We narrowed to 83,772 results for: MYC
-
Plasmid#57397PurposeLocalization: Lysosome Membrane, Excitation: 505 / 569, Emission: 516 / 581DepositorInsertLAMP1
TagsmEos2ExpressionMammalianMutationD50EPromoterCMVAvailable SinceFeb. 5, 2015AvailabilityIndustry, Academic Institutions, and Nonprofits -
pJRT.Agro.CLCrVA.010
Plasmid#31810DepositorInsertJRTCLCrVA.010
UseAgrobacterium binaryAvailable SinceNov. 13, 2012AvailabilityAcademic Institutions and Nonprofits only -
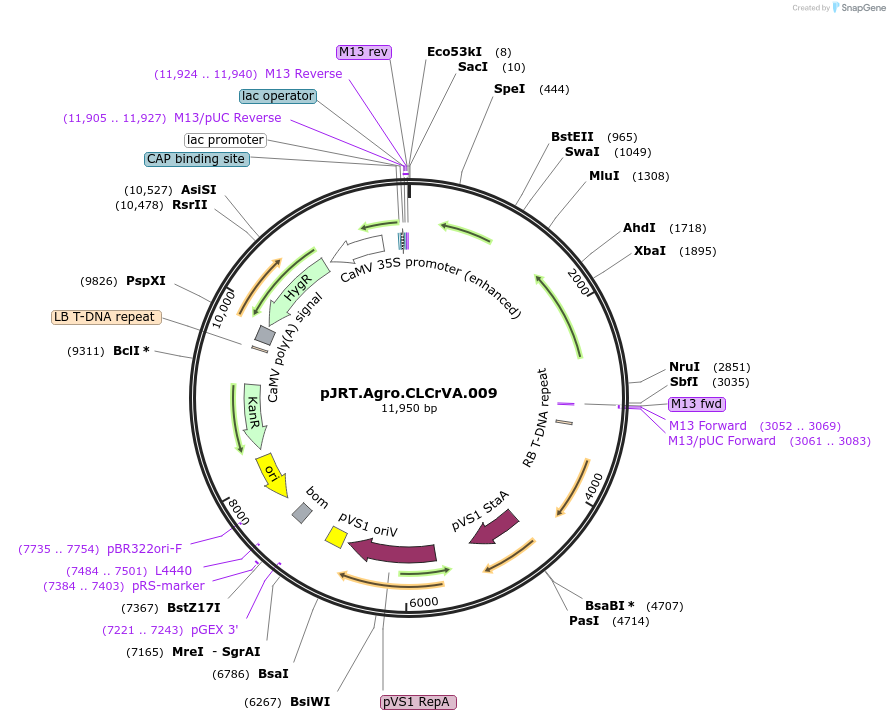
pJRT.Agro.CLCrVA.009
Plasmid#37972DepositorInsertpJRTCLCrVA.009
UseAgrobacterium binaryAvailable SinceOct. 15, 2012AvailabilityAcademic Institutions and Nonprofits only -
mEos4b-Actin-C-18
