We narrowed to 763 results for: streptococcus -(crispr) -(cas9)
-
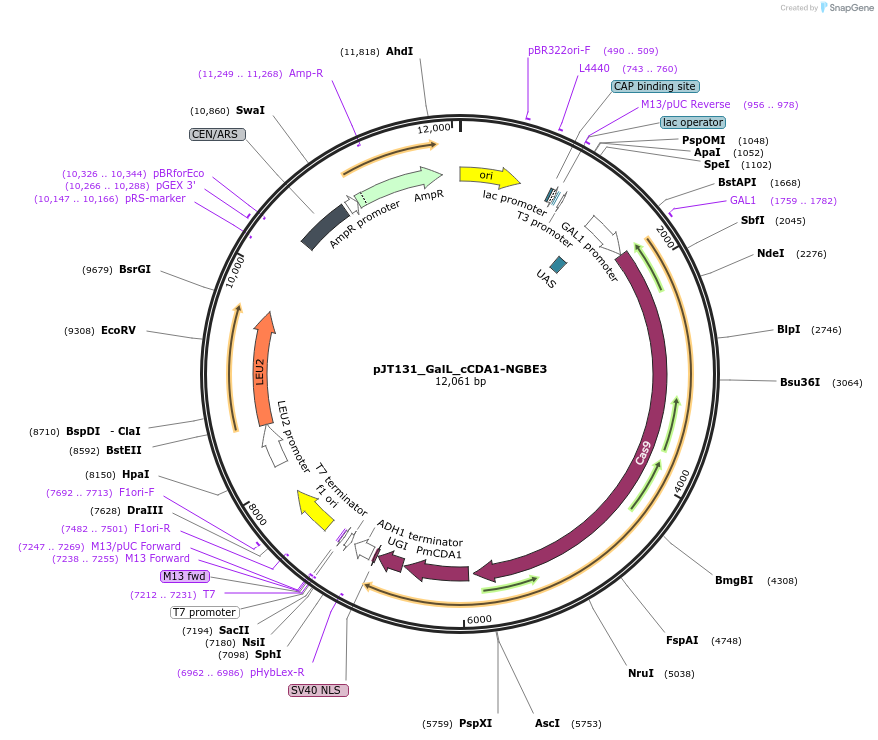
Plasmid#145095PurposeExpressing base editor cCDA1-NGBE3 in yeast cellsDepositorInsertcCDA1-NGBE3
UseCRISPRExpressionYeastMutationspCas9(D10A/L1111R/D1135V/G1218R/E1219F/A1322R/R1…PromoterGalLAvailable SinceJuly 15, 2020AvailabilityAcademic Institutions and Nonprofits only -
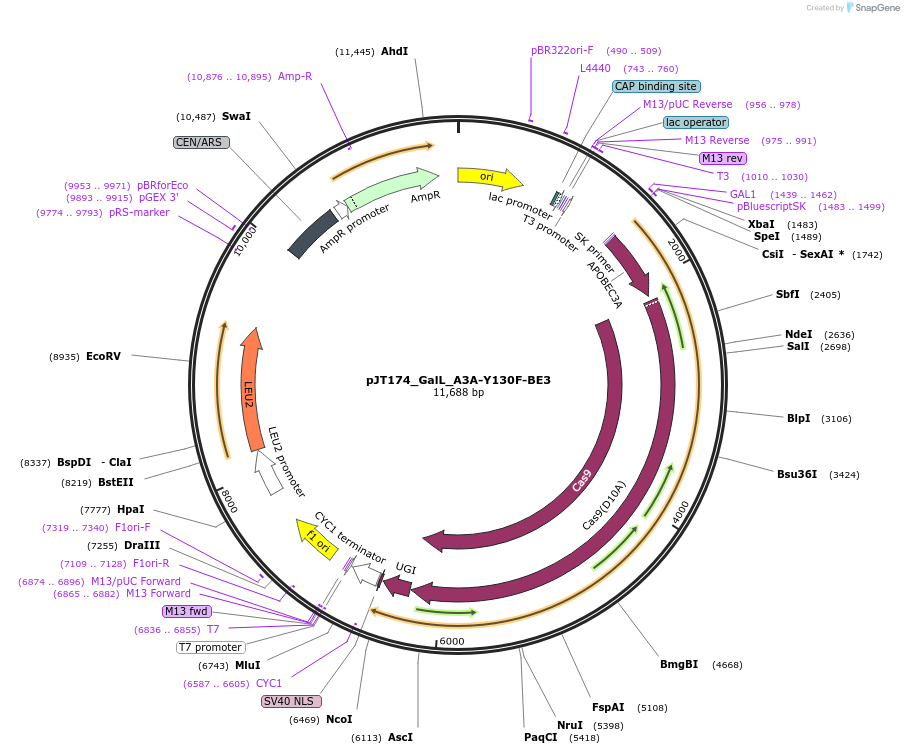
pJT174_GalL_A3A-Y130F-BE3
Plasmid#145123PurposeExpressing base editor A3A-Y130F-BE3 in yeast cellsDepositorInsertA3A(Y130F)-BE3
UseCRISPRExpressionYeastMutationA3A(Y130F); spCas9(D10A)PromoterGalLAvailable SinceJuly 22, 2020AvailabilityAcademic Institutions and Nonprofits only -
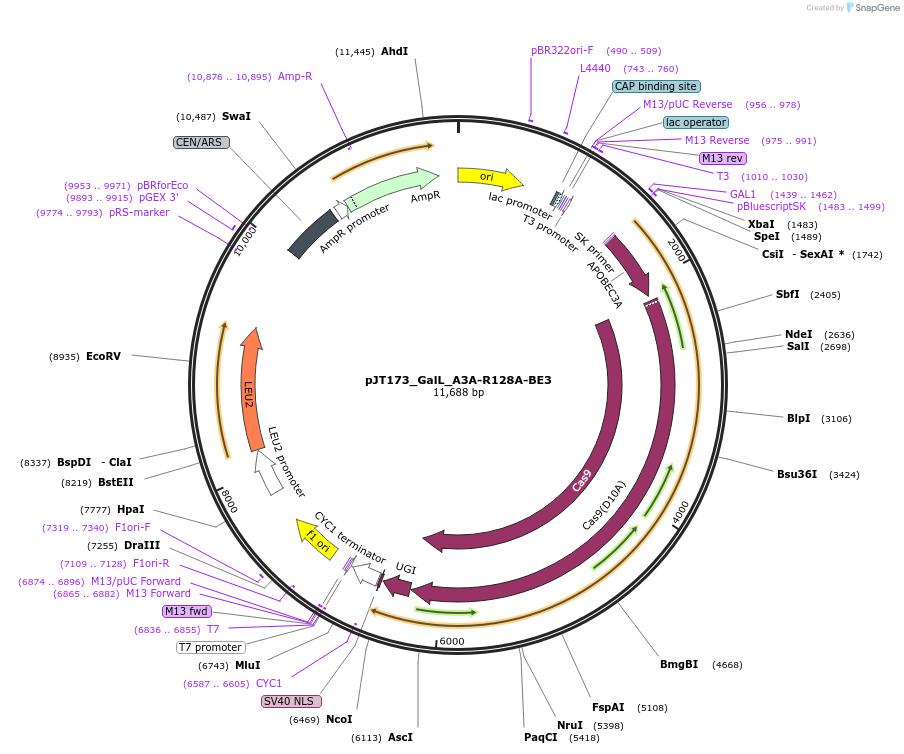
pJT173_GalL_A3A-R128A-BE3
Plasmid#145122PurposeExpressing base editor A3A-R128A-BE3 in yeast cellsDepositorInsertA3A(R128A)-BE3
