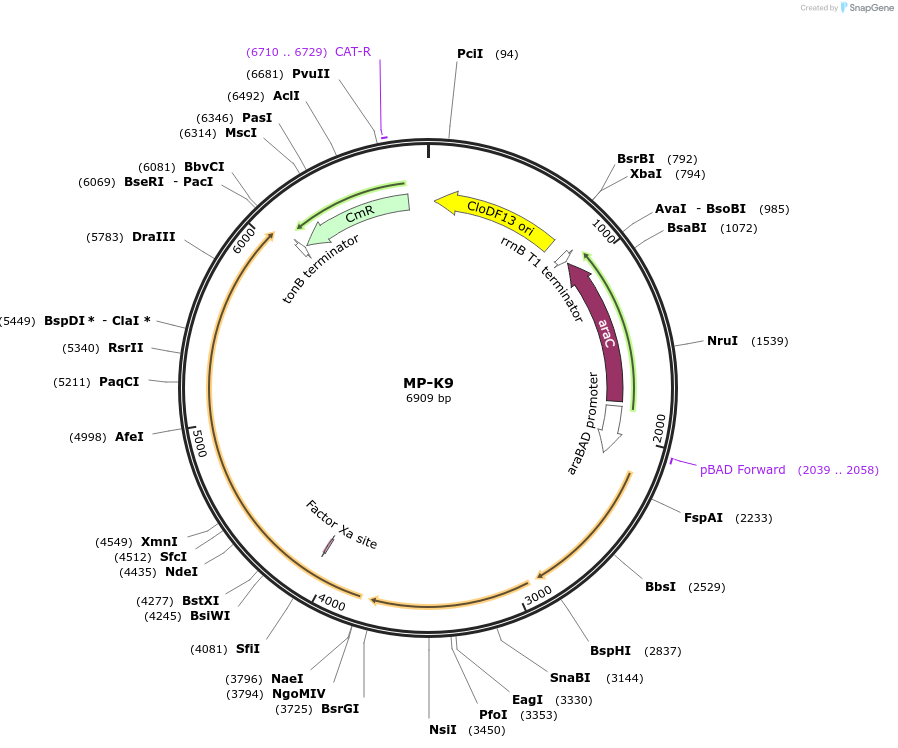
MP-K9
Plasmid#69653PurposeUntargetted Bacterial MutagenesisDepositorInsertdnaQ926 dam mutSdN
ExpressionBacterialPromoterE. coli PBADAvailable SinceOct. 8, 2015AvailabilityAcademic Institutions and Nonprofits only -
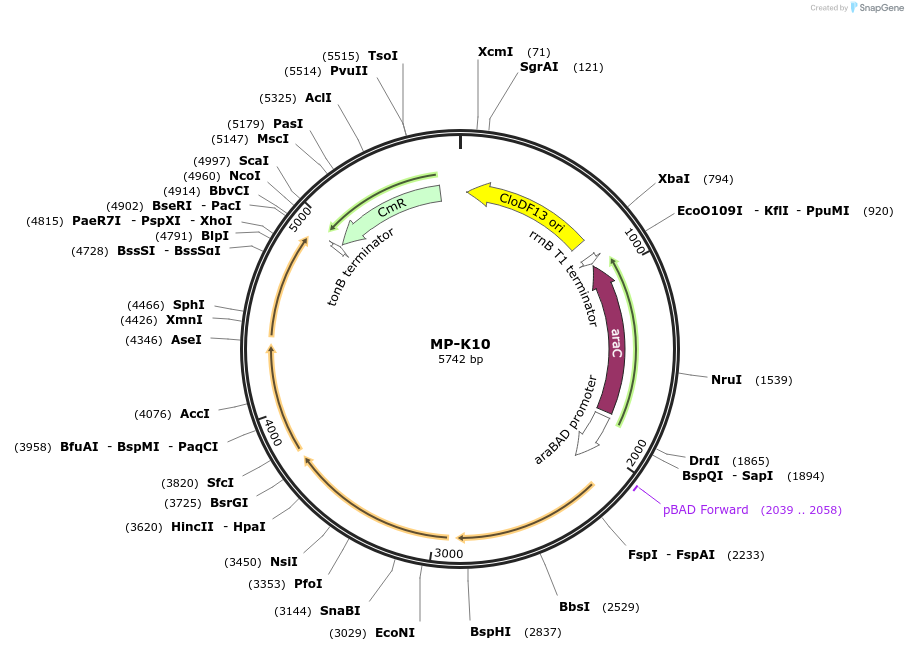
MP-K10
Plasmid#69654PurposeUntargetted Bacterial MutagenesisDepositorInsertdnaQ926 dam seqA emrR
ExpressionBacterialPromoterE. coli PBADAvailable SinceOct. 8, 2015AvailabilityAcademic Institutions and Nonprofits only -
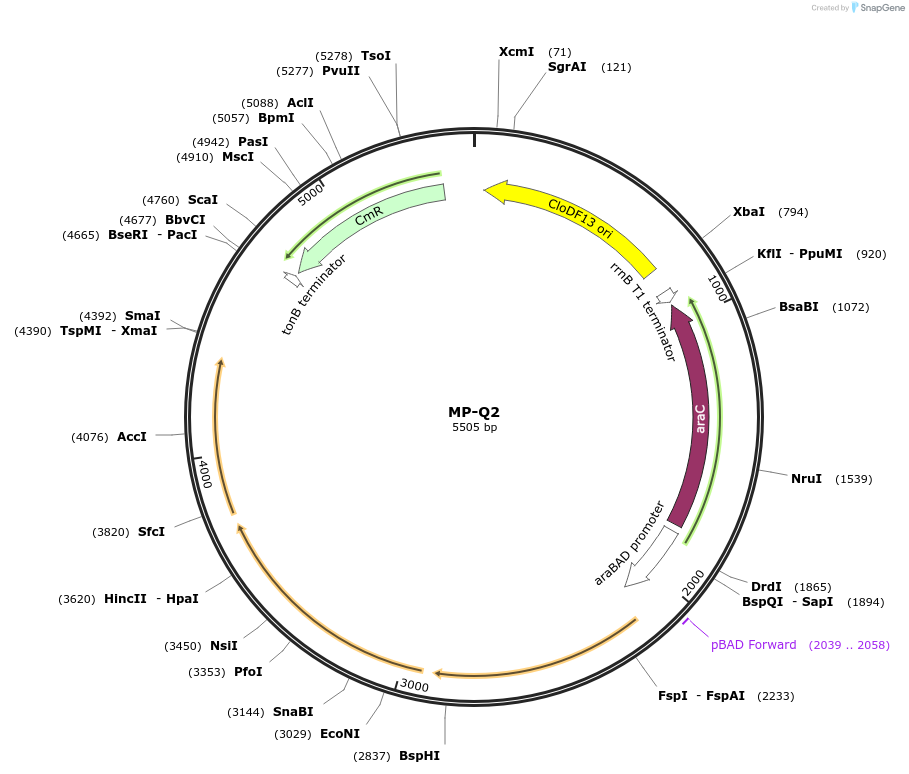
MP-Q2
Plasmid#69672PurposeUntargetted Bacterial MutagenesisDepositorInsertdnaQ926 dam seqA cchA
ExpressionBacterialPromoterE. coli PBADAvailable SinceOct. 2, 2015AvailabilityAcademic Institutions and Nonprofits only -
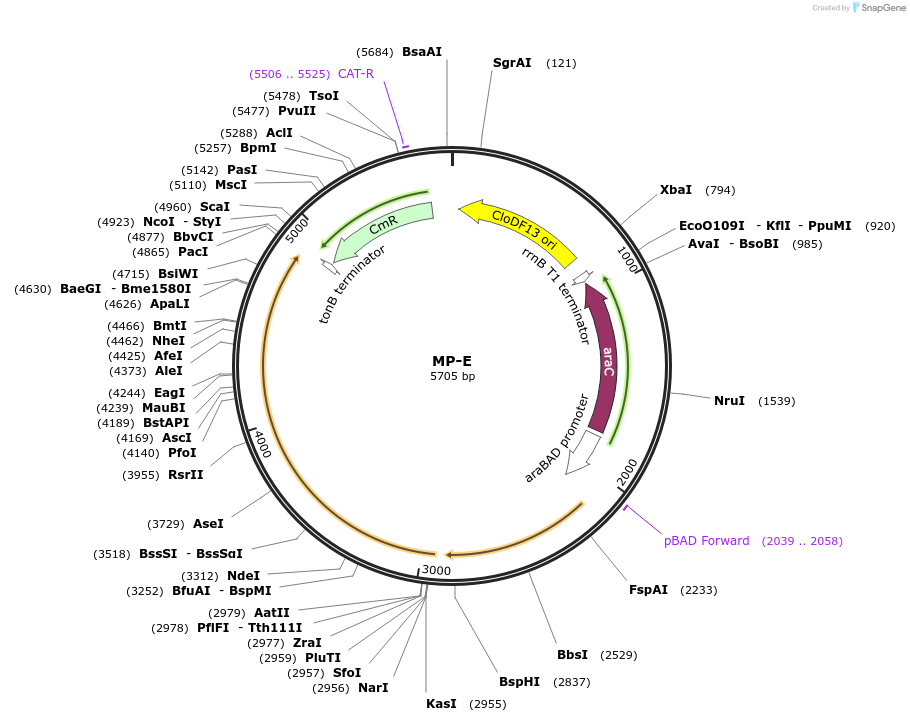
MP-E
