We narrowed to 6,009 results for: pCas
-
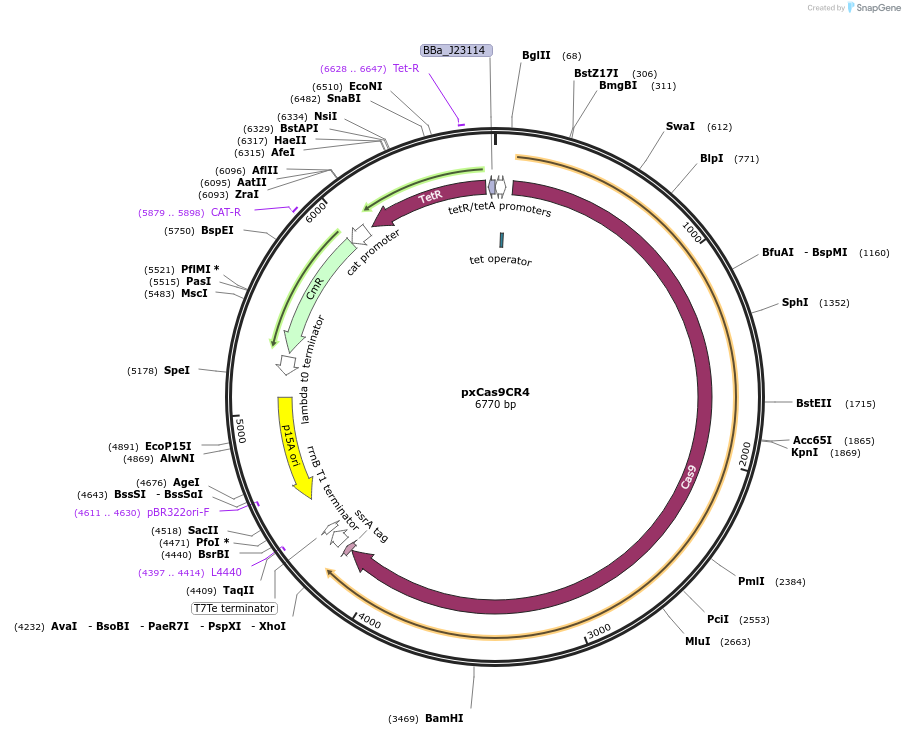
Plasmid#111656PurposePCas9CR4 with the xCas9 mutations allowing NG PAM sitesDepositorInsertTetR and xCas9
UseSynthetic BiologyTagsssrAExpressionBacterialMutationxCas9 mutationsAvailable SinceJuly 30, 2018AvailabilityAcademic Institutions and Nonprofits only -
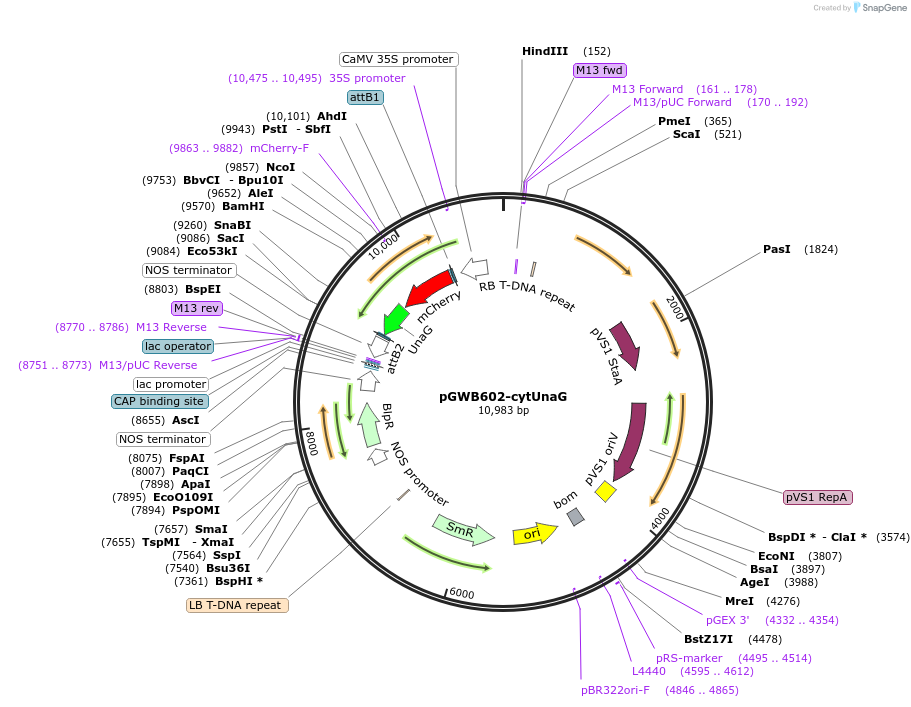
pGWB602-cytUnaG
Plasmid#220101PurposeFor Agrobacterium-mediated transformation. Cytosol-localized mCherry-UnaG.DepositorInsertmCherry-UnaG
ExpressionPlantAvailable SinceOct. 15, 2025AvailabilityAcademic Institutions and Nonprofits only -
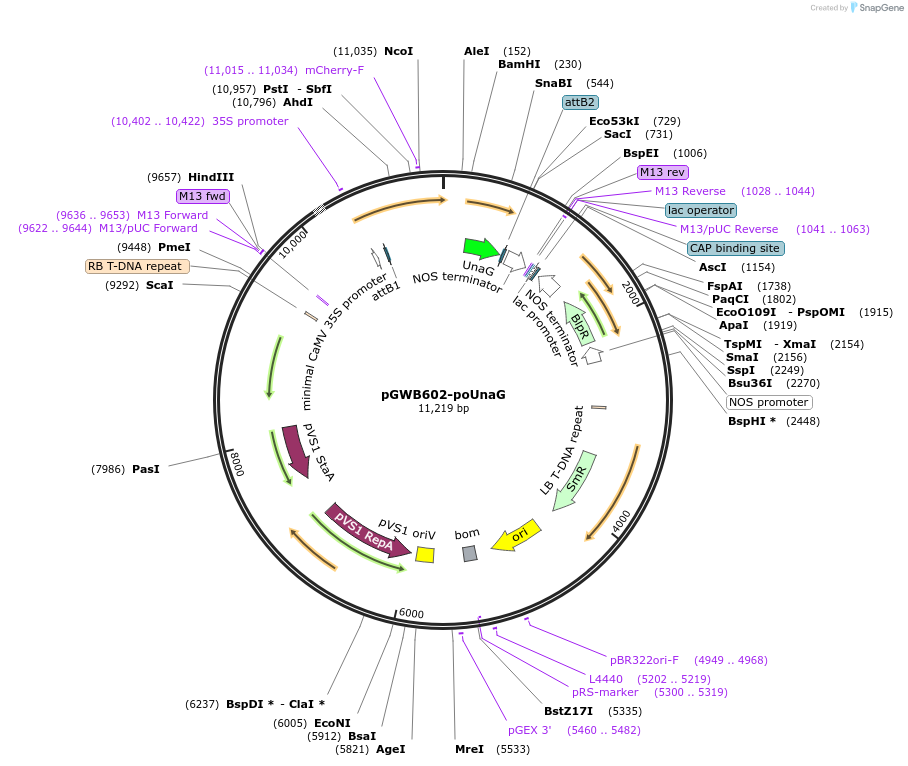
pGWB602-poUnaG
Plasmid#220103PurposeFor Agrobacterium-mediated transformation. Peroxisome-localized mCherry-UnaG.DepositorInsertmCherry-UnaG
