We narrowed to 8,892 results for: AMPH
-
Plasmid#127143DepositorAvailable SinceJuly 26, 2019AvailabilityAcademic Institutions and Nonprofits only
-
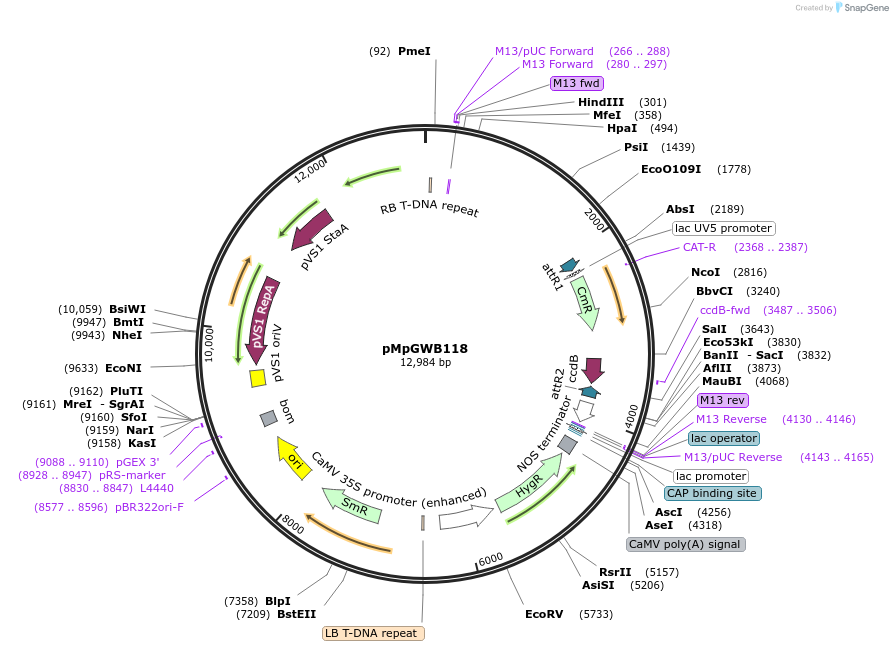
pMpGWB118
Plasmid#68572PurposeGateway binary vector designed for transgenic research with Marchantia polymorpha as well as other plantsDepositorTypeEmpty backboneTagsModified EAR motif plant-specific repression doma…ExpressionPlantPromoterMpEF1alpha promoterAvailable SinceJune 24, 2016AvailabilityAcademic Institutions and Nonprofits only -
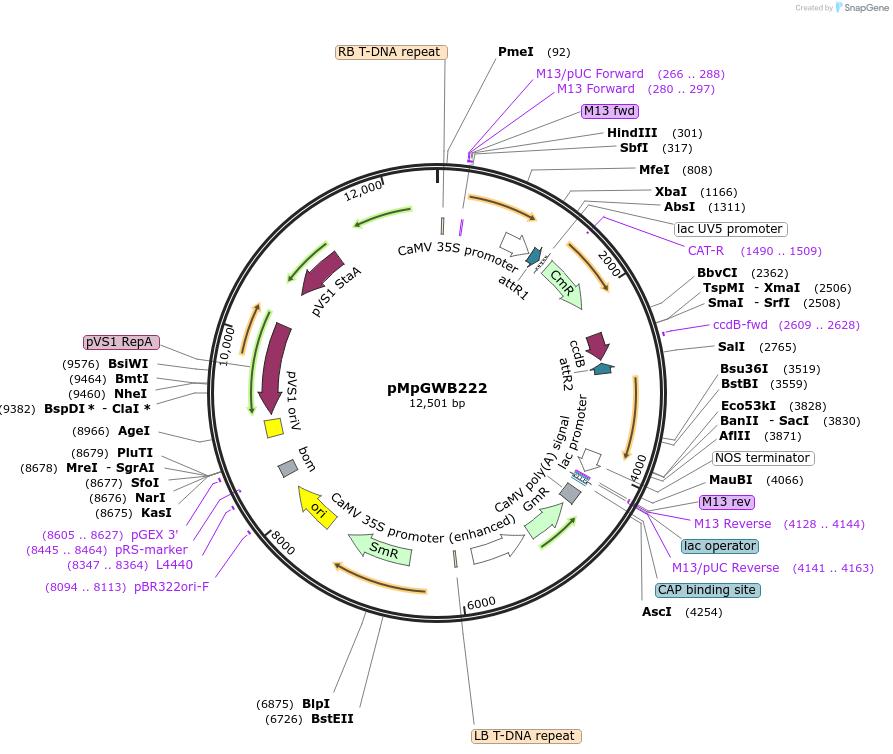
pMpGWB222
Plasmid#68613PurposeGateway binary vector designed for transgenic research with Marchantia polymorpha as well as other plantsDepositorTypeEmpty backboneTagsModified EAR motif plant-specific repression doma…ExpressionPlantPromoterCauliflower mosaic virus 35S promoterAvailable SinceJuly 13, 2016AvailabilityAcademic Institutions and Nonprofits only -
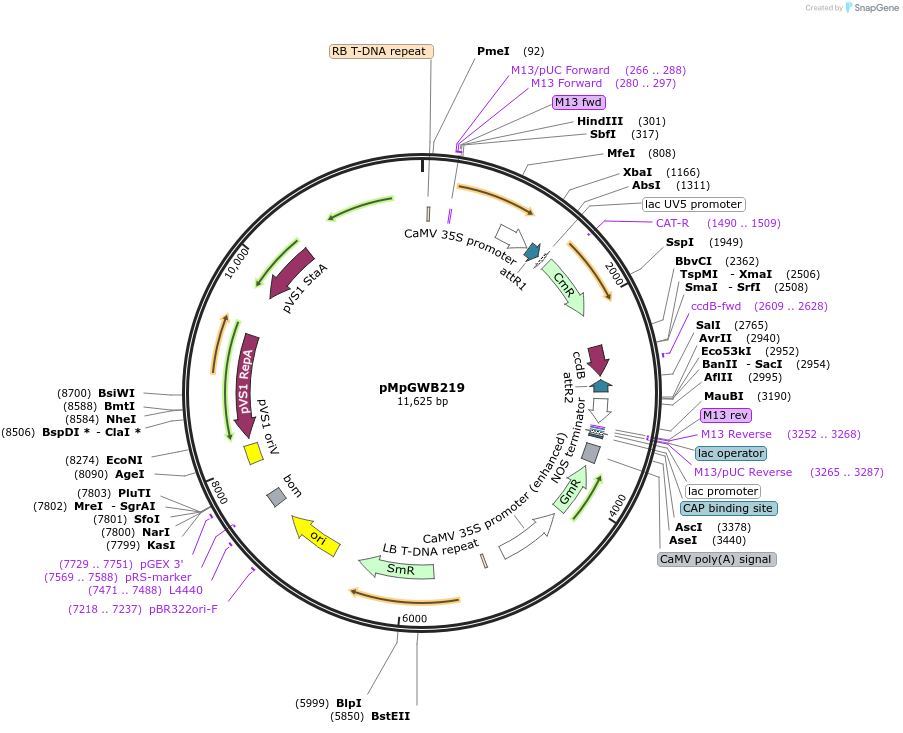
pMpGWB219
Plasmid#68610PurposeGateway binary vector designed for transgenic research with Marchantia polymorpha as well as other plantsDepositorTypeEmpty backboneTagsModified EAR motif plant-specific repression doma…ExpressionPlantPromoterCauliflower mosaic virus 35S promoterAvailable SinceJune 24, 2016AvailabilityAcademic Institutions and Nonprofits only -