UseCRISPRExpressionYeastMutationA3A(R128A); spCas9(D10A)PromoterGalLAvailable SinceJuly 22, 2020AvailabilityAcademic Institutions and Nonprofits only -
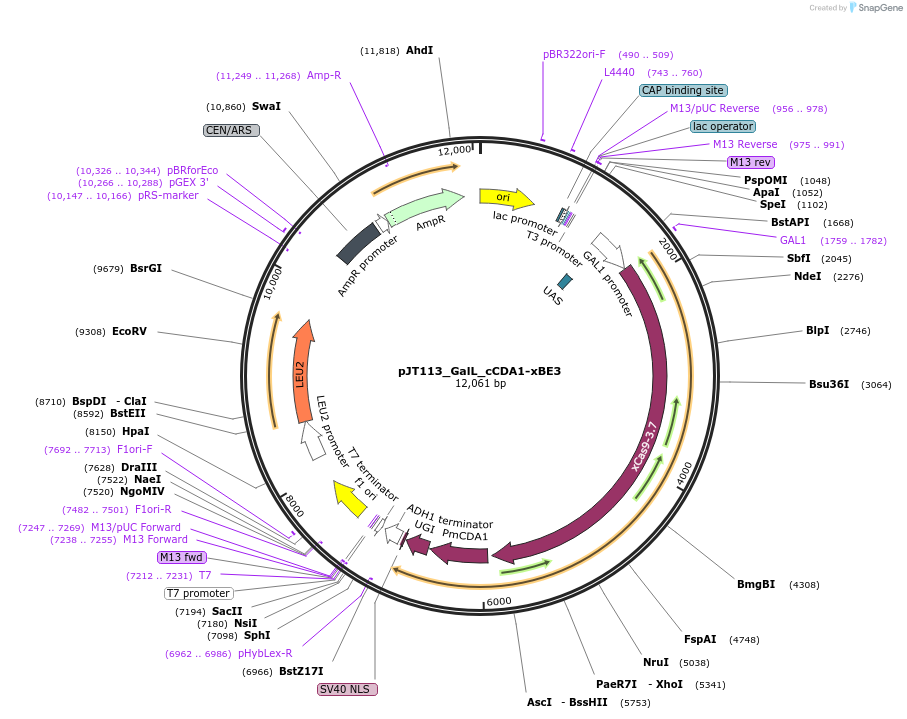
pJT113_GalL_cCDA1-xBE3
Plasmid#145079PurposeExpressing base editor cCDA1-xBE3 in yeast cellsDepositorInsertcCDA1-xBE3
UseCRISPRExpressionYeastMutationspCas9(D10A, A262T, R324L, S409I, E480K, E543D, M…PromoterGalLAvailable SinceJuly 15, 2020AvailabilityAcademic Institutions and Nonprofits only -
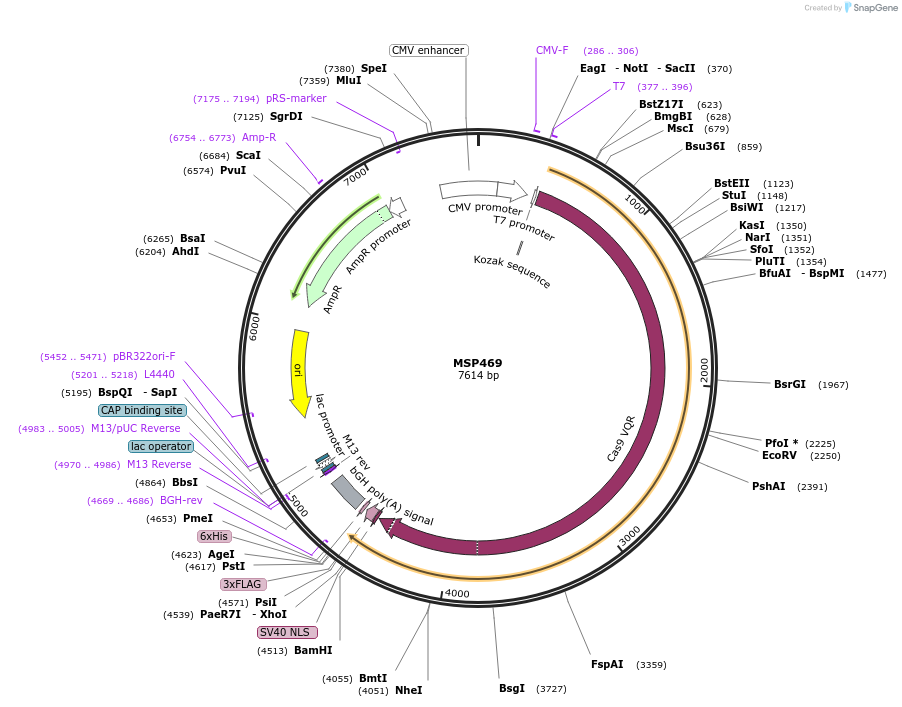
MSP469
Plasmid#65771PurposeHuman expression vector for SpCas9 VQR variant: CMV-T7-humanSpCas9(D1135V/R1335Q/T1337R)-NLS-3xFLAG (VQR variant)DepositorInsertmammalian codon-optimized Streptococcus pyogenes Cas9 (D1135V/R1335Q/T1337R)-NLS-3XFlag
UseCRISPRTags3x FLAG and NLSExpressionMammalianMutationD1135V, R1335Q, and T1337R mutations in Cas9PromoterCMV & T7Available SinceJune 22, 2015AvailabilityAcademic Institutions and Nonprofits only -
