We narrowed to 8,755 results for: SARS
-
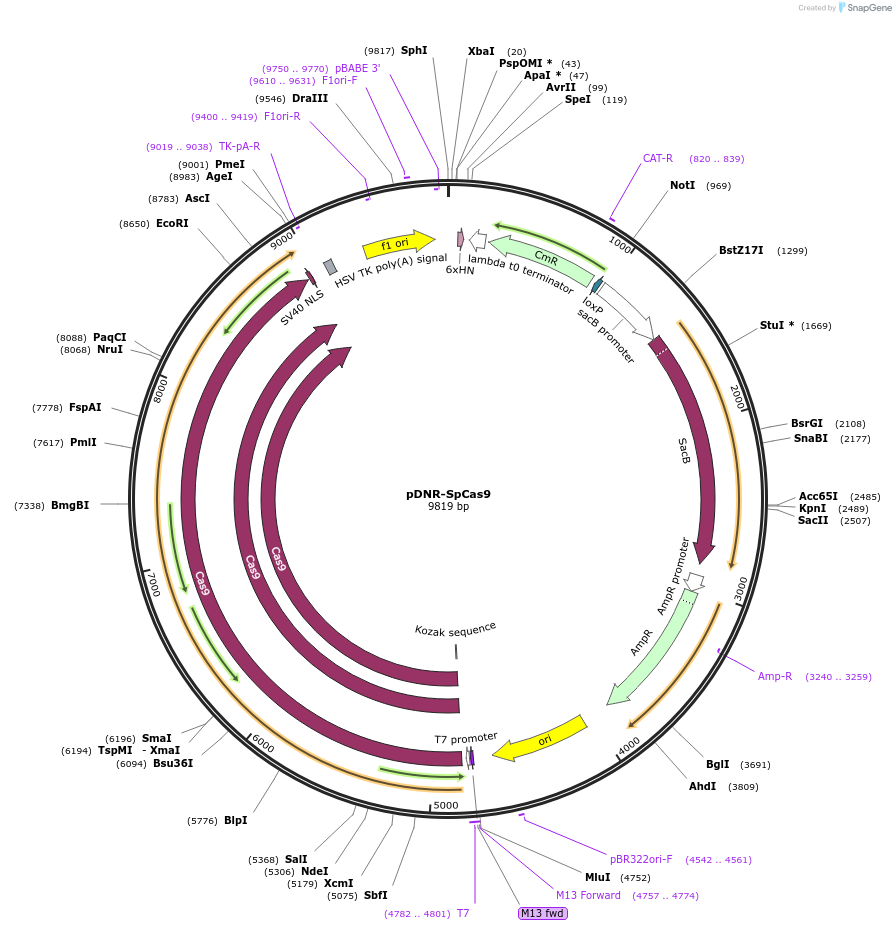
Plasmid#80425PurposeUsed for in vitro transcription of SpCas9 mRNA.DepositorInsertSpCas9
UseCRISPRTagsnoneMutationcodon optimized for humanPromoterT7Available SinceAug. 15, 2016AvailabilityAcademic Institutions and Nonprofits only -
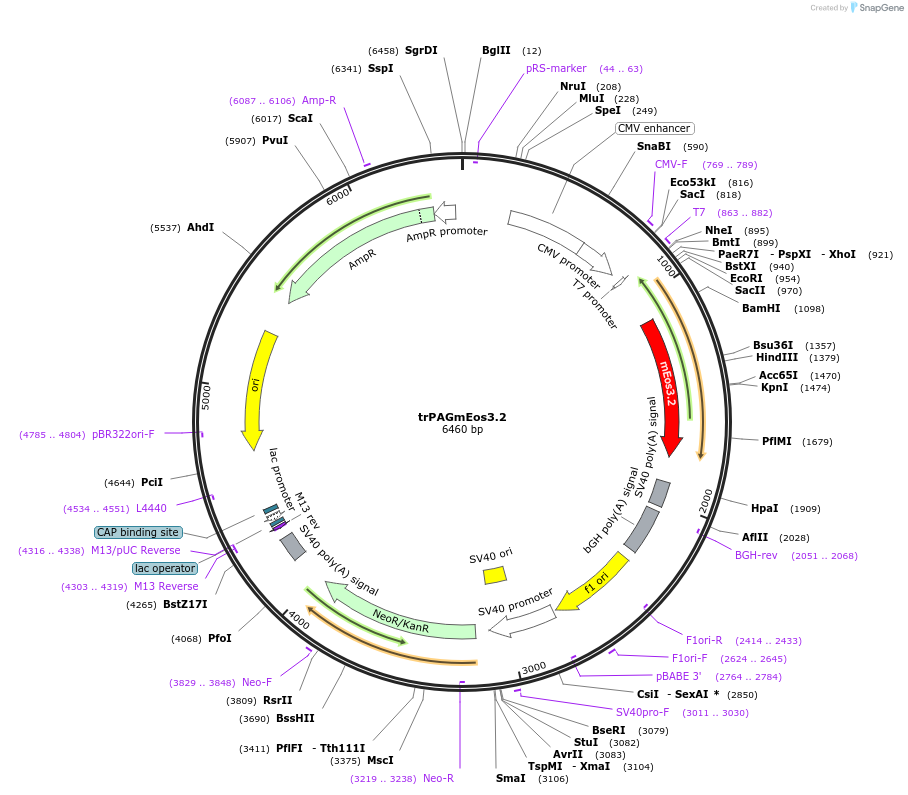
trPAGmEos3.2
Plasmid#203752PurposeEncodes the transmembrane domain from PAG/CSK, including its 2 palmitoylation sites, with mEos3.2 fused on the C terminus. Used as a fluorescent plasma membrane probe.DepositorAvailable SinceAug. 7, 2023AvailabilityAcademic Institutions and Nonprofits only -
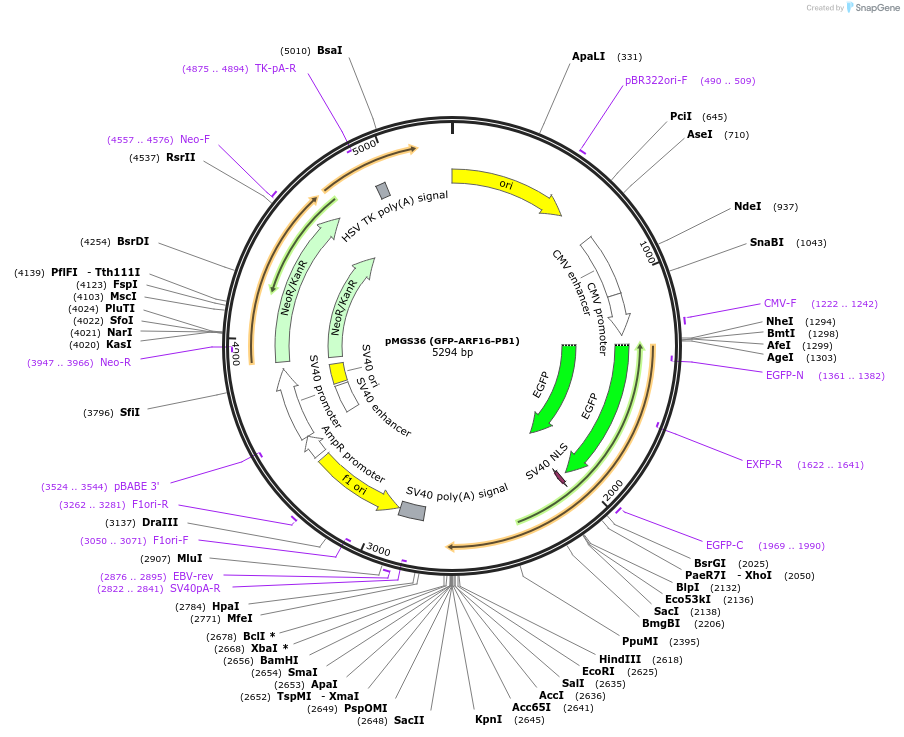
pMGS36 (GFP-ARF16-PB1)
Plasmid#126581PurposeConstruct to express GFP-ARF16-PB1 under the control of the CMV promoterDepositorInsertOsARF16-PB1 domain