Plasmid#69636PurposeUntargetted Bacterial MutagenesisDepositorInsertdnaQ926 dnaX36
ExpressionBacterialPromoterE. coli PBADAvailable SinceSept. 21, 2015AvailabilityAcademic Institutions and Nonprofits only -
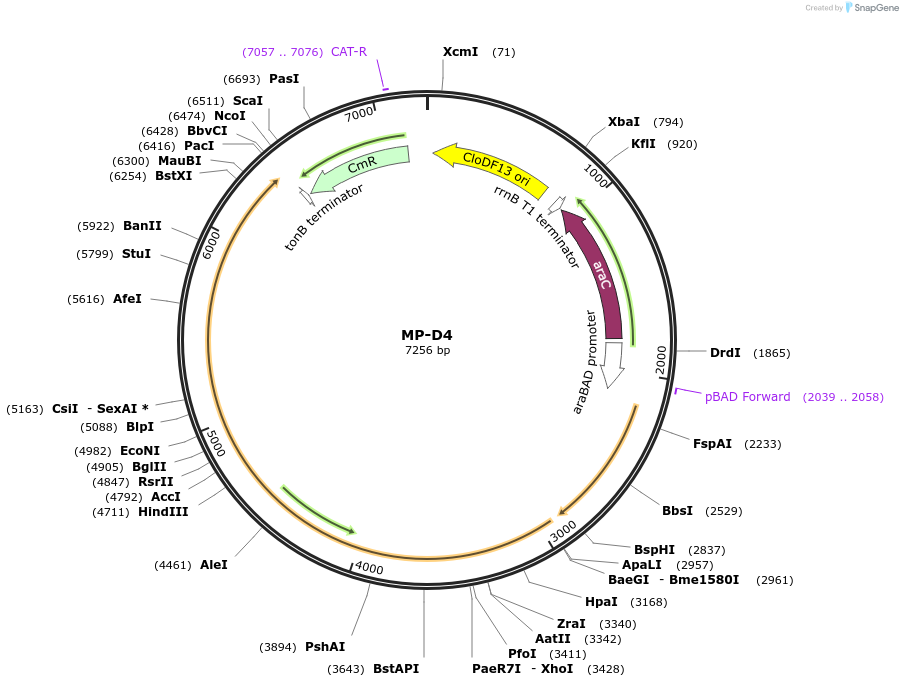
MP-D4
Plasmid#69635PurposeUntargetted Bacterial MutagenesisDepositorInsertdnaQ926 dnaE1026
ExpressionBacterialPromoterE. coli PBADAvailable SinceSept. 21, 2015AvailabilityAcademic Institutions and Nonprofits only -
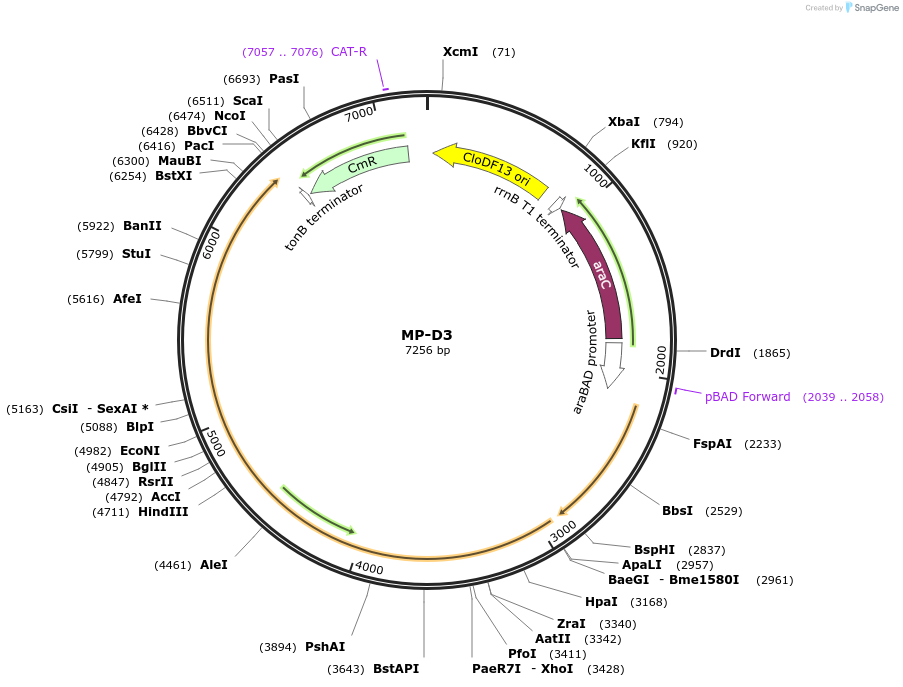
MP-D3
Plasmid#69634PurposeUntargetted Bacterial MutagenesisDepositorInsertdnaQ926 dnaE486
ExpressionBacterialPromoterE. coli PBADAvailable SinceSept. 21, 2015AvailabilityAcademic Institutions and Nonprofits only -
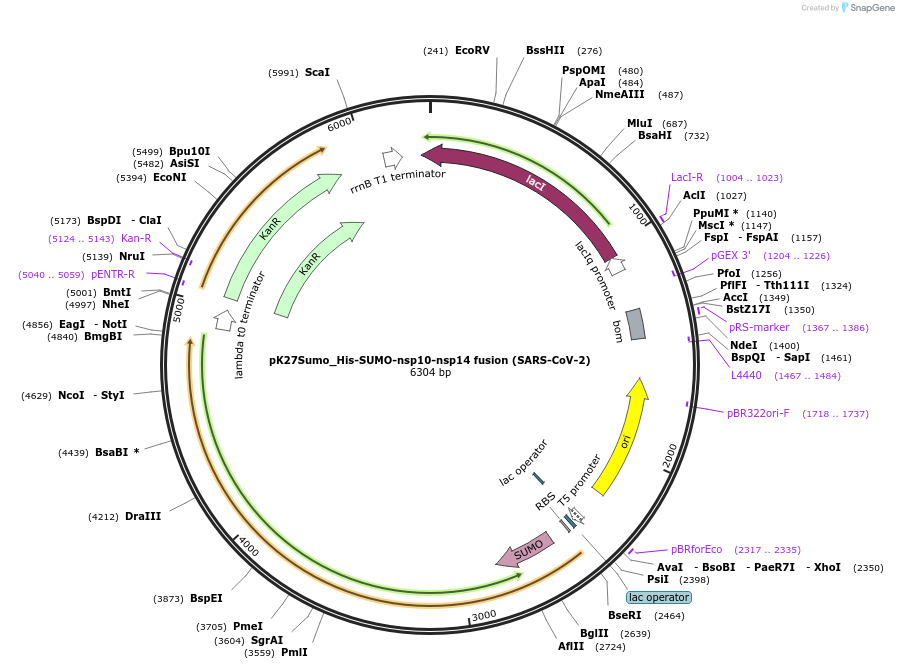
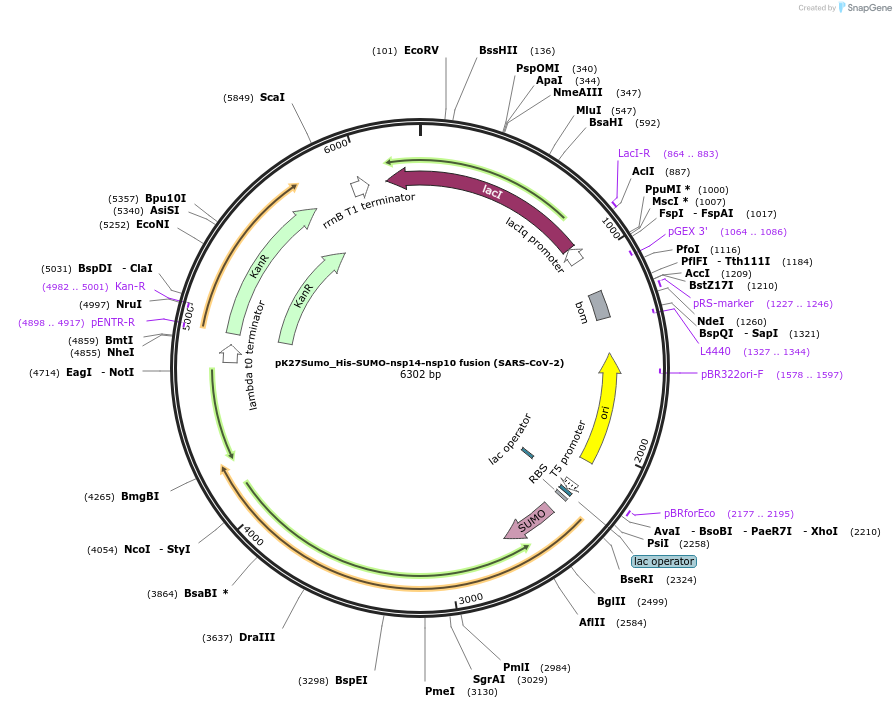
pK27Sumo_His-SUMO-nsp10-nsp14 fusion (SARS-CoV-2)