ExpressionPlantAvailable SinceOct. 11, 2024AvailabilityAcademic Institutions and Nonprofits only -
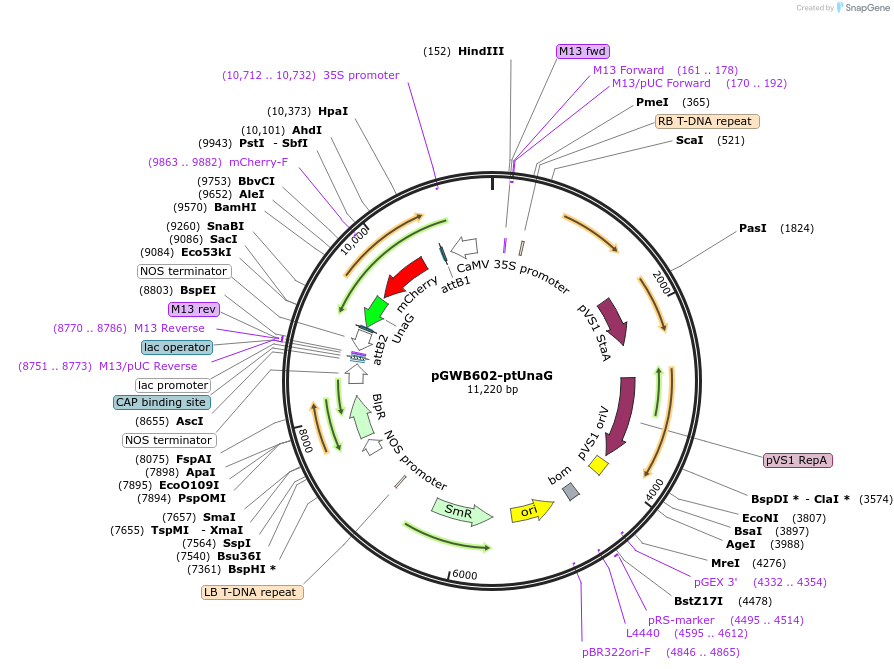
pGWB602-ptUnaG
Plasmid#221660PurposeFor Agrobacterium-mediated transformation. Plastid(stroma)-localized mCherry-UnaG.DepositorInsertmCherry-UnaG
ExpressionPlantAvailable SinceOct. 11, 2024AvailabilityAcademic Institutions and Nonprofits only -
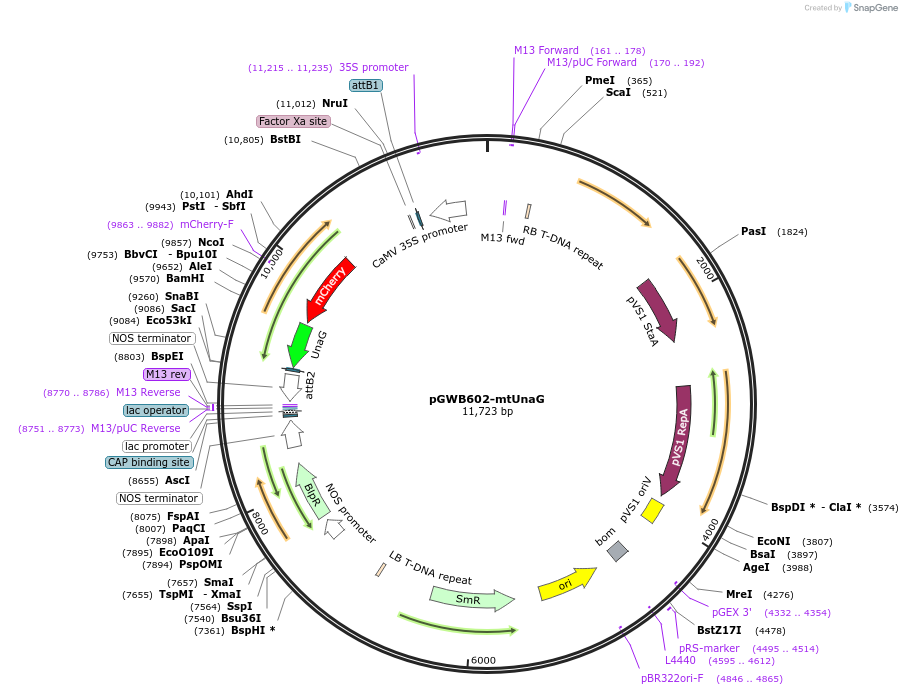
pGWB602-mtUnaG
Plasmid#221664PurposeFor Agrobacterium-mediated transformation. Mitochondria-localized mCherry-UnaGDepositorInsertmCherry-UnaG flanked with an N-terminal presequence of F1-ATPase γ-subunit
ExpressionPlantAvailable SinceOct. 11, 2024AvailabilityAcademic Institutions and Nonprofits only -
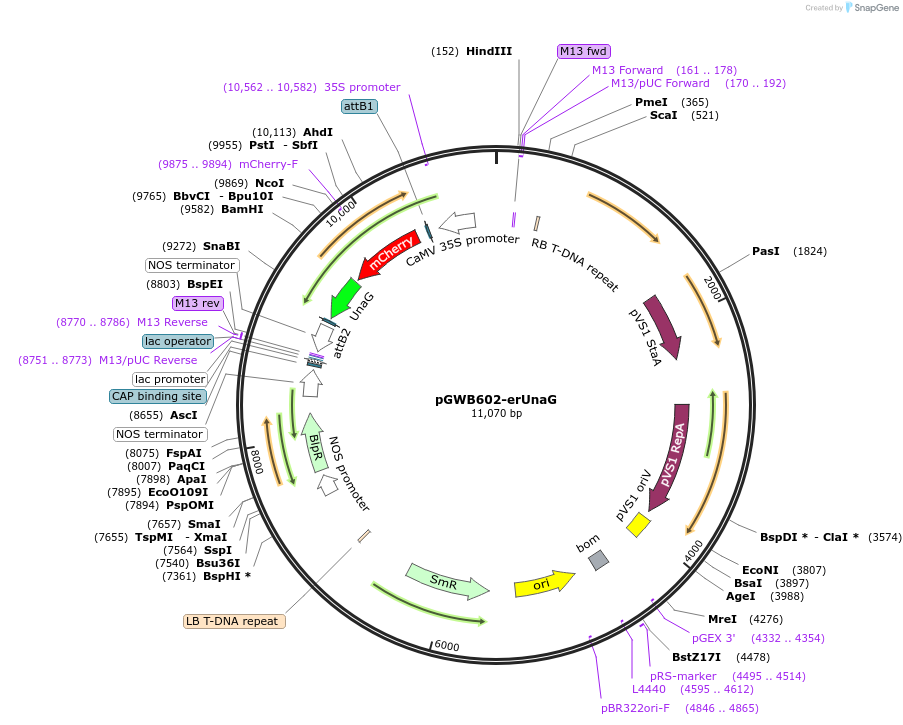
pGWB602-erUnaG
