We narrowed to 832 results for: HL
-
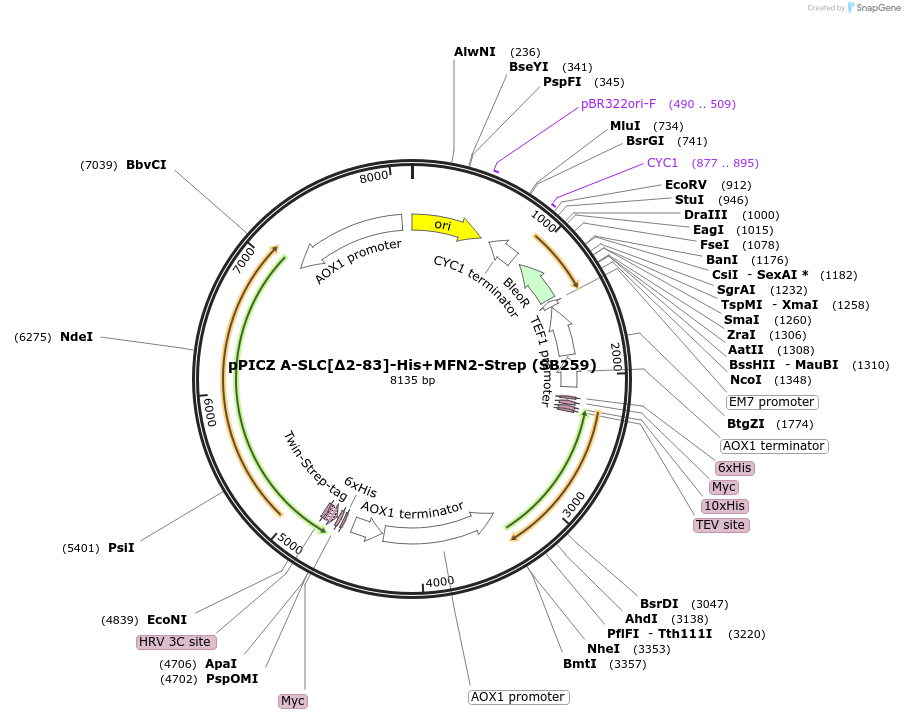
Plasmid#227608PurposeInducible coexpression of His-tagged SLC25A46 in its N-terminal region (∆2-83) and Strep-tagged MFN2 in Pichia pastorisDepositorTags10xHis and Twin-StrepTagExpressionYeastMutationaa 2-83 deletedPromoterAOX1Available SinceDec. 6, 2024AvailabilityAcademic Institutions and Nonprofits only
-
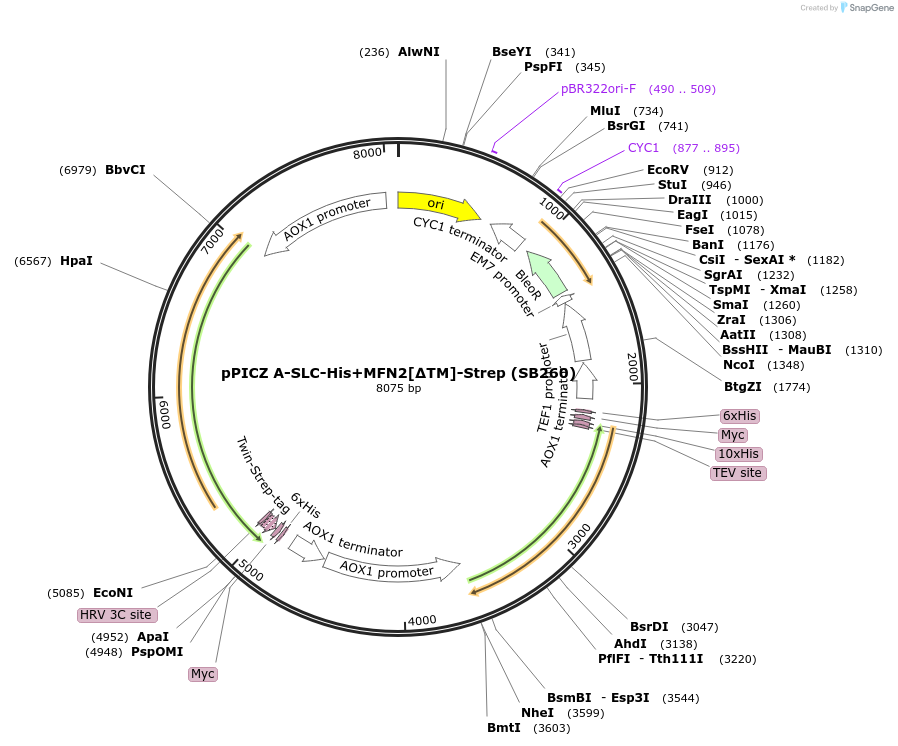
pPICZ A-SLC-His+MFN2[∆TM]-Strep (SB260)
Plasmid#227609PurposeInducible coexpression of His-tagged SLC25A46 and Strep-tagged MFN2 lacking its transmembrane region in Pichia pastorisDepositorTags10xHis and Twin-StrepTagExpressionYeastMutationcontains aa 2-544, 651-757 only [TM deleted]PromoterAOX1Available SinceDec. 6, 2024AvailabilityAcademic Institutions and Nonprofits only -
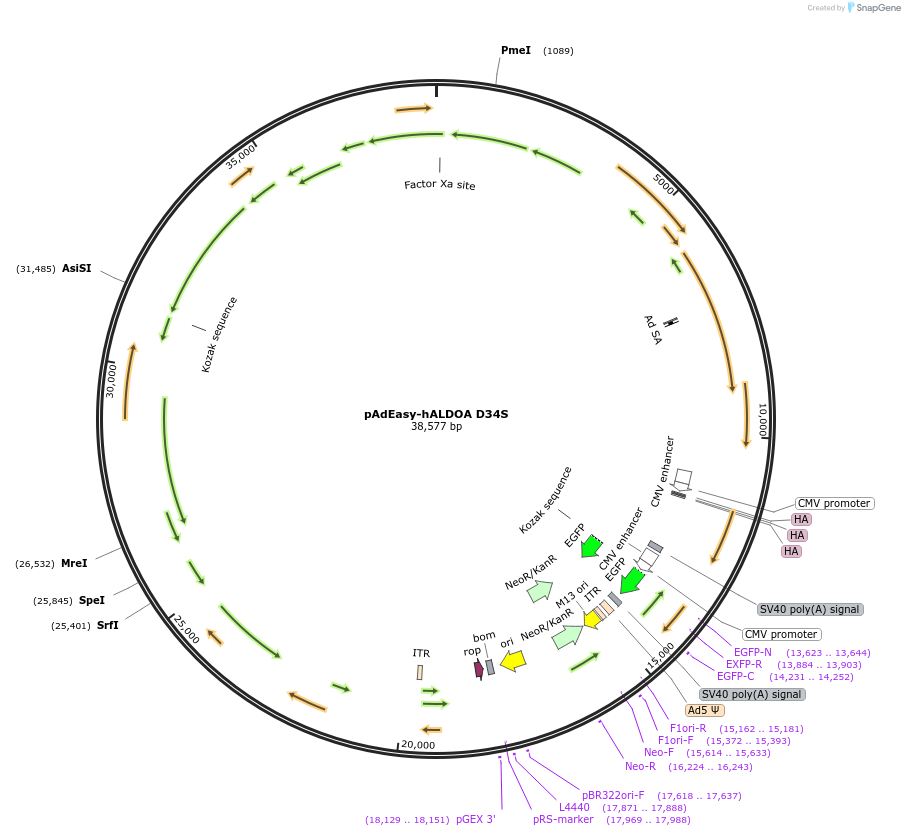
pAdEasy-hALDOA D34S
Plasmid#210813PurposeTo reduce the catalytic constant (kcat) for conversion of FBP to dihydroxyacetone phosphate (DHAP) and glyceraldehyde-3-phosphate (G3P)DepositorInsertALDOA (ALDOA Human)
