We narrowed to 8,782 results for: set
-
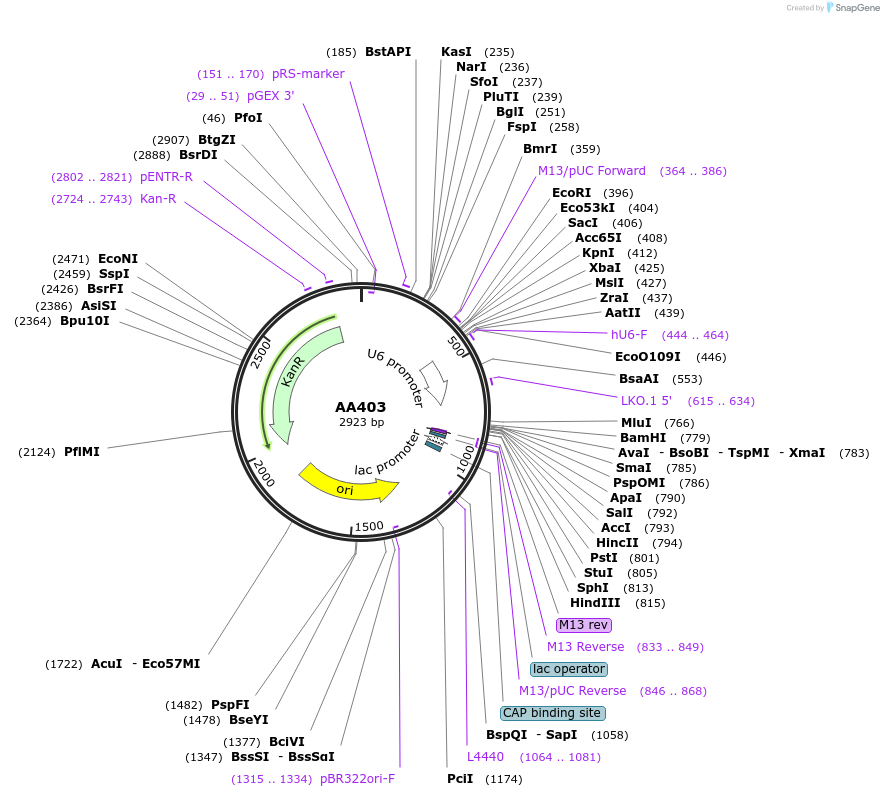
Plasmid#215960PurposeFragmid fragment: (guide cassette) guide expressionDepositorHas ServiceCloning Grade DNAInsertU6_v1; DR_v0 [Cas7-11]; BsmBI_v16; BsmBI_v12
UseCRISPR; FragmentAvailable SinceApril 29, 2024AvailabilityAcademic Institutions and Nonprofits only -
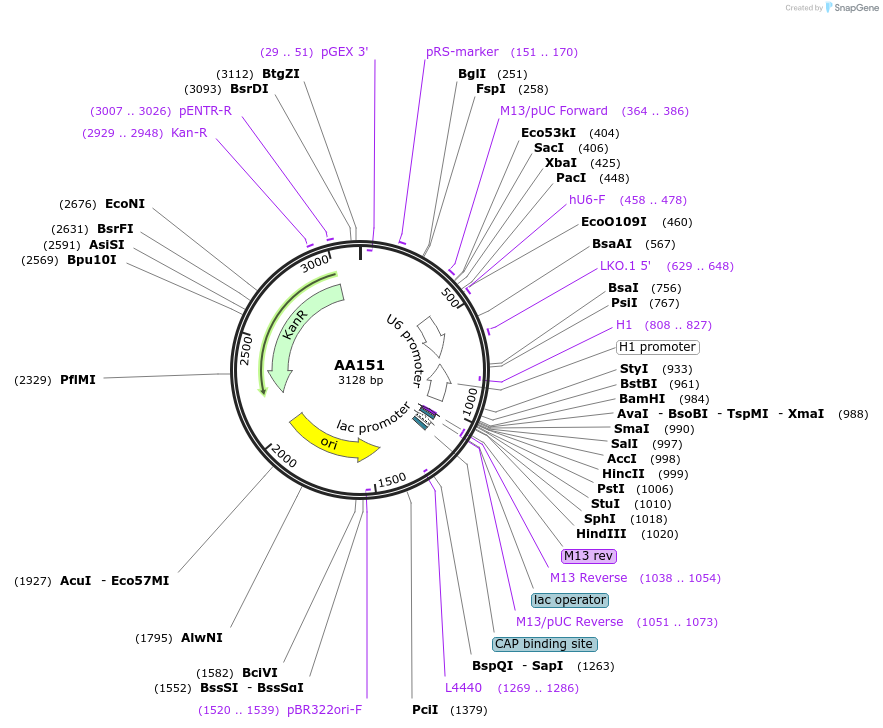
AA151
Plasmid#215938PurposeFragmid fragment: (guide cassette) for dual expression sgRNADepositorHas ServiceCloning Grade DNAInsertU6_v1; BsmBI_v0; BsmBI_v4; rev{H1_v1}
UseCRISPR; FragmentAvailable SinceApril 29, 2024AvailabilityAcademic Institutions and Nonprofits only -
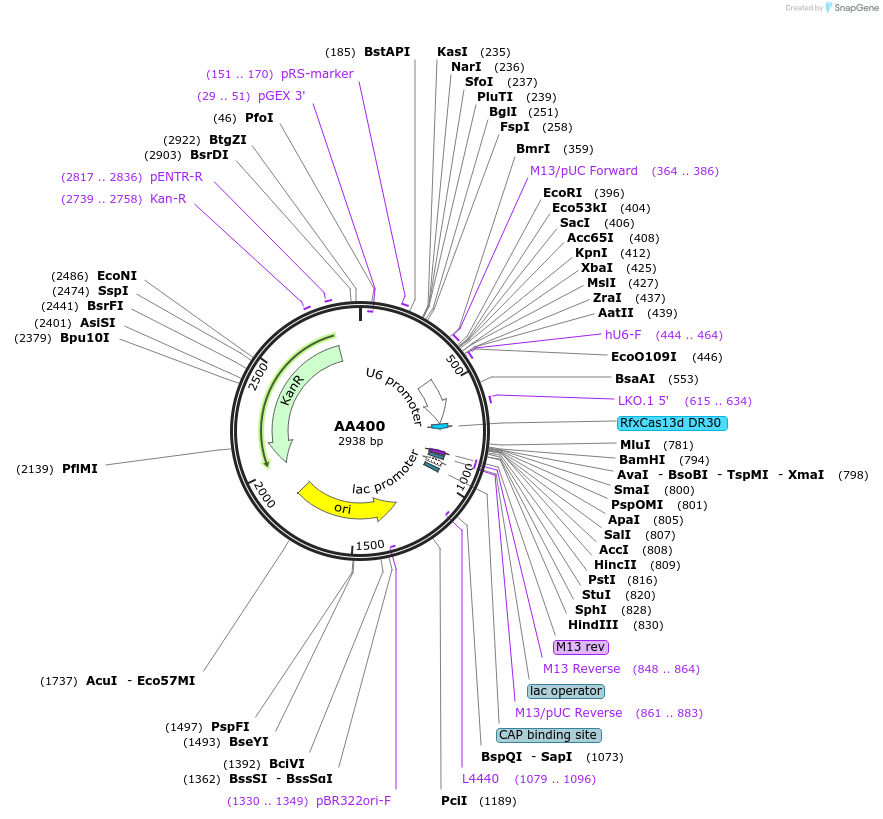
AA400
Plasmid#215958PurposeFragmid fragment: (guide cassette) guide expressionDepositorHas ServiceCloning Grade DNAInsertU6_v1; DR_v1 [Rfx]; BsmBI_v1; BsmBI_v12
UseCRISPR; FragmentAvailable SinceApril 29, 2024AvailabilityAcademic Institutions and Nonprofits only -
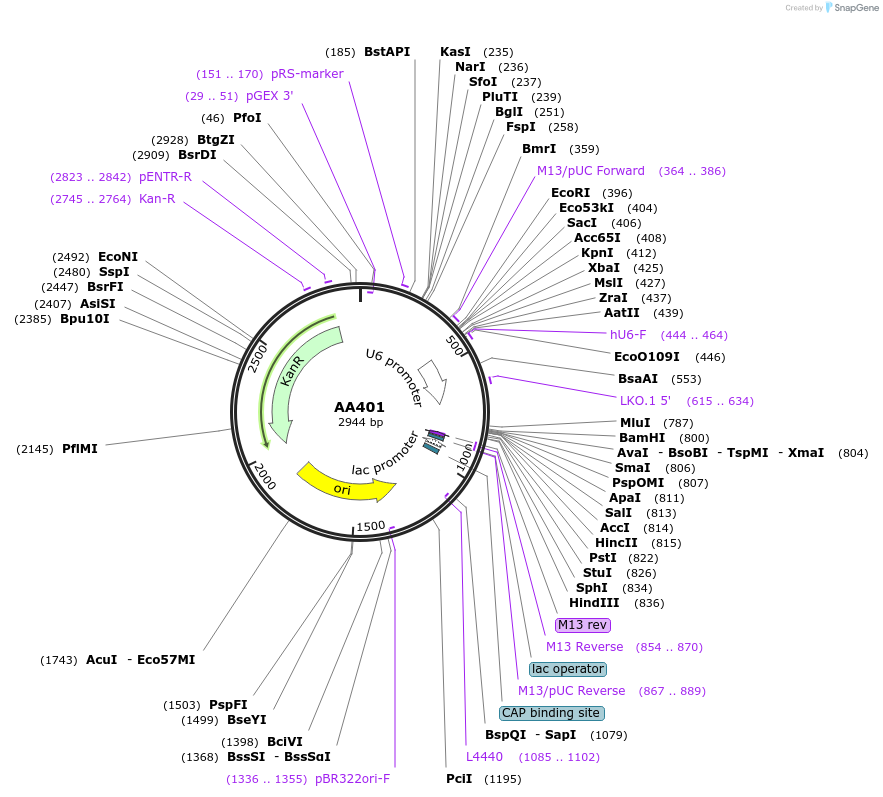
AA401
Plasmid#215959PurposeFragmid fragment: (guide cassette) guide expressionDepositorHas ServiceCloning Grade DNAInsertU6_v1; DR_v1 [Dj]; BsmBI_v1; BsmBI_v12
UseCRISPR; FragmentAvailable SinceApril 29, 2024AvailabilityAcademic Institutions and Nonprofits only -
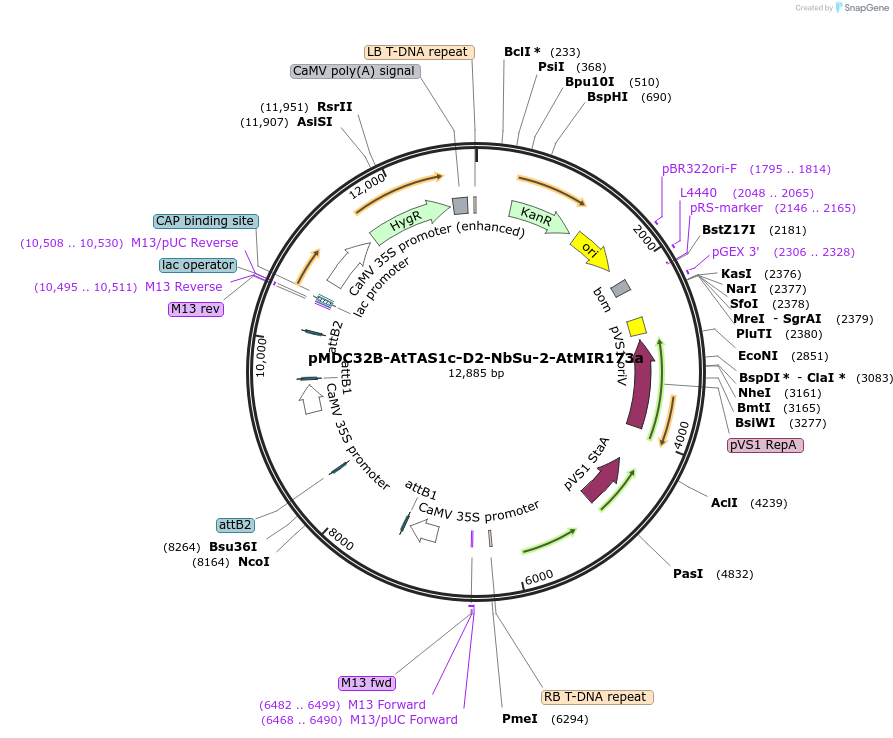
pMDC32B-AtTAS1c-D2-NbSu-2-AtMIR173a
