We narrowed to 5,231 results for: codon optimized
-
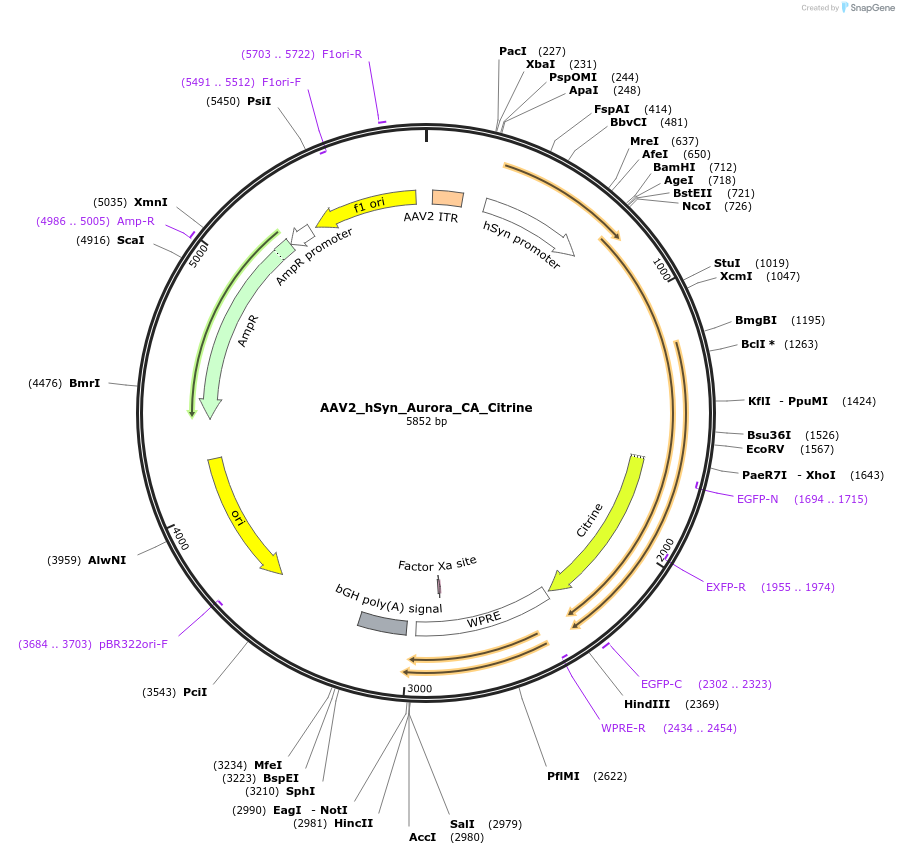
Plasmid#98219PurposeSlow cycling (open for minutes) step-function artificial anion conducting channelrhodopsin (aACR). High light sensitivity. Activation with green light, inactivation max with 625 nm (accelarates closure to ms). Not recommended for in vivo use. Codon optimized for mammalian expression.DepositorInsertSynthetic construct Aurora_C128A gene
UseAAVTagsCitrineExpressionMammalianMutationV59S, E83N, E90Q, E101S, V117R, E123S, C128A, P24…Promoterhuman synapsinAvailable SinceAug. 22, 2017AvailabilityAcademic Institutions and Nonprofits only -
pLJM60-FLAG-SLC38A9.1 Y71A
Plasmid#71865Purposelentiviral stable expressionDepositorInsertSLC38A9.1 (SLC38A9 Human)
UseLentiviralTagsFLAGExpressionMammalianMutationcodon optimized, Tyrosine 71 mutated to AlaninePromoterCMVAvailable SinceFeb. 2, 2016AvailabilityAcademic Institutions and Nonprofits only -
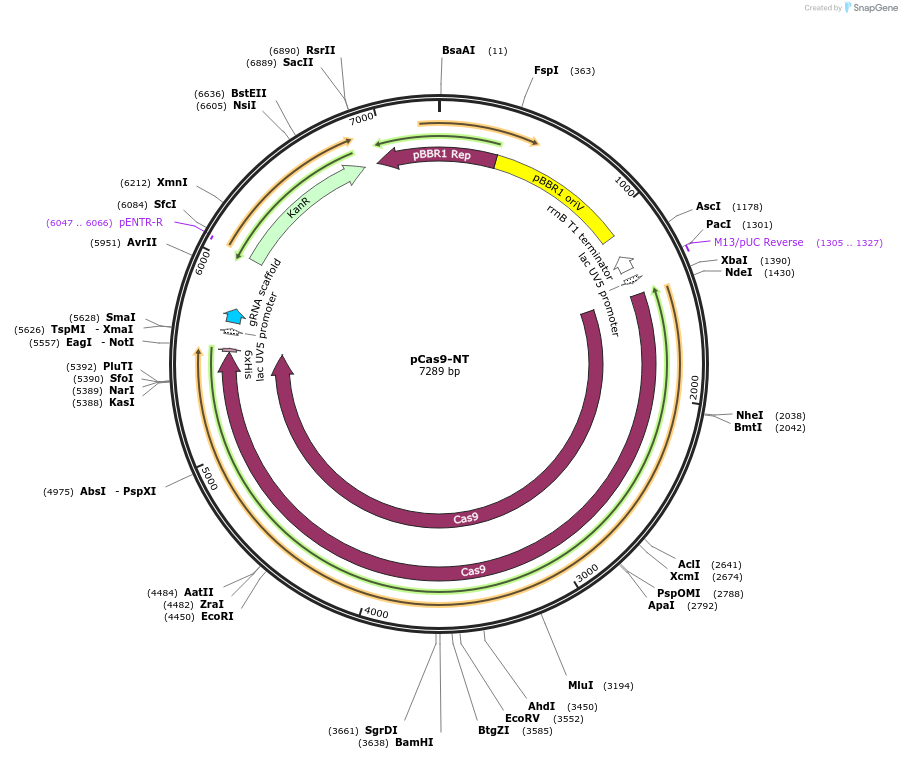
pCas9-NT
Plasmid#235165PurposeExpresses the codon harmonized Streptococcus pyogenes cas9 gene and a NT sgRNADepositorInsertSpCas9
ExpressionBacterialAvailable SinceAug. 22, 2025AvailabilityAcademic Institutions and Nonprofits only -
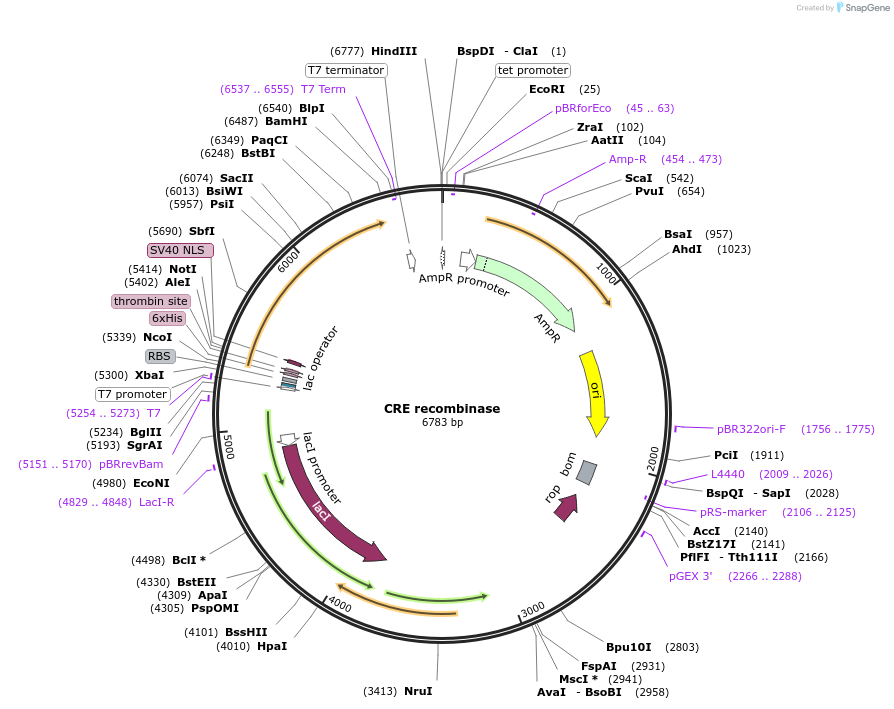
CRE recombinase
Plasmid#62730PurposeExpresses Cre recombinase protein in bacterial cellsDepositorInsertCre recombinase