UseAdenoviralTagsHAMutationresidues 34 of ALDOA mutated to serinePromoterCMVAvailable SinceDec. 22, 2023AvailabilityAcademic Institutions and Nonprofits only -
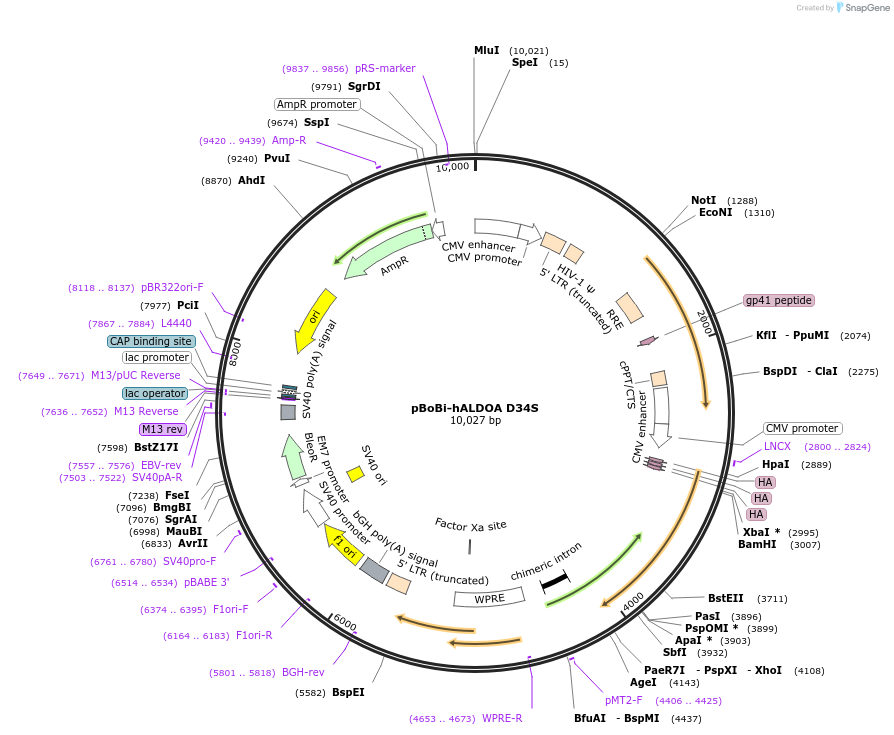
pBoBi-hALDOA D34S
Plasmid#210807PurposeTo reduce the catalytic constant (kcat) for conversion of FBP to dihydroxyacetone phosphate (DHAP) and glyceraldehyde-3-phosphate (G3P)DepositorInsertALDOA (ALDOA Human)
UseLentiviralTagsHAMutationresidues 34 of ALDOA mutated to serinePromoterCMVAvailable SinceDec. 22, 2023AvailabilityAcademic Institutions and Nonprofits only -
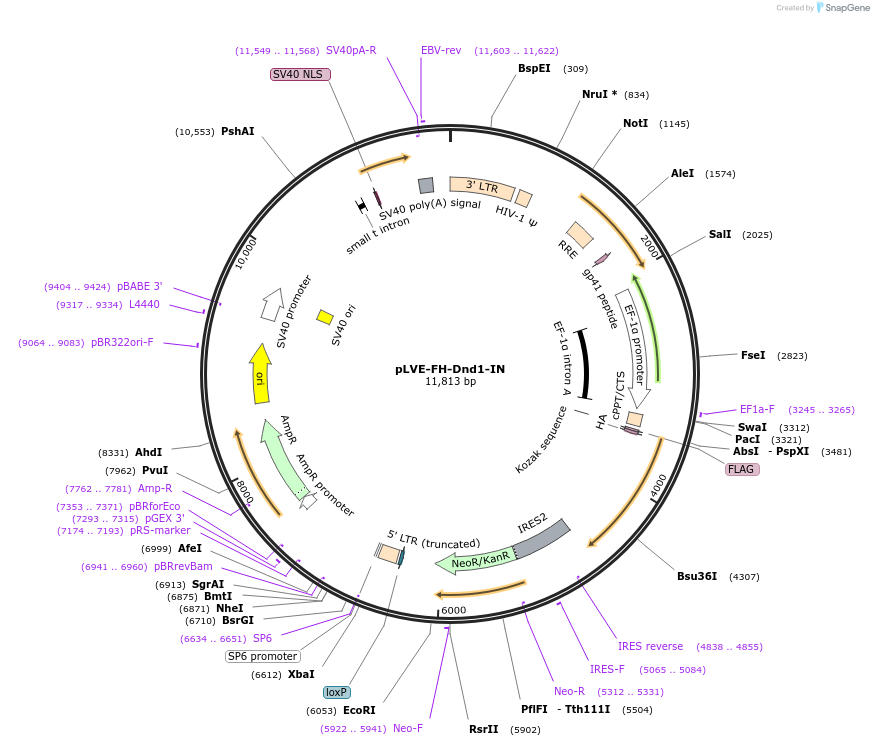
pLVE-FH-Dnd1-IN
Plasmid#70072Purpose2nd gen lentiviral vector. Cre-removable and tet/dox-controllable expression of FLAG-HA-tagged murine Dnd1 in mammalian cells.DepositorInsertDnd1 (Dnd1 Mouse)
UseCre/Lox and LentiviralTagsFLAG and HAExpressionMammalianPromoterEF1alphaAvailable SinceNov. 22, 2017AvailabilityAcademic Institutions and Nonprofits only -
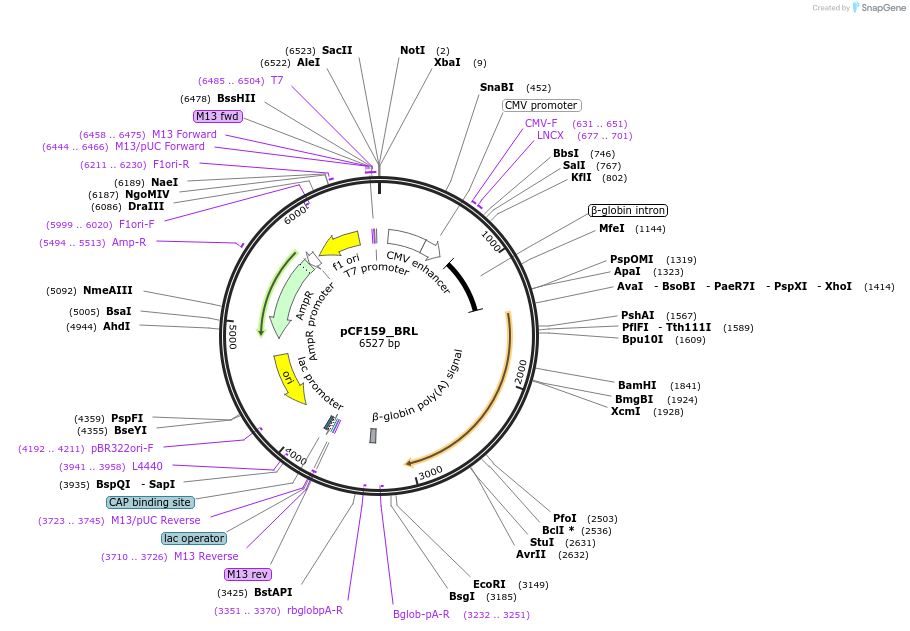
pCF159_BRL
