We narrowed to 1,801 results for: nif
-
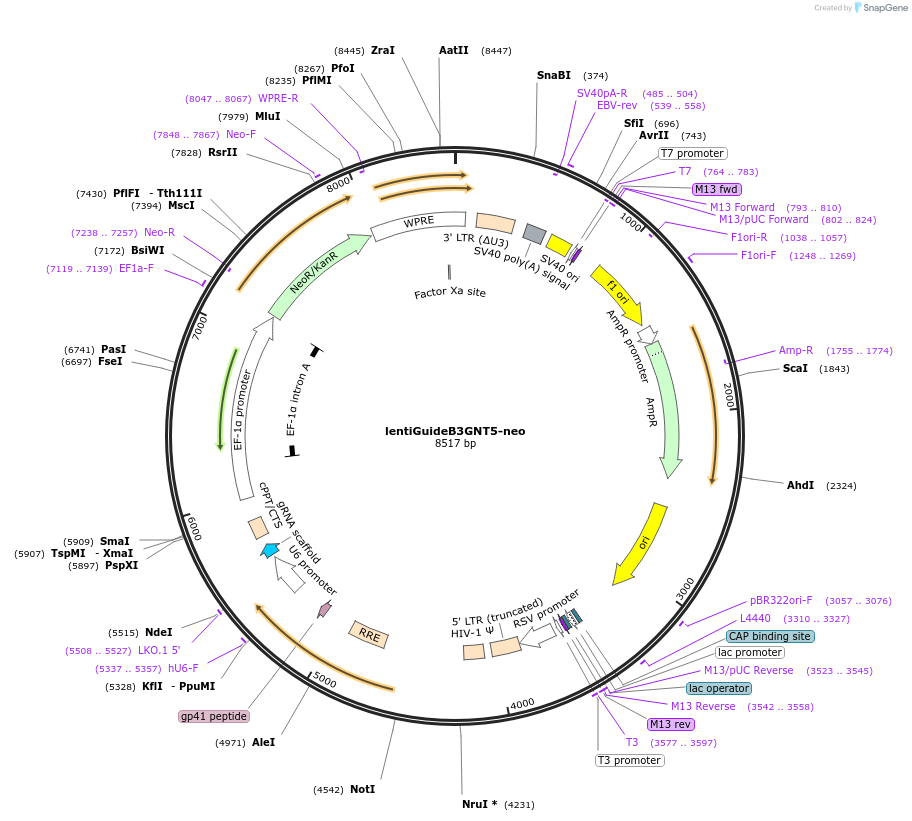
Plasmid#220867Purposeencodes CRISPR sgRNA targeting human B3GNT5DepositorInsertB3GNT5 (B3GNT5 Human)
UseLentiviralAvailable SinceSept. 3, 2024AvailabilityAcademic Institutions and Nonprofits only -
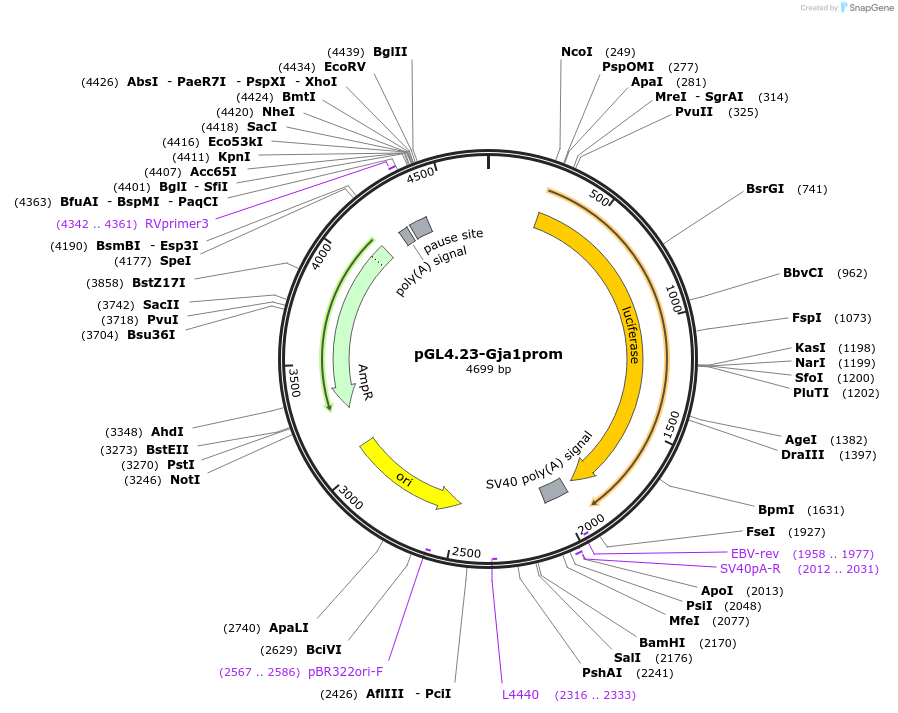
pGL4.23-Gja1prom
Plasmid#188114PurposeExpresses luciferase under control of Gja1 promoter in transfected cellsDepositorInsertGja1 promoter (Gja1 Mouse)
UseLuciferasePromoterGja1 endogenous promoter up to gene start codonAvailable SinceSept. 28, 2023AvailabilityAcademic Institutions and Nonprofits only -
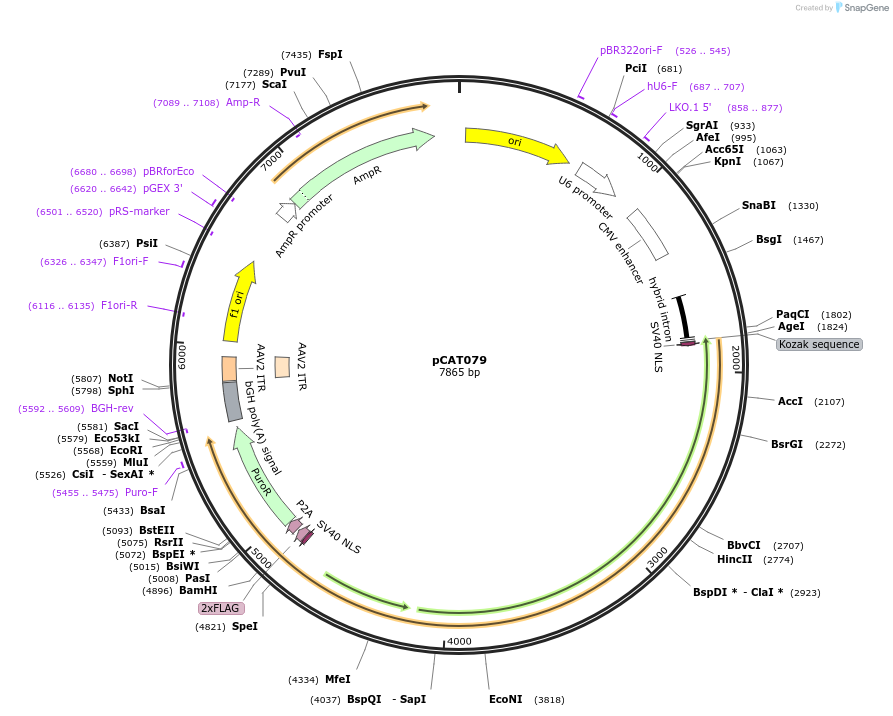
pCAT079
Plasmid#180509PurposePlasmid expressing mammalian codon optimized wt DpbCasX, sgRNAv2 scaffold, and restriction sites to clone in new spacersDepositorInsertsCasX sgRNAv2
DpbCasX
UseCRISPRTags2x FLAG and SV40 NLSExpressionMammalianPromoterCAG and U6Available SinceJune 10, 2022AvailabilityAcademic Institutions and Nonprofits only -