Plasmid#169161PurposeTo express nsp10-nsp14 fusion protein in E. coliDepositorInsert14His-SUMO-nsp10-GGSGGS-nsp14 (ORF1ab Synthetic, SARS-CoV-2)
Tags14His-SUMO and Fusion protein of SARS-CoV-2 nsp10…ExpressionBacterialMutationCodon optimised for E. coliPromoterT5Available SinceJuly 6, 2021AvailabilityAcademic Institutions and Nonprofits only -
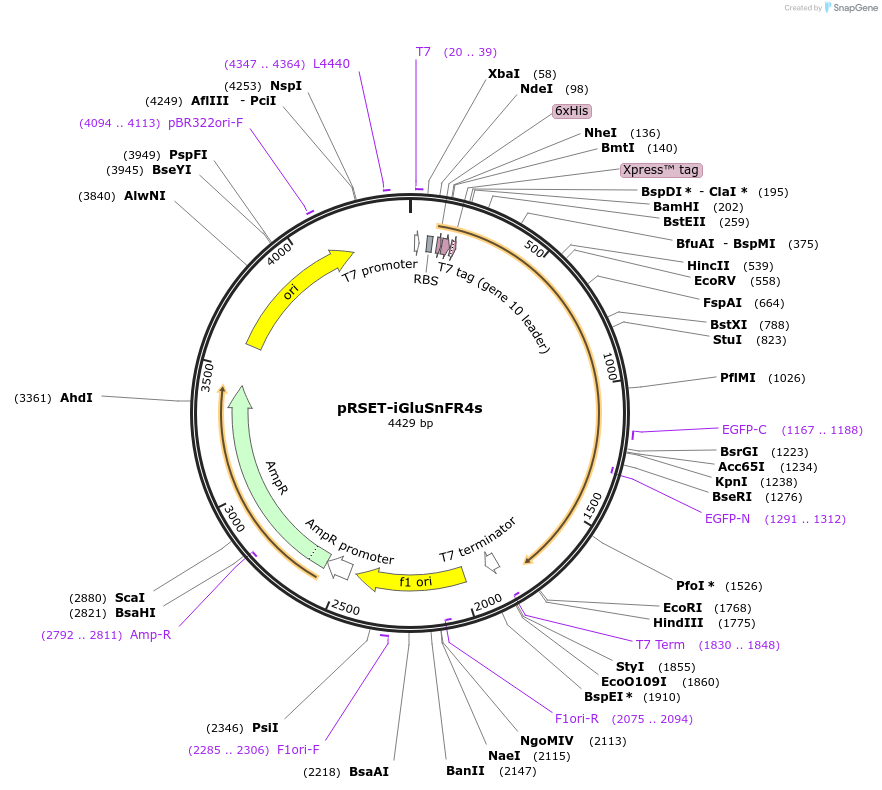
pRSET-iGluSnFR4s
Plasmid#234443PurposeBacterial expression of glutamate sensor with improved sensitivity and slow deactivation kinetics (4s = slow)DepositorInsertiGluSnFR4s
Tags6xHis-Xpress epiExpressionBacterialPromoterT7Available SinceMarch 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
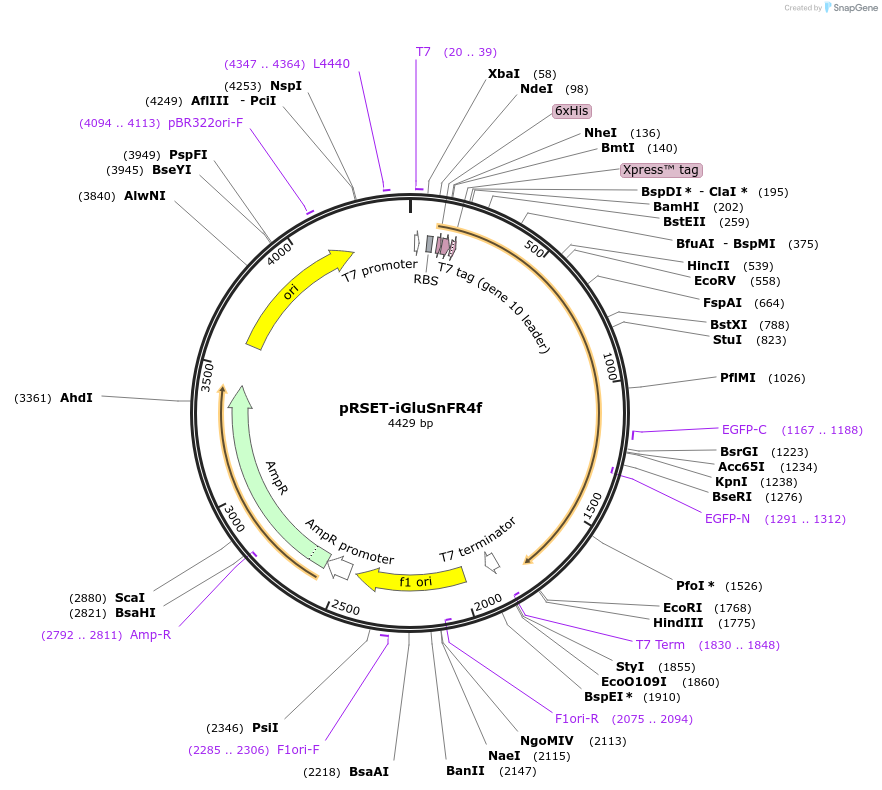
pRSET-iGluSnFR4f
Plasmid#234444PurposeBacterial expression of glutamate sensor with with improved sensitivity and fast deactivation kinetics (4f = fast)DepositorInsertiGluSnFR4f