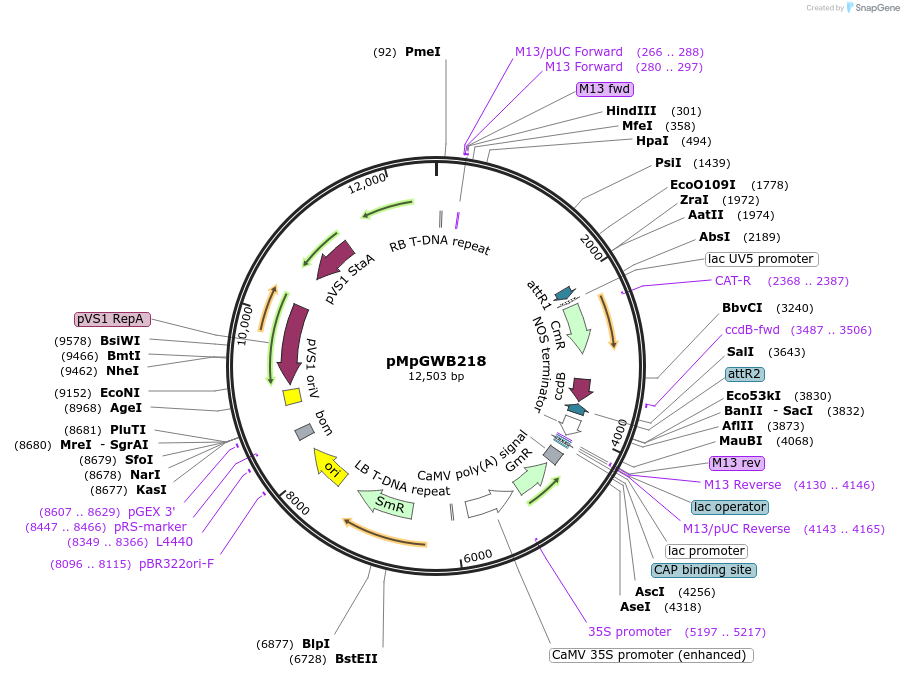
pMpGWB218
Plasmid#68609PurposeGateway binary vector designed for transgenic research with Marchantia polymorpha as well as other plantsDepositorTypeEmpty backboneTagsModified EAR motif plant-specific repression doma…ExpressionPlantPromoterMpEF1alpha promoterAvailable SinceJune 24, 2016AvailabilityAcademic Institutions and Nonprofits only -
pChlr-1SE
Plasmid#52140PurposeEncodes module 1 with amino acids 12S-16E for recognition of nt 8GDepositorInsertPumilio homology domain amino acids 25-60 (PUM1 Human)
ExpressionBacterialMutationChanged 16Q to 16E in Module 1Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-2SE
Plasmid#52144PurposeEncodes module 2 with amino acids 12S-16E for recognition of nt 7GDepositorInsertPumilio homology domain amino acids 61-96 (PUM1 Human)
ExpressionBacterialMutationChanged 12N to 12S and 16Q to 16E in Module 2Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-3SE
Plasmid#52148PurposeEncodes module 3 with amino acids 12S-16E for recognition of nt 6GDepositorInsertPumilio homology domain amino acids 97-132 (PUM1 Human)
ExpressionBacterialMutationChanged 12C to 12S and 16Q to 16E in Module 3Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
