We narrowed to 1,440 results for: pUC57
-
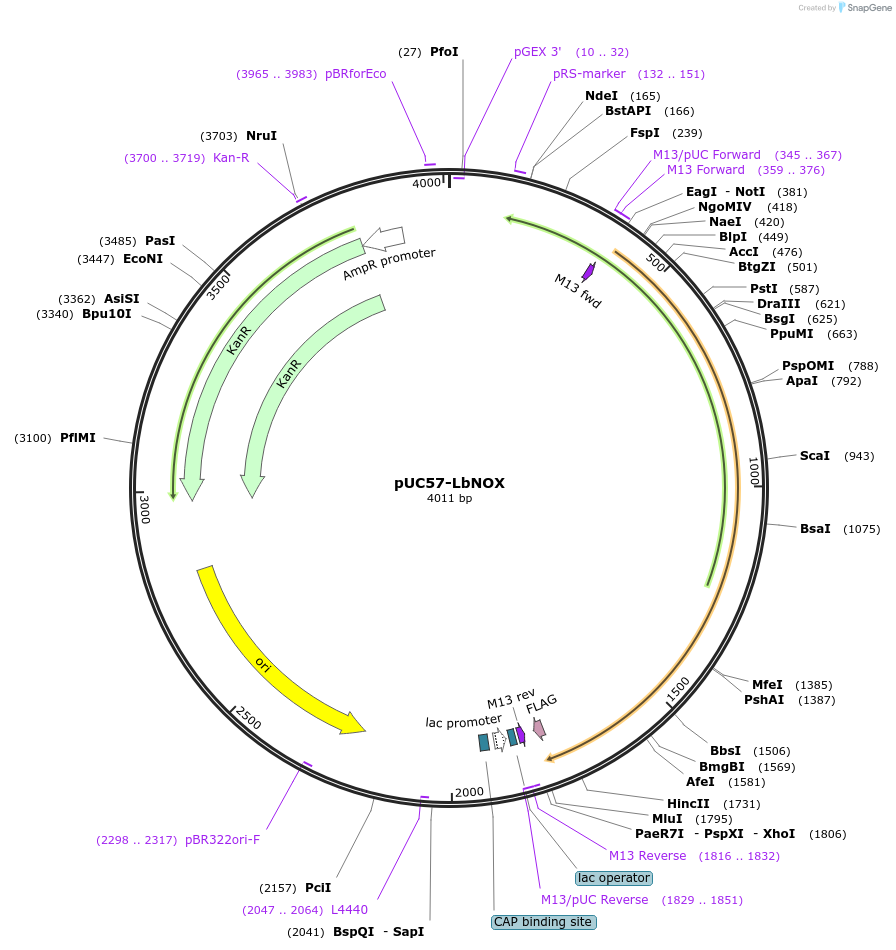
Plasmid#75285PurposeThis plasmid contains human codon optimized sequence of LbNOX which is a water-forming NADH oxidase that can be used to increase NAD+/NADH ratio in mammalian cells.DepositorInsertLbNOX
TagsC-terminal FLAG tag with linkerMutationHuman codon optimizedAvailable SinceMay 17, 2016AvailabilityAcademic Institutions and Nonprofits only -
pUC57-mitoLbNOX
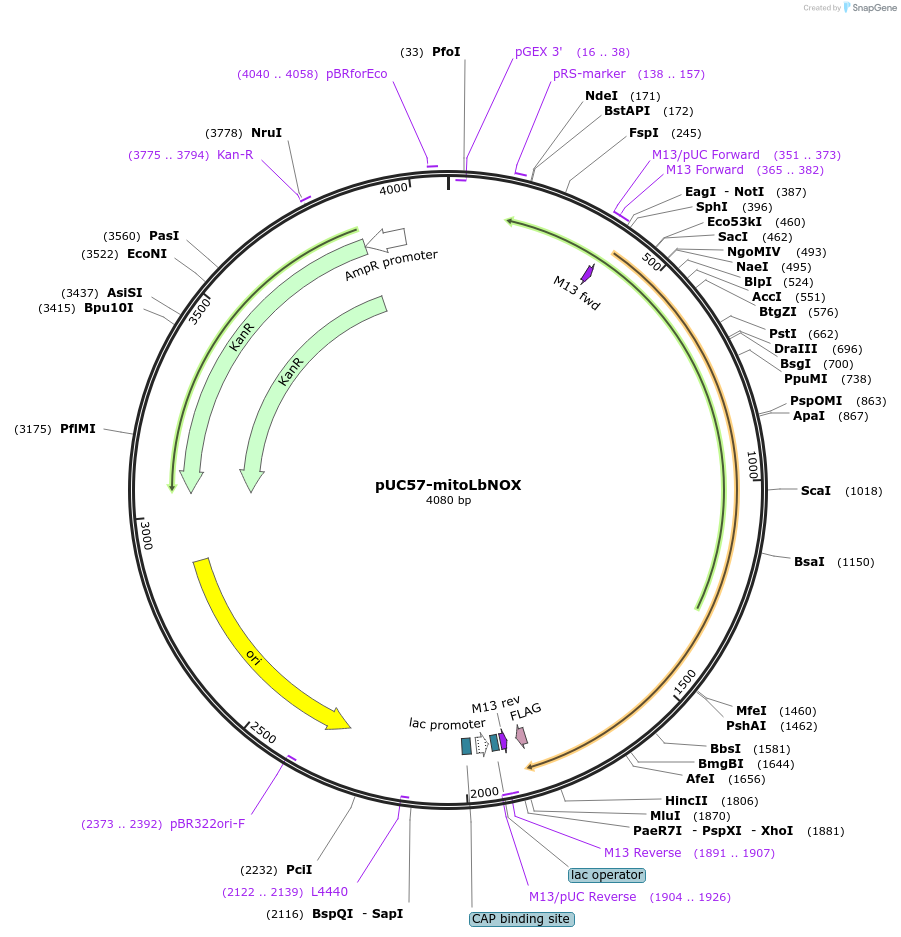
Plasmid#74448PurposeThis plasmid contains human codon optimized sequence of mitoLbNOX which is a water-forming NADH oxidase that can be used to increase NAD+/NADH ratio in mammalian cells.DepositorInsertmitoLbNOX
TagsFlag tag with linker and mitochondrial targeting …MutationHuman codon optimizedAvailable SinceJune 3, 2016AvailabilityAcademic Institutions and Nonprofits only -
pUC57-mitoTPNOX
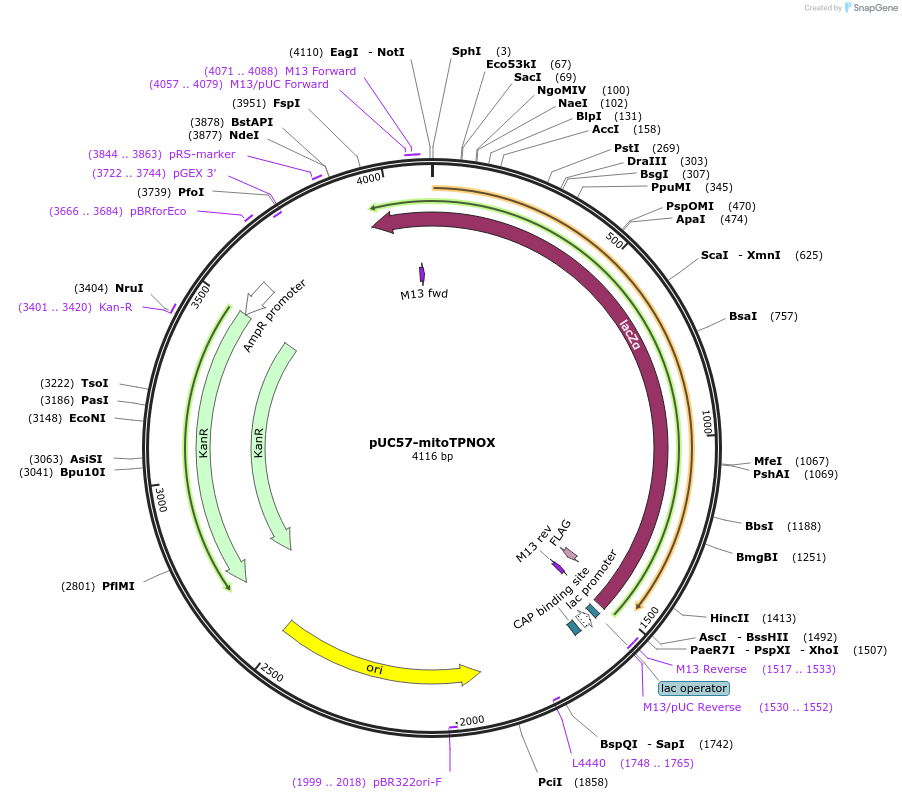