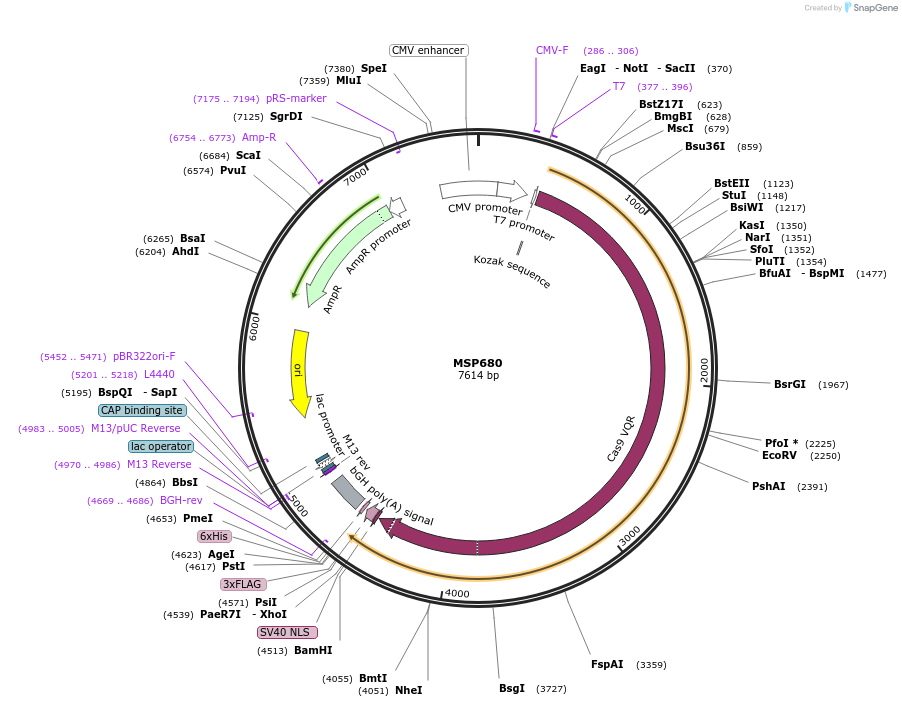
MSP680
Plasmid#65772PurposeHuman expression vector for SpCas9 EQR variant: CMV-T7-humanSpCas9(D1135E/R1335Q/T1337R)-NLS-3xFLAG (EQR variant)DepositorInsertmammalian codon-optimized Streptococcus pyogenes Cas9 (D1135E/R1335Q/T1337R)-NLS-3XFlag
UseCRISPRTags3x FLAG and NLSExpressionMammalianMutationD1135E, R1335Q, and T1337R mutations in Cas9PromoterCMV & T7Available SinceJune 22, 2015AvailabilityAcademic Institutions and Nonprofits only -
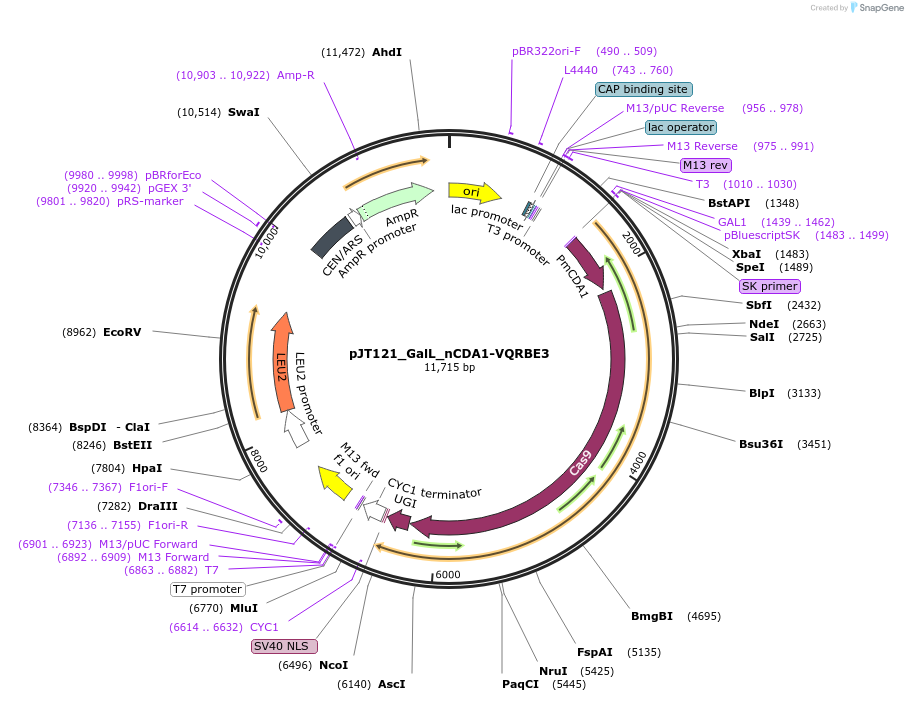
pJT121_GalL_nCDA1-VQRBE3
Plasmid#145086PurposeExpressing base editor nCDA1-VQRBE3 in yeast cellsDepositorInsertnCDA1-VQRBE3
UseCRISPRExpressionYeastMutationspCas9(D10A, D1135V, R1335Q, T1337R)PromoterGalLAvailable SinceJuly 15, 2020AvailabilityAcademic Institutions and Nonprofits only -