Plasmid#213401PurposePlant expression vector (2x35S) for expressing a syn-tasiRNA against Nicotiana benthamiana SULFUR gene from AtTAS1c precursorDepositorInsertArabidopsis TAS1c with a syn-tasiRNA sequence at D2 for silencing N. benthamiana SULFUR gene. MIR173 cassette.
ExpressionPlantPromoter2x35SAvailable SinceFeb. 9, 2024AvailabilityAcademic Institutions and Nonprofits only -
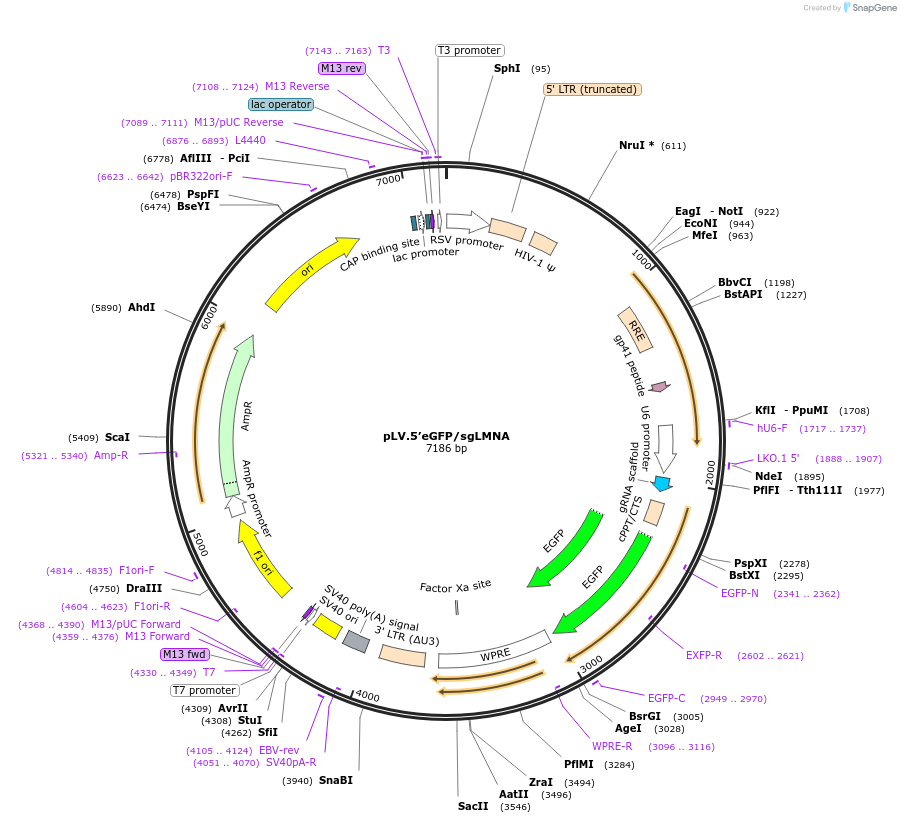
pLV.5’eGFP/sgLMNA
Plasmid#170546PurposeExpresses a sgRNA for 5' tagging to LMNA and contains a cassette with eGFP without homology armsDepositorInsertsgRNA targeting 5'- end of LMNA
UseCRISPR and LentiviralAvailable SinceJan. 12, 2024AvailabilityAcademic Institutions and Nonprofits only -
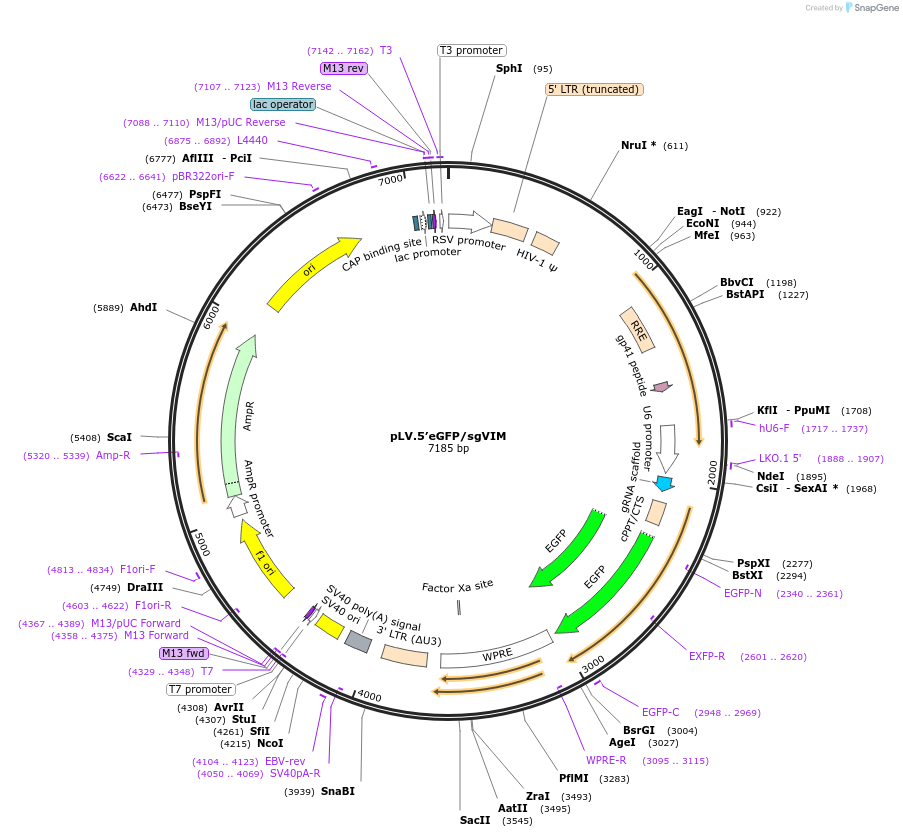
pLV.5’eGFP/sgVIM
Plasmid#170548PurposeExpresses a sgRNA for 5' tagging to VIM and contains a cassette with eGFP without homology armsDepositorInsertsgRNA targeting 5'- end of VIM
UseCRISPR and LentiviralPromoterU6Available SinceJan. 12, 2024AvailabilityAcademic Institutions and Nonprofits only -
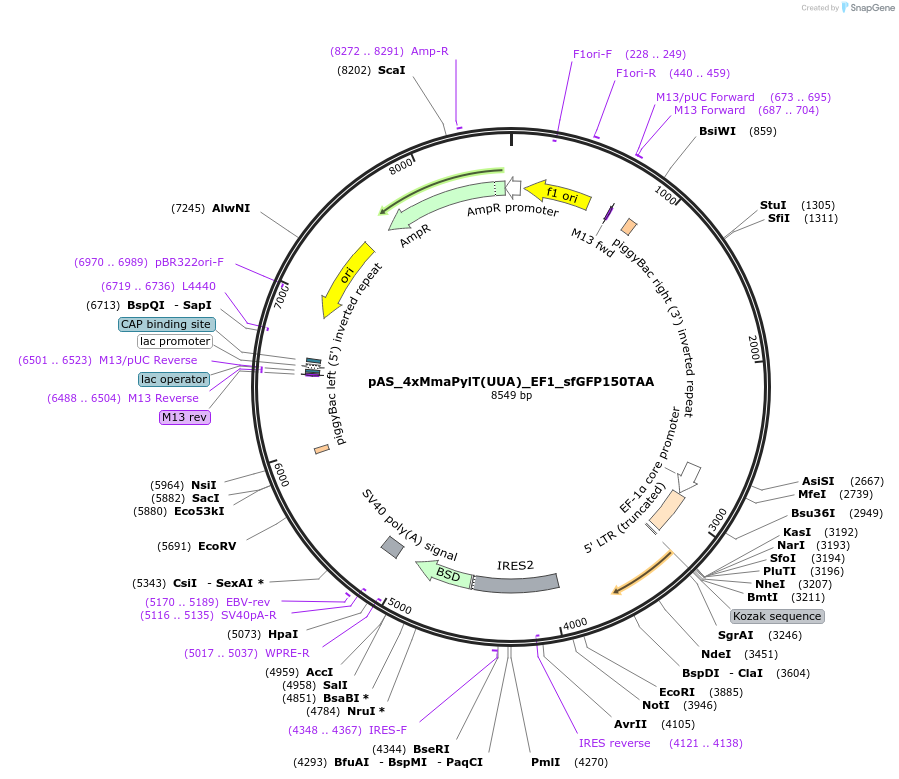
pAS_4xMmaPylT(UUA)_EF1_sfGFP150TAA
Plasmid#174897Purposeochre suppression reporter sfGFP150TAA expression, with MmaPylT(UUA) ochre suppressor tRNA cassetteDepositorInsertsf GFP
ExpressionMammalianMutation150TAA in sfGFPPromoterEF1Available SinceNov. 7, 2023AvailabilityAcademic Institutions and Nonprofits only -
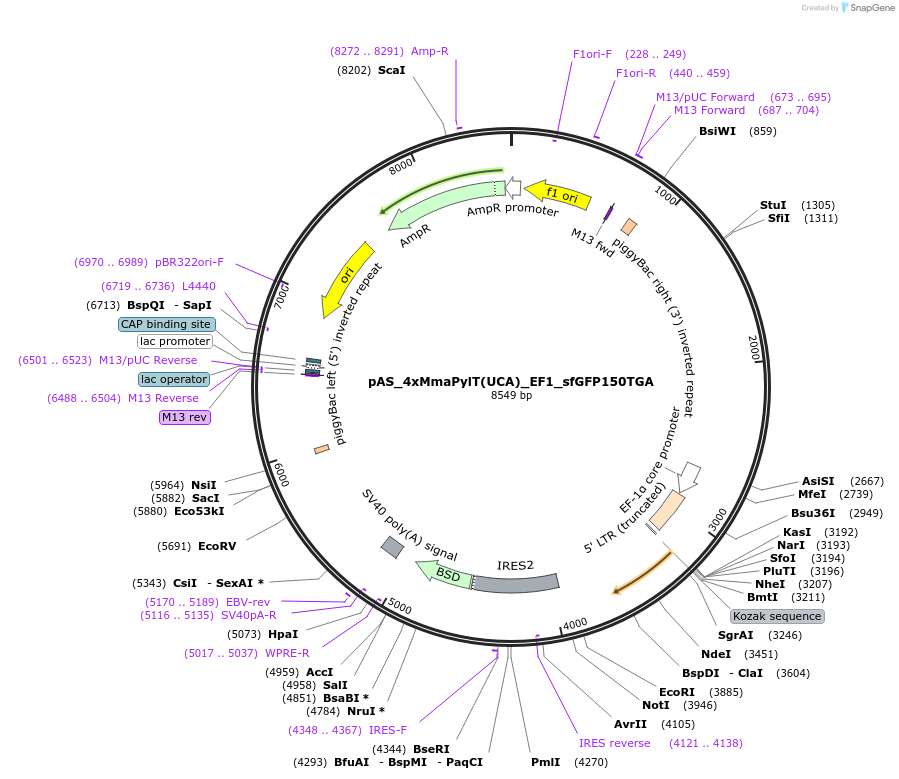
pAS_4xMmaPylT(UCA)_EF1_sfGFP150TGA