Plasmid#57500PurposeLocalization: Actin, Excitation: 505 / 569, Emission: 516 / 581DepositorAvailable SinceMarch 4, 2015AvailabilityIndustry, Academic Institutions, and Nonprofits -
Reduced Expression Alpha-5-Integrin-GFP
Plasmid#30450DepositorInsertalpha 5 integrin (ITGA5 Human)
TagsEGFPExpressionMammalianMutation91-544 of enhancer region of CMV promoter removed…Available SinceAug. 29, 2011AvailabilityAcademic Institutions and Nonprofits only -
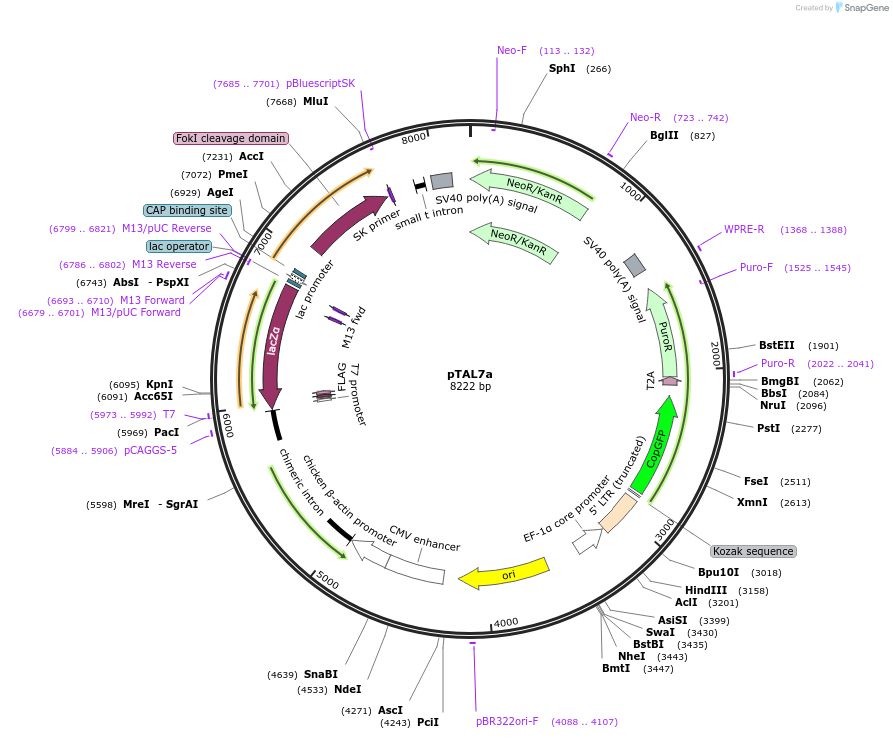
pTAL7a
Plasmid#48705PurposeTALEN expression in mammalian cellsDepositorTypeEmpty backboneUseTALENTagsFokI nucleaseExpressionMammalianPromoterCAGAvailable SinceNov. 12, 2013AvailabilityAcademic Institutions and Nonprofits only -