Tags6xHis-Xpress epiExpressionBacterialPromoterT7Available SinceMarch 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
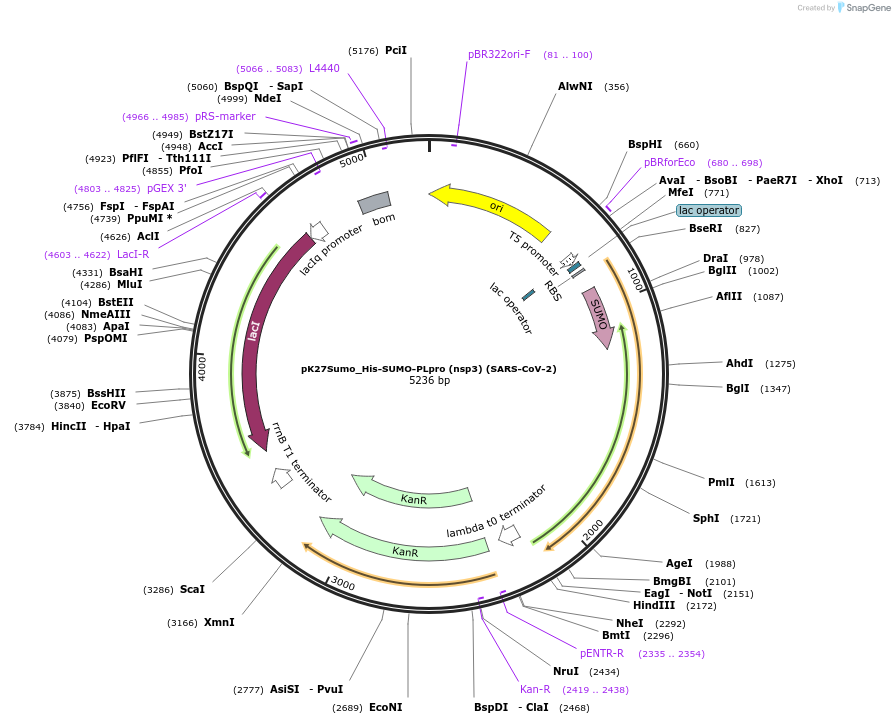
pK27Sumo_His-SUMO-PLpro (nsp3) (SARS-CoV-2)
Plasmid#169193PurposeTo express SARS-CoV-2 Papain-like protease in E. coliDepositorInsert14His-SUMO-nsp3_E746-K1060 (pp1ab_E1564-K1878) (ORF1ab Synthetic, SARS-CoV-2)
Tags14His-SUMOExpressionBacterialMutationCodon optimised for E. coliPromoterT5Available SinceJuly 6, 2021AvailabilityAcademic Institutions and Nonprofits only -
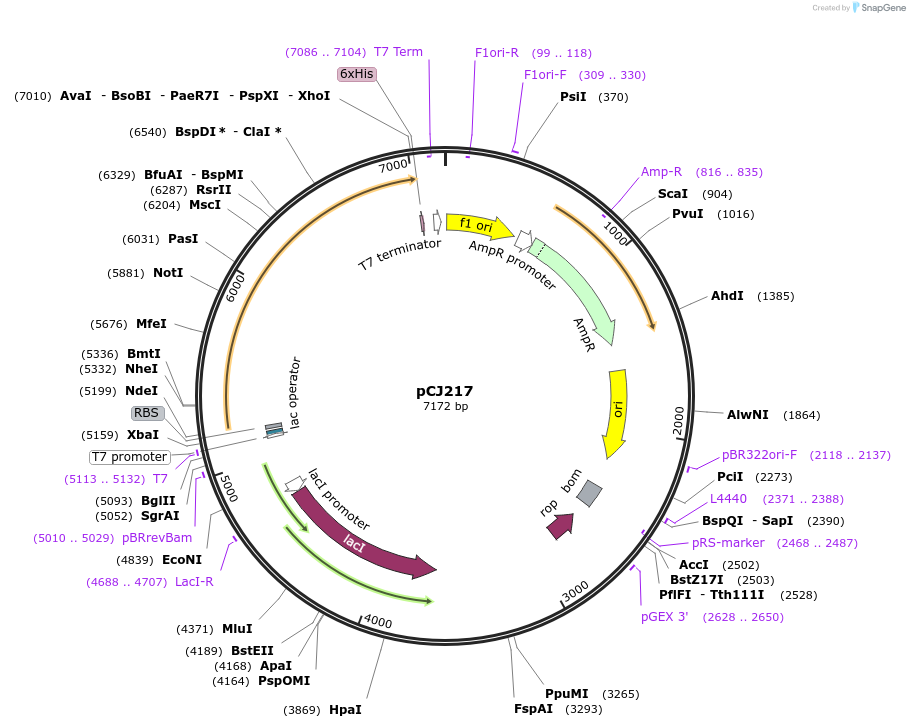
pCJ217
Plasmid#162685PurposepET-21b(+) based plasmid for expression of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating two point mutations, G489C and S530C, to introduce a 6th disulfide bond (from PETase).DepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), with mutations to introduce a 6th disulfide bond from PETase
