We narrowed to 10,386 results for: Coli
-
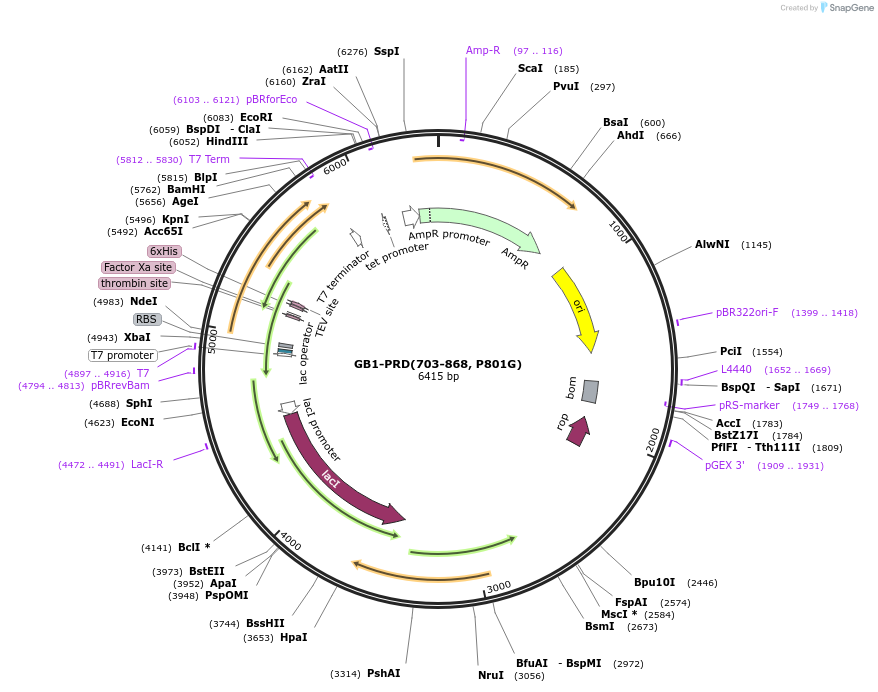
Plasmid#180029PurposeCodon-optimized bacterial expression plasmid. Expresses GB1-His6-TEV-ALIX(703-868, P801G). P801G mutation facilitates expression in E. coli.DepositorInsertGB1-His6-TEV-ALIX(703-868, P801G) (PDCD6IP Human)
TagsGB1-His6-TEVExpressionBacterialMutationP801GAvailable SinceJune 5, 2023AvailabilityAcademic Institutions and Nonprofits only -
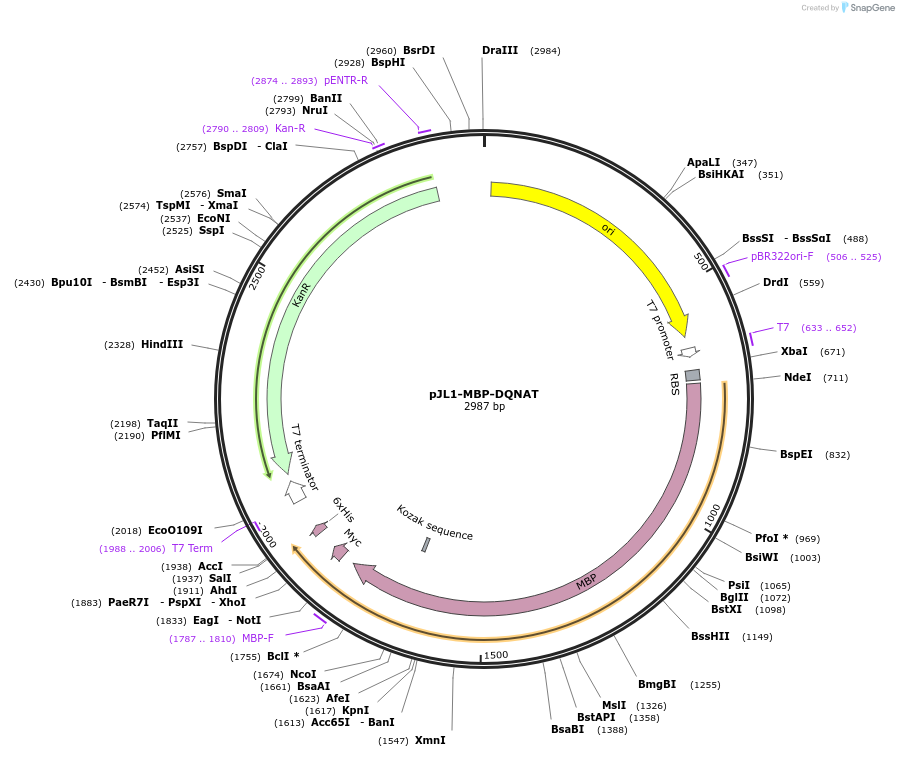
pJL1-MBP-DQNAT
Plasmid#198215PurposepJL1 plasmid encoding E. coli maltose binding protein (MBP) modified at the C-terminus with 30 amino acids containing one optimal DQNAT sequon followed by a 6x-His tagDepositorInsertMBP_DQNAT
Tags6x His Tag and DQNAT glycosylation tagExpressionBacterialPromoterT7Available SinceMay 23, 2023AvailabilityAcademic Institutions and Nonprofits only -
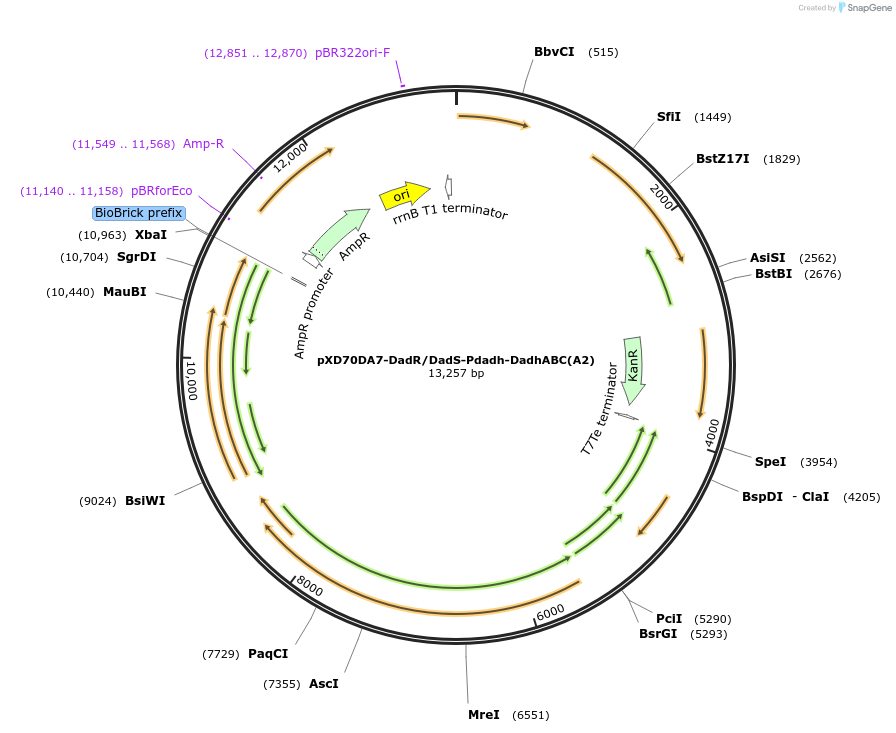
pXD70DA7-DadR/DadS-Pdadh-DadhABC(A2)