Dronpa-Alpha-V-Integrin-25
Plasmid#57261PurposeLocalization: Focal Adhesions, Excitation: 503, Emission: 518DepositorAvailable SinceJan. 13, 2015AvailabilityIndustry, Academic Institutions, and Nonprofits -
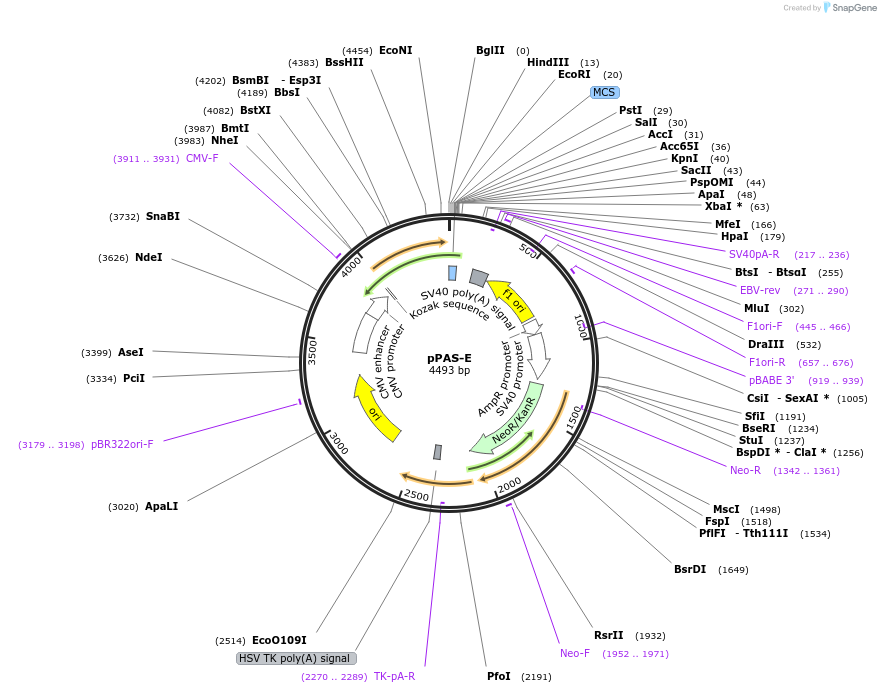
pPAS-E
Plasmid#39867DepositorInsertiRFP protein PAS domain
TagsE coilExpressionMammalianPromoterCMVAvailable SinceJuly 26, 2013AvailabilityAcademic Institutions and Nonprofits only -
mPA-GFP-ER-5
Plasmid#57132PurposeLocalization: Endoplasmic Reticulum, Excitation: 400 / 504, Emission: 515 / 517DepositorAvailable SinceFeb. 5, 2015AvailabilityIndustry, Academic Institutions, and Nonprofits -
mPA-GFP-ER-3
