We narrowed to 61,648 results for: Son
-
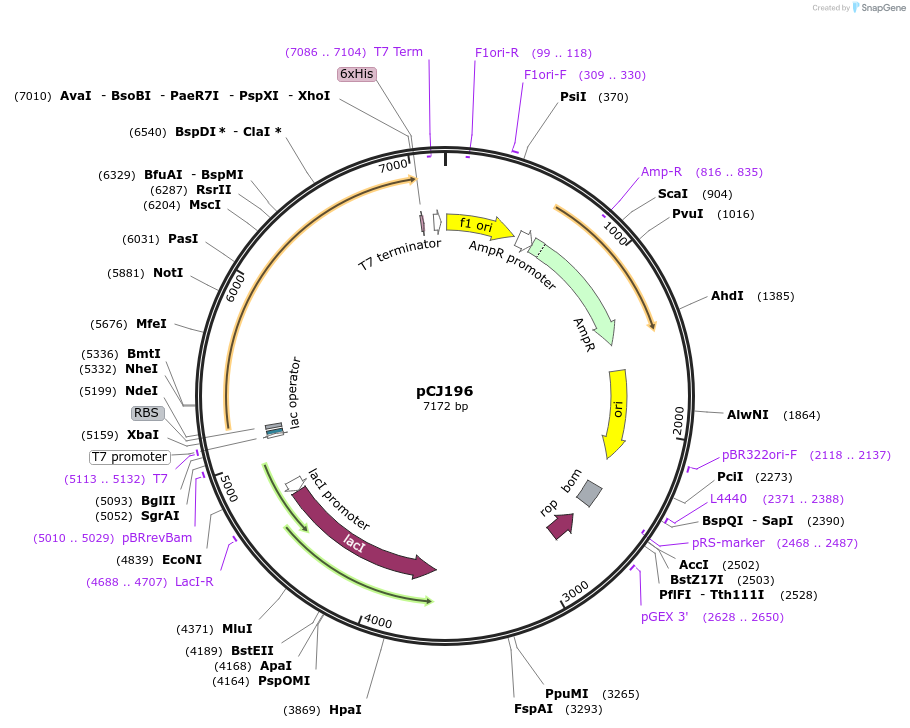
Plasmid#162669PurposepET-21b(+) based plasmid for expression of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating catalytic mutation, S225ADepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1)
TagsHisExpressionBacterialMutationS225A; codon optimized for expression in E. coli …PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
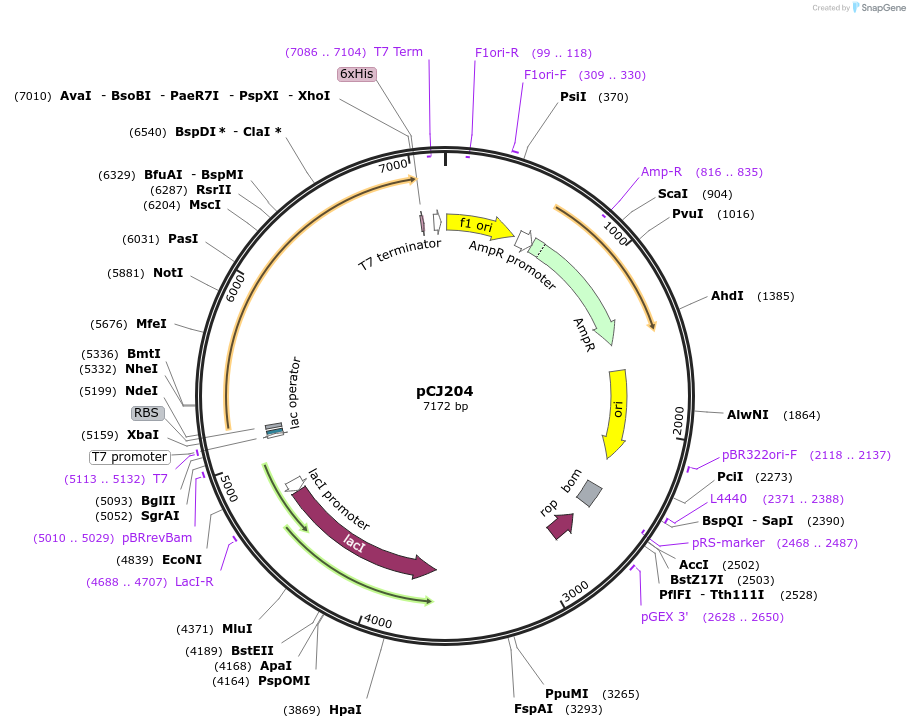
pCJ204
Plasmid#162677PurposepET-21b(+) based plasmid for expression of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating C224H and C529F mutations.DepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1)
TagsHisExpressionBacterialMutationC224H, C529F; codon optimized for expression in E…PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
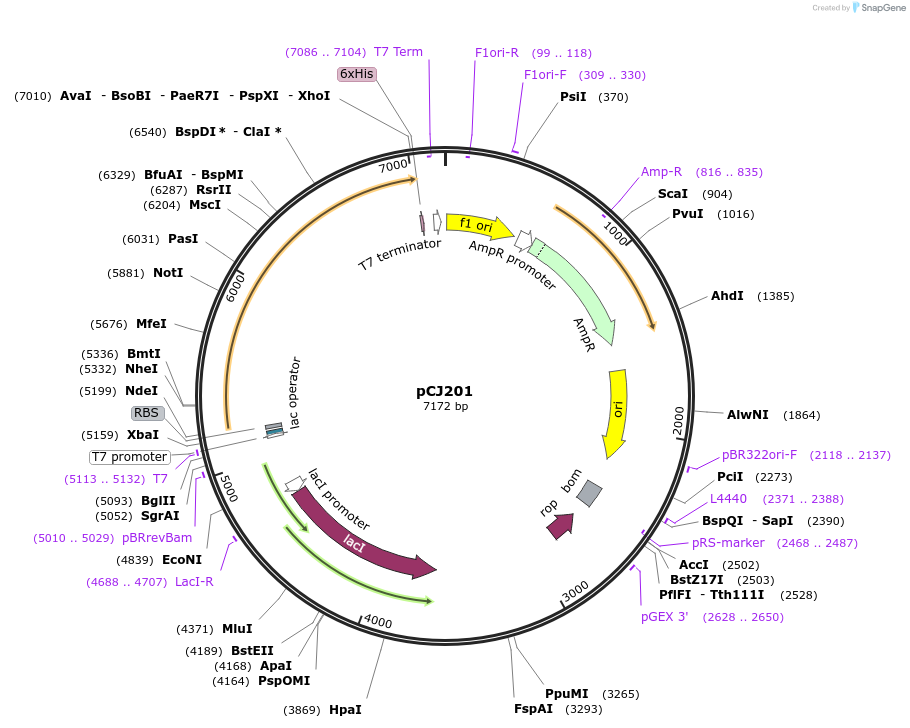
pCJ201
Plasmid#162674PurposepET-21b(+) based plasmid for expression of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating C224W and C529SS mutations.DepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1)