Plasmid#225962PurposeCMV-Intron-BRL (env protein). Expresses BRL (BaEVRLess) env for VLP production.DepositorInsertCMV-Intron-BRL (env protein)
ExpressionMammalianAvailable SinceOct. 30, 2024AvailabilityAcademic Institutions and Nonprofits only -
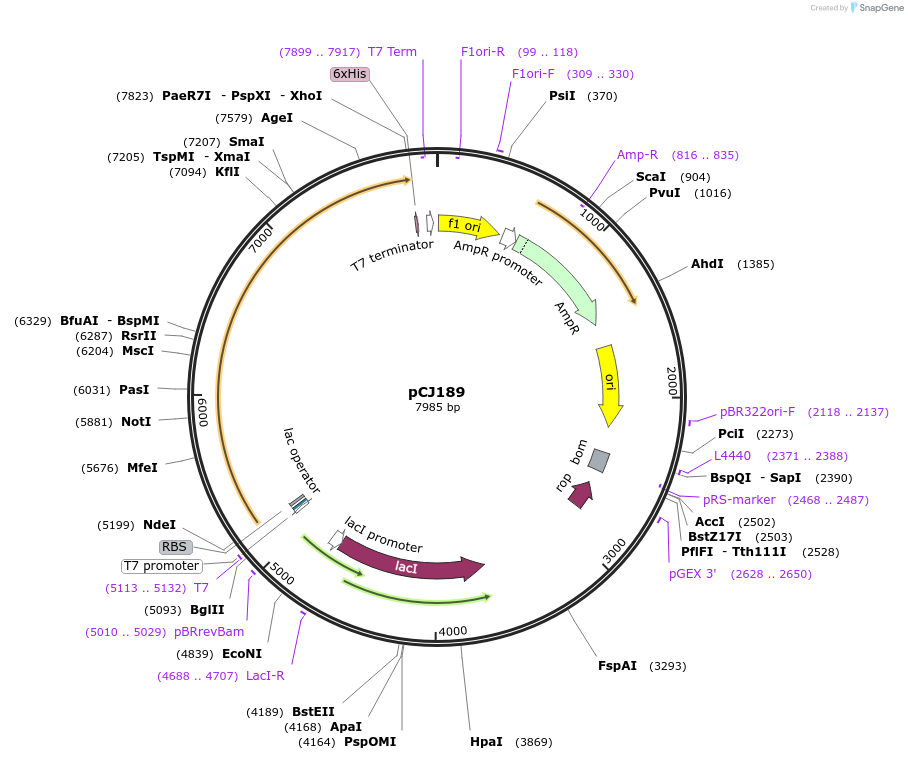
pCJ189
Plasmid#162666PurposepET-21b(+) based plasmid for expresion of MHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by an 8 residue Gly/Ser linker; codon optimized for expression in E. coli K12, with C-terminal His tag.DepositorInsertMHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by an 8 residue Gly/Ser linker
TagsHisExpressionBacterialMutationcodon optimized for expression in E. coli K12PromoterT7/lacAvailable SinceFeb. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
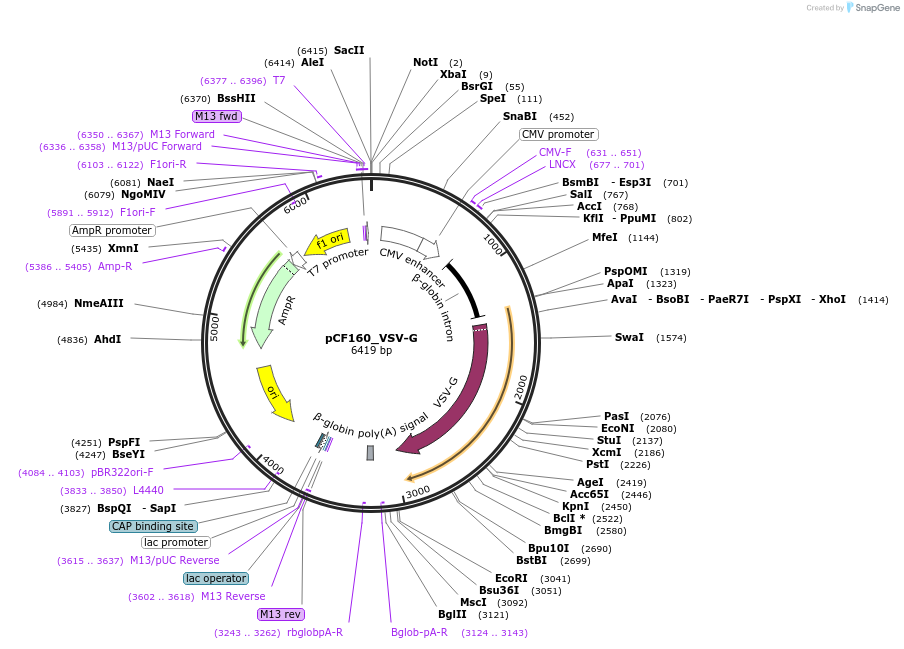
pCF160_VSV-G
Plasmid#225963PurposeCMV-Intron-VSVG (env protein). Expresses VSV-G env for VLP production.DepositorInsertCMV-Intron-VSVG (env protein)
ExpressionMammalianAvailable SinceOct. 30, 2024AvailabilityAcademic Institutions and Nonprofits only -
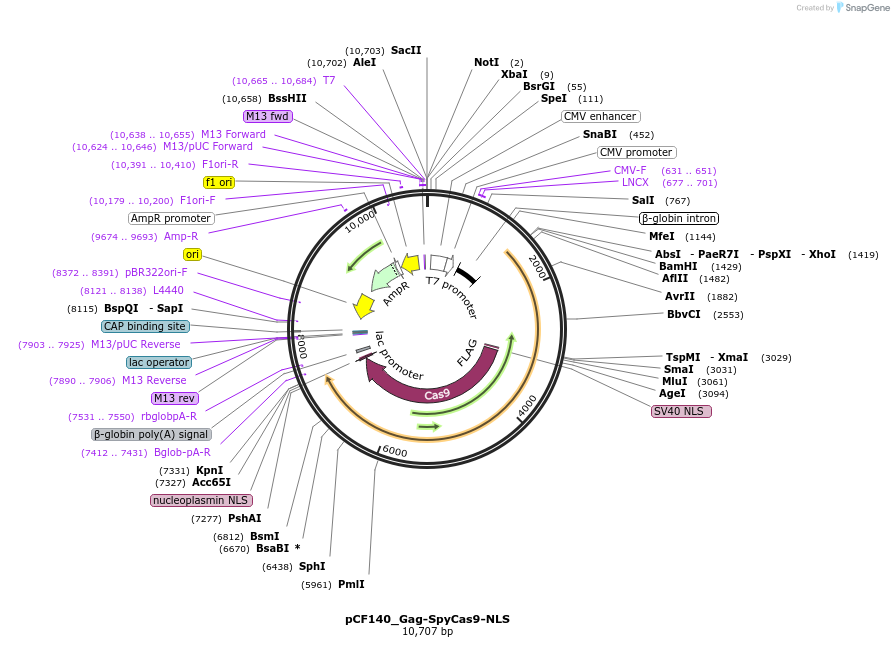
