We narrowed to 6,481 results for: siae
-
Plasmid#60400PurposeExpression of chimeric yeast beta tubulin (TUB2) - human beta6 tubulin Cterminal tail (TUBB1) using a GAL promoterDepositorInsertTUB2
ExpressionYeastMutationTUB2 amino acids 429 - end, replaced with human T…PromoterGALAvailable SinceMay 6, 2015AvailabilityAcademic Institutions and Nonprofits only -
pRS424-TUB2 - human TUBB3 Cterminal tail
Plasmid#60397PurposeExpression of chimeric yeast beta tubulin (TUB2) - human beta3 tubulin Cterminal tail (TUBB3) using a GAL promoterDepositorInsertTUB2
ExpressionYeastMutationC127S; TUB2 amino acids 429 - end, replaced with …PromoterGALAvailable SinceMay 6, 2015AvailabilityAcademic Institutions and Nonprofits only -
pRevLCav2FeS
Plasmid#60808Purposeto disrupt iron sulfur cluster in yeast Rev3DepositorInsertREV3 (REV3 Budding Yeast)
ExpressionYeastMutationcontains 647 C-terminal amino acids and 335 bp of…Available SinceMarch 9, 2015AvailabilityAcademic Institutions and Nonprofits only -
pRS424-TUB2 - human TUBB7 Cterminal tail
Plasmid#60399PurposeExpression of chimeric yeast beta tubulin (TUB2) - human beta7 tubulin Cterminal tail (TUBB7) using a GAL promoterDepositorInsertTUB2
ExpressionYeastMutationTUB2 amino acids 429 - end, replaced with human T…PromoterGALAvailable SinceJan. 26, 2015AvailabilityAcademic Institutions and Nonprofits only -
pRS424-TUB2 - human TUBB8 Cterminal tail
Plasmid#60401PurposeExpression of chimeric yeast beta tubulin (TUB2) - human beta8 tubulin Cterminal tail (TUBB8) using a GAL promoterDepositorInsertTUB2
ExpressionYeastMutationTUB2 amino acids 429 - end, replaced with human T…PromoterGALAvailable SinceJan. 26, 2015AvailabilityAcademic Institutions and Nonprofits only -
pRS424-TUB2 - human TUBB4a Cterminal tail