TagsHisExpressionBacterialMutationC224W, C529SS; codon optimized for expression in …PromoterT7/lacAvailable SinceJan. 6, 2021AvailabilityAcademic Institutions and Nonprofits only -
mKusabira Orange-N1
Plasmid#55046PurposeEmpty backbone for fusing gene of interest to N-term of mKODepositorTypeEmpty backboneTagsmKOExpressionMammalianPromoterCMVAvailable SinceSept. 17, 2014AvailabilityAcademic Institutions and Nonprofits only -
CRASP-Seq CBC Backbone
Pooled Library#232068PurposeThe CRASP-Seq backbone is designed to accommodate the cloning of diverse gRNA libraries. It includes a cell barcode (CBC), which enhances its functionality by enabling library preparation.DepositorExpressionMammalianSpeciesHomo sapiensUseCRISPR and LentiviralAvailable SinceOct. 28, 2025AvailabilityAcademic Institutions and Nonprofits only -
CRASP-Seq BE Tiling v1 empty
Pooled Library#232070PurposeTiling library to conduct CRASP-Seq experiments combined with high-throughput base editing for precise and scalable investigations into protein regions that affect splicing regulation.DepositorExpressionMammalianSpeciesHomo sapiensUseCRISPR and LentiviralAvailable SinceOct. 28, 2025AvailabilityAcademic Institutions and Nonprofits only -