Plasmid#221663PurposeFor Agrobacterium-mediated transformation. Endoplasmic reticulum-localized mCherry-UnaG.DepositorInsertmCherry-UnaG flanked with an N-terminal signal peptide or field pumpkin 25 albumin
ExpressionPlantAvailable SinceOct. 15, 2024AvailabilityAcademic Institutions and Nonprofits only -
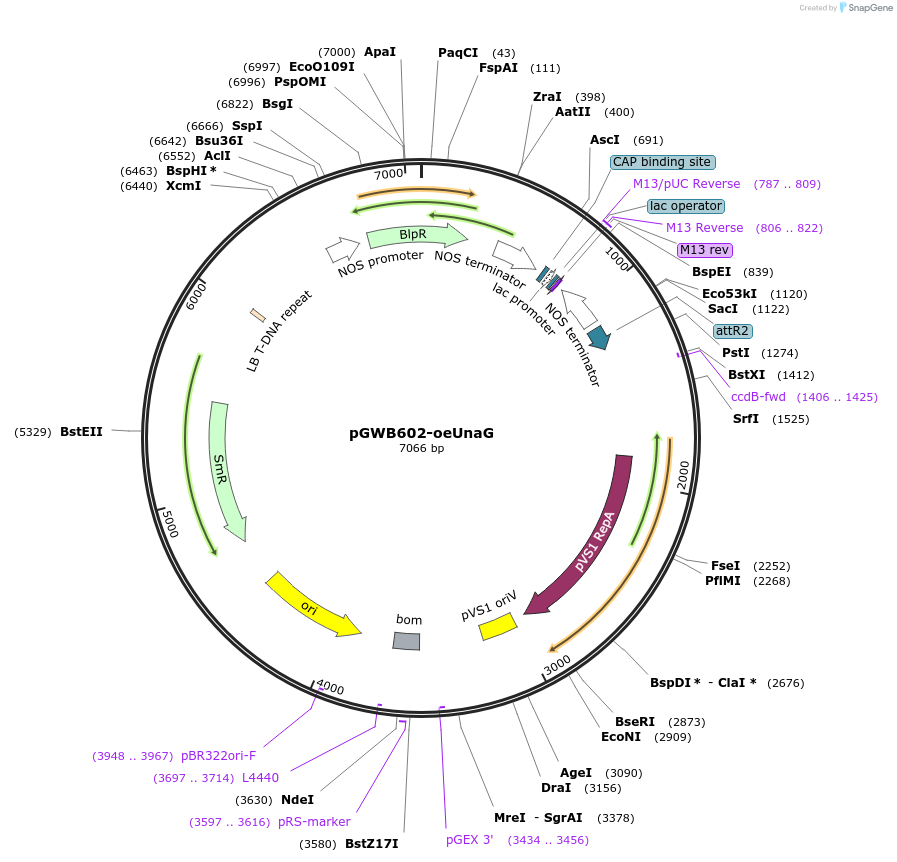
pGWB602-oeUnaG
Plasmid#221662PurposeFor Agrobacterium-mediated transformation. Plastid(outer envelope membrane)-localized mCherry-UnaG.DepositorInsertmCherry-UnaG flanked with an N-terminal sequence of Arabidopsis OUTRE ENVELOPE PROTEIN7
ExpressionPlantAvailable SinceOct. 15, 2024AvailabilityAcademic Institutions and Nonprofits only -
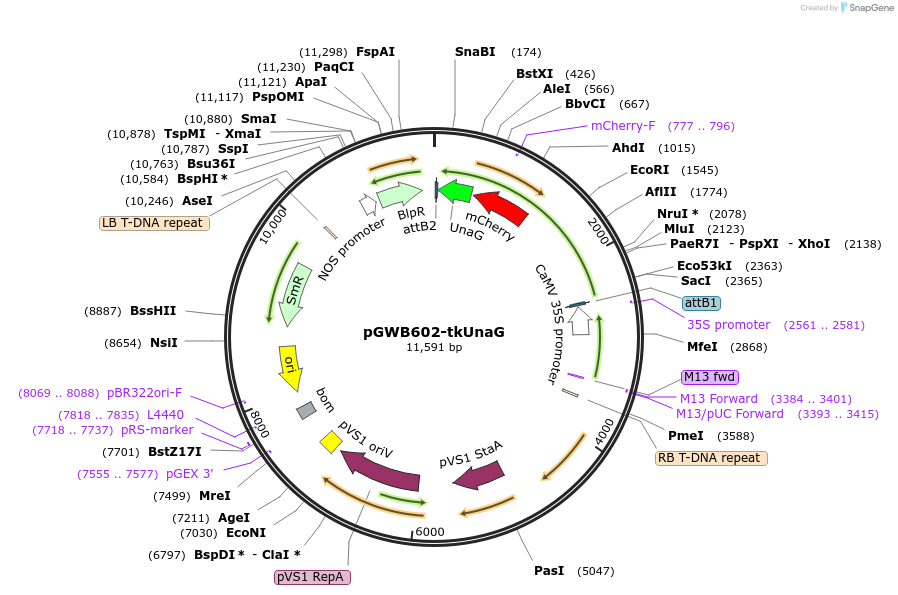
pGWB602-tkUnaG
