We narrowed to 18,940 results for: multi
-
-
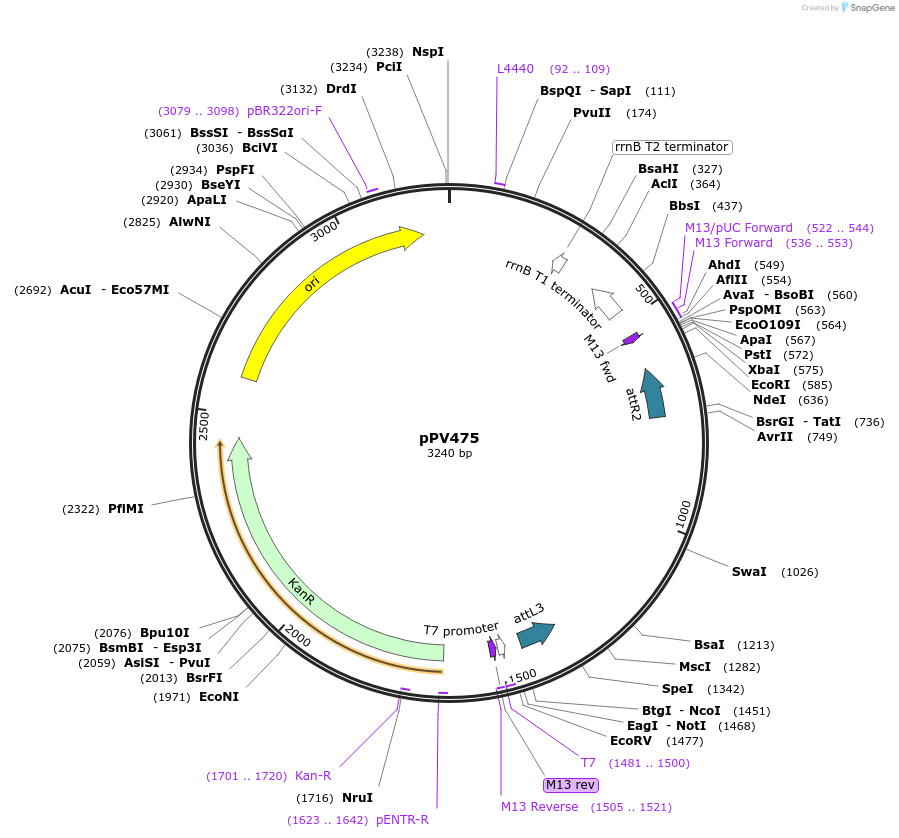
pEvol-pAzFRS.2.t1
Plasmid#73546PurposetRNA synthetase/tRNA pair for the in vivo incorporation of several non-standard amino acids, into proteins in E coli in response to the amber codon, TAGDepositorInsertspAzFRS.2.t1
pAzFRS.2.t1
ExpressionBacterialAvailable SinceMarch 25, 2016AvailabilityAcademic Institutions and Nonprofits only -
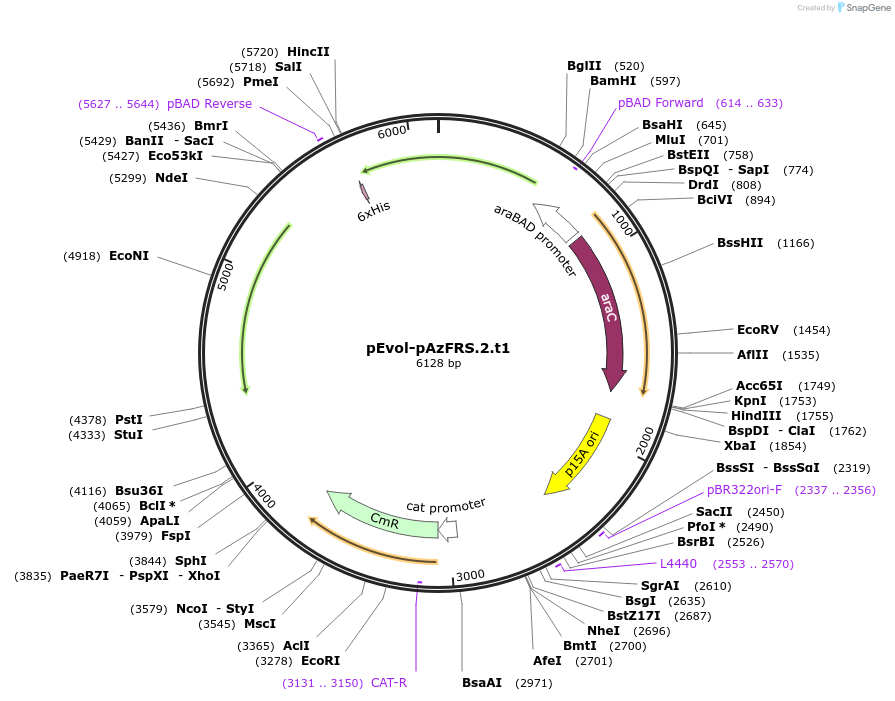
pEvol-pAzFRS.1.t1
Plasmid#73547PurposetRNA synthetase/tRNA pair for the in vivo incorporation of p-azido-l-phenylalanine, into proteins in E coli in response to the amber codon, TAGDepositorInsertspAzFRS.1.t1
pAzFRS.1.t1
ExpressionBacterialAvailable SinceMarch 31, 2016AvailabilityAcademic Institutions and Nonprofits only -
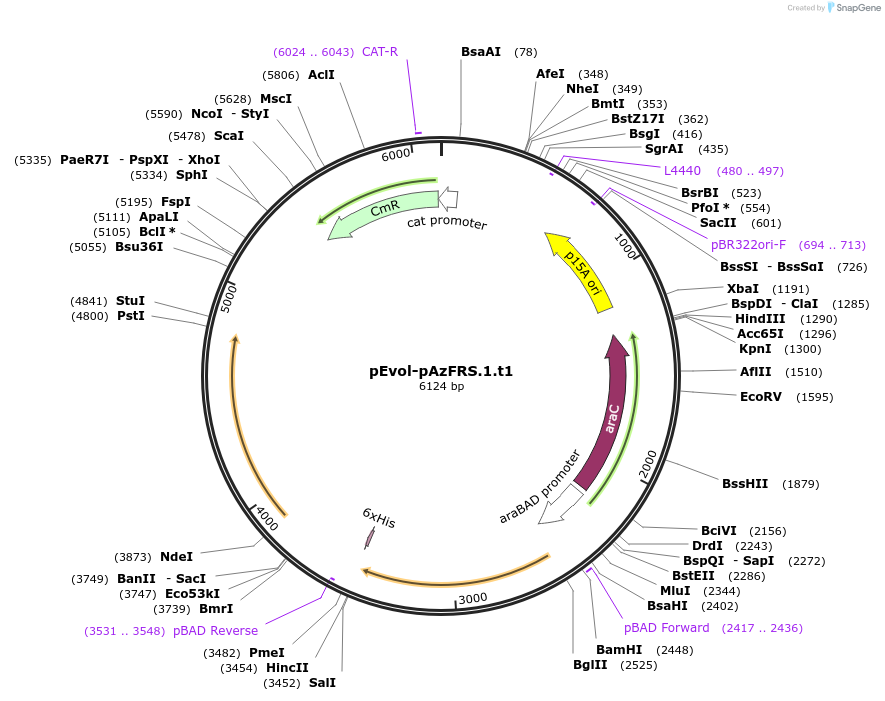
pEvol-pAcFRS.2.t1