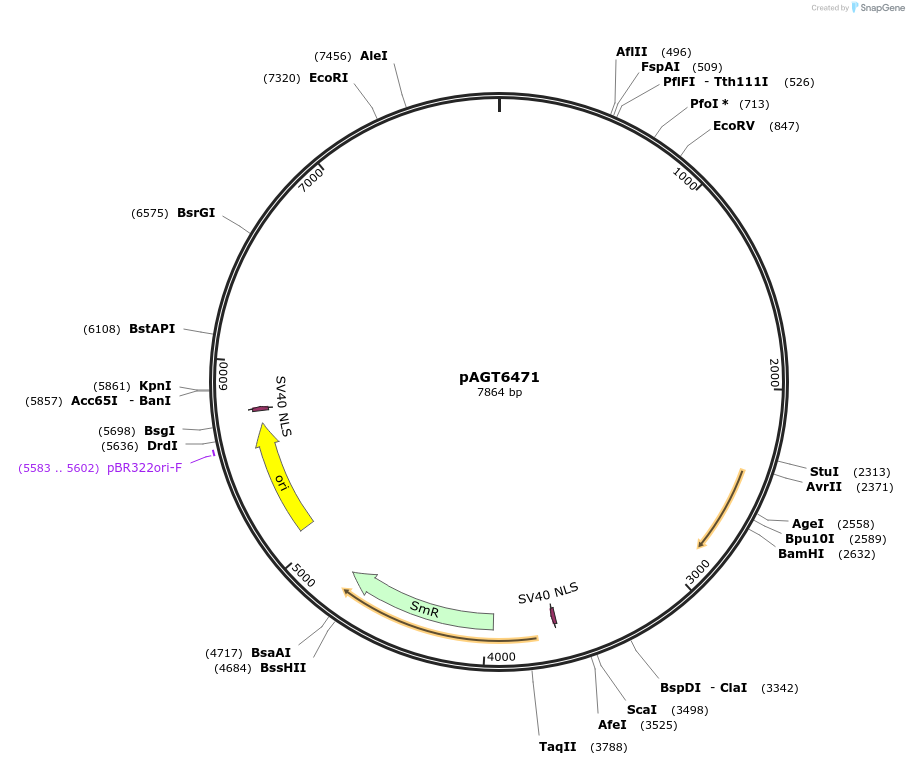
pAGT6471
Plasmid#219428PurposeModule of intronized NLS-SpCas9-NLS for N-terminal and C-terminal tag fusions (AGGT_N-SpCas9i-N_TTCG)DepositorInsertAGGT_NLS-SpCas9i-NLS(no stop)_TTCG
UseMoclo compatible level 0 moduleAvailable SinceJuly 2, 2024AvailabilityAcademic Institutions and Nonprofits only -
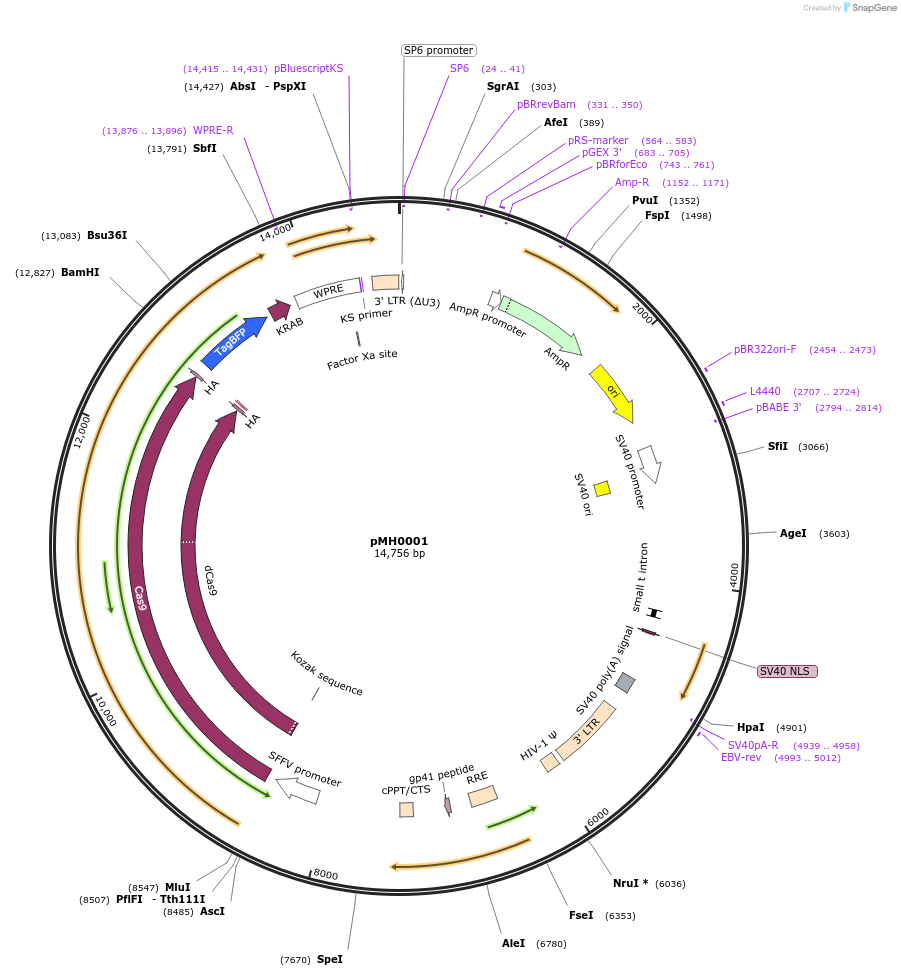
pMH0001
Plasmid#85969PurposeUCOE-SFFV-dCas9-BFP-KRABDepositorInsertdCas9
UseLentiviralTagsBFP-KRABExpressionMammalianMutationD10A, H840APromoterSFFVAvailable SinceFeb. 3, 2017AvailabilityAcademic Institutions and Nonprofits only -
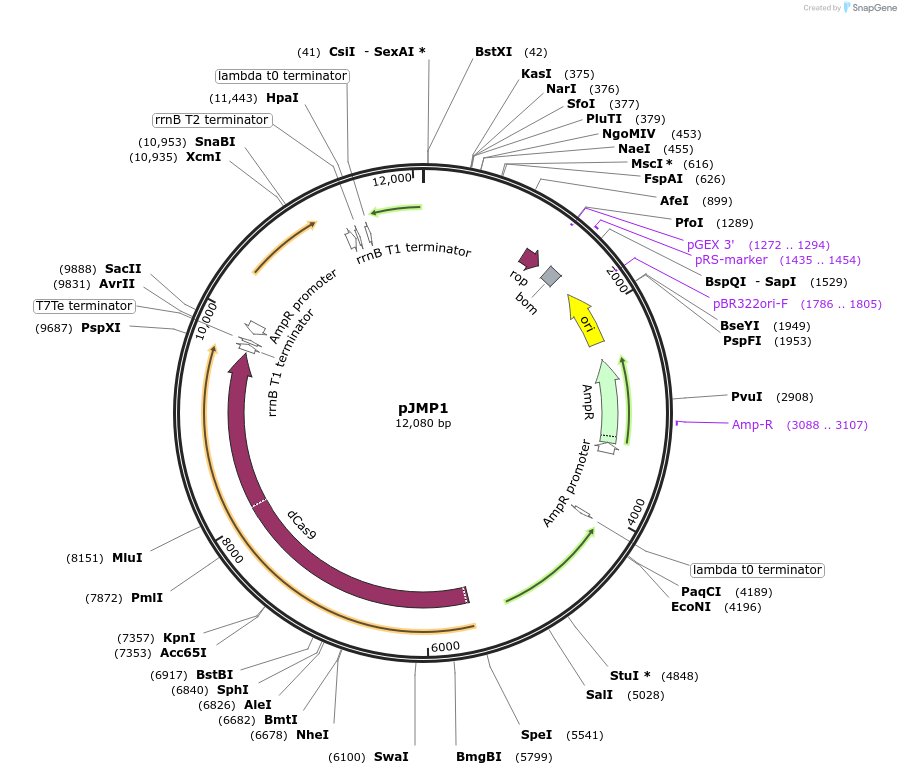
pJMP1
Plasmid#79873PurposeBacillus subtilis dCas9 expression vector; integrates into lacA/ganADepositorInsertdCas9