Plasmid#221661PurposeFor Agrobacterium-mediated transformation. Plastid(thylakoid membrane)-localized mCherry-UnaG.DepositorInsertmCherry-UnaG flanked with the full-length coding sequence for Arabidopsis BESTROPHIN-LIKE PROTEIN2
ExpressionPlantAvailable SinceOct. 11, 2024AvailabilityAcademic Institutions and Nonprofits only -
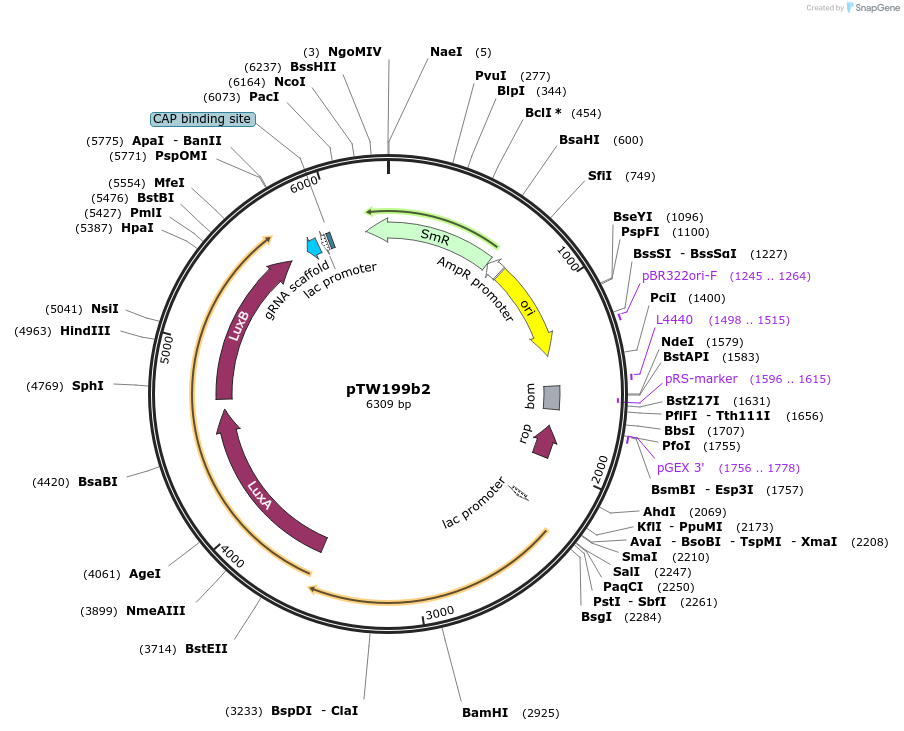
pTW199b2
Plasmid#136928PurposeDirected evolution of SpCas9DepositorInsertgIII
UseCRISPRExpressionBacterialMutationnoneAvailable SinceFeb. 11, 2020AvailabilityAcademic Institutions and Nonprofits only -
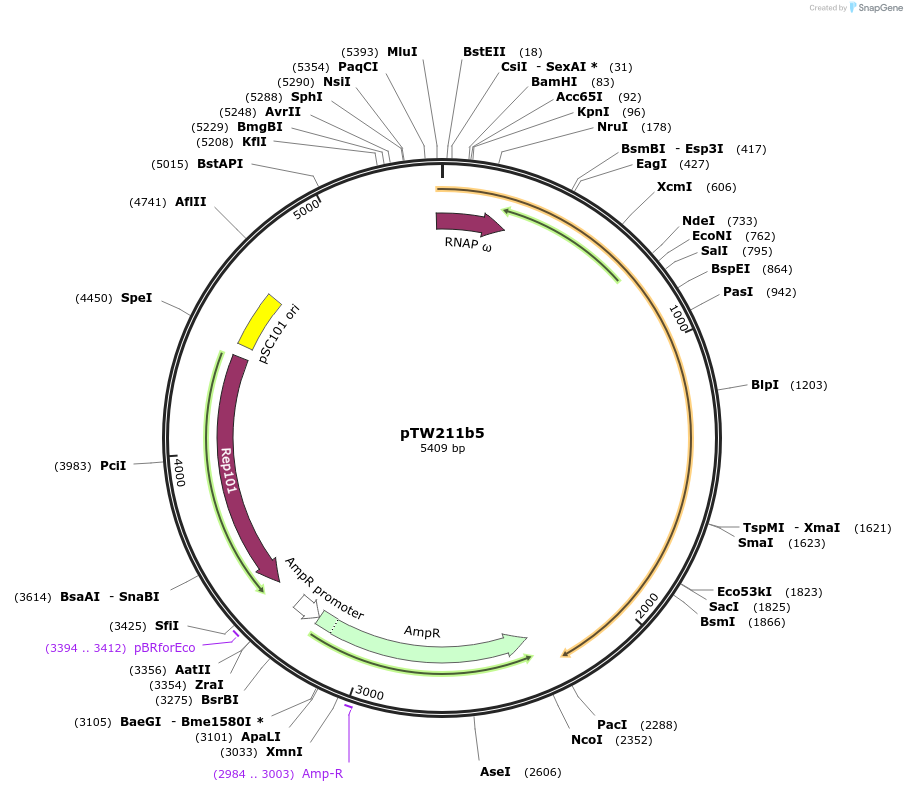
pTW211b5
Plasmid#136927PurposeDirected evolution of SpCas9DepositorInsertCas9(N)-NpuN
UseCRISPRExpressionBacterialMutationsee manuscriptAvailable SinceFeb. 11, 2020AvailabilityAcademic Institutions and Nonprofits only -
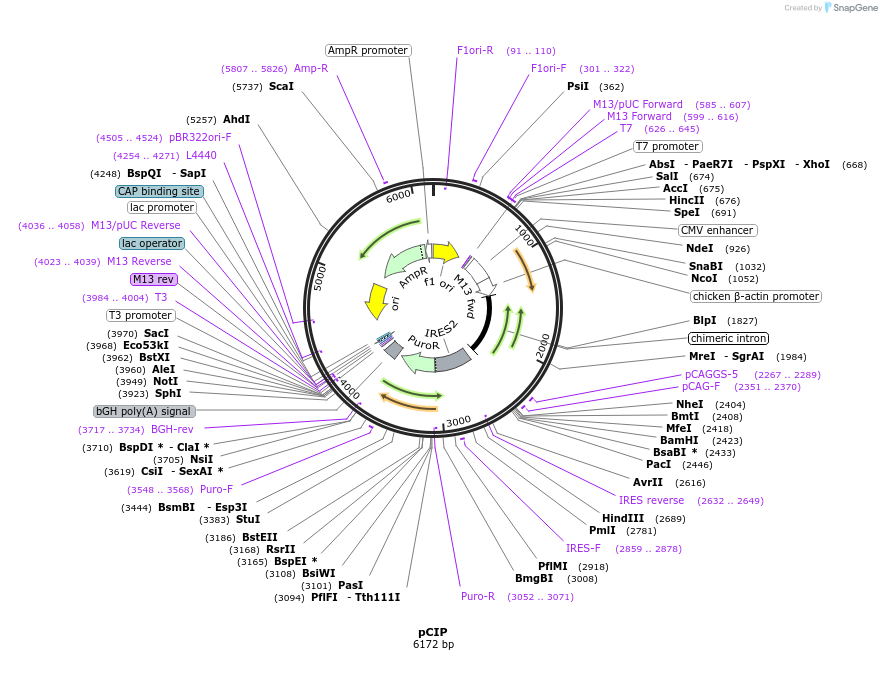
pCIP