Plasmid#73544PurposetRNA synthetase/tRNA pair for the in vivo incorporation of several non-standard amino acids, into proteins in E coli in response to the amber codon, TAGDepositorInsertspAcFRS.2.t1
pAcFRS.2.t1
ExpressionBacterialAvailable SinceMarch 25, 2016AvailabilityAcademic Institutions and Nonprofits only -
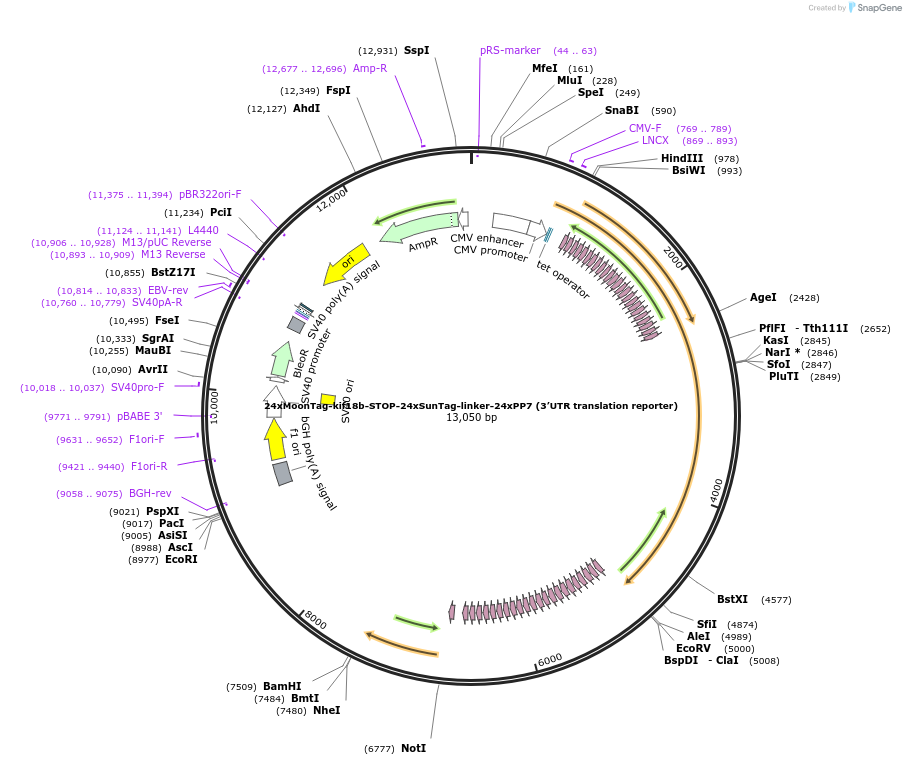
24xMoonTag-kif18b-STOP-24xSunTag-linker-24xPP7 (3’UTR translation reporter)
Plasmid#128605PurposeExpresses 24xgp41 peptide array (MoonTag peptide array), followed by kif18b gene, STOP codon, 24xgcn4 peptide array (SunTag peptide array), linker and 24 repeats of PP7 hairpins.DepositorInsertsExpressionMammalianPromoterPcmvAvailable SinceJan. 21, 2020AvailabilityAcademic Institutions and Nonprofits only -
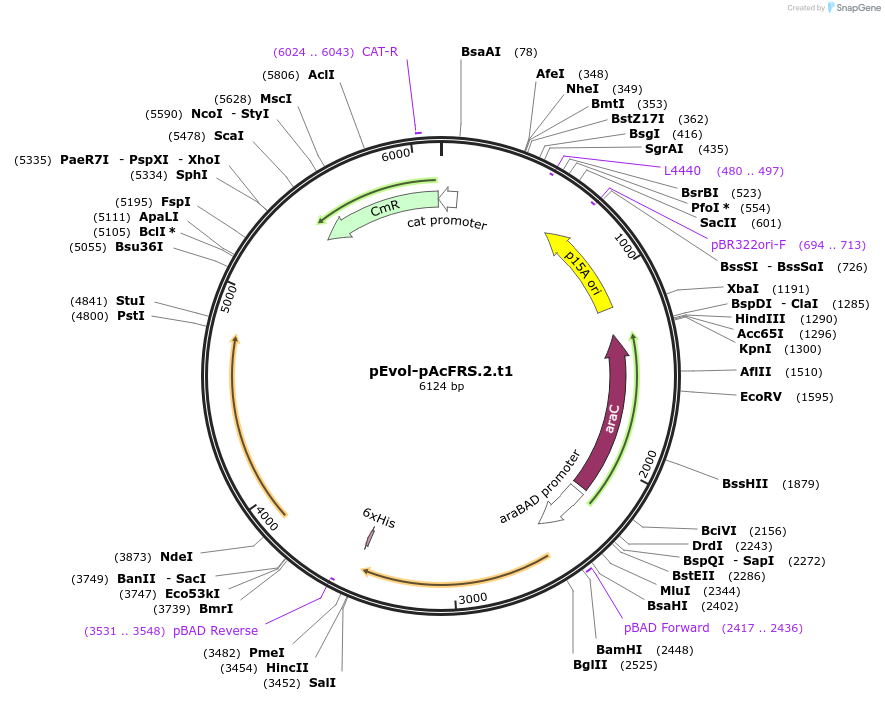
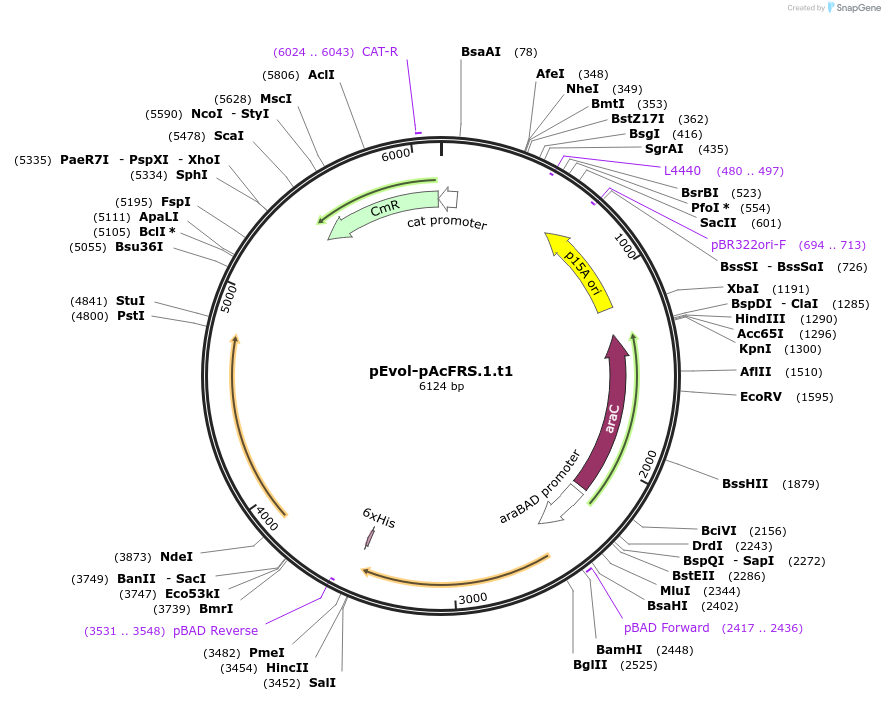
pEvol-pAcFRS.1.t1