Plasmid#174898Purposeopal suppression reporter sfGFP150TGA expression, with MmaPylT(UCA) opal suppressor tRNA cassetteDepositorInsertsfGFP
ExpressionMammalianMutation150TGA in sfGFPPromoterEF1Available SinceNov. 7, 2023AvailabilityAcademic Institutions and Nonprofits only -
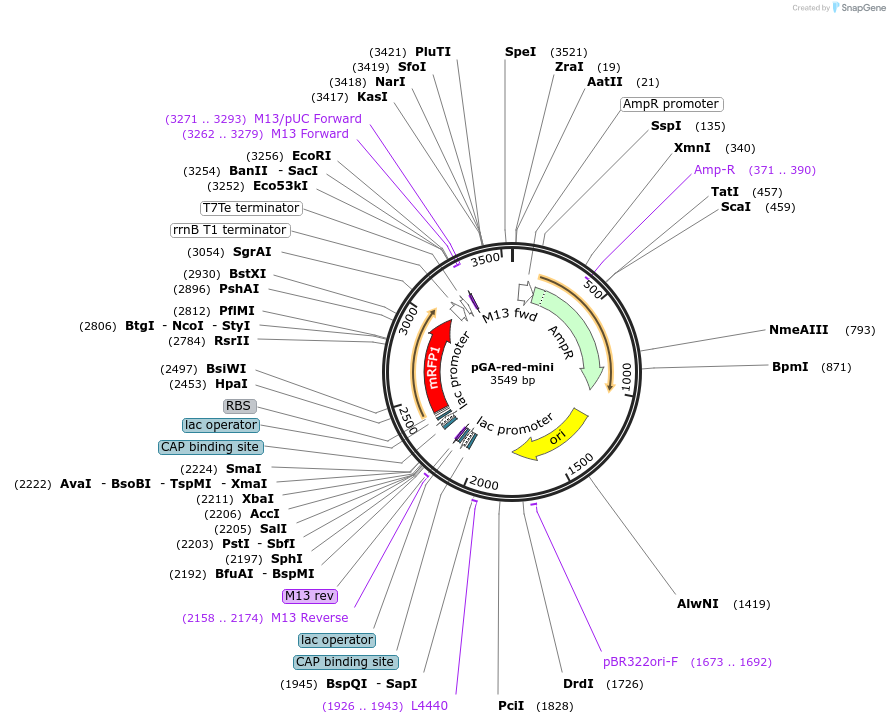
pGA-red-mini
Plasmid#196338PurposeEasy-MISE toolkit Level 1 destination plasmid for cassette cloning with BsaI-based Golden Gate Assembly; allowing fast postitive clones test with mRFP1 fluorescent protein-based red/white screeningDepositorTypeEmpty backboneUseSynthetic Biology; Golden gate assembly backboneAvailable SinceMay 15, 2023AvailabilityAcademic Institutions and Nonprofits only -
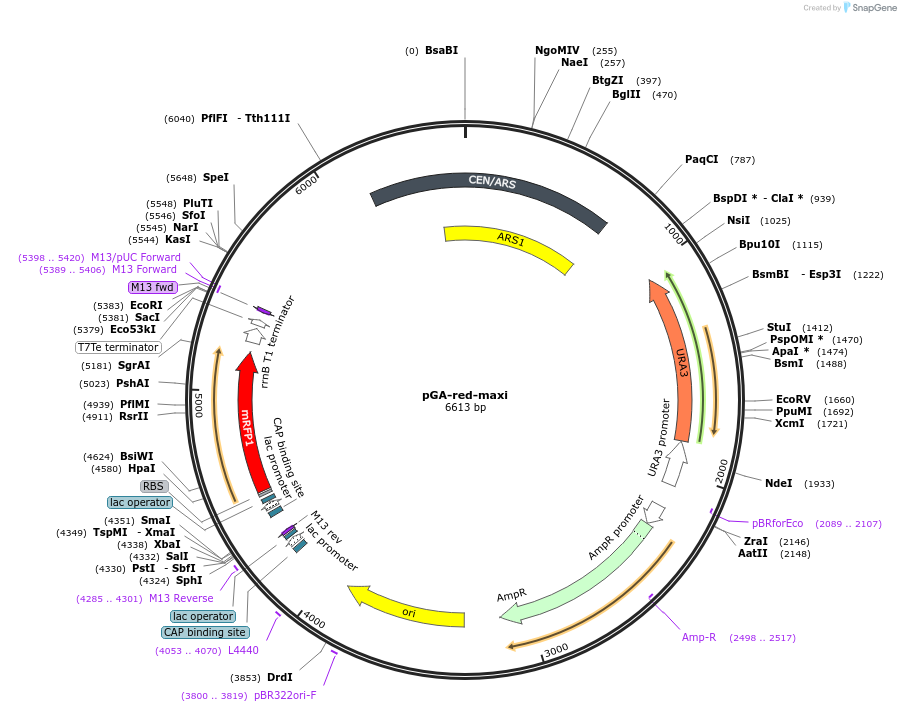
pGA-red-maxi
Plasmid#196337PurposeEasy-MISE toolkit Level 1 destination plasmid for cassette cloning with BsaI-based Golden Gate Assembly; allowing fast postitive clones test with mRFP1 fluorescent protein-based red/white screeningDepositorTypeEmpty backboneUseSynthetic Biology; Golden gate assembly backboneAvailable SinceMay 15, 2023AvailabilityAcademic Institutions and Nonprofits only -
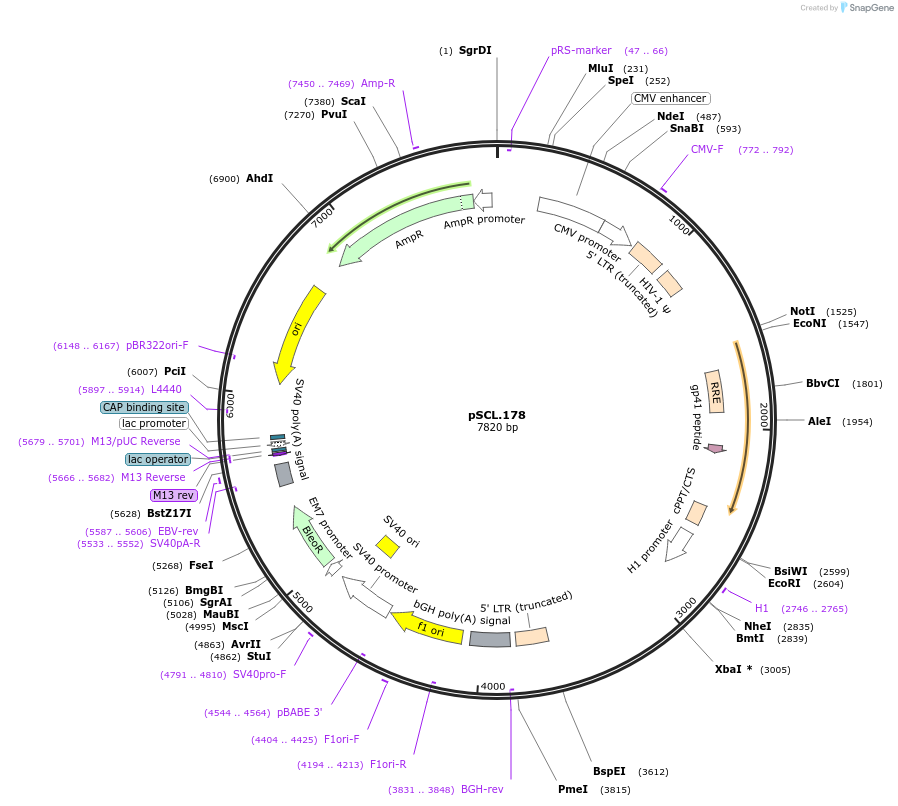
pSCL.178
Plasmid#184999PurposeExpress -Eco1 FANCF editing ncRNA and gRNADepositorInsertEco1: FANCF targeting and editing ncRNA-gRNA (a1/a2 length: 27 v1), expressed constitutively from a pol III (H1) promoter
TagsSV40NLSExpressionMammalianMutationFANCF donor, a1/a2 length extended for 27 bpPromoterH1Available SinceNov. 4, 2022AvailabilityAcademic Institutions and Nonprofits only -