UseCre/LoxTagsHis tag and NLSExpressionBacterialMutationCodon optimazed for e.coliPromoterT7Available SinceApril 24, 2015AvailabilityAcademic Institutions and Nonprofits only -
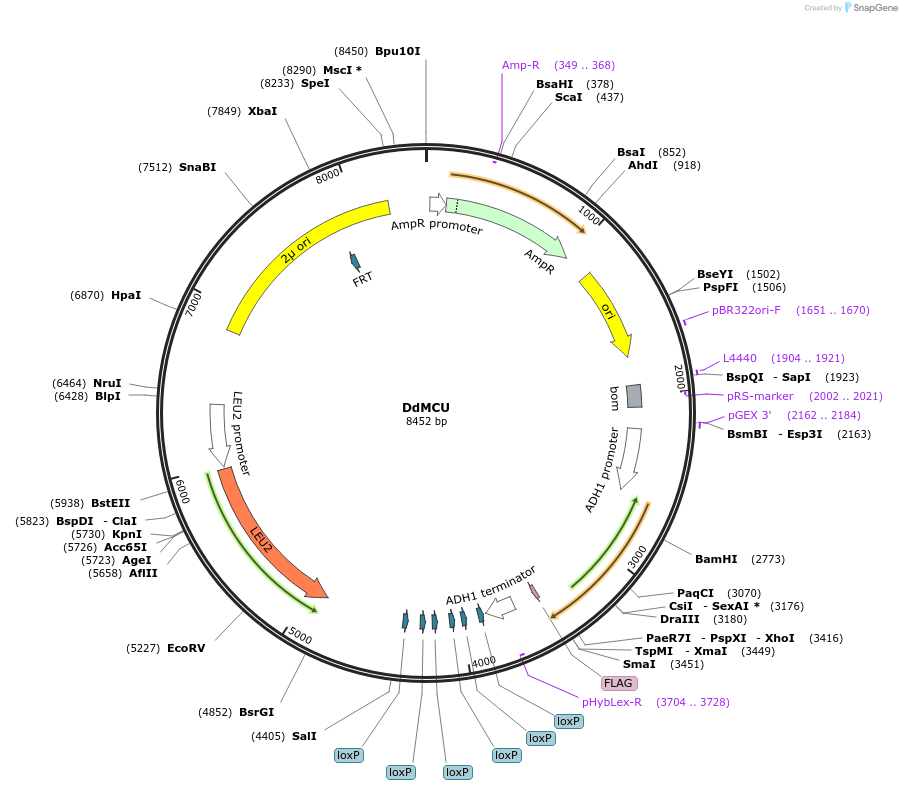
DdMCU
Plasmid#67642PurposeDdMCU-Flag in pACT2 vector human codon optimisedDepositorInsertDdMCU
TagsFlag tag with linkerExpressionYeastPromoterADH1Available SinceSept. 3, 2015AvailabilityAcademic Institutions and Nonprofits only -
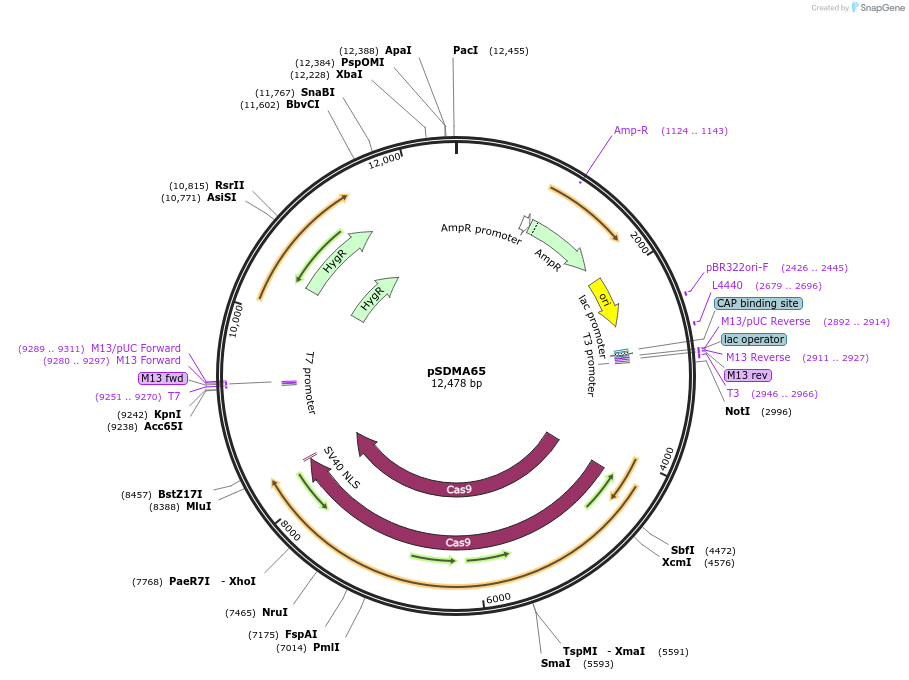
pSDMA65
Plasmid#128354PurposeCodon optimised Streptococcus pyogenes CAS9 ORF regulated by the Cryptococcus neoformans TEF1 promoter and terminator.DepositorTypeEmpty backboneUseCRISPRPromoterACT1Available SinceAug. 19, 2019AvailabilityAcademic Institutions and Nonprofits only -
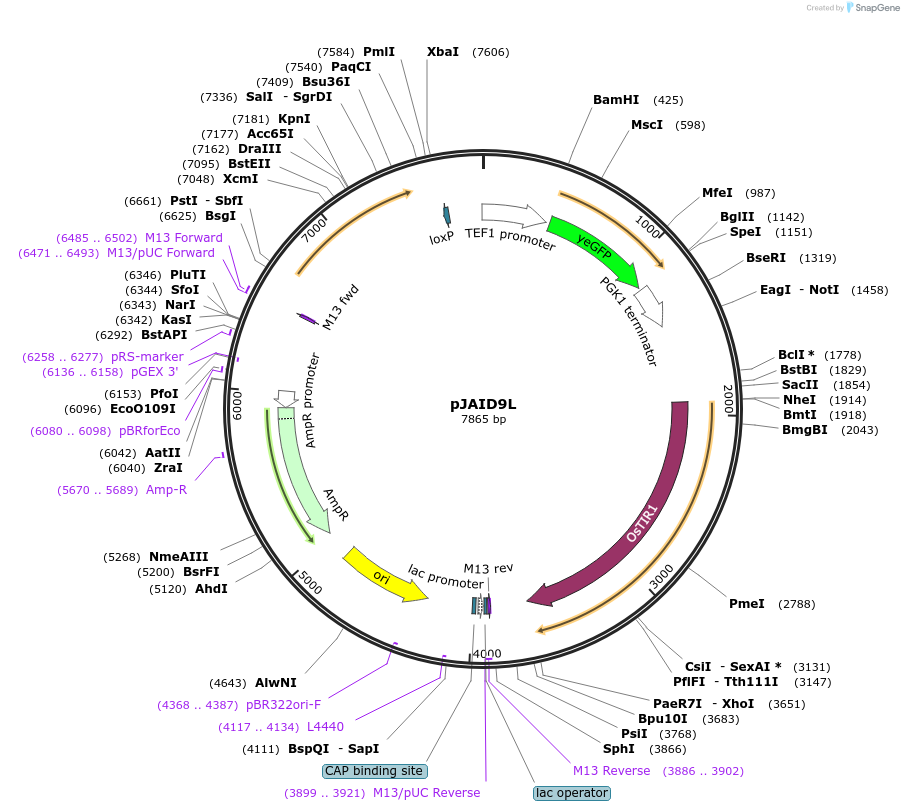
pJAID9L
Plasmid#165052PurposeIntegrative expression of yEGFP and rice auxin receptor gene OsTIR1. OsTIR1 is codon-optimised for expression in S. cerevisiaeDepositorInsertP-URA3>KlURA3>T-AgTEF1-P-TEF1>yEGFP>T-PGK1-P-ACS2>OsTIR1opt>T-URA3
ExpressionYeastMutationWTAvailable SinceMarch 12, 2021AvailabilityAcademic Institutions and Nonprofits only -
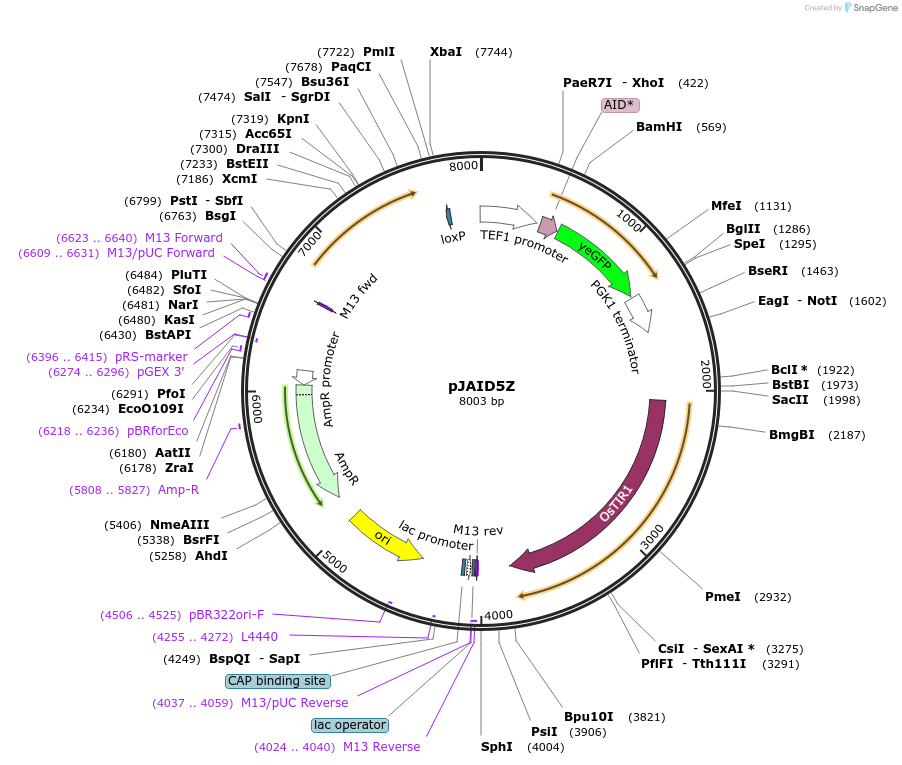
pJAID5Z
Plasmid#165047PurposeIntegrative expression of AID* (mini auxin inducible degron)-yEGFP and rice auxin receptor gene OsTIR1.DepositorInsertP-URA3>KlURA3>T-AgTEF1-P-TEF1>AID*>yEGFP>T-PGK1-P-ACS2>OsTIR1opt>T-URA3