Plasmid#60402PurposeExpression of chimeric yeast beta tubulin (TUB2) - human beta4a tubulin Cterminal tail (TUBB4a) using a GAL promoterDepositorInsertTUB2
ExpressionYeastMutationTUB2 amino acids 429 - end, replaced with human T…PromoterGALAvailable SinceJan. 26, 2015AvailabilityAcademic Institutions and Nonprofits only -
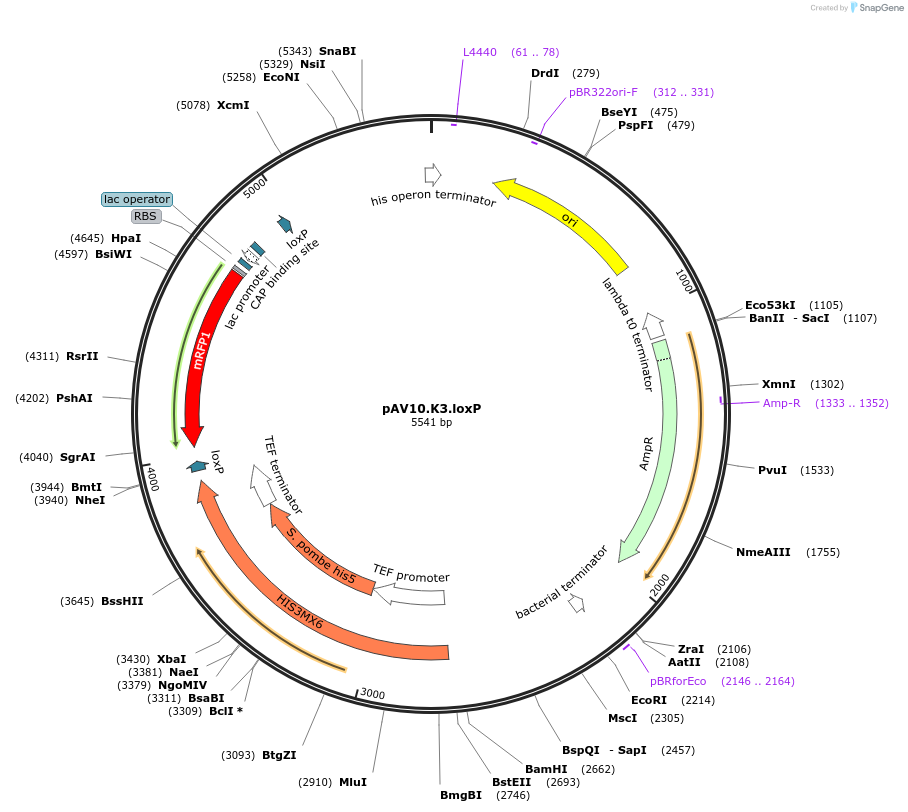
pAV10.K3.loxP
Plasmid#63202PurposepCACS backbone (Amp), SpHIS5, contains an RFP flanked by BsaI sites with CAGT & TTTT overhangs for TU assembly using yGG. LoxP flanked TU. Integration into YKL162C-A (NotI).DepositorTypeEmpty backboneUseSynthetic BiologyExpressionBacterial and YeastAvailabilityAcademic Institutions and Nonprofits only -
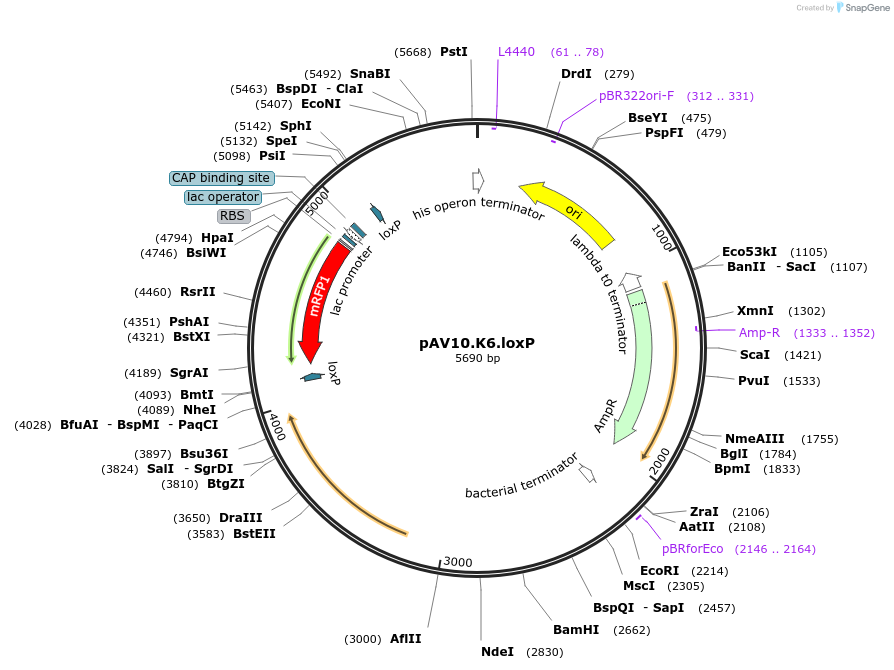
pAV10.K6.loxP
Plasmid#63203PurposepCACS backbone (Amp), KlURA3, contains an RFP flanked by BsaI sites with CAGT & TTTT overhangs for TU assembly using yGG. LoxP flanked