Plasmid#73545PurposetRNA synthetase/tRNA pair for the in vivo incorporation of several non-standard amino acids, into proteins in E coli in response to the amber codon, TAGDepositorInsertspAcFRS.1.t1
pAcFRS.1.t1
ExpressionBacterialAvailable SinceMarch 25, 2016AvailabilityAcademic Institutions and Nonprofits only -
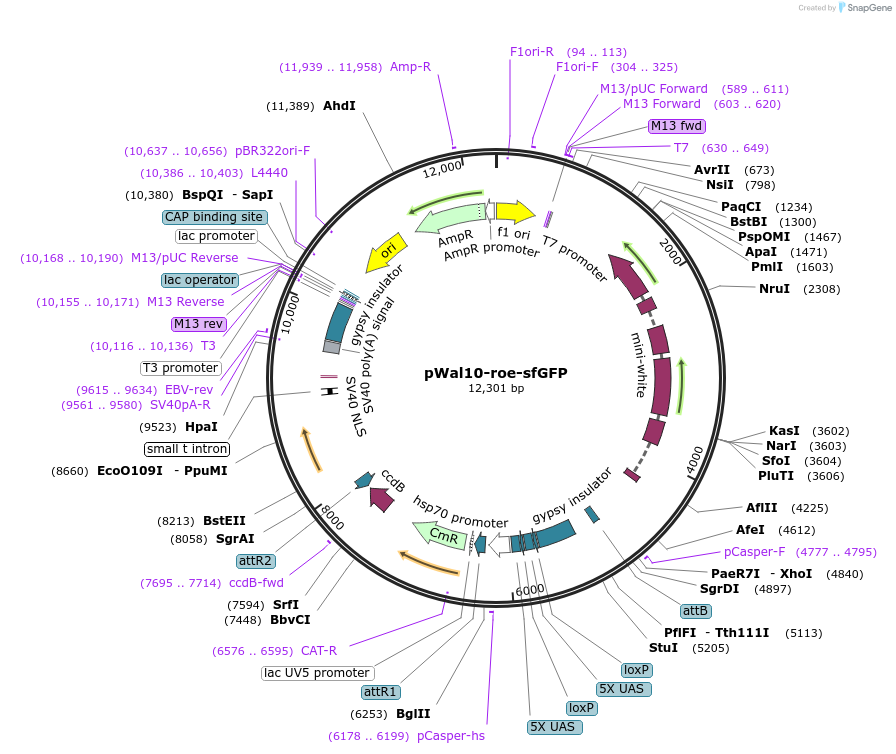
pWal10-roe-sfGFP
Plasmid#240239PurposeGateway destination plasmid to generate C-terminal sfGFP fusions under control of UAS promoter. Can be used to generate transgenic fly lines by white+ fly eye selection and phiC31 integration.DepositorTypeEmpty backboneUseGateway CloningExpressionInsectAvailable SinceJuly 11, 2025AvailabilityAcademic Institutions and Nonprofits only -
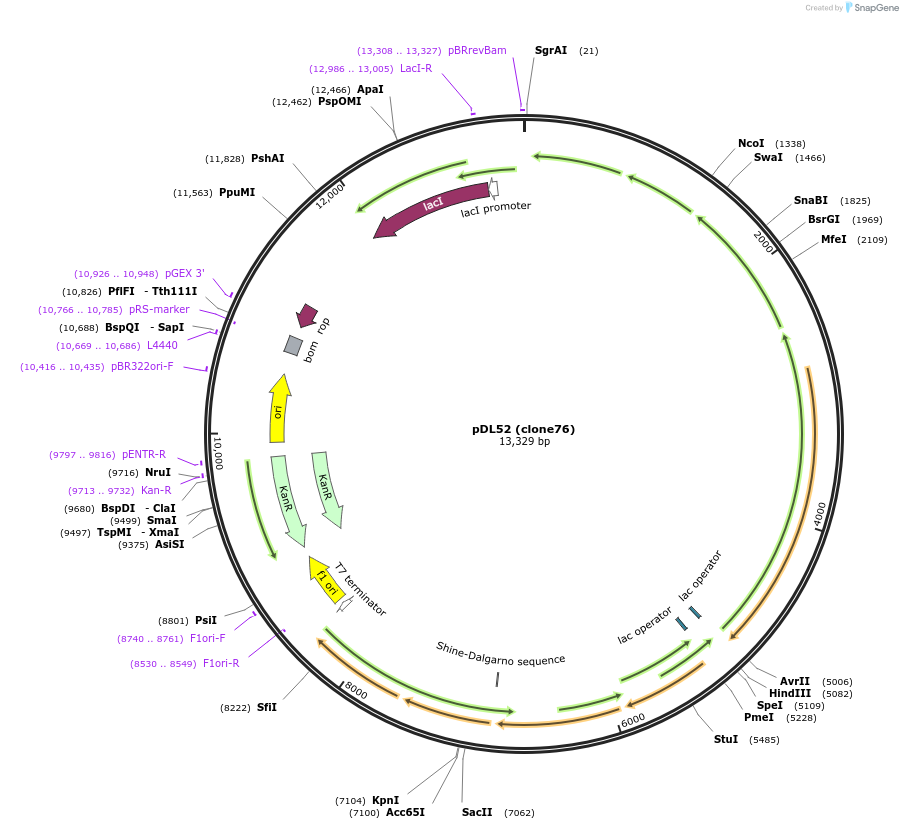
pDL52 (clone76)
Plasmid#188579PurposeHigh-yield production of coenzyme F420 in E. coliDepositorInsertsribA