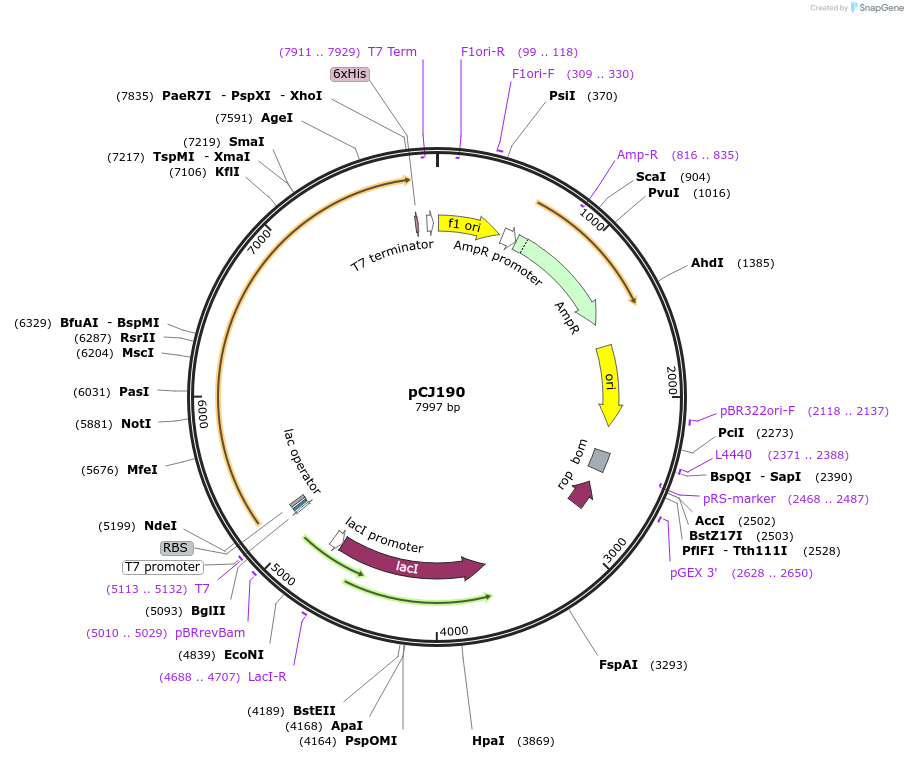
pCJ190
Plasmid#162667PurposepET-21b(+) based plasmid for expresion of MHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by a 12 residue Gly/Ser linker; codon optimized for expression in E. coli K12, with C-terminal His tag.DepositorInsertMHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by a 12 residue Gly/Ser linker
TagsHisExpressionBacterialMutationcodon optimized for expression in E. coli K12PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
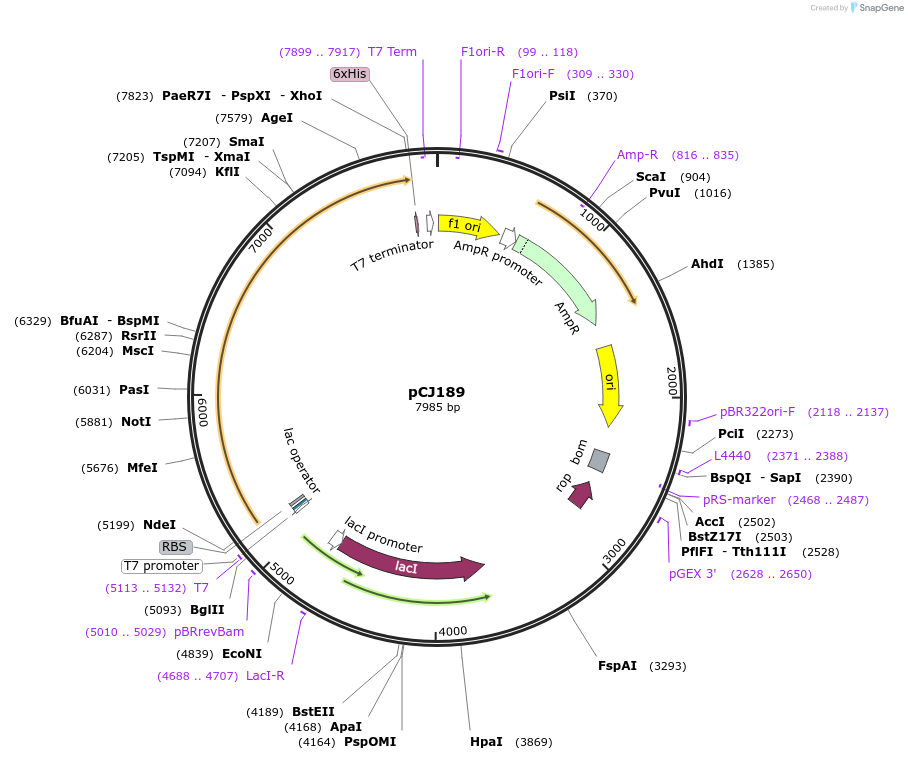
pCJ189
Plasmid#162666PurposepET-21b(+) based plasmid for expresion of MHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by an 8 residue Gly/Ser linker; codon optimized for expression in E. coli K12, with C-terminal His tag.DepositorInsertMHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by an 8 residue Gly/Ser linker
TagsHisExpressionBacterialMutationcodon optimized for expression in E. coli K12PromoterT7/lacAvailable SinceFeb. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
