We narrowed to 23,722 results for: ESC;
-
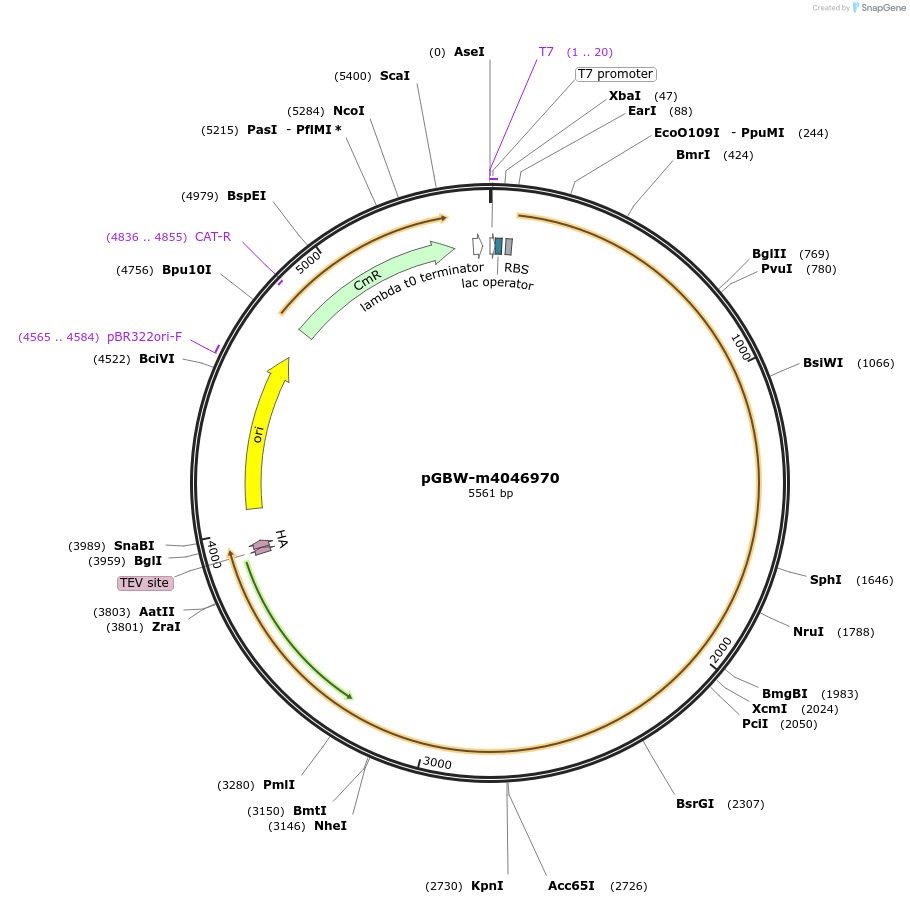
Plasmid#145753PurposeBacterial Expression plasmid for SARS-CoV-2 surface glycoprotein (Spike protein)DepositorInsertSARS-CoV-2 S (surface glycoprotein) (S Escherichia coli str. K-12 substr. MG1655; Severe acute respiratory syndrome coronavirus 2)
TagsCleavable TEV;HAExpressionBacterialMutationEscherichia coli recode 1PromoterPromoter | pT7 ; Operator | lacO ; RBS | T7_rbs ;…Available SinceApril 29, 2020AvailabilityIndustry, Academic Institutions, and Nonprofits -
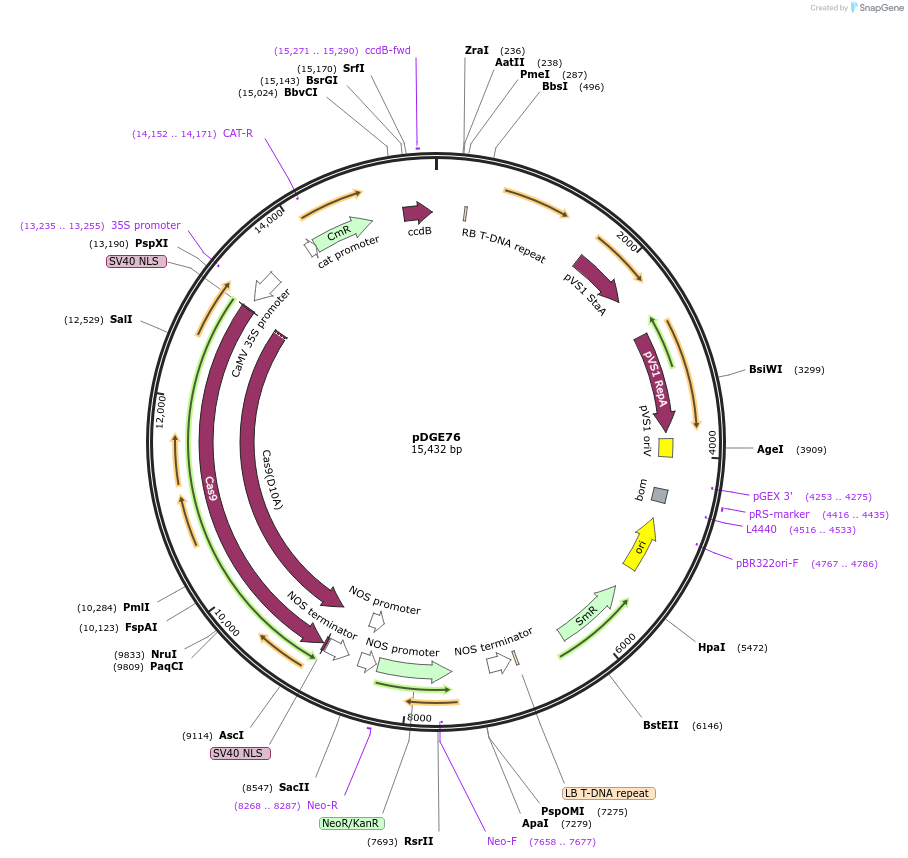
pDGE76
Plasmid#80578Purposerecipient plasmid [multiplexing, nickase]; p35S:Cas9 D10A, nptIIDepositorInsertspnos:nptII-tmos
p2x35S:Cas9 D10A-tnos
BsaI-ccdB_CmR-BsaI
UseCRISPRExpressionPlantMutationD10AAvailable SinceSept. 9, 2016AvailabilityAcademic Institutions and Nonprofits only -
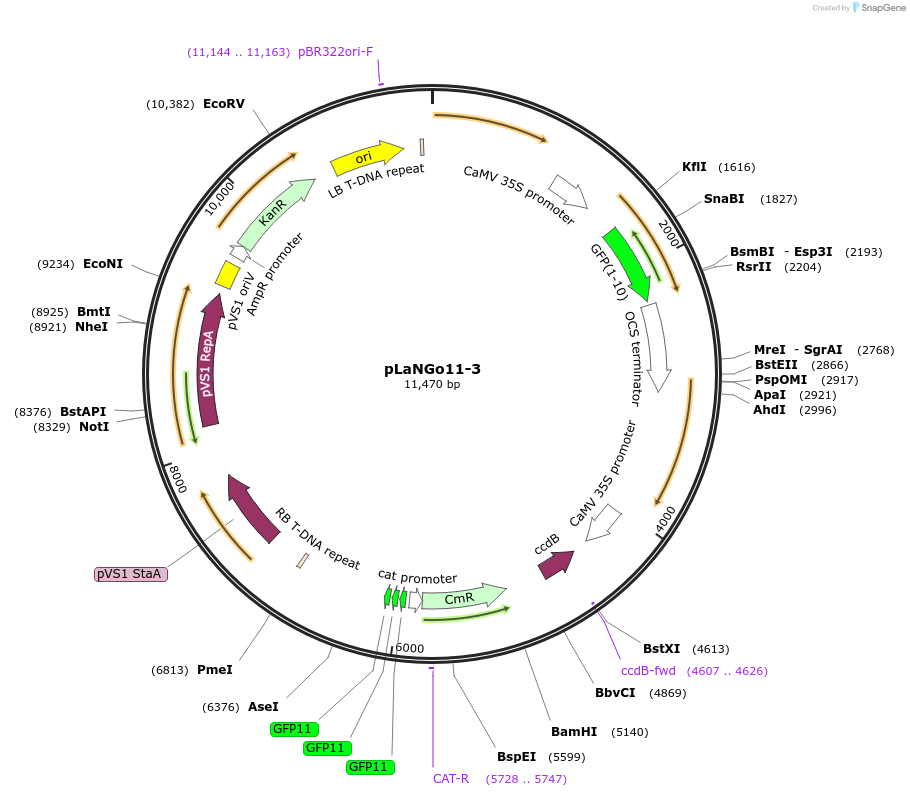
pLaNGo11-3
