We narrowed to 69,184 results for: Eras
-
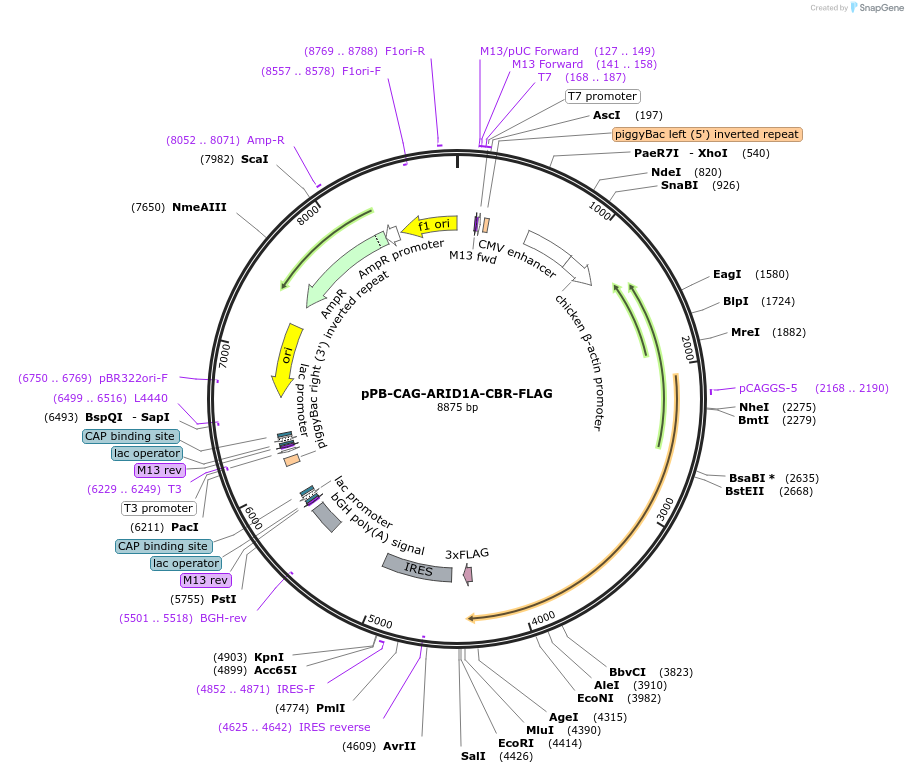
Plasmid#242212PurposeExpression of core binding region (CBR) of human ARID1A using piggyBac transposition. ∆IDR1, ∆ARID, and ∆IDR2 (deletion of a.a 2-1610)DepositorInsertARID1A (ARID1A Human)
UsePiggybacTagsFLAGExpressionMammalianMutationCore binding region (CBR) only of ARID1A. Deleted…PromoterCAGAvailable SinceSept. 9, 2025AvailabilityAcademic Institutions and Nonprofits only -
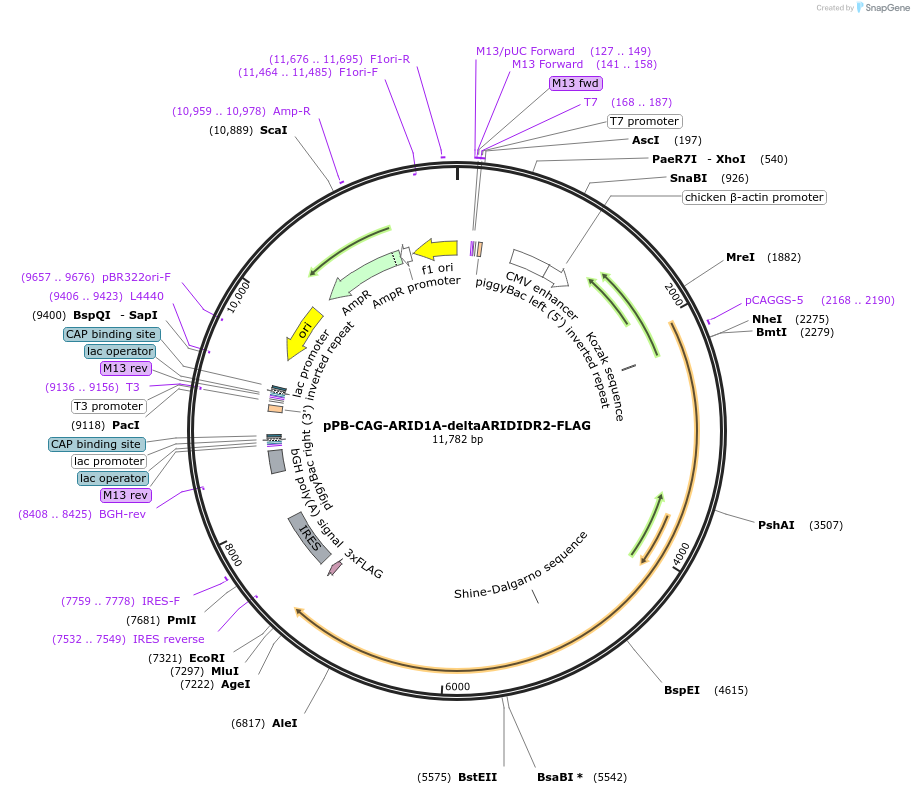
pPB-CAG-ARID1A-deltaARIDIDR2-FLAG
Plasmid#242214PurposeExpression of human ARID1A ∆ARID and ∆IDR2 (deletion of a.a. 971-1610) using piggyBac transpositionDepositorInsertARID1A (ARID1A Human)
UsePiggybacTagsFLAGExpressionMammalianMutationDeleted ARID and IDR2 from ARID1A (deletion of a.…PromoterCAGAvailable SinceSept. 9, 2025AvailabilityAcademic Institutions and Nonprofits only -