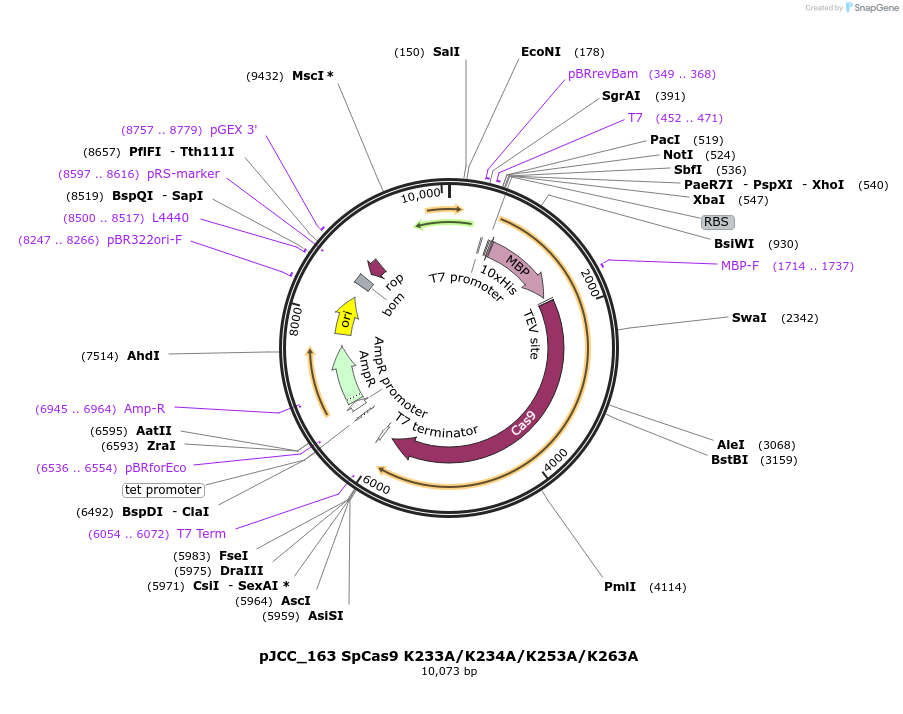
pJCC_163 SpCas9 K233A/K234A/K253A/K263A
Plasmid#179527PurposeFor bacterial expression of SpCas9 K233A/K234A/K253A/K263A (helix-rolling basic patch mutant) with an N-terminal His-MBP tagDepositorInsertSpCas9 K233A/K234A/K253A/K263A
Tags10X His, TEV protease cleavage site, and maltose-…ExpressionBacterialMutationchanged lysines 233, 234, 253, and 263 to alanineAvailable SinceApril 19, 2022AvailabilityAcademic Institutions and Nonprofits only -
pJSC119 - Bacterial expression plasmid for SpCas9 + N497A/R661A/Q695A variant
Plasmid#101207PurposeBacterial expression plasmid for SpCas9 + N497A/R661A/Q695A variantDepositorInsertSpCas9 variant N497A/R661A/Q695A
Tags10x His, MBP, and TEV siteExpressionBacterialMutationN497A, R661A and Q695APromoterT7Available SinceNov. 7, 2017AvailabilityAcademic Institutions and Nonprofits only -
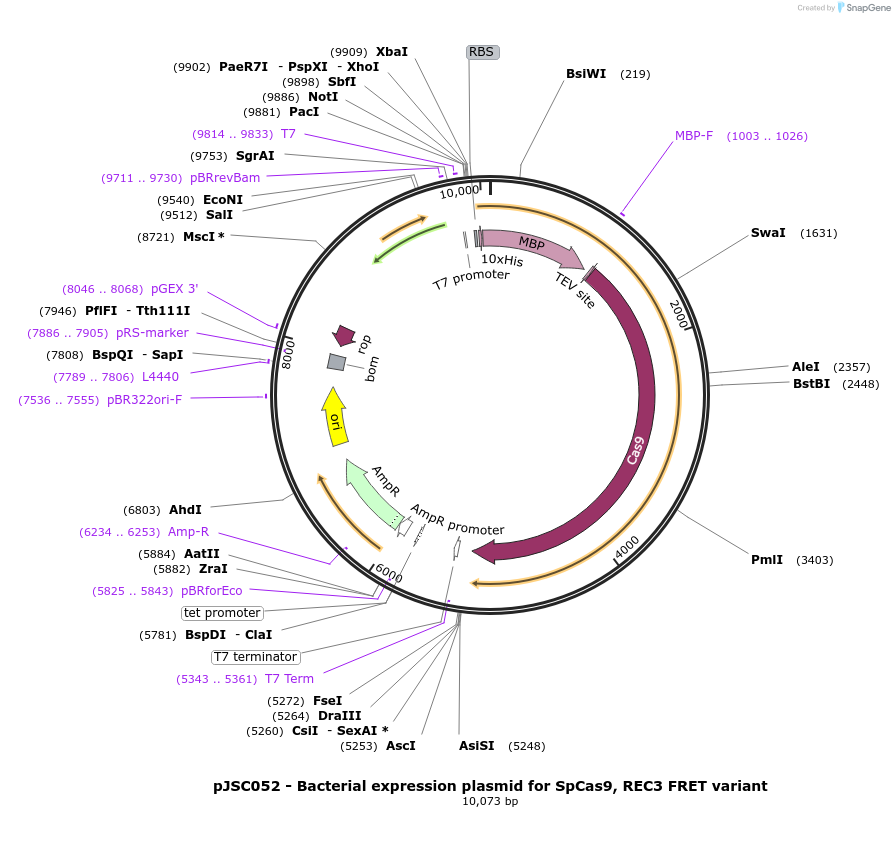
pJSC052 - Bacterial expression plasmid for SpCas9, REC3 FRET variant