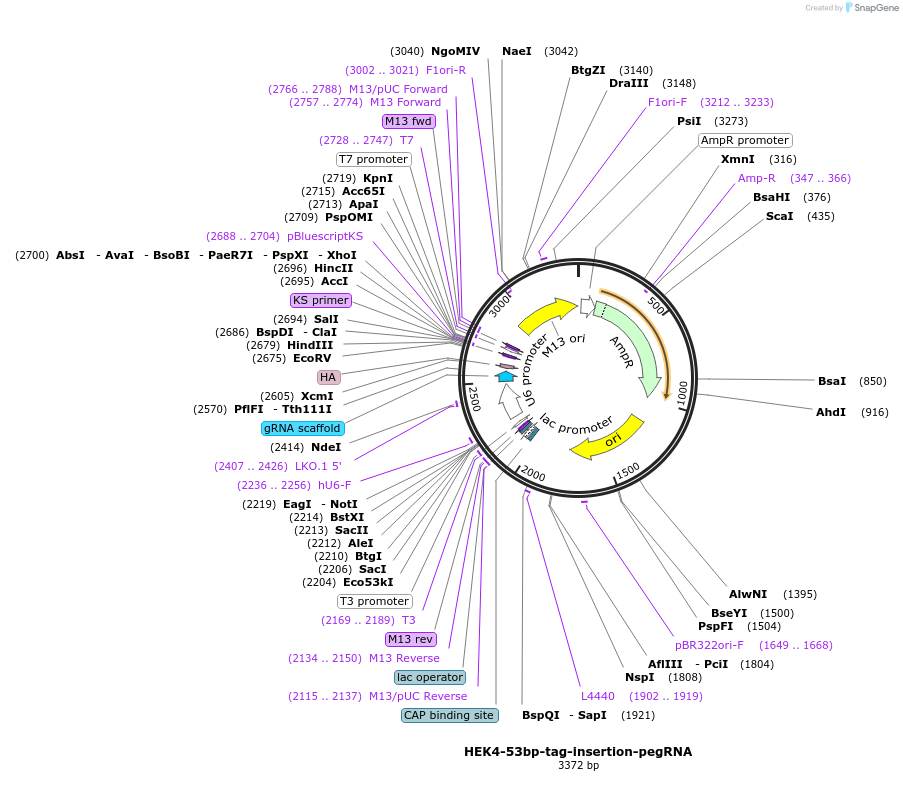
HEK4-53bp-tag-insertion-pegRNA
Plasmid#227673PurposePegRNA for 53 bp insertionDepositorAvailable SinceOct. 22, 2024AvailabilityAcademic Institutions and Nonprofits only -
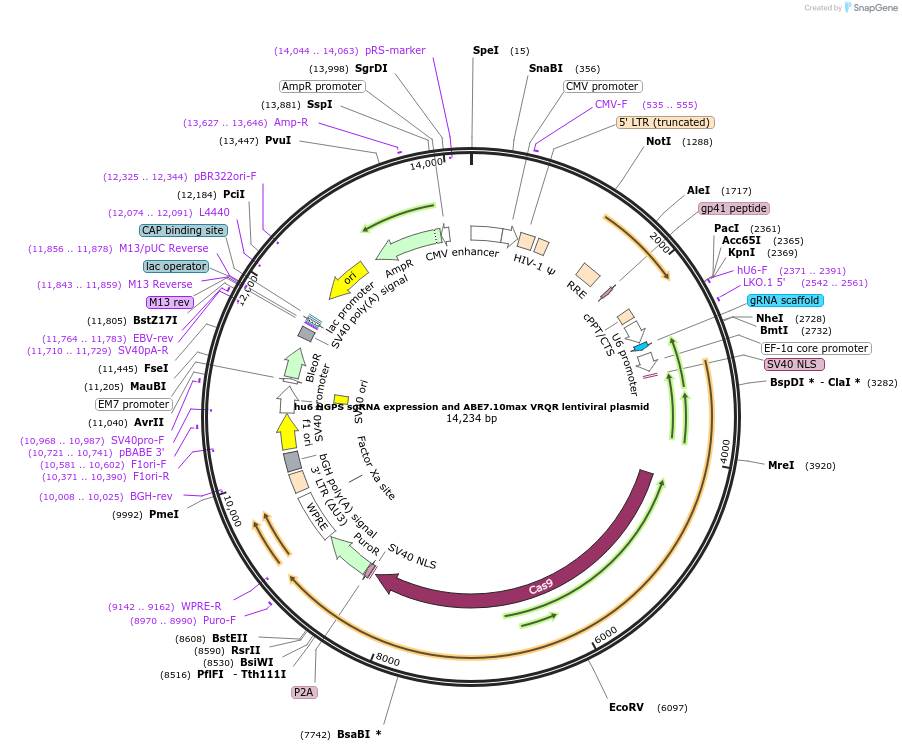
hu6 HGPS sgRNA expression and ABE7.10max VRQR lentiviral plasmid
Plasmid#154429Purposehu6 HGPS sgRNA expression and ABE7.10max VRQR lentiviral plasmidDepositorInserthu6 HGPS sgRNA expression and ABE7.10max VRQR lentiviral plasmid
UseCRISPR and LentiviralExpressionMammalianMutationVRQR point mutations in SpCas9Available SinceFeb. 19, 2021AvailabilityAcademic Institutions and Nonprofits only -
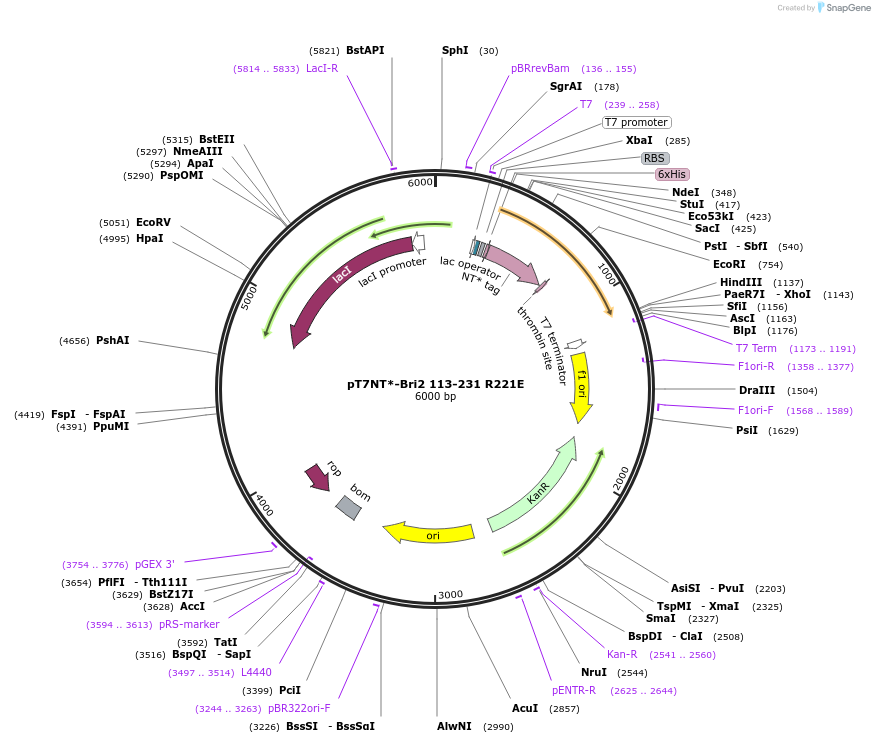
pT7NT*-Bri2 113-231 R221E
Plasmid#138134PurposeExpresseds human Bri2 BRICHOS R221E mutant in E. coliDepositorInserthuman Bri2 BRICHOS R221E (ITM2B Human)
TagsNT* tag derived from spider silk proteinsExpressionBacterialPromoterT7Available SinceMay 28, 2020AvailabilityAcademic Institutions and Nonprofits only -
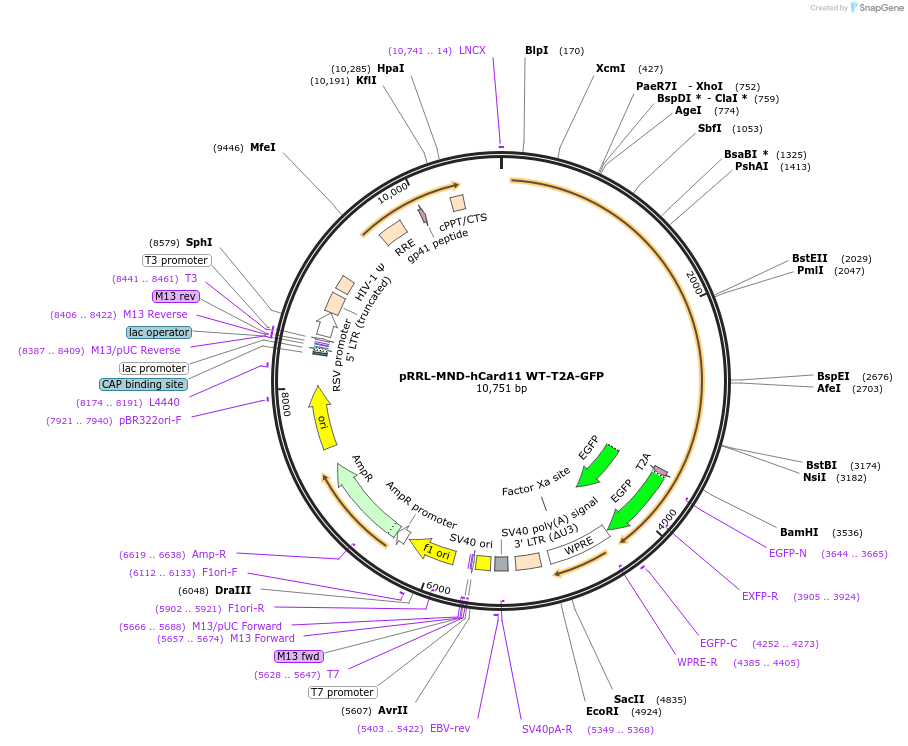