UseCRISPRExpressionBacterialMutationD10A, H840APromoterxylAAvailable SinceOct. 20, 2016AvailabilityAcademic Institutions and Nonprofits only -
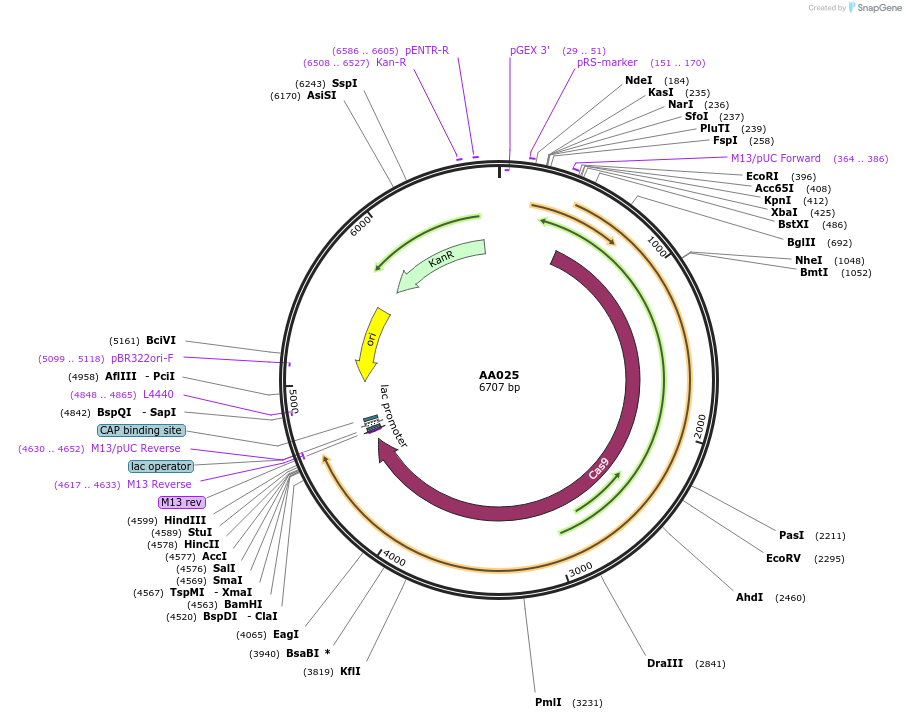
AA025
Plasmid#216005PurposeFragmid fragment: (Cas protein) NG PAMDepositorHas ServiceCloning Grade DNAInsertCas9-NG_v1.1 [Sp]
UseCRISPR; FragmentAvailable SinceApril 29, 2024AvailabilityAcademic Institutions and Nonprofits only -
AA027
Plasmid#216007PurposeFragmid fragment: (Cas protein) nearly PAM-lessDepositorHas ServiceCloning Grade DNAInsertCas9-SpRY_v1.1 [Sp]
UseCRISPR; FragmentAvailable SinceApril 29, 2024AvailabilityAcademic Institutions and Nonprofits only -
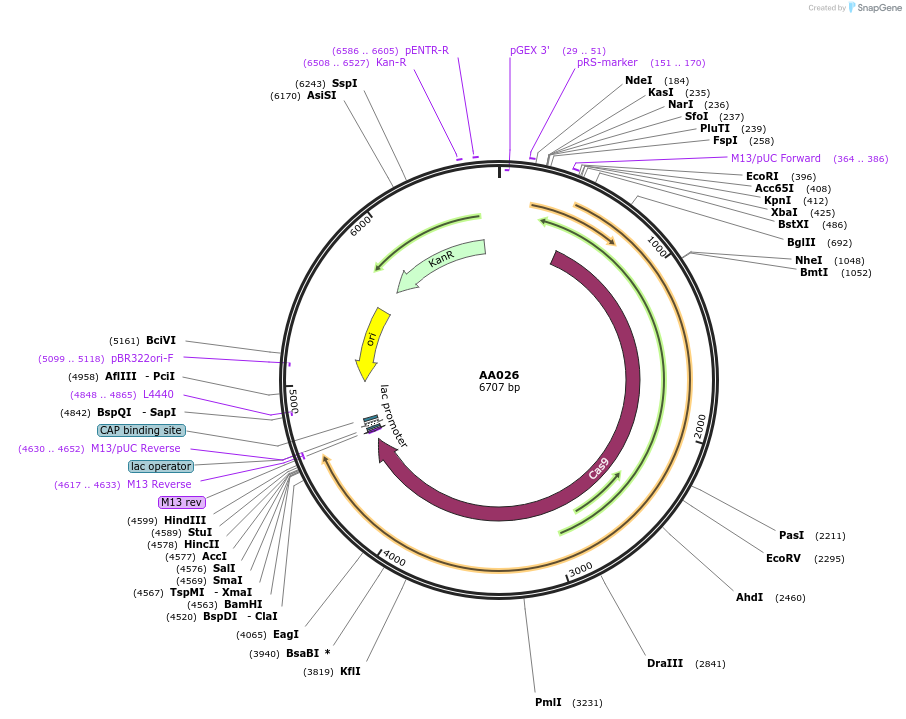
AA026
Plasmid#216006PurposeFragmid fragment: (Cas protein) NG PAMDepositorHas ServiceCloning Grade DNAInsertCas9-SpG_v1.1 [Sp]
UseCRISPR; FragmentAvailable SinceApril 29, 2024AvailabilityAcademic Institutions and Nonprofits only -