Plasmid#79009PurposeFor expression in mouse ESCs or other eukaryotic cells. Contains pCAGGS-IRES-Puro-bpA, with a NheI-MfeI cloning site in between pCAGGS and IRES.DepositorTypeEmpty backboneExpressionMammalianPromoterpCAGGsAvailable SinceJuly 5, 2016AvailabilityAcademic Institutions and Nonprofits only -
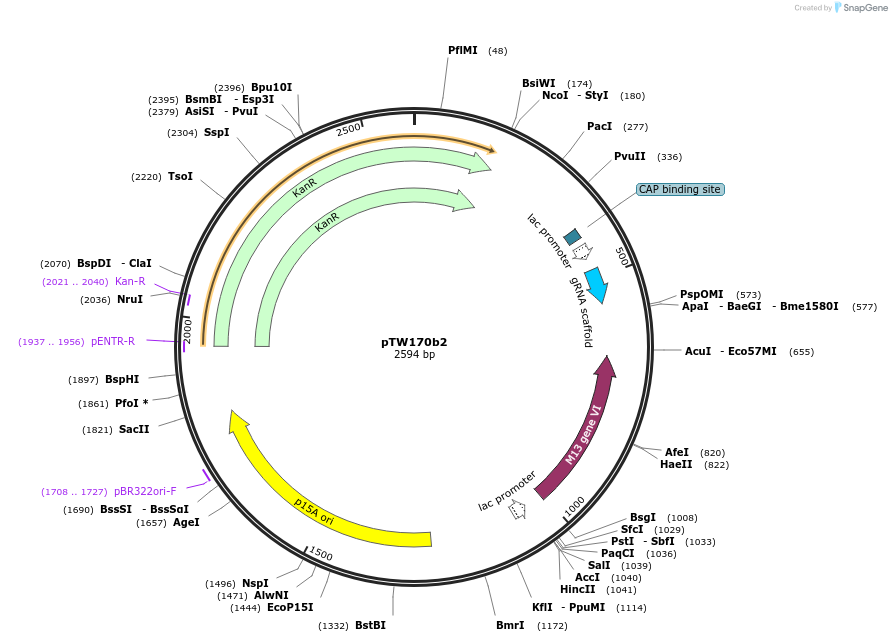
pTW170b2
Plasmid#136929PurposeDirected evolution of SpCas9DepositorInsertgVI
UseCRISPRExpressionBacterialMutationnoneAvailable SinceFeb. 20, 2020AvailabilityAcademic Institutions and Nonprofits only -
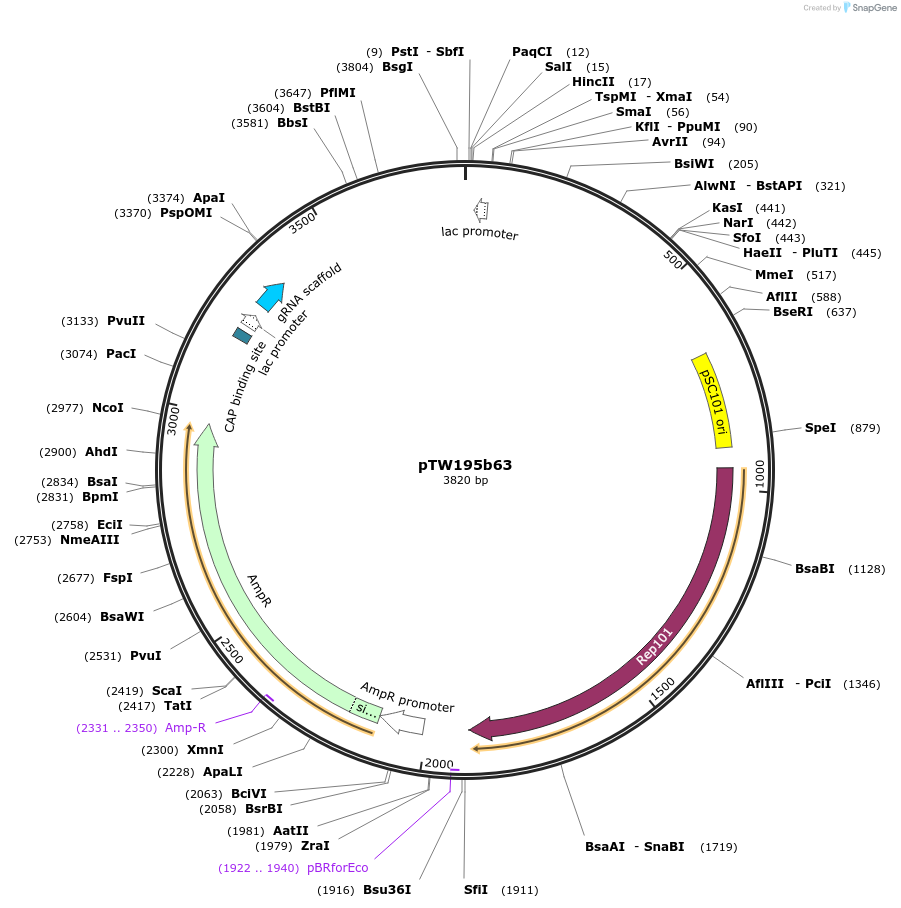
pTW195b63
Plasmid#136931PurposeDirected evolution of SpCas11DepositorInsertgIII(N)-npuN
UseCRISPRExpressionBacterialMutationnoneAvailable SinceFeb. 11, 2020AvailabilityAcademic Institutions and Nonprofits only -