Plasmid#57131PurposeLocalization: Endoplasmic Reticulum, Excitation: 400 / 504, Emission: 515 / 517DepositorAvailable SinceFeb. 5, 2015AvailabilityIndustry, Academic Institutions, and Nonprofits -
Dendra2-VASP-5
Plasmid#57744PurposeLocalization: Actin/Focal Adhesions, Excitation: 490 / 553, Emission: 507 / 573DepositorInsertVASP
TagsDendra2ExpressionMammalianMutationA209T in VASPPromoterCMVAvailable SinceJan. 6, 2015AvailabilityIndustry, Academic Institutions, and Nonprofits -
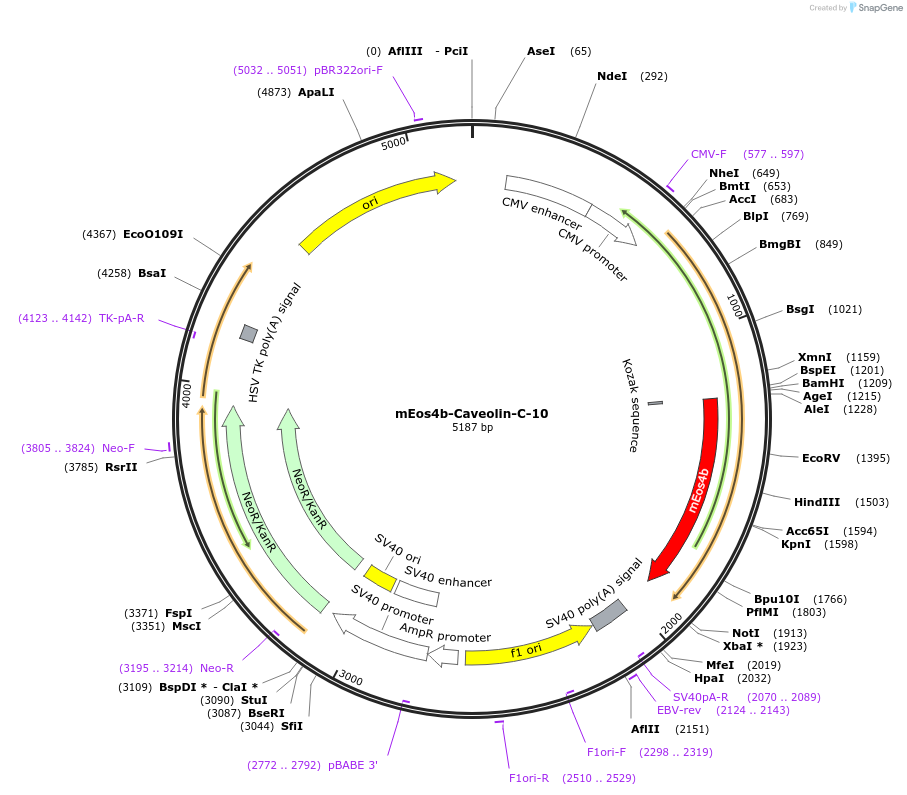
mEos4b-Caveolin-C-10
Plasmid#57503PurposeLocalization: Membrane, Excitation: 505 / 569, Emission: 516 / 581DepositorAvailable SinceMarch 4, 2015AvailabilityIndustry, Academic Institutions, and Nonprofits -