ExpressionYeastMutationOsTIR1 is codon-optimised for expression in S. ce…Available SinceMarch 12, 2021AvailabilityAcademic Institutions and Nonprofits only -
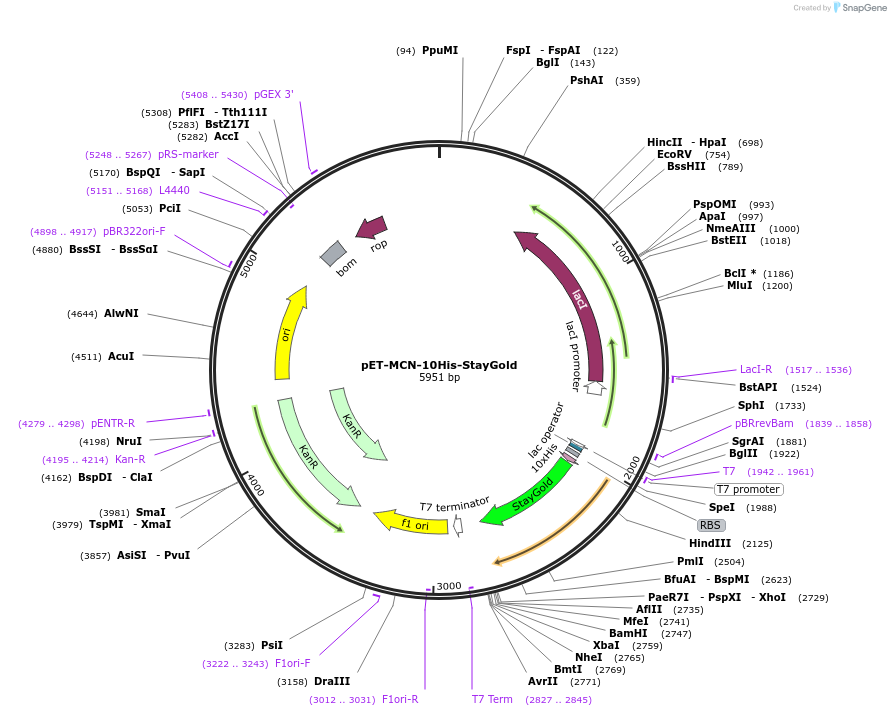
pET-MCN-10His-StayGold
Plasmid#211360PurposeBacterial expression of the bright and photostable StayGold fluorescent protein. Codon optimised for expression in Escherichia coli. Contains a N-terminal 10xHis tag.DepositorInsertStayGold
Tags10xHis-tagExpressionBacterialPromoterT7Available SinceDec. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
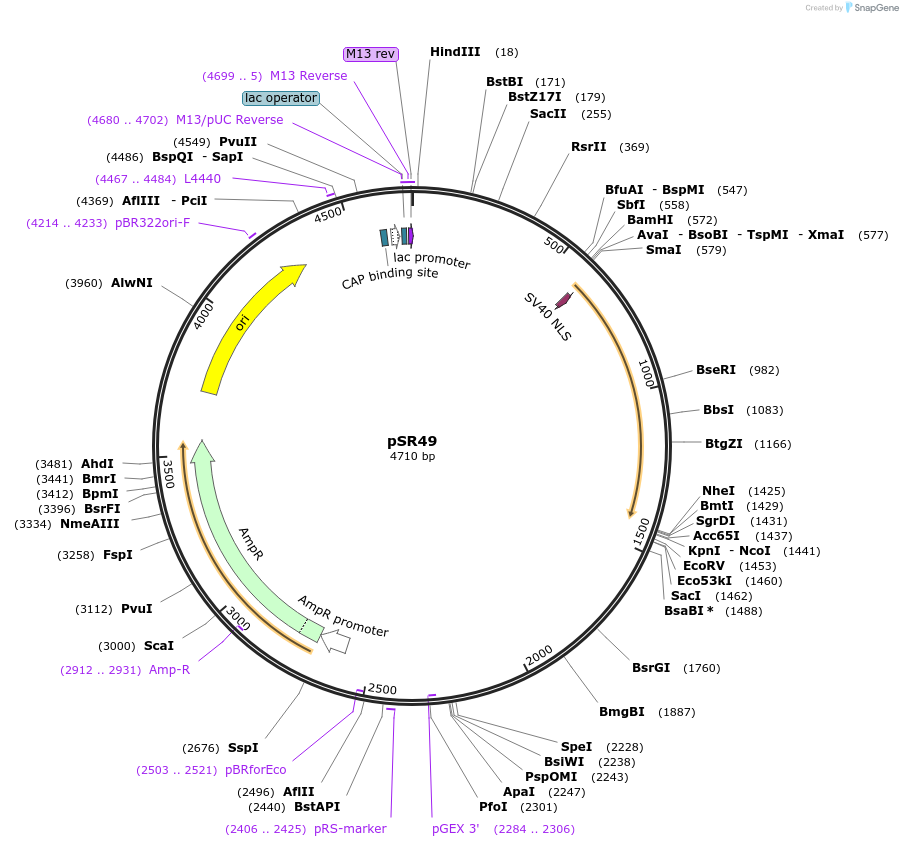
pSR49
Plasmid#69259Purposephlh-8::tagBFP2DepositorInsertTagBFP2
UseCre/LoxExpressionWormMutationcodon-optimzed index 1.0, TagBFP2 contains 2 sing…Promoterhlh-8Available SinceOct. 14, 2015AvailabilityAcademic Institutions and Nonprofits only -
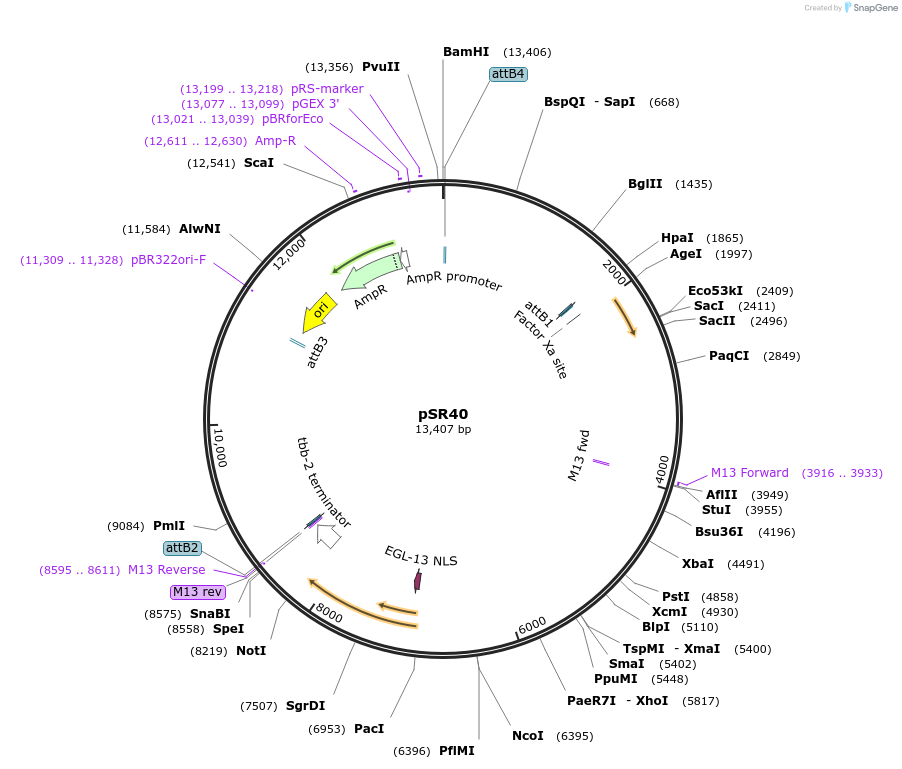
pSR40
Plasmid#69257Purposeplin-31::CRE for integration on ttTi14024, Chr XDepositorInsertCre recombinase
UseCre/Lox; MossciExpressionWormMutationcodon-optimzed index 1.0Promoterlin-31Available SinceOct. 14, 2015AvailabilityAcademic Institutions and Nonprofits only -
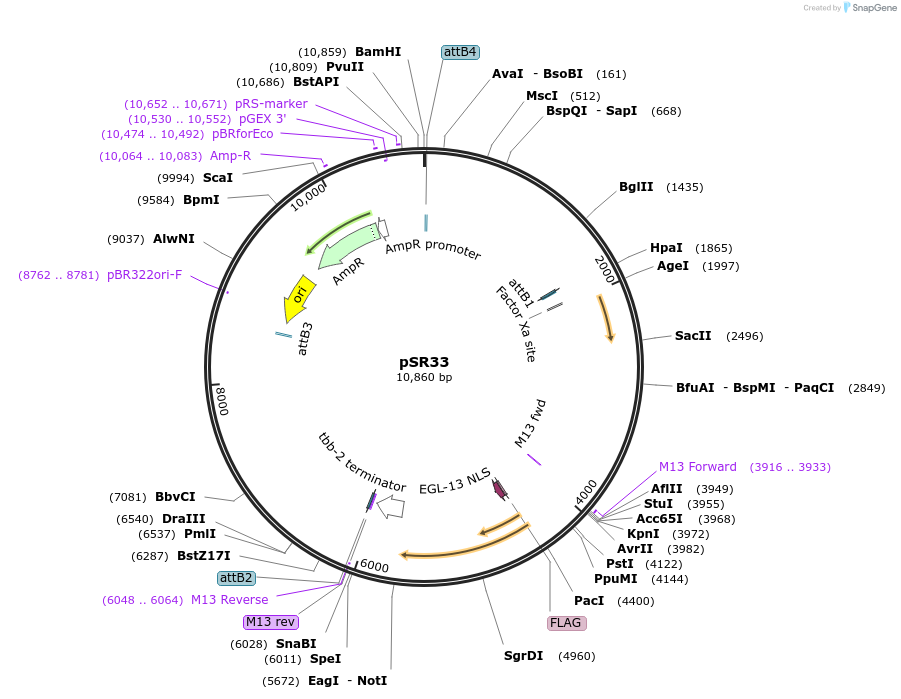
pSR33
Plasmid#69254Purposephsp16.2::CRE for integration on ttTi14024, Chr XDepositorInsertCre recombinase