ribD
yigB
fbiC
cofC
cofD
cofI
cofE
ExpressionBacterialMutationCodon optimization, synthetic ribosome binding si…PromoterT7 variantsAvailable SinceSept. 7, 2022AvailabilityAcademic Institutions and Nonprofits only -
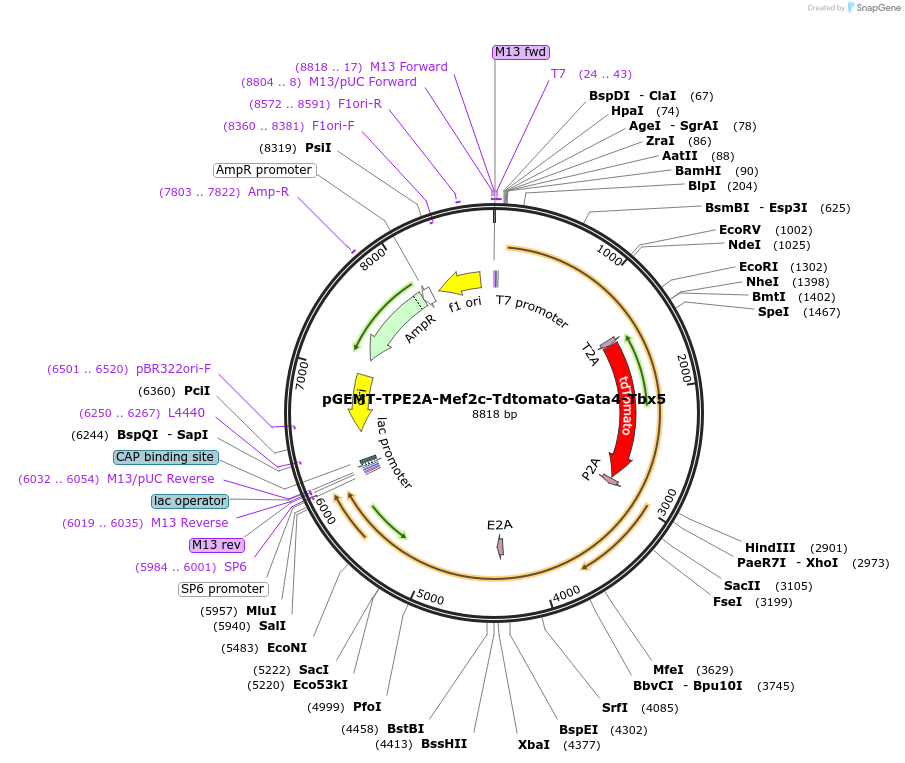
pGEMT-TPE2A-Mef2c-Tdtomato-Gata4-Tbx5
Plasmid#111818PurposeQuadcistronic construct: Co-express four genes by three 2A peptides -- T2A, P2A and E2ADepositorAvailable SinceAug. 20, 2018AvailabilityAcademic Institutions and Nonprofits only -
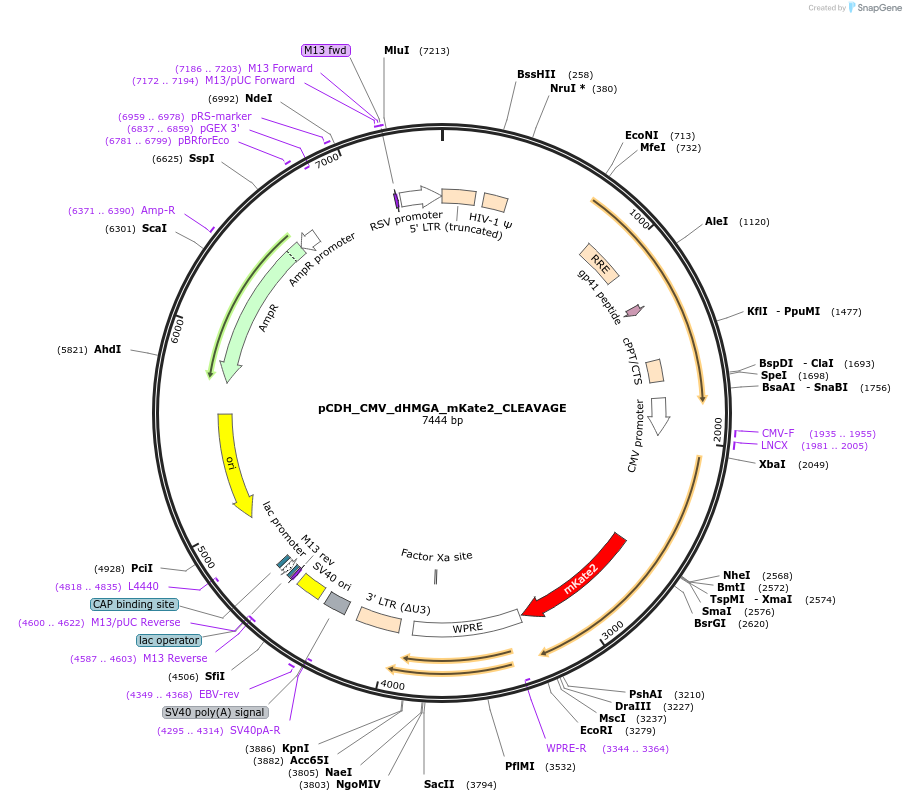
pCDH_CMV_dHMGA_mKate2_CLEAVAGE
Plasmid#167337PurposeExpresses mKate2 fused to TFAM dHMGA mutant in mammalian cellsDepositorInsertTFAM (TFAM Human)
UseLentiviralTagsmKate2ExpressionMammalianMutationTruncation mutant containing MTS (1-42) as well a…PromoterCMVAvailable SinceApril 15, 2021AvailabilityAcademic Institutions and Nonprofits only -