Plasmid#191630PurposeE. coli - Eggerthella lenta shuttle plasmid (KanR), native promoter-dopamine dehydroxylase from dopamine-metabolizing E. lenta A2 strain with transcriptional regulators DadR/DadSDepositorInsertDopamine dehydroxylase
ExpressionBacterialAvailable SinceJan. 5, 2023AvailabilityAcademic Institutions and Nonprofits only -
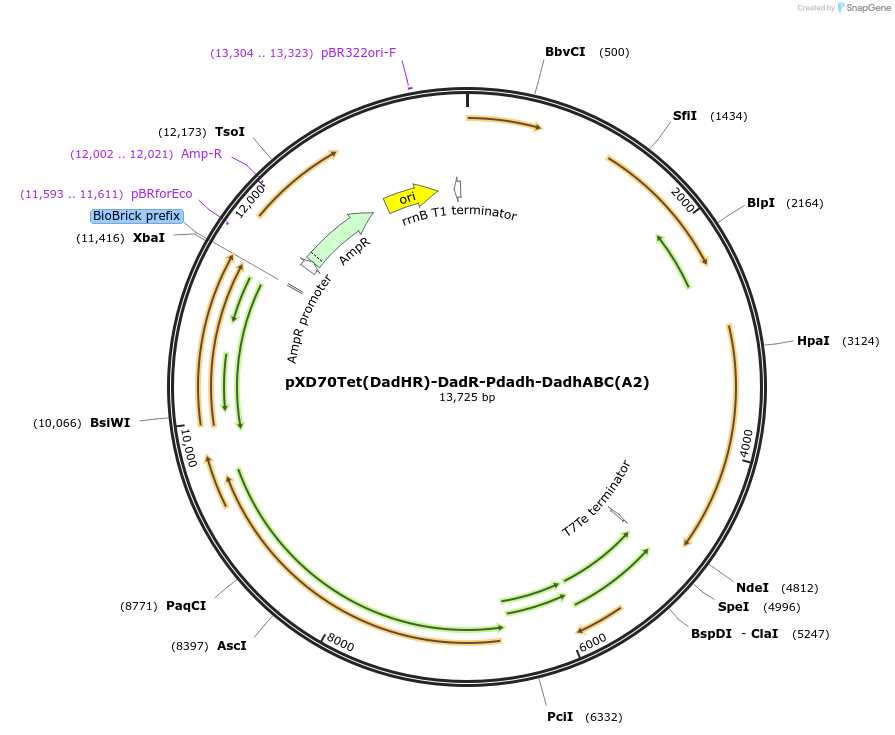
pXD70Tet(DadHR)-DadR-Pdadh-DadhABC(A2)
Plasmid#191645PurposeE. coli - Eggerthella lenta shuttle plasmid (TetR), native promoter-dopamine dehydroxylase from dopamine-metabolizing E. lenta A2 strain with transcriptional regulator DadRDepositorInsertDopamine dehydroxylase
ExpressionBacterialAvailable SinceJan. 5, 2023AvailabilityAcademic Institutions and Nonprofits only -
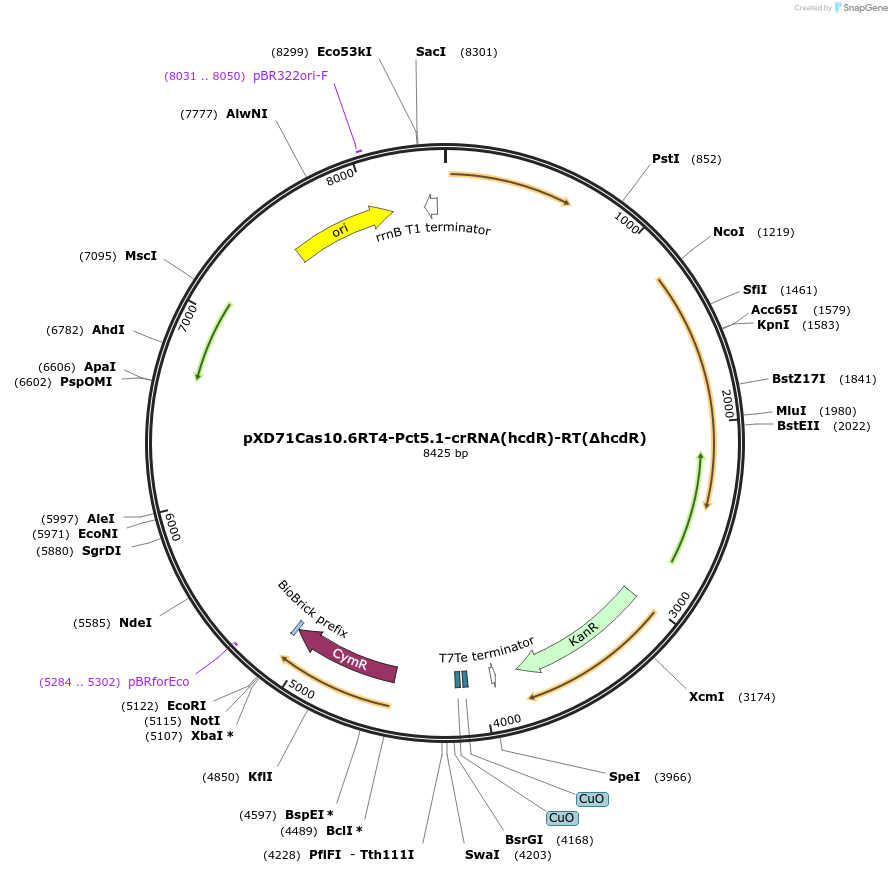
pXD71Cas10.6RT4-Pct5.1-crRNA(hcdR)-RT(ΔhcdR)