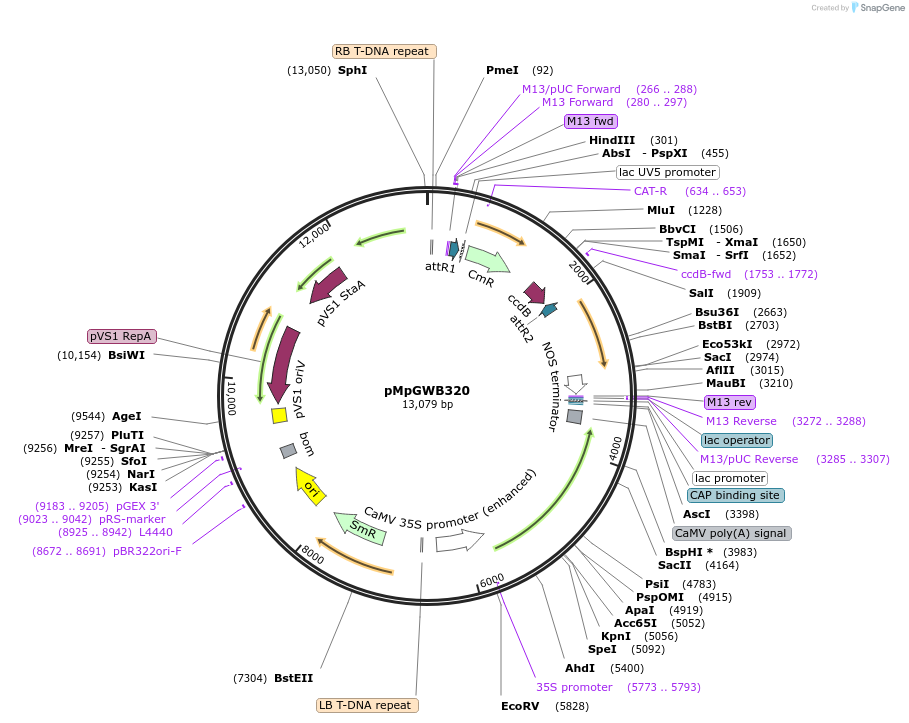
pMpGWB320
Plasmid#68648PurposeGateway binary vector designed for transgenic research with Marchantia polymorpha as well as other plantsDepositorTypeEmpty backboneTagsModified EAR motif plant-specific repression doma…ExpressionPlantAvailable SinceJune 24, 2016AvailabilityAcademic Institutions and Nonprofits only -
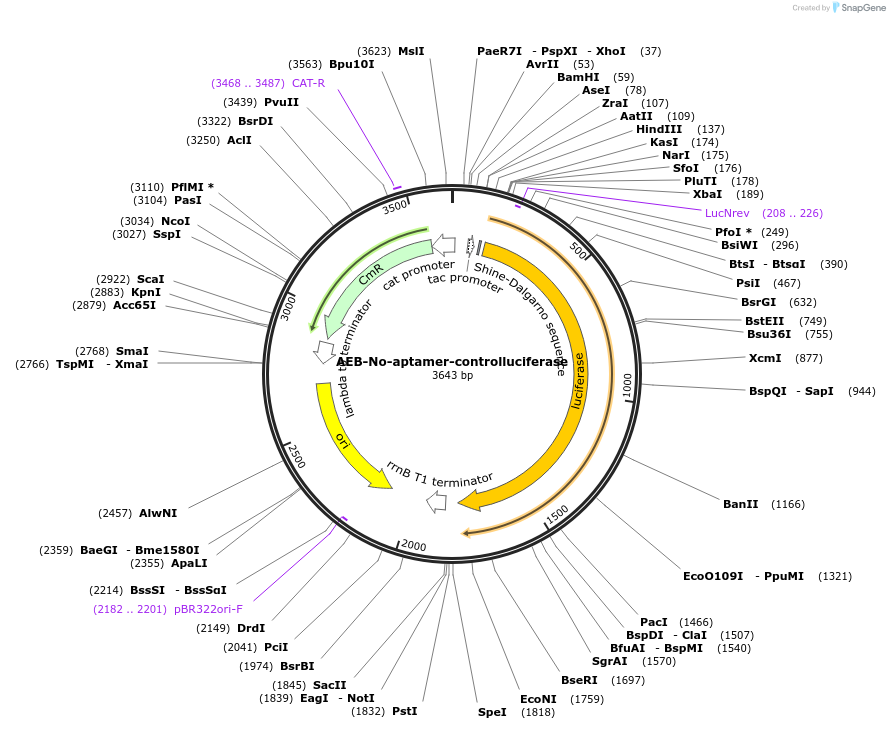
AEB-No-aptamer-controlluciferase
Plasmid#63853PurposeThis pFTV1 vector contains a no-aptamer control mRNA encoding luciferase gene. It should be used when testing AEB-Dopa5, AEB-T4-2 riboswitches to measure their non-specific ligand effect on luciferase expression.DepositorInsertNo-aptamer control mRNA
UseSynthetic BiologyPromotertacAvailable SinceFeb. 19, 2016AvailabilityAcademic Institutions and Nonprofits only -
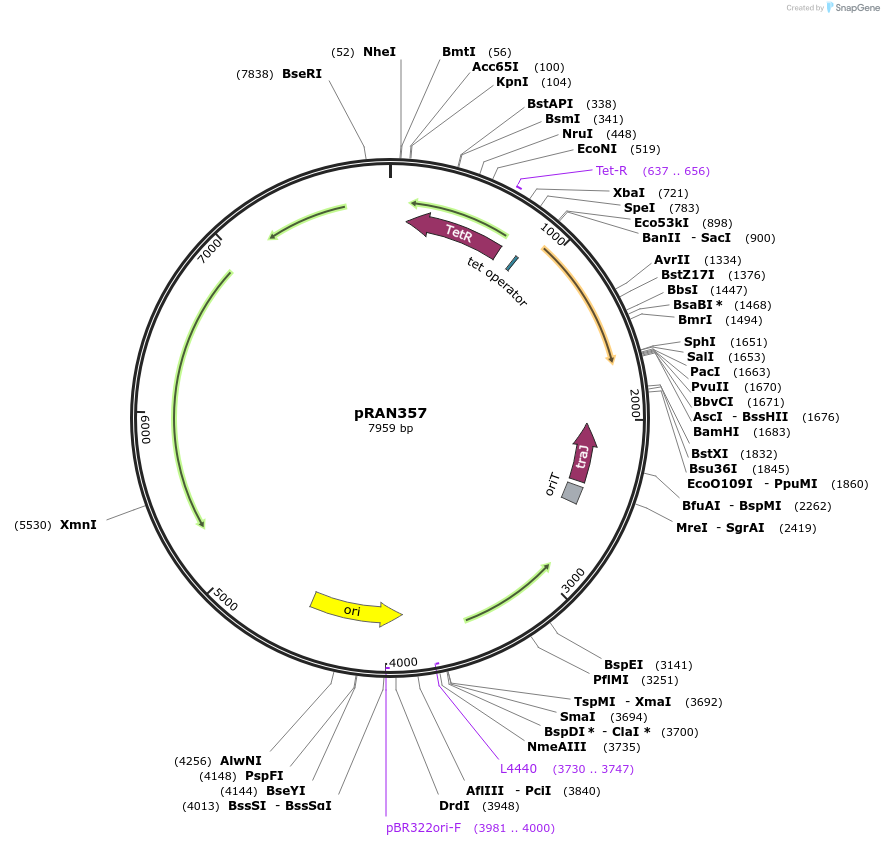
pRAN357
Plasmid#120898PurposeE. coli-C. difficile shuttle vector: tetracycline-inducible promoter (Ptet) driving expression of cyan fluorescent protein (CfpOpt), multiple-cloning site for fusion to N-terminus of target proteinDepositorInsertcyan fluorescent protein, codon-optimized for low GC organisms, multiple-cloning site for protein fusion
ExpressionBacterialPromoterPtetAvailable SinceFeb. 12, 2019AvailabilityAcademic Institutions and Nonprofits only -
