We narrowed to 15,149 results for: RING
-
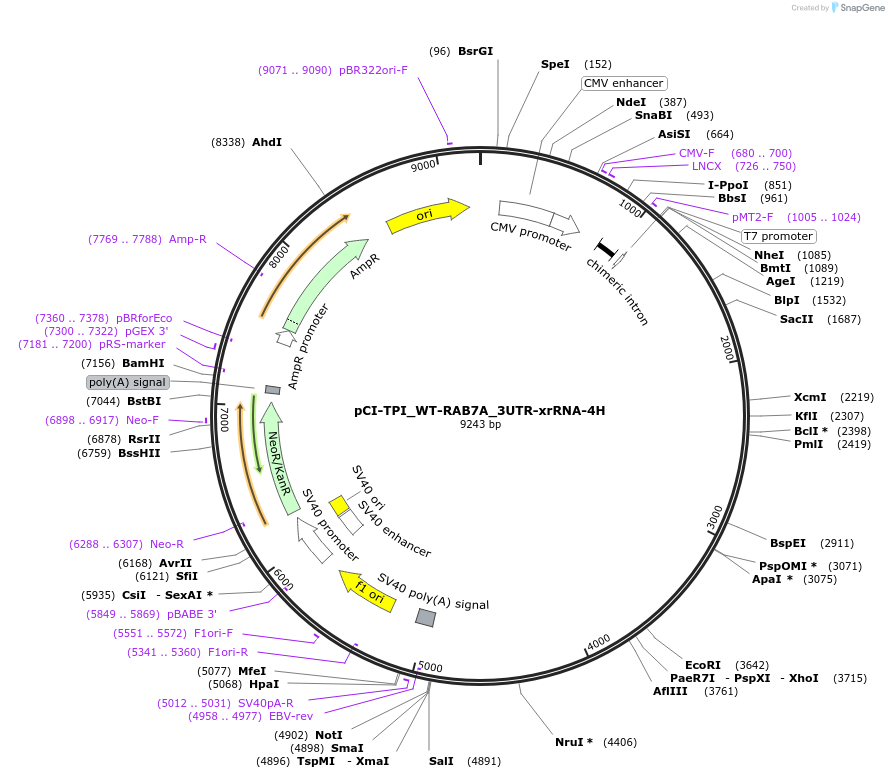
Plasmid#108372PurposeExpresses wild type TPI reporter; contains partial RAB7A control 3' UTR; enables detection of 5'-3' decay intermediates (xrFrag) and allows detection via northern blot (4H binding sites)DepositorInsertExpressionMammalianPromoterCMVAvailable SinceMay 3, 2018AvailabilityAcademic Institutions and Nonprofits only
-
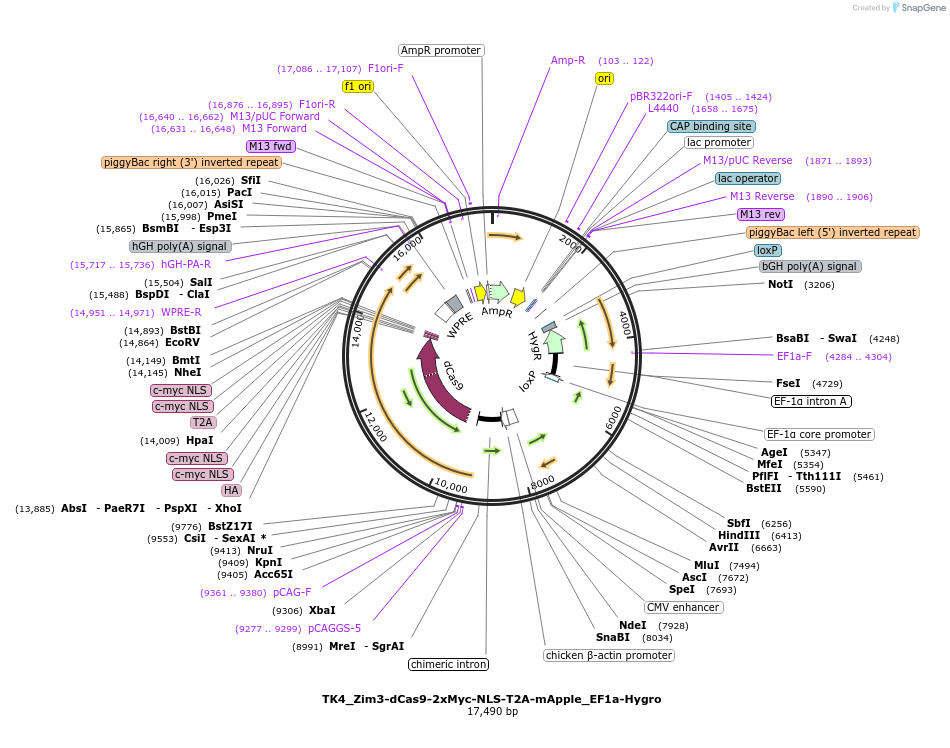
TK4_Zim3-dCas9-2xMyc-NLS-T2A-mApple_EF1a-Hygro
Plasmid#254456PurposeTK4 PiggyBac plasmid for stable expression of Zim3-dCas9 during human pluripotent stem cell differentiationDepositorInsertsZim3-dCas9-2xMyc-NLS-T2A-mApple
hygroR
UsePiggybacTagsT2A-mAppleExpressionMammalianPromoterCAG and EF1aAvailable SinceApril 24, 2026AvailabilityAcademic Institutions and Nonprofits only -
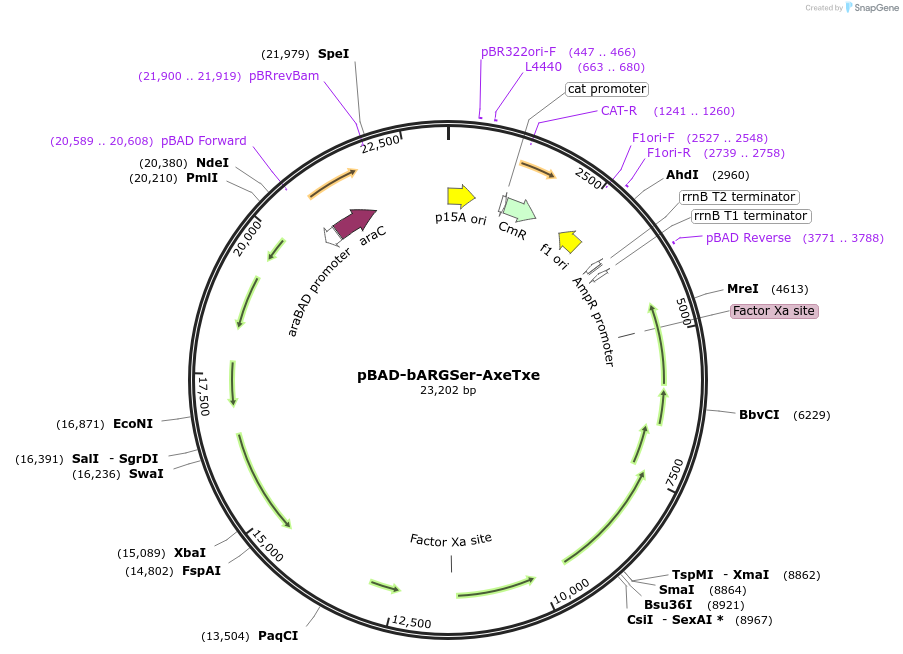
pBAD-bARGSer-AxeTxe
Plasmid#192473PurposeSecond-generation bacterial acoustic reporter gene derived from SerratiaDepositorInsertsbARGSer
AxeTxe