TagsGFPExpressionMammalianAvailable SinceJune 10, 2019AvailabilityAcademic Institutions and Nonprofits only -
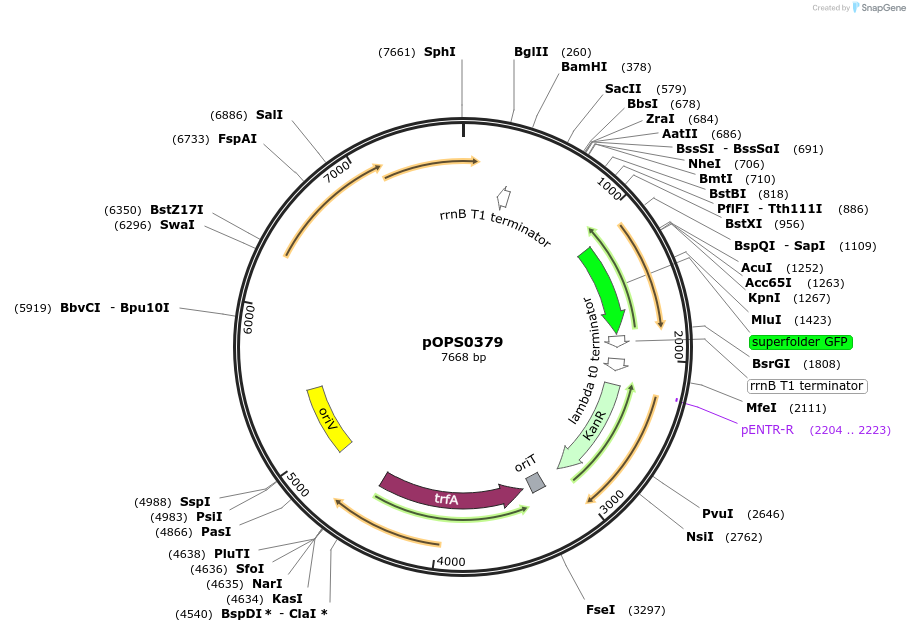
pOPS0379
Plasmid#115510PurposeSuitable for experiments in environments where there is no antibiotic selection present. Symbiosis-induced nifH promoter from Rhizobium leguminosarum biovar vicia 3841 (Rlv3841pNifH) expressing sfGFPDepositorInsertRlv3841pNifH(pOGG082) sfGFP (pOGG037) and T-pharma (pOGG003) assembled in pOGG026
ExpressionBacterialMutationDomesticated for Golden Gate cloningAvailable SinceOct. 3, 2018AvailabilityAcademic Institutions and Nonprofits only -
RCAS-K-ras
Plasmid#11549DepositorInsertv-Ki-ras2 Kirsten rat sarcoma viral oncogene homolog (Kras Mouse)
UseRetroviralMutationG12DAvailable SinceApril 5, 2006AvailabilityAcademic Institutions and Nonprofits only -
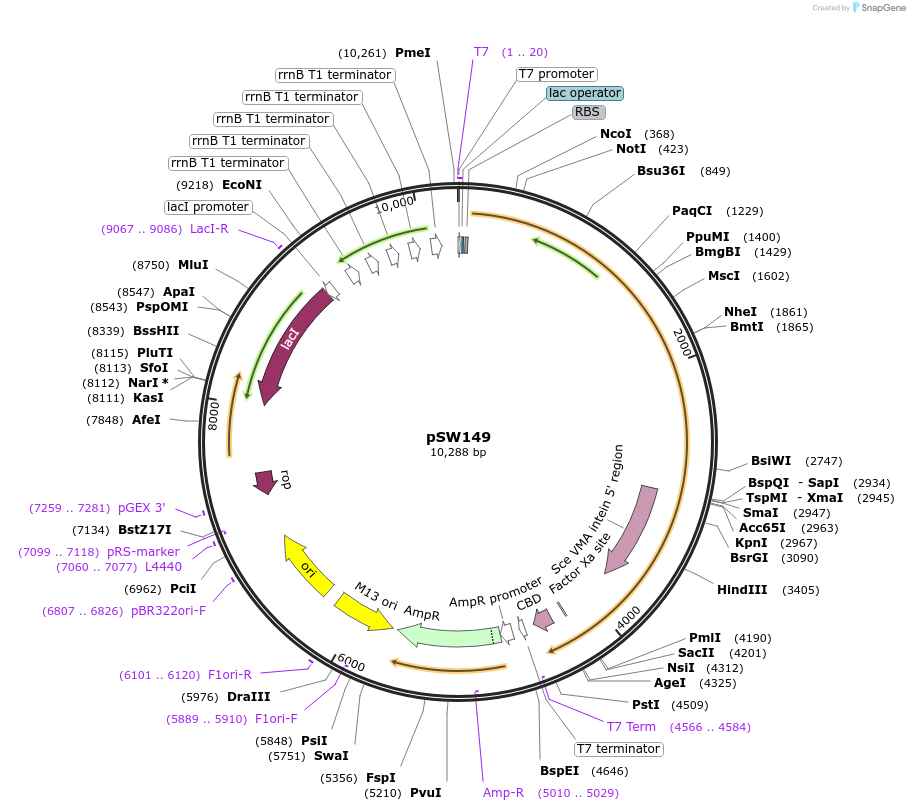
pSW149