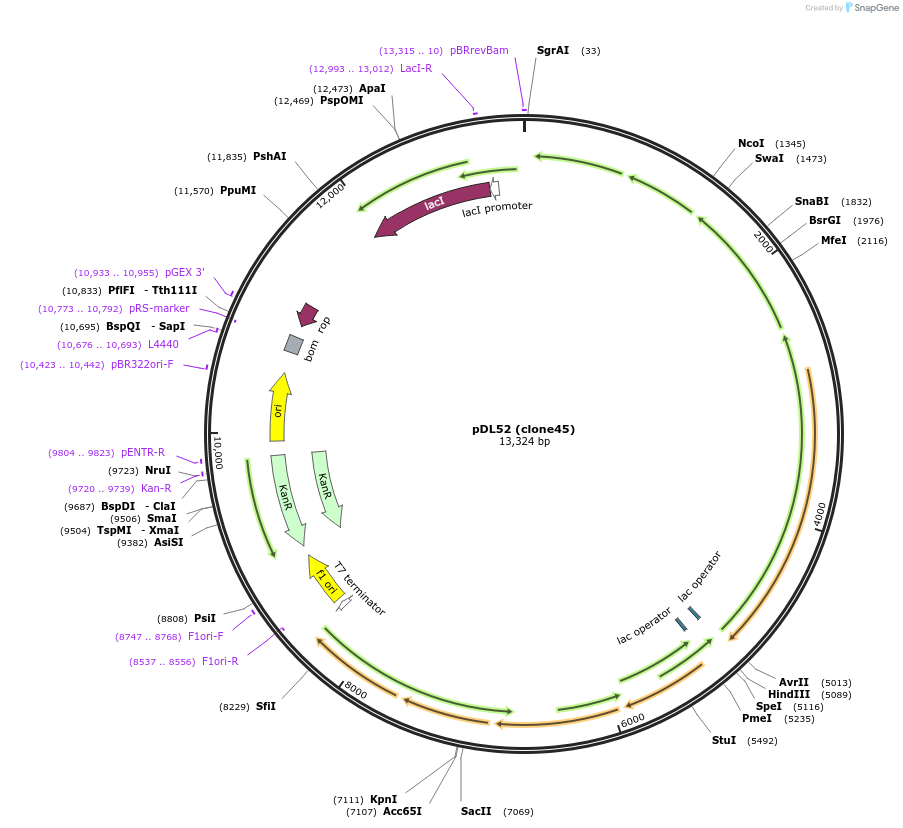
pDL52 (clone45)
Plasmid#188578PurposeHigh-yield production of coenzyme F420 in E. coliDepositorInsertsribA
ribD
yigB
fbiC
cofC
cofD
cofI
cofE
ExpressionBacterialMutationCodon optimization, synthetic ribosome binding si…PromoterT7 variantsAvailable SinceSept. 7, 2022AvailabilityAcademic Institutions and Nonprofits only -
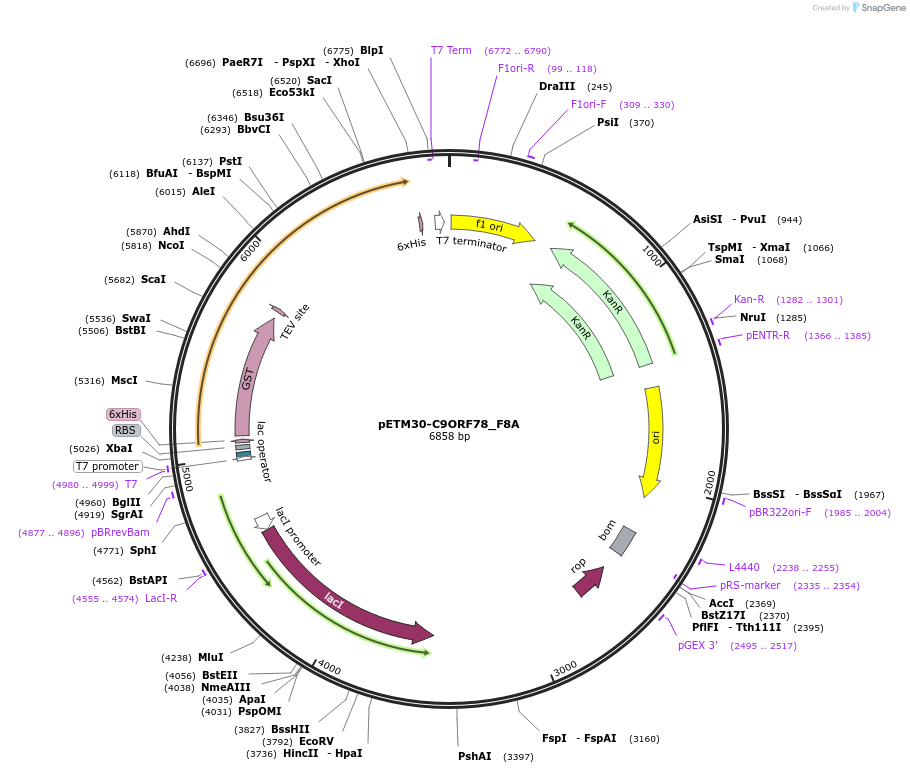
pETM30-C9ORF78_F8A
Plasmid#193369PurposeExpression of C9ORF78 F8A variant in E.coli with N-terminal His6-GST-tag which is cleavable with TEV proteaseDepositorInsertC9ORF78_F8A (C9orf78 Human)
TagsHis6-GST-TEVExpressionBacterialMutationF8A, the mutation is located in the binding inter…PromoterT7Available SinceJan. 24, 2023AvailabilityAcademic Institutions and Nonprofits only -
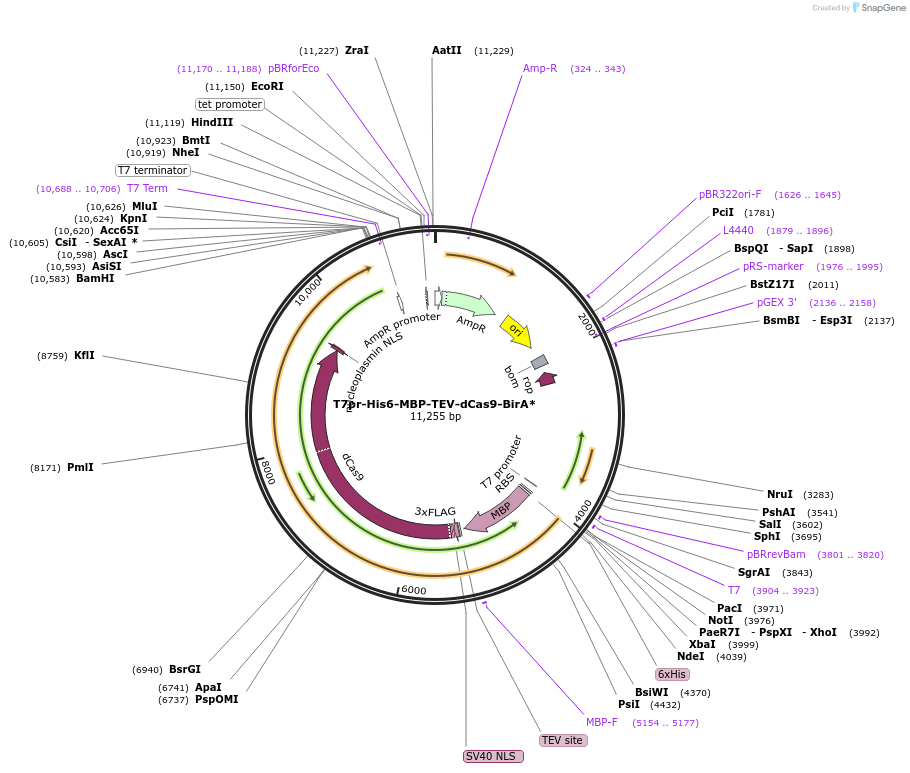
T7pr-His6-MBP-TEV-dCas9-BirA*