TU. Integration into YKL162C-A (NotI or BciVI).DepositorTypeEmpty backboneUseSynthetic BiologyExpressionBacterial and YeastAvailabilityAcademic Institutions and Nonprofits only -
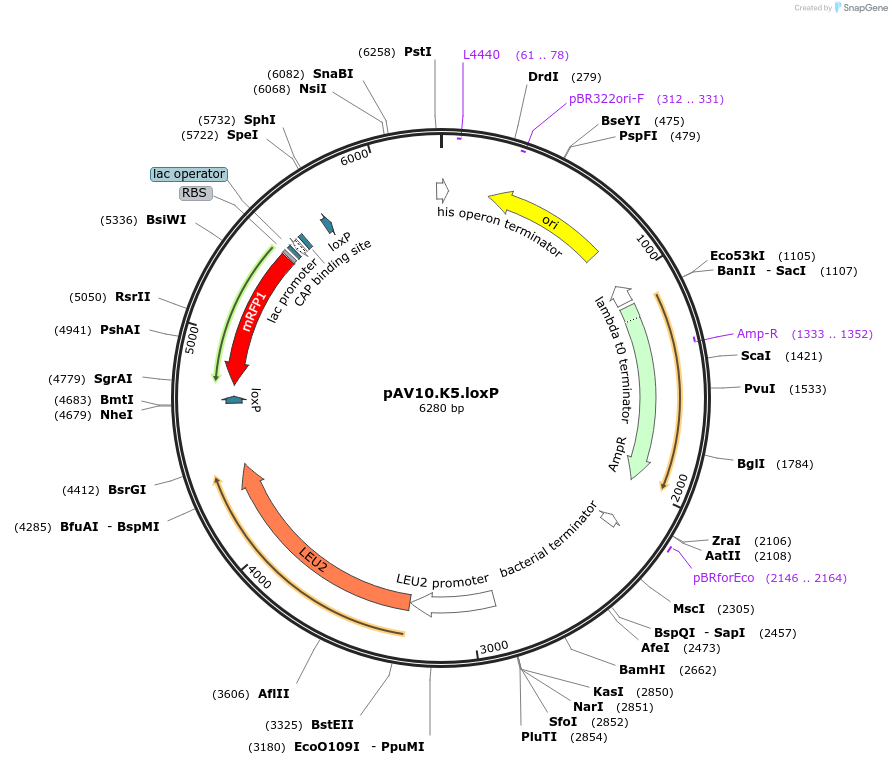
pAV10.K5.loxP
Plasmid#63204PurposepCACS backbone (Amp), LEU2, contains an RFP flanked by BsaI sites with CAGT & TTTT overhangs for TU assembly using yGG. LoxP flanked TU. Integration into YKL162C-A (NotI or BciVI).DepositorTypeEmpty backboneUseSynthetic BiologyExpressionBacterial and YeastAvailabilityAcademic Institutions and Nonprofits only -
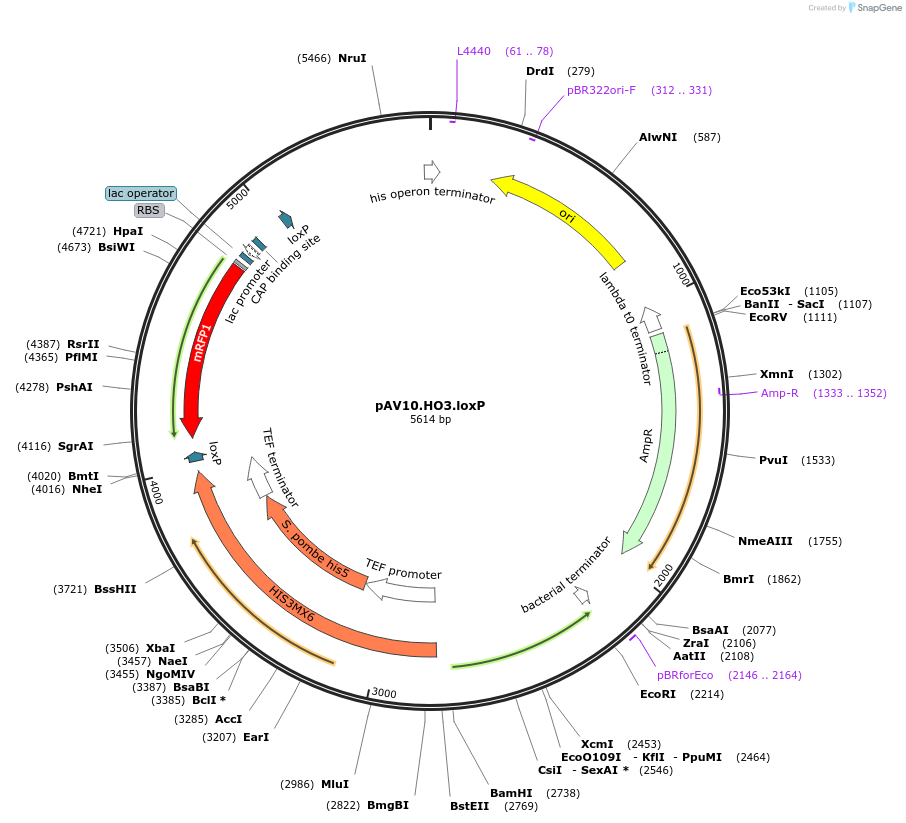
pAV10.HO3.loxP
Plasmid#63205PurposepCACS backbone (Amp), SpHIS5, contains an RFP flanked by BsaI sites with CAGT & TTTT overhangs for TU assembly using yGG. LoxP flanked TU. Integration into HO locus (NotI).DepositorTypeEmpty backboneUseSynthetic BiologyExpressionBacterial and YeastAvailabilityAcademic Institutions and Nonprofits only -