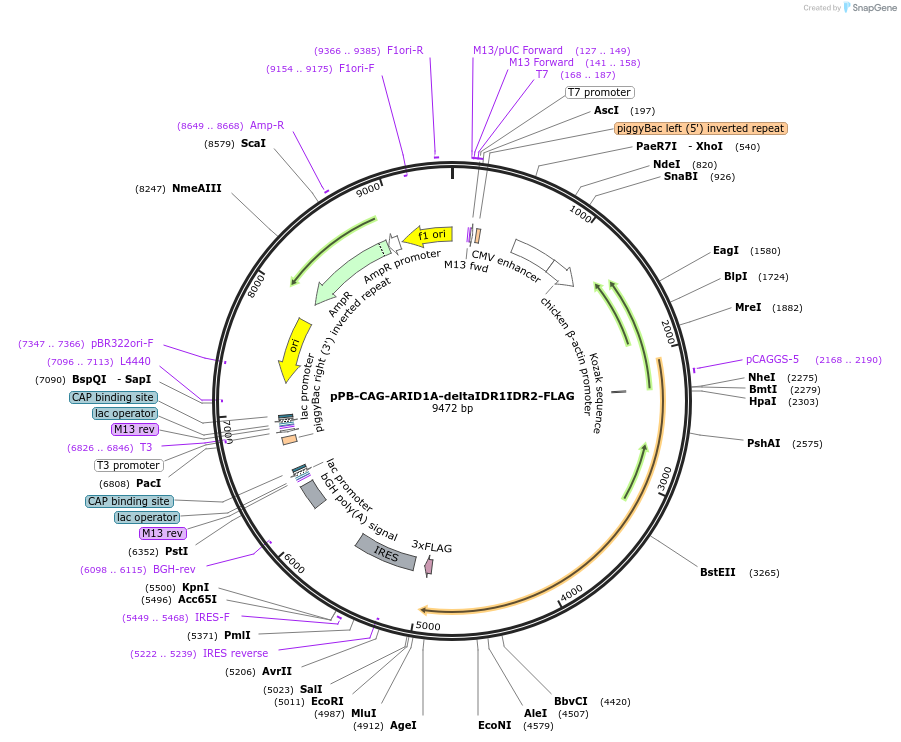
pPB-CAG-ARID1A-deltaIDR1IDR2-FLAG
Plasmid#242213PurposeExpression of human ARID1A ∆IDR1 and ∆IDR2 (deletion of a.a. 2-970 and a.a. 1170-1610) using piggyBac transpositionDepositorInsertARID1A (ARID1A Human)
UsePiggybacTagsFLAGExpressionMammalianMutationDeleted IDR1 and IDR2 from ARID1A (deletion of a.…PromoterCAGAvailable SinceSept. 9, 2025AvailabilityAcademic Institutions and Nonprofits only -
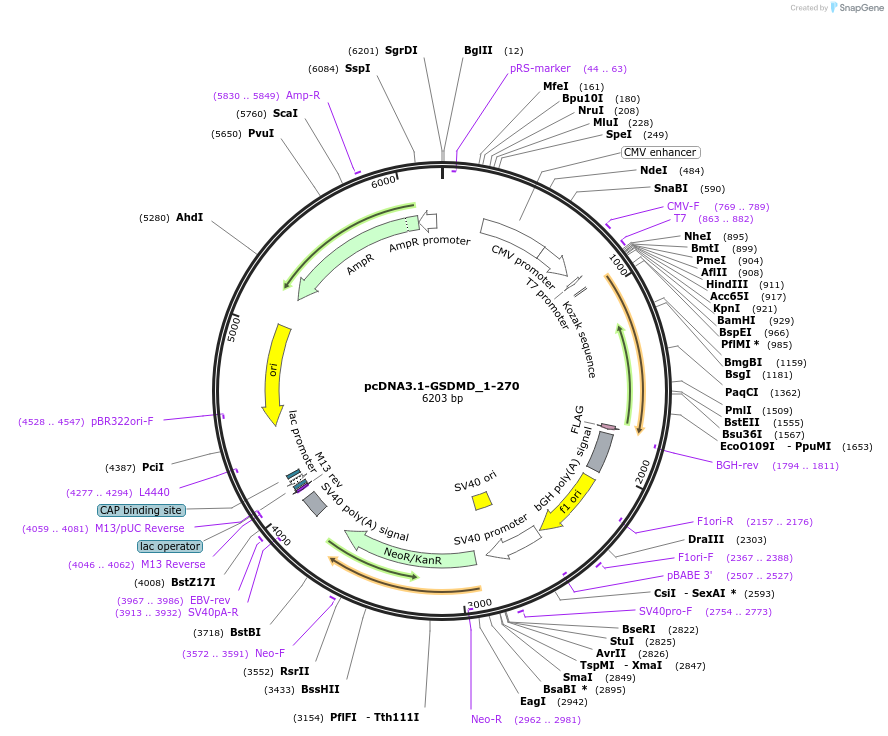
pcDNA3.1-GSDMD_1-270
Plasmid#218229PurposeGasdermin1-270 is a pore-forming GSDMD formed by SARS-CoV-2 3CLpro cleavage. Transfection in HEK293 cells showed GSDMD 1-270 pores secrete 3CLpro, nucleocapsid, and in excess, kill cells by pyroptosisDepositorInsertGSDMD (GSDMD Human)