Plasmid#87854PurposeThis plasmid contains human codon optimized sequence of Lactobacillus brevis mutant of a water-forming oxidase that can be used to increase NADP+/NADPH ratio in mammalian cells.DepositorInsertmitoTPNOX
TagsC-terminal FLAG tag with a linker and mitochondri…ExpressionBacterialAvailable SinceAug. 8, 2017AvailabilityAcademic Institutions and Nonprofits only -
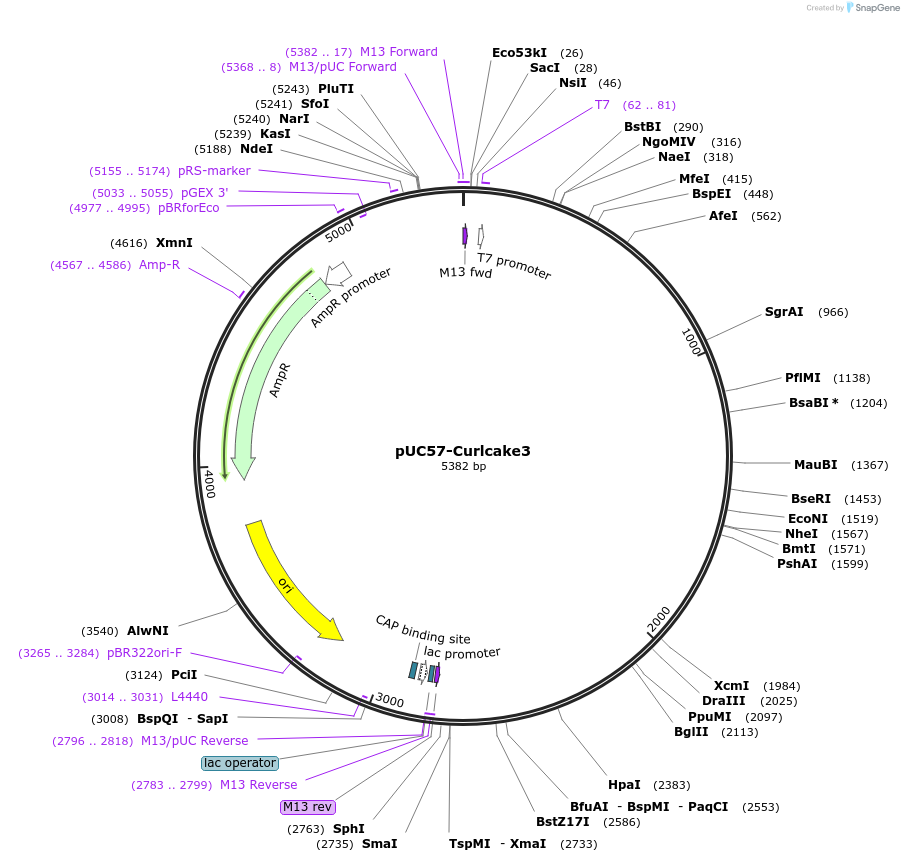
pUC57-Curlcake3
Plasmid#139342PurposeExpresses Curlcake3 (CC3) in E.coliDepositorInsertCurlcake3 with T7 promoter
ExpressionBacterialPromoterT7Available SinceDec. 15, 2020AvailabilityAcademic Institutions and Nonprofits only -
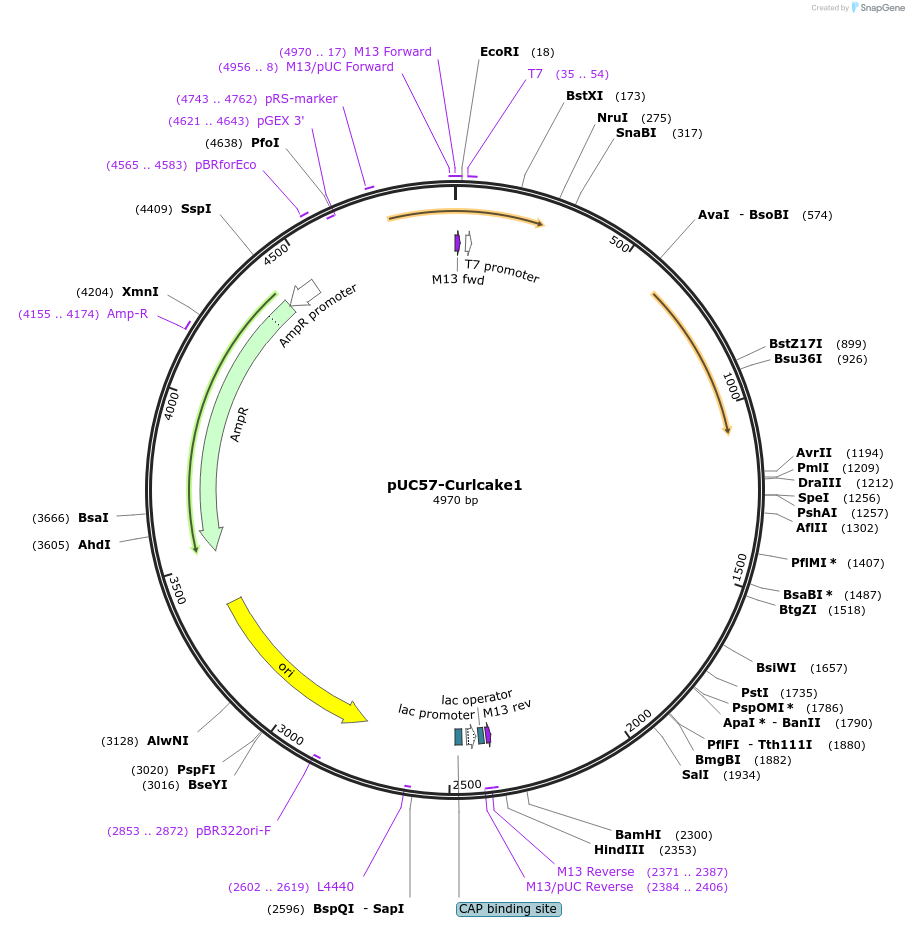
pUC57-Curlcake1
Plasmid#139340PurposeExpresses Curlcake1 (CC1) in E.coliDepositorInsertCurlcake1 with T7 promoter
ExpressionBacterialPromoterT7Available SinceDec. 15, 2020AvailabilityAcademic Institutions and Nonprofits only -
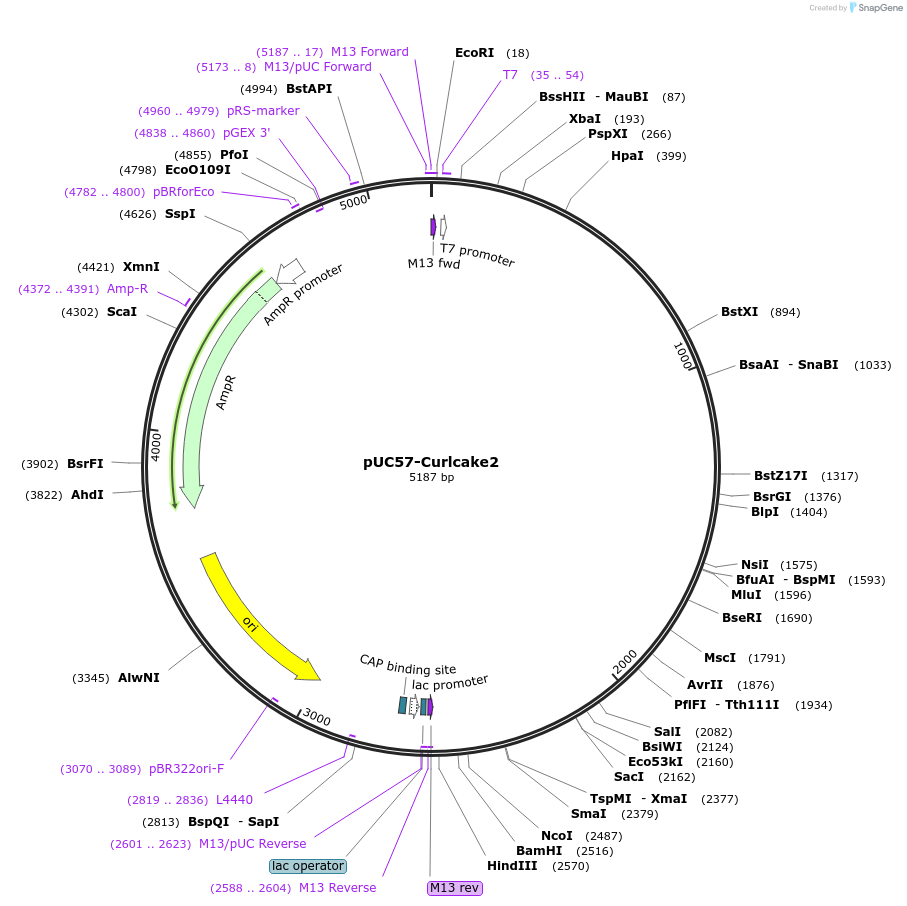
pUC57-Curlcake2
Plasmid#139341PurposeExpresses Curlcake2 (CC2) in E.coliDepositorInsertCurlcake2 with T7 promoter