Plasmid#101201PurposeBacterial expression plasmid for SpCas9, REC3 FRET variantDepositorInsertSpCas9 variant C80S/C574S/S701C/S960C
Tags10x His, MBP, and TEV siteExpressionBacterialMutationC80S, C574S, S701C and S960CPromoterT7Available SinceOct. 30, 2017AvailabilityAcademic Institutions and Nonprofits only -
pJSC077 - Bacterial expression plasmid for SpCas9 + K855A, HNH FRET variant
Plasmid#101214PurposeBacterial expression plasmid for SpCas9 + K855A, HNH FRET variantDepositorInsertSpCas9 variant C80S/C574S/S355C/S867C/K855A
Tags10x His, MBP, and TEV siteExpressionBacterialMutationC80S, C574S, S355C, S867C and K855APromoterT7Available SinceOct. 25, 2017AvailabilityAcademic Institutions and Nonprofits only -
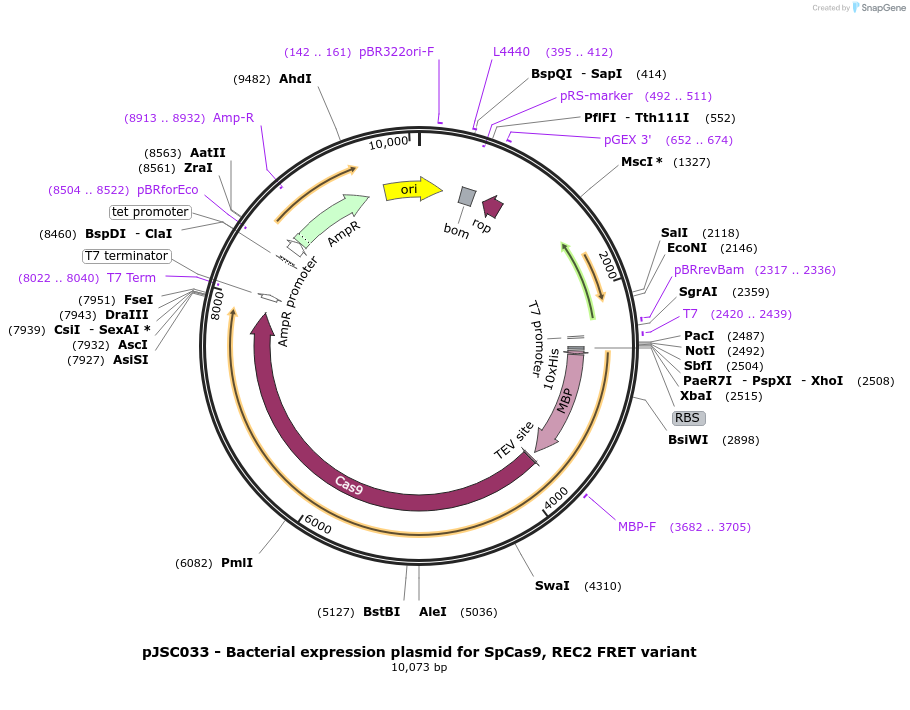
pJSC033 - Bacterial expression plasmid for SpCas9, REC2 FRET variant
Plasmid#101200PurposeBacterial expression plasmid for SpCas9, REC2 FRET variantDepositorInsertSpCas9 variant C80S/C574S/E60C/D273C