Plasmid#122248PurposepTYB2 containing TIF4631 gene, expresses yeast eIF4G1 as an intein-chitin-binding domain fusion for purification of untagged full-length protein, with or without a fluorescent label on the C-terminus.DepositorInsertTIF4631 (TIF4631 Budding Yeast)
TagsIntein-Chitin-binding domainExpressionBacterialPromoterT7Available SinceJuly 22, 2019AvailabilityAcademic Institutions and Nonprofits only -
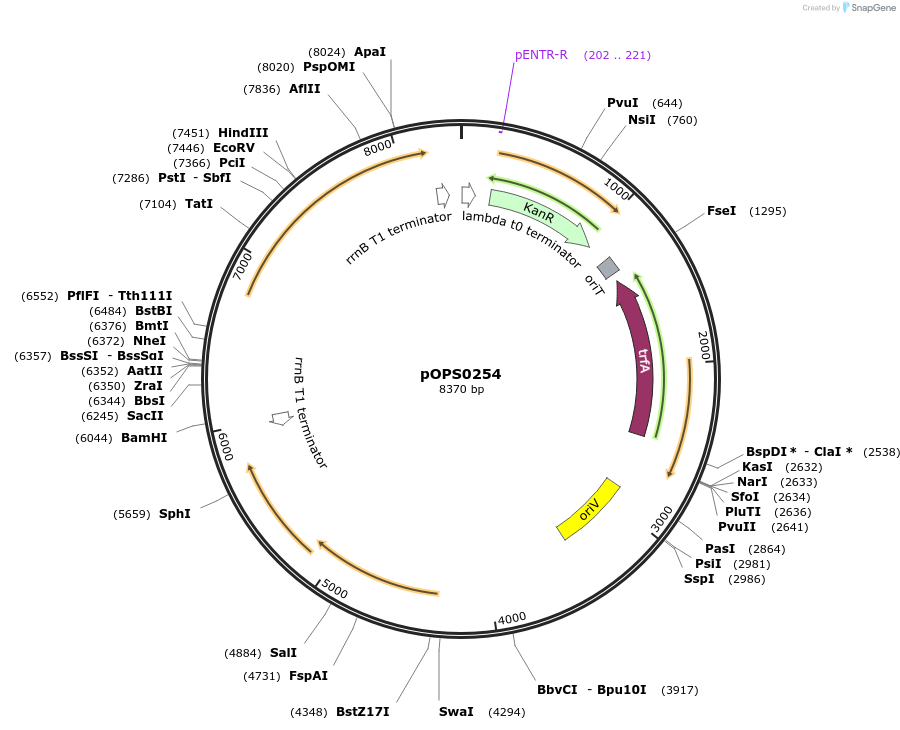
pOPS0254
Plasmid#115506PurposeReporter plasmid suitable for experiments in environments where there is no antibiotic selection present. Symbiosis-induced nifH promoter from Rhizobium leguminosarum biovar vicia 3841 (Rlv3841pNifH) expressing celBDepositorInsertRlv3841pNifH pOGG082, celB pOGG050, T-pharma pOGG003 assembled in pOGG026
ExpressionBacterialMutationDomesticated for Golden Gate cloningAvailable SinceOct. 3, 2018AvailabilityAcademic Institutions and Nonprofits only -