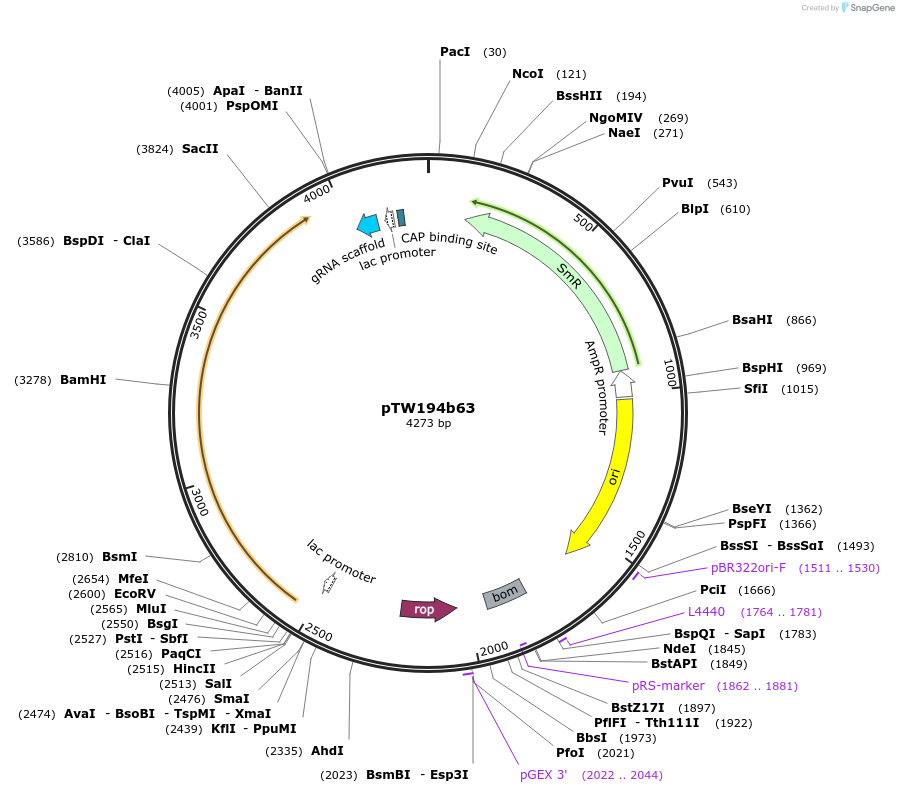
pTW194b63
Plasmid#136930PurposeDirected evolution of SpCas10DepositorInsertnpuC-gIII(C)
UseCRISPRExpressionBacterialMutationnoneAvailable SinceFeb. 11, 2020AvailabilityAcademic Institutions and Nonprofits only -
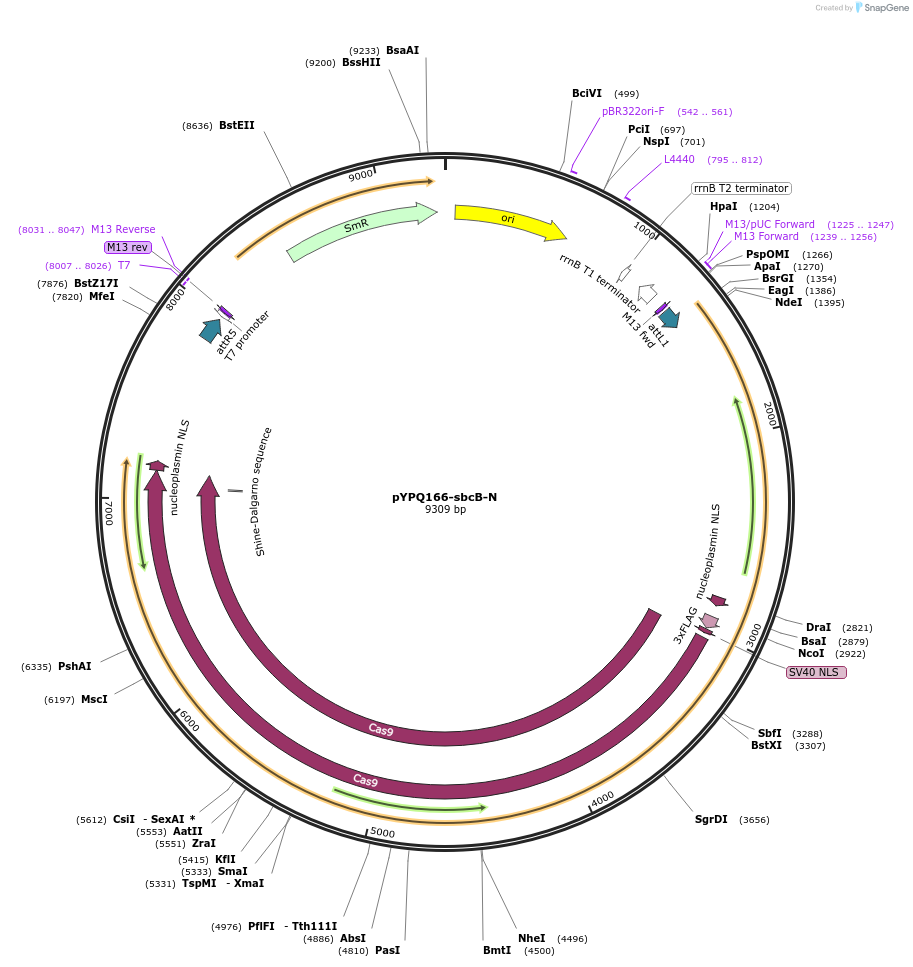
pYPQ166-sbcB-N
Plasmid#220305PurposeGateway entry plasmid (attL1& attR5) expressing sbcB exonuclease fused to the N-terminus of zSpCas9, connected by a flexible XTEN linker, without promoterDepositorInsertsbcB-XTEN linker-zSpCas9
UseCRISPR; Gateway compatible sbcb-xten linker- zspc…ExpressionPlantAvailable SinceJan. 7, 2026AvailabilityAcademic Institutions and Nonprofits only -
-
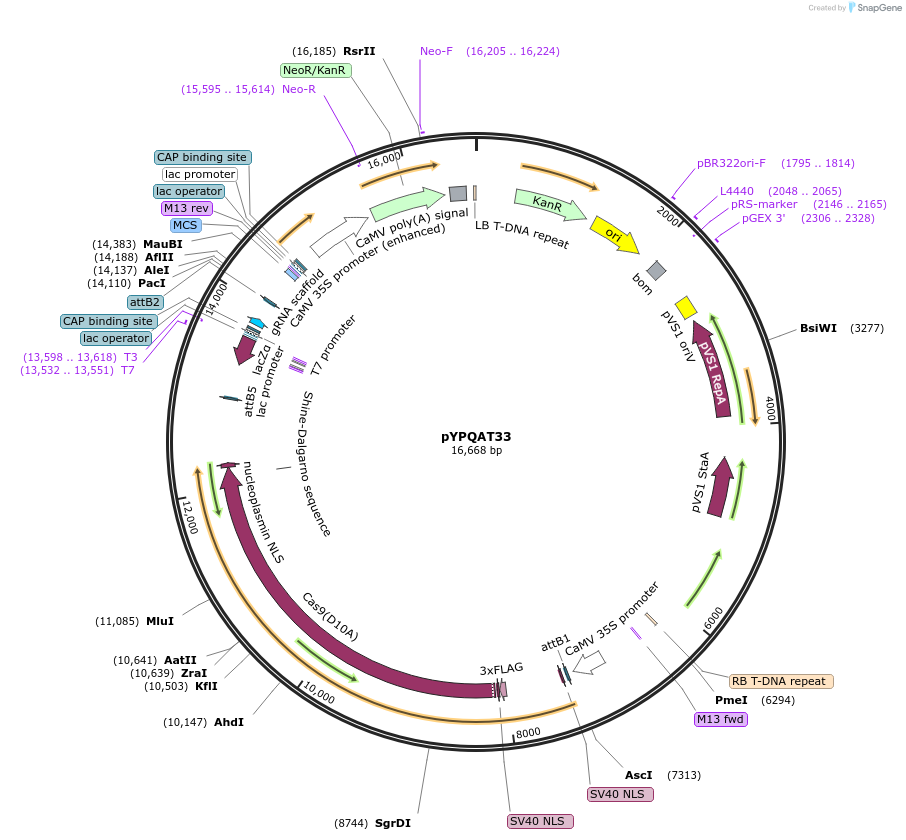
pYPQAT33