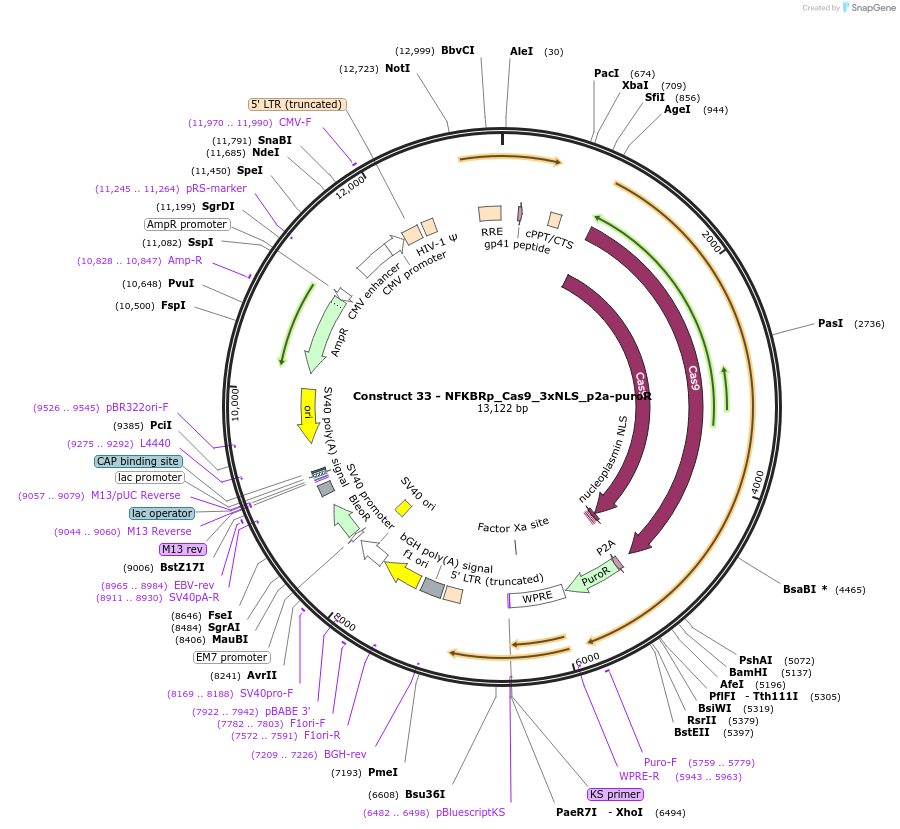
Construct 33 - NFKBRp_Cas9_3xNLS_p2a-puroR
Plasmid#81255PurposeLentiviral construct for building stable cell lines expressing Cas9 in the presence of TNFa via puromycin selectionDepositorInsertCas9-3xNLS
UseLentiviral and Synthetic BiologyExpressionMammalianAvailable SinceSept. 14, 2016AvailabilityAcademic Institutions and Nonprofits only -
PS-CFP2-Mito-7
Plasmid#57240PurposeLocalization: Cytoskeleton, Excitation: 400 / 490, Emission: 468 / 511DepositorAvailable SinceApril 1, 2015AvailabilityIndustry, Academic Institutions, and Nonprofits -
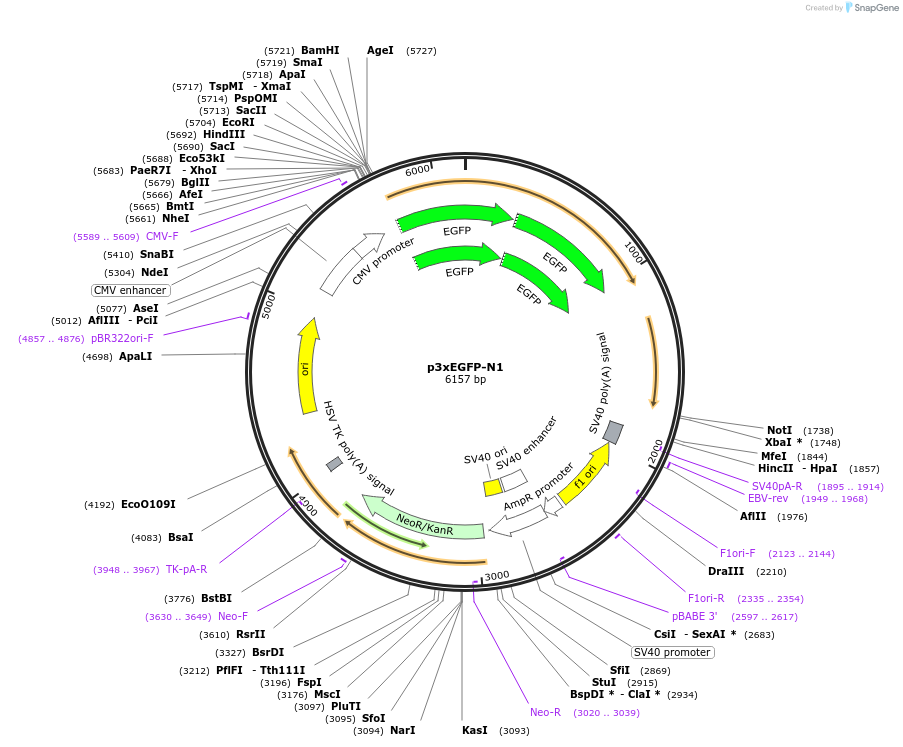
p3xEGFP-N1
Plasmid#122163PurposeExpression of 3xEGFPDepositorInsert3xEGFP