Plasmid#191646PurposeE. coli - Eggerthella lenta shuttle plasmid (KanR), cumate-inducible promoter Pct5.1-crRNA(hcdR-targeting spacer) hcdR-deleting repair template, used for hcdR deletionDepositorInsertCymR
UseCRISPRExpressionBacterialAvailable SinceJan. 5, 2023AvailabilityAcademic Institutions and Nonprofits only -
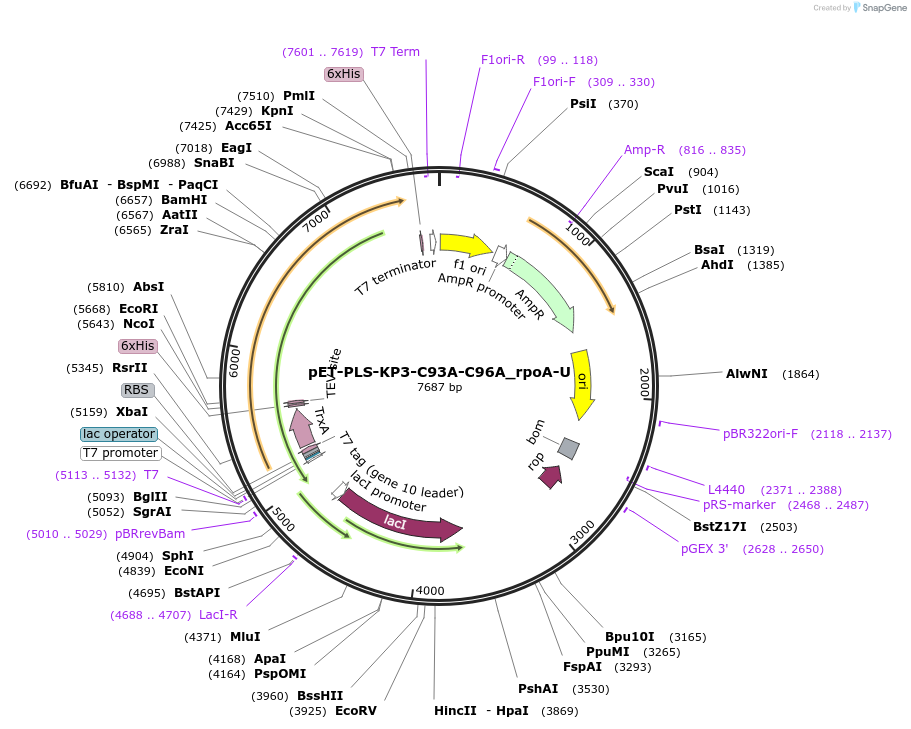
pET-PLS-KP3-C93A-C96A_rpoA-U
Plasmid#190962PurposeTest the U-to-C editing activity of KP3:C93A/C96A on AtrpoA editing site, in E. coliDepositorInsertPLS-KP3
TagsHisExpressionBacterialMutationDYW-KP:C93A/C96APromoterT7Available SinceOct. 31, 2022AvailabilityAcademic Institutions and Nonprofits only -
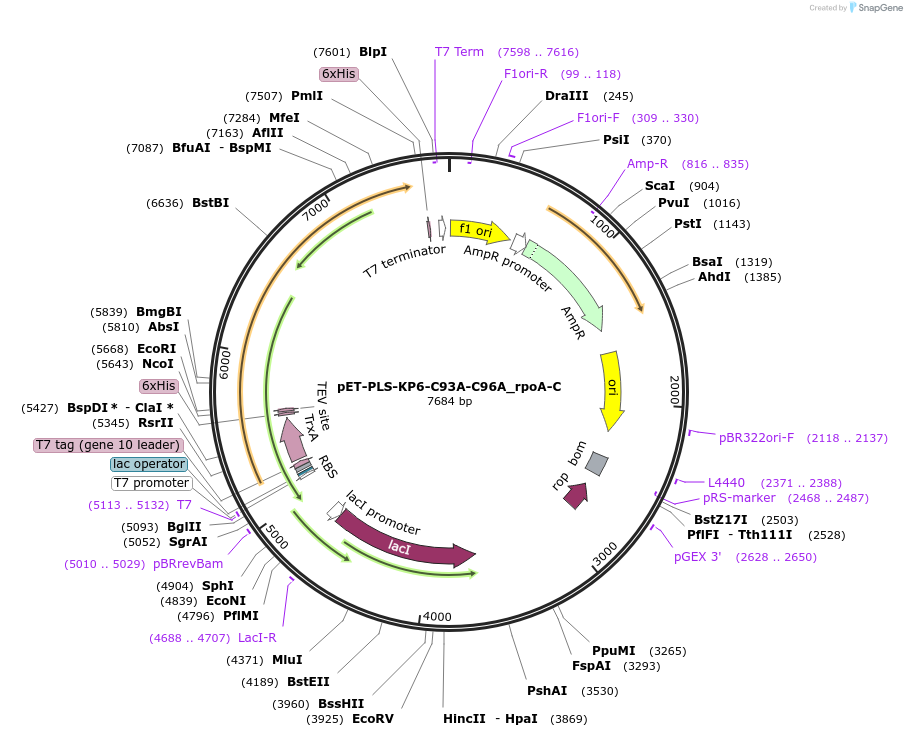
pET-PLS-KP6-C93A-C96A_rpoA-C
Plasmid#190970PurposeTest the C-to-U editing activity of KP6:C93A/C96A on AtrpoA editing site, in E. coliDepositorInsertPLS-KP6
TagsHisExpressionBacterialMutationDYW-KP:C93A/C96APromoterT7Available SinceOct. 31, 2022AvailabilityAcademic Institutions and Nonprofits only -
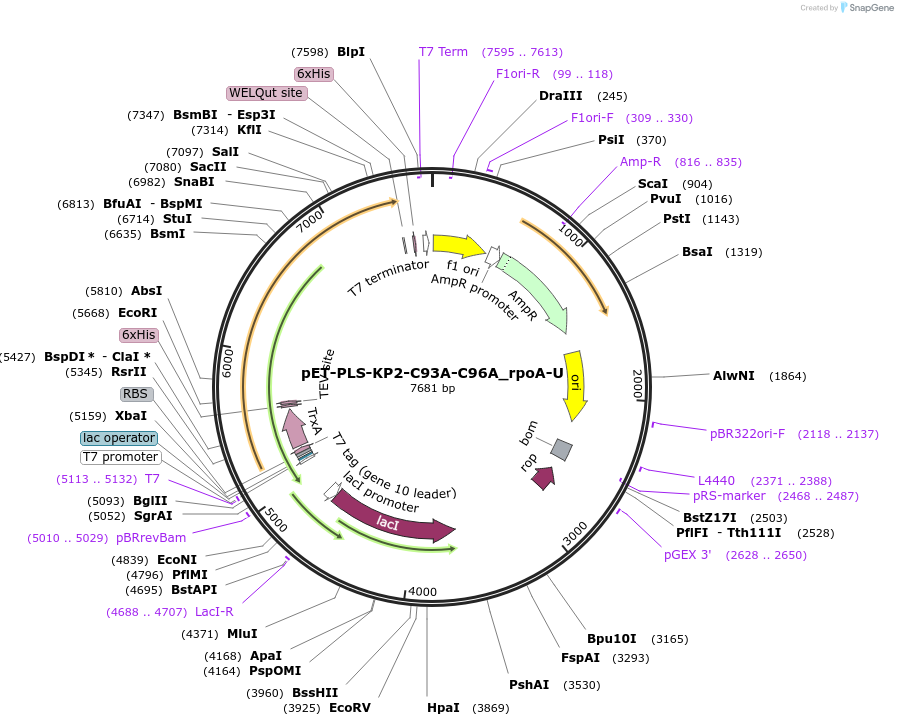