TagsflagExpressionMammalianMutationconstruct only codes for amino acids 1-270Available SinceJuly 1, 2025AvailabilityIndustry, Academic Institutions, and Nonprofits -
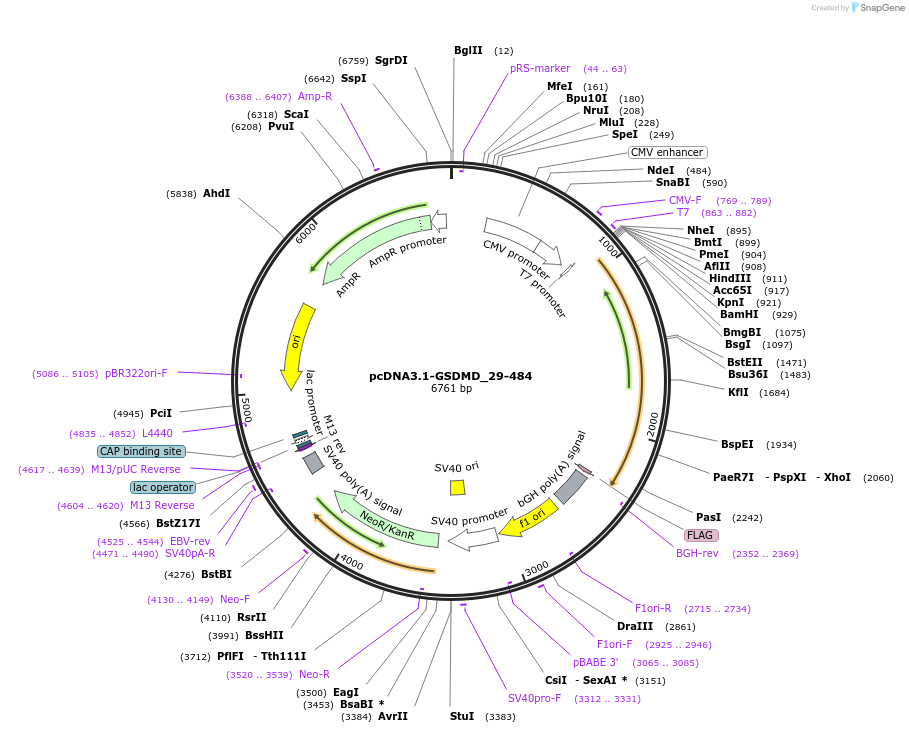
pcDNA3.1-GSDMD_29-484
Plasmid#218231PurposeGasdermin-D_29-484 is a non-functional GSDMD formed by SARS-CoV-2 3CLpro cleavage in the N-terminal domain. Transfection in HEK293 cells showed no pore-forming activity or induction of pyroptosisDepositorInsertGSDMD (GSDMD Human)
TagsflagExpressionMammalianMutationconstruct only codes for amino acids 29-484Available SinceJuly 1, 2025AvailabilityIndustry, Academic Institutions, and Nonprofits -
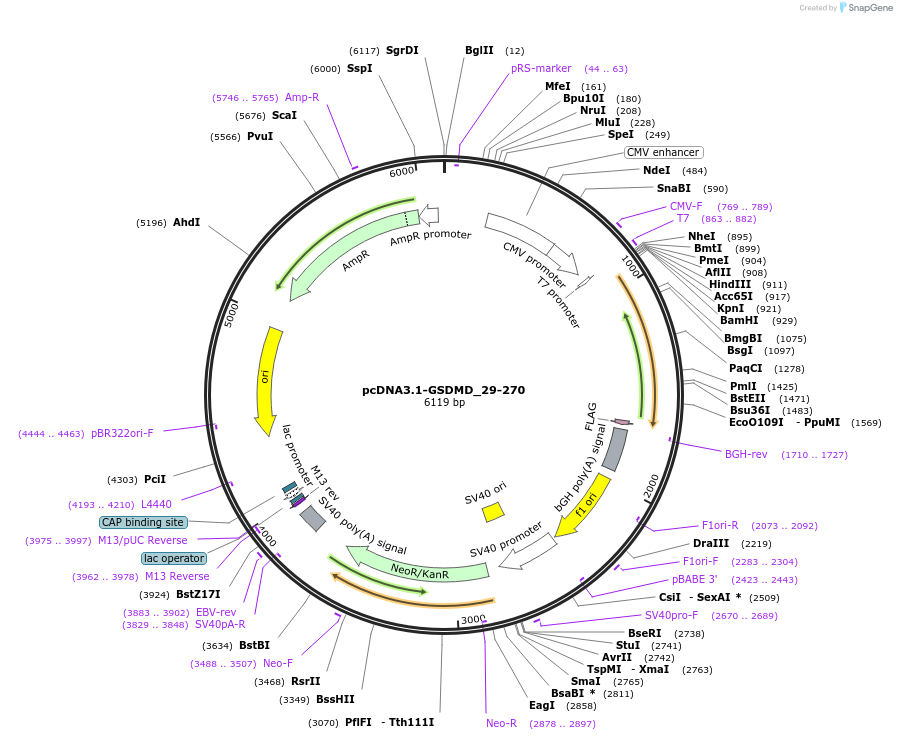
pcDNA3.1-GSDMD_29-270
Plasmid#218232PurposeGasdermin-D_29-270 is a non-functional GSDMD formed by two SARS-CoV-2 3CLpro cleavages. Transfection in HEK293 cells showed no pore-forming activity or induction of pyroptosisDepositorInsertGSDMD (GSDMD Human)