ExpressionBacterialPromoterT7Available SinceDec. 15, 2020AvailabilityAcademic Institutions and Nonprofits only -
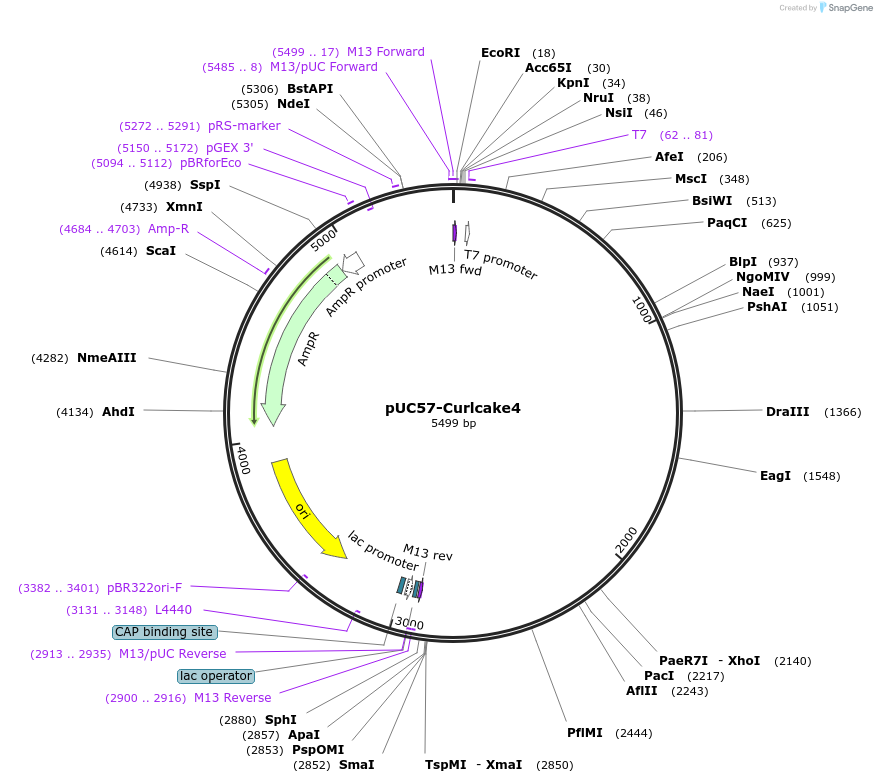
pUC57-Curlcake4
Plasmid#139343PurposeExpresses Curlcake4 (CC4) in E.coliDepositorInsertCurlcake4 with T7 promoter
ExpressionBacterialPromoterT7Available SinceDec. 15, 2020AvailabilityAcademic Institutions and Nonprofits only -
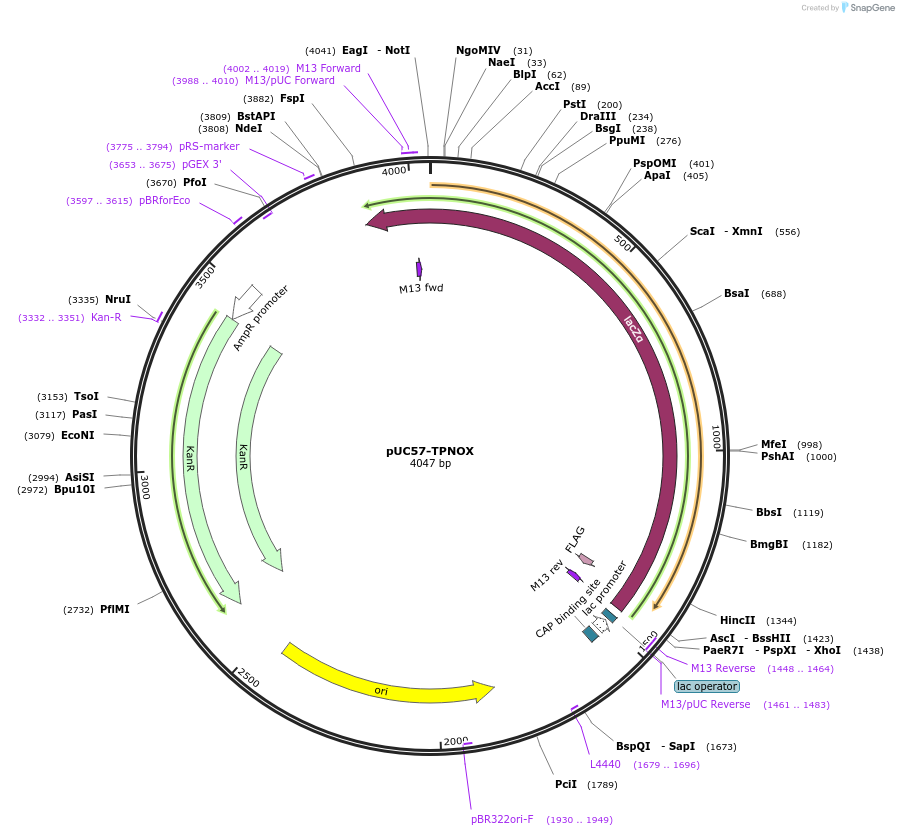
pUC57-TPNOX
Plasmid#87853PurposeThis plasmid contains human codon optimized sequence of Lactobacillus brevis mutant of a water-forming oxidase that can be used to increase NADP+/NADPH ratio in mammalian cells.DepositorInsertTPNOX
TagsC-terminal FLAG tag with a linkerExpressionBacterialAvailable SinceAug. 8, 2017AvailabilityAcademic Institutions and Nonprofits only -
pUC57-CmYLCV
Plasmid#205245PurposeFor plant prime editing in wheat plants or monocotyledons protoplastsDepositorInsertCmYLCV-csy4_binding_site-esgRNA-tevopreQ1-csy4_binding_site