pCF140_Gag-SpyCas9-NLS
Plasmid#225959PurposeCMV-Intron-Gag-SpyCas9-NLS. Expresses Gag-SpyCas9-NLS for VLP production.DepositorInsertCMV-Intron-Gag-SpyCas9-NLS
ExpressionMammalianAvailable SinceOct. 30, 2024AvailabilityAcademic Institutions and Nonprofits only -
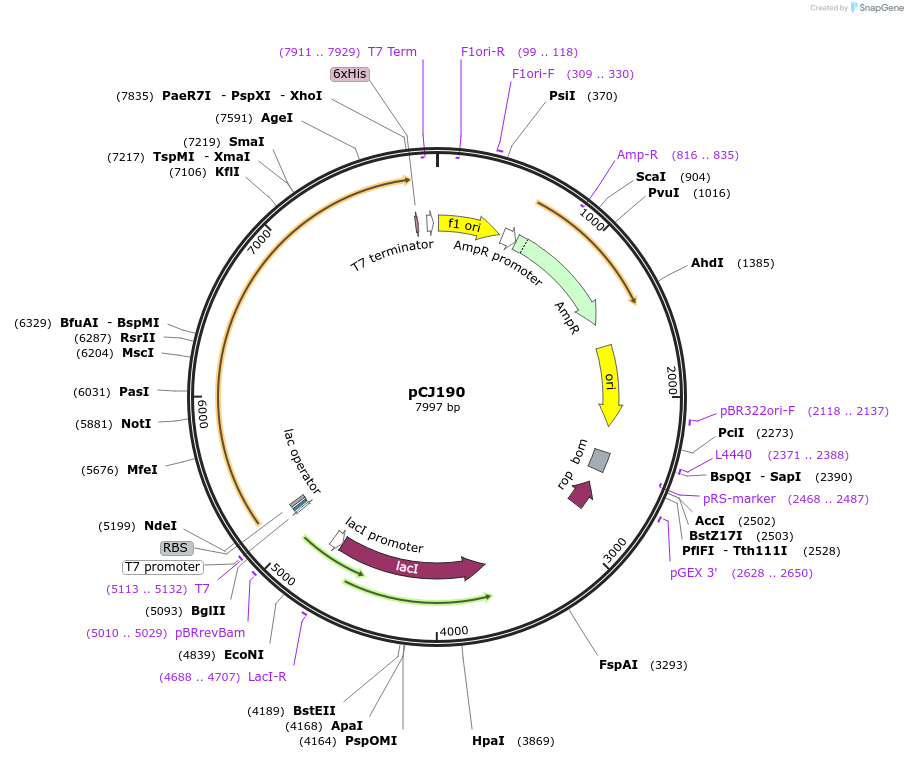
pCJ190
Plasmid#162667PurposepET-21b(+) based plasmid for expresion of MHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by a 12 residue Gly/Ser linker; codon optimized for expression in E. coli K12, with C-terminal His tag.DepositorInsertMHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by a 12 residue Gly/Ser linker
TagsHisExpressionBacterialMutationcodon optimized for expression in E. coli K12PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
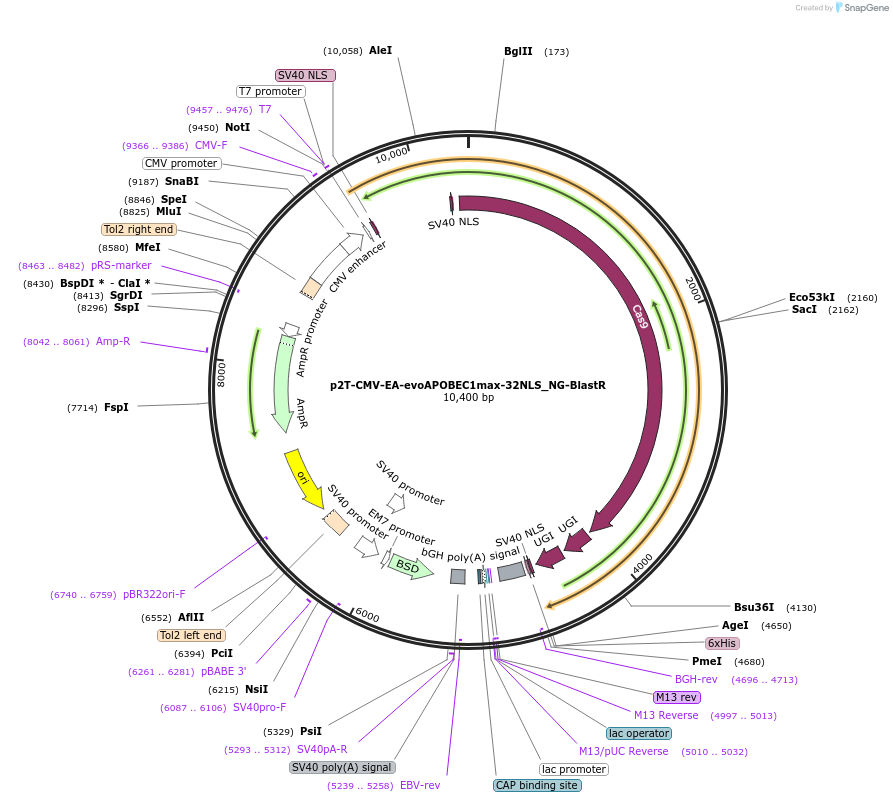
p2T-CMV-EA-evoAPOBEC1max-32NLS_NG-BlastR
Plasmid#232722PurposeC-to-T base editorDepositorInsertevoAPOBEC1max H47E+S48A-32NLS linker-nCas9(NG)-UGI-UGI
ExpressionMammalianMutationwithin evoAPOBEC1: H47E+S48A; within Cas9 D10AAvailable SinceMarch 3, 2025AvailabilityAcademic Institutions and Nonprofits only -