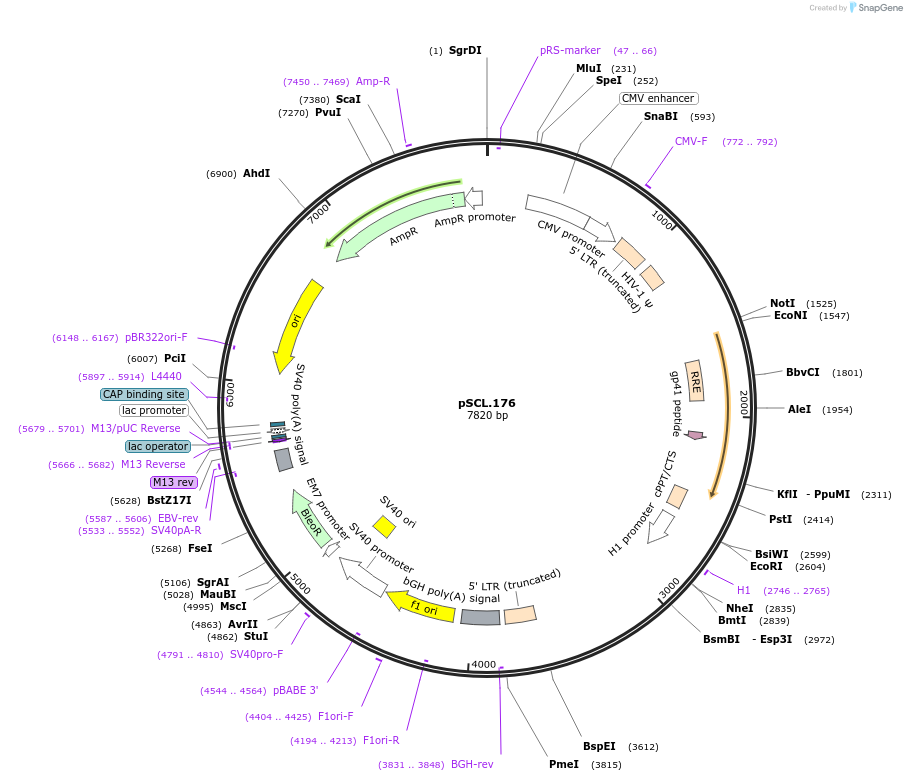
pSCL.176
Plasmid#184997PurposeExpress -Eco1 RNF2 editing ncRNA and gRNADepositorInsertEco1: RNF2 targeting and editing ncRNA-gRNA (a1/a2 length: 27 v1), expressed constitutively from a pol III (H1) promoter
TagsSV40NLSExpressionMammalianMutationRNF2 donor, a1/a2 length extended for 27 bpPromoterH1Available SinceNov. 4, 2022AvailabilityAcademic Institutions and Nonprofits only -
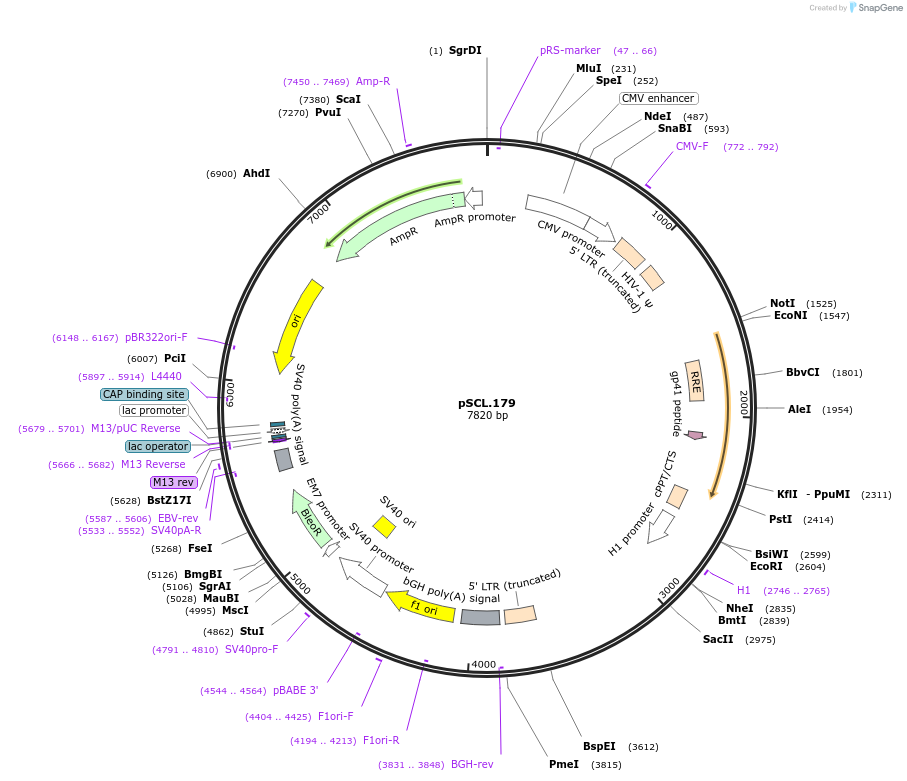
pSCL.179
Plasmid#185000PurposeExpress -Eco1 HEK4 editing ncRNA and gRNADepositorInsertEco1: HEK4 targeting and editing ncRNA-gRNA (a1/a2 length: 27 v1), expressed constitutively from a pol III (H1) promoter
TagsSV40NLSExpressionMammalianMutationHEK4 donor, a1/a2 length extended for 27 bpPromoterH1Available SinceNov. 3, 2022AvailabilityAcademic Institutions and Nonprofits only -
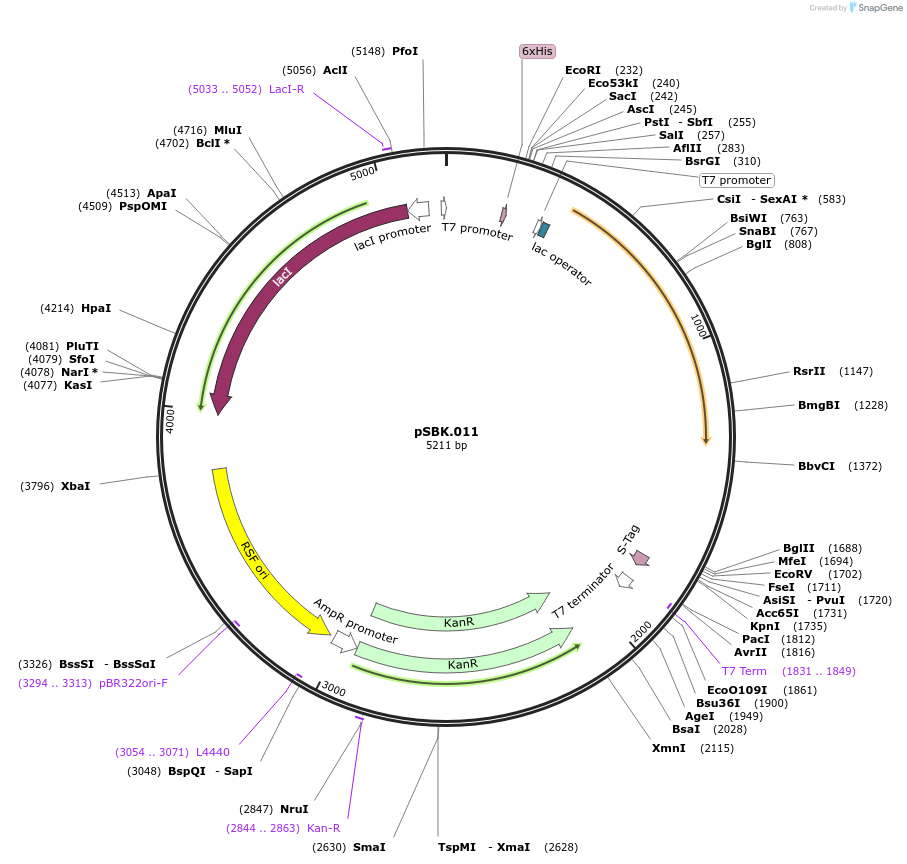
pSBK.011
Plasmid#187211PurposeExpresses retron Eco1 ncRNA v35 (long a1/a2, barcode 3 [CTG-CAG]), and Cas1 + Cas2DepositorInsertsRetron Eco1 ncRNA v35 (long a1/a2, barcode 3 [CTG-CAG])
Cas1 + Cas2
UseSynthetic BiologyExpressionBacterialPromoterT7/LacAvailable SinceSept. 28, 2022AvailabilityAcademic Institutions and Nonprofits only -
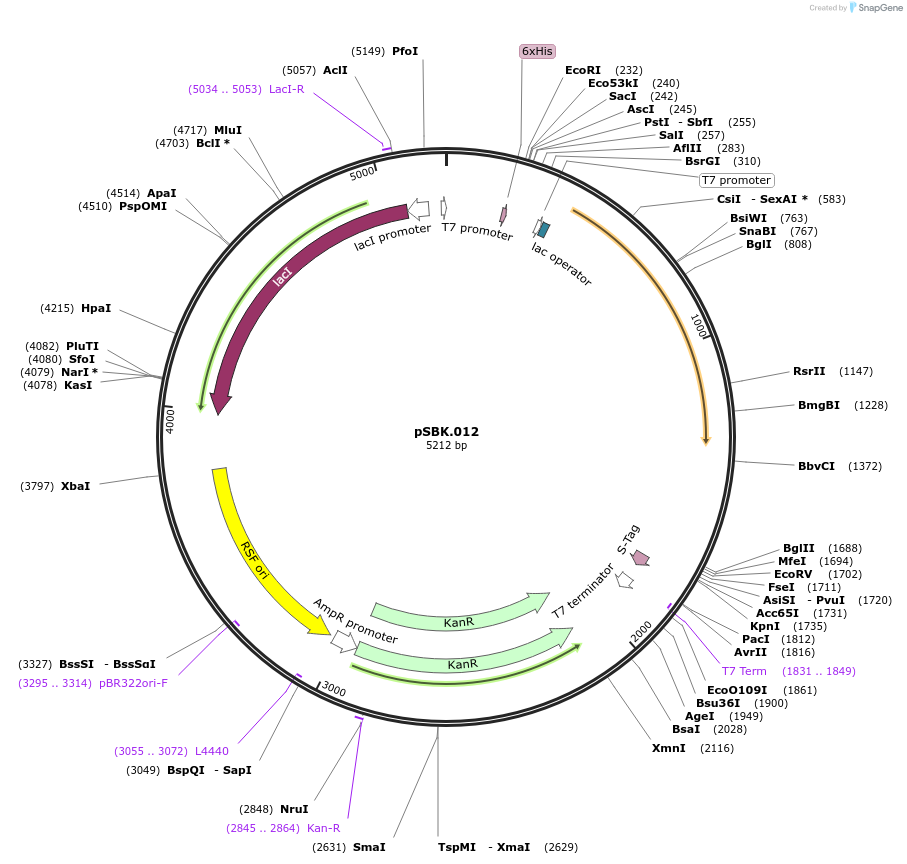
pSBK.012
Plasmid#187212PurposeExpresses retron Eco1 ncRNA v35 (long a1/a2, barcode 4 [GTG-CAC]), and Cas1 + Cas2DepositorInsertsRetron Eco1 ncRNA v35 (long a1/a2, barcode 4 [GTG-CAC])