UseCRISPRExpressionPlantPromoterCmYLCVAvailable SinceSept. 27, 2023AvailabilityAcademic Institutions and Nonprofits only -
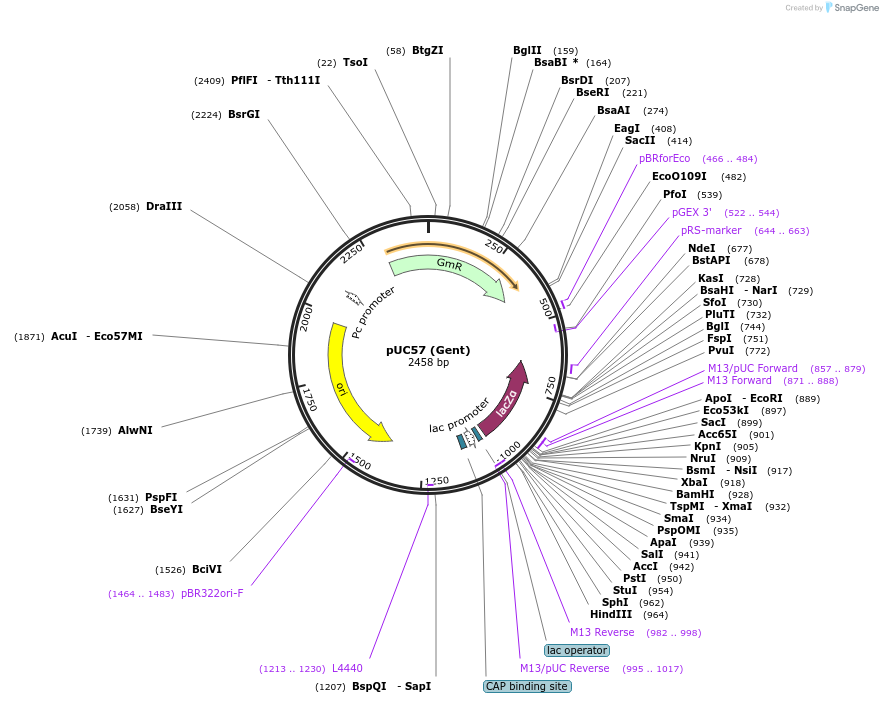
pUC57 (Gent)
Plasmid#54338DepositorTypeEmpty backboneUseCloning vectorAvailable SinceJune 4, 2014AvailabilityAcademic Institutions and Nonprofits only -
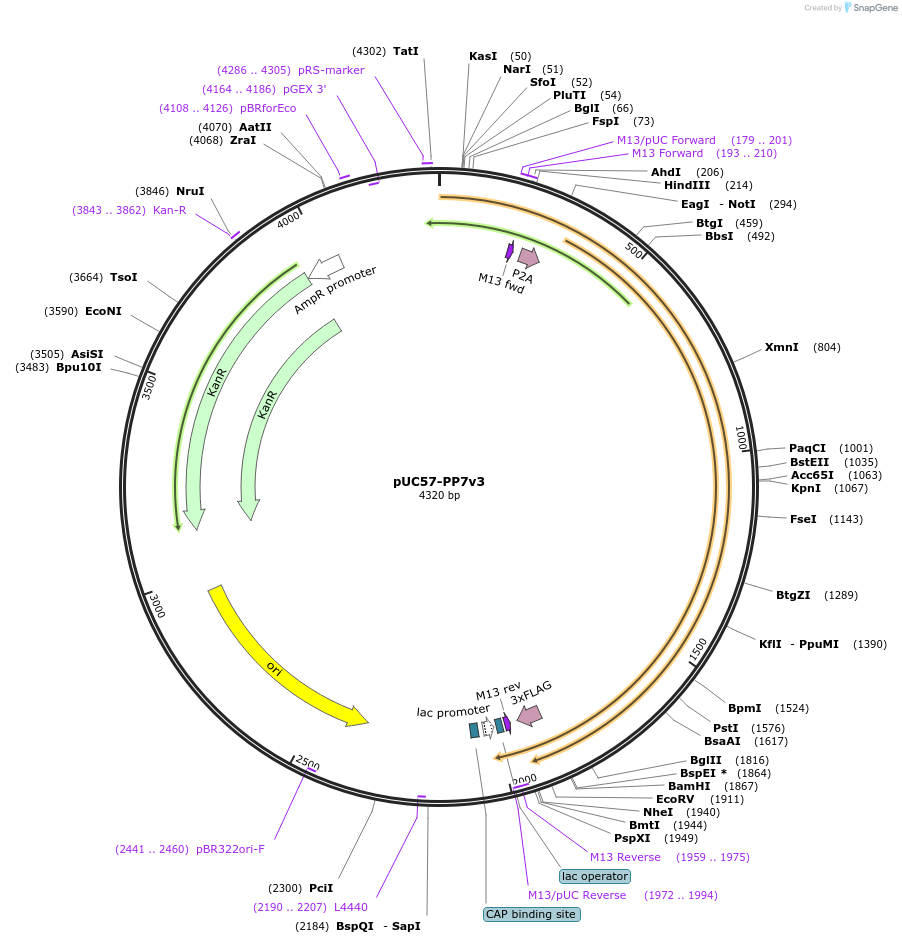
pUC57-PP7v3
Plasmid#107230PurposeExpresses 24x optimized PP7 stem loops (version 3) in bacterial cellsDepositorInsert24xPP7v3
Tags3x FLAG-tag and P2A siteExpressionBacterialAvailable SinceMarch 29, 2018AvailabilityAcademic Institutions and Nonprofits only -
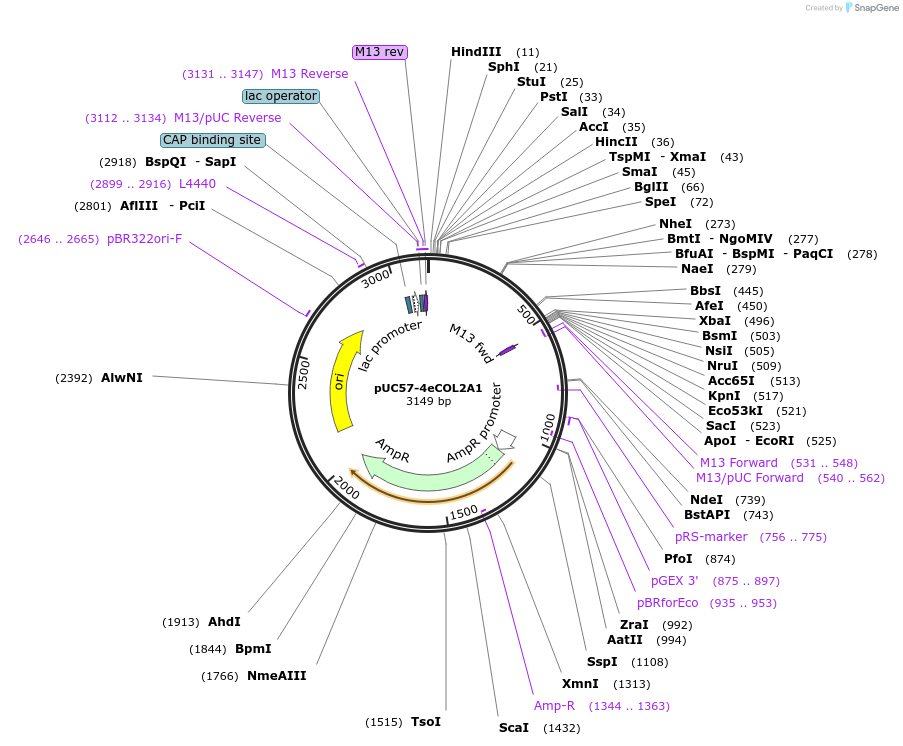
pUC57-4eCOL2A1
Plasmid#97211PurposeFor cloning of a COL2A1-based, chondrogenesis-responsive promoter, and subcloning this insert into new vectorsDepositorInsert4 repeats of COL2A1 intronic enhancer (+2126/+2174) upstream of core promoter (-164/+37) (COL2A1 Human)