ExpressionMammalianPromoterCMVAvailable SinceFeb. 21, 2019AvailabilityAcademic Institutions and Nonprofits only -
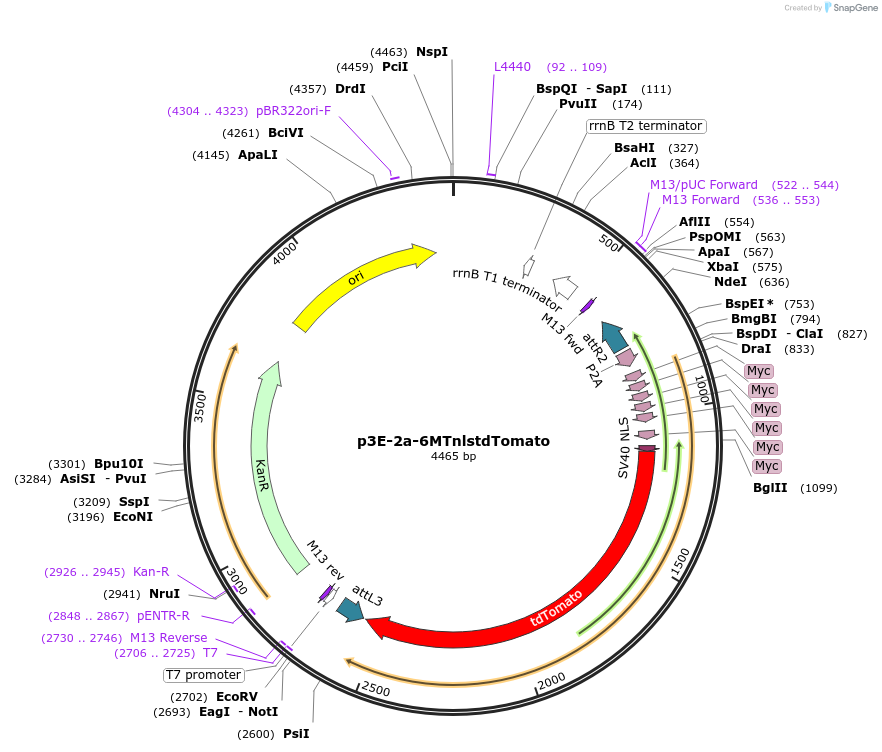
p3E-2a-6MTnlstdTomato
Plasmid#67699Purpose3' entry clone with 6xmyc nuclear localized tdTomatoDepositorInserttdTomato
Tags2A-6xMyc-NLSAvailable SinceAug. 6, 2015AvailabilityAcademic Institutions and Nonprofits only -
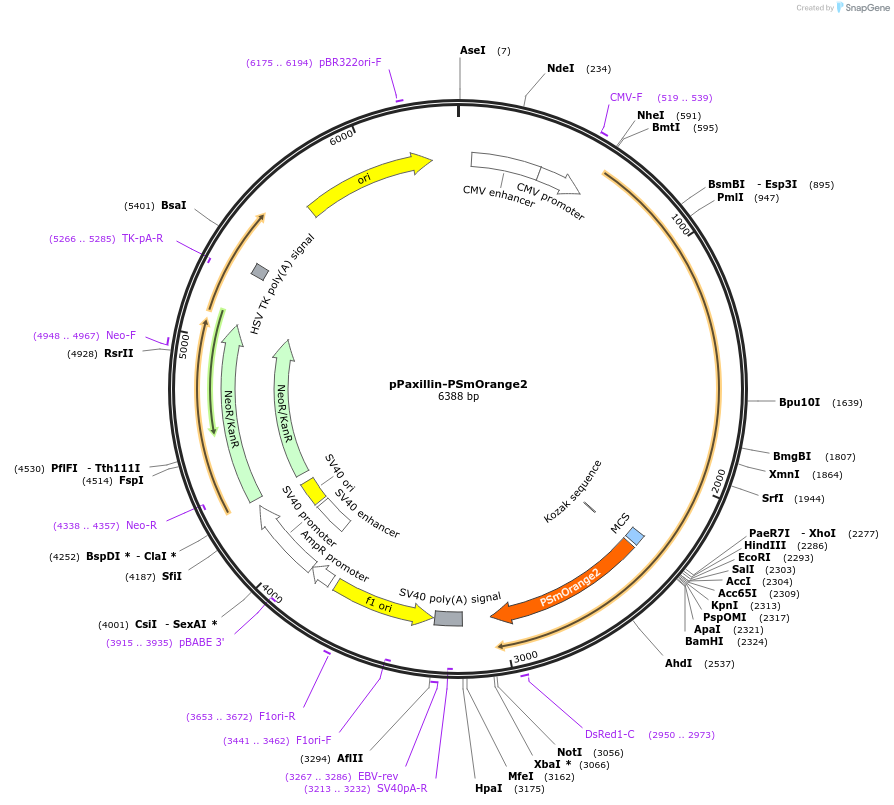
pPaxillin-PSmOrange2
Plasmid#34964DepositorInsertPSmOrange2
TagsPaxillinExpressionMammalianMutationR17H, R36H, F64I, Q194L, A224S, G226A (mutations …Available SinceAug. 20, 2012AvailabilityAcademic Institutions and Nonprofits only -
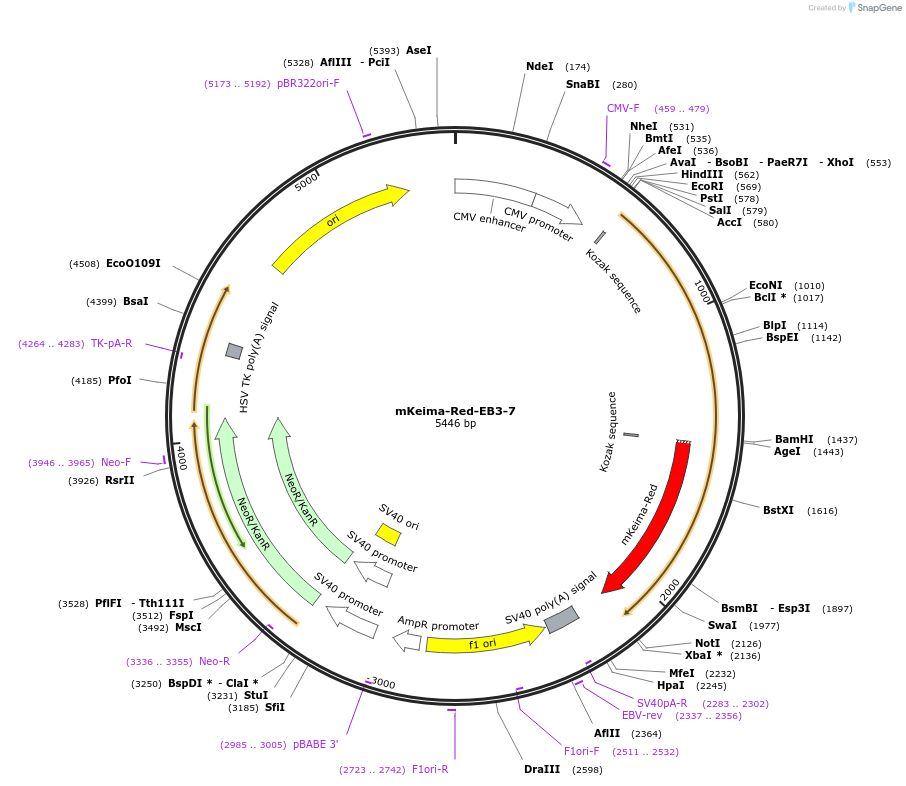