UseCre/Lox; MossciExpressionWormMutationcodon-optimzed index 1.0Promoterhsp16.2Available SinceFeb. 12, 2016AvailabilityAcademic Institutions and Nonprofits only -
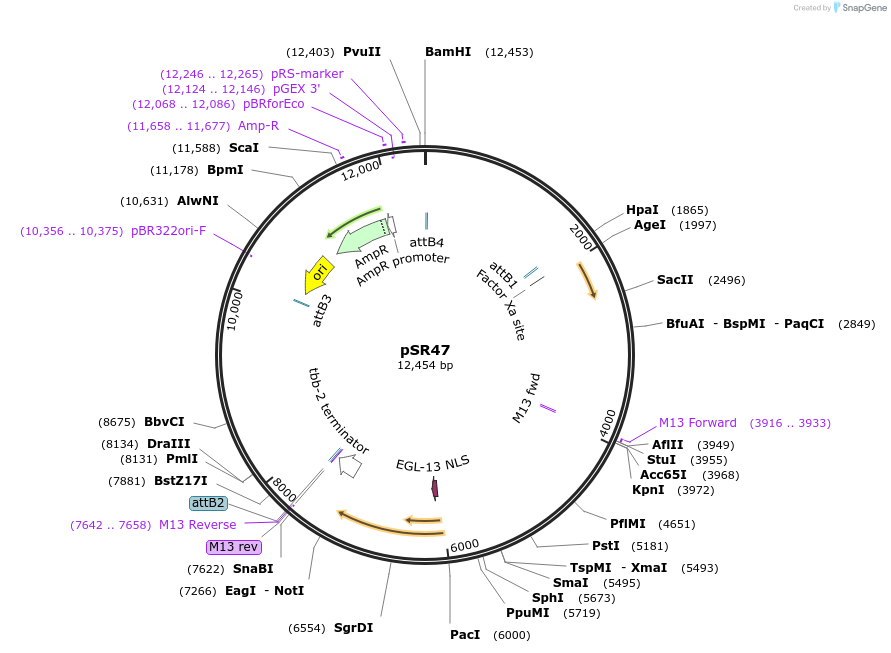
pSR47
Plasmid#69258Purposepmyo-3::CRE for integration on ttTi14024, Chr XDepositorInsertCre recombinase
UseCre/Lox; MossciExpressionWormMutationcodon-optimzed index 1.0Promotermyo-3Available SinceOct. 20, 2015AvailabilityAcademic Institutions and Nonprofits only -
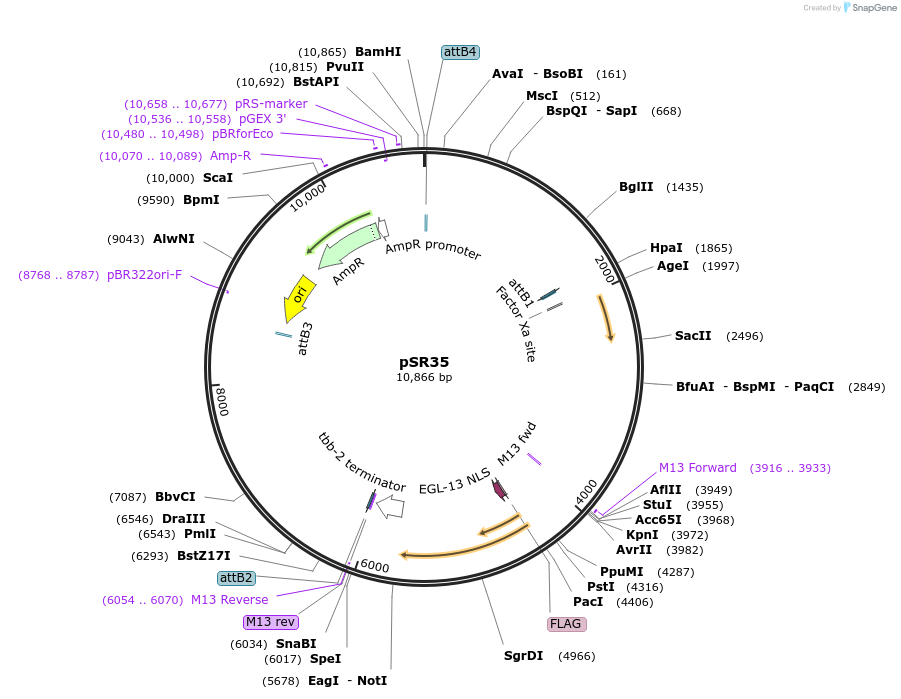
pSR35
Plasmid#69256Purposephsp16.41::CRE for integration on ttTi14024, Chr XDepositorInsertCre recombinase
UseCre/Lox; MossciExpressionWormMutationcodon-optimzed index 1.0Promoterhsp16.41Available SinceFeb. 12, 2016AvailabilityAcademic Institutions and Nonprofits only -
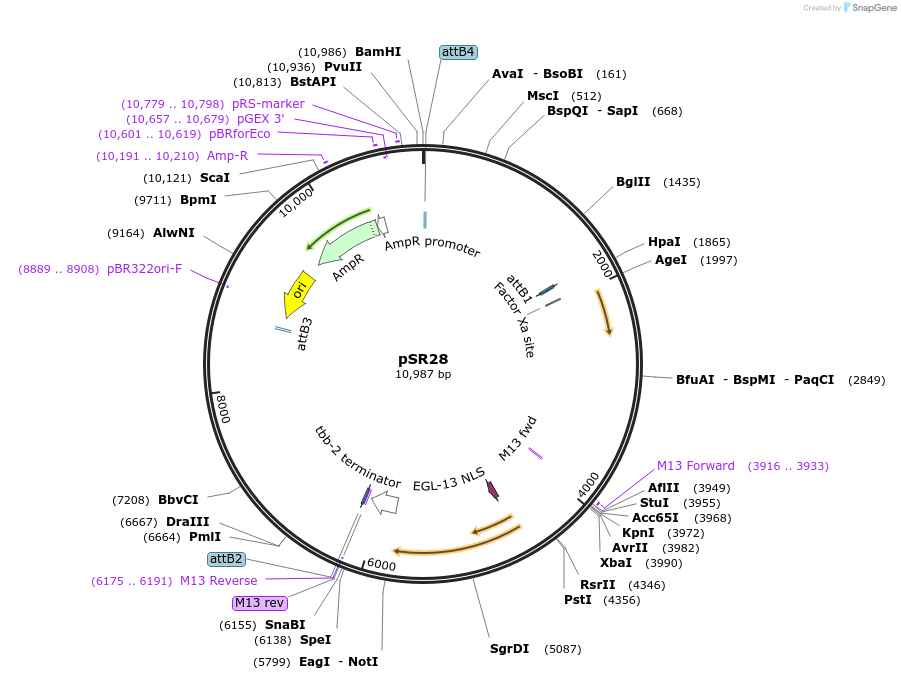
pSR28
Plasmid#69253Purposephlh-8::CRE for integration on ttTi14024, Chr XDepositorInsertCre recombinase
UseCre/Lox; MossciExpressionWormMutationcodon-optimzed index 1.0Promoterhlh-8Available SinceOct. 14, 2015AvailabilityAcademic Institutions and Nonprofits only -
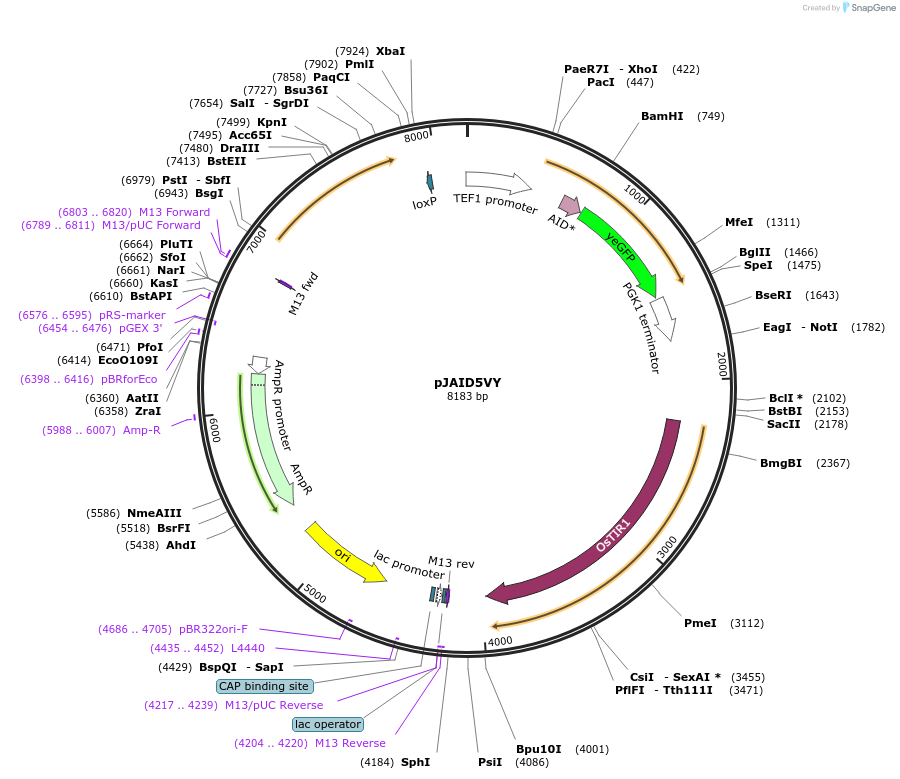
pJAID5VY
Plasmid#165048PurposeIntegrative expression of CUP1-AID* (mini auxin inducible degron)-yEGFP and rice auxin receptor gene OsTIR1.DepositorInsertP-URA3>KlURA3>T-AgTEF1-P-TEF1>CUP1-AID*>yEGFP>T-PGK1-P-ACS2>OsTIR1opt>T-URA3