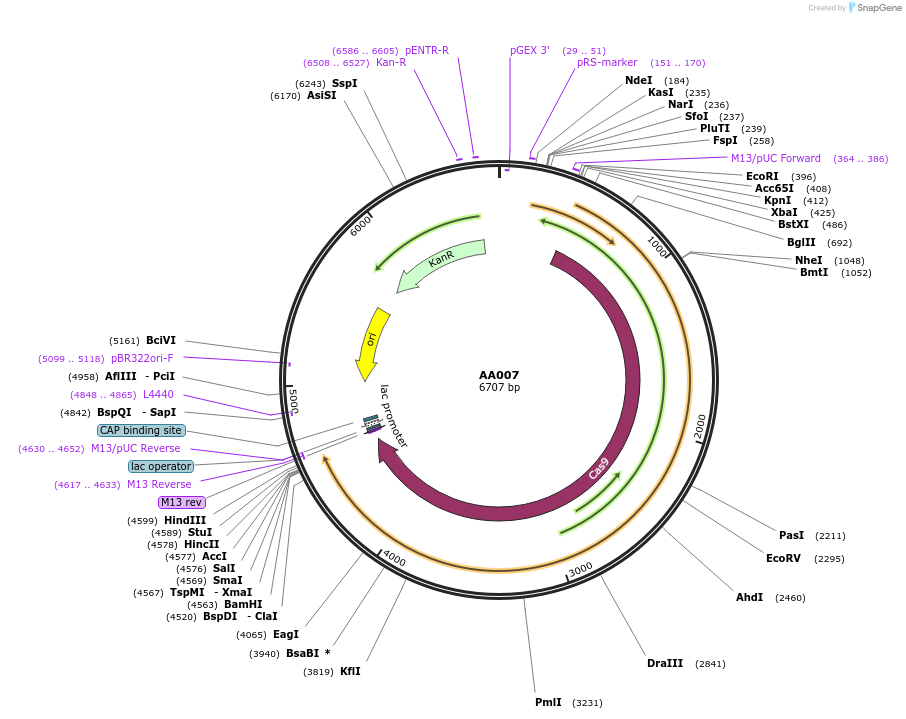
AA007
Plasmid#216002PurposeFragmid fragment: (Cas protein) canonical Cas9; NGG PAMDepositorHas ServiceCloning Grade DNAInsertCas9_v1.1 [Sp]
UseCRISPR; FragmentAvailable SinceApril 29, 2024AvailabilityAcademic Institutions and Nonprofits only -
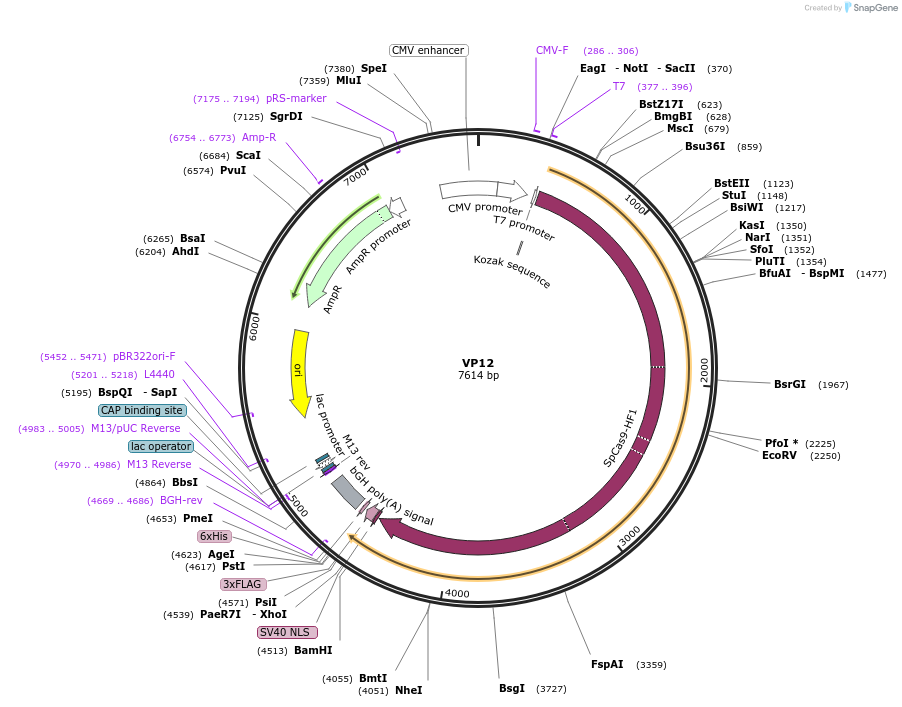
VP12
Plasmid#72247PurposeHuman expression plasmid for SpCas9-HF1 variant: CMV-T7-humanSpCas9-HF1(N497A, R661A, Q695A, Q926A)-NLS-3xFLAGDepositorInsertmammalian codon-optimized Streptococcus pyogenes Cas9 HF1(N497A/R661A/Q695A/Q926A)-NLS-3xFlag
UseCRISPRTags3x FLAG and NLSExpressionMammalianMutationN497A, R661A, Q695A and Q926APromoterCMVAvailable SinceJan. 6, 2016AvailabilityAcademic Institutions and Nonprofits only -
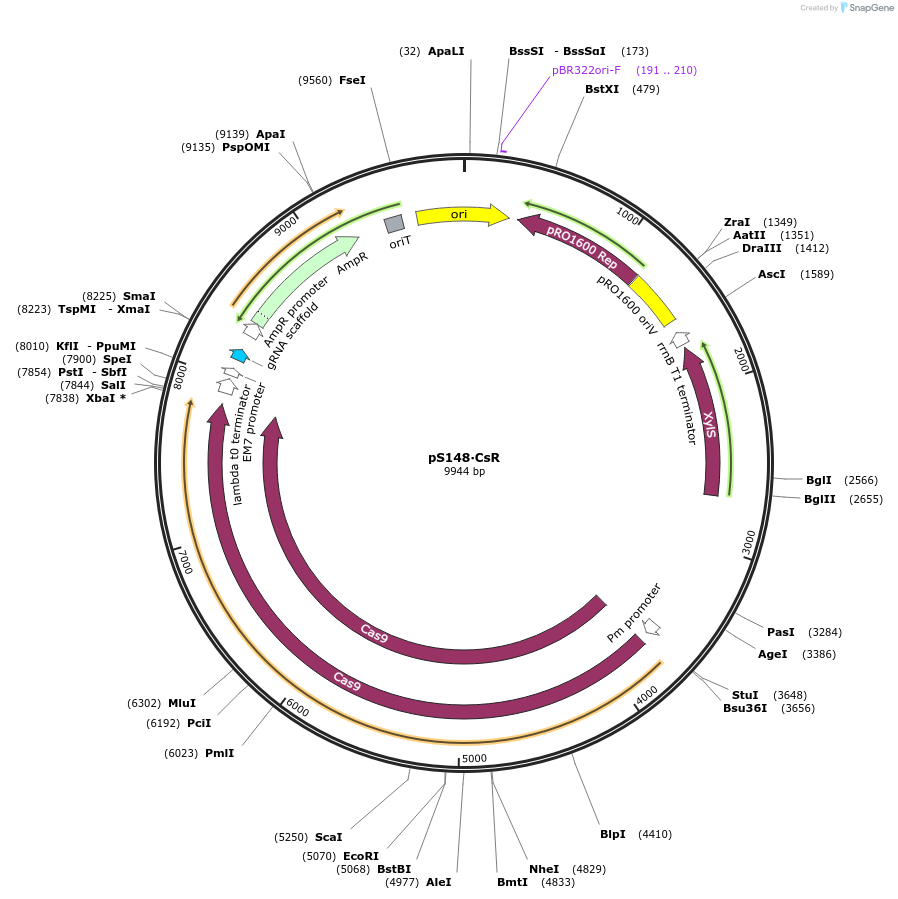
pS148∙CsR