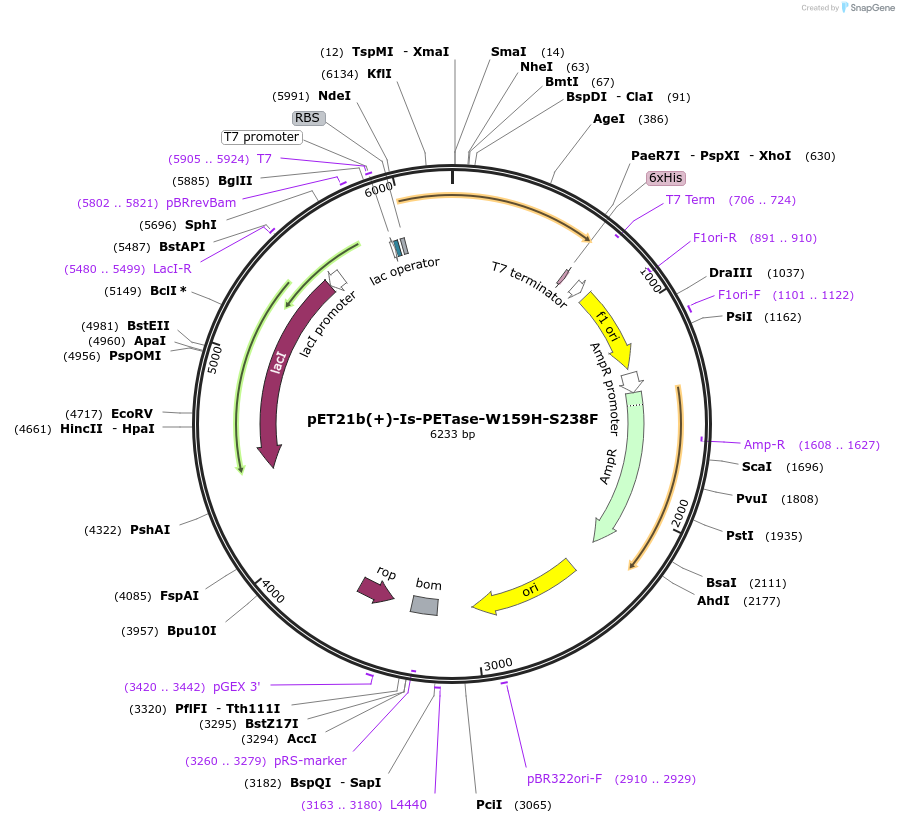
pET21b(+)-Is-PETase-W159H-S238F
Plasmid#112203PurposepET-21b(+) based plasmid for expression of PETase from Ideonella sakaiensis 201-F6 (Genbank GAP38373.1) with W159H and S238F mutations, codon optimized for expression in E. coli K12DepositorInsertPETase gene from Ideonella sakaiensis 201-F6 with W159H and S238F mutations, codon optimized for expression in E. coli K12
Tags6XHISExpressionBacterialMutationW159H, S238FPromoterN/AAvailable SinceJuly 30, 2018AvailabilityAcademic Institutions and Nonprofits only -
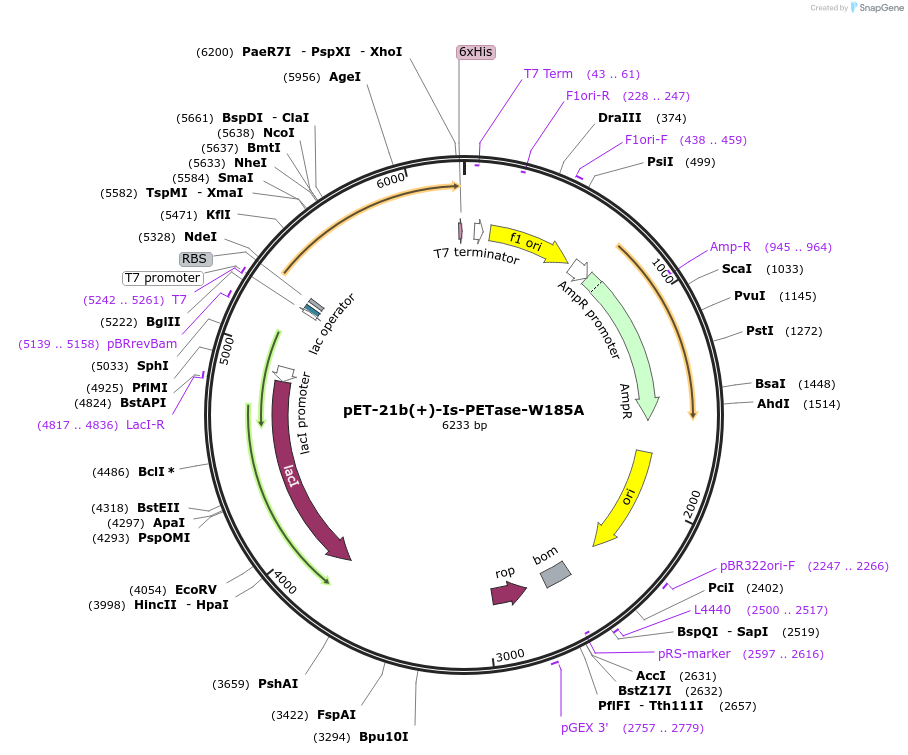
pET-21b(+)-Is-PETase-W185A
Plasmid#112204PurposepET-21b(+) based plasmid for expression of PETase from Ideonella sakaiensis 201-F6 (Genbank GAP38373.1) with W185A mutations, codon optimized for expression in E. coli K12DepositorInsertPETase gene from Ideonella sakaiensis 201-F6 with W185A mutation, codon optimized for expression in E. coli K12