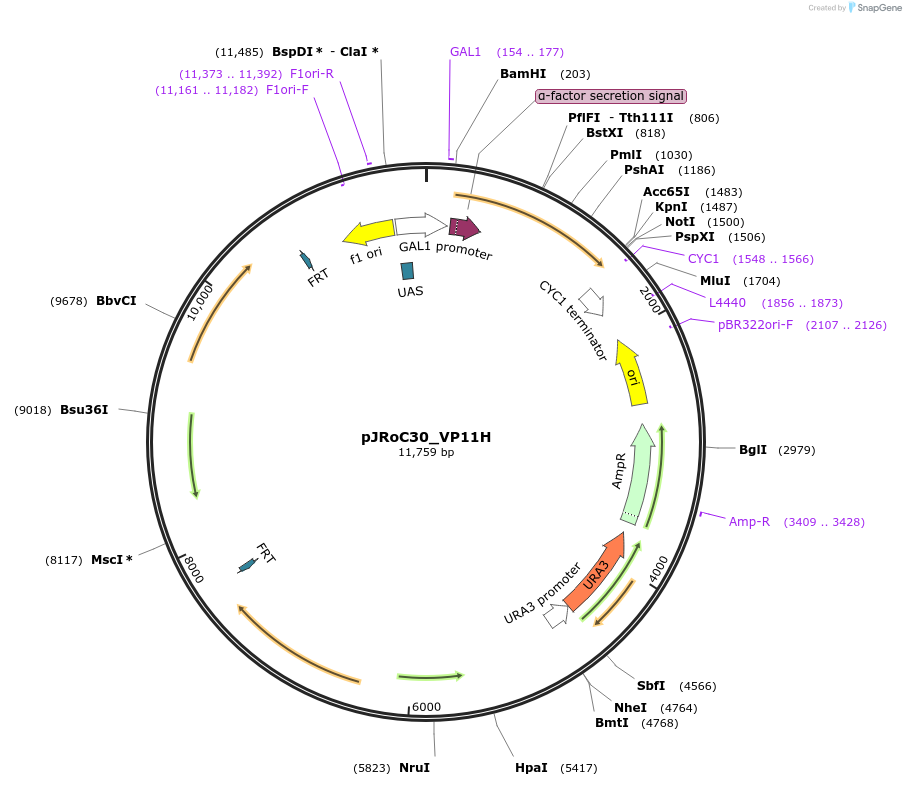
pJRoC30_VP11H
Plasmid#188063PurposePROSS-designed versatile peroxidase 11HDepositorInsertPROSS-designed versatile peroxidase from Pleurotus ostreatus
TagsS. cerevisiae native α-factor prepro-leader signa…ExpressionYeastMutation38 PROSS mutations, described in publicationAvailable SinceAug. 23, 2022AvailabilityAcademic Institutions and Nonprofits only -
pMAL_dstβL
Plasmid#73805PurposePlasmid for selection for insertion and homodimerizationDepositorInsertsCAT Marker
ToxR
beta lactamase
spectomycin resistance
CLS TM domain
TagsPart Of the dstbl chiemra, Part of the dsTbL chim…ExpressionBacterialPromoterNaN and ToxR promotorAvailable SinceMarch 14, 2016AvailabilityAcademic Institutions and Nonprofits only -
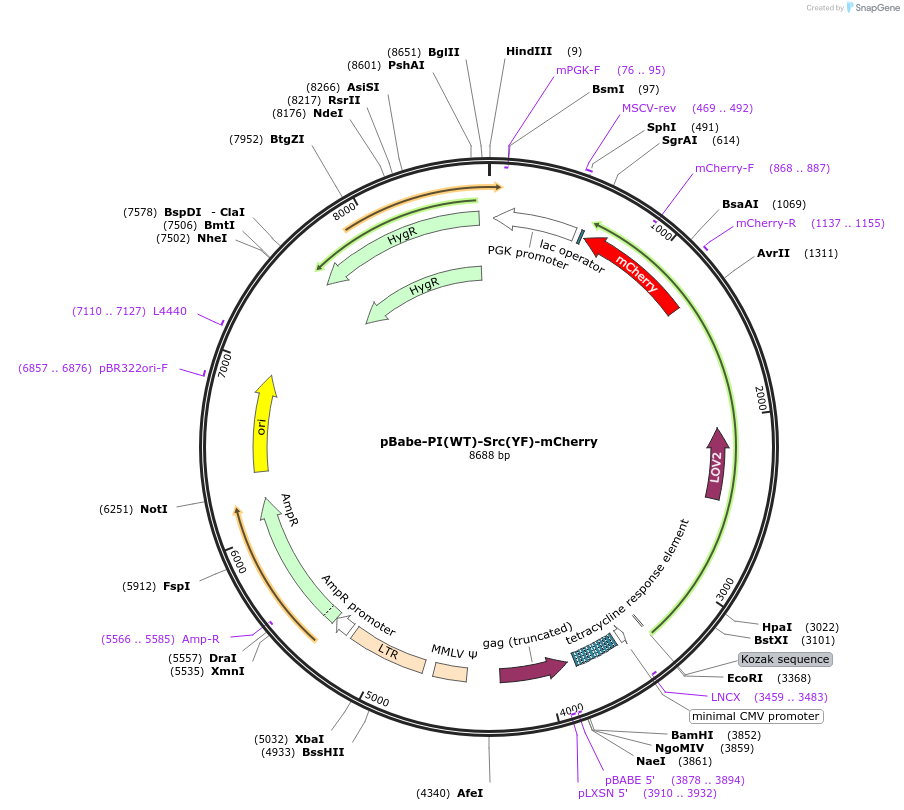
pBabe-PI(WT)-Src(YF)-mCherry