Plasmid#159994PurposeBacterial expression of dCas9-BirA*DepositorInsertdCas9-BirA*
Tags6X-His, MBP, 3X-FLAG, NLSExpressionBacterialMutationD10A, H840APromoterT7Available SinceOct. 20, 2020AvailabilityAcademic Institutions and Nonprofits only -
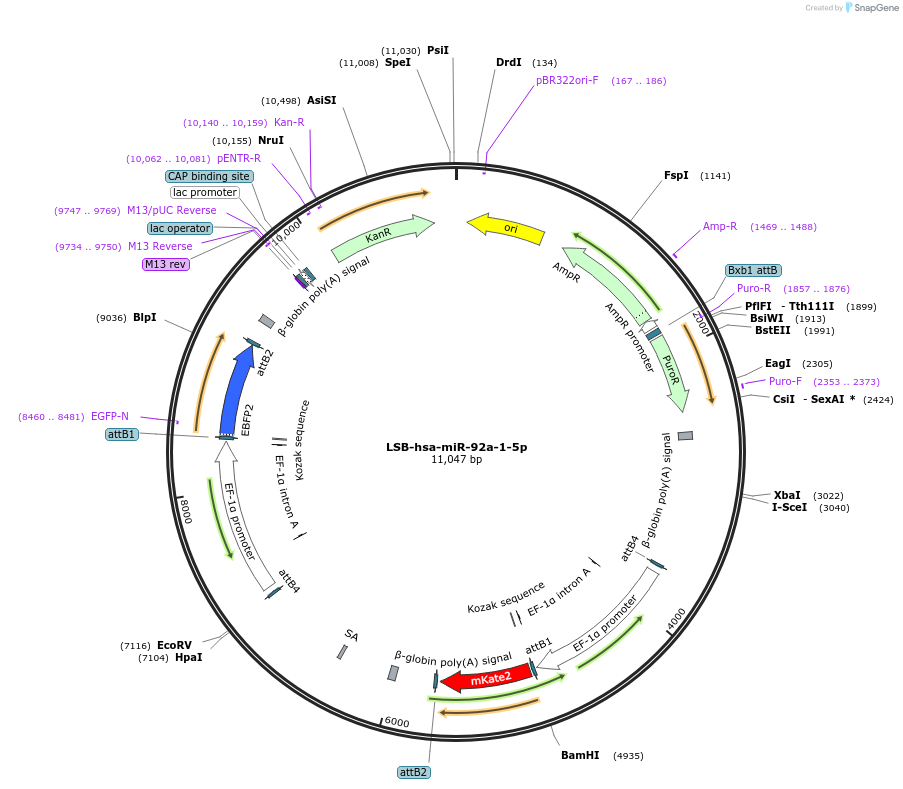
LSB-hsa-miR-92a-1-5p
Plasmid#103750PurposeUsed to sense miRNA activity using fluorescence based tools. hsa-miR-92a-1-5p target in 3' UTR of of mKate. miRNA activity can be measured by the repression of mKate2 relative to EBFP2 fluorescent proteins . Plasmid is inherently unstable; please see Depositor CommentsDepositorInserthsa-miR-92a-1-5p target (MIR92A1 Human)
UseSynthetic BiologyExpressionMammalianPromoterEF-1aAvailable SinceNov. 26, 2018AvailabilityAcademic Institutions and Nonprofits only -
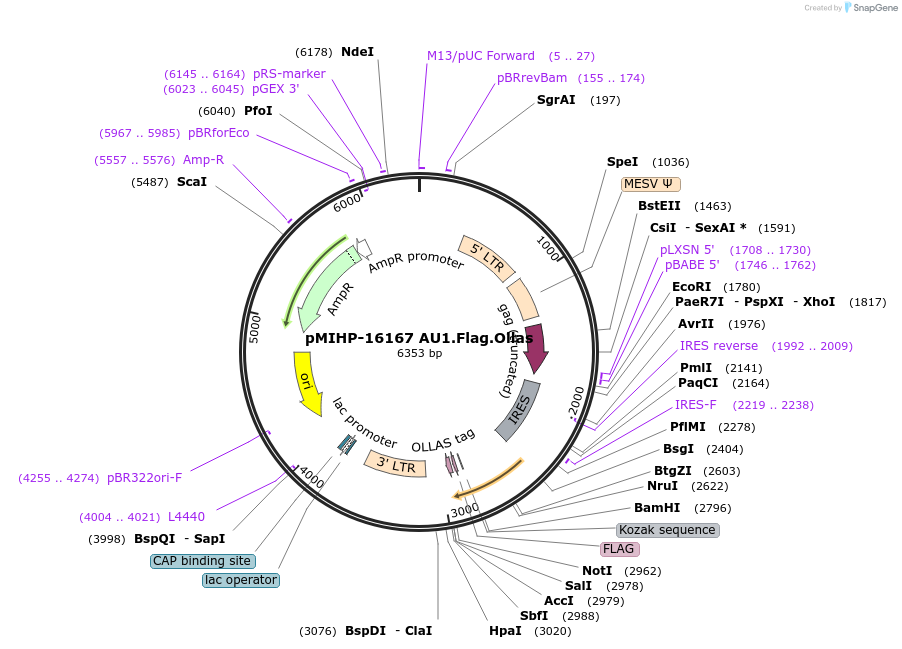
pMIHP-16167 AU1.Flag.Ollas