ExpressionBacterialMutationthe gene Ser39006_001280 was deleted from the clu…PromoterNative promoter from Enterococcus faecium and pBADAvailable SinceJan. 3, 2023AvailabilityAcademic Institutions and Nonprofits only -
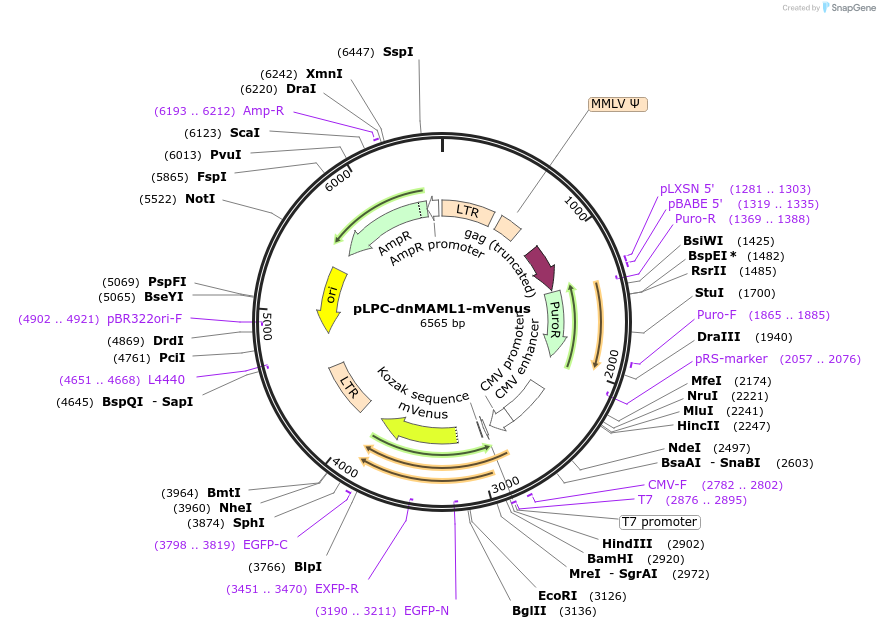
pLPC-dnMAML1-mVenus
Plasmid#171023PurposeRetroviral vector for CMV promoter driven expression of dominant negative (DN) MAML1 (Notch pathway) fused to mVenus fluorescent protein (to be used in conjunction with Phoenix packaging cells).DepositorInsertMAML1 Dominant Negative (MAML1 Human)
UseRetroviralTagsmVenusExpressionMammalianMutationamino acids 12-74 only, hence conferring dominant…PromoterCMVAvailable SinceApril 5, 2025AvailabilityAcademic Institutions and Nonprofits only -
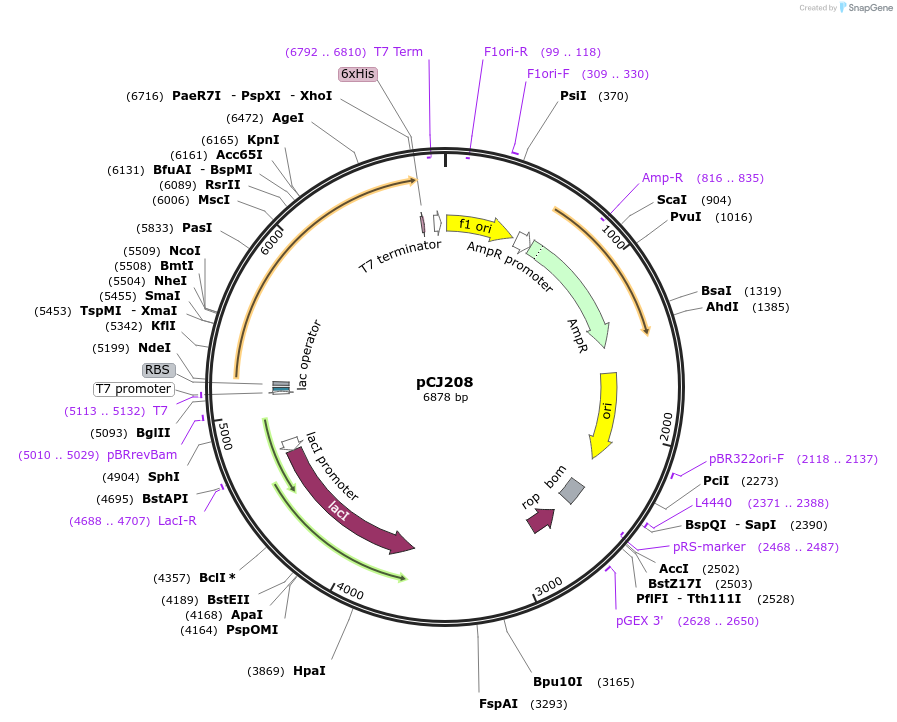
pCJ208
Plasmid#162681PurposeLidded variant of PETase from Ideonella sakaiensis 201-F6 (Genbank GAP38373.1) removing a seven-residue loop of PETase (Trp185:Phe191, PETase numbering) and replacing it with Gly251:Thr472 from MHETase; codon optimized for expression in E. coli K12, with C-terminal His tag.DepositorInsertPETase from Ideonella sakaiensis 201-F6 (Genbank GAP38373.1) incorporating the MHETase lid