Plasmid#223405PurposeT-DNA vector for SpCas9-D10A based A-to-G base editing for dicot plants; NGG PAM; SpCas9D10A-ABE was driven by 2x35s and the sgRNA was driven by AtU3 promoter; Kanamycin for plants selection.DepositorInsert2x35s-ecTadA8e-SpCas9-D10A-AtU3-gRNA scaffold
UseCRISPRTags3x FLAG and NLSExpressionPlantAvailable SinceApril 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
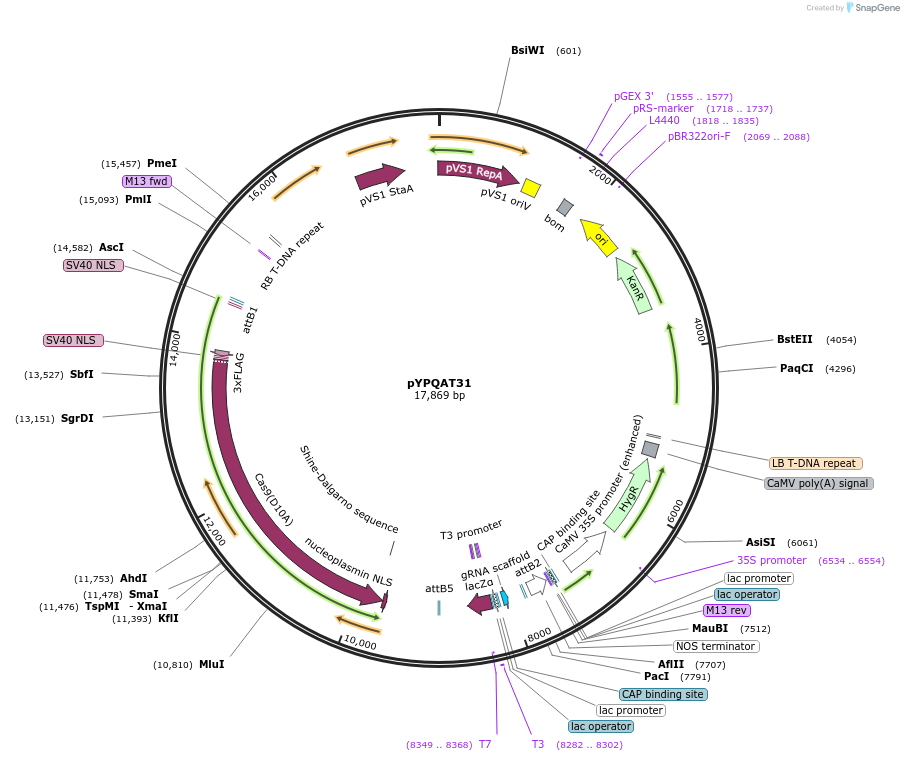
pYPQAT31
Plasmid#223403PurposeT-DNA vector for SpCas9-D10A based A-to-G base editing for dicot plants; NGG PAM; SpCas9D10A-ABE was driven by AtUBQ10 and the sgRNA was driven by AtU3 promoter; Hygromycin for plants selection.DepositorInsertAtUBQ10-ecTadA8e-SpCas9-D10A-AtU3-gRNA scaffold
UseCRISPRTags3x FLAG and NLSExpressionPlantAvailable SinceApril 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
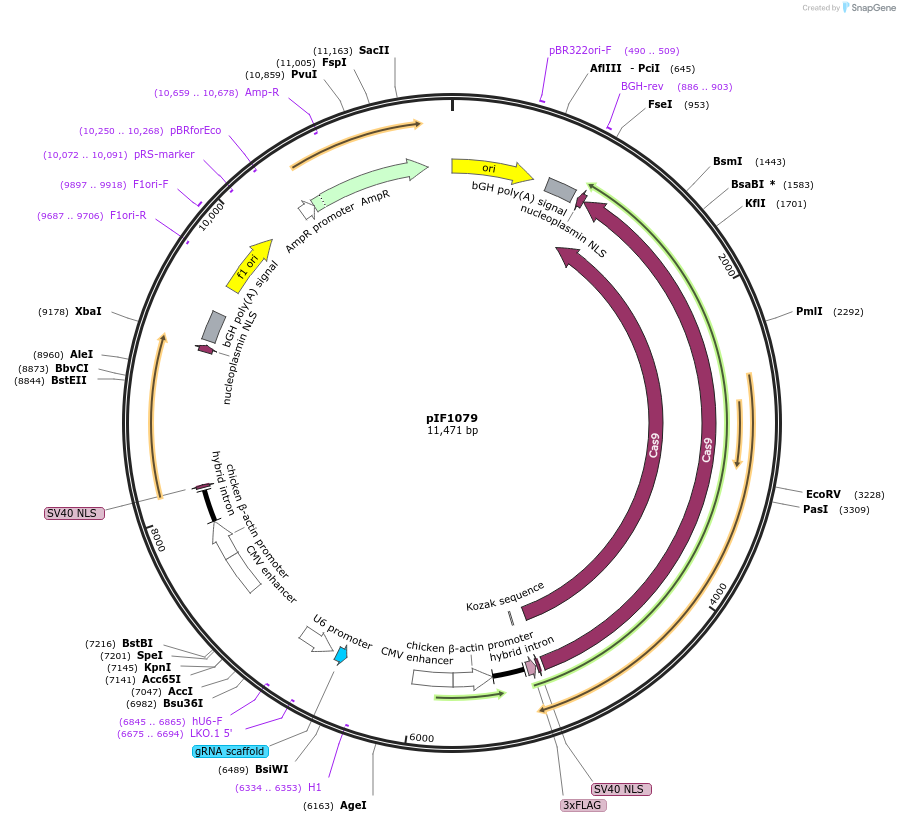
pIF1079