mKeima-Red-EB3-7
Plasmid#56015PurposeLocalization: MT End Binding Protein, Excitation: 440, Emission: 620DepositorAvailable SinceApril 1, 2015AvailabilityAcademic Institutions and Nonprofits only -
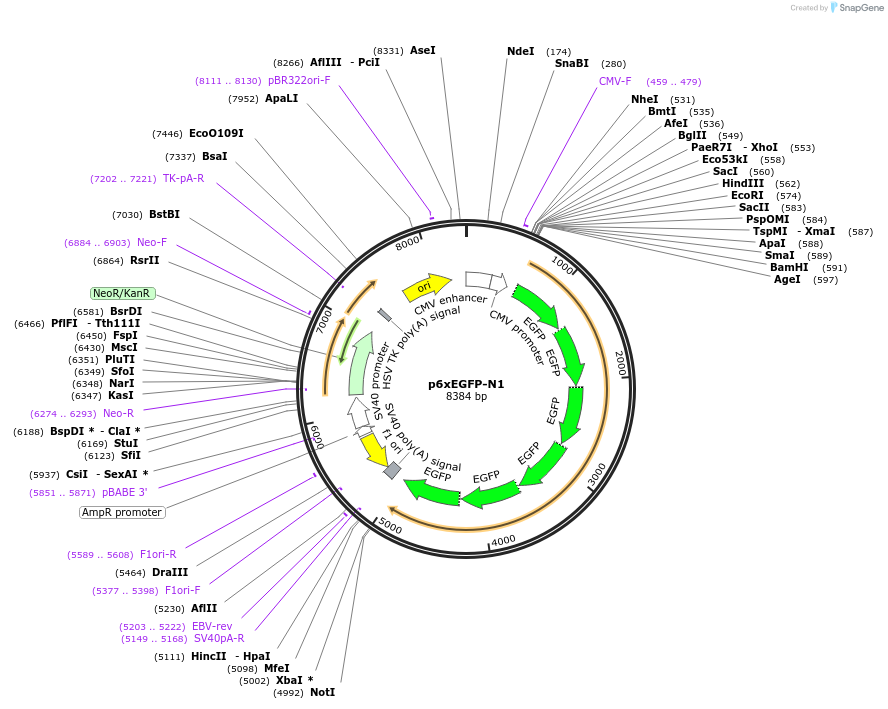
p6xEGFP-N1
Plasmid#122166PurposeExpression of 6xEGFPDepositorInsert6xEGFP
ExpressionMammalianPromoterCMVAvailable SinceFeb. 22, 2019AvailabilityAcademic Institutions and Nonprofits only -
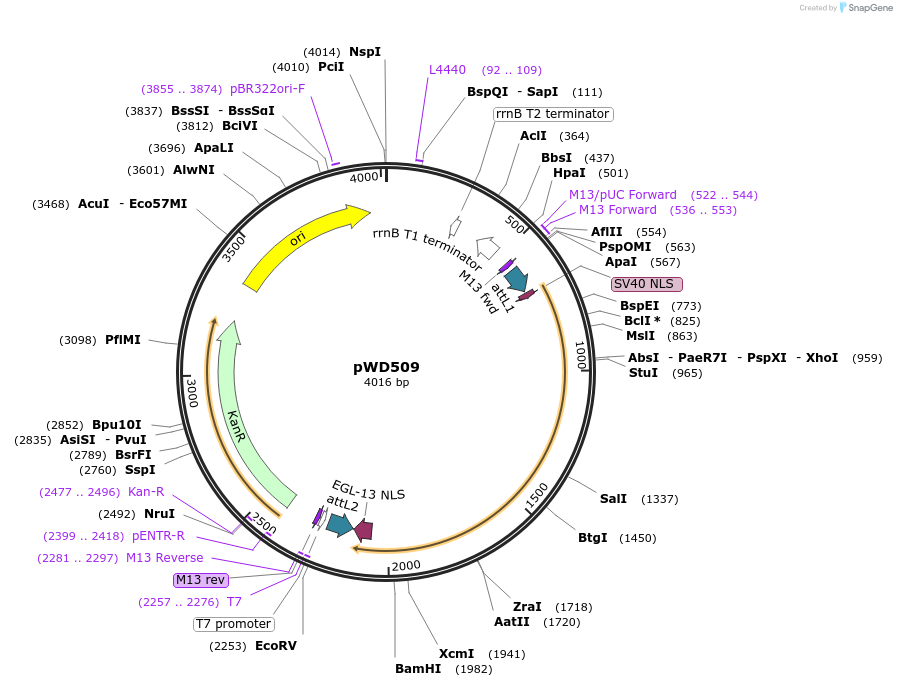
pWD509
Plasmid#73739PurposeGateway compatible [1-2] Entry Vector encoding 2xNLS-FLP-D5DepositorInsert2xNLS-FLP-D5
ExpressionWormMutationG5DAvailable SinceMarch 25, 2016AvailabilityAcademic Institutions and Nonprofits only