pET-PLS-KP2-C93A-C96A_rpoA-U
Plasmid#190958PurposeTest the U-to-C editing activity of KP2:C93A/C96A on AtrpoA editing site, in E. coliDepositorInsertPLS-KP2
TagsHisExpressionBacterialMutationDYW-KP:C93A/C96APromoterT7Available SinceOct. 28, 2022AvailabilityAcademic Institutions and Nonprofits only -
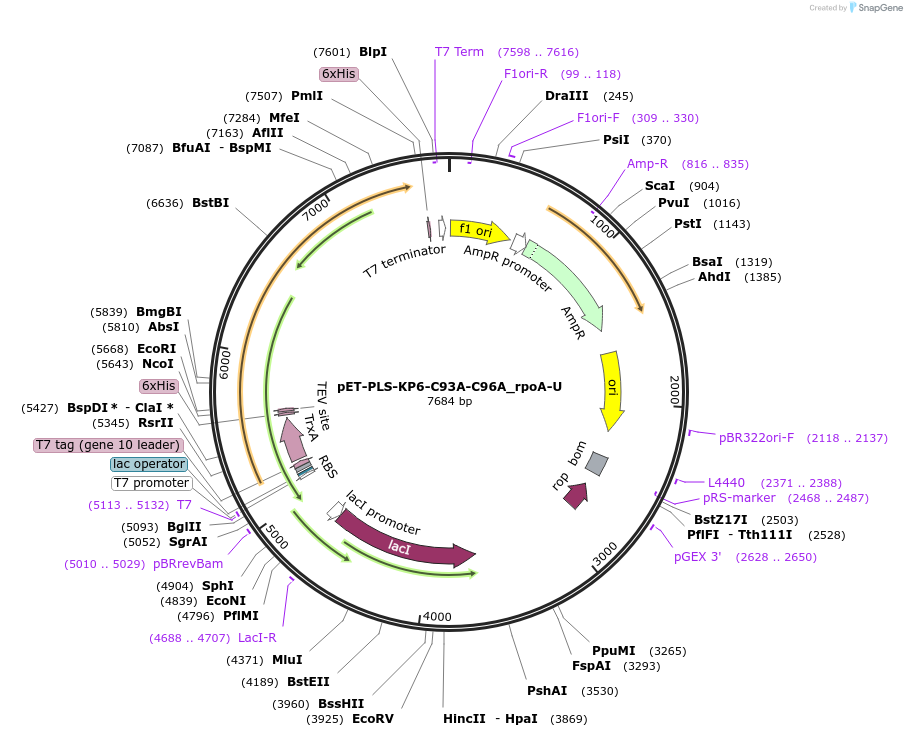
pET-PLS-KP6-C93A-C96A_rpoA-U
Plasmid#190971PurposeTest the U-to-C editing activity of KP6:C93A/C96A on AtrpoA editing site, in E. coliDepositorInsertPLS-KP6
TagsHisExpressionBacterialMutationDYW-KP:C93A/C96APromoterT7Available SinceOct. 28, 2022AvailabilityAcademic Institutions and Nonprofits only -
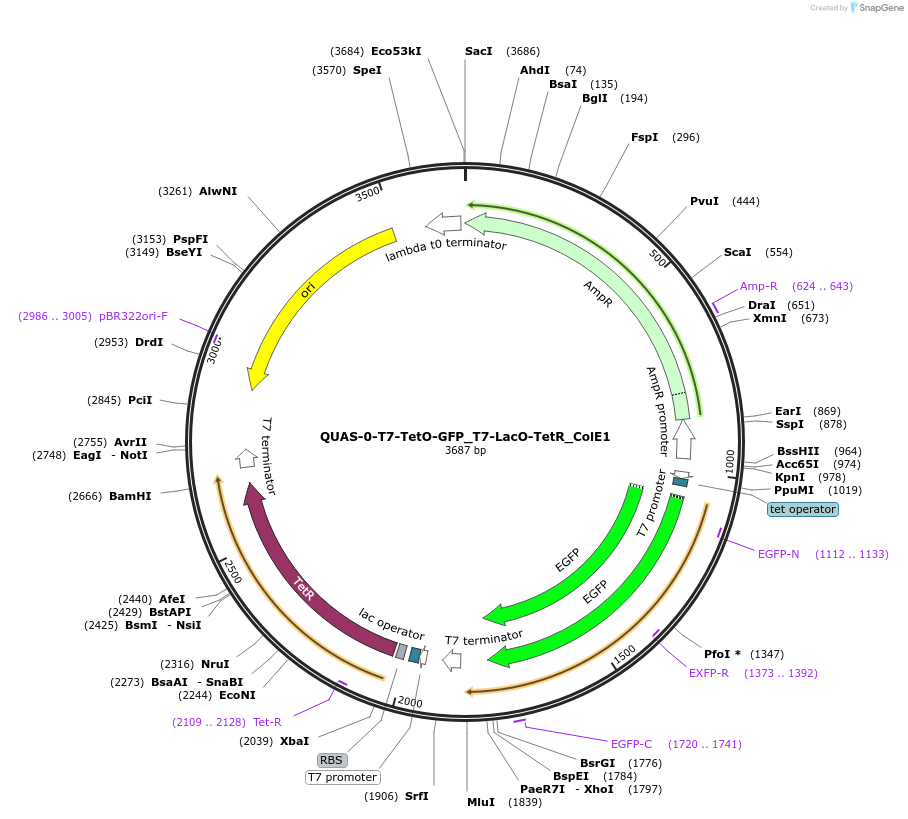
QUAS-0-T7-TetO-GFP_T7-LacO-TetR_ColE1
Plasmid#171663PurposeQUAS directly upstream of a T7 promoter with a TetO operator. Expression of TetR in BL21(DE3) E. coli. GFP is expressed according to presence of aTc. Contains the ColE1 origin of replication.DepositorInsertQUAS-T7-TetO-GFP ssrA-T7 Stop- T7-LacO-TetR-T7 Stop