TagsHisExpressionBacterialMutationG489C, S530C; codon optimized for E. coli K12PromoterT7/lacAvailable SinceJan. 6, 2021AvailabilityAcademic Institutions and Nonprofits only -
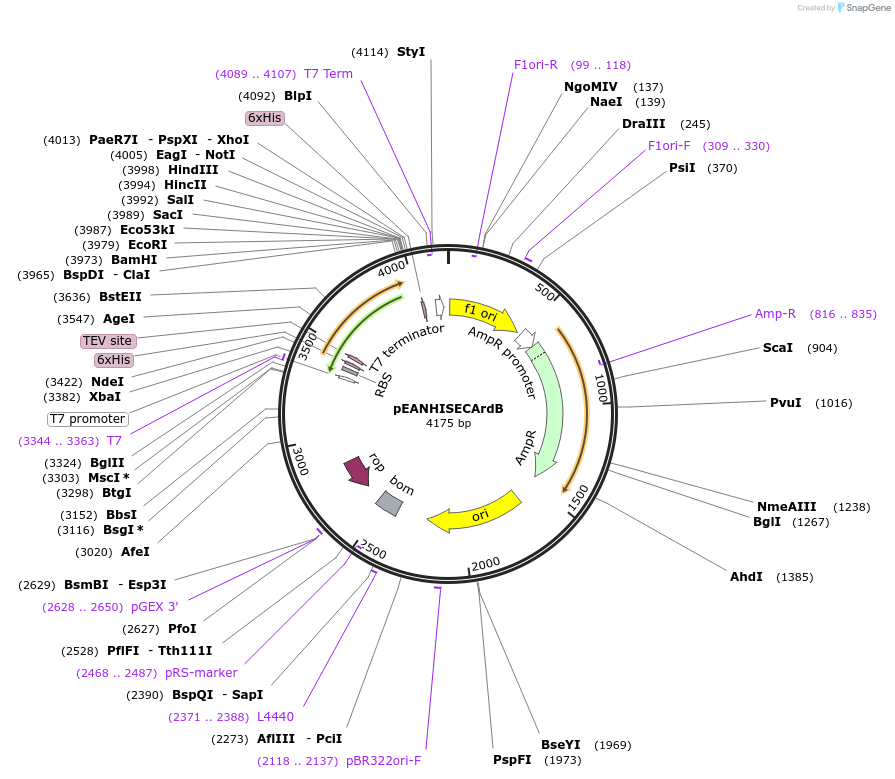
pEANHISECArdB
Plasmid#96967PurposeExpression of E. coli ArdB protein wirh an N-terminal TEV protease cleavable His tagDepositorInsertE. coli ArdB
TagsN-ter TEV cleavable 6HisExpressionBacterialMutationnonePromoterT7Available SinceAug. 17, 2017AvailabilityAcademic Institutions and Nonprofits only -
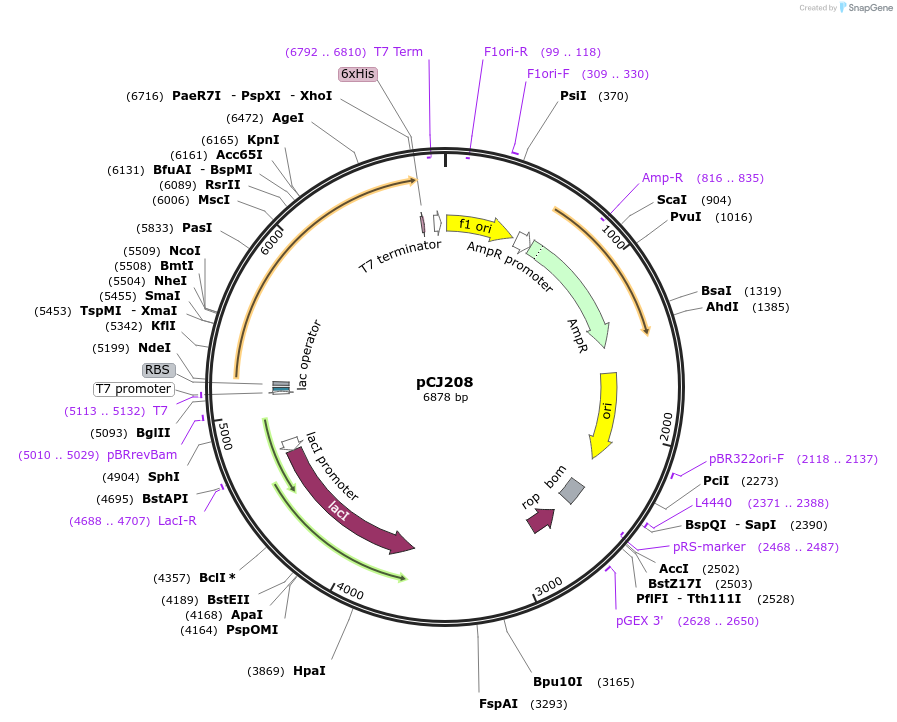
pCJ208
Plasmid#162681PurposeLidded variant of PETase from Ideonella sakaiensis 201-F6 (Genbank GAP38373.1) removing a seven-residue loop of PETase (Trp185:Phe191, PETase numbering) and replacing it with Gly251:Thr472 from MHETase; codon