Plasmid#126054PurposeA self-assembling split-GFP based Golden Gate vector to determine protein targeting to plastids in plant cellDepositorInsertschloroplastic FNR transit peptide
ccdB cassette (BsaI)
GFP11x3 (5 aa linker)
UseBinaryTagsGFP1-10ExpressionPlantAvailable SinceAug. 26, 2019AvailabilityAcademic Institutions and Nonprofits only -
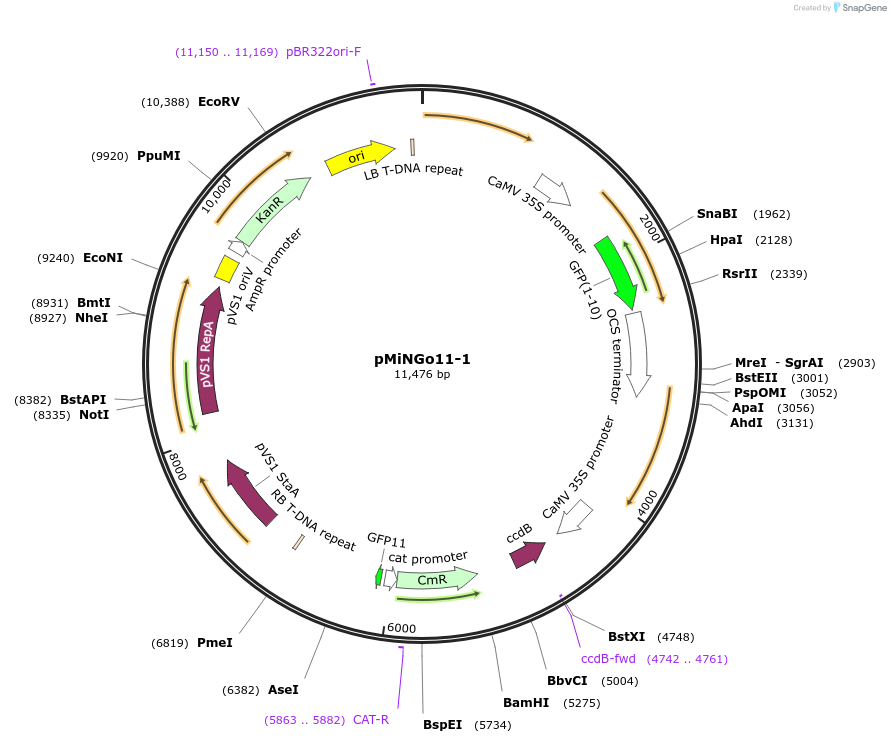
pMiNGo11-1
Plasmid#126056PurposeA self-assembling split-GFP based Golden Gate vector to determine protein targeting to mitochondria in plant cellDepositorInsertsmitochondria Rieske iron sulphur protein (1-100 aa)
ccdB cassette (BsaI)
GFP11
UseBinaryTagsGFP1-10ExpressionPlantAvailable SinceJune 27, 2019AvailabilityAcademic Institutions and Nonprofits only -
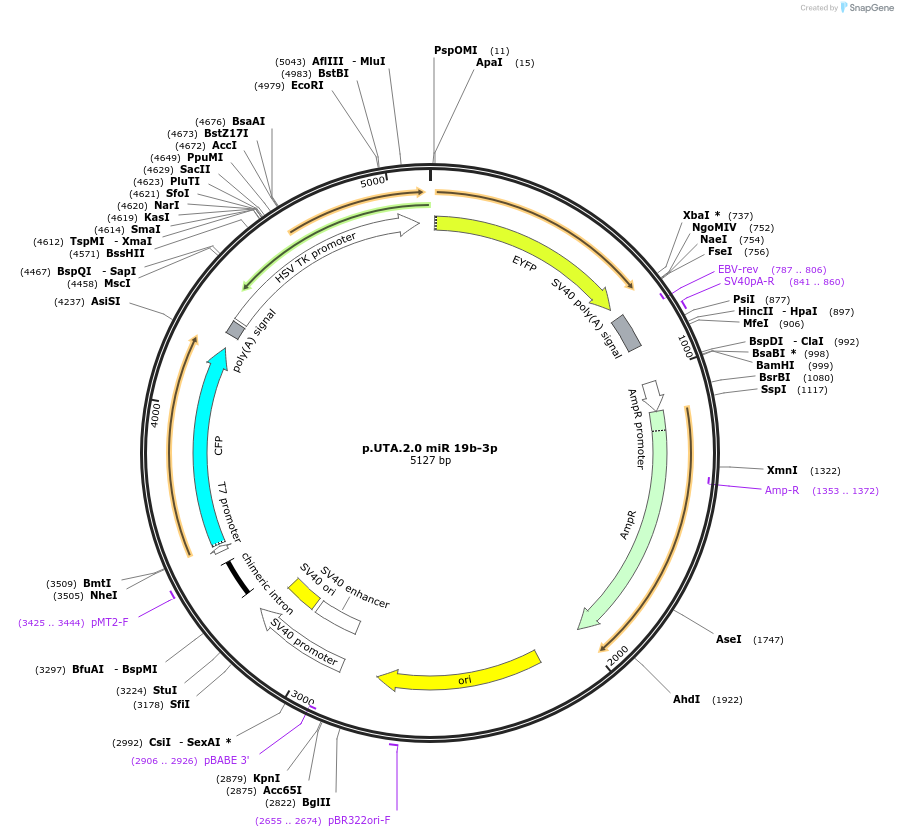
p.UTA.2.0 miR 19b-3p
Plasmid#82442PurposeDual Fluorescent endogenous miRNA sensor. Perfect complementary region of hsa-miR-19b-3pDepositorInserthsa-miR-19-3p