TagsHisExpressionBacterialMutationReplacement of W185-F191 with G251-T472 from MHET…PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
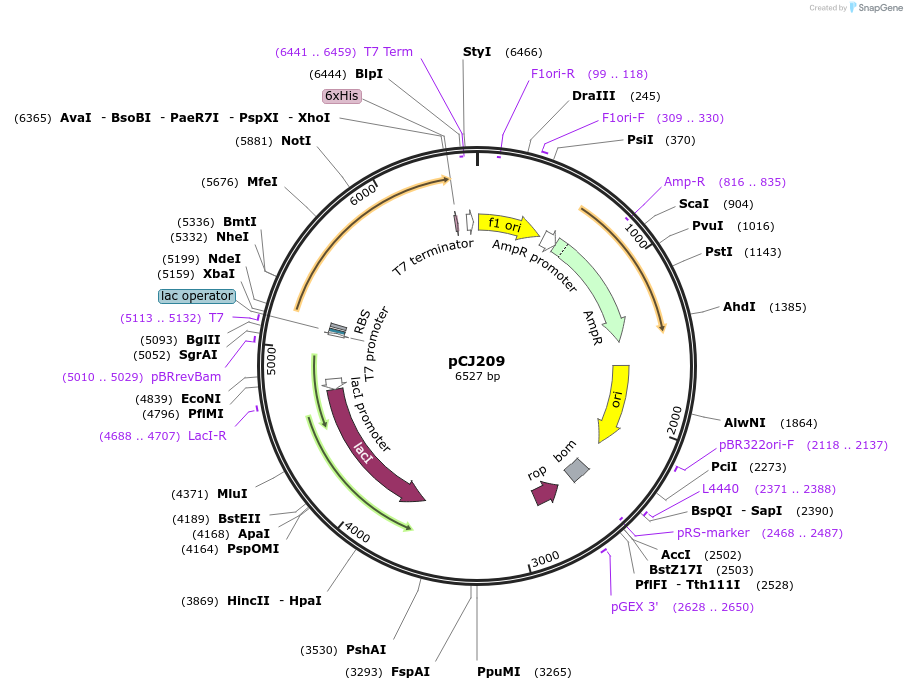
pCJ209
Plasmid#162682PurposeLidless variant of MHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1) replacing Gly251:Thr472 of MHETase with the 7-residue loop (Trp185:Phe191, PETase numbering) of PETase , codon optimized for expression in E. coli K12, with C-terminal His tag.DepositorInsertMHETase from Ideonella sakaiensis 201-F6 (Genbank GAP38911.1) with the lid removed
TagsHisExpressionBacterialMutationReplacement of G251-T472 with W185-F191 from PETa…PromoterT7/lacAvailable SinceJan. 6, 2021AvailabilityAcademic Institutions and Nonprofits only -
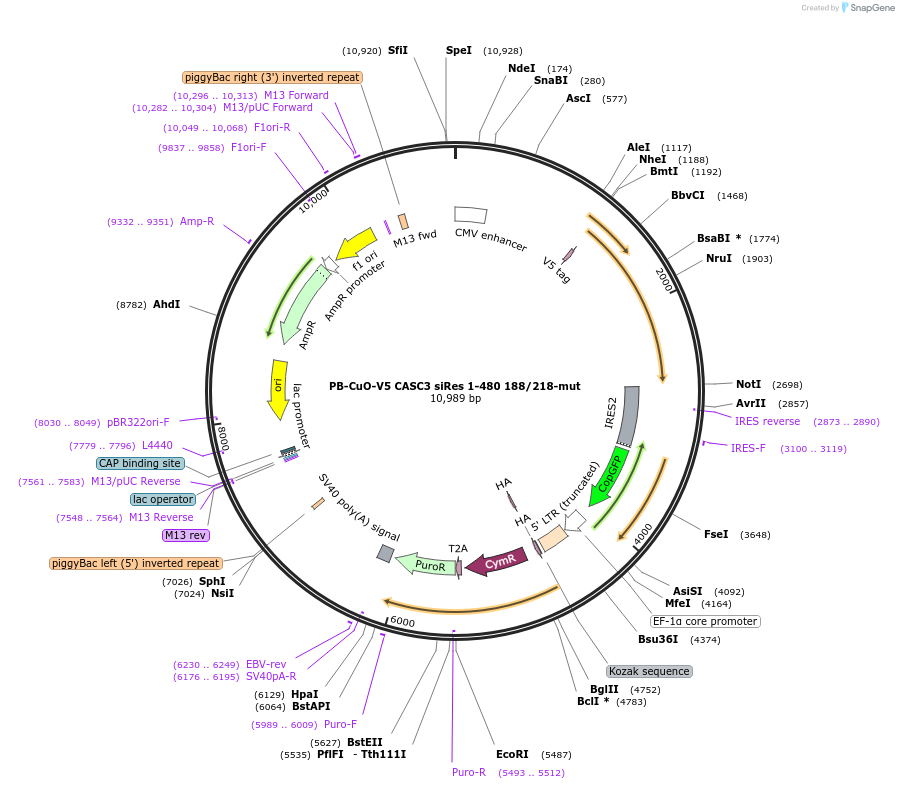
PB-CuO-V5 CASC3 siRes 1-480 188/218-mut
Plasmid#158542PurposeFor PiggyBac-mediated integration of V5-tagged siRNA-resistant EJC-binding deficient CASC3 1-480DepositorInsertCASC3; BTZ; MLN51 (CASC3 Human)