TagsssrA degradation tag (DAS+4)ExpressionBacterialPromoterT7Available SinceOct. 14, 2022AvailabilityAcademic Institutions and Nonprofits only -
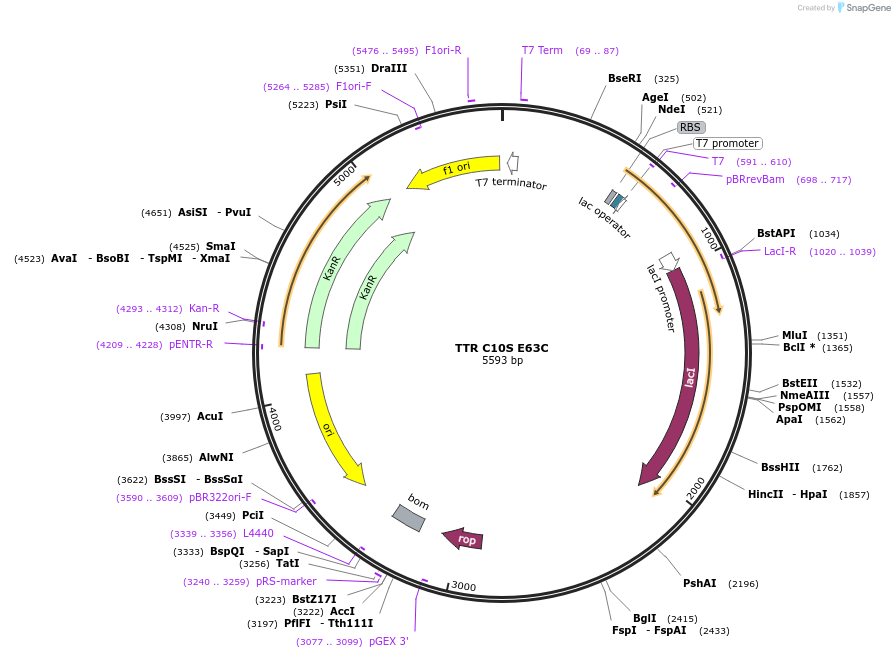
TTR C10S E63C
Plasmid#190009PurposeExpression of human TTR in E coliDepositorAvailable SinceSept. 27, 2022AvailabilityAcademic Institutions and Nonprofits only -
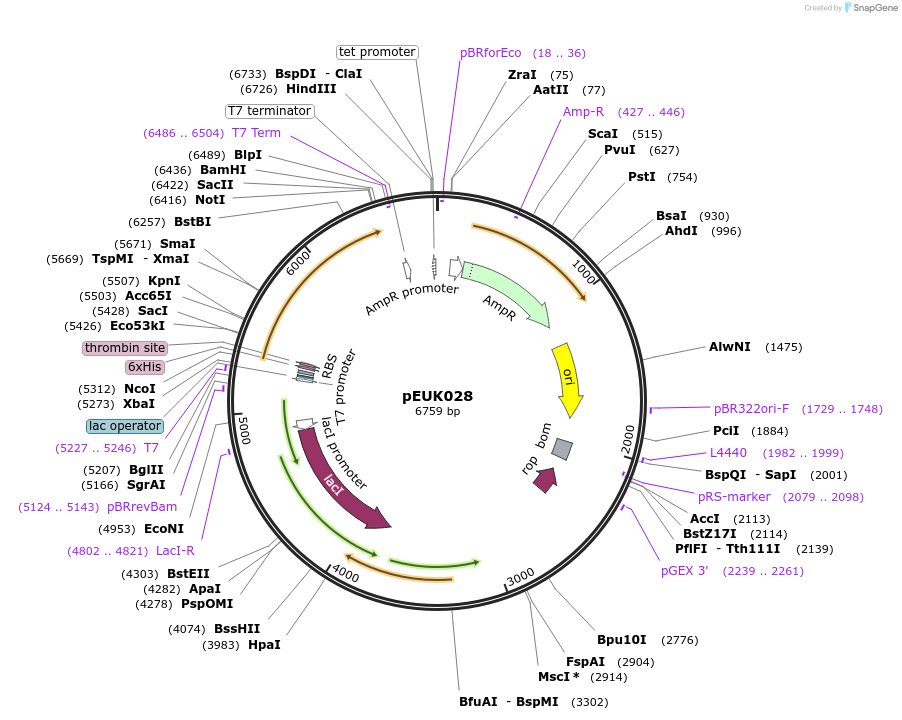
pEUK028
Plasmid#180826PurposeT7-inducible expression construct to produce SwGdmA (SWIT_RS16490) in E. coliDepositorArticleInsertSwGdmA (SWIT_RS16490 Sphingomonas wittichii RW1, Synthetic)
Tags6X HisExpressionBacterialPromoterT7Available SinceJune 28, 2022AvailabilityAcademic Institutions and Nonprofits only -
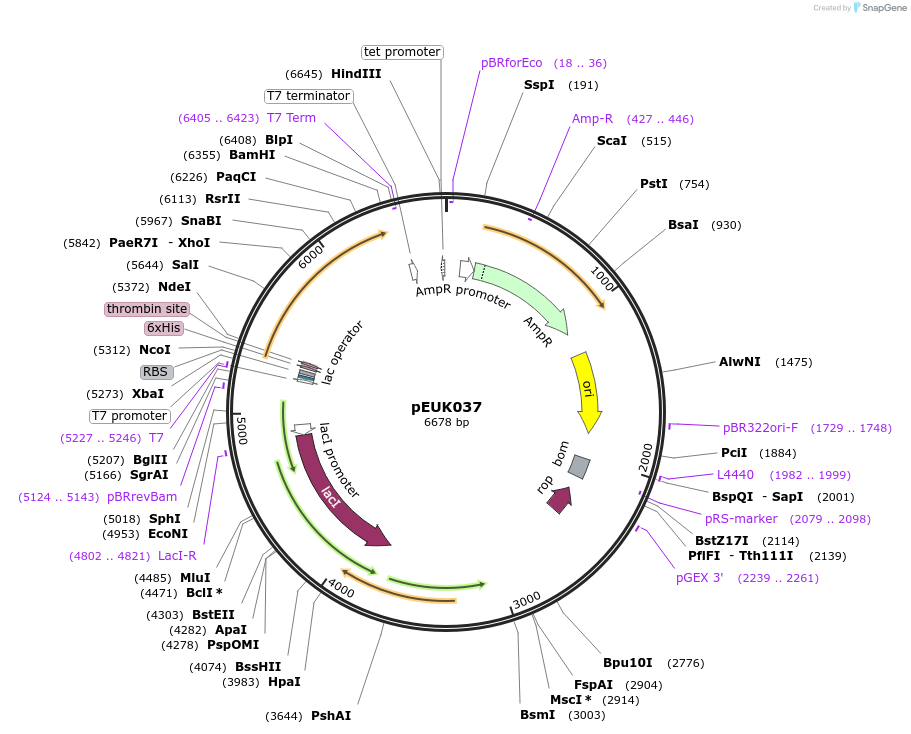
pEUK037
Plasmid#180829PurposeT7-inducible expression construct to produce CnGdmB (CNE_RS35790) in E. coliDepositorArticleAvailable SinceJune 28, 2022AvailabilityAcademic Institutions and Nonprofits only -