pRRL-MND-hCard11 WT-T2A-GFP
Plasmid#163787PurposeExpression of Wild-type CARD11DepositorAvailable SinceFeb. 1, 2021AvailabilityAcademic Institutions and Nonprofits only -
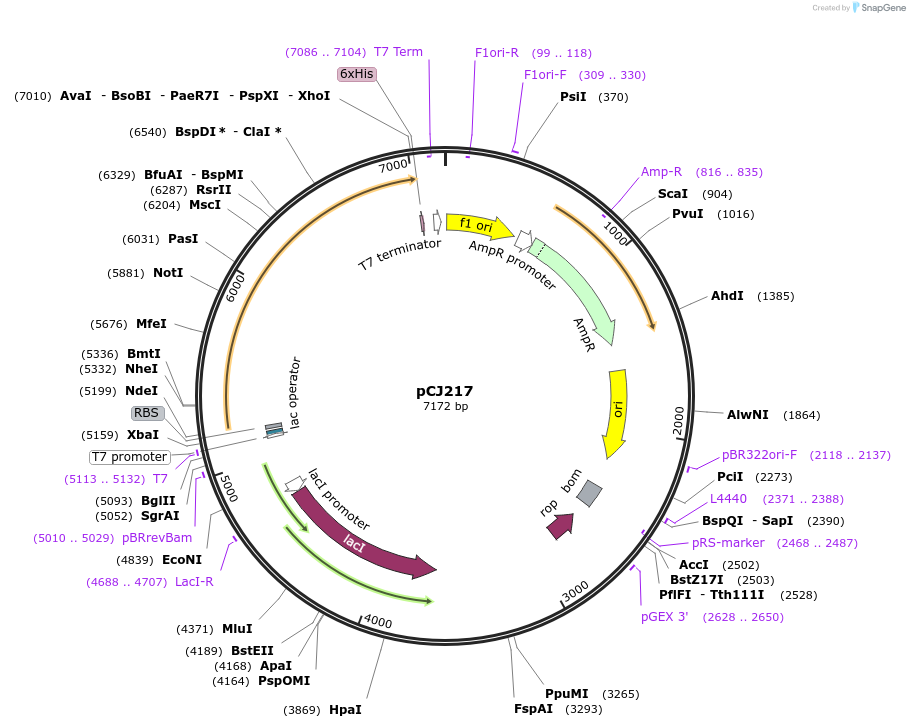
pCJ217
Plasmid#162685PurposepET-21b(+) based plasmid for expression of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating two point mutations, G489C and S530C, to introduce a 6th disulfide bond (from PETase).DepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), with mutations to introduce a 6th disulfide bond from PETase
TagsHisExpressionBacterialMutationG489C, S530C; codon optimized for E. coli K12PromoterT7/lacAvailable SinceJan. 6, 2021AvailabilityAcademic Institutions and Nonprofits only -
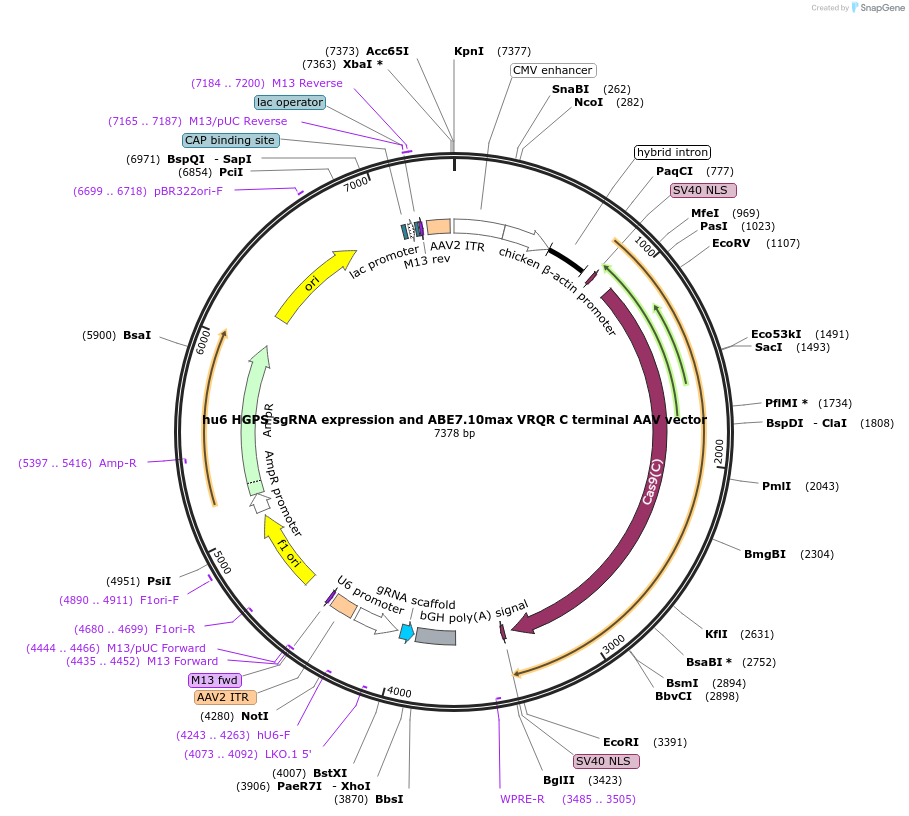
hu6 HGPS sgRNA expression and ABE7.10max VRQR C terminal AAV vector
Plasmid#154430Purposehu6 HGPS sgRNA expression and ABE7.10max VRQR C terminal AAV vectorDepositorInserthu6 HGPS sgRNA expression and ABE7.10max VRQR C terminal AAV vector