optimized for expression in E. coli K12, with C-terminal His tag.DepositorInsertPETase from Ideonella sakaiensis 201-F6 (Genbank GAP38373.1) incorporating the MHETase lid
TagsHisExpressionBacterialMutationReplacement of W185-F191 with G251-T472 from MHET…PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
pK27Sumo_His-SUMO-nsp14-nsp10 fusion (SARS-CoV-2)
Plasmid#169160PurposeTo express nsp14-nsp10 fusion protein in E. coliDepositorInsert14His-SUMO-nsp14-GGSGGS-nsp10 (ORF1ab Synthetic, SARS-CoV-2)
Tags14His-SUMO and Fusion protein of SARS-CoV-2 nsp14…ExpressionBacterialMutationCodon optimised for E. coliPromoterT5Available SinceJuly 6, 2021AvailabilityAcademic Institutions and Nonprofits only -
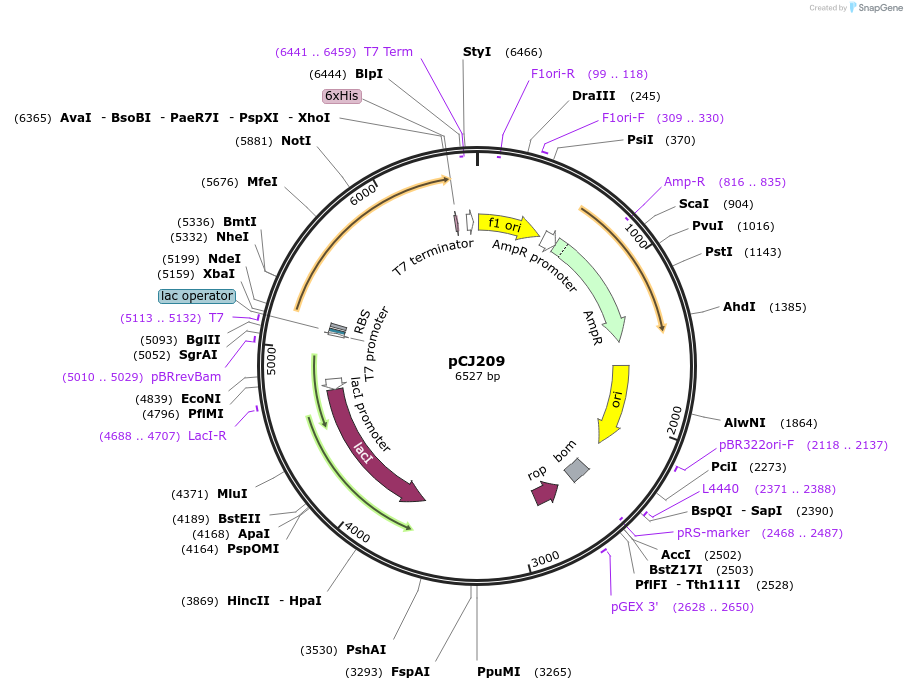
pCJ209