MutationComplementary region for hsa-miR-19-3p as sensorAvailable SinceMarch 24, 2017AvailabilityAcademic Institutions and Nonprofits only -
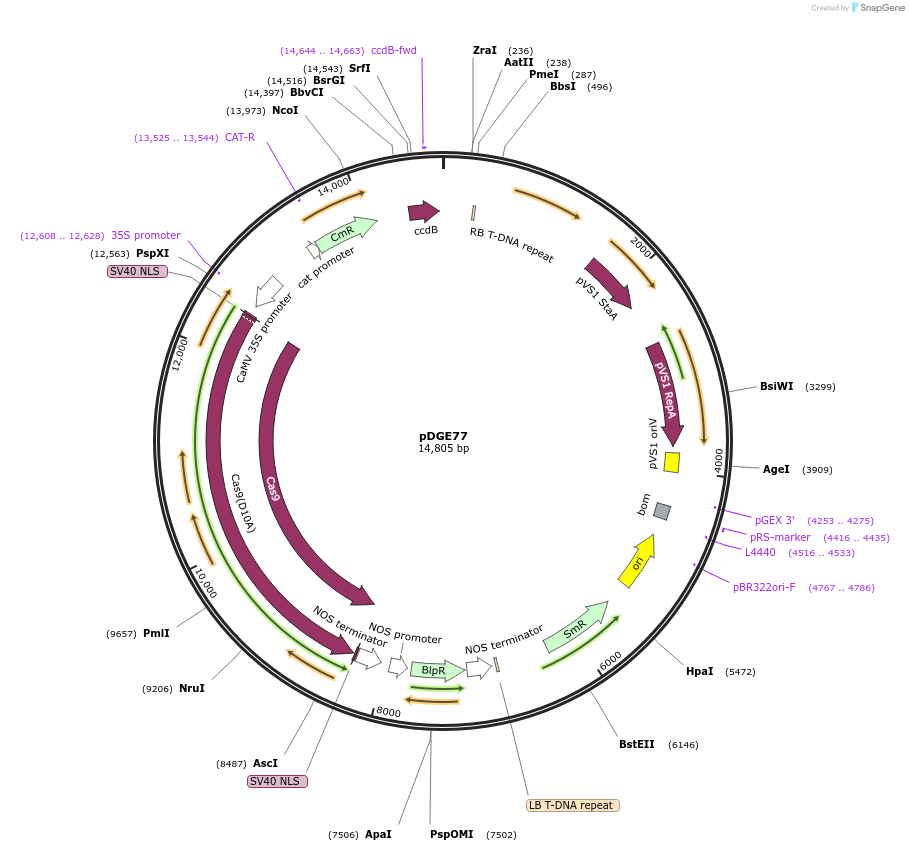
pDGE77
Plasmid#80579Purposerecipient plasmid [multiplexing, nickase]; p35S:Cas9 D10A, BASTADepositorInsertspnos:PAT-tnos
p2x35S:Cas9 D10A-tnos
BsaI-ccdB_CmR-BsaI
UseCRISPRExpressionPlantMutationD10AAvailable SinceSept. 9, 2016AvailabilityAcademic Institutions and Nonprofits only -
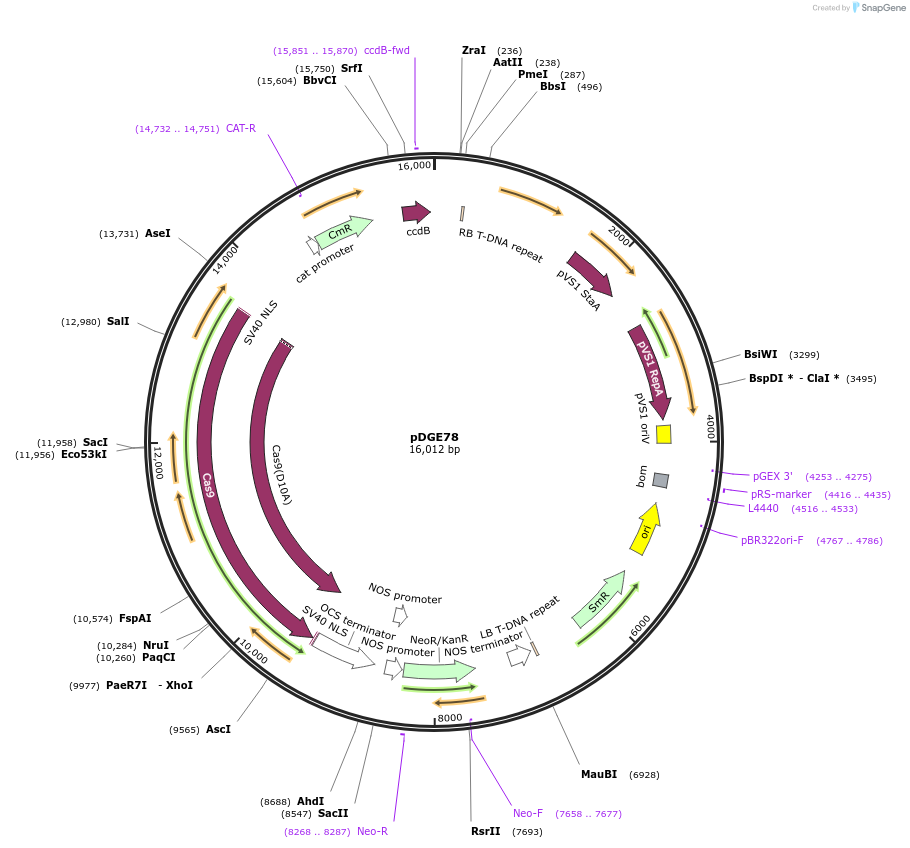
pDGE78
Plasmid#80580Purposerecipient plasmid [multiplexing, nickase]; pPcUbi:Cas9 D10A, nptIIDepositorInsertspnos:nptII-tmos
pPcUbi:Cas9 D10A-tnos
BsaI-ccdB_CmR-BsaI
UseCRISPRExpressionPlantMutationD10AAvailable SinceSept. 9, 2016AvailabilityAcademic Institutions and Nonprofits only -
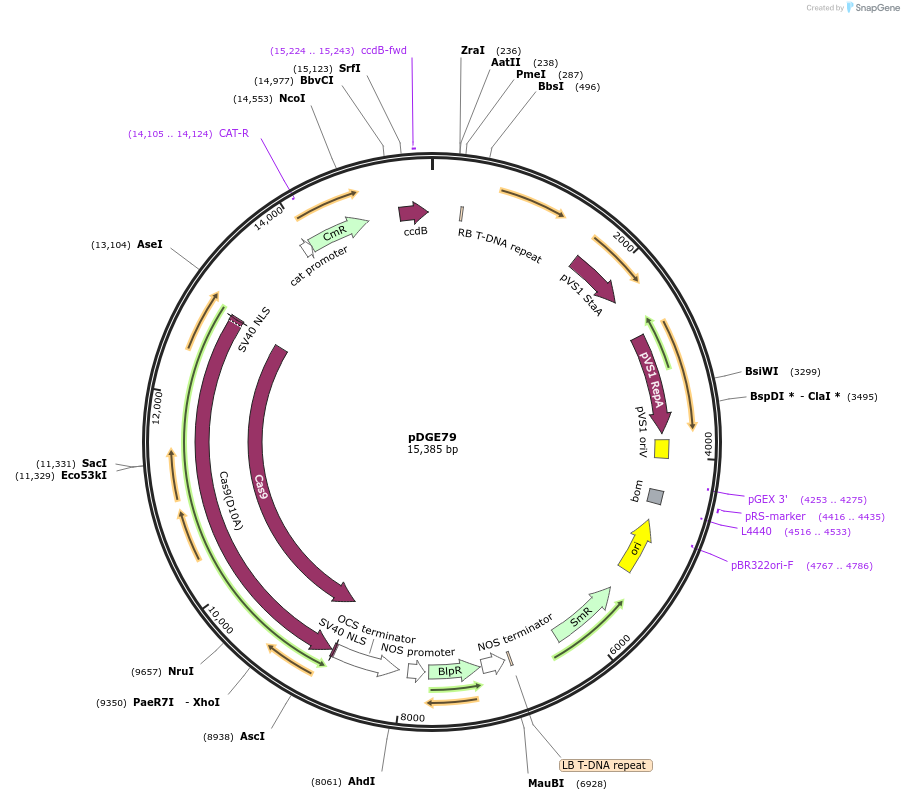
pDGE79
Plasmid#80581Purposerecipient plasmid [multiplexing, nickase]; pPcUbi:Cas9 D10A, BASTADepositorInsertspnos:PAT-tnos