Plasmid#237445PurposeExpression SpCas9 and Efe1-RT from bidirectional CAG promoters. SpCas9 sgRNA and Efe1 non-coding expression is driven by U6 and H1 respecitvely.DepositorInsertsSpCas9
Efe1-RT
EMX1 sgRNA
Efe1-ncRNA targeting EMX1
Tags3xFLAGExpressionMammalianPromoterCAG, H1, and U6Available SinceJune 24, 2025AvailabilityAcademic Institutions and Nonprofits only -
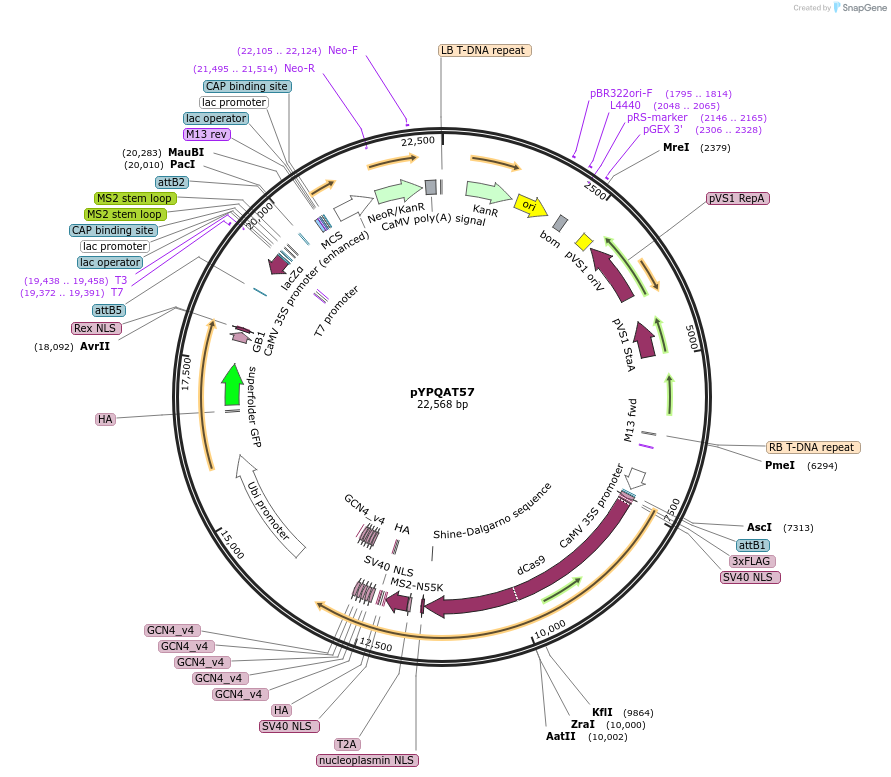
pYPQAT57
Plasmid#223429PurposeT-DNA vector for dSpCas9 mediated gene activation for dicot plants; NGG PAM; dSpCas9-Act3.0 was driven by 2x35s and the sgRNA was driven by AtU3 promoter; Kanamycin for plants selection.DepositorInsert2x35s-dSpCas9-Act3.0-AtU3-gRNA scaffold 2.0
UseCRISPRTags3x FLAG and NLSExpressionPlantAvailable SinceApril 10, 2025AvailabilityAcademic Institutions and Nonprofits only