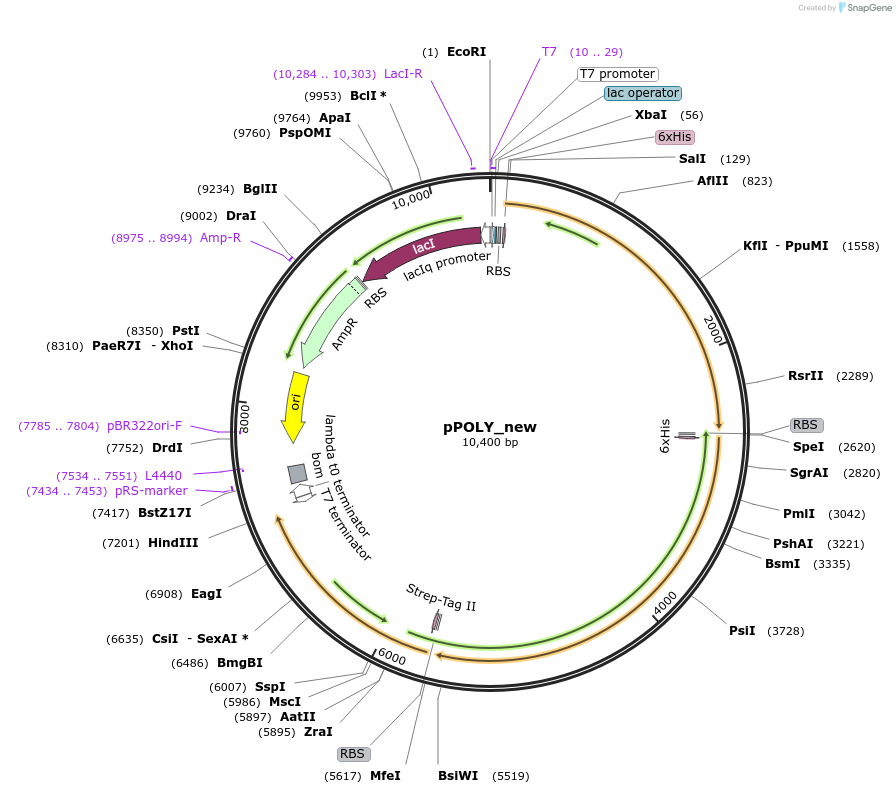
pPOLY_new
Plasmid#179276PurposeExpresses CuCBP, CcCDP and BaSP tricistronically in E. coli BL21(DE3)DepositorInsertscellodextrin phosphorylase
cellobiose phosphorylase
sucrose phosphorylase (C8077_RS00435 Bifidobacterium alodescentis)
UseSynthetic BiologyTags6xHIS and Strep-Tag IIExpressionBacterialPromoterT7 lacOAvailable SinceApril 13, 2022AvailabilityAcademic Institutions and Nonprofits only -
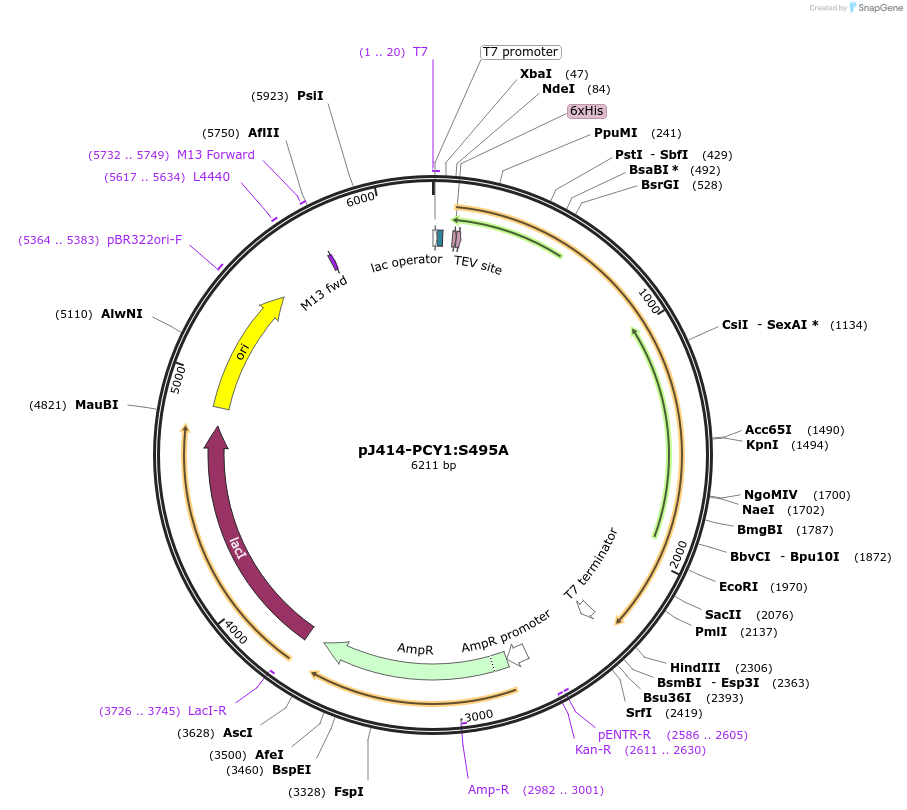
pJ414-PCY1:S495A
Plasmid#98151PurposeExpresses PCY1:S495A from Saponaria vaccaria possesing a N-terminal His6-tag (TEV cleavable) in E. coli.DepositorInsertPeptide cyclase 1:S495A
TagsHis6-TEVExpressionBacterialMutationChanged serine 495 to alaninePromoterT7Available SinceMarch 1, 2022AvailabilityAcademic Institutions and Nonprofits only -
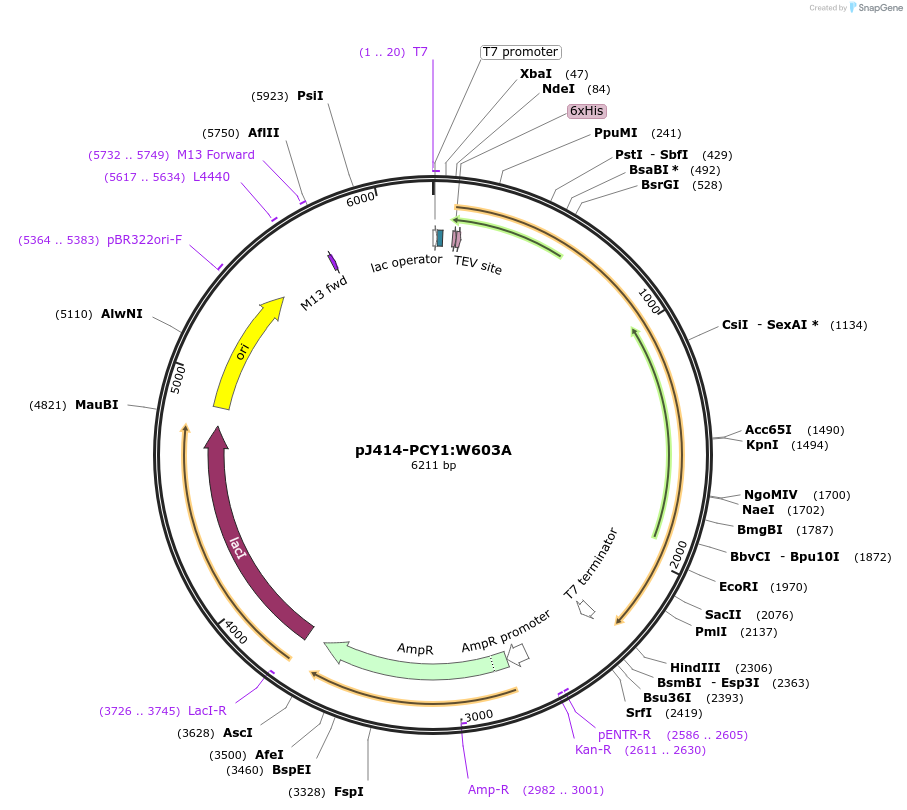