UseAAV and CRISPRExpressionMammalianMutationVRQR point mutations in SpCas9Available SinceFeb. 19, 2021AvailabilityAcademic Institutions and Nonprofits only -
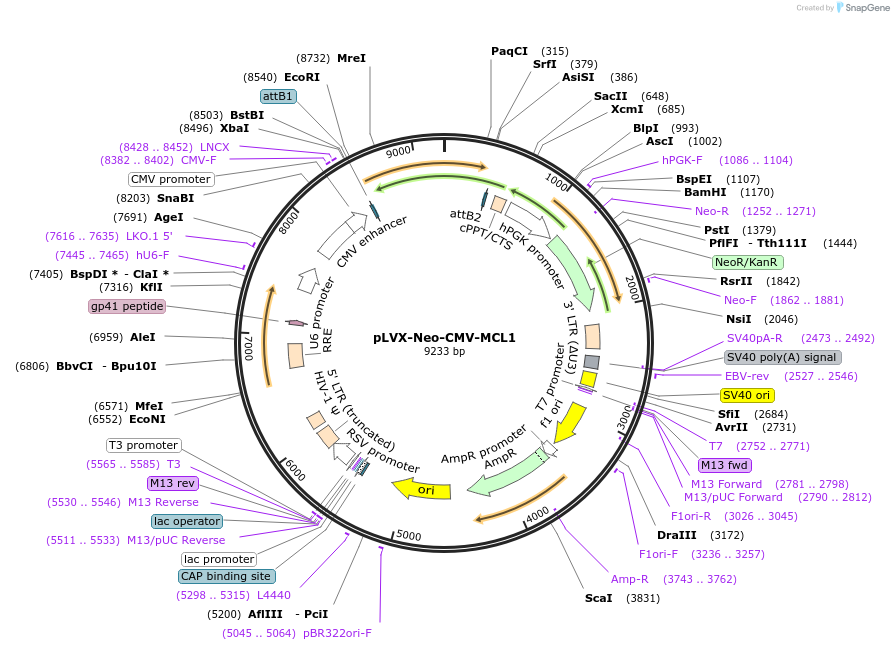
pLVX-Neo-CMV-MCL1
Plasmid#202668PurposeMCL1 overexpression vector with G418 resistanceDepositorInsertMCL1 apoptosis regulator, BCL2 family member (MCL1 Human)
UseLentiviralExpressionMammalianPromoterCMVAvailable SinceMarch 18, 2024AvailabilityAcademic Institutions and Nonprofits only -
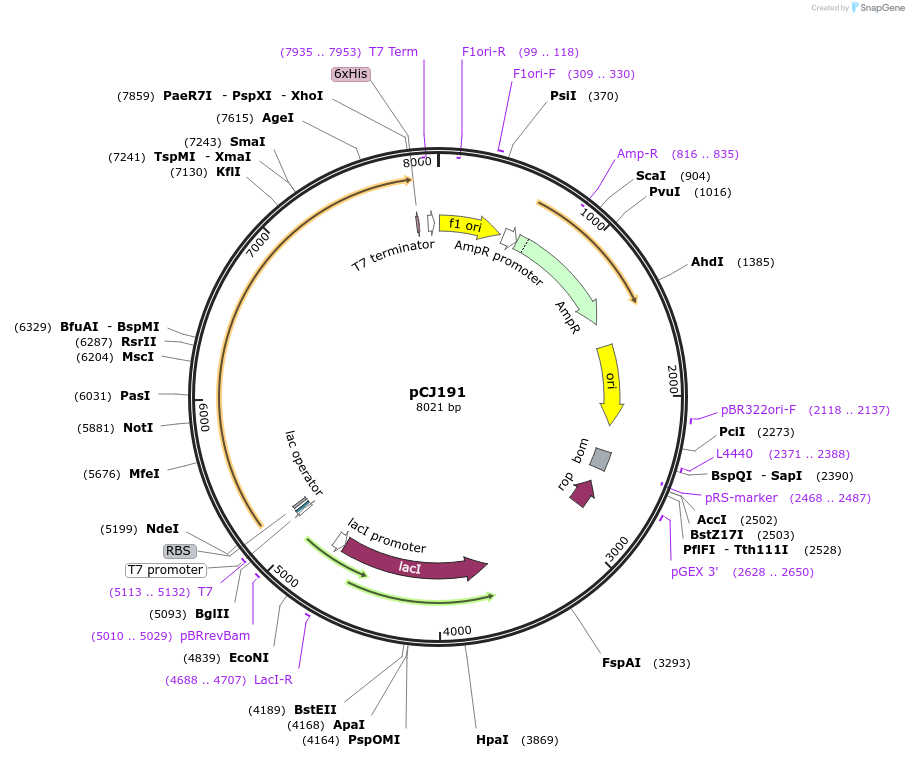
pCJ191
Plasmid#162668PurposepET-21b(+) based plasmid for expresion of MHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by a 20 residue Gly/Ser linker; codon optimized for expression in E. coli K12, with C-terminal His tag.DepositorInsertMHETase (Genbank GAP38911.1) linked to PETase (Genbank GAP38373.1) from Ideonella sakaiensis by a 20 residue Gly/Ser linker
TagsHisExpressionBacterialMutationcodon optimized for expression in E. coli K12PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
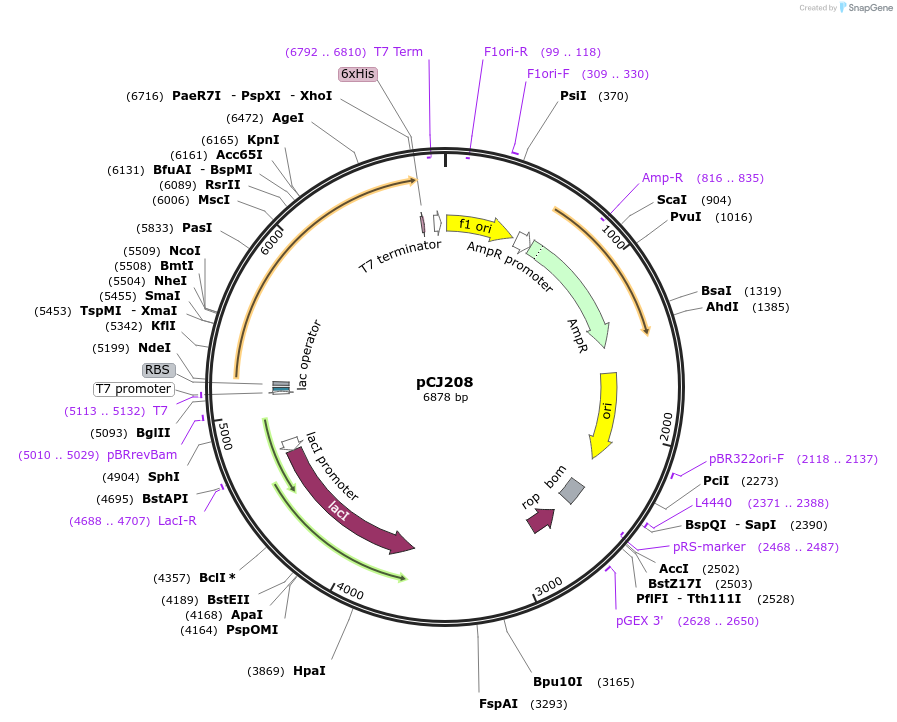
pCJ208