Plasmid#241479PurposeAU1.Flag.Ollas Flow Code Plasmid This plasmid is to be used with mRNAs.DepositorTypeEmpty backboneUseRetroviralTagsAU1, Flag, and OllasAvailable SinceNov. 12, 2025AvailabilityAcademic Institutions and Nonprofits only -
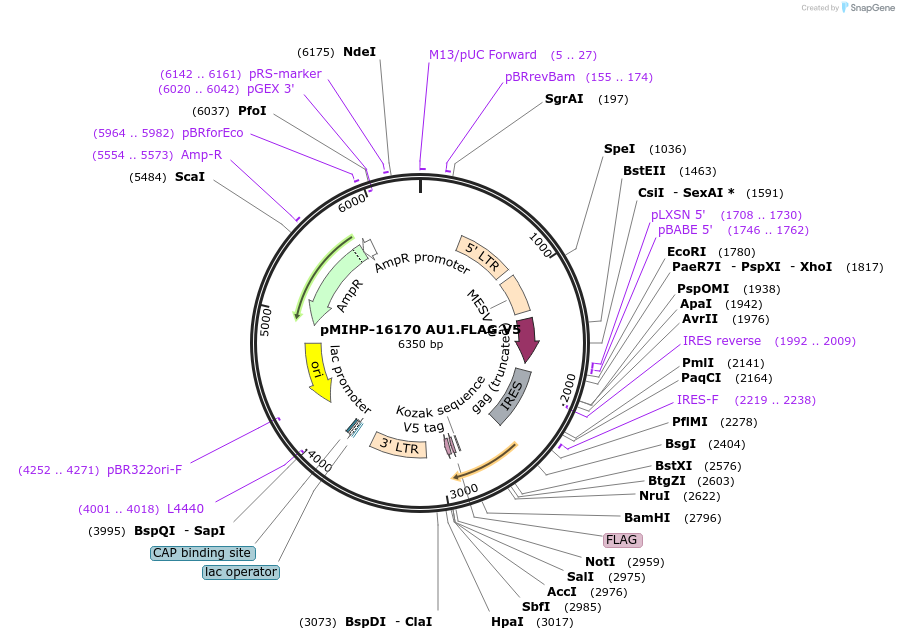
pMIHP-16170 AU1.FLAG.V5
Plasmid#241480PurposeAU1.FLAG.V5 Flow Code Plasmid This plasmid is to be used with mRNAs.DepositorTypeEmpty backboneUseRetroviralTagsAU1, FLAG, and V5Available SinceNov. 12, 2025AvailabilityAcademic Institutions and Nonprofits only -
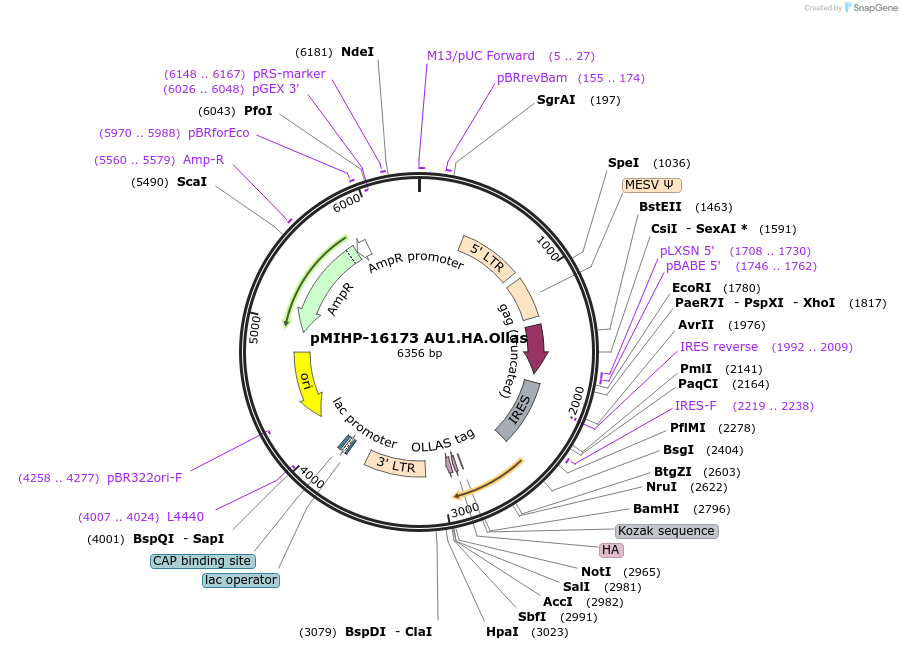
pMIHP-16173 AU1.HA.Ollas
Plasmid#241481PurposeAU1.HA.Ollas Flow Code Plasmid This plasmid is to be used with mRNAs.DepositorTypeEmpty backboneUseRetroviralTagsAU1, HA, and OllasAvailable SinceNov. 12, 2025AvailabilityAcademic Institutions and Nonprofits only -