Plasmid#87357PurposePI(WT): Photo-inhibitable/blue light sensitive; YF: Y535F const. active mutationDepositorInsertsTagsmCherryExpressionBacterial and MammalianMutationY535FPromoterCMVAvailable SinceApril 27, 2017AvailabilityAcademic Institutions and Nonprofits only -
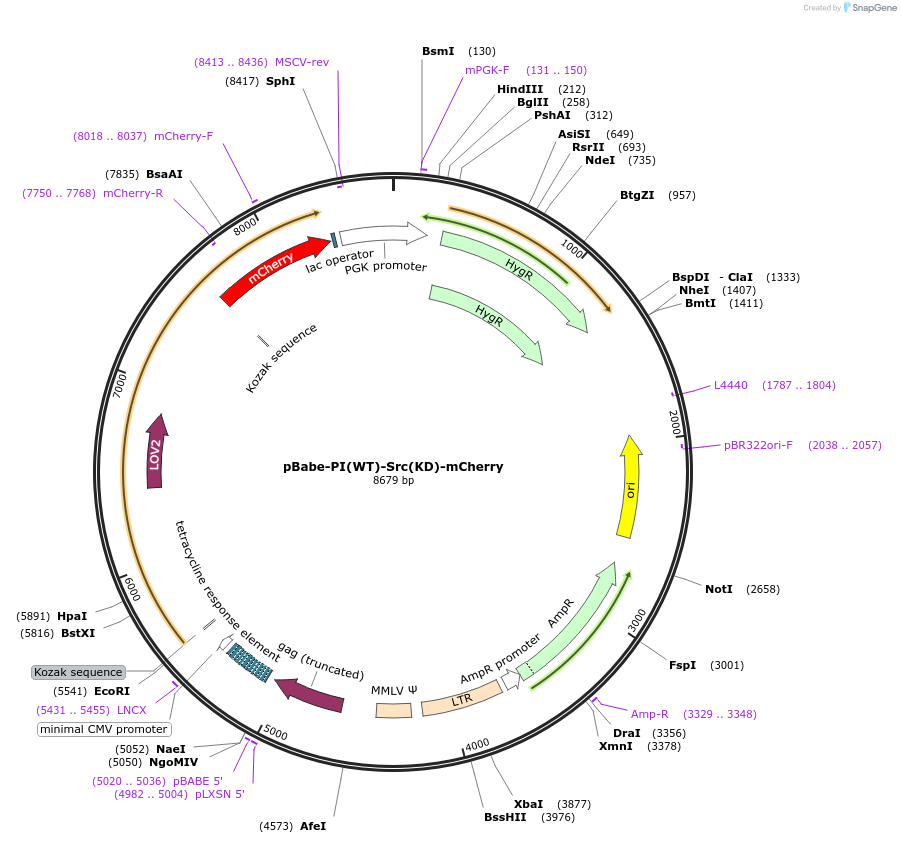
pBabe-PI(WT)-Src(KD)-mCherry
Plasmid#87358PurposePI(WT): Photo-inhibitable/blue light sensitive; kinase dead Src mutationDepositorInsertsTagsmCherryExpressionBacterial and MammalianMutationD388RPromoterCMVAvailable SinceApril 27, 2017AvailabilityAcademic Institutions and Nonprofits only -
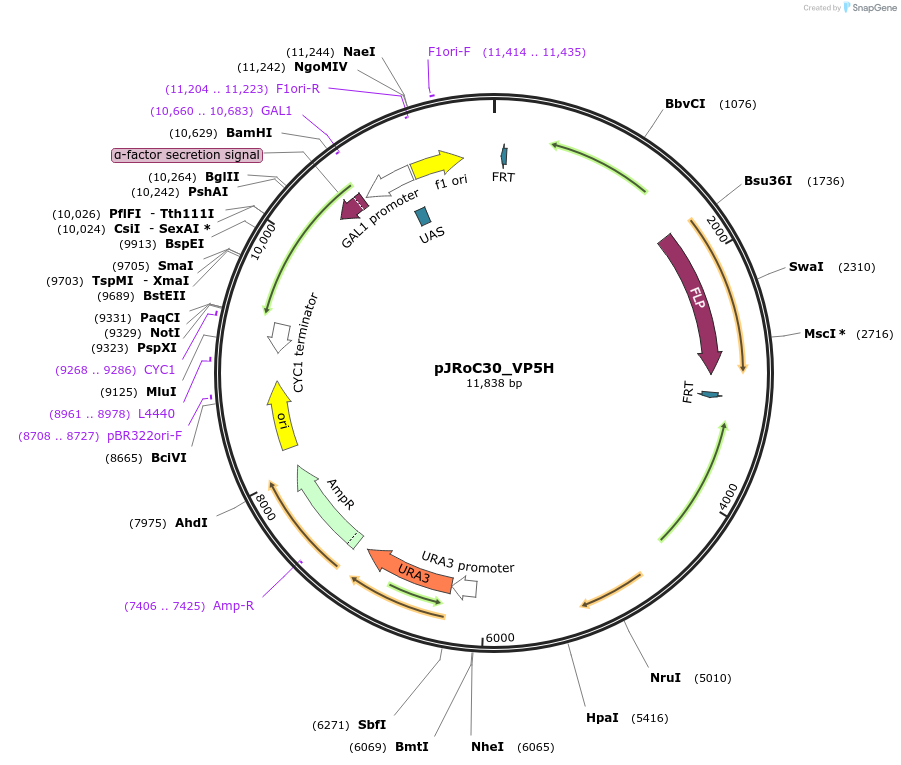
pJRoC30_VP5H
Plasmid#188061PurposePROSS-designed versatile peroxidase 5HDepositorInsertPROSS-designed versatile peroxidase from Pleurotus ostreatus
TagsS. cerevisiae native α-factor prepro-leader signa…ExpressionYeastMutation43 PROSS mutations, described in publicationAvailable SinceAug. 23, 2022AvailabilityAcademic Institutions and Nonprofits only -