pChlr-1SR
Plasmid#52142PurposeEncodes module 1 with amino acids 12S-16R for recognition of nt 8CDepositorInsertPumilio homology domain amino acids 25-60 (PUM1 Human)
ExpressionBacterialMutationChanged 16Q to 16R in Module 1Available SinceMay 5, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-1NQ
Plasmid#52141PurposeEncodes module 1 with amino acids 12N-16Q for recognition of nt 8UDepositorInsertPumilio homology domain amino acids 25-60 (PUM1 Human)
ExpressionBacterialMutationChanged 12S to 12N in Module 1Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-2CQ
Plasmid#52143PurposeEncodes module 2 with amino acids 12C-16Q for recognition of nt 7ADepositorInsertPumilio homology domain amino acids 61-96 (PUM1 Human)
ExpressionBacterialMutationChanged 12N to 12C in Module 2Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-2SR
Plasmid#52146PurposeEncodes module 2 with amino acids 12S-16R for recognition of nt 7CDepositorInsertPumilio homology domain amino acids 61-96 (PUM1 Human)
ExpressionBacterialMutationChanged 12N to 12S and 16Q to 16R in Module 2Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-3NQ
Plasmid#52149PurposeEncodes module 3 with amino acids 12N-16Q for recognition of nt 6UDepositorInsertPumilio homology domain amino acids 97-132 (PUM1 Human)