TagsV5ExpressionMammalianMutationAmino acids 1-480, C-terminal deletion, F188D, W2…PromoterCMVAvailable SinceOct. 21, 2020AvailabilityAcademic Institutions and Nonprofits only -
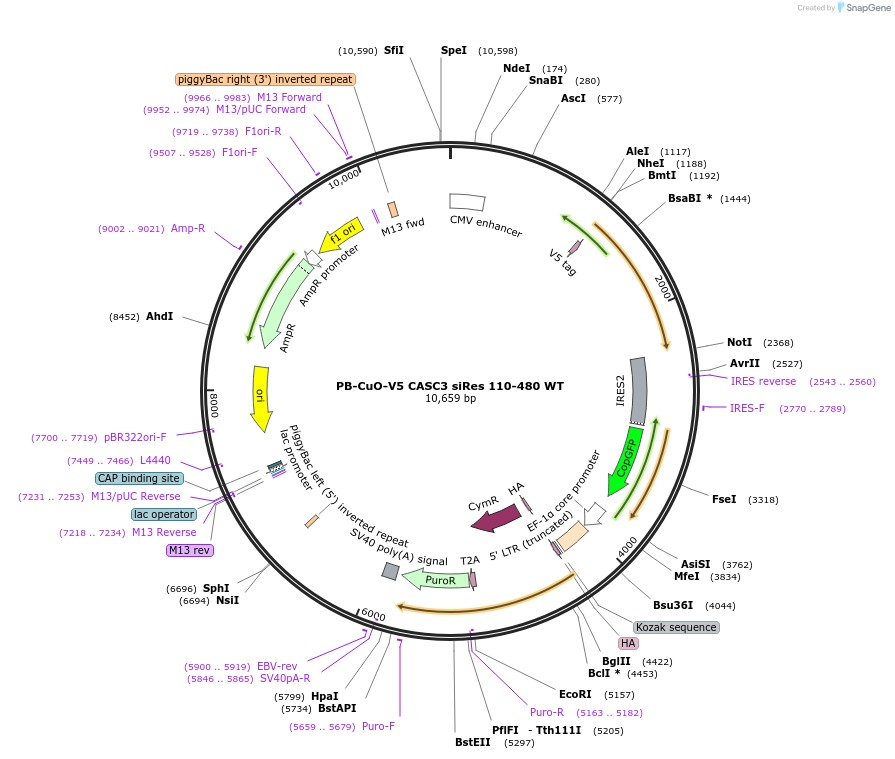
PB-CuO-V5 CASC3 siRes 110-480 WT
Plasmid#158543PurposeFor PiggyBac-mediated integration of V5-tagged siRNA-resistant CASC3 110-480DepositorInsertCASC3; BTZ; MLN51 (CASC3 Human)
TagsV5ExpressionMammalianMutationAmino acids 110-480, C-terminal deletionPromoterCMVAvailable SinceOct. 21, 2020AvailabilityAcademic Institutions and Nonprofits only -
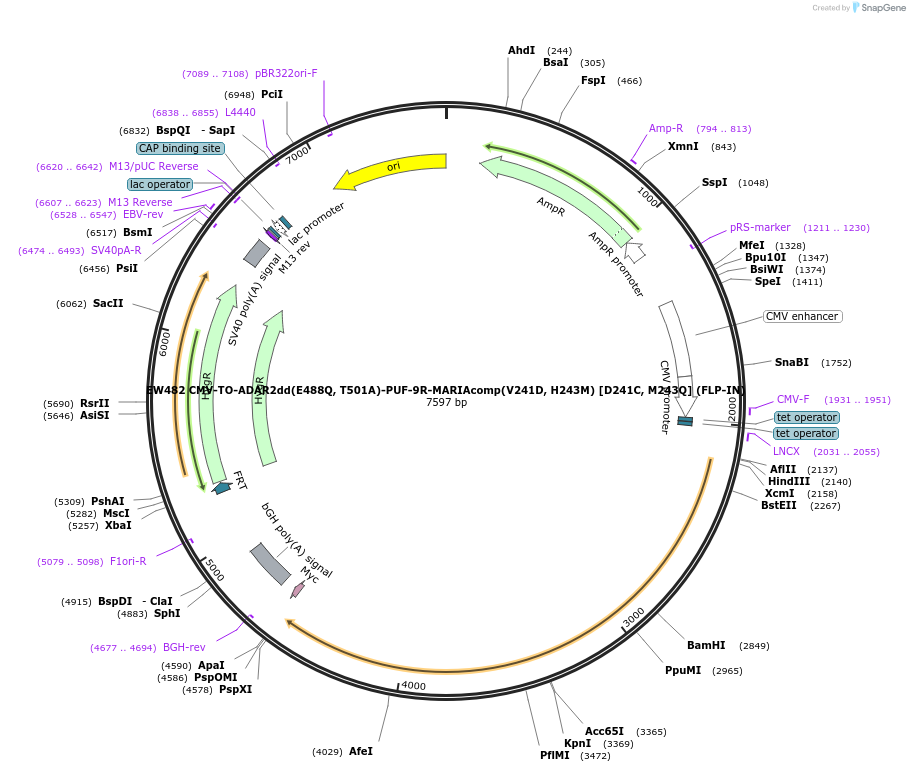
EW482 CMV-TO-ADAR2dd(E488Q, T501A)-PUF-9R-MARIAcomp(V241D, H243M) [D241C, M243Q] (FLP-IN)
Plasmid#236128PurposePlasmid encoding ADAR2dd attached to PUF-9R with deimmunizing and function-restoring mutations under control of CMV promoter with two TetR binding sitesDepositorInsertADAR2dd-PUF-9R (ADARB1 Synthetic, Human)