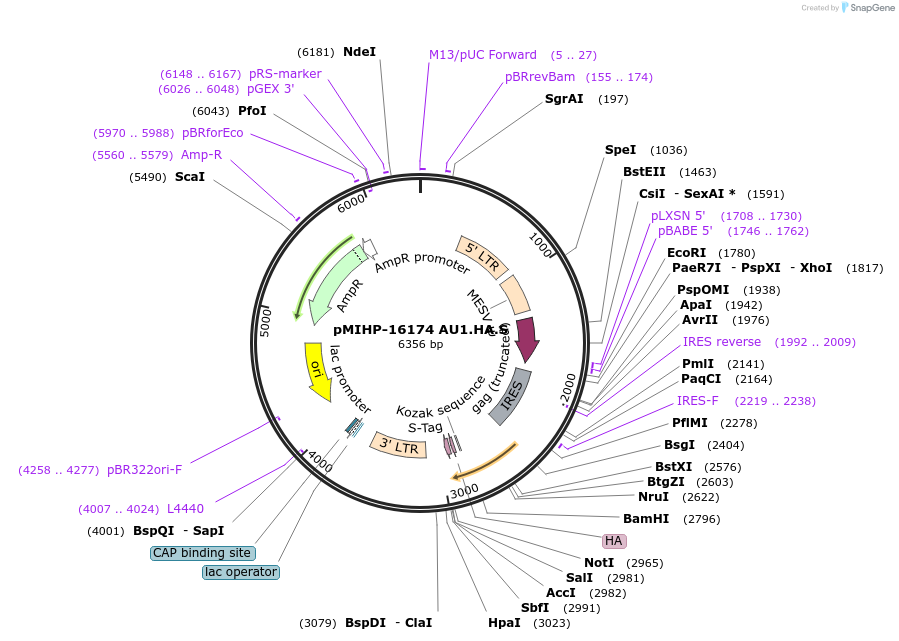
pMIHP-16174 AU1.HA.S
Plasmid#241482PurposeAU1.HA.S Flow Code Plasmid This plasmid is to be used with mRNAs.DepositorTypeEmpty backboneUseRetroviralTagsAU1, HA, and SAvailable SinceNov. 12, 2025AvailabilityAcademic Institutions and Nonprofits only -
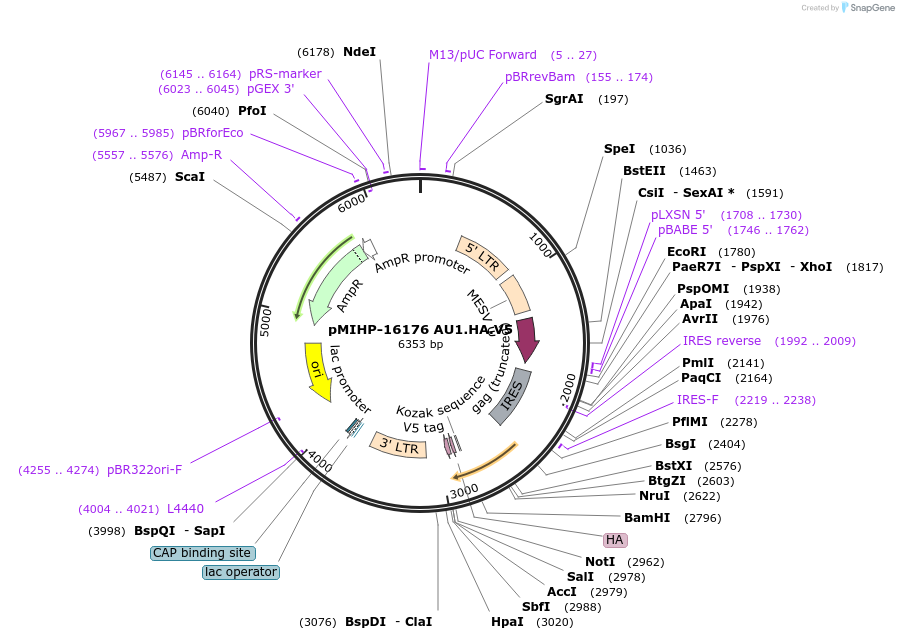
pMIHP-16176 AU1.HA.V5
Plasmid#241483PurposeAU1.HA.V5 Flow Code Plasmid This plasmid is to be used with mRNAs.DepositorTypeEmpty backboneUseRetroviralTagsAU1, HA, and V5Available SinceNov. 12, 2025AvailabilityAcademic Institutions and Nonprofits only -
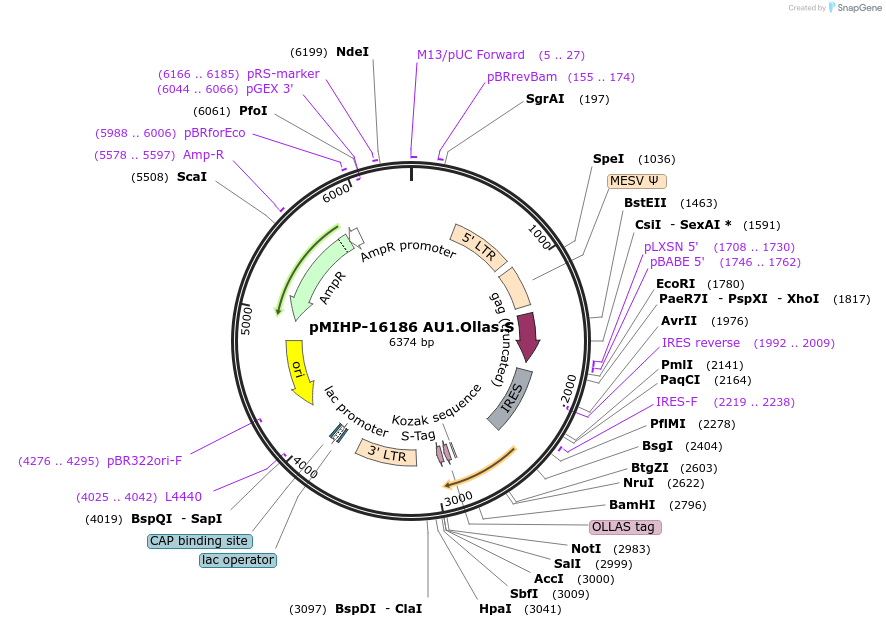
pMIHP-16186 AU1.Ollas.S
Plasmid#241484PurposeAU1.Ollas.S Flow Code Plasmid This plasmid is to be used with mRNAs.DepositorTypeEmpty backboneUseRetroviralTagsAU1, Ollas, and SAvailable SinceNov. 12, 2025AvailabilityAcademic Institutions and Nonprofits only