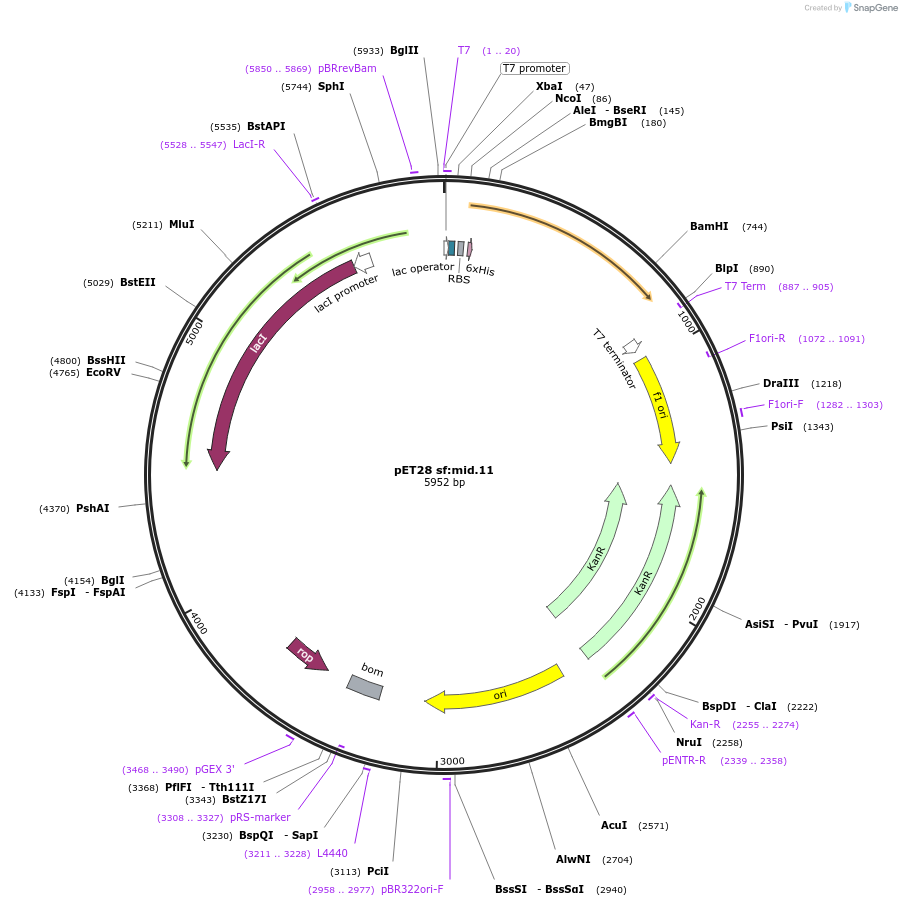
pET28 sf:mid.11
Plasmid#191921PurposeEncodes a GFP design, sf:mid.11DepositorInsertsf:mid.11
TagsHis TagExpressionBacterialMutationQ69M Y145F T167I L220V V224IPromoterT7Available SinceDec. 15, 2022AvailabilityAcademic Institutions and Nonprofits only -
pET28 sf:mid.7
Plasmid#191933PurposeEncodes a GFP design, sf:mid.7DepositorInsertsf:mid.7
TagsHis TagExpressionBacterialMutationS72T Y145F V224IPromoterT7Available SinceDec. 15, 2022AvailabilityAcademic Institutions and Nonprofits only -
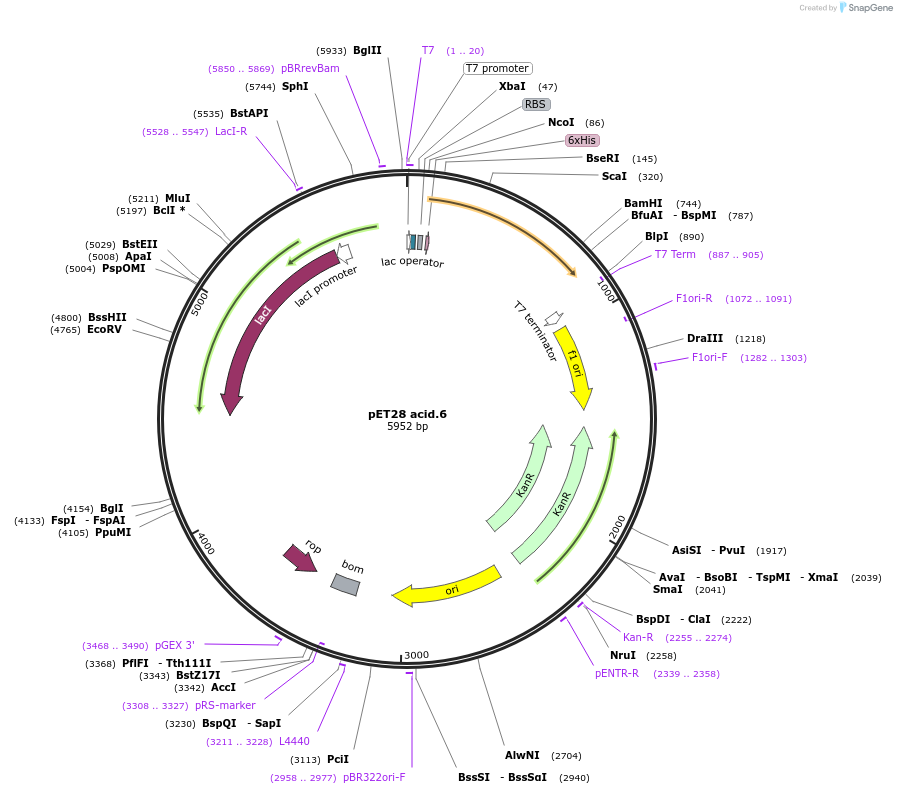
pET28 acid.6
Plasmid#191912PurposeEncodes a GFP design, acid.6DepositorInsertacid.6
TagsHis TagExpressionBacterialMutationQ69L S72A Y145M T167V H181LPromoterT7Available SinceDec. 14, 2022AvailabilityAcademic Institutions and Nonprofits only -