ExpressionMammalianMutationE488Q, T501A in ADAR2dd; V241C, H243Q in PUF-9RPromoterCMV promoter with two TetR binding sitesAvailable SinceApril 9, 2025AvailabilityAcademic Institutions and Nonprofits only -
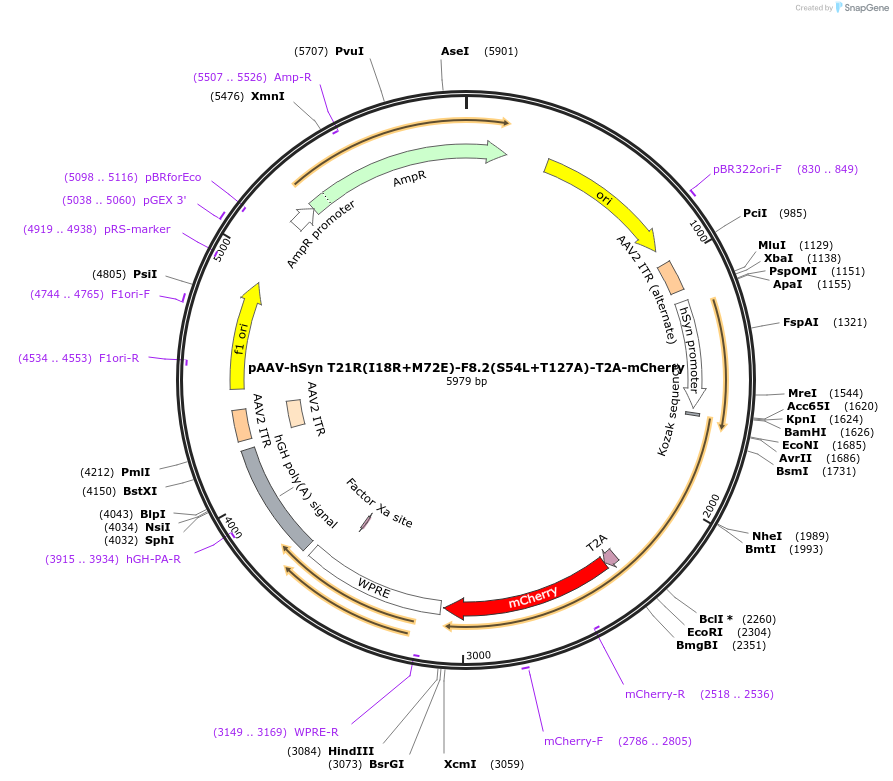
pAAV-hSyn T21R(I18R+M72E)-F8.2(S54L+T127A)-T2A-mCherry
Plasmid#225591PurposeAAV transgene plasmid with hSyn promoter for expression of catalytically dead TRIM21 RING(I18R+M72E)-F8.2(S54L+T127A) anti-tau degrader with a self-cleaving T2A-mCherry fluorescent expression reporterDepositorInsertT21R(I18R+M72E)-F8.2(S54L+T127A)-T2A-mCherry
UseAAVTagsmCherryExpressionMammalianPromoterhSynAvailable SinceFeb. 11, 2025AvailabilityAcademic Institutions and Nonprofits only -
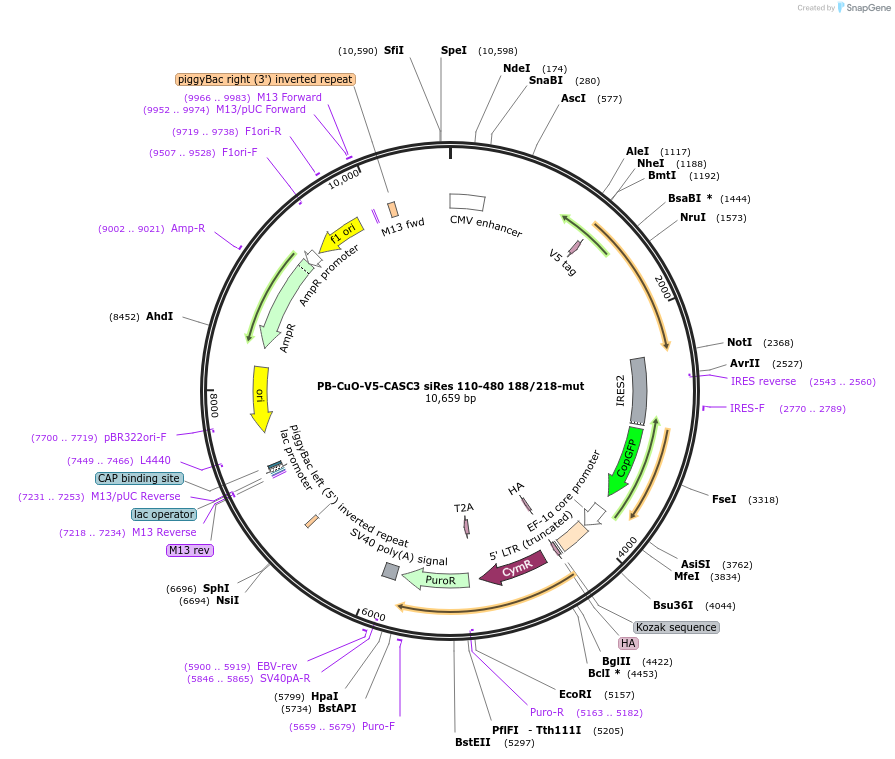
PB-CuO-V5-CASC3 siRes 110-480 188/218-mut
Plasmid#158544PurposeFor PiggyBac-mediated integration of V5-tagged siRNA-resistant EJC-binding deficient CASC3 110-480DepositorInsertCASC3; BTZ; MLN51 (CASC3 Human)
TagsV5ExpressionMammalianMutationAmino acids 110-480, C-terminal deletion, F188D, …PromoterCMVAvailable SinceOct. 21, 2020AvailabilityAcademic Institutions and Nonprofits only -