pPcUbi:Cas9 D10A-tnos
BsaI-ccdB_CmR-BsaI
UseCRISPRExpressionPlantMutationD10AAvailable SinceSept. 9, 2016AvailabilityAcademic Institutions and Nonprofits only -
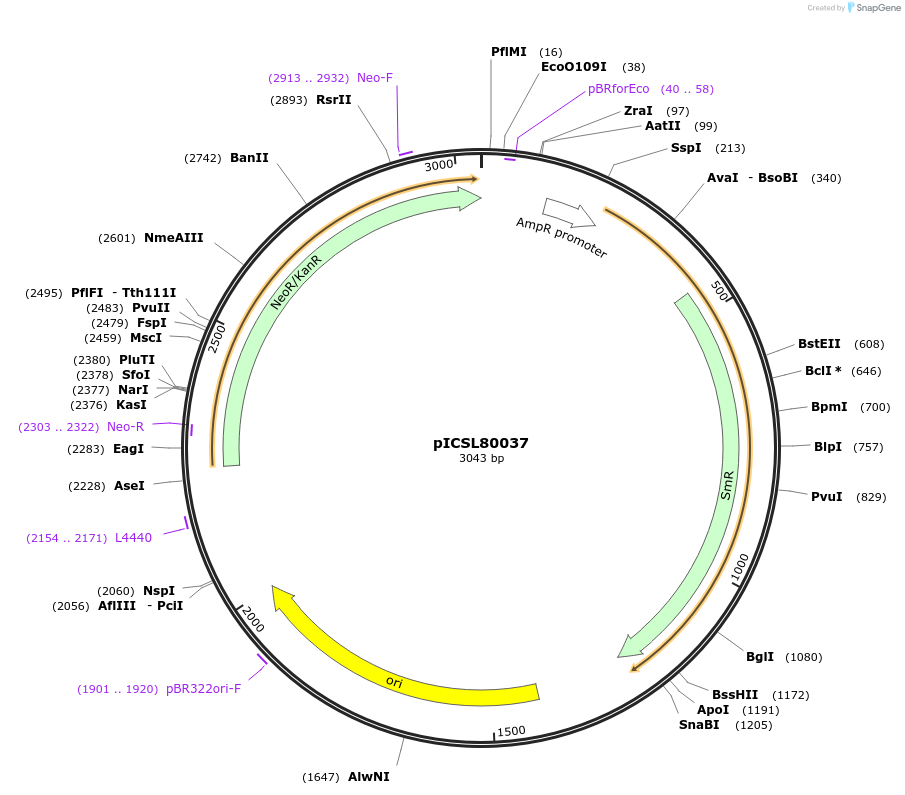
pICSL80037
Plasmid#68260PurposeLevel 0 Golden Gate Part: Coding sequence, neomycin phophotransferase II (Escherichia coli)DepositorInsertCoding sequence, neomycin phophotransferase II (Escherichia coli)
UseSynthetic BiologyAvailable SinceAug. 20, 2015AvailabilityAcademic Institutions and Nonprofits only -
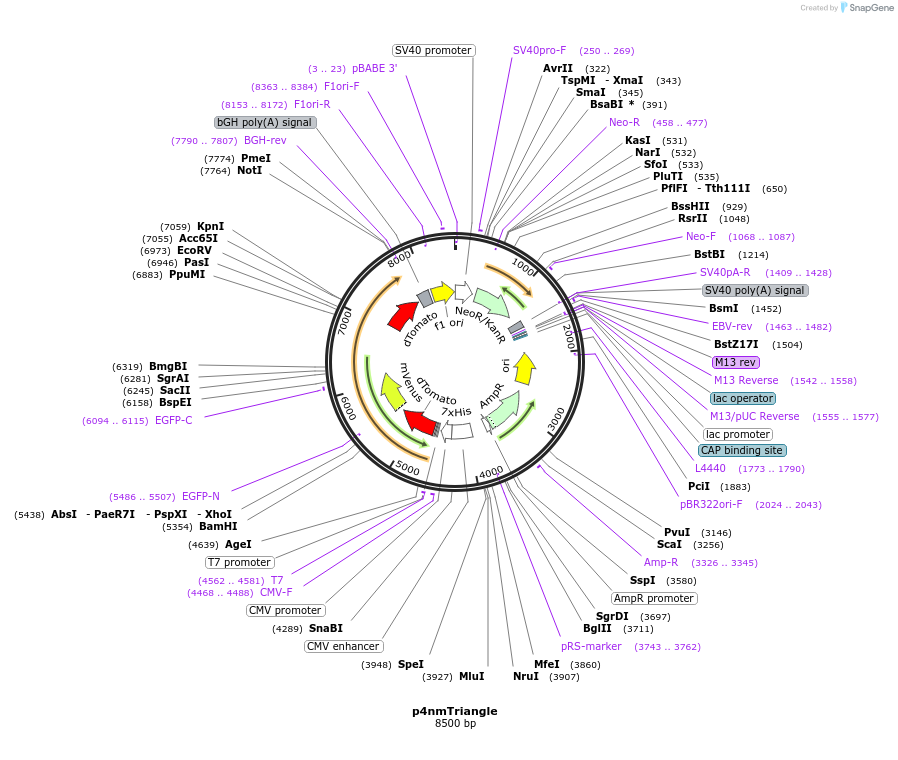
p4nmTriangle
Plasmid#250946PurposeExpresses a putative triangle with dTomato-mVenus-mVenusHP-dTomato in mammalian cells. Each fluorescent protein is separated by a 4nm ER/K helix.DepositorInsert4nm Triangle
Tags6HisExpressionMammalianPromoterCMVAvailable SinceApril 27, 2026AvailabilityAcademic Institutions and Nonprofits only -
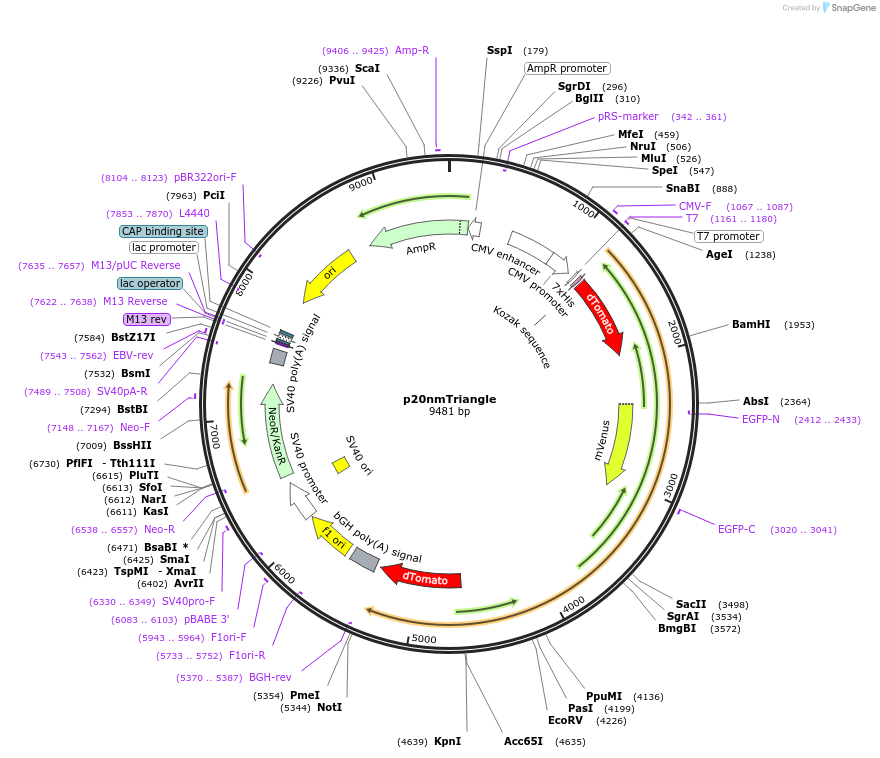
p20nmTriangle