Tags6XHISExpressionBacterialMutationW185APromoterN/AAvailable SinceJuly 18, 2018AvailabilityAcademic Institutions and Nonprofits only -
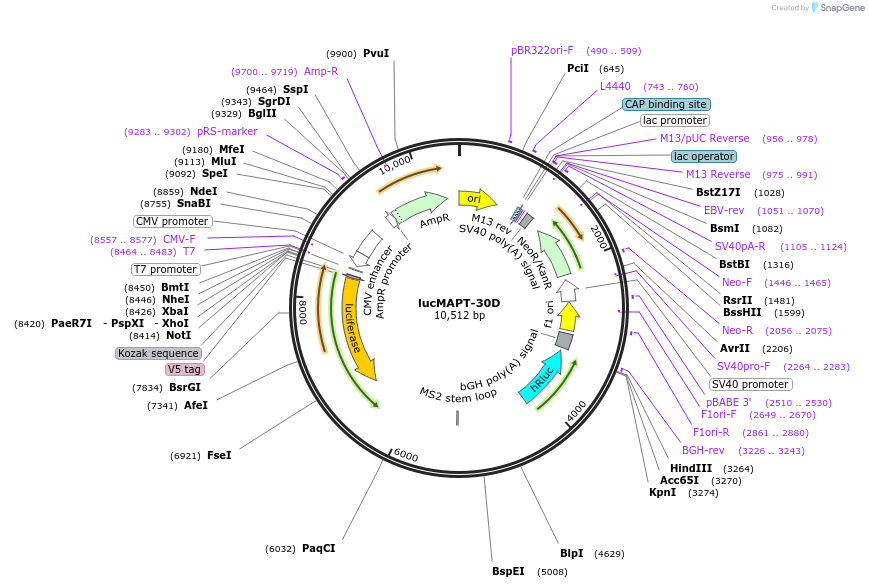
lucMAPT-30D
Plasmid#214672PurposeMAPT Exon 10 reporter containing the MS2 hairpin 30 base pairs downstream of the 5′ splice siteDepositorInsertMAPT (MAPT Human)
TagsFirefly Luciferase, Renilla Luciferase, and V5ExpressionMammalianMutationContains Exon 9, Exon 10 with a mutation to intro…PromoterCMVAvailable SinceMarch 15, 2024AvailabilityAcademic Institutions and Nonprofits only -
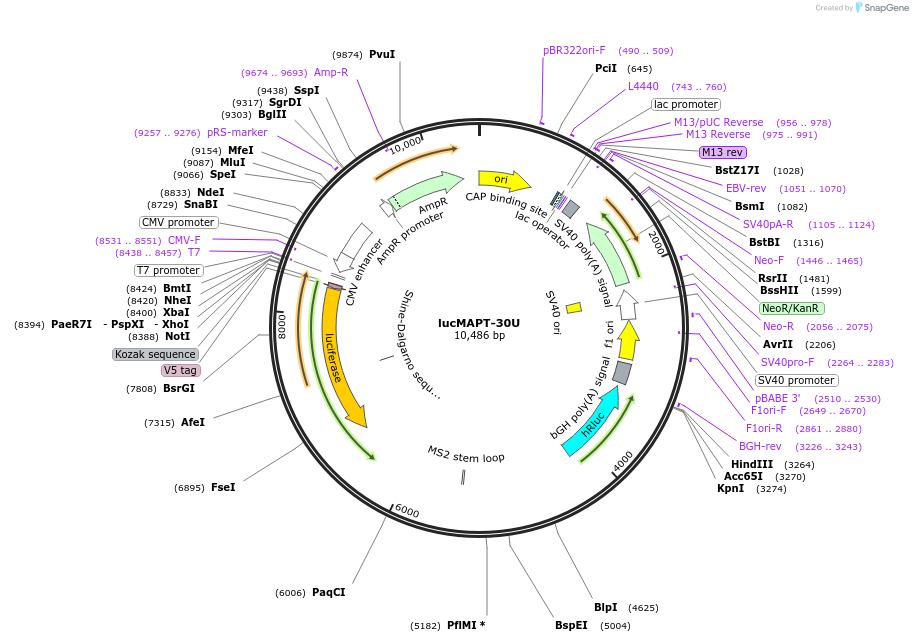
lucMAPT-30U
Plasmid#214673PurposeMAPT Exon 10 reporter containing the MS2 hairpin 30 base pairs upstream of the 3′ splice siteDepositorInsertMAPT (MAPT Human)
TagsFirefly Luciferase, Renilla Luciferase, and V5ExpressionMammalianMutationContains Exon 9, Exon 10 with a mutation to intro…PromoterCMVAvailable SinceMarch 15, 2024AvailabilityAcademic Institutions and Nonprofits only -
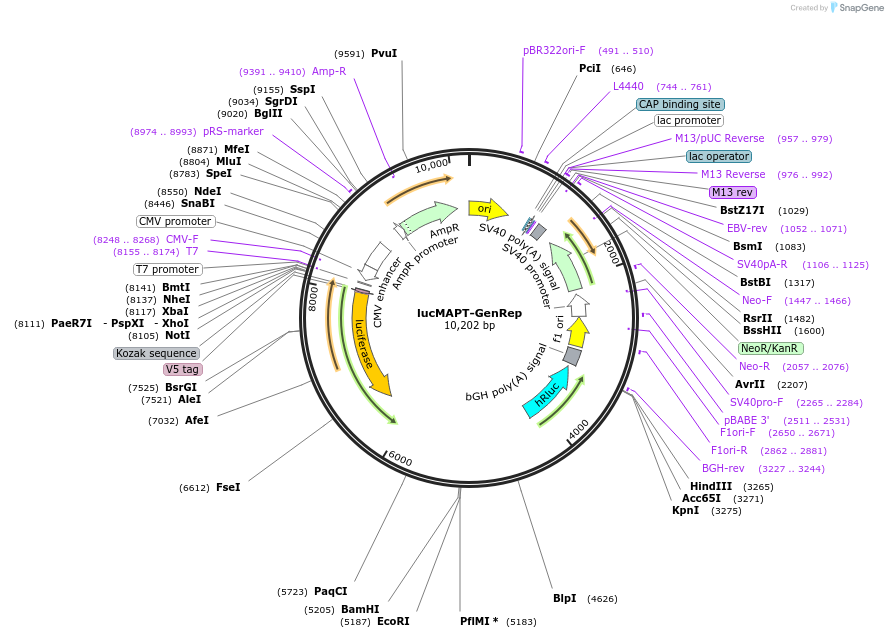
lucMAPT-GenRep