Tags10x His, MBP, and TEV siteExpressionBacterialMutationC80S, C574S, E60C and D273CPromoterT7Available SinceOct. 4, 2017AvailabilityAcademic Institutions and Nonprofits only -
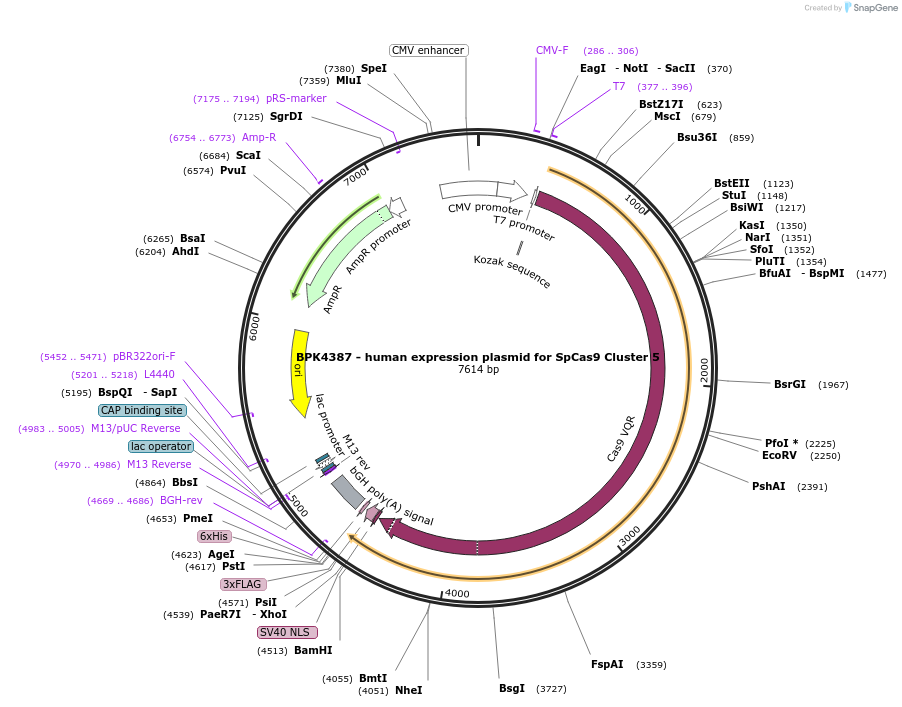
BPK4387 - human expression plasmid for SpCas9 Cluster 5
Plasmid#101191PurposeHuman expression plasmid for SpCas9 Cluster 5 variant: CMV-T7-hSpCas9-Cluster5(K918A, V922A, R925A)-NLS(SV40)-3xFLAGDepositorInserthSpCas9-Cluster5(K918A/V922A/R925A)
Tags3x FLAG and NLS (SV40)ExpressionMammalianMutationK918A/V922A/R925APromoterCMVAvailable SinceSept. 29, 2017AvailabilityAcademic Institutions and Nonprofits only -
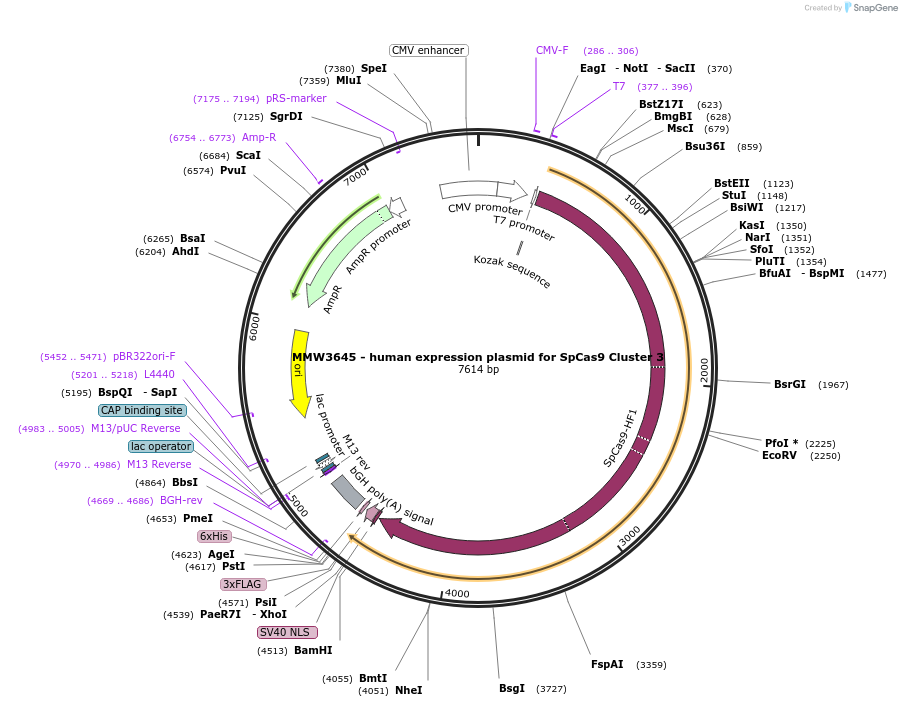
MMW3645 - human expression plasmid for SpCas9 Cluster 3
Plasmid#101187PurposeHuman expression plasmid for SpCas9 Cluster 3 variant: CMV-T7-hSpCas9-Cluster3(T657A, G658A, W659A, R661A)-NLS(SV40)-3xFLAGDepositorInserthSpCas9-Cluster3(T657A/G658A/W659A/R661A)
Tags3x FLAG and NLS (SV40)ExpressionMammalianMutationT657A/G658A/W659A/R661APromoterCMVAvailable SinceSept. 29, 2017AvailabilityAcademic Institutions and Nonprofits only -