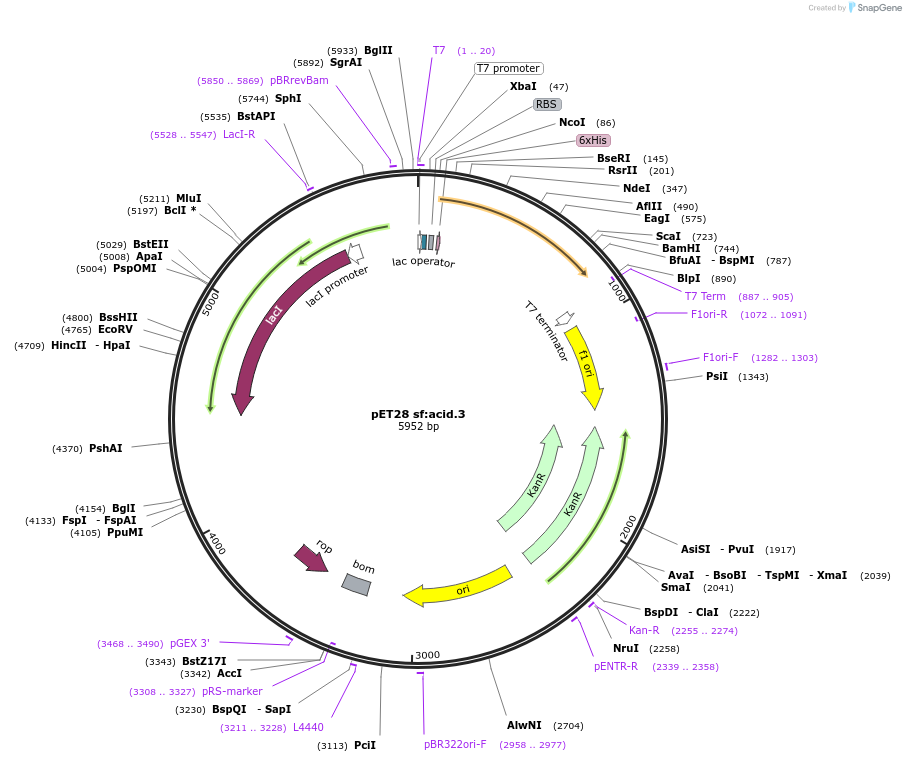
pET28 sf:acid.3
Plasmid#191926PurposeEncodes a GFP design, sf:acid.3DepositorInsertsf:acid.3
TagsHis TagExpressionBacterialMutationT65S Q69L S72A T108V Y145M V224IPromoterT7Available SinceDec. 5, 2022AvailabilityAcademic Institutions and Nonprofits only -
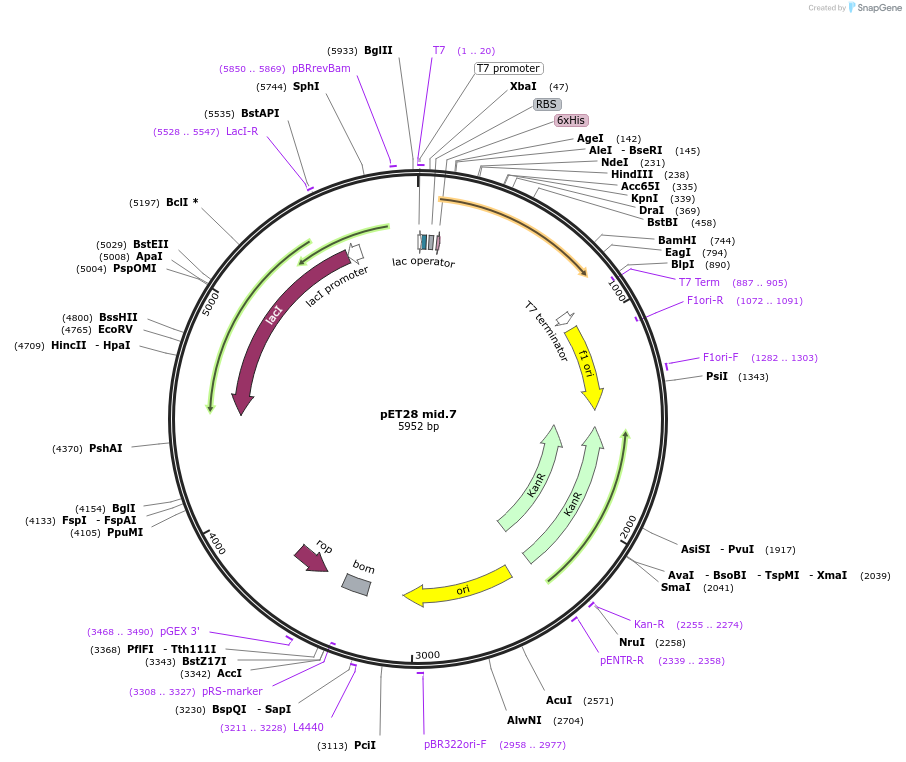
pET28 mid.7
Plasmid#191900PurposeEncodes a GFP design, mid.7DepositorInsertmid.7
TagsHis TagExpressionBacterialMutationS72T Y145F V224IPromoterT7Available SinceDec. 5, 2022AvailabilityAcademic Institutions and Nonprofits only -
pGL3-24p3/LCN2-282-LUC
Plasmid#46865Purposemurine 24p3 minimal promoter as a luciferase reporterDepositorInsert24p3/lipocalin 2 proximal promoter
UseLuciferasePromoter24p3 minimal promoterAvailable SinceSept. 13, 2013AvailabilityAcademic Institutions and Nonprofits only -
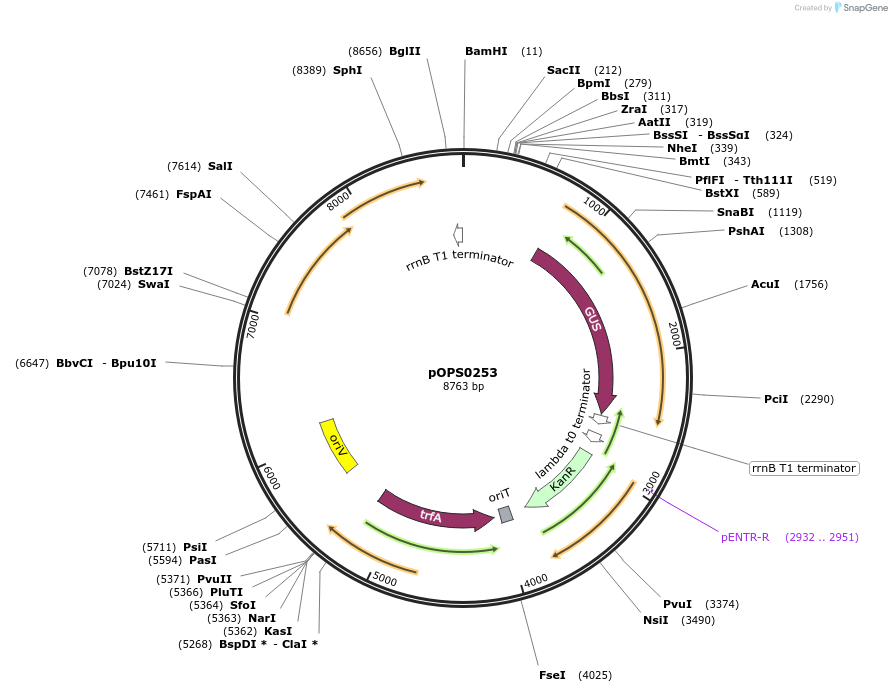
pOPS0253