Cas1 + Cas2
UseSynthetic BiologyExpressionBacterialPromoterT7/LacAvailable SinceSept. 28, 2022AvailabilityAcademic Institutions and Nonprofits only -
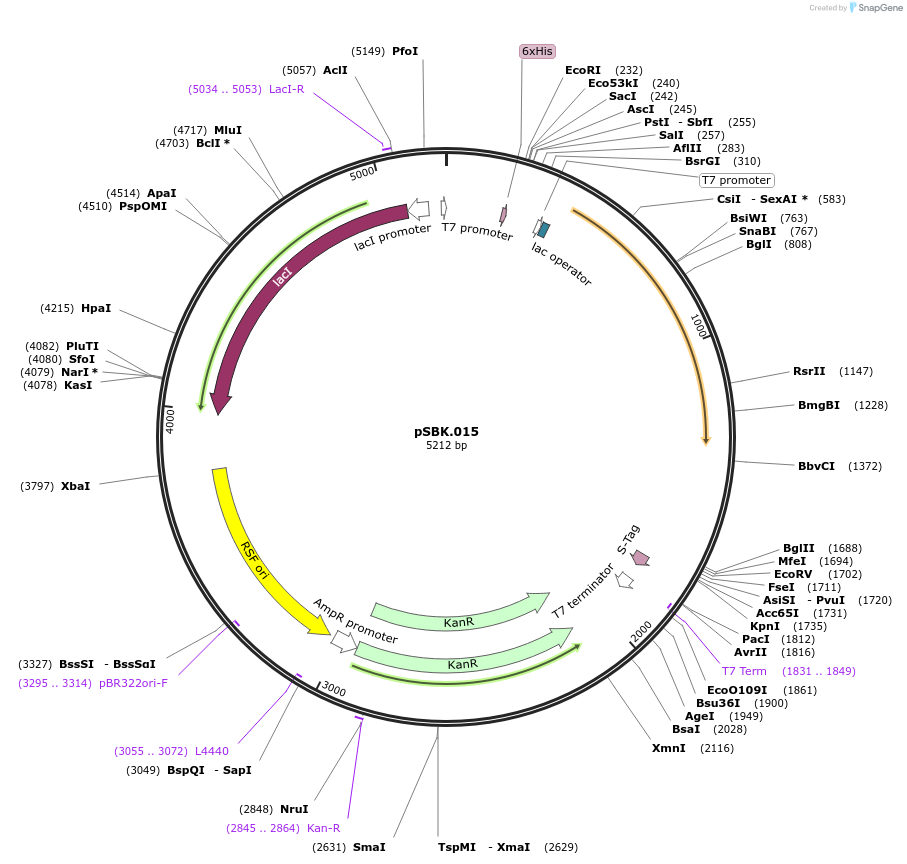
pSBK.015
Plasmid#187216PurposeExpresses retron Eco1 ncRNA v35 (long a1/a2, barcode 7 [GAG-CTC]), and Cas1 + Cas2DepositorInsertsRetron Eco1 ncRNA v35 (long a1/a2, barcode 7 [GAG-CTC])
Cas1 + Cas2
UseSynthetic BiologyExpressionBacterialPromoterT7/LacAvailable SinceSept. 28, 2022AvailabilityAcademic Institutions and Nonprofits only -
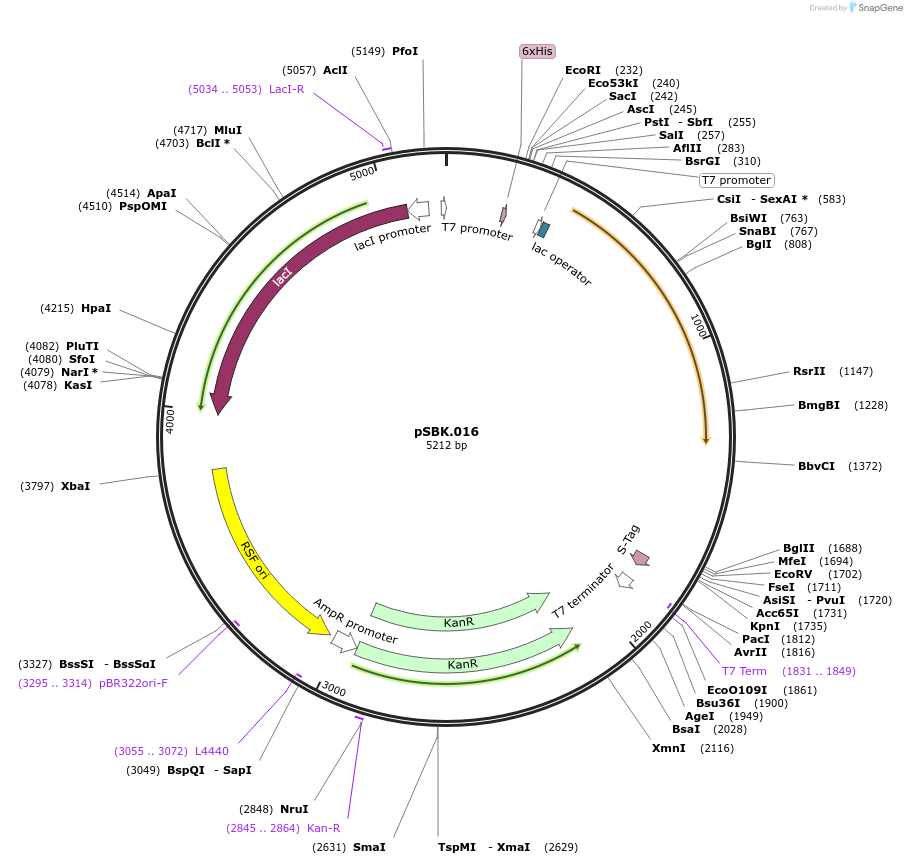
pSBK.016
Plasmid#187217PurposeExpresses retron Eco1 ncRNA v35 (long a1/a2, barcode 8 [GCT-TGC]), and Cas1 + Cas2DepositorInsertsRetron Eco1 ncRNA v35 (long a1/a2, barcode 8 [GCA-TGC])
Cas1 + Cas2
UseSynthetic BiologyExpressionBacterialPromoterT7/LacAvailable SinceSept. 28, 2022AvailabilityAcademic Institutions and Nonprofits only -
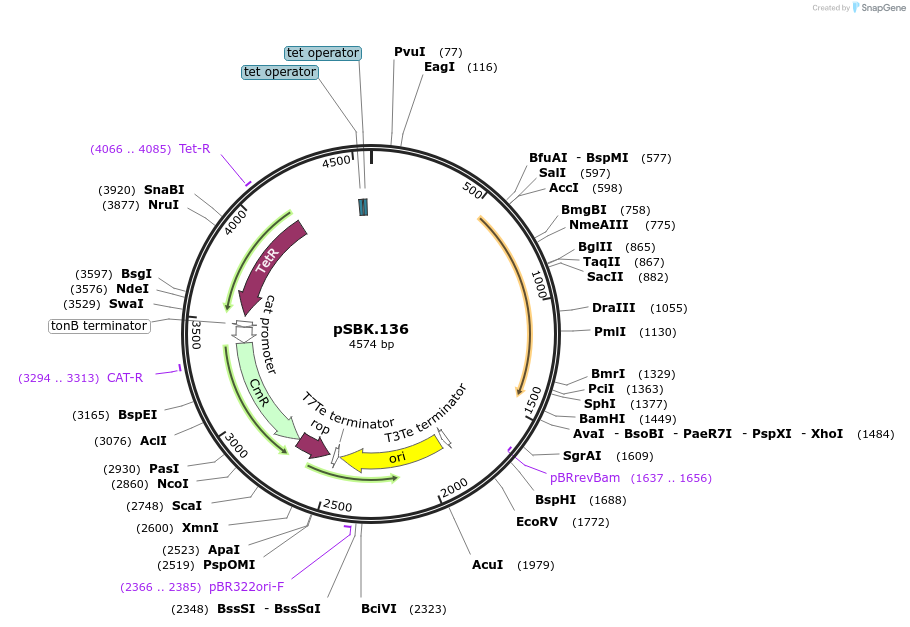
pSBK.136
Plasmid#187220PurposeExpresses Eco1 ncRNA v35 (long a1/a2) and Eco1 ncRNA v35 (long a1/a2, barcode 6) from promoters pSalTTC and pTet*, respectivelyDepositorInsertRetron Eco1 ncRNA v35 (long a1/a2); Retron Eco1 ncRNA v35 (long a1/a2, barcode 6)
ExpressionBacterialPromoterA: pSalTTC; B: pTet*Available SinceSept. 28, 2022AvailabilityAcademic Institutions and Nonprofits only -
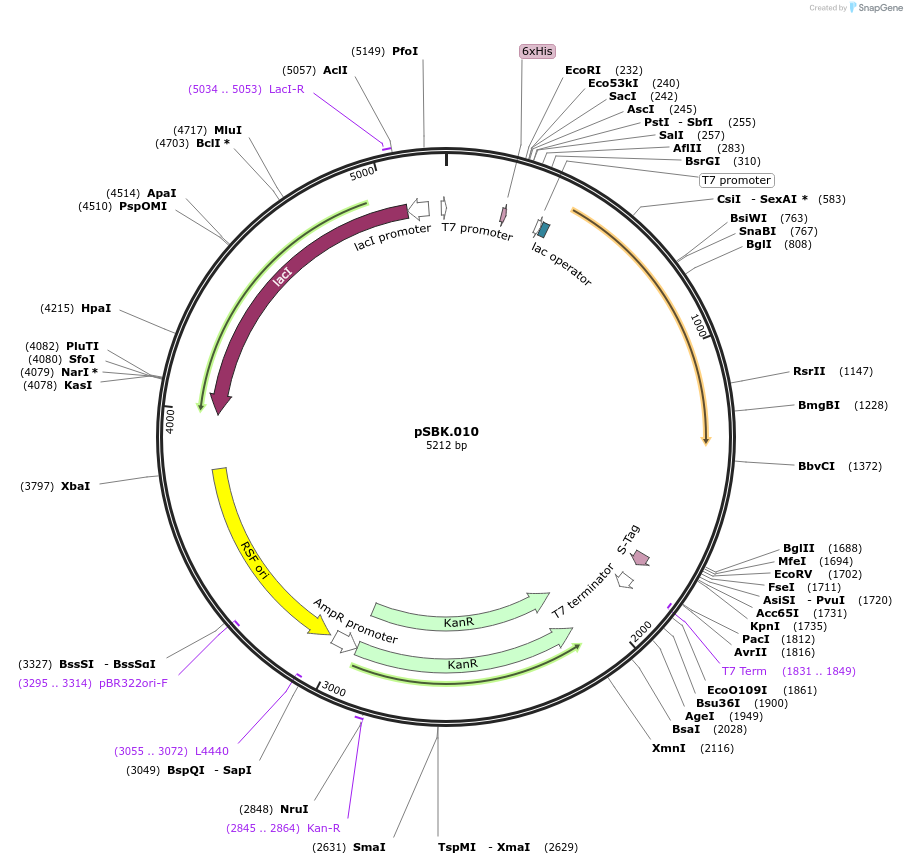
pSBK.010