TagsflagExpressionMammalianMutationconstruct only codes for amino acids 29-270Available SinceJuly 1, 2025AvailabilityIndustry, Academic Institutions, and Nonprofits -
pMiR-CIP2A-stop-mut-miR-375-mutA-E
Plasmid#53714PurposeLuciferase reporter assay for CIP2A with mutated firefly stop codon that also has five miR-375 binding site mutationsDepositorInsertCIP2A (CIP2A Human)
UseLuciferaseMutationMutation on the stop codon of firefly luciferase …Available SinceJuly 2, 2014AvailabilityAcademic Institutions and Nonprofits only -
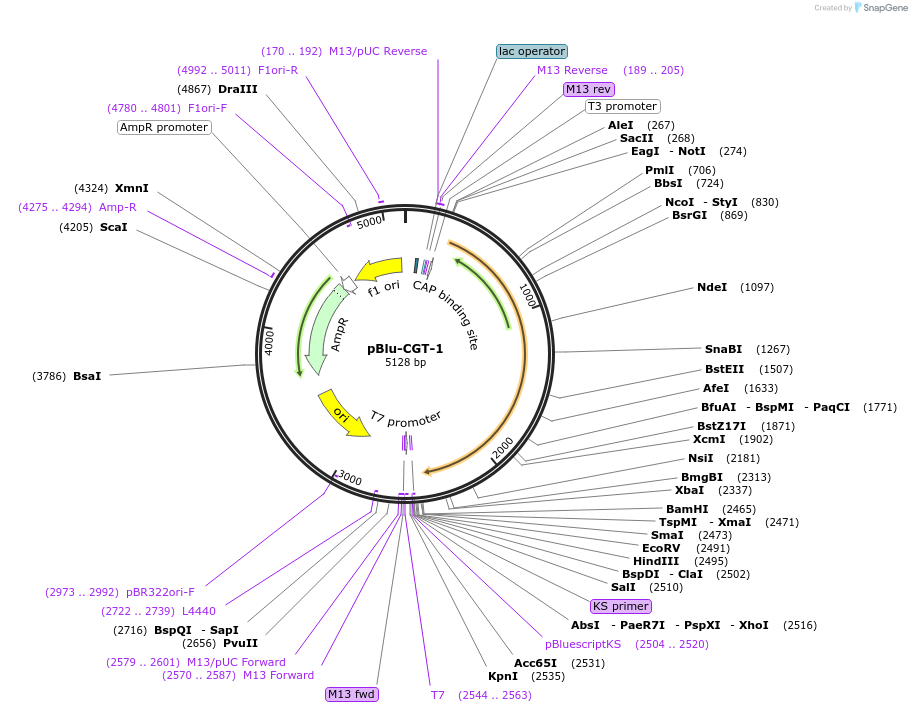
pBlu-CGT-1
Plasmid#170158PurposeExpresses bacterial Cyclomaltodextrin Glucanotransferase from Paenibacillus sp. c36 in E.coliDepositorInsertCyclomaltodextrin Glucanotransferase
ExpressionBacterialPromoterTacAvailable SinceMay 10, 2023AvailabilityAcademic Institutions and Nonprofits only -
Syn61Δ3(ev5) ΔrecA (ev1)
Bacterial Strain#189857PurposeLaboratory adapted, genomically recoded E. coli strain without TCG, TCA, or TAG codons and deleted serT, serU, prfA, and recA genes. Evolved for ~270 generations in LB at 37 °C.DepositorBacterial ResistanceStreptomycinSpeciesSyntheticAvailable SinceJan. 27, 2023AvailabilityAcademic Institutions and Nonprofits only -
MG1655-tMMR
Bacterial Strain#62392PurposeE. coli MG1655 temperature-sensitive mismatch repair deficient strain, harboring mutS(A134V) mutL(G62S), and nadA::Tn10 pglΔ8 [λ cI857 Δ(cro-bioA)]} for λ Red expressionDepositorBacterial ResistanceTetracyclineSpeciesEscherichia coliAvailable SinceMarch 4, 2015AvailabilityAcademic Institutions and Nonprofits only