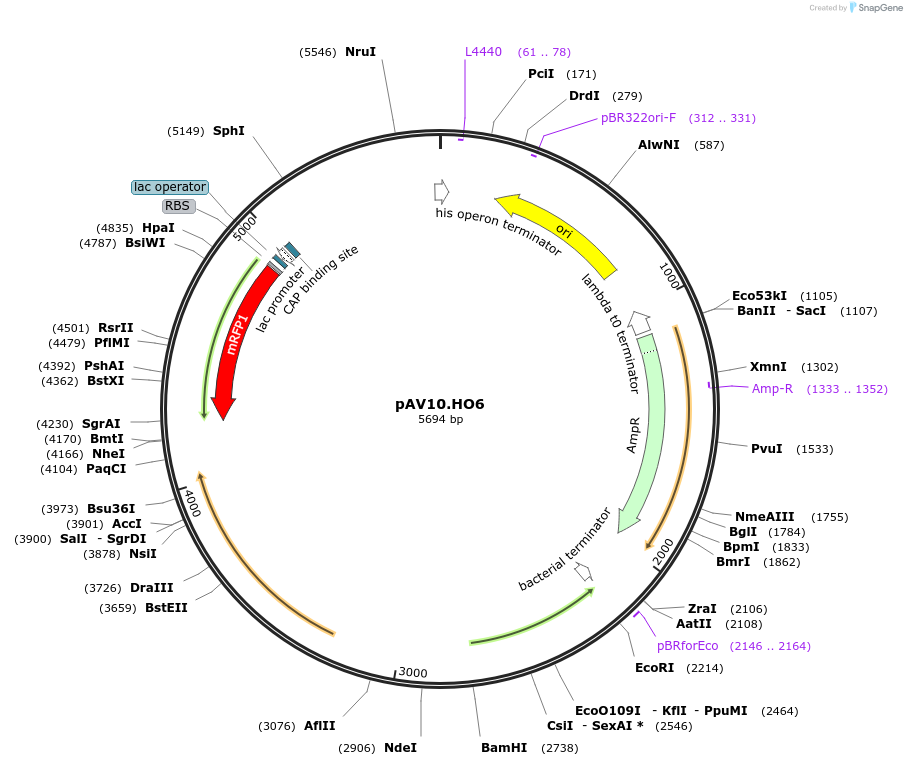
pAV10.HO6
Plasmid#63211PurposepCACS backbone (Amp), KlURA3, contains an RFP flanked by BsaI sites with CAGT & TTTT overhangs for TU assembly using yGG. Integration into HO locus (NotI or BciVI).DepositorTypeEmpty backboneUseSynthetic BiologyExpressionBacterial and YeastAvailabilityAcademic Institutions and Nonprofits only -
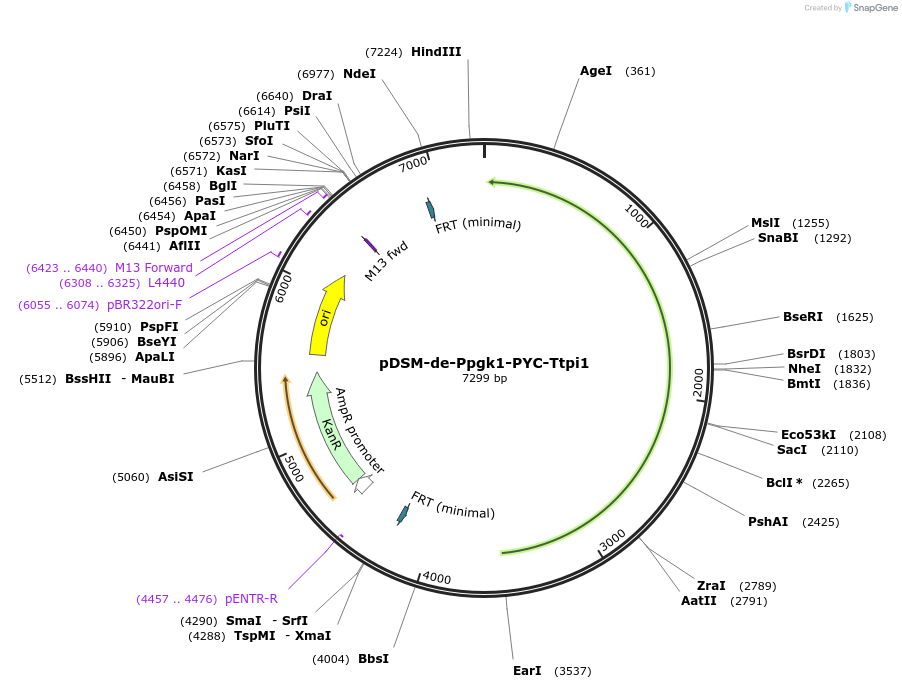
pDSM-de-Ppgk1-PYC-Ttpi1
Plasmid#127735PurposeYeast pathway position 5. PYC transcription unit with the PGK1 promoter and TPI1 terminator.DepositorInsertpyruvate carboxylase (PYC1 Budding Yeast, Synthetic)
UseSynthetic BiologyExpressionYeastPromoterPpgk1AvailabilityAcademic Institutions and Nonprofits only -
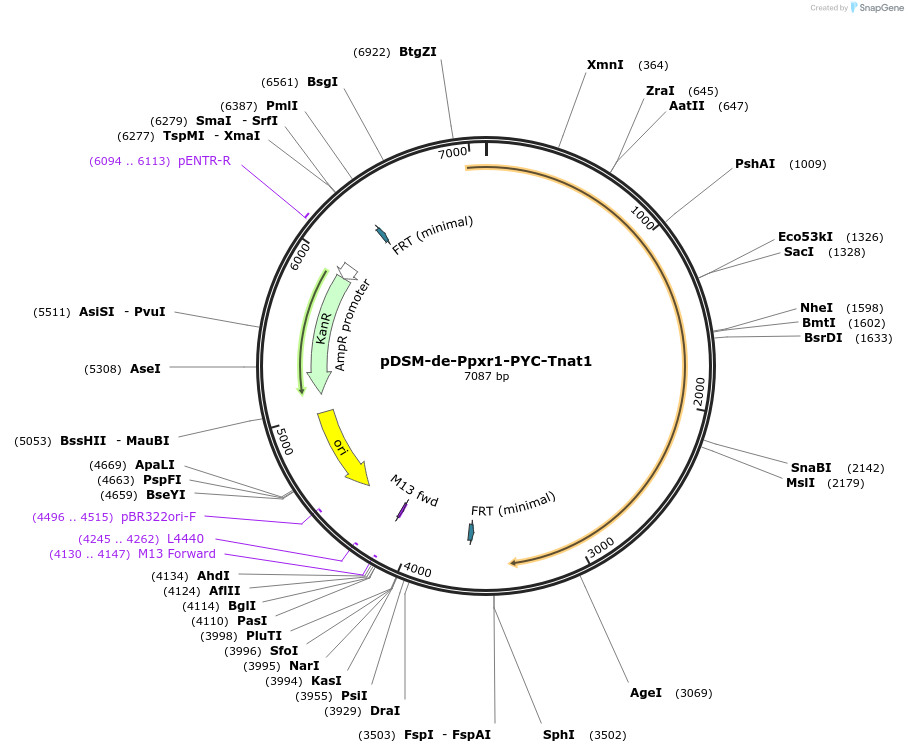
pDSM-de-Ppxr1-PYC-Tnat1