pJ414-PCY1:W603A
Plasmid#98153PurposeExpresses PCY1:W603A from Saponaria vaccaria possesing a N-terminal His6-tag (TEV cleavable) in E. coli.DepositorInsertPeptide cyclase 1:W603A
TagsHis6-TEVExpressionBacterialMutationChanged tryptophan 603 to alaninePromoterT7Available SinceMarch 1, 2022AvailabilityAcademic Institutions and Nonprofits only -
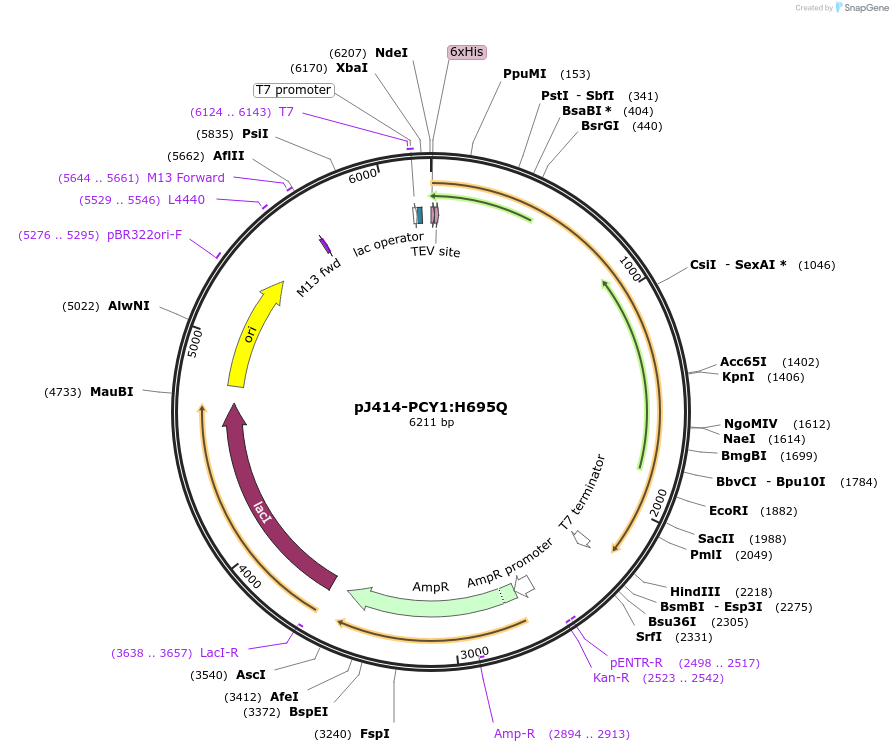
pJ414-PCY1:H695Q
Plasmid#98154PurposeExpresses PCY1:H695Q from Saponaria vaccaria possesing a N-terminal His6-tag (TEV cleavable) in E. coli.DepositorInsertPeptide cyclase 1:H695Q
TagsHis6-TEVExpressionBacterialMutationChanged histidine 695 to glutaminePromoterT7Available SinceMarch 1, 2022AvailabilityAcademic Institutions and Nonprofits only -
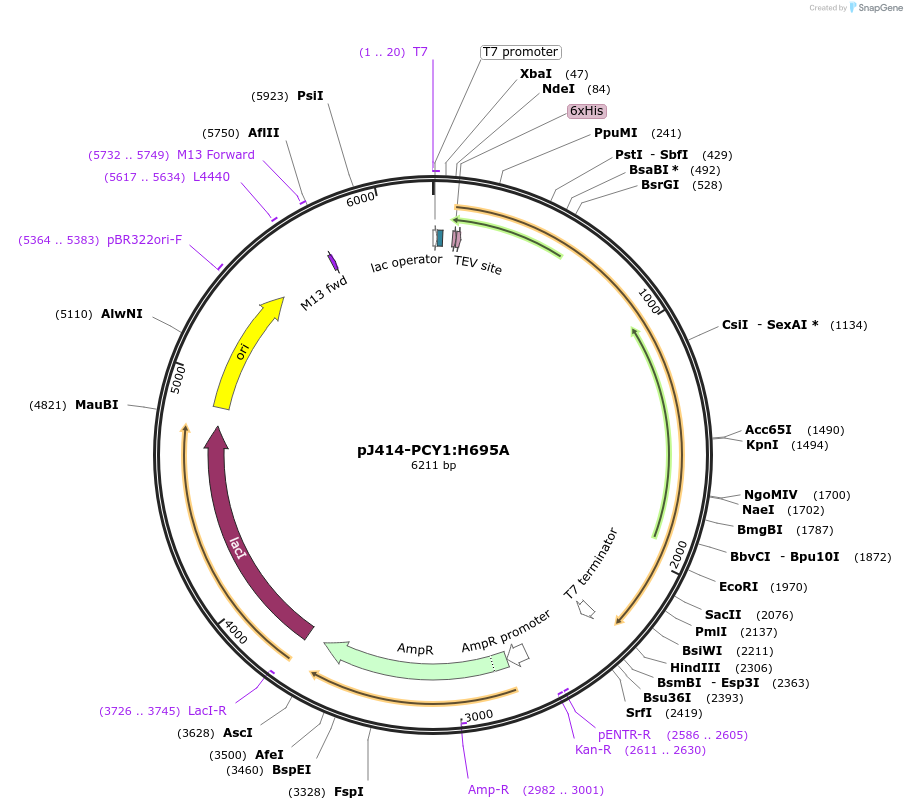
pJ414-PCY1:H695A
Plasmid#98155PurposeExpresses PCY1H695A from Saponaria vaccaria possesing a N-terminal His6-tag (TEV cleavable) in E. coli.DepositorInsertPeptide cyclase 1:H695A