ExpressionYeastMutationOsTIR1 is codon-optimised for expression in S. ce…Available SinceMarch 12, 2021AvailabilityAcademic Institutions and Nonprofits only -
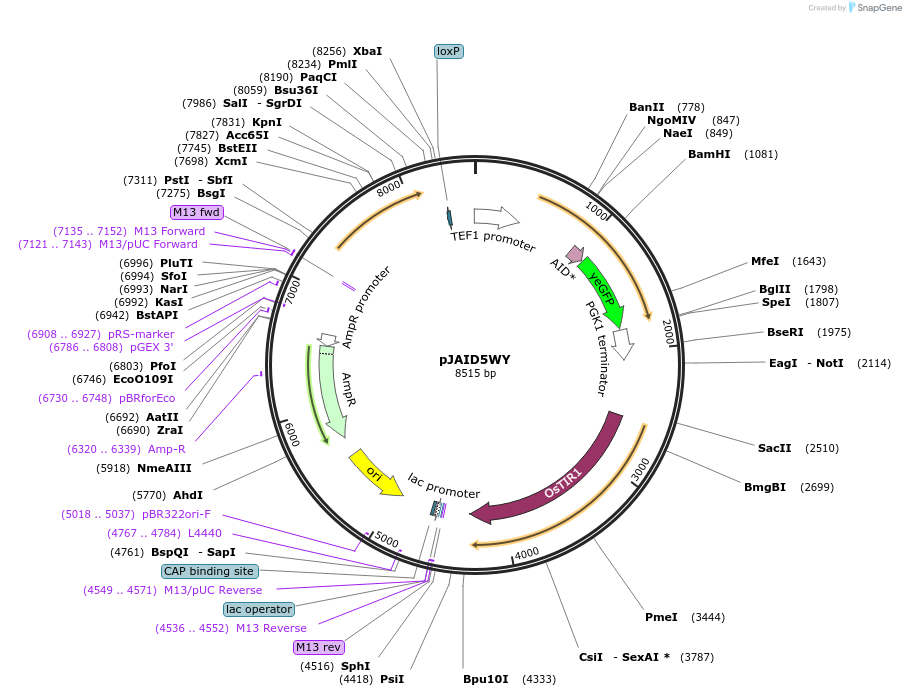
pJAID5WY
Plasmid#165050PurposeIntegrative expression of SOD1-AID* (mini auxin inducible degron)-yEGFP and rice auxin receptor gene OsTIR1.DepositorInsertP-URA3>KlURA3>T-AgTEF1-P-TEF1>SOD1-AID*>yEGFP>T-PGK1-P-ACS2>OsTIR1opt>T-URA3
ExpressionYeastMutationOsTIR1 is codon-optimised for expression in S. ce…Available SinceMarch 16, 2021AvailabilityAcademic Institutions and Nonprofits only -
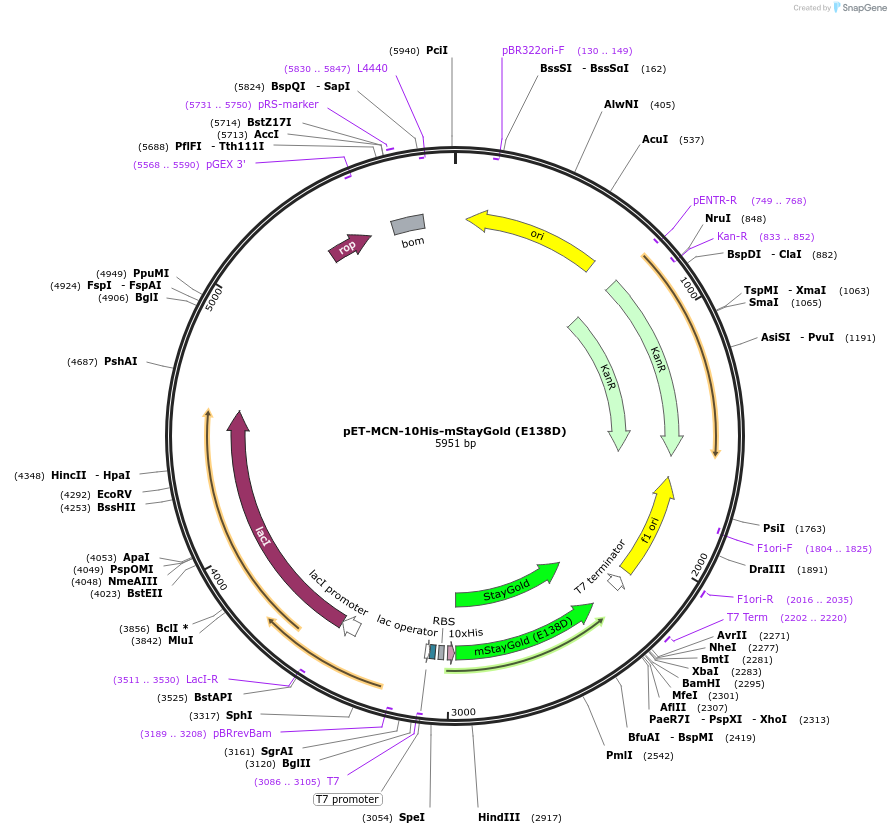
pET-MCN-10His-mStayGold (E138D)
Plasmid#211361PurposeBacterial expression of the bright and photostable monomeric StayGold fluorescent protein (E138D). Codon optimised for expression in Escherichia coli. Contains a N-terminal 10xHis tag.DepositorInsertmStayGold
Tags10xHis-tagExpressionBacterialMutationE138DPromoterT7Available SinceDec. 20, 2023AvailabilityAcademic Institutions and Nonprofits only -
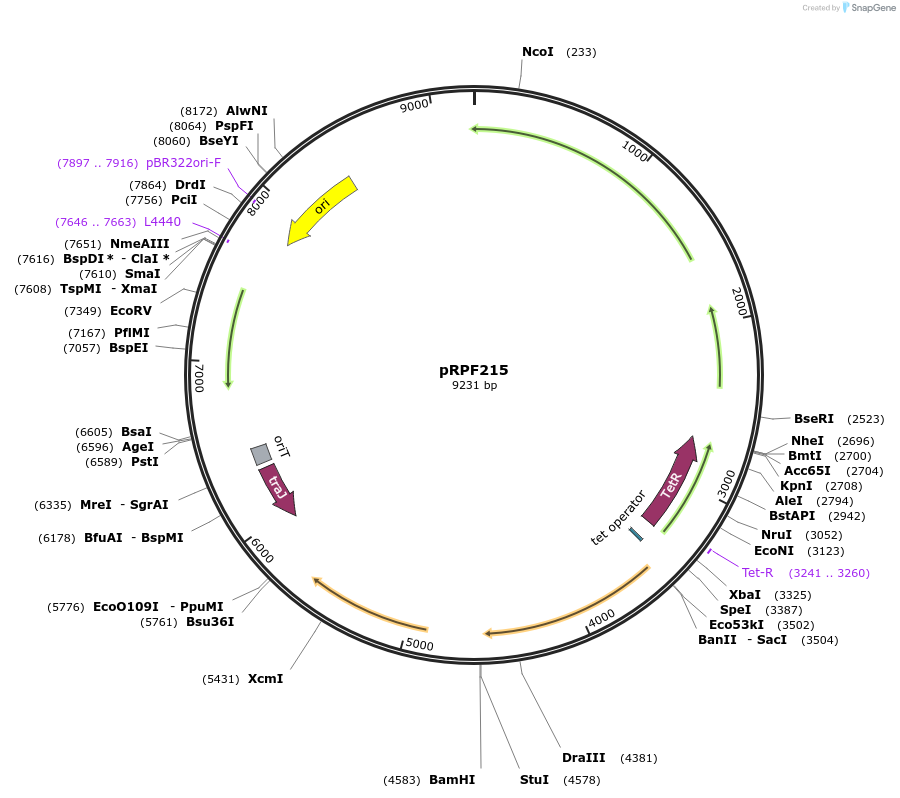
pRPF215
Plasmid#106377PurposeC. difficile transposon mutagenesis system. Shuttle vector containing tet-inducible Himar1 transposase and custom ErmR transposonDepositorInsertPtet-himar1-Ter(slpA)-transposon