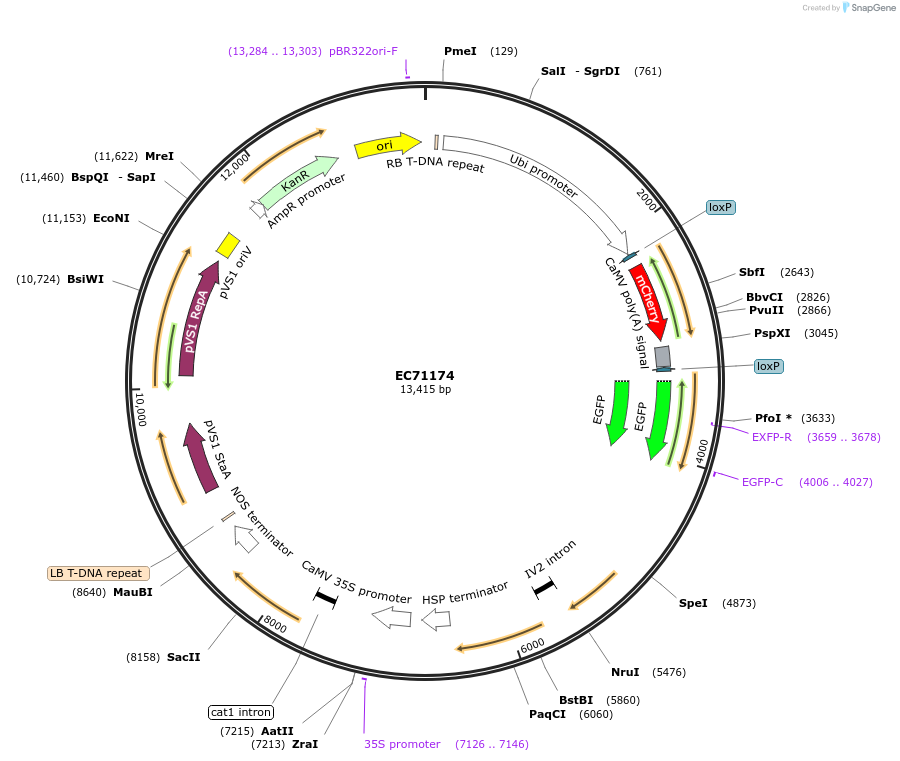
EC71174
Plasmid#154067PurposeLevel 2 Golden Gate vector, pL2M-HvHSP17-CREU5-UBQ-loxmCHERRYER-eGFPERDepositorInsertsHS Cre recombinase
Ubiquitin promoter driving mCherry with the 35S terminator, flanked by lox sites, followed by eGFP with the Actin terminator
HYGROMYCIN Resistance Cassette
UseCre/Lox and Synthetic Biology; Golden gateExpressionBacterial and PlantMutationThe Arabidopsis thaliana U5 small nuclear ribonuc…Promoter35S promoter, Hordeum vulgare HSP 17 promoter use…Available SinceSept. 29, 2020AvailabilityAcademic Institutions and Nonprofits only -
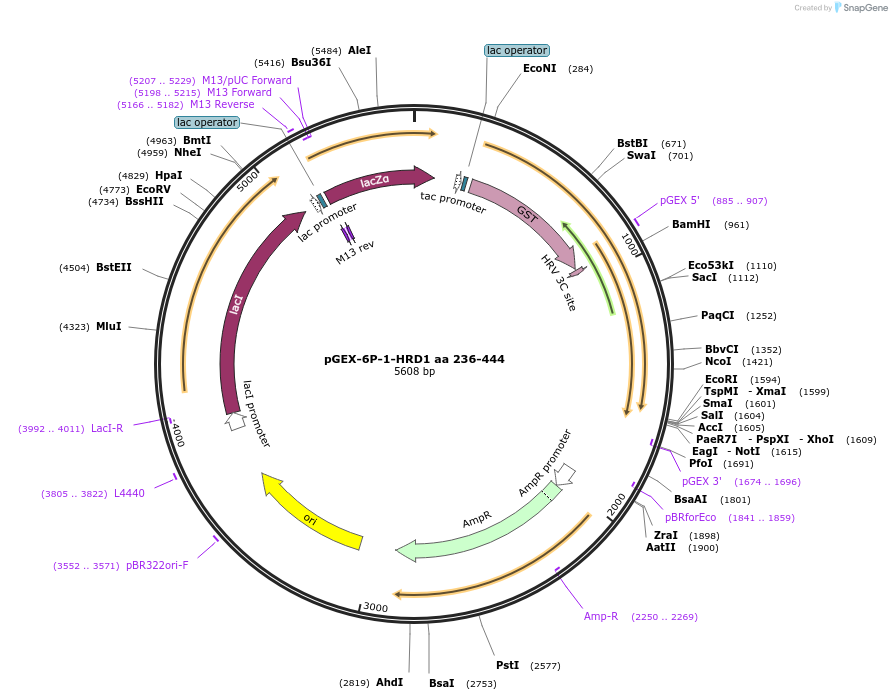
pGEX-6P-1-HRD1 aa 236-444
Plasmid#185349PurposeRING domain-containing GST fusion protein for ubiquitination assays.DepositorAvailable SinceJuly 10, 2023AvailabilityAcademic Institutions and Nonprofits only -
LV2btCRISPRko.v1
Pooled Library#213927PurposeUses lentivirus to deliver the library into hard-to-transduce cellsDepositorExpressionMammalianUseCRISPR and LentiviralAvailable SinceApril 9, 2024AvailabilityAcademic Institutions and Nonprofits only -