Plasmid#250947PurposeExpresses a putative triangle with dTomato-mVenus-mVenusHP-dTomato in mammalian cells. Each fluorescent protein is separated by a 20nm ER/K helix.DepositorInsert20nm Triangle
Tags6HisExpressionMammalianPromoterCMVAvailable SinceApril 27, 2026AvailabilityAcademic Institutions and Nonprofits only -
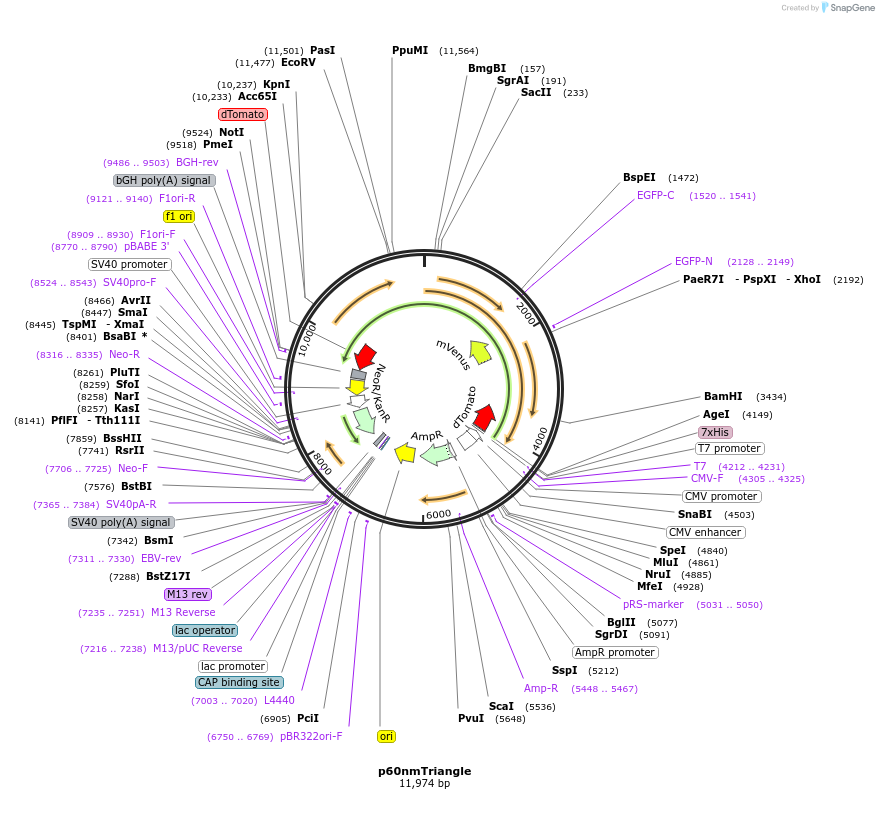
p60nmTriangle
Plasmid#250948PurposeExpresses a putative triangle with dTomato-mVenus-mVenusHP-dTomato in mammalian cells. Each fluorescent protein is separated by a 60nm ER/K helix.DepositorInsert60nm Triangle
Tags6HisExpressionMammalianPromoterCMVAvailable SinceApril 27, 2026AvailabilityAcademic Institutions and Nonprofits only -
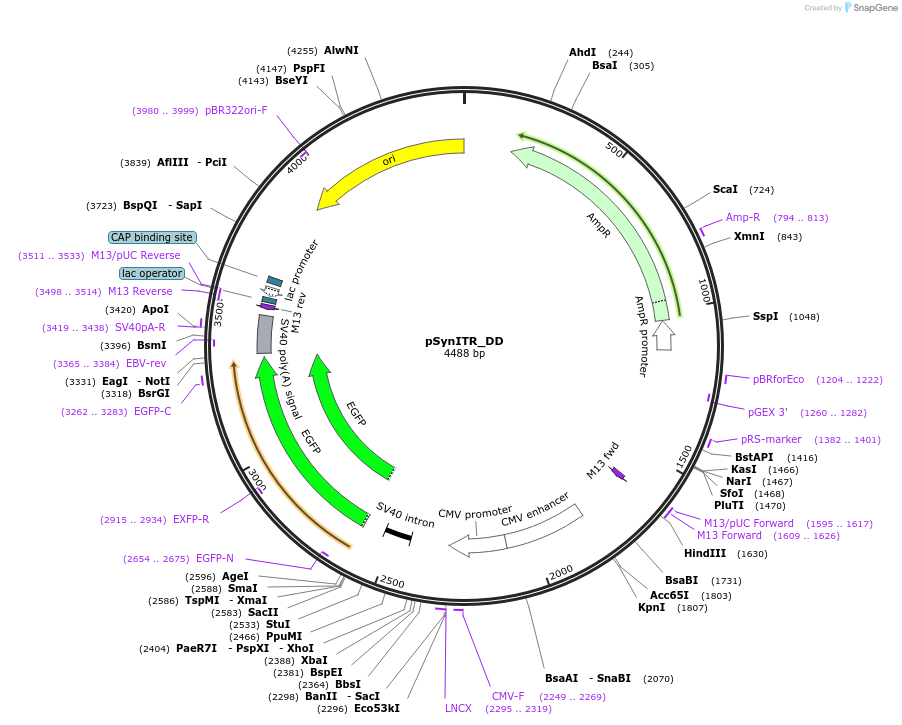
pSynITR_DD
Plasmid#253227PurposeThis is a DD-ITR AAV Production plasmid that contains the SynITR sequence as published in PMID: 39868534DepositorInsertsEnhanced Green Fluorescent Protein
SynITR