Plasmid#197858PurposeExpresses Streptococcus pyogenes' cas9 and a tailored sgRNA for counterselection in gram-negative bacteriaDepositorInsertgRNA scaffold - Ap/Cb resistance cassette - incP oriT
UseCRISPR and Synthetic BiologyPromoterPEM7Available SinceOct. 10, 2023AvailabilityAcademic Institutions and Nonprofits only -
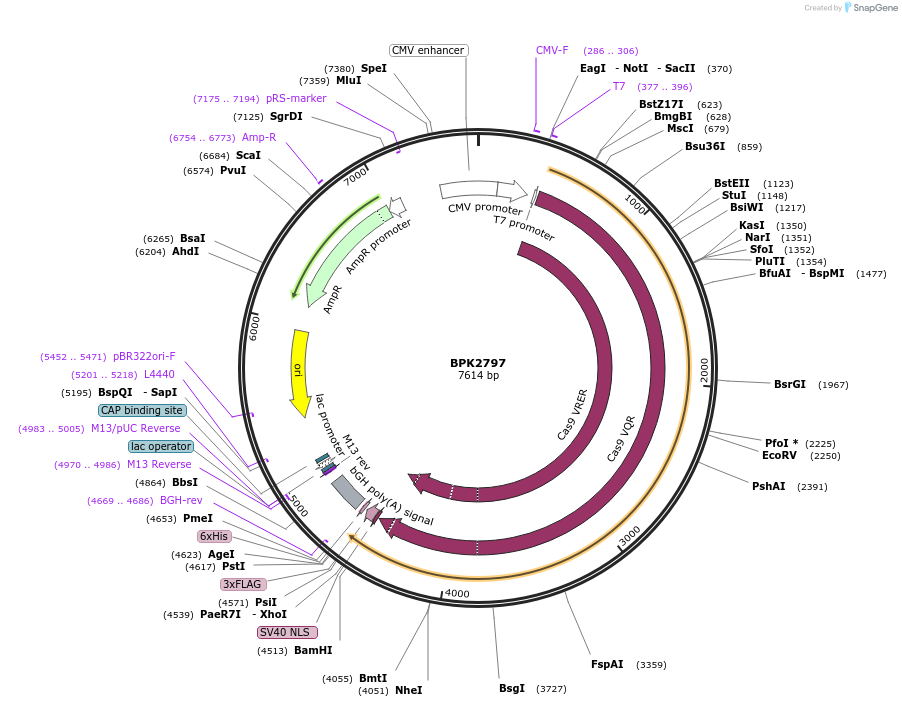
BPK2797
Plasmid#72251PurposeHuman expression plasmid for SpCas9-VRQR variant: CMV-T7-humanSpCas9-VRQR(D1135V, G1218R, R1335Q, T1337R)-NLS-3xFLAGDepositorInsertmammalian codon-optimized Streptococcus pyogenes Cas9 VRQR(D1135V/G1218R/R1335Q/T1337R)-NLS-3xFlag
UseCRISPRTags3x FLAG and NLSExpressionMammalianMutationD1135V, G1218R, R1335Q, and T1337RPromoterCMVAvailable SinceJan. 6, 2016AvailabilityAcademic Institutions and Nonprofits only -
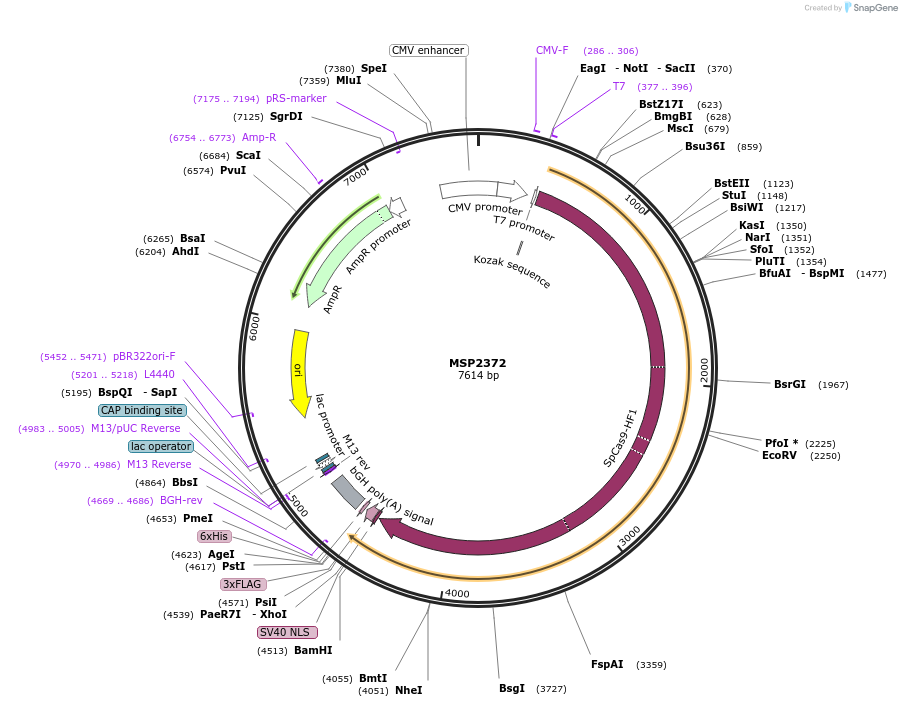
MSP2372