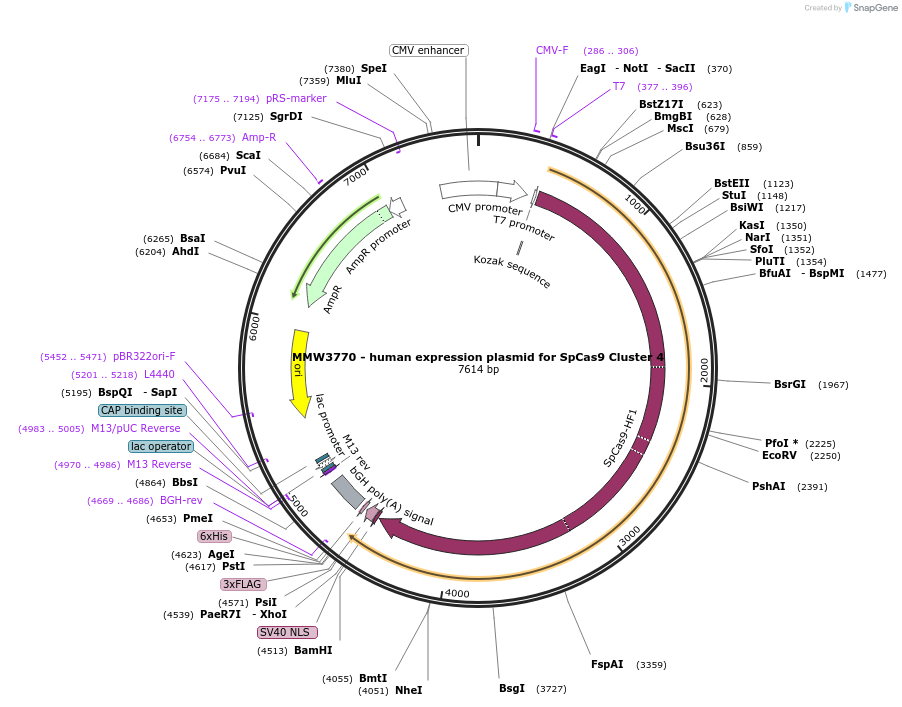
MMW3770 - human expression plasmid for SpCas9 Cluster 4
Plasmid#101189PurposeHuman expression plasmid for SpCas9 Cluster 4 variant: CMV-T7-hSpCas9-Cluster4(F491A, M495A, T496A, N497A)-NLS(SV40)-3xFLAGDepositorInserthSpCas9-Cluster4(F491A/M495A/T496A/N497A)
Tags3x FLAG and NLS (SV40)ExpressionMammalianMutationF491A/M495A/T496A/N497APromoterCMVAvailable SinceSept. 29, 2017AvailabilityAcademic Institutions and Nonprofits only -
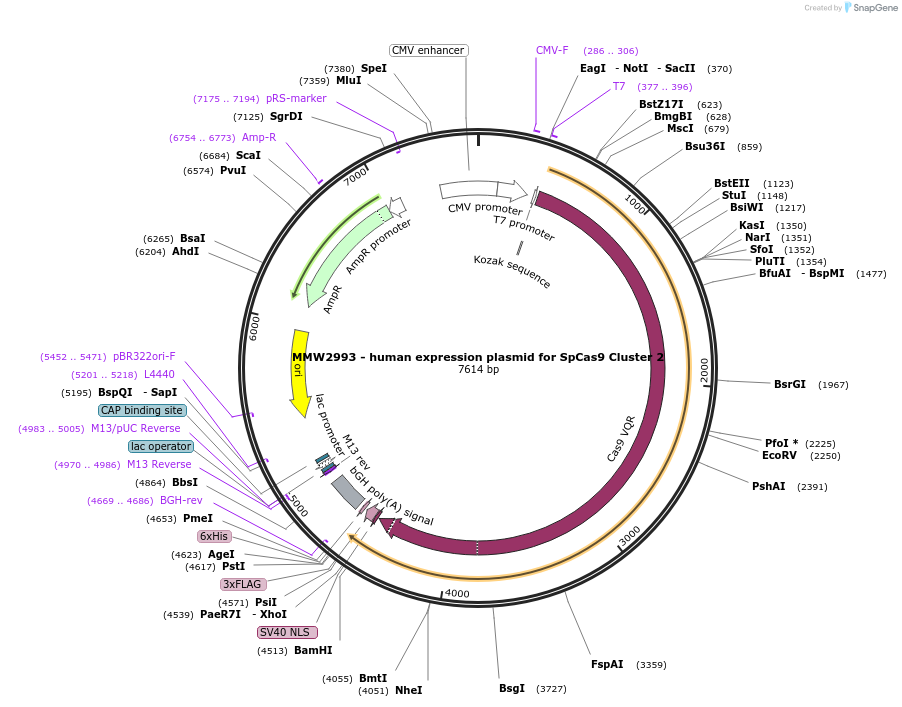
MMW2993 - human expression plasmid for SpCas9 Cluster 2
Plasmid#101185PurposeHuman expression plasmid for SpCas9 Cluster 2 variant: CMV-T7-hSpCas9-Cluster2(G582A, V583A, E584A, D585A, N588A)-NLS(SV40)-3xFLAGDepositorInserthSpCas9-Cluster2(G582A/V583A/E584A/D585A/N588A)
Tags3x FLAG and NLS (SV40)ExpressionMammalianMutationG582A/V583A/E584A/D585A/N588APromoterCMVAvailable SinceSept. 29, 2017AvailabilityAcademic Institutions and Nonprofits only -