UseAAVExpressionMammalianAvailable SinceMarch 11, 2026AvailabilityAcademic Institutions and Nonprofits only -
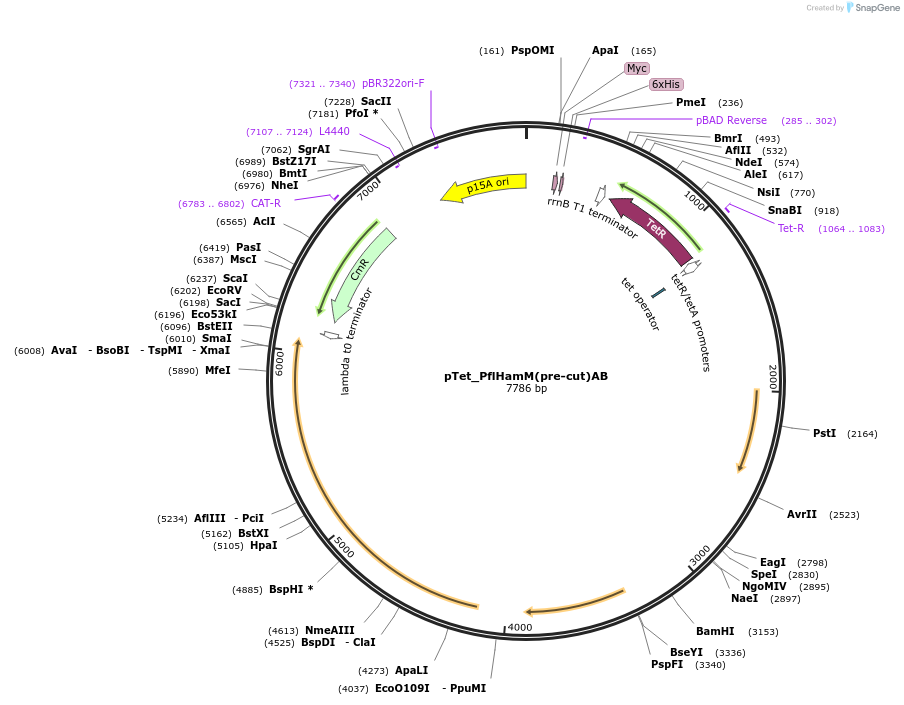
pTet_PflHamM(pre-cut)AB
Plasmid#248942PurposeHachiman from P. fluorescens under pTet control with HamM programmed breaks in the peptide backbone at I118 and I218DepositorInsertPflHamM(pre-cut)AB
ExpressionBacterialMutationHamM programmed breaks in the peptide backbone at…Available SinceFeb. 23, 2026AvailabilityAcademic Institutions and Nonprofits only -
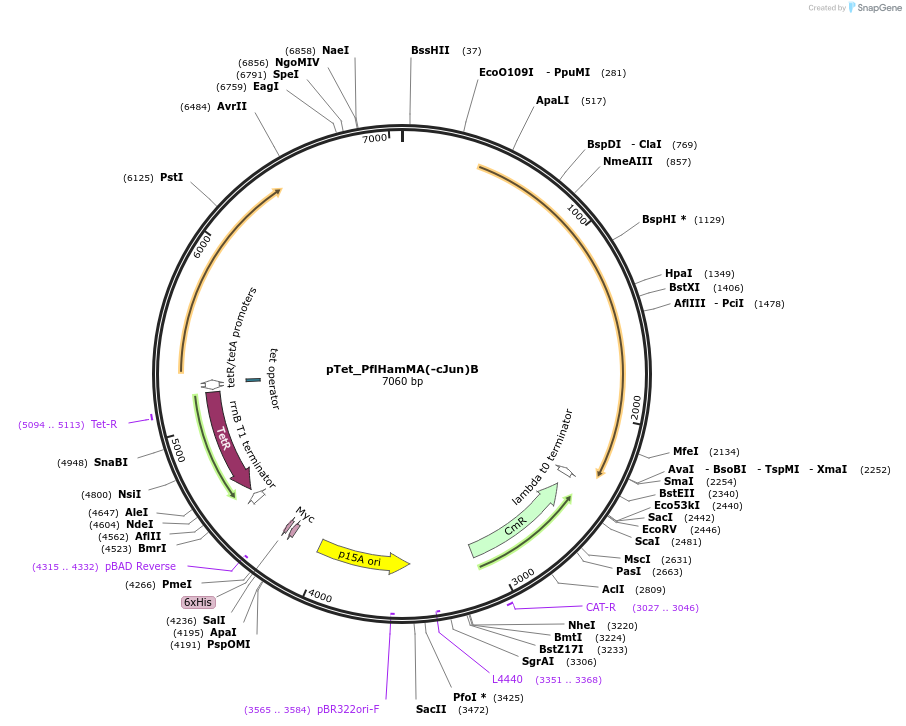
pTet_PflHamMA(-cJun)B
Plasmid#248940PurposeHachiman from P. fluorescens under pTet control with cJun fused to the protease domain and natural linkerDepositorInsertflHamMA(-cJun)B
ExpressionBacterialMutationc-Jun insertion, HamA domain deletionAvailable SinceFeb. 23, 2026AvailabilityAcademic Institutions and Nonprofits only -
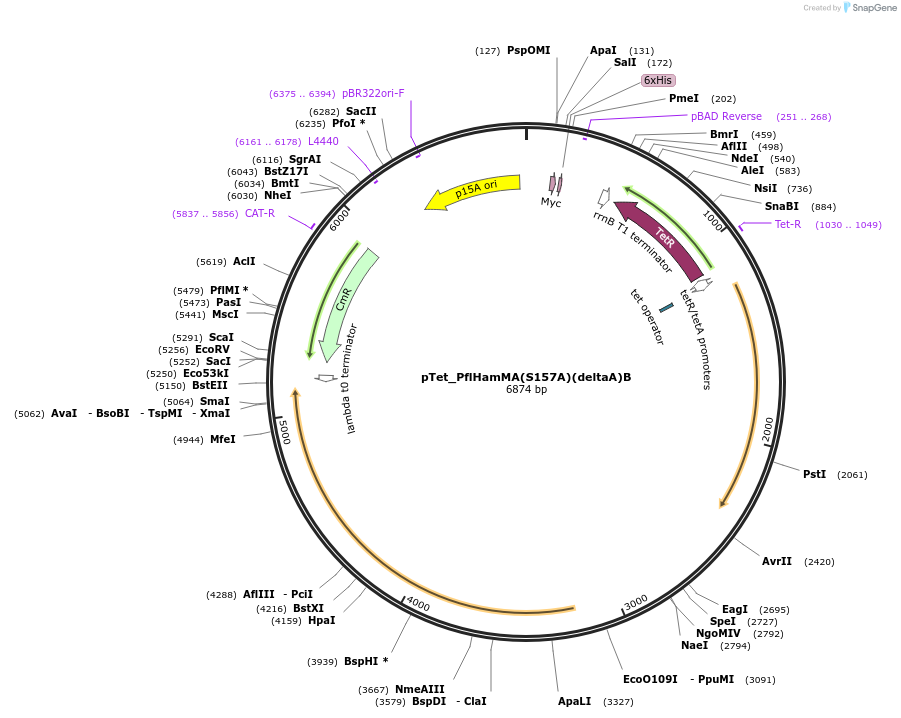
pTet_PflHamMA(S157A)(deltaA)B