UseVector is designed for cloning and generation of …PromoterCOL2A1 regulatory elementsAvailable SinceOct. 13, 2017AvailabilityAcademic Institutions and Nonprofits only -
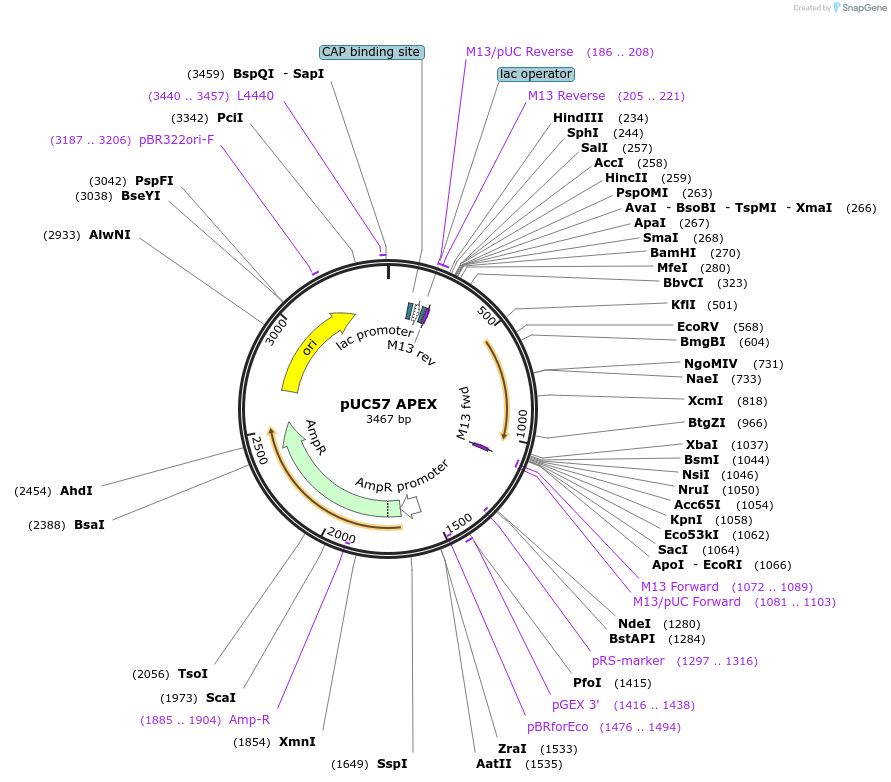
pUC57 APEX
Plasmid#40306PurposeA monomeric and improved-activity variant of pea ascorbate peroxidase. Used for electron microscopy and live-cell proteomic mapping. This construct has been replaced by APEX2 (construct 49386).DepositorInsertpea APEX (Synthetic)
UseUnspecifiedMutationK14D, W41F, E112K (relative to wild-type)Available SinceSept. 5, 2012AvailabilityAcademic Institutions and Nonprofits only -
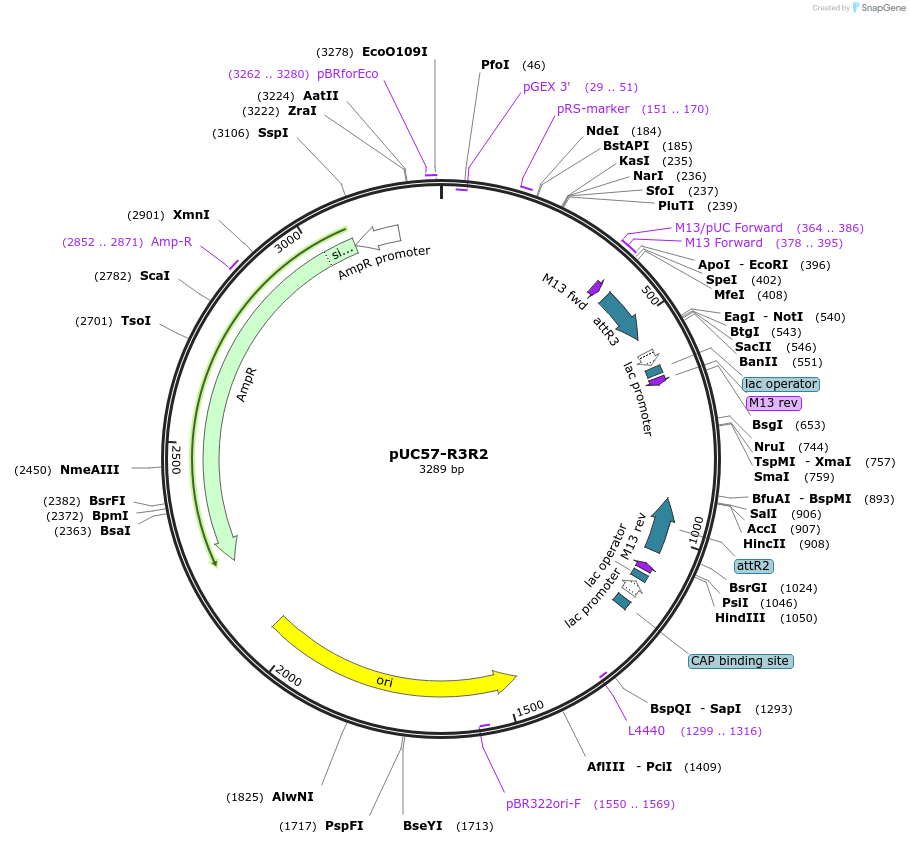
pUC57-R3R2
Plasmid#105109Purposecreate your own 2in1 constructsDepositorTypeEmpty backboneUseOtherAvailable SinceFeb. 22, 2018AvailabilityAcademic Institutions and Nonprofits only -
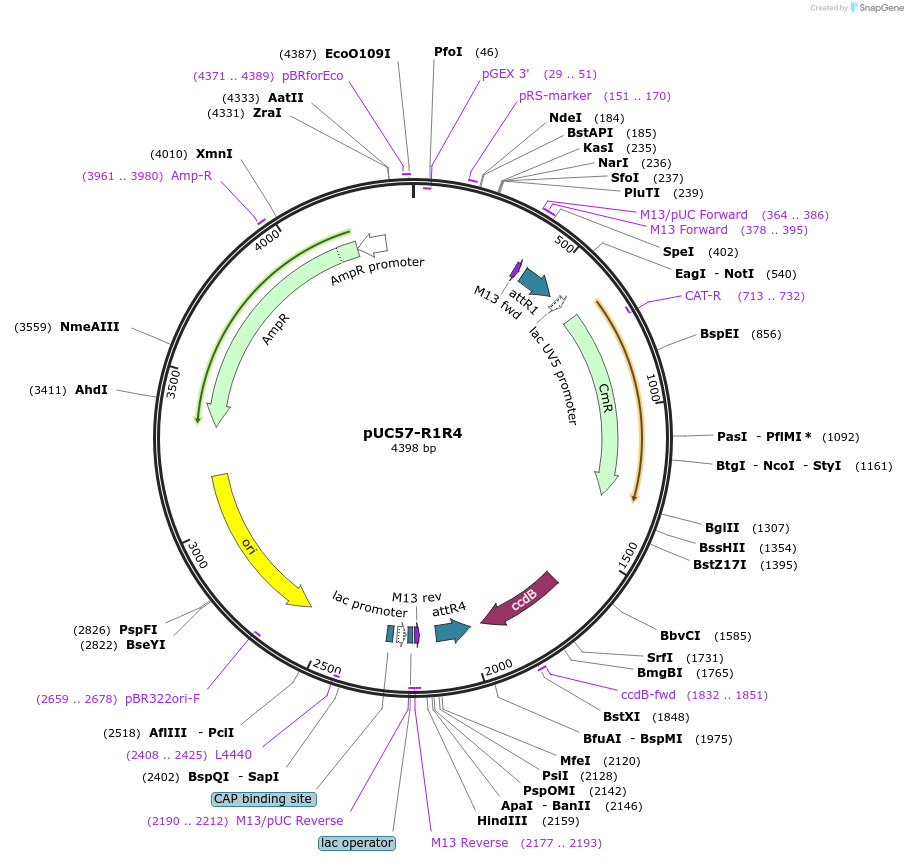
pUC57-R1R4
Plasmid#105110Purposecreate your own 2in1 constructsDepositorTypeEmpty backboneUseOtherAvailable SinceFeb. 22, 2018AvailabilityAcademic Institutions and Nonprofits only -