Plasmid#187210PurposeExpresses retron Eco1 ncRNA v35 (long a1/a2, barcode 2 [GCT-AGC]), and Cas1 + Cas2DepositorInsertsRetron Eco1 ncRNA v35 (long a1/a2, barcode 2 [GCT-AGC])
Cas1 + Cas2
UseSynthetic BiologyExpressionBacterialPromoterT7/LacAvailable SinceSept. 28, 2022AvailabilityAcademic Institutions and Nonprofits only -
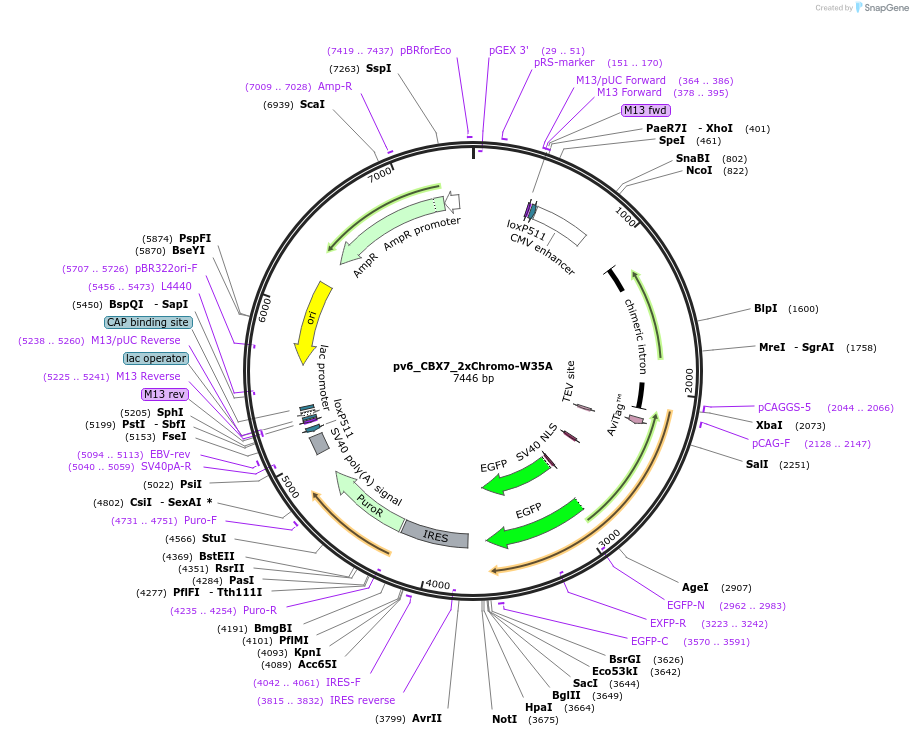
pv6_CBX7_2xChromo-W35A
Plasmid#179397PurposeCell line generation via recombination-mediated cassette exchange (RMCE) and stable expression of CBX7_2xChromo(W35A)DepositorInsertCBX7_2xChromo(W35A)
TagsAviTag and EGFPExpressionMammalianMutationChromodomain W35APromoterCAGGSAvailable SinceSept. 27, 2022AvailabilityAcademic Institutions and Nonprofits only -
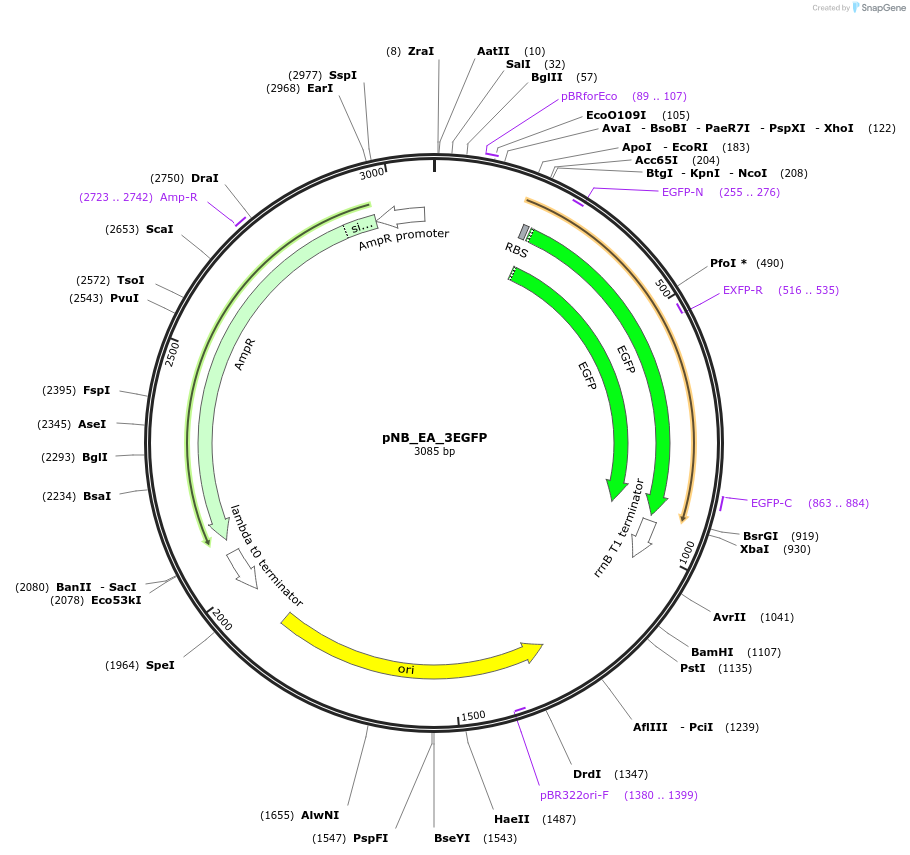
pNB_EA_3EGFP
Plasmid#185967PurposeExpresses EGFP from PR promoter in E.coli Dh5alpha cellDepositorInsertPr-Egfp cassette in forward direction in Network Brick
UseSynthetic BiologyPromoterPRAvailable SinceAug. 29, 2022AvailabilityAcademic Institutions and Nonprofits only -
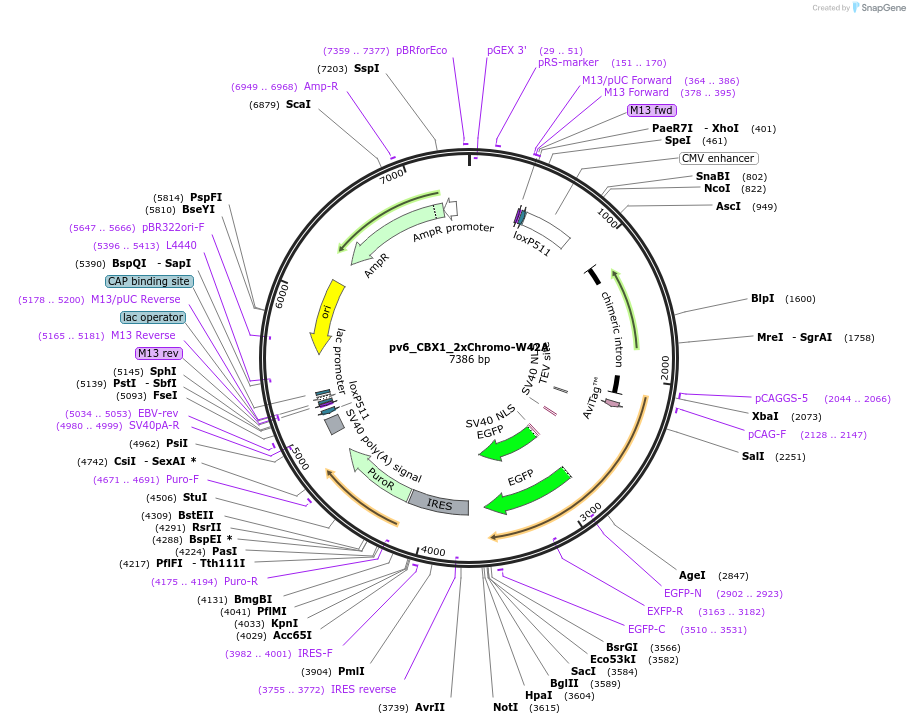
pv6_CBX1_2xChromo-W42A
Plasmid#179394PurposeCell line generation via recombination-mediated cassette exchange (RMCE) and stable expression of CBX1_2xChromo(W42A)DepositorInsertCBX1_2xChromo(W42A)
TagsAviTag and EGFPExpressionMammalianMutationChromodomain W42APromoterCAGGSAvailable SinceAug. 29, 2022AvailabilityAcademic Institutions and Nonprofits only -