TagsHis6-TEVExpressionBacterialMutationChanged histidine 695 to alaninePromoterT7Available SinceMarch 1, 2022AvailabilityAcademic Institutions and Nonprofits only -
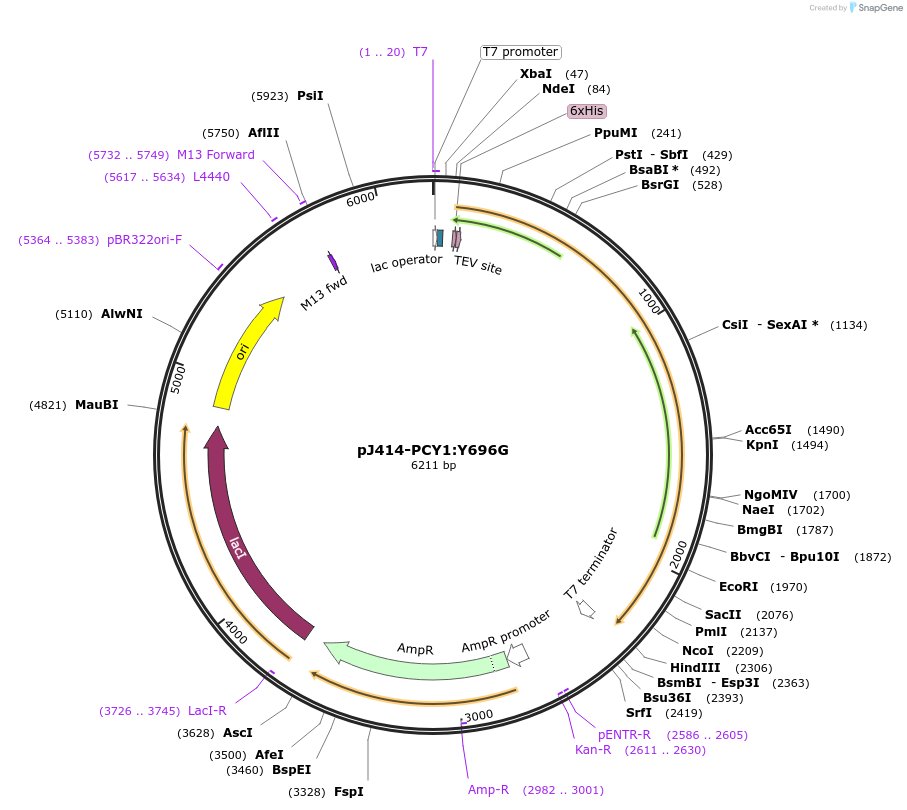
pJ414-PCY1:Y696G
Plasmid#98156PurposeExpresses PCY1:R696G from Saponaria vaccaria possesing a N-terminal His6-tag (TEV cleavable) in E. coli.DepositorInsertPeptide cyclase 1:Y696G
TagsHis6-TEVExpressionBacterialMutationChanged tyrosine 696 to glycinePromoterT7Available SinceMarch 1, 2022AvailabilityAcademic Institutions and Nonprofits only -
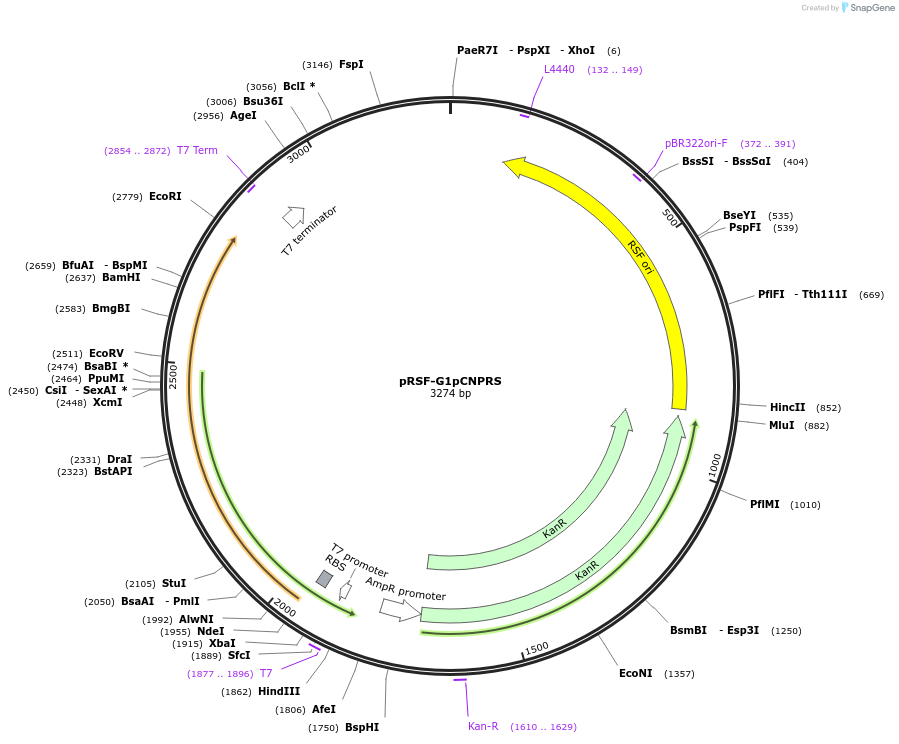
pRSF-G1pCNPRS
Plasmid#174719PurposetRNA synthetase/tRNA pair for the in vivo incorporation of L-3-(2-cyano-5-pyridyl)alanine (pCNP), into proteins in E. coli in response to the amber (TAG) codonDepositorInserttRNA synthetase
ExpressionBacterialPromoterT7Available SinceFeb. 10, 2022AvailabilityAcademic Institutions and Nonprofits only