Plasmid#127736PurposeYeast pathway position 5. PYC transcription unit with the PXR1 promoter and NAT1 terminator.DepositorInsertpyruvate carboxylase (PYC1 Budding Yeast, Synthetic)
UseSynthetic BiologyExpressionYeastPromoterPpxr1AvailabilityAcademic Institutions and Nonprofits only -
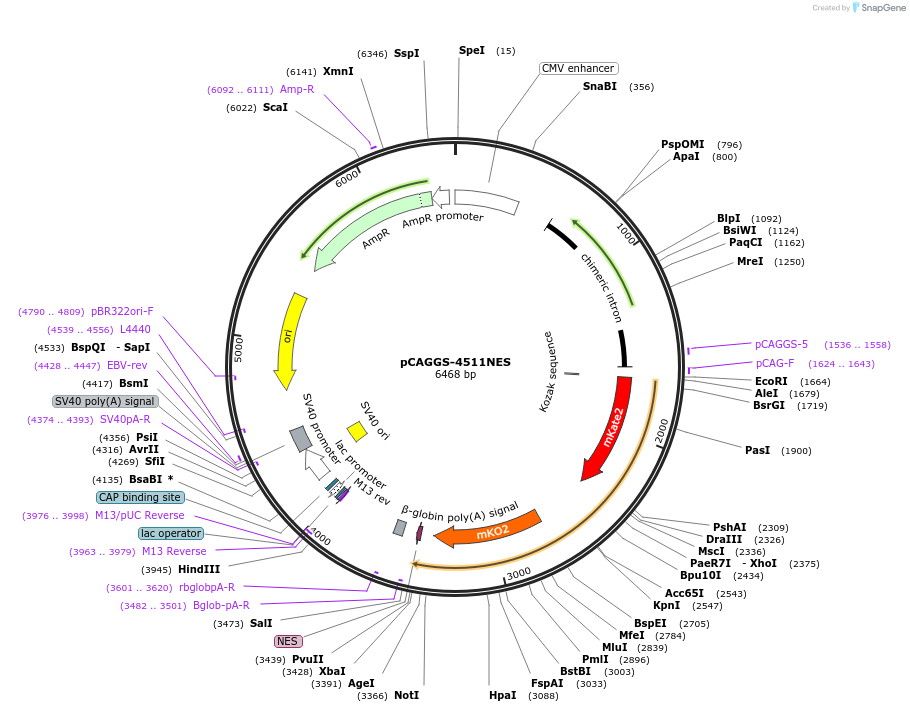
pCAGGS-4511NES
Plasmid#138375PurposeBooster-ERK, a Red-Shifted Genetically Encoded FRET Biosensor for ERK activity.DepositorInsertBooster-ERK
ExpressionMammalianPromoterCAGAvailable SinceMarch 4, 2020AvailabilityAcademic Institutions and Nonprofits only -
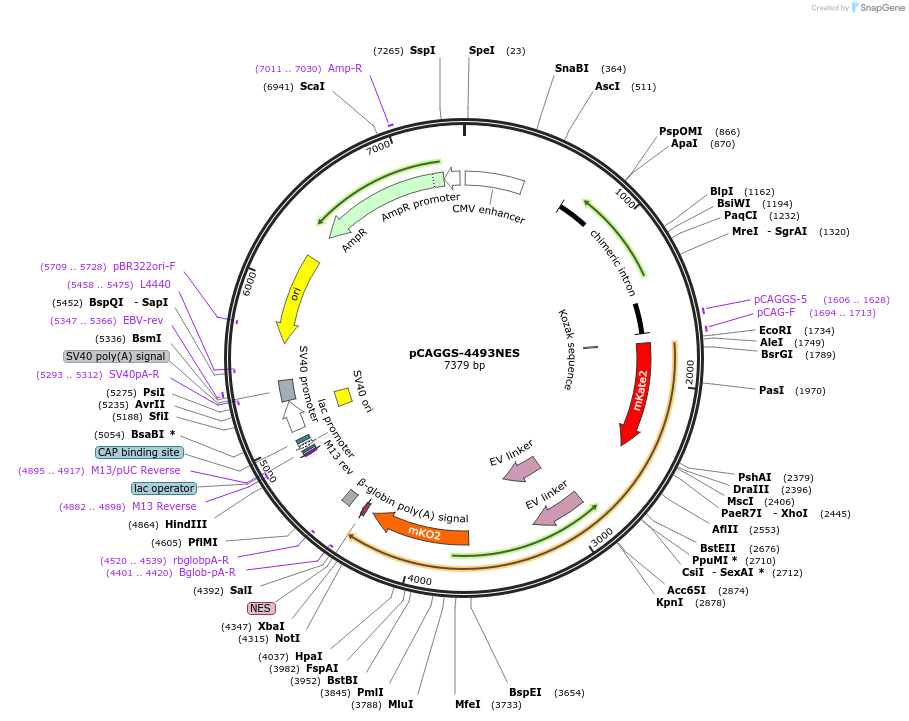
pCAGGS-4493NES
Plasmid#138373PurposeBooster-PKA, a Red-Shifted Genetically Encoded FRET Biosensor for monitoring PKA activity.DepositorInsertBooster-PKA
ExpressionMammalianPromoterCAGAvailable SinceMarch 4, 2020AvailabilityAcademic Institutions and Nonprofits only -
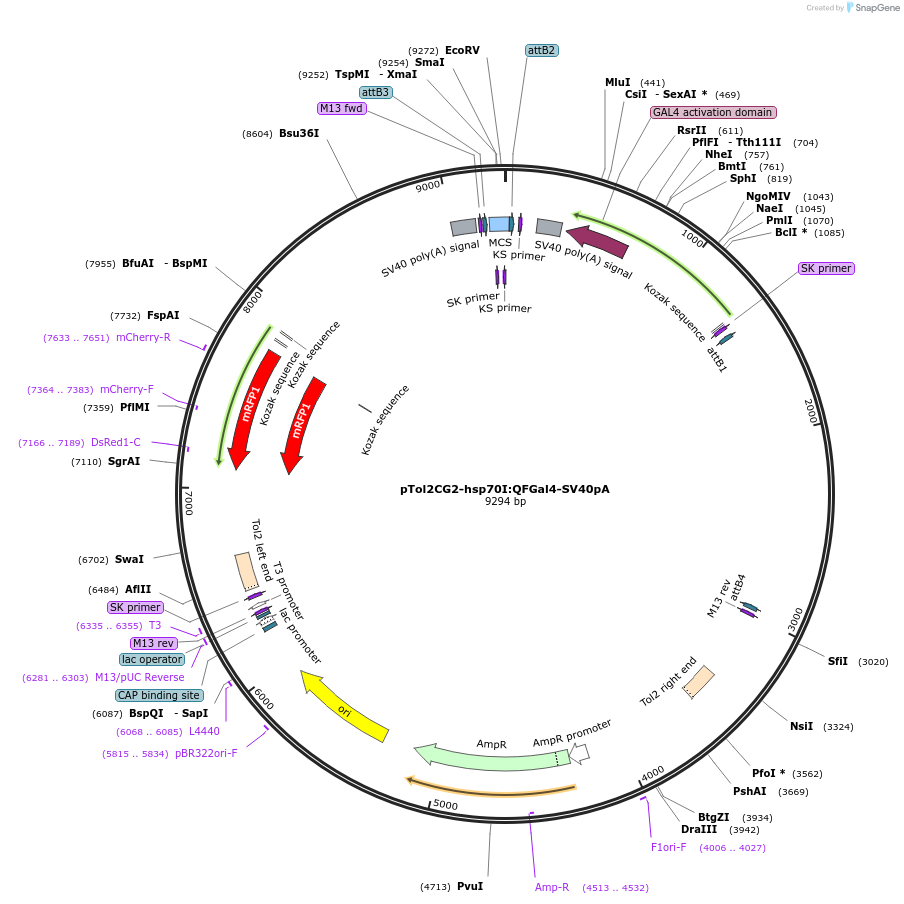
pTol2CG2-hsp70I:QFGal4-SV40pA