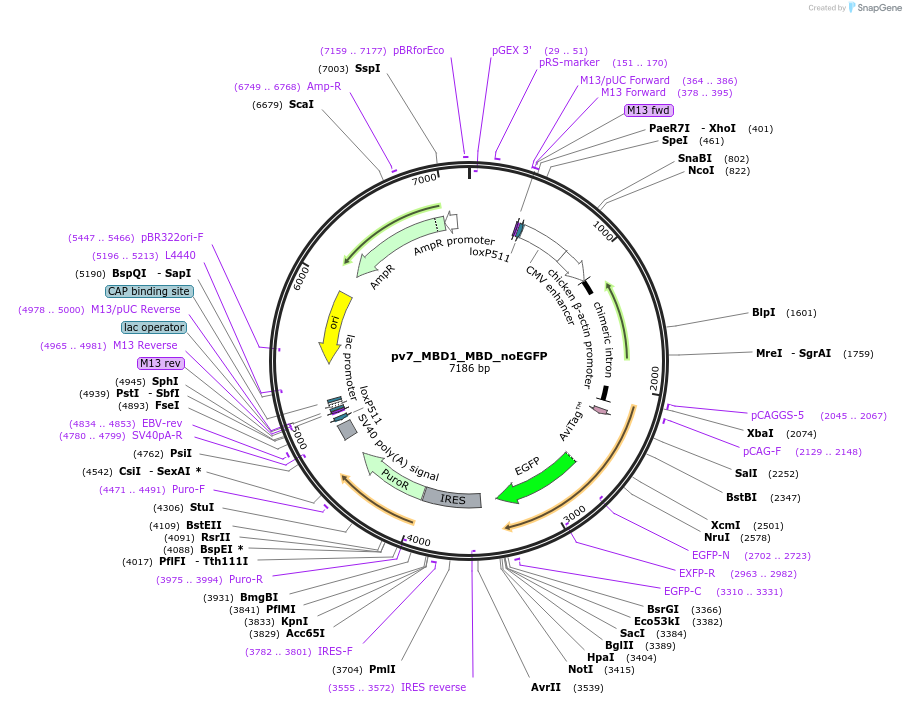
pv7_MBD1_MBD_noEGFP
Plasmid#179406PurposeCell line generation via recombination-mediated cassette exchange (RMCE) and stable expression of MBD1_MBDDepositorInsertMBD1_MBD
TagsAviTagExpressionMammalianPromoterCAGGSAvailable SinceAug. 29, 2022AvailabilityAcademic Institutions and Nonprofits only -
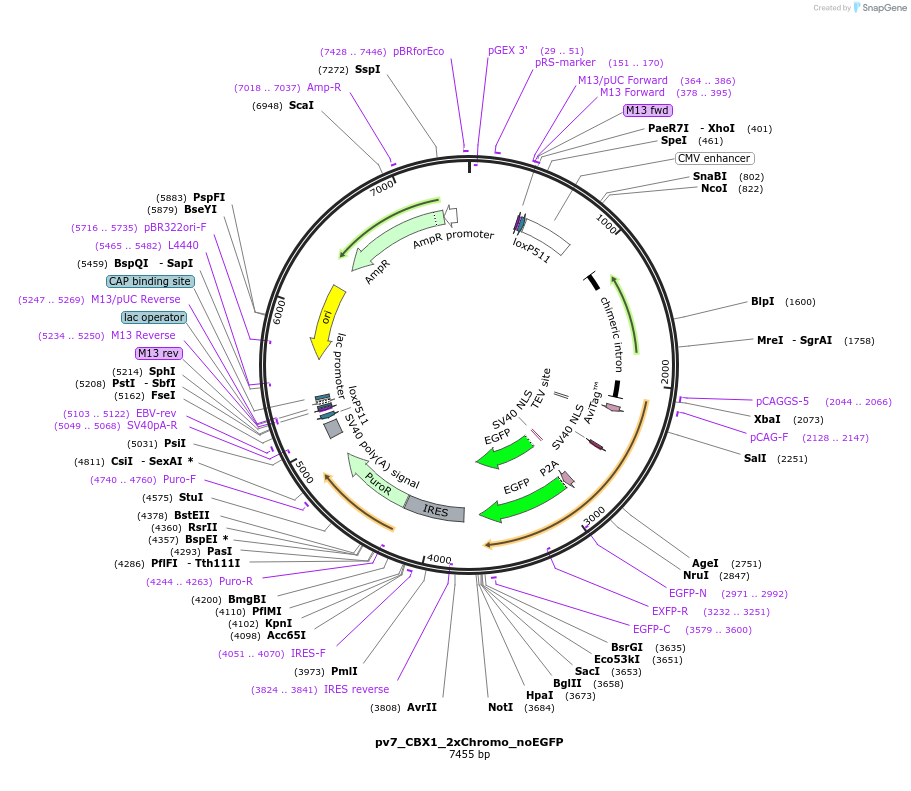
pv7_CBX1_2xChromo_noEGFP
Plasmid#179395PurposeCell line generation via recombination-mediated cassette exchange (RMCE) and stable expression of CBX1_2xChromo_noEGFPDepositorInsertCBX1_2xChromo_noEGFP
TagsAviTagExpressionMammalianPromoterCAGGSAvailable SinceAug. 26, 2022AvailabilityAcademic Institutions and Nonprofits only -
pv6_MBD1_MBD-R22A
Plasmid#179405PurposeCell line generation via recombination-mediated cassette exchange (RMCE) and stable expression of MBD1_MBD(R22A)DepositorInsertMBD1_MBD(R22A)
TagsAviTag and EGFPExpressionMammalianMutationMBD R22APromoterCAGGSAvailable SinceAug. 26, 2022AvailabilityAcademic Institutions and Nonprofits only -
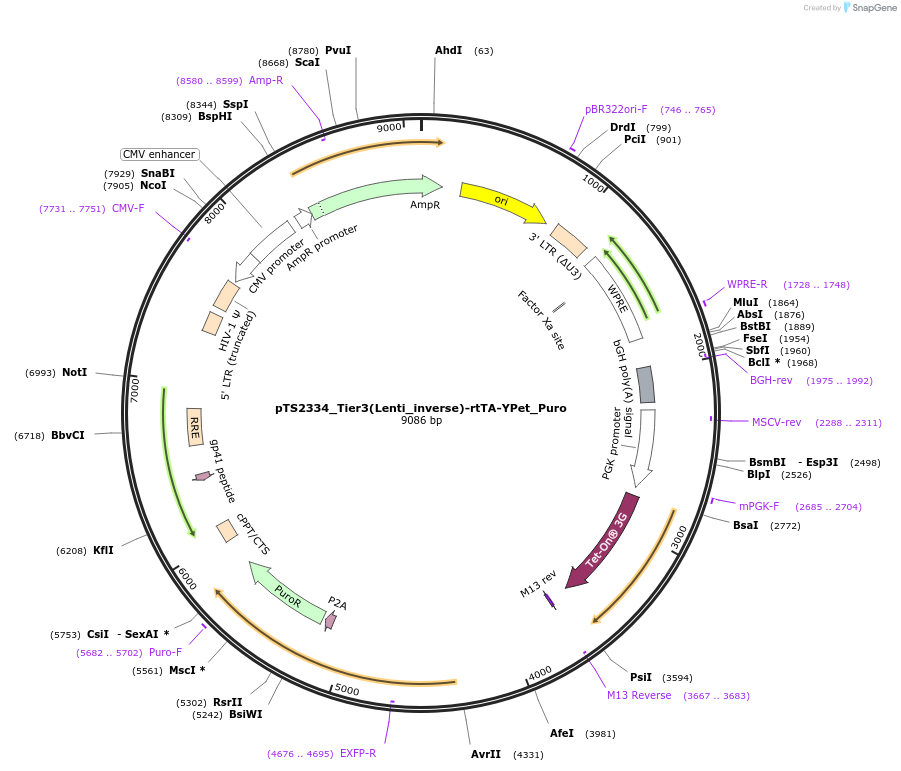
pTS2334_Tier3(Lenti_inverse)-rtTA-YPet_Puro
Plasmid#169665PurposeTier-3 vector for lentiviral stable integration of the rtTA and puromyine - Ypet selection cassette in inverted direction with polyA (PRPBSA-YPet-p2a-PuroR::PPGK-rtTA-pA::A3-pA)DepositorInsertPPGK-driven rtTA expression and PRPBSA-driven YPet and PuroR expression