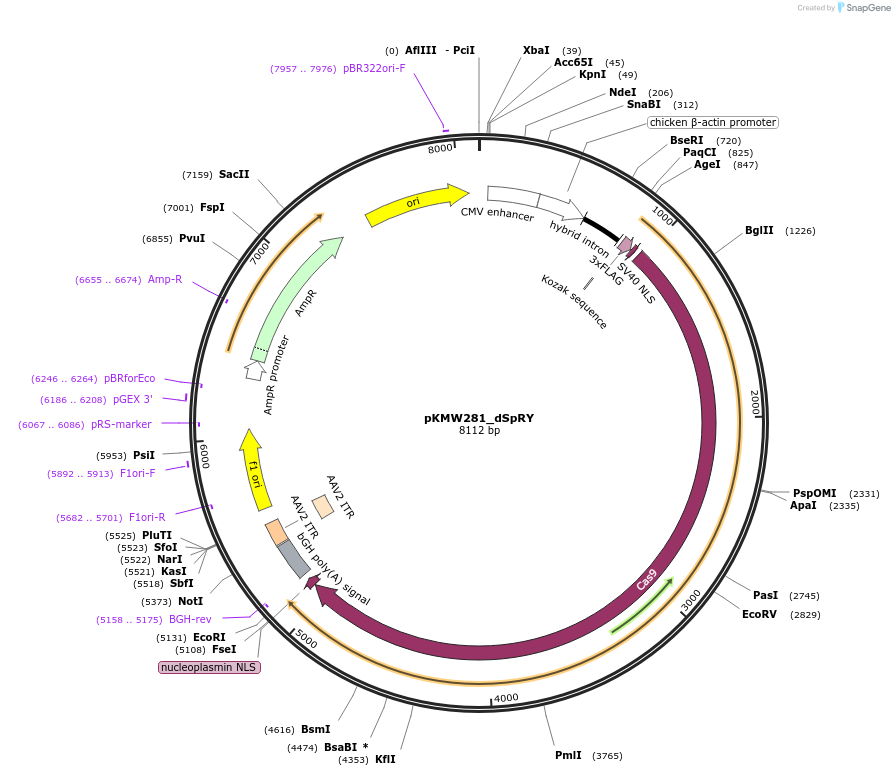
pKMW281_dSpRY
Plasmid#244822PurposeMammalian expression of catalytically inactive SpRY Cas9 (dSpRY)DepositorInsertdSpRY Cas9
UseCRISPRTags3xFLAG-SV40 NLS and Nucleoplasmin NLSExpressionMammalianMutationD10A/A61R/H840A/L1111R/D1135L/S1136W/G1218K/E1219…PromoterCAGAvailable SinceSept. 17, 2025AvailabilityAcademic Institutions and Nonprofits only -
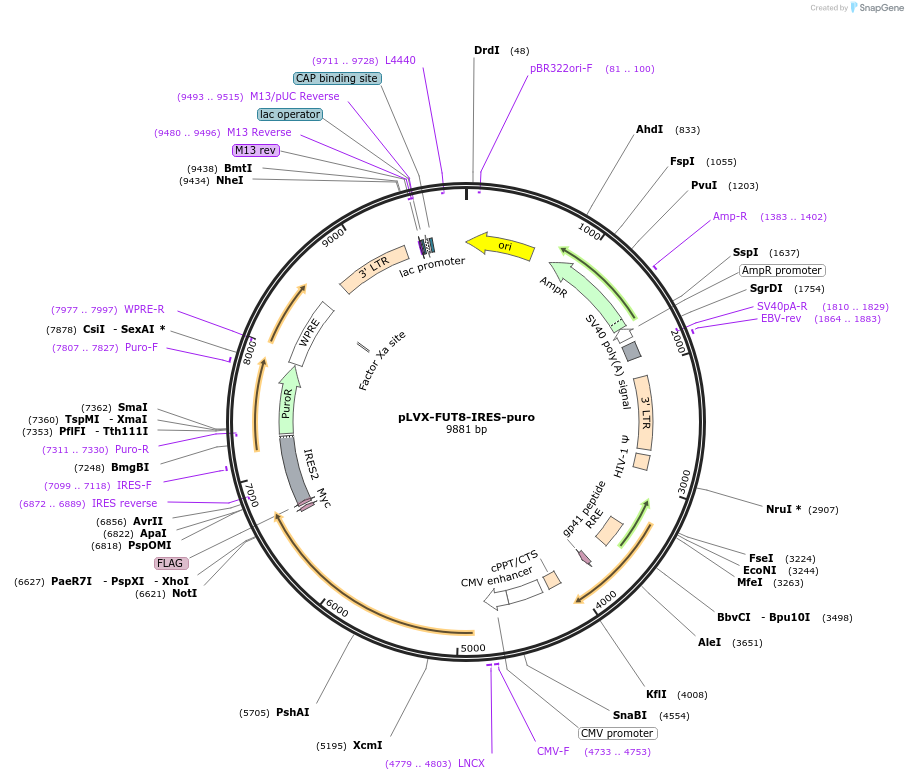
pLVX-FUT8-IRES-puro
Plasmid#250319PurposeExpression of human FUT8 with C-terminal myc and FLAG tags in mammalian cellsDepositorInsertFUT8 (FUT8 Human)
UseLentiviralTagsmyc and FLAGExpressionMammalianMutationsynonymous mutations to DNA encoding Asp306PromoterCMV promoterAvailable SinceMarch 5, 2026AvailabilityAcademic Institutions and Nonprofits only -
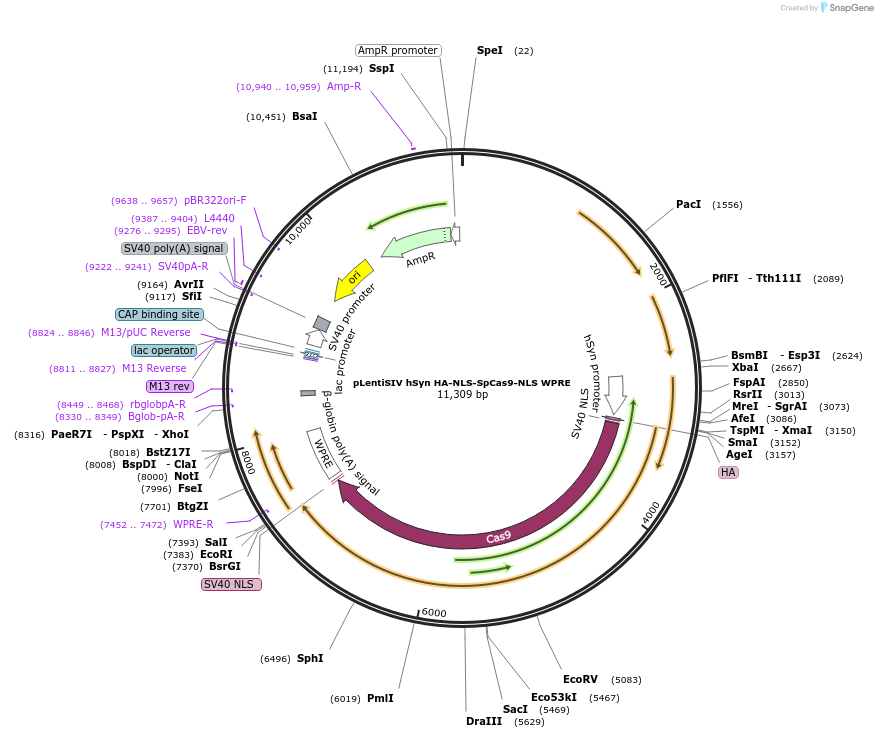
pLentiSIV hSyn HA-NLS-SpCas9-NLS WPRE
Plasmid#236242PurposeLentiviral expression of SpCas9DepositorInsertSpCas9