Plasmid#115505PurposeSuitable for experiments in environments where there is no antibiotic selection present. Symbiosis-induced nifH promoter from Rhizobium leguminosarum biovar vicia 3841 (Rlv3841pNifH) expressing gusA.DepositorInsertRlv3841pNifH pOGG082, gusA pOGG083, T-pharma pOGG003 assembled in pOGG026
ExpressionBacterialMutationDomesticated for Golden Gate cloningAvailable SinceOct. 3, 2018AvailabilityAcademic Institutions and Nonprofits only -
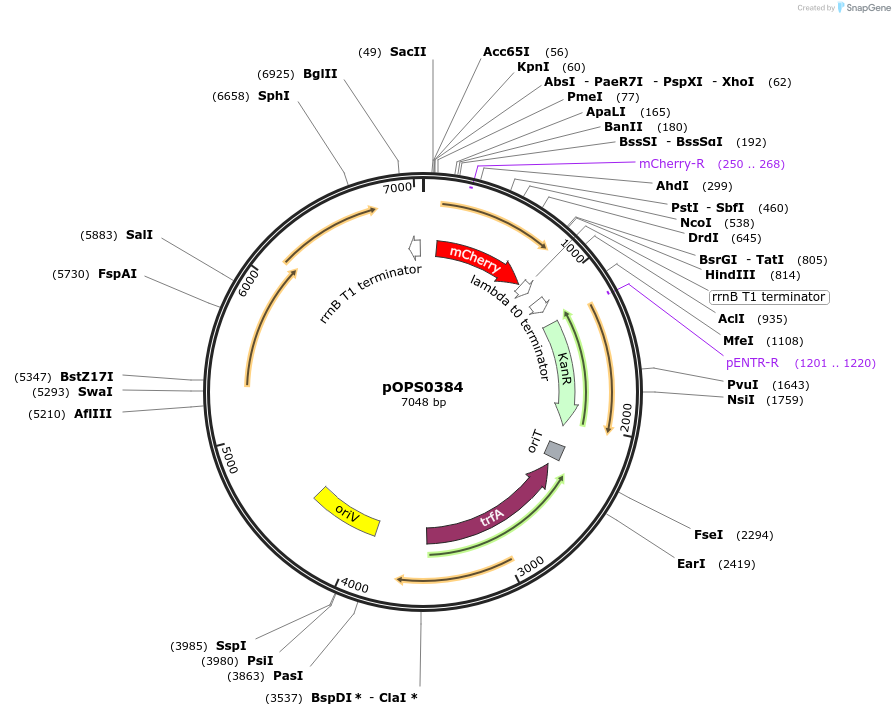
pOPS0384
Plasmid#133234PurposeReporter plasmid for R. leguminosarum suitable for experiments in environments where there is no antibiotic selection present.DepositorInsertconstructed with NoP (pOGG113), mCherry (EC15071) and T-pharma (pOGG003)
ExpressionBacterialMutationDomesticated for Golden Gate cloningAvailable SinceApril 8, 2020AvailabilityAcademic Institutions and Nonprofits only