UseE. coli - c. difficile shuttle vectorMutationCodon optimised for C. difficilePromoterPtetAvailable SinceMarch 8, 2018AvailabilityAcademic Institutions and Nonprofits only -
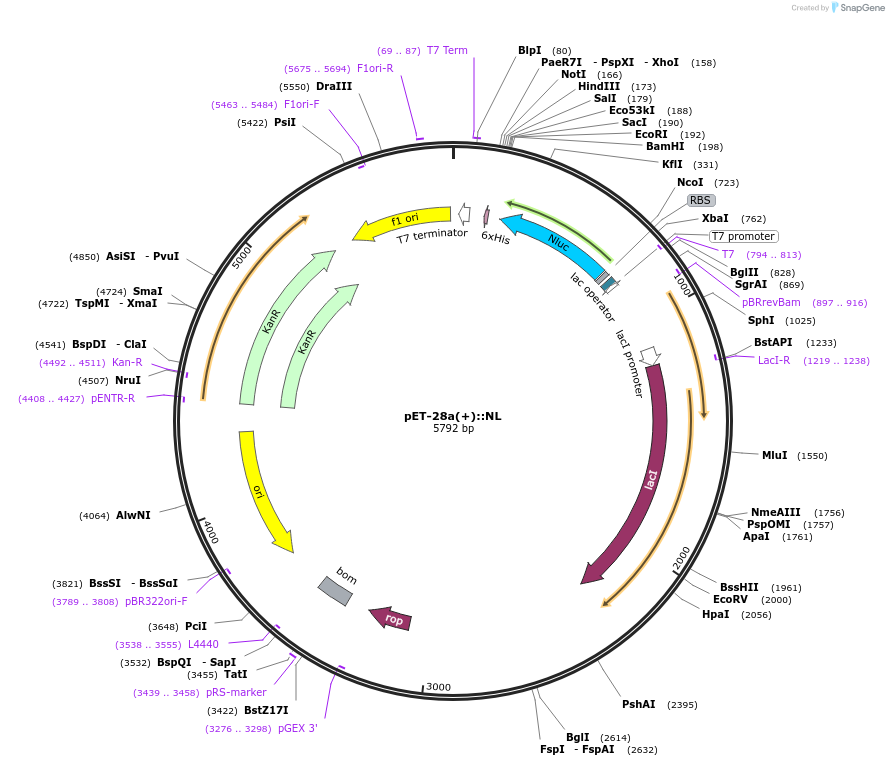
pET-28a(+)::NL
Plasmid#141289PurposeNLUC in pET-28 a (+) backbone for bacterial IPTG inducible expression (KmR)DepositorInsertNLUC
UseLuciferaseExpressionBacterialMutationEngineered for high stability (t1/2 = 11.5 days a…Promoterpromoter for bacteriophage T7 RNA polymeraseAvailable SinceFeb. 17, 2021AvailabilityAcademic Institutions and Nonprofits only -
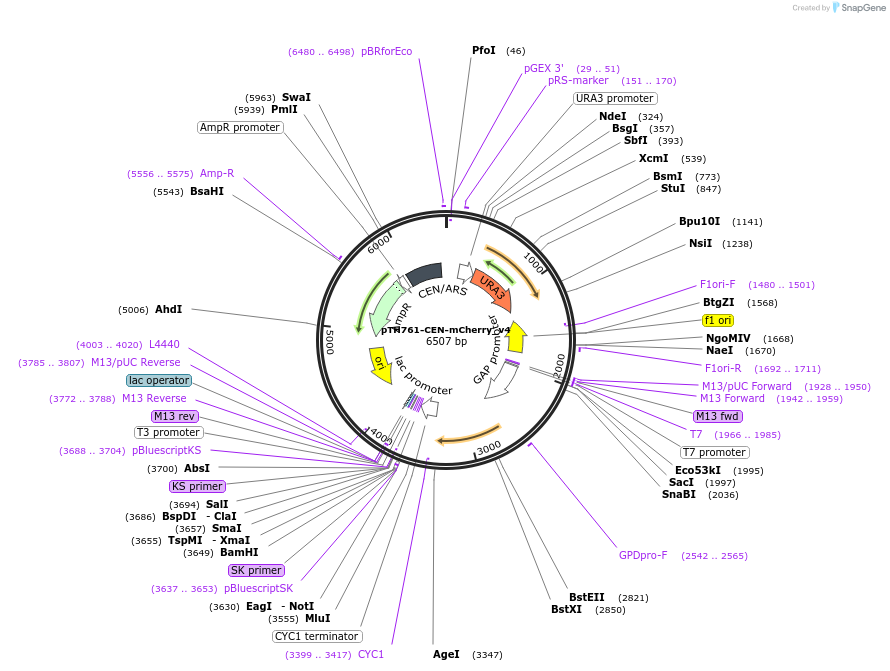
pTH761-CEN-mCherry_v4
Plasmid#40599DepositorInsertmCherry (Red Fluorescent Protein variant) with yeast promoter and terminator sequences
ExpressionYeastMutationmCherry ORF partially codon optimised for yeast e…PromoterGPD (TDH3)Available SinceOct. 18, 2012AvailabilityAcademic Institutions and Nonprofits only -
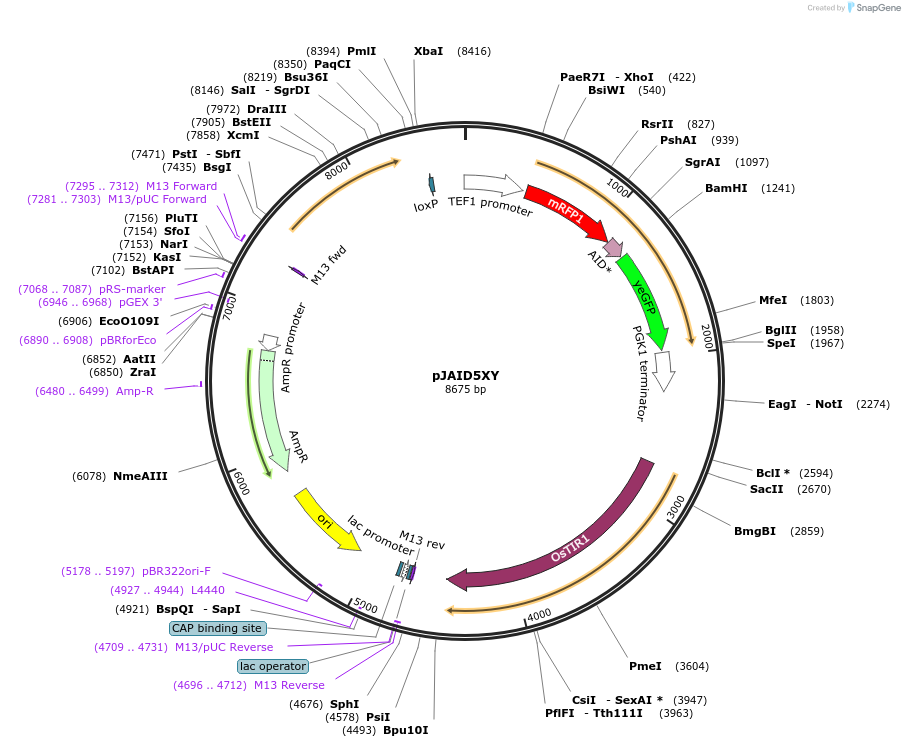
pJAID5XY
Plasmid#165051PurposeIntegrative expression of mRFP-CUP1-AID* (mini auxin inducible degron)-yEGFP and rice auxin receptor gene OsTIR1.DepositorInsertP-URA3>KlURA3>T-AgTEF1-P-TEF1>mRFP-AID*>yEGFP>T-PGK1-P-ACS2>OsTIR1opt>T-URA3
ExpressionYeastMutationOsTIR1 is codon-optimised for expression in S. ce…Available SinceMarch 12, 2021AvailabilityAcademic Institutions and Nonprofits only -
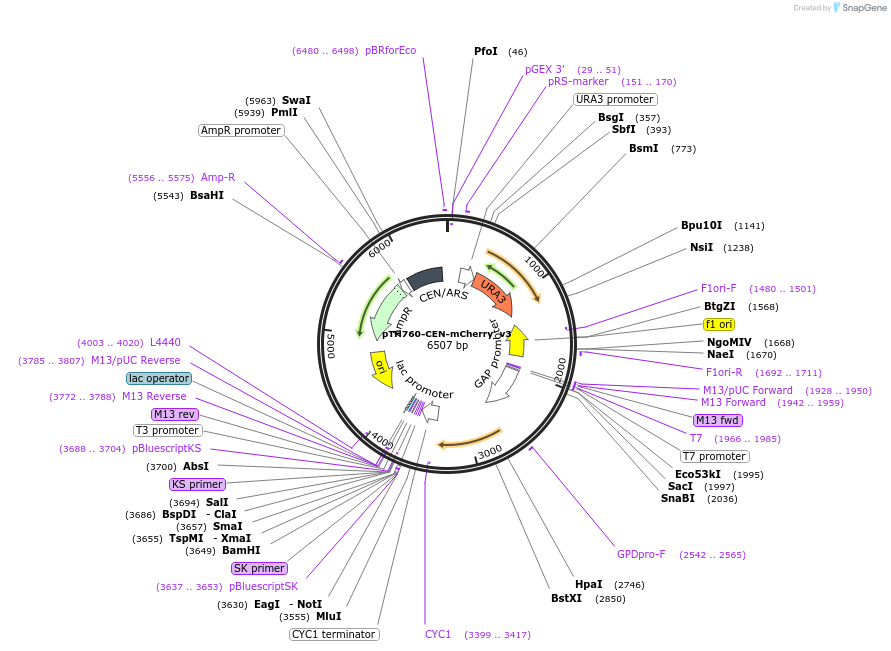
pTH760-CEN-mCherry_v3