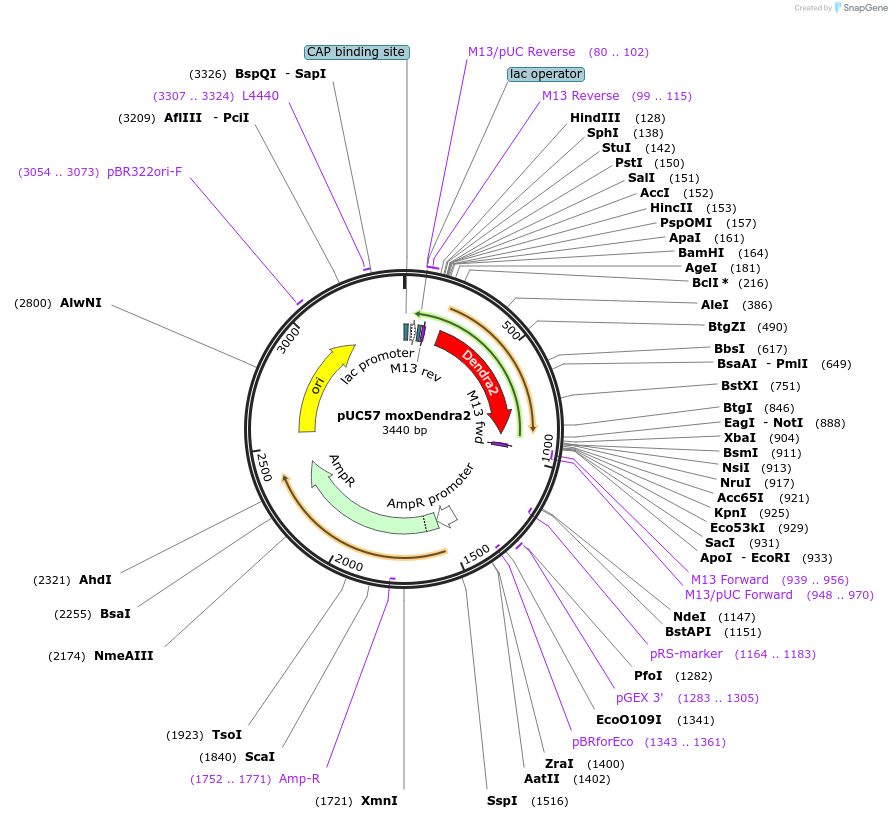
pUC57 moxDendra2
Plasmid#89791Purposebacterial expression of moxDendra2DepositorInsertmoxDendra2
UseCloning vectorAvailable SinceApril 24, 2017AvailabilityAcademic Institutions and Nonprofits only -
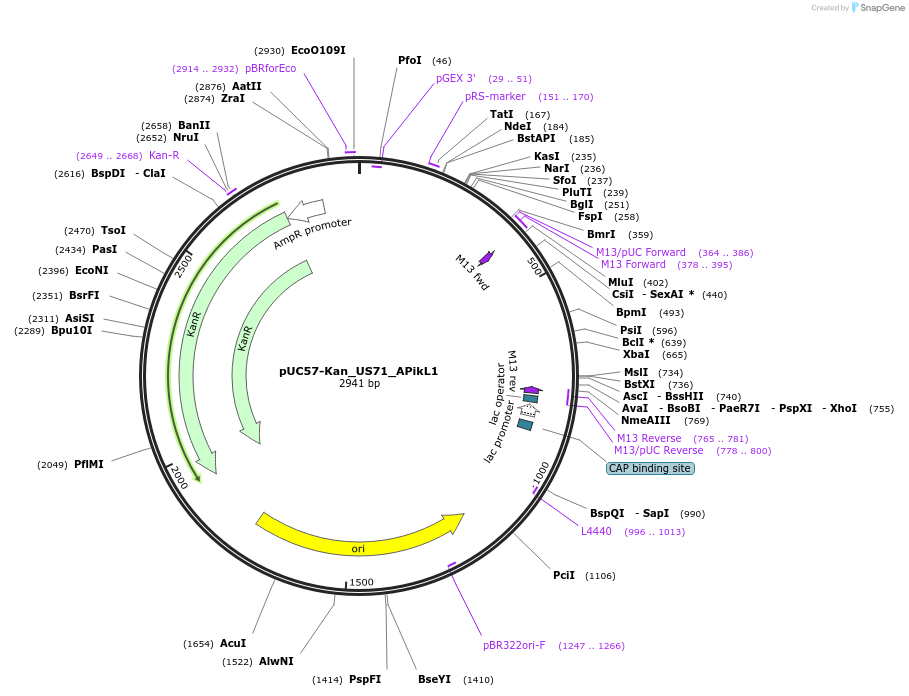
pUC57-Kan_US71_APikL1
Plasmid#123646PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertUS71_APikL1
UseGolden gateAvailable SinceSept. 12, 2019AvailabilityAcademic Institutions and Nonprofits only -
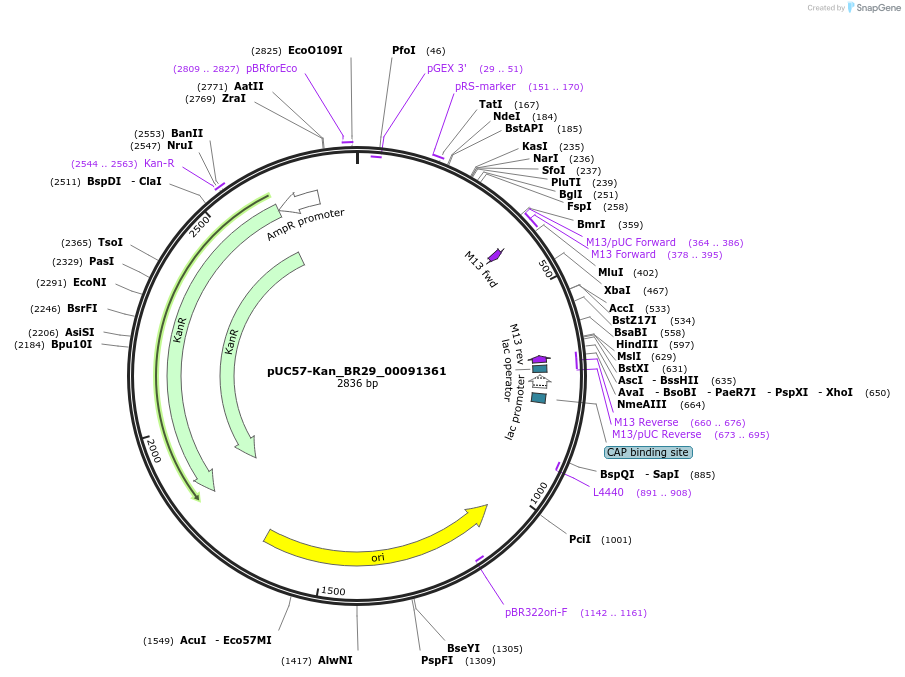
pUC57-Kan_BR29_00091361
Plasmid#121209PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertBR29_00091361
UseGolden gateMutationMagnaporthe griseaAvailable SinceMarch 7, 2019AvailabilityAcademic Institutions and Nonprofits only -
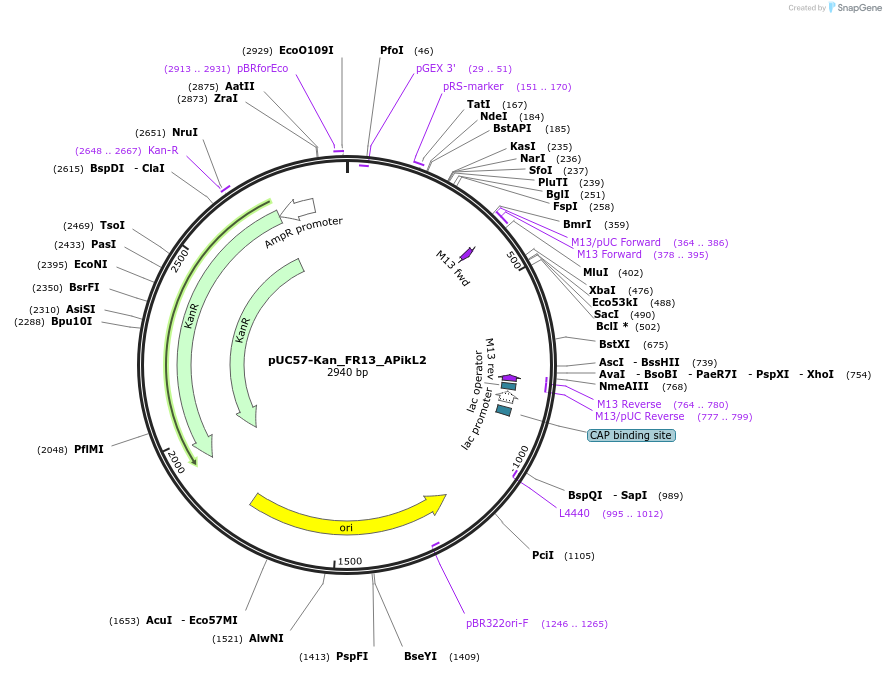
pUC57-Kan_FR13_APikL2
Plasmid#123648PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertFR13_APikL2