ExpressionMammalianPromoterPhCMV / PPGK / RPBSAAvailable SinceJune 28, 2022AvailabilityAcademic Institutions and Nonprofits only -
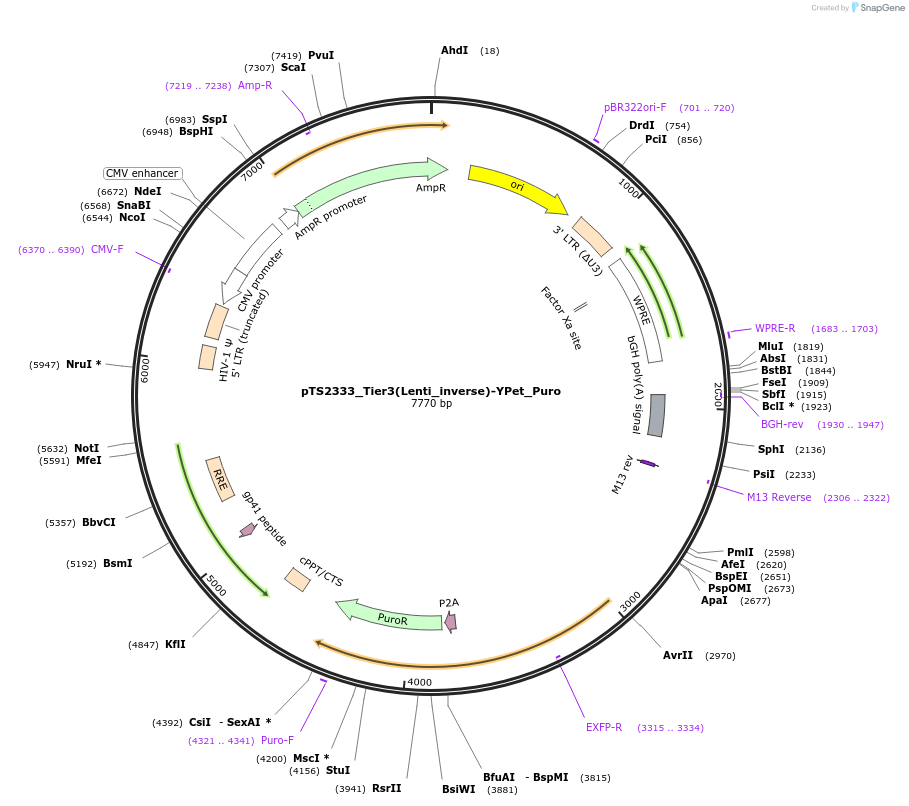
pTS2333_Tier3(Lenti_inverse)-YPet_Puro
Plasmid#169664PurposeTier-3 vector for stable integration via lentiviral transduction in inverted direction with polyA carrying a constitutive expression cassette for puromyine and Ypet (PCMV-5'LTR-Psi-gag-env-PRPBSA-YPet-p2A-PuroR-pA::A2-pA::A3-pA-WPRE-3'LTR)DepositorInsertPRPBSA-driven YPet and PuroR expression
ExpressionMammalianPromoterPhCMV / RPBSAAvailable SinceJune 28, 2022AvailabilityAcademic Institutions and Nonprofits only -
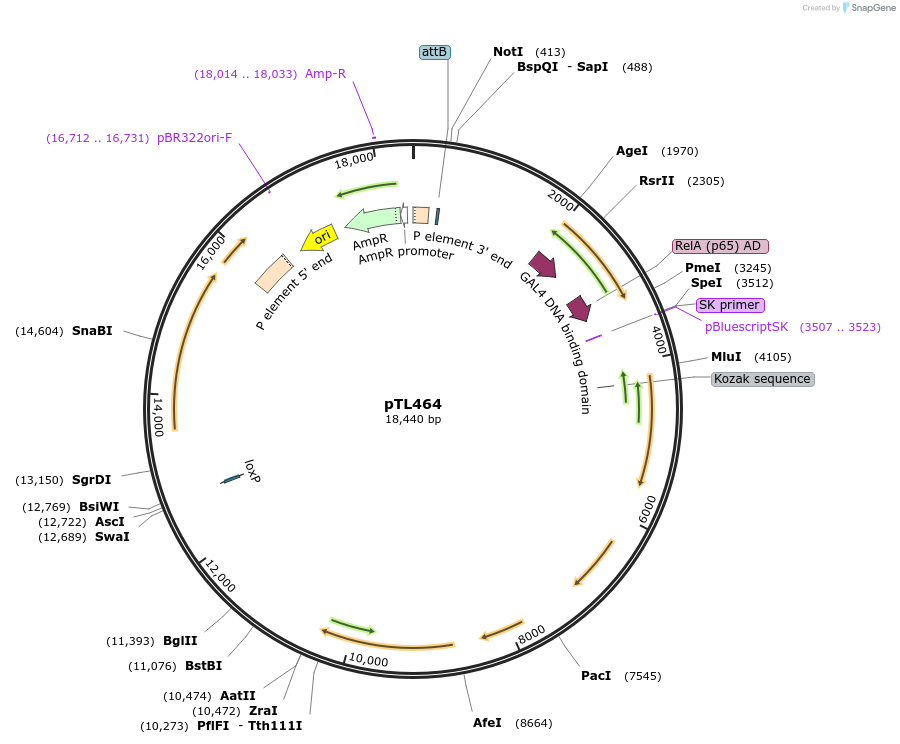
pTL464
Plasmid#69881Purposea dCirl transcriptional reporter allele that contains an optimized gal4.2::p65 cassette at the start codon of the genomic dCirl ORFDepositorInsertCIRL (Cirl Fly)
UsePhic31-integration vectorAvailable SinceJune 24, 2022AvailabilityAcademic Institutions and Nonprofits only -
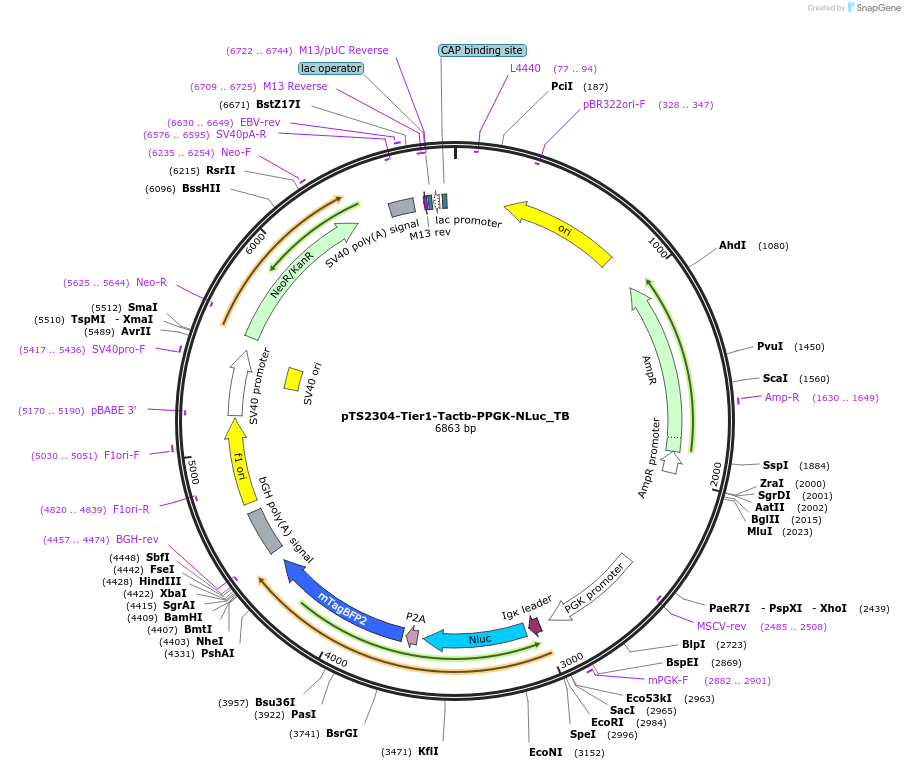
pTS2304-Tier1-Tactb-PPGK-NLuc_TB
Plasmid#169549PurposeTier-1 expression vector encoding PmPGK1-driven secreted Nluc coupled to fluorescent reporter mTagBFP2 that is insulated via a 5' Tactb insulator sequence (Tactb-PmPGK1-IgkSS-Nluc-p2A-mTagBFP2-pA).DepositorInsertTactb-modulated PPGK-driven NanoLuc Luciferase - mTagBFP expression cassette
ExpressionMammalianPromoterPPGKAvailable SinceJune 22, 2022AvailabilityAcademic Institutions and Nonprofits only