PBbtCRISPRko.v1
Pooled Library#213928PurposeUses the PiggyBac transposon system to deliver the library into hard to transduce cellsDepositorExpressionMammalianUseCRISPR and LentiviralAvailable SinceApril 9, 2024AvailabilityAcademic Institutions and Nonprofits only -
p11-3’_8xN_PAM-site1 (RTW554)
Pooled Library#160132PurposePlasmid library harboring a randomized 8nt PAM 3’ of target site 1. Can be used to profile CRISPR-Cas PAM requirements of enzymes that recognize 3’ PAMs (e.g. Cas9)DepositorSpeciesSyntheticUseCRISPRAvailable SinceFeb. 8, 2021AvailabilityAcademic Institutions and Nonprofits only -
p11-5’_10xN_PAM-site3 (RTW572)
Pooled Library#160134PurposePlasmid library harboring a randomized 10nt PAM 5’ of target site 3. Can be used to profile CRISPR-Cas PAM requirements of enzymes that recognize 5’ PAMs (e.g. Cas12a)DepositorSpeciesSyntheticUseCRISPRAvailable SinceFeb. 8, 2021AvailabilityAcademic Institutions and Nonprofits only -
Pdest-EF1α-EGFP-32bp barcode polyA library
Pooled Library#211610PurposeBarcoded retrovirus plasmid library for capturing the transcriptome and viral barcode sequences of individual cells to infer their identities and lineage information.DepositorSpeciesSyntheticUseRetroviralAvailable SinceAug. 29, 2024AvailabilityAcademic Institutions and Nonprofits only -
p11-5’_10xN_PAM-site4 (RTW574)
Pooled Library#160135PurposePlasmid library harboring a randomized 10nt PAM 5’ of target site 4. Can be used to profile CRISPR-Cas PAM requirements of enzymes that recognize 5’ PAMs (e.g. Cas12a)DepositorSpeciesSyntheticUseCRISPRAvailable SinceFeb. 8, 2021AvailabilityAcademic Institutions and Nonprofits only -
p11-3’_8xN_PAM-site2 (RTW555)
Pooled Library#160133PurposePlasmid library harboring a randomized 8nt PAM 3’ of target site 2. Can be used to profile CRISPR-Cas PAM requirements of enzymes that recognize 3’ PAMs (e.g. Cas9)DepositorSpeciesSyntheticUseCRISPRAvailable SinceFeb. 8, 2021AvailabilityAcademic Institutions and Nonprofits only