Plasmid#40598DepositorInsertmCherry (Red Fluorescent Protein variant) with yeast promoter and terminator sequences
ExpressionYeastMutationmCherry ORF partially codon optimised for yeast e…PromoterGPD (TDH3)Available SinceOct. 18, 2012AvailabilityAcademic Institutions and Nonprofits only -
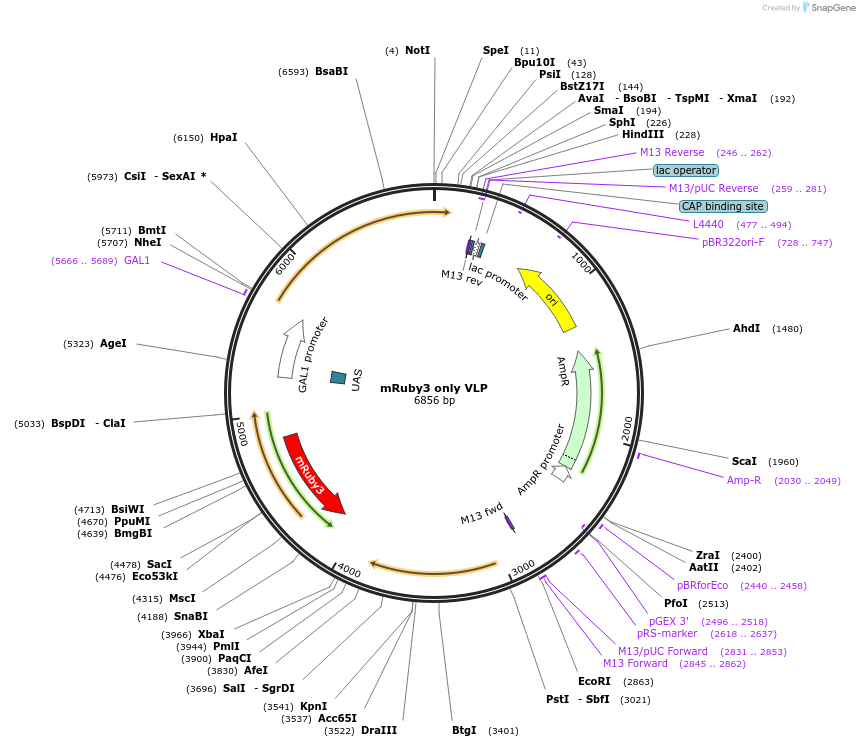
mRuby3 only VLP
Plasmid#182429PurposeYIp for expressing Murine polyomavirus deltaVP1 (GAL1 promoter) and mRuby3 tagged with VP2C (GAL10 promoter). mRuby3 was codon-optimised for E. coli. Contains uracil marker (K. lactis URA3).DepositorInsertsVP2C-mRuby3
deltaVP1
UseSynthetic BiologyExpressionYeastPromoterGAL1 and GAL10Available SinceDec. 6, 2022AvailabilityAcademic Institutions and Nonprofits only -
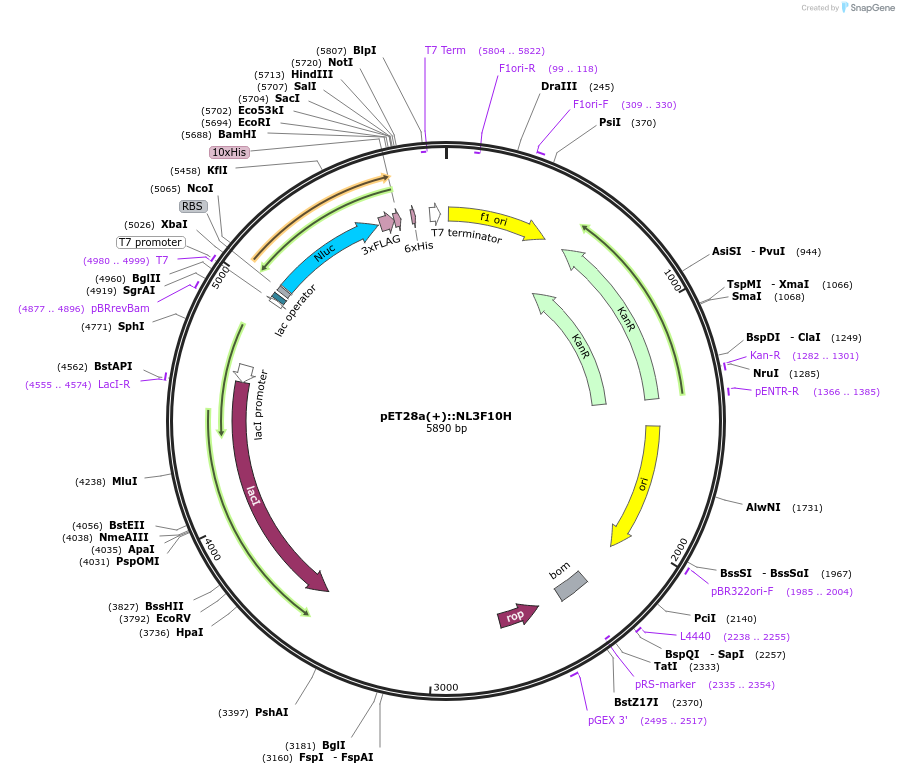
pET28a(+)::NL3F10H
Plasmid#141290PurposeNLUC:3xFlag:10xHis in pET-28 a (+) backbone for bacterial IPTG inducible expression (KmR)DepositorInsertNLUC3F10H
UseLuciferaseTagsFlag (x3) and His (x10)ExpressionBacterialMutationEngineered for high stability (t1/2 = 11.5 days a…Promoterpromoter for bacteriophage T7 RNA polymeraseAvailable SinceFeb. 17, 2021AvailabilityAcademic Institutions and Nonprofits only -
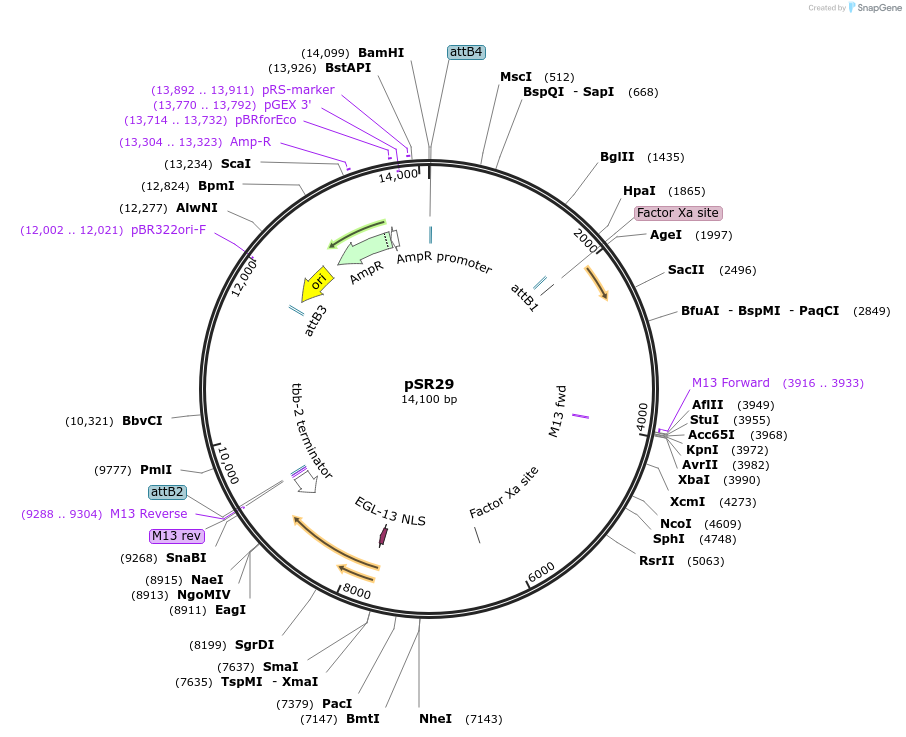
pSR29
Plasmid#69255Purposepelt-2::CRE for integration on ttTi14024, Chr XDepositorInsertCre recombinase