Plasmid#162681PurposeLidded variant of PETase from Ideonella sakaiensis 201-F6 (Genbank GAP38373.1) removing a seven-residue loop of PETase (Trp185:Phe191, PETase numbering) and replacing it with Gly251:Thr472 from MHETase; codon optimized for expression in E. coli K12, with C-terminal His tag.DepositorInsertPETase from Ideonella sakaiensis 201-F6 (Genbank GAP38373.1) incorporating the MHETase lid
TagsHisExpressionBacterialMutationReplacement of W185-F191 with G251-T472 from MHET…PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
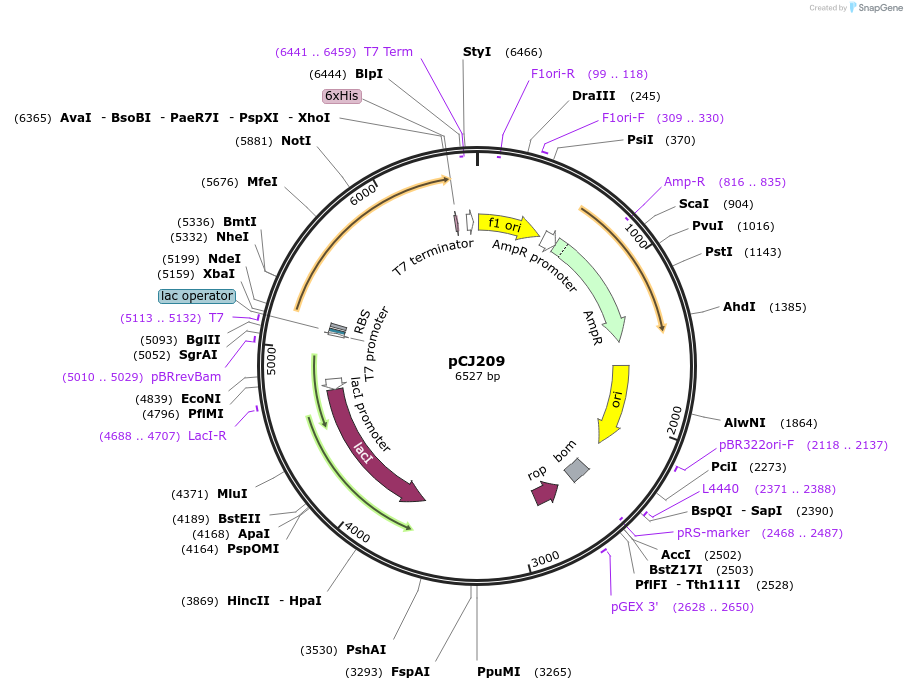
pCJ209
Plasmid#162682PurposeLidless variant of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1) replacing Gly251:Thr472 of MHETase with the 7-residue loop (Trp185:Phe191, PETase numbering) of PETase , codon optimized for expression in E. coli K12, with C-terminal His tag.DepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1) with the lid removed
TagsHisExpressionBacterialMutationReplacement of G251-T472 with W185-F191 from PETa…PromoterT7/lacAvailable SinceJan. 6, 2021AvailabilityAcademic Institutions and Nonprofits only -
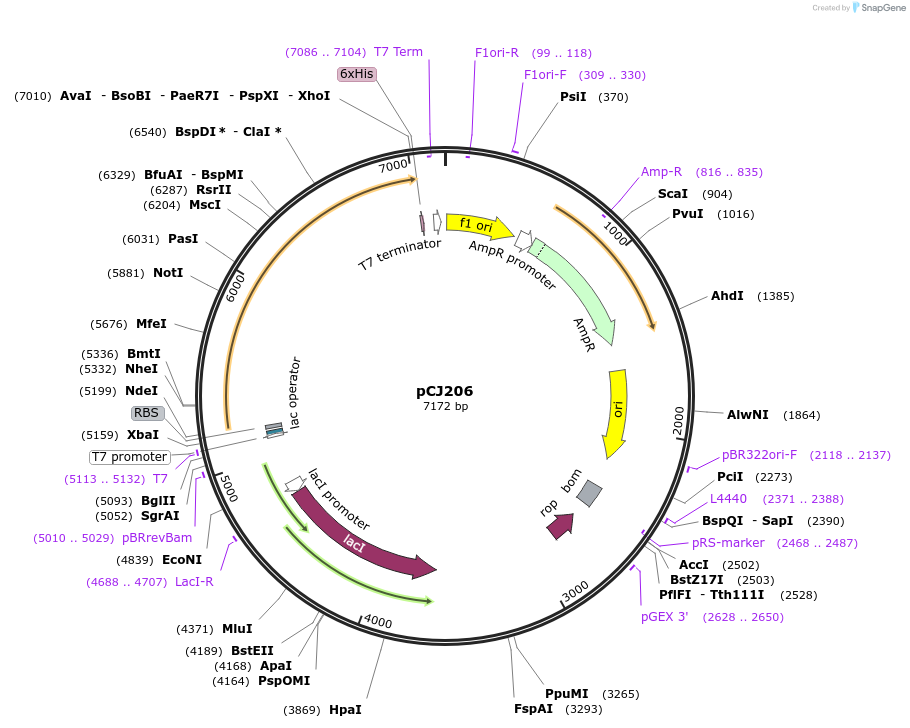
pCJ206