Plasmid#73196PurposeHuman expression plasmid for SpCas9(R661A/Q695A/Q926A) variant: CMV-T7-humanSpCas9(R661A, Q695A, Q926A)-NLS-3xFLAGDepositorInsertmammalian codon-optimized Streptococcus pyogenes Cas9 (R661A/Q695A/Q926A)-NLS-3xFlag
UseCRISPRTags3x FLAG and NLSExpressionMammalianMutationR661A, Q695A, and Q926A in SpCas9PromoterCMVAvailable SinceFeb. 22, 2016AvailabilityAcademic Institutions and Nonprofits only -
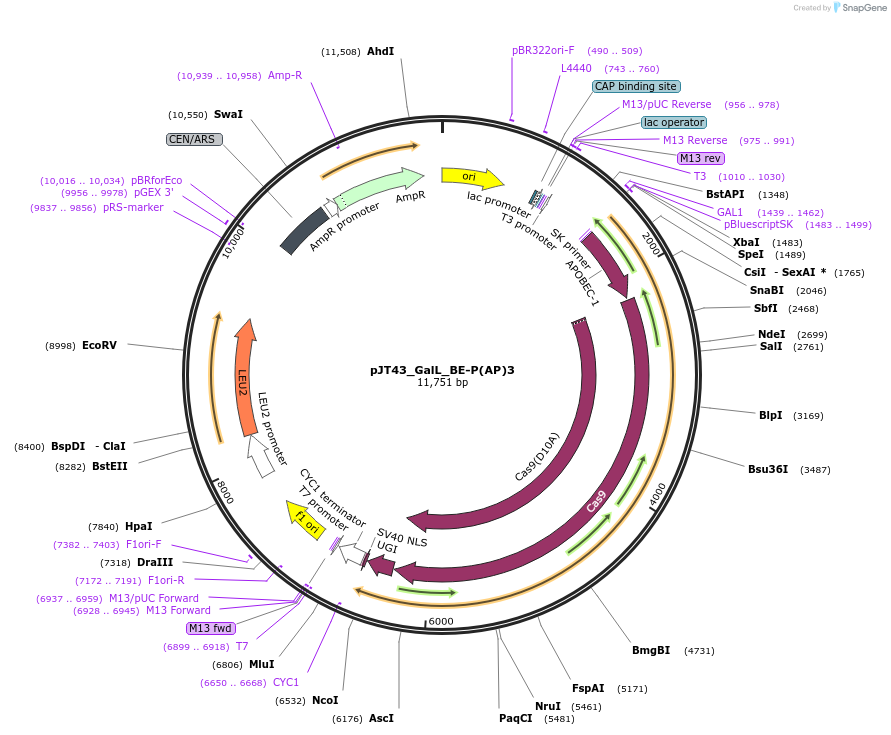
pJT43_GalL_BE-P(AP)3
Plasmid#145036PurposeExpresses BE-P(AP)3 in yeast cellsDepositorInsertBE-(AP)3
UseCRISPRExpressionYeastMutationspCas9(D10A)PromoterGalLAvailable SinceJuly 9, 2020AvailabilityAcademic Institutions and Nonprofits only -
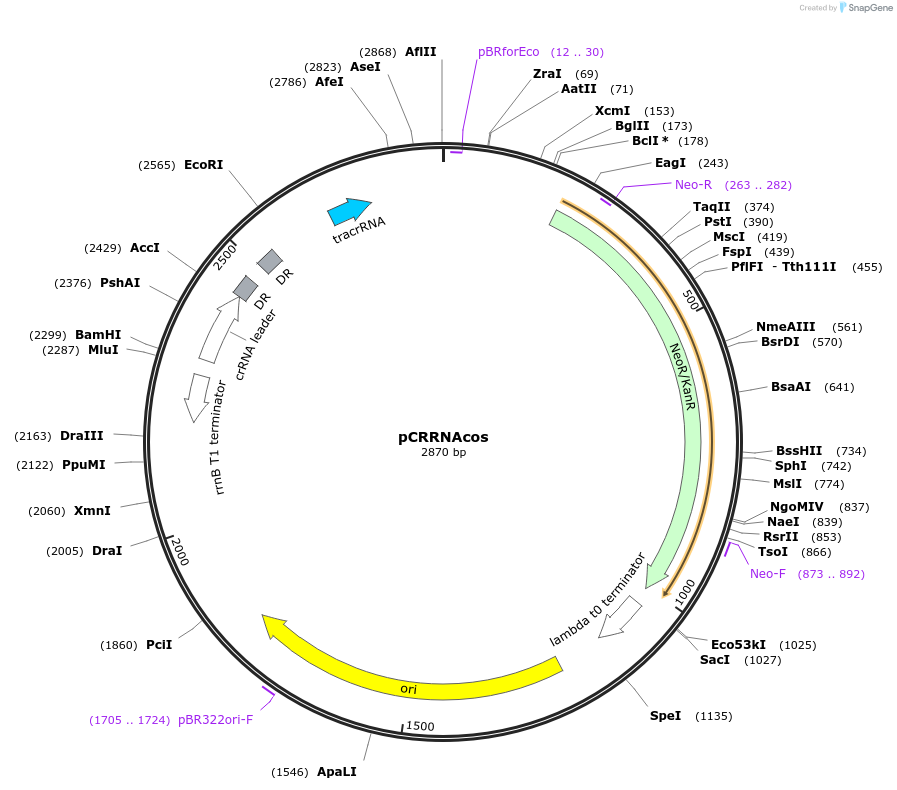
pCRRNAcos
Plasmid#78492PurposeTranscription of Streptococcus pyogenes crRNA and tracrRNA under the wildtype crRNA promoter.DepositorInserttracrRNA
ExpressionBacterialAvailable SinceApril 4, 2017AvailabilityAcademic Institutions and Nonprofits only