Plasmid#155120PurposeTol2 vector for heat-shock inducible expression of QFGal4. Contains cmlc2:mRFP-SV40pA transgenesis markerDepositorInserthsp70l:QFGal4 (hsp70l Budding Yeast, Neurospora crassa, Zebrafish)
UseZebrafish expressionPromoterhsp70lAvailable SinceSept. 14, 2020AvailabilityAcademic Institutions and Nonprofits only -
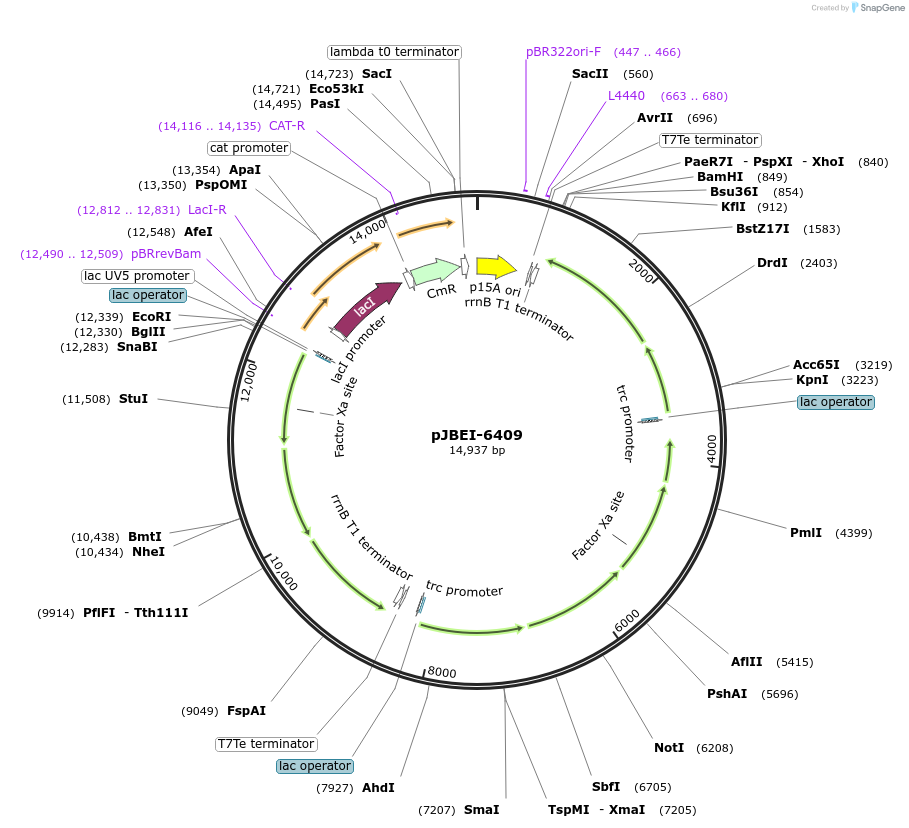
pJBEI-6409
Plasmid#47048PurposeBglBrick plasmid (=pBbA5c-MTSAe-T1f-MBI(f)-T1002i-Ptrc-trGPPS(co)-LS) coding for MEV pathway enzymes to produce limonene from glucose in E. coliDepositorInsertCodon-optimized sequences for MEV pathway expression in E. coli to produce limonene
UseSynthetic Biology; BglbrickExpressionBacterialMutationR364S mutation in MKPromoterPlacUV5Available SinceSept. 12, 2013AvailabilityAcademic Institutions and Nonprofits only -
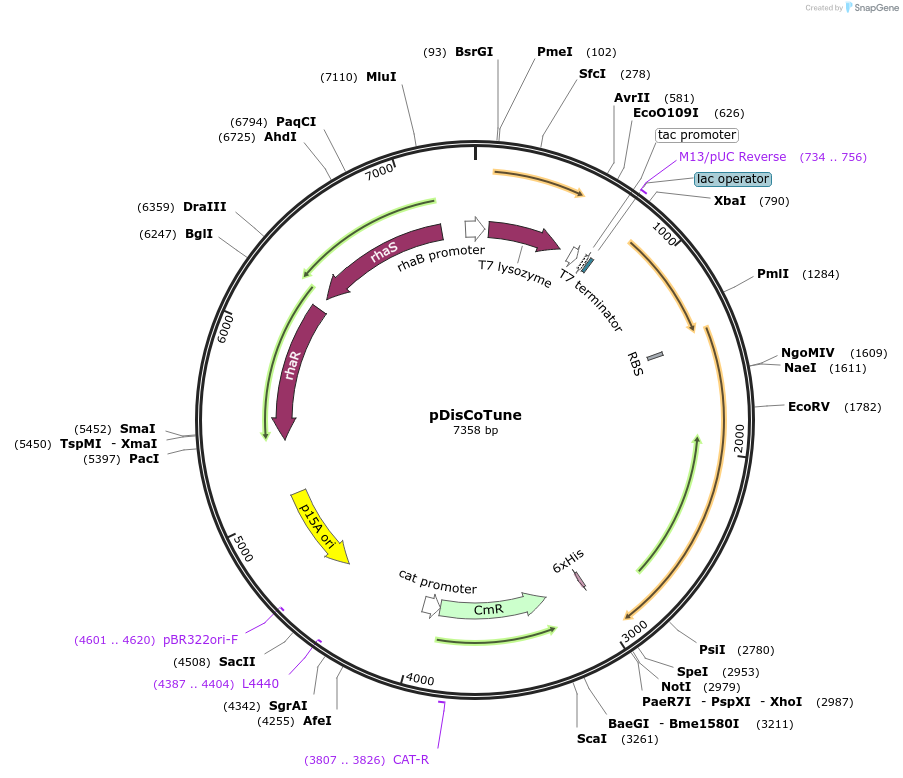
pDisCoTune