Plasmid#162679PurposepET-21b(+) based plasmid for expression of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), codon optimized for expression in E. coli K12, with C-terminal His tag, incorporating the E226T mutation to the putative lipase box.DepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1), incorporating the E226T mutation to the putative lipase box
TagsHisExpressionBacterialMutationE226T; codon optimized for expression in E. coli …PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
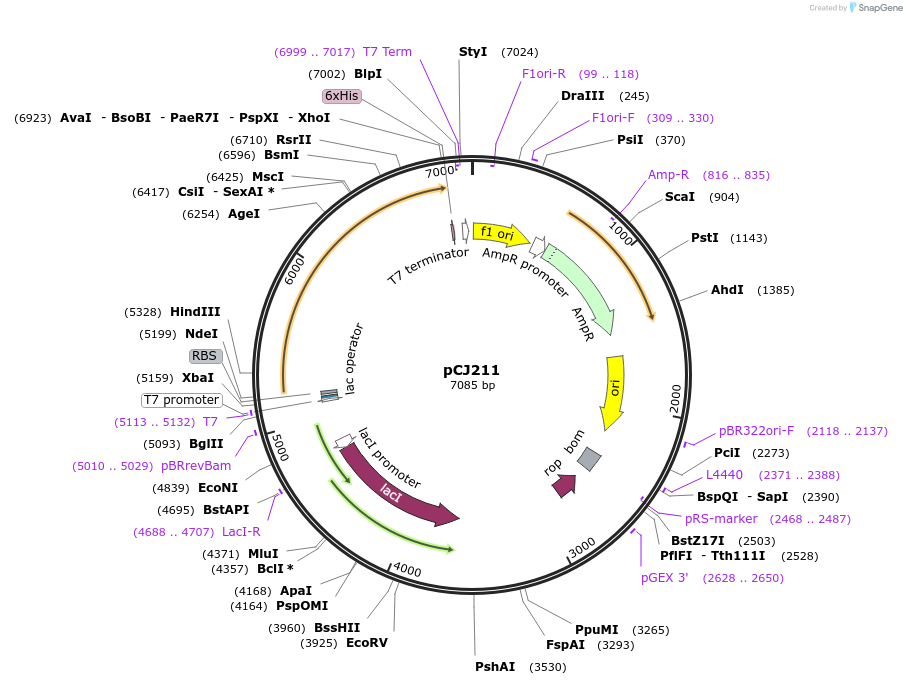
pCJ211
Plasmid#162684PurposepET-21b(+) based plasmid for expression of the putative MHETase from Hydrogenophaga sp. PML113 (Genbank WP_083293388.1) with deletion of the predicted signal peptide (Lys2:Gly19), with C-terminal His tag, codon optimized for expression in E. coli K12.DepositorInsertPutative MHETase from Hydrogenophaga sp. PML113 (Genbank WP_083293388.1) without 19 residue signal peptide
TagsHisExpressionBacterialMutationdeltaK2:G19; codon optimized for expression in E.…PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only