Plasmid#248939PurposeHachiman from P. fluorescens under pTet control with the dHamA domain removed and with a protease S157A mutationDepositorInsertPflHamMA(S157A)(deltaA)B
ExpressionBacterialMutationS157A, HamA domain deletionAvailable SinceFeb. 23, 2026AvailabilityAcademic Institutions and Nonprofits only -
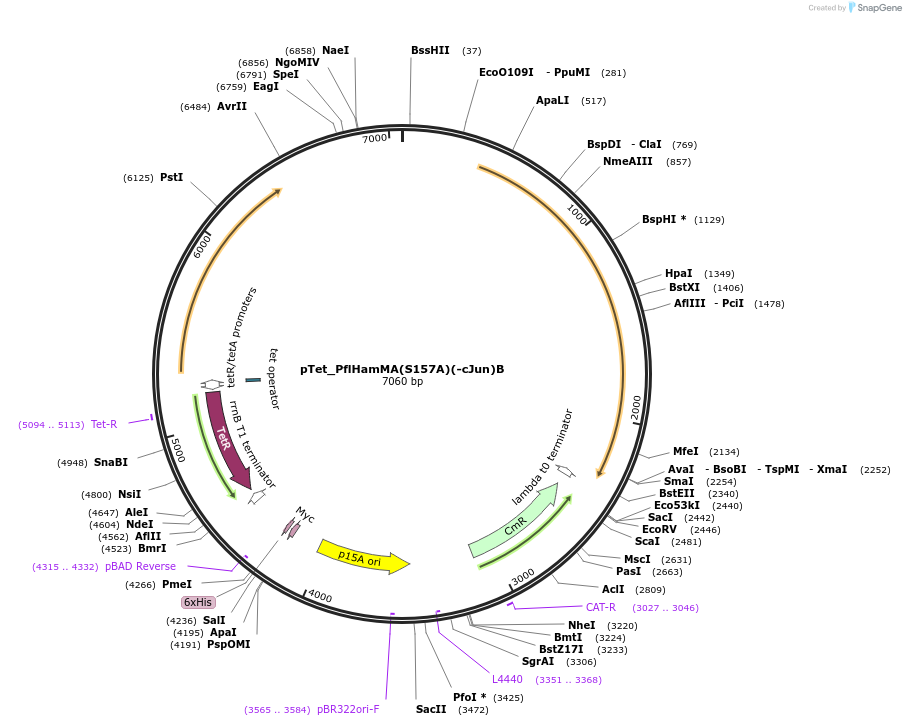
pTet_PflHamMA(S157A)(-cJun)B
Plasmid#248941PurposeHachiman from P. fluorescens under pTet control with cJun fused to the protease domain and natural linker and the protease mutated (S157A)DepositorInsertPflHamMA(S157A)(-cJun)B
ExpressionBacterialMutationS157A, c-Jun insertion, HamA domain deletionAvailable SinceFeb. 19, 2026AvailabilityAcademic Institutions and Nonprofits only -
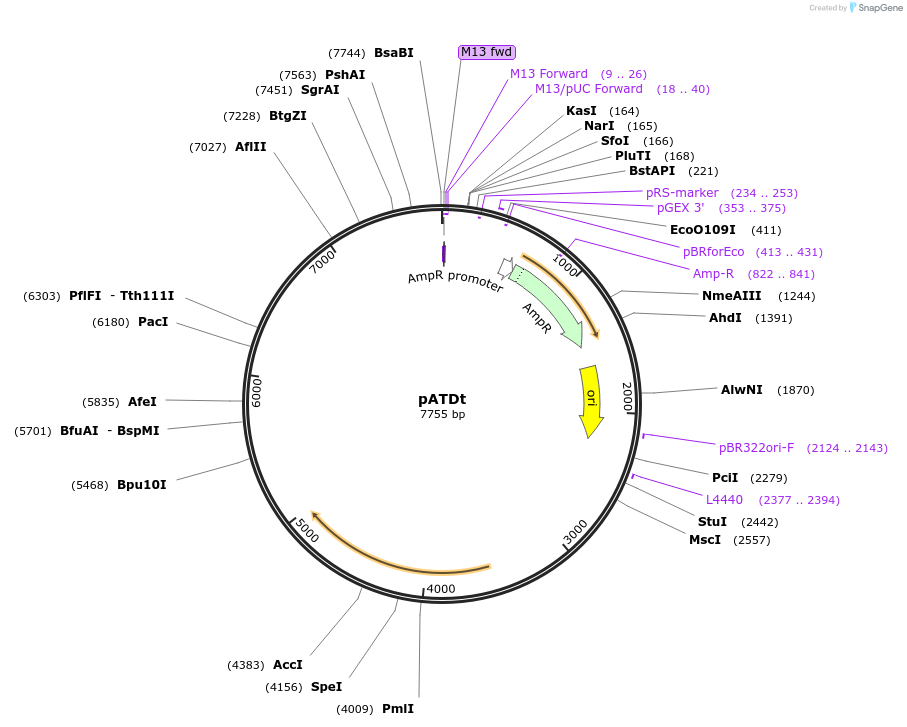
pATDt
Plasmid#249499PurposeExpresses tdTomato – Monodirectional chloroplast expression vector (psbH Rescue) in Chlamydomonas Reinhardtii photosynthetically deficient strain CC4388DepositorInserttdTomato (red fluorescence)