Plasmid#173713PurposePromotes expression of disulfide-rich protein in E. coli T7 expression system. The plasmid encodes expression of the folding factors Erv1p and hPDI controlled by the Ptac promoter.DepositorInsertsExpressionBacterialAvailable SinceNov. 24, 2021AvailabilityAcademic Institutions and Nonprofits only -
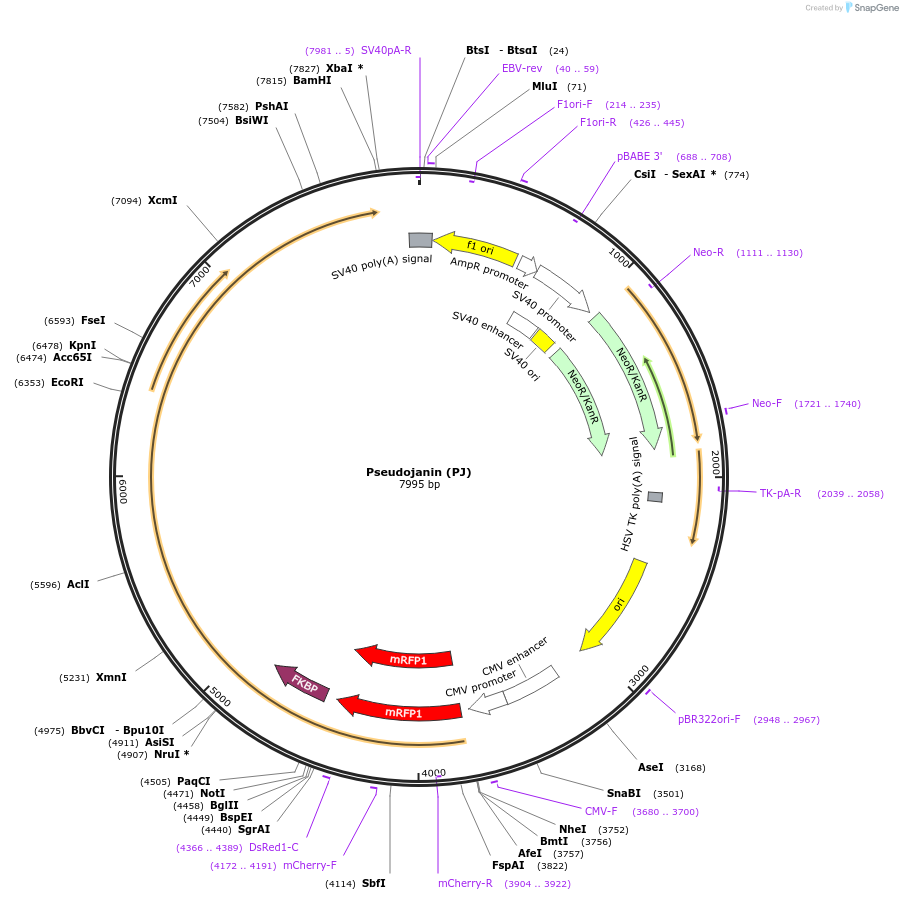
Pseudojanin (PJ)
Plasmid#37999DepositorAvailable SinceJuly 13, 2012AvailabilityAcademic Institutions and Nonprofits only -
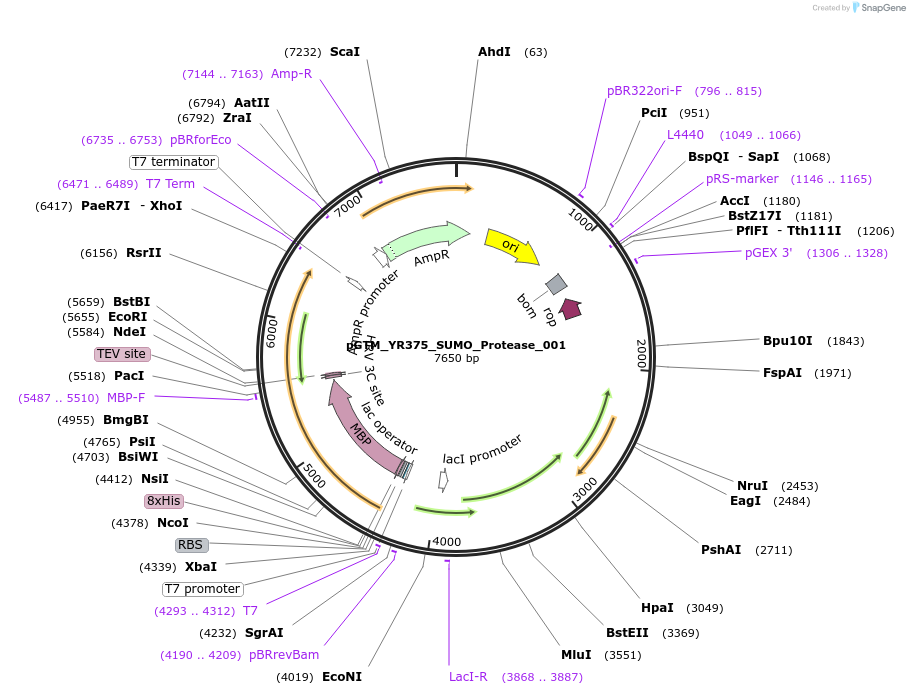
pGTM_YR375_SUMO_Protease_001
Plasmid#190063PurposeExpresses the yeast Ulp1 Sumo protease in active form as a TEV protease-cleavable MBP fusionDepositorInsertULP1_YEAST (ULP1 Budding Yeast)
Tags8X His-MBP (Maltose Binding Protein)ExpressionBacterialMutationConstruct contains residues 347 – 621PromoterT7 lacAvailable SinceJan. 30, 2025AvailabilityIndustry, Academic Institutions, and Nonprofits