Plasmid#162682PurposeLidless variant of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1) replacing Gly251:Thr472 of MHETase with the 7-residue loop (Trp185:Phe191, PETase numbering) of PETase , codon optimized for expression in E. coli K12, with C-terminal His tag.DepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1) with the lid removed
TagsHisExpressionBacterialMutationReplacement of G251-T472 with W185-F191 from PETa…PromoterT7/lacAvailable SinceJan. 6, 2021AvailabilityAcademic Institutions and Nonprofits only -
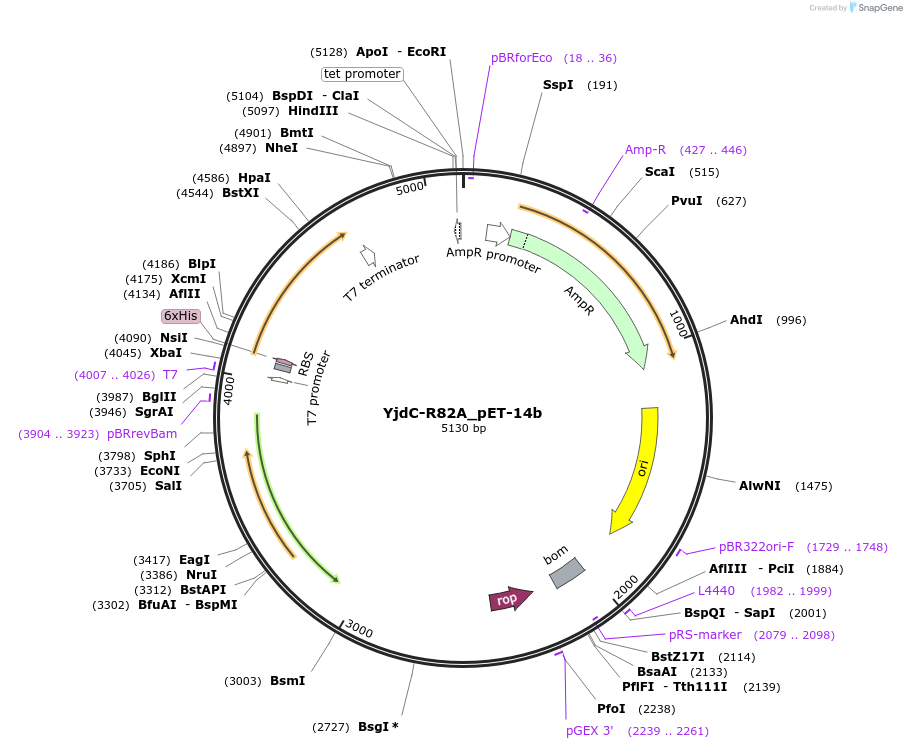
YjdC-R82A_pET-14b
Plasmid#233276PurposeOverexpress his-tag YjdC protein with a site directed mutagenesis, arginine at position 82 was substituted with alanine (R82A)DepositorInsertYjdC (yjdC )
ExpressionBacterialMutationArginine was replaced with Alanine at position 82…PromoterT7 promoterAvailable SinceMarch 17, 2025AvailabilityAcademic Institutions and Nonprofits only -