UseChlamydomonas reinhardtii chloroplast genomeExpressionBacterialMutationtdTomato results from the fusion of two copies of…PromoterThe psbD/ 5′ UTR, a light-activated chloroplast p…Available SinceJan. 21, 2026AvailabilityAcademic Institutions and Nonprofits only -
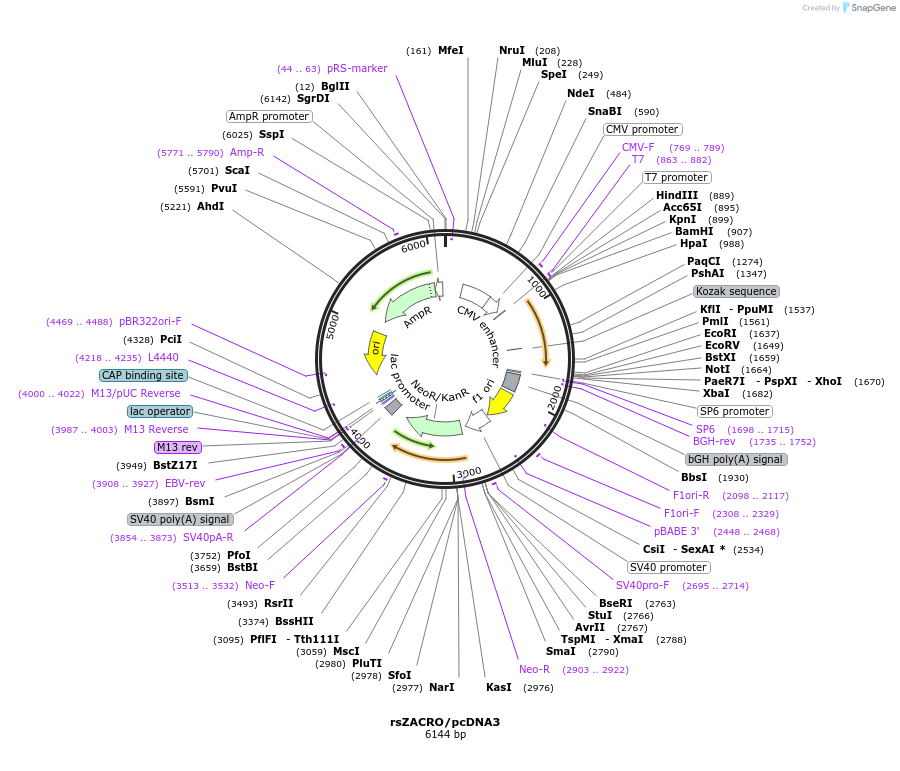
rsZACRO/pcDNA3
Plasmid#236199PurposeMammalian expression of rsZACRO, a positively reversibly photoswitchable fluorescent proteinDepositorInsertrsZACRO
ExpressionMammalianAvailable SinceMay 30, 2025AvailabilityAcademic Institutions and Nonprofits only -
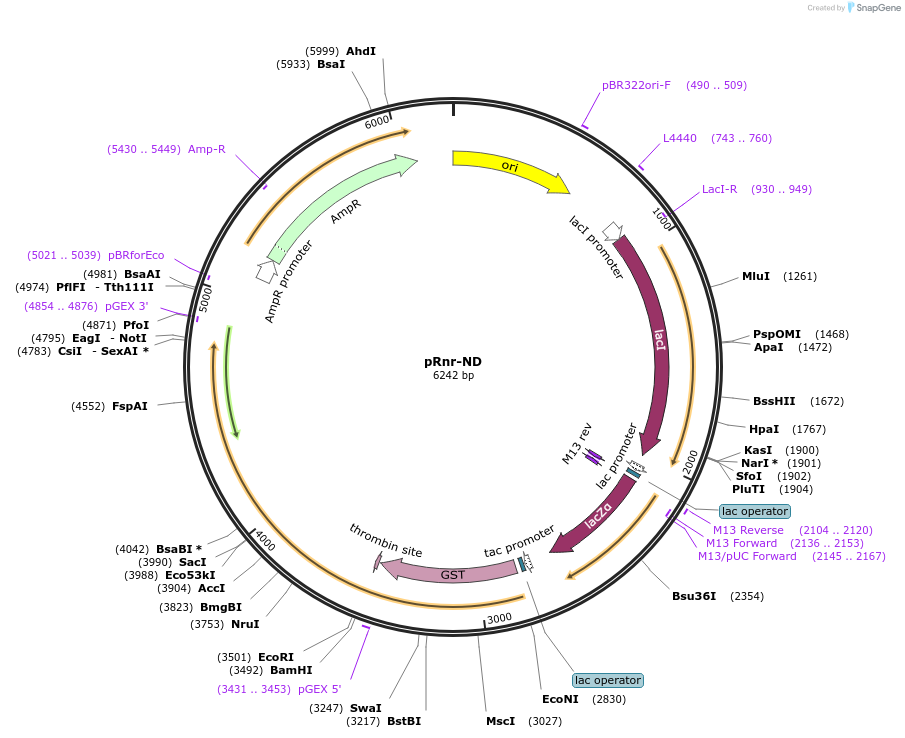
pRnr-ND
Plasmid#232579PurposeExpression of the Rnr nuclease domain (residues 649 to 1929) as a GST-fusion.DepositorInsertRnr nuclease domain (residues 649 to 1929)
TagsGSTExpressionBacterialAvailable SinceMay 12, 2025AvailabilityIndustry, Academic Institutions, and Nonprofits