ExpressionBacterialMutationChanged 12C to 12N in Module 3Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-3SR
Plasmid#52150PurposeEncodes module 3 with amino acids 12S-16R for recognition of nt 6CDepositorInsertPumilio homology domain amino acids 97-132 (PUM1 Human)
ExpressionBacterialMutationChanged 12C to 12S and 16Q to 16R in Module 3Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-4CQ
Plasmid#52151PurposeEncodes module 4 with amino acids 12C-16Q for recognition of nt 5ADepositorInsertPumilio homology domain amino acids 133-168 (PUM1 Human)
ExpressionBacterialMutationChanged 12N to 12C in Module 4Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-4SYE
Plasmid#52156PurposeEncodes module 4 with amino acids 12S-13Y-16E for recognition of nt 5GDepositorInsertD43951 (PUM1 Human)
ExpressionBacterialMutationChanged 12N to 12S, 13H to 13Y, and 16Q to 16E in…Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only -
pChlr-4NYQ
Plasmid#52157PurposeEncodes module 4 with amino acids 12N-13Y-16Q for recognition of nt 5UDepositorInsertPumilio homology domain amino acids 133-168 (PUM1 Human)
ExpressionBacterialMutationChanged 13H to 13Y in Module 4Available SinceApril 28, 2014AvailabilityAcademic Institutions and Nonprofits only