UseGolden gateAvailable SinceMay 22, 2019AvailabilityAcademic Institutions and Nonprofits only -
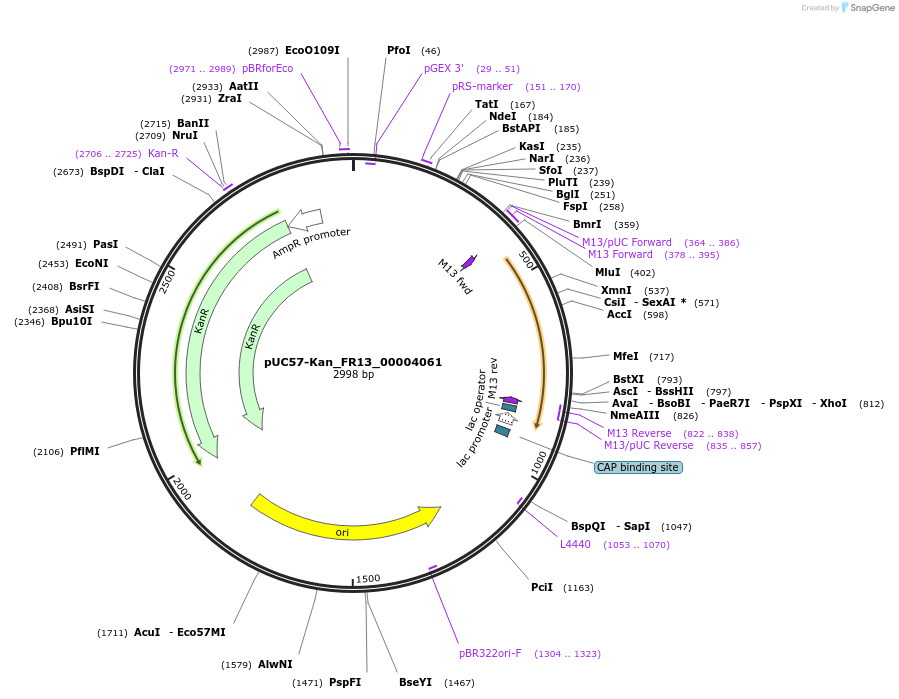
pUC57-Kan_FR13_00004061
Plasmid#121292PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertFR13_00004061
UseGolden gateMutationMagnaporthe oryzaeAvailable SinceMarch 12, 2019AvailabilityAcademic Institutions and Nonprofits only -
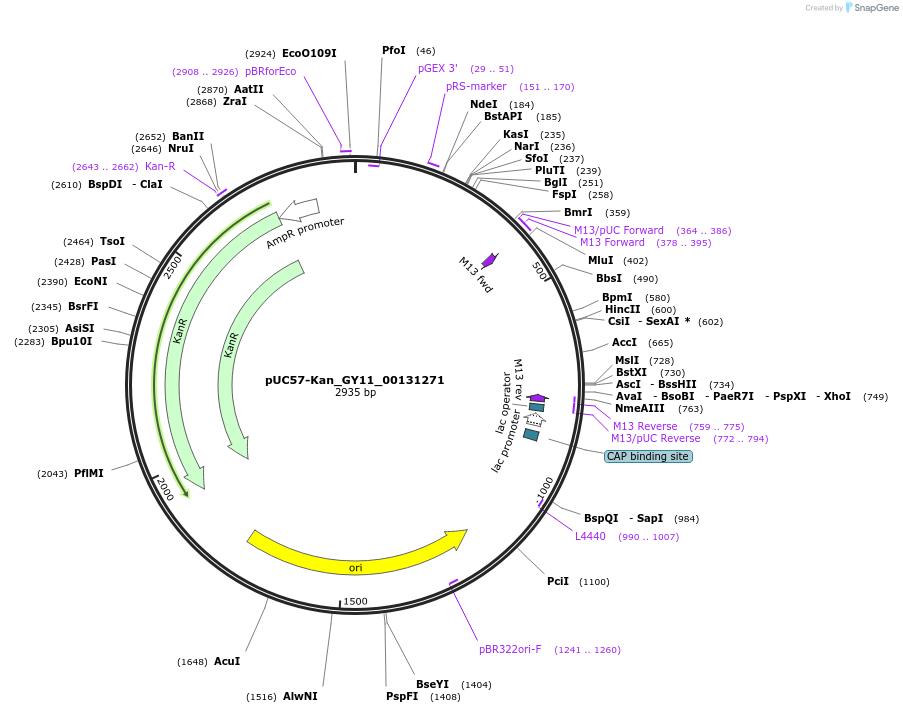
pUC57-Kan_GY11_00131271
Plasmid#121355PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertGY11_00131271
UseGolden gateMutationMagnaporthe oryzaeAvailable SinceMarch 14, 2019AvailabilityAcademic Institutions and Nonprofits only -
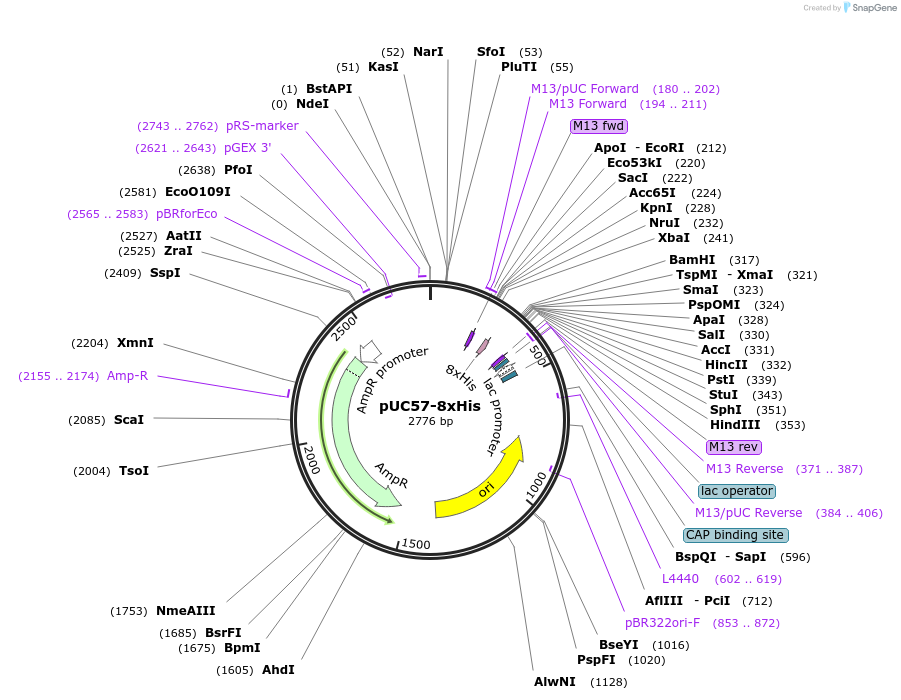
pUC57-8xHis
Plasmid#101699Purpose8xHis in domestication vectorDepositorInsertAATG-[8xHis]-AGCC
UseYeast expressionAvailable SinceJuly 24, 2018AvailabilityAcademic Institutions and Nonprofits only -
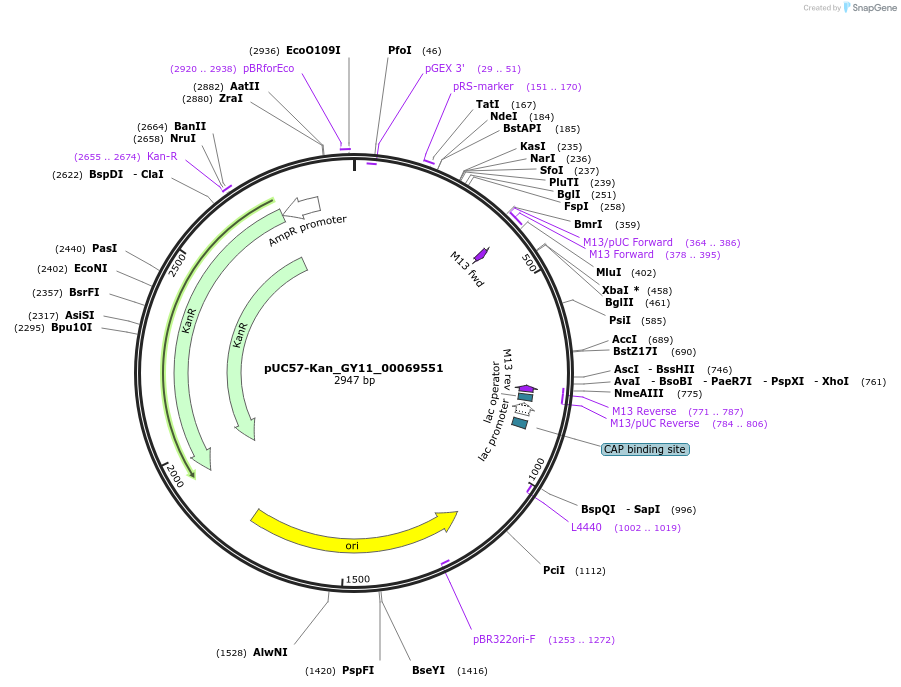
pUC57-Kan_GY11_00069551
Plasmid#121341PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertGY11_00069551