UseLentiviralPromoterhuman Synapsin 1Available SinceMay 1, 2025AvailabilityAcademic Institutions and Nonprofits only -
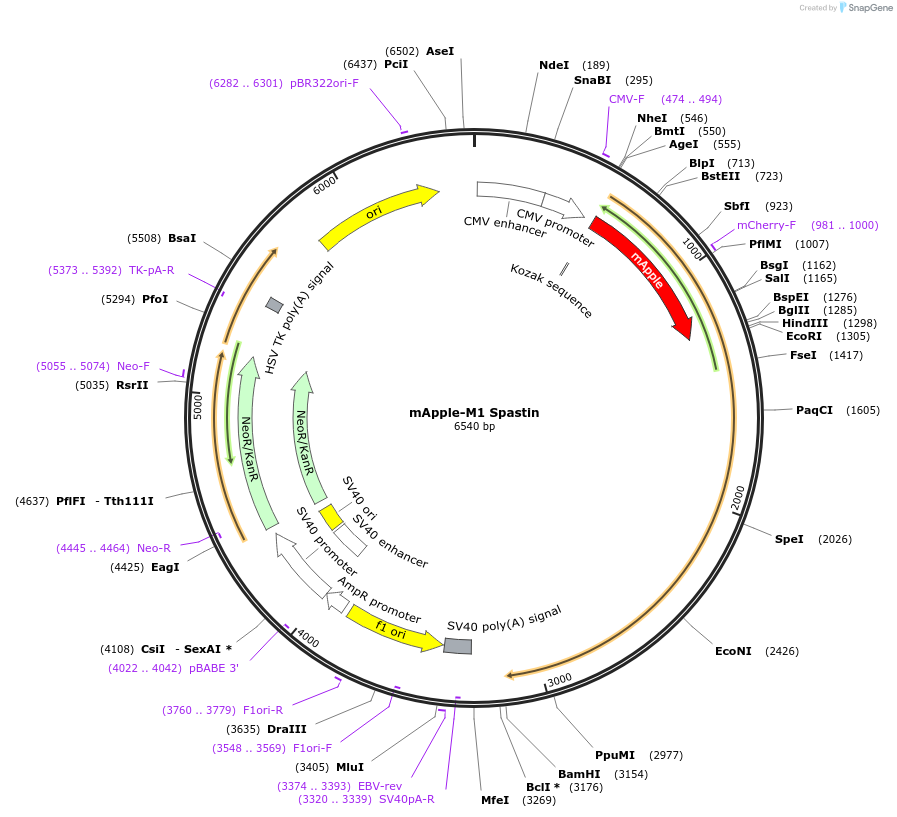
mApple-M1 Spastin
Plasmid#134461PurposeM1 Spastin expressionDepositorInsertM1 Spastin (SPAST Human)
ExpressionMammalianMutationMet to Ala change at amino acid 87PromoterCMVAvailable SinceNov. 25, 2019AvailabilityAcademic Institutions and Nonprofits only -
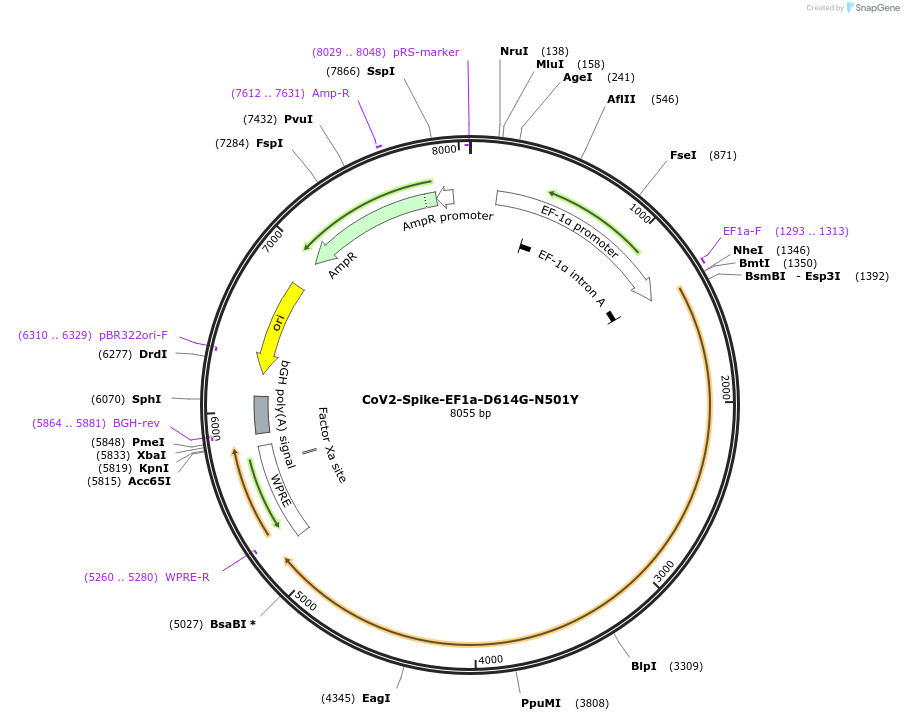
CoV2-Spike-EF1a-D614G-N501Y
Plasmid#177939PurposeTransient mammalian expression of codon optimized SARS-CoV-2 Spike proteins (D614G, N501Y). This version gives lower expression of Spike and is more convenient for generating VLPs.DepositorAvailable SinceNov. 16, 2021AvailabilityAcademic Institutions and Nonprofits only -
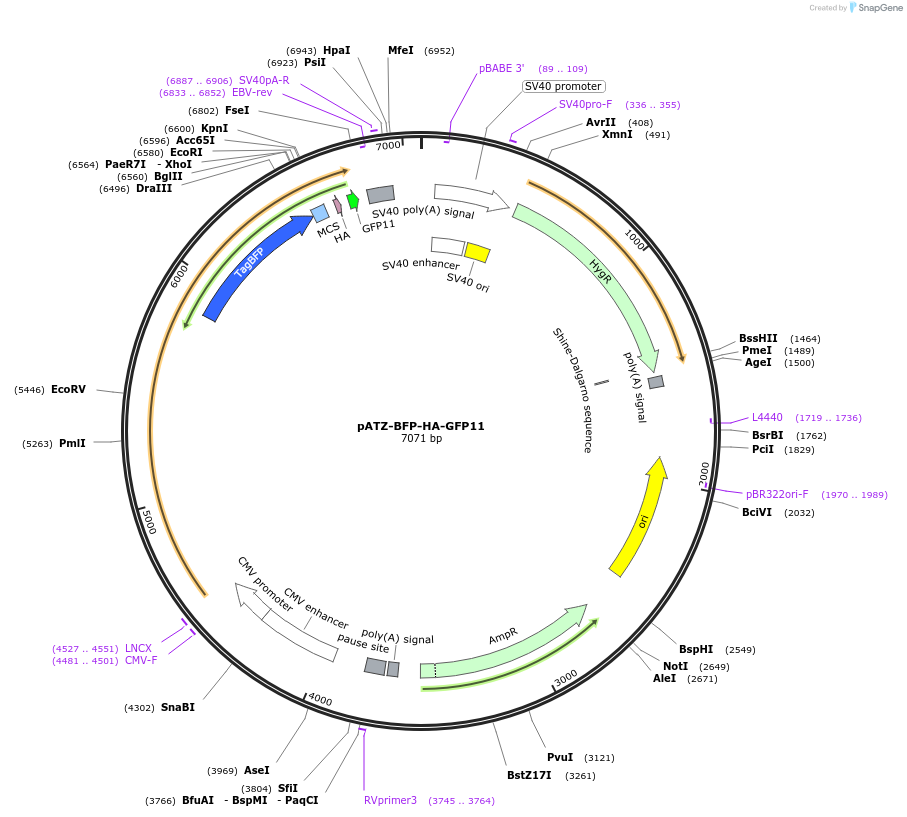
pATZ-BFP-HA-GFP11
Plasmid#177721PurposeER protein alpha-1-antitrypsin containing E342K mutation (ATZ mutant) fused to blue fluorescence protein (BFP) tag protein, an HA tag and the eleventh β-strand of GFPDepositorInsertalpha-1-antitrypsin (SERPINA1 Human)
TagsGFP11, HA, and TagBFPExpressionBacterial and MammalianMutationE342K (ATZ mutant)Available SinceJan. 26, 2022AvailabilityAcademic Institutions and Nonprofits only -
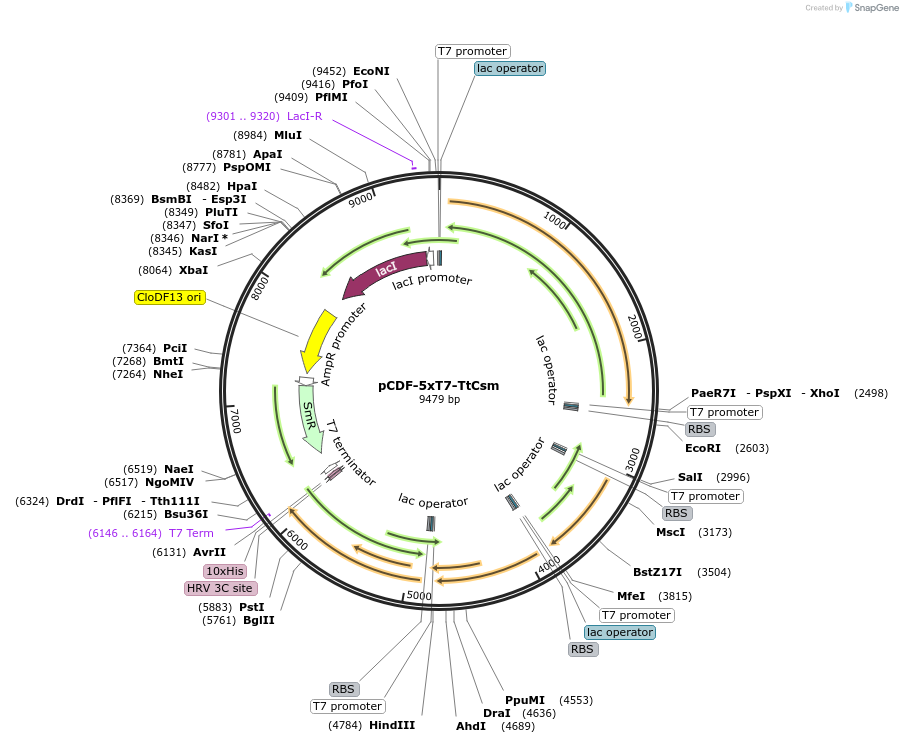
pCDF-5xT7-TtCsm
Plasmid#128572PurposeFor expression of T. thermophilus Csm in E. coli (Csm1-5)DepositorInsertsCsm1
Csm2, Csm3, Csm4, Csm5
TagsHRV 3C protease cleavage site and 10xHis tag on C…ExpressionBacterialMutationCodon optimized for expression in E. coli and Cod…PromoterT7 promoter, lac operatorAvailable SinceAug. 6, 2019AvailabilityAcademic Institutions and Nonprofits only -
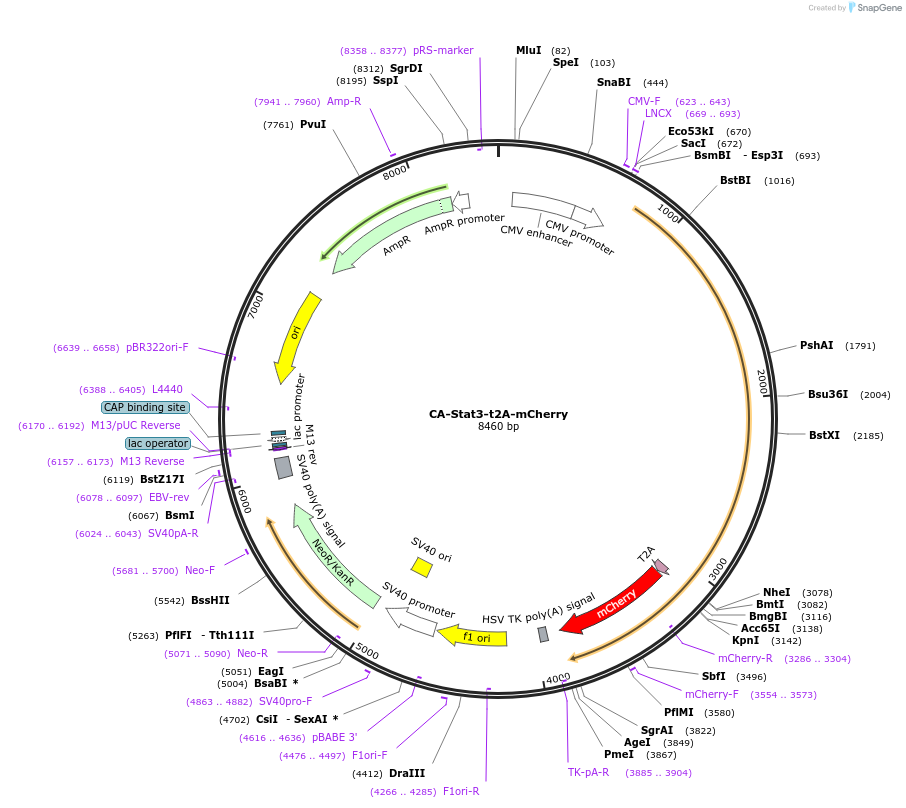
CA-Stat3-t2A-mCherry
Plasmid#127540PurposeExpresses constitutively active (CA) mouse Stat3 and mCherry via t2A linker.DepositorInsertStat3 (Stat3 Mouse)
ExpressionMammalianMutationA662C and N664C. Residues changed to Cys renders …PromoterCMVAvailable SinceJuly 25, 2019AvailabilityAcademic Institutions and Nonprofits only