UseCre/Lox; MossciExpressionWormMutationcodon-optimzed index 1.0, tbb-2 UTR is in wrong o…Promoterelt-2Available SinceOct. 20, 2015AvailabilityAcademic Institutions and Nonprofits only -
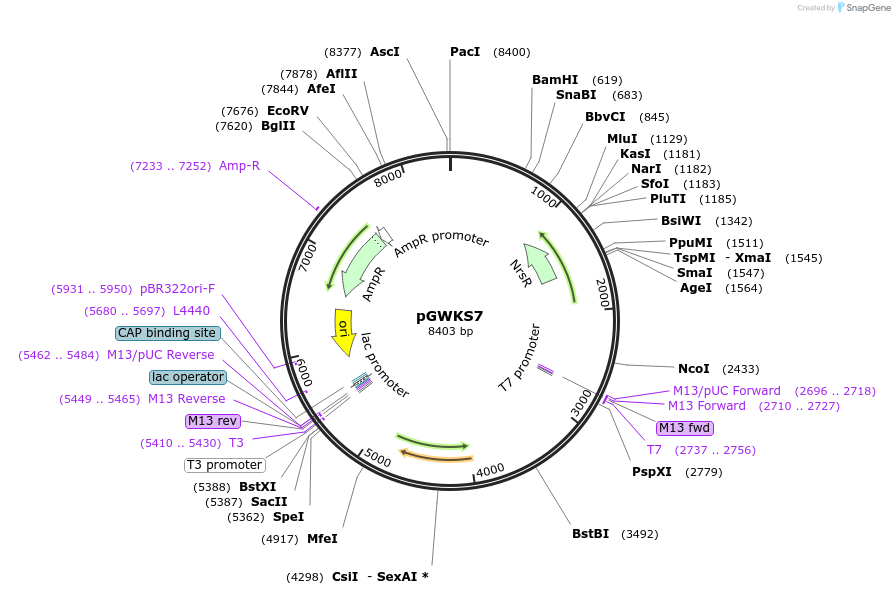
pGWKS7
Plasmid#139414PurposeFluoroescent marker for use in Cryptococcus neoformans. Contains a codon optimised mRuby3 marker.DepositorInsertmRuby3
ExpressionYeastPromoterTEF1Available SinceMay 4, 2020AvailabilityAcademic Institutions and Nonprofits only -
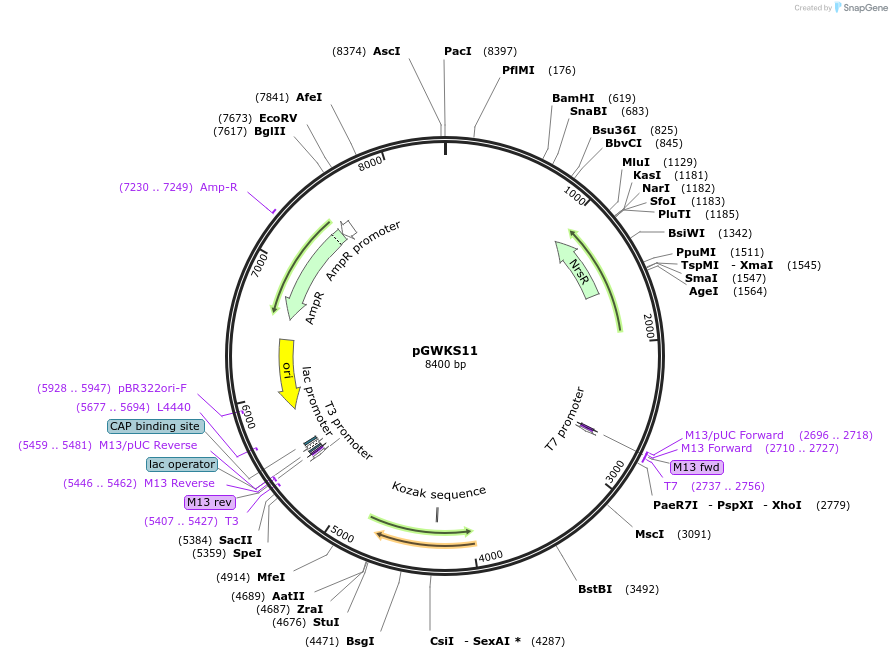
pGWKS11
Plasmid#139418PurposeFluoroescent marker for use in Cryptococcus neoformans. Contains a codon optimised mNeonGreen marker.DepositorInsertmNeonGreen
ExpressionYeastPromoterTEF1Available SinceMay 4, 2020AvailabilityAcademic Institutions and Nonprofits only -
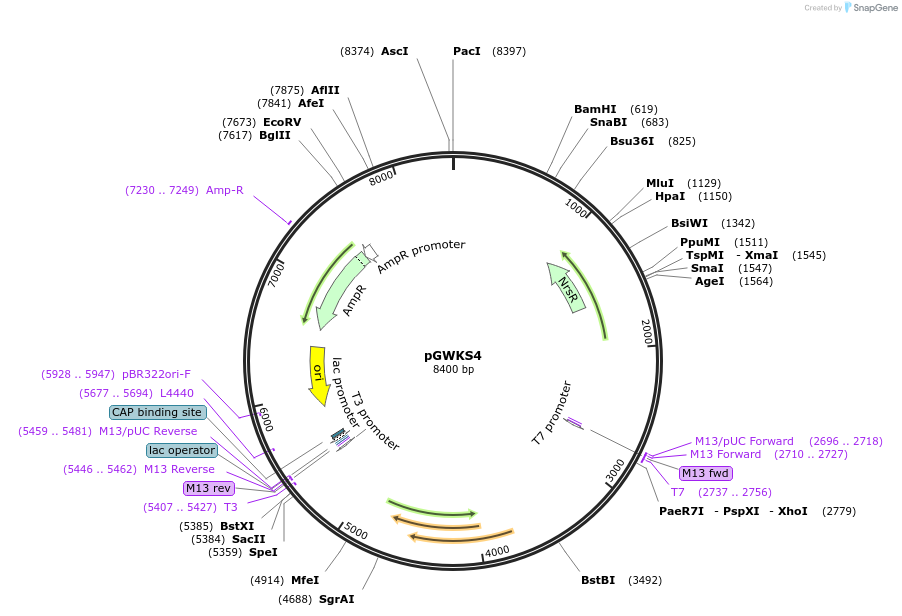
pGWKS4
Plasmid#139411PurposeFluoroescent marker for use in Cryptococcus neoformans. Contains a codon optimised mCherry marker.DepositorInsertmCherry
ExpressionYeastPromoterTEF1Available SinceMay 4, 2020AvailabilityAcademic Institutions and Nonprofits only -
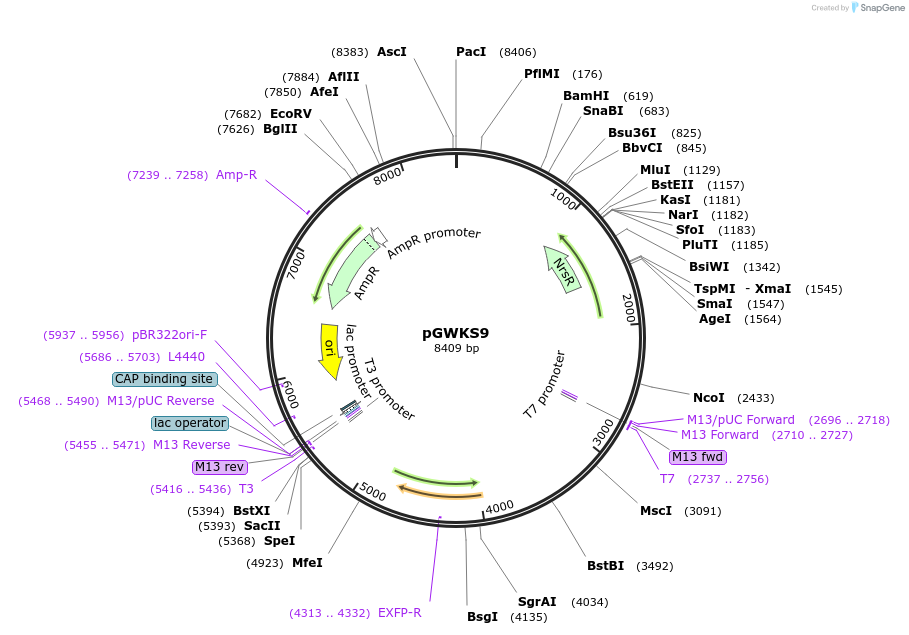
pGWKS9
Plasmid#139416PurposeFluoroescent marker for use in Cryptococcus neoformans. Contains a codon optimised mTurquoise2 marker.DepositorInsertmTurquoise2
ExpressionYeastPromoterTEF1Available SinceOct. 6, 2020AvailabilityAcademic Institutions and Nonprofits only