UseGolden gateMutationMagnaporthe oryzaeAvailable SinceMarch 14, 2019AvailabilityAcademic Institutions and Nonprofits only -
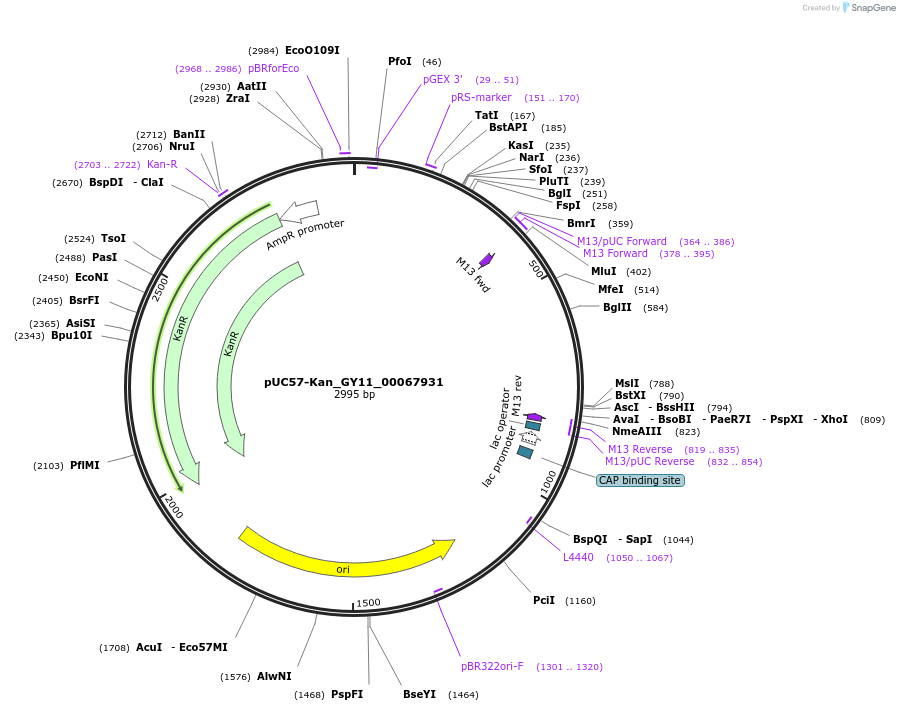
pUC57-Kan_GY11_00067931
Plasmid#121340PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertGY11_00067931
UseGolden gateMutationMagnaporthe oryzaeAvailable SinceMarch 14, 2019AvailabilityAcademic Institutions and Nonprofits only -
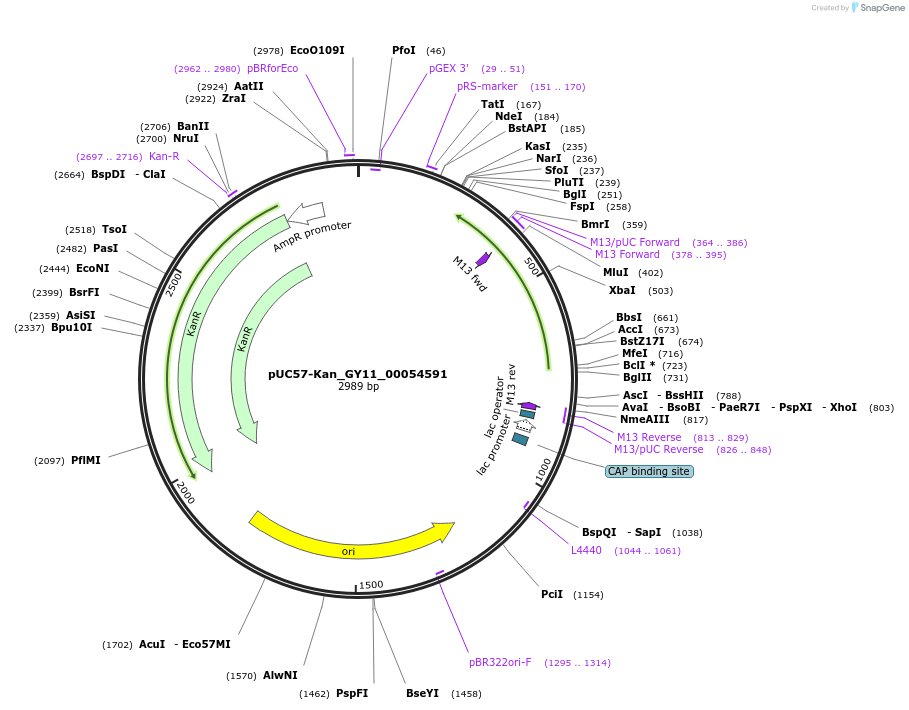
pUC57-Kan_GY11_00054591
Plasmid#121338PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertGY11_00054591
UseGolden gateMutationMagnaporthe oryzaeAvailable SinceMarch 14, 2019AvailabilityAcademic Institutions and Nonprofits only -
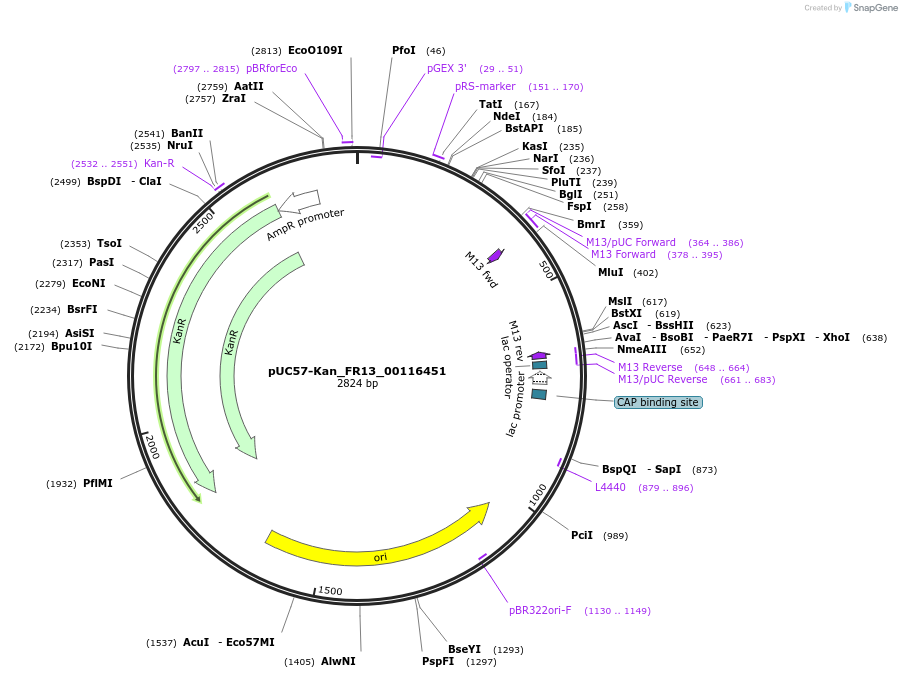
pUC57-Kan_FR13_00116451
Plasmid#121327PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertFR13_00116451
UseGolden gateMutationMagnaporthe oryzaeAvailable SinceMarch 14, 2019AvailabilityAcademic Institutions and Nonprofits only -