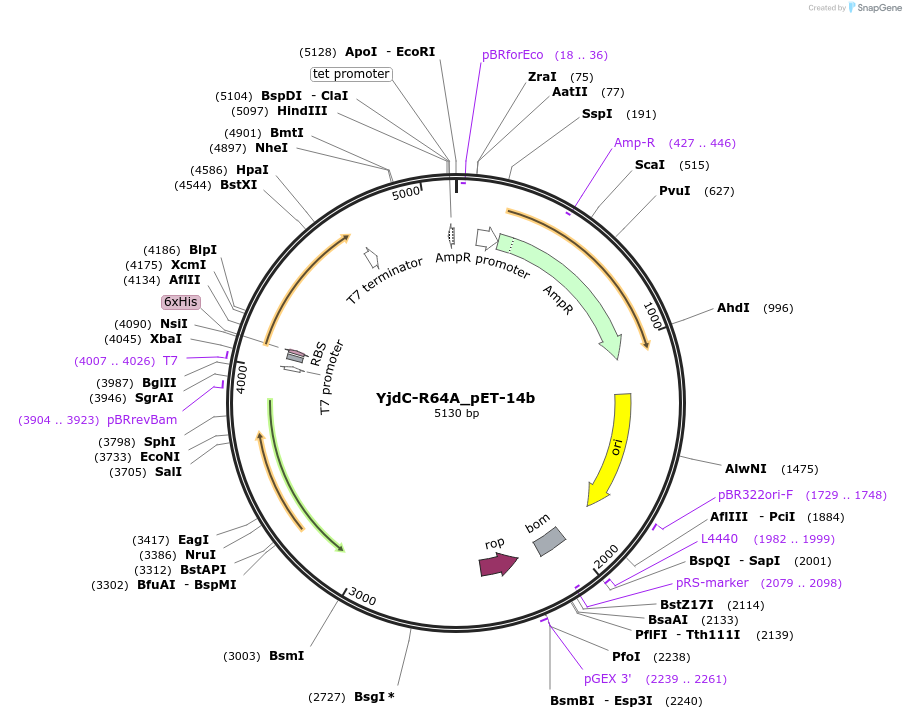
YjdC-R64A_pET-14b
Plasmid#233263PurposeOverexpress his-tag YjdC protein with a site directed mutagenesis, arginine at position 64 was substituted with alanine (R64A)DepositorInsertYjdC (yjdC )
ExpressionBacterialMutationArginine was replaced with Alanine at position 64…PromoterT7 promoterAvailable SinceMarch 17, 2025AvailabilityAcademic Institutions and Nonprofits only -
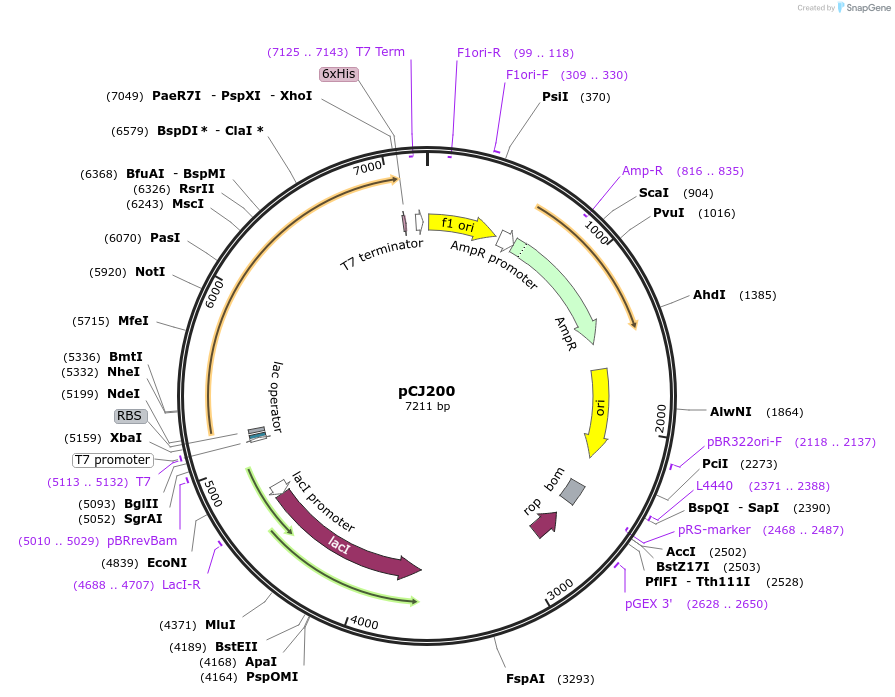
pCJ200
Plasmid#162673PurposepET-21b(+) based plasmid for expression of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating S136C and a sequence Tyr75:Phe104 to introduce a 6th disulfide bond as in AoFaeB-2.DepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1)
TagsHisExpressionBacterialMutationS136C and a sequence Y75-F104 to introduce a 6th…PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only