Plasmid#214674PurposeMAPT Exon 10 reporter, with Exon 10 and 100bp flanking intronic regions replaced with a BamHI/EcoRI cloning site for testing different exonsDepositorInsertMAPT (MAPT Human)
TagsFirefly Luciferase, Renilla Luciferase, and V5ExpressionMammalianMutationContains Exon 9 and Exon 11, with a BamHI EcoRI c…PromoterCMVAvailable SinceMarch 15, 2024AvailabilityAcademic Institutions and Nonprofits only -
mCardinal-Farnesyl-5
Plasmid#56159PurposeLocalization: Plasma Membrane, Excitation: 604, Emission: 659DepositorInsertFarnesyl (HRAS Human)
TagsmCardinalExpressionMammalianMutationencodes final 20 aa of NM_001130442.1PromoterCMVAvailable SinceFeb. 27, 2015AvailabilityIndustry, Academic Institutions, and Nonprofits -
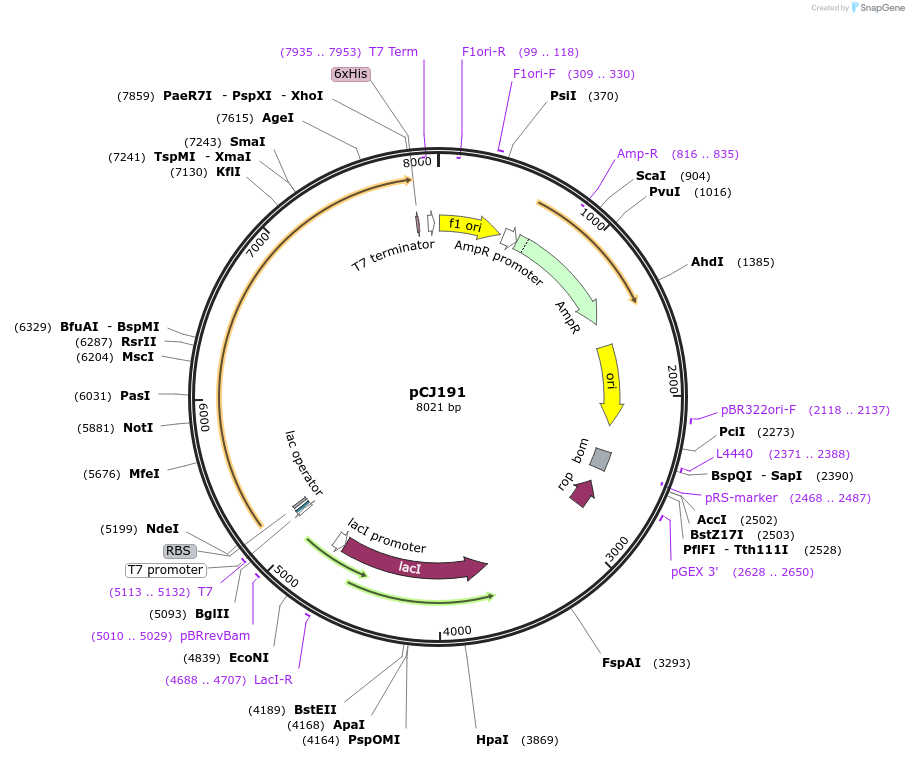
pCJ191
Plasmid#162668PurposepET-21b(+) based plasmid for expresion of MHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by a 20 residue Gly/Ser linker; codon optimized for expression in E. coli K12, with C-terminal His tag.DepositorInsertMHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by a 20 residue Gly/Ser linker
TagsHisExpressionBacterialMutationcodon optimized for expression in E. coli K12PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
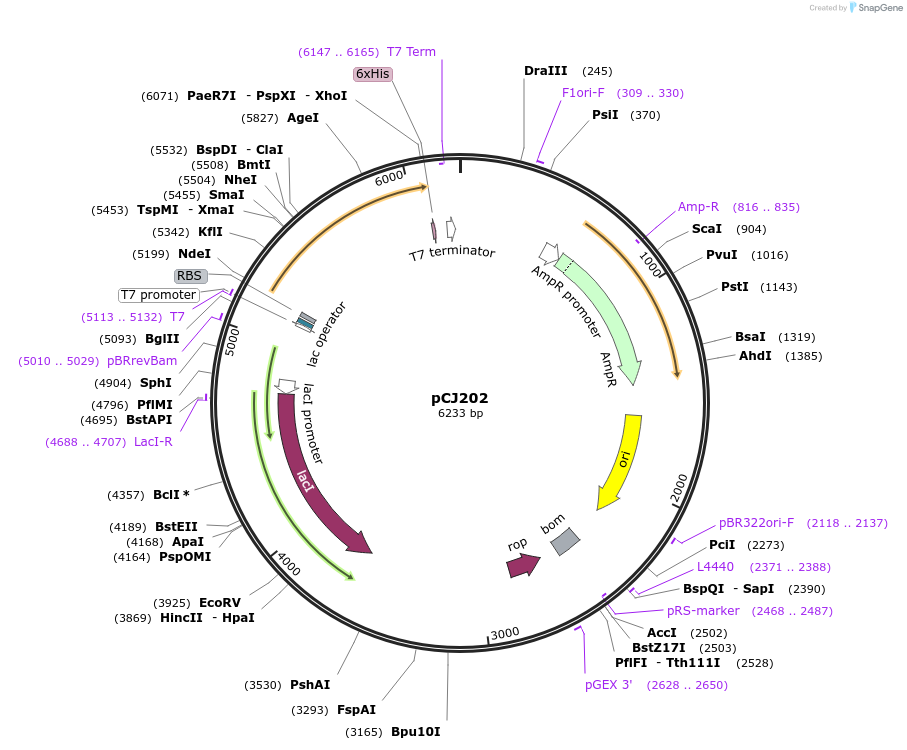
pCJ202
Plasmid#162675PurposepET-21b(+) based plasmid for expression of PETase from Ideonella sakaiensis 201-F6 (Genbank GAP38373.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating W159C and S238C mutations.DepositorInsertPETase from Ideonella sakaiensis 201-F6 (Genbank GAP38373.1)
TagsHisExpressionBacterialMutationW159C, S238C; codon optimized for expression in E…PromoterT7/lacAvailable SinceFeb. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
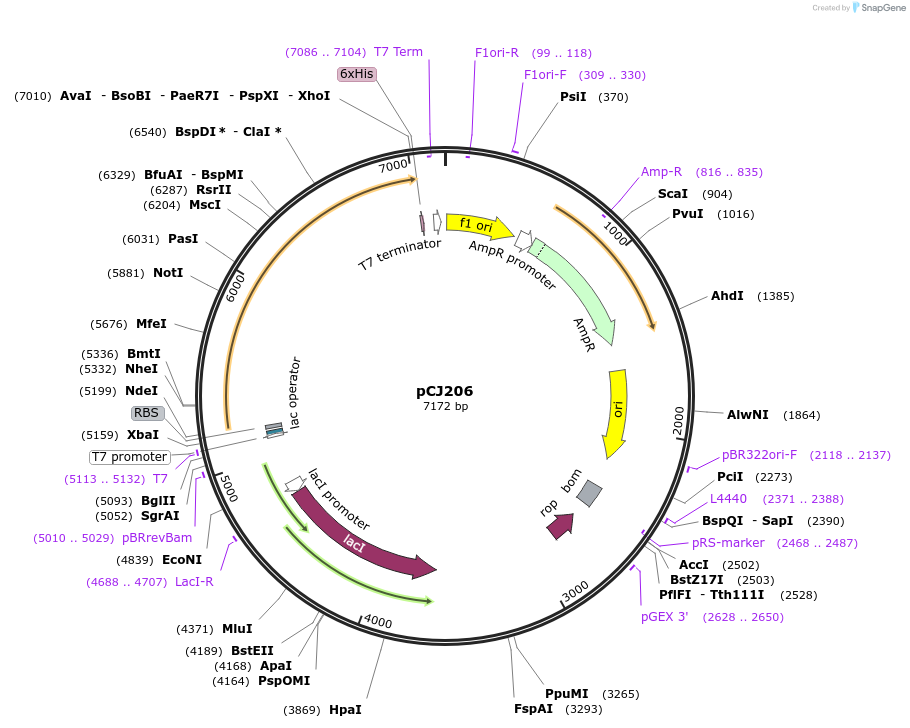
pCJ206
Plasmid#162679PurposepET-21b(+) based plasmid for expression of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating the E226T mutation to the putative lipase box.DepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), incorporating the E226T mutation to the putative lipase box
TagsHisExpressionBacterialMutationE226T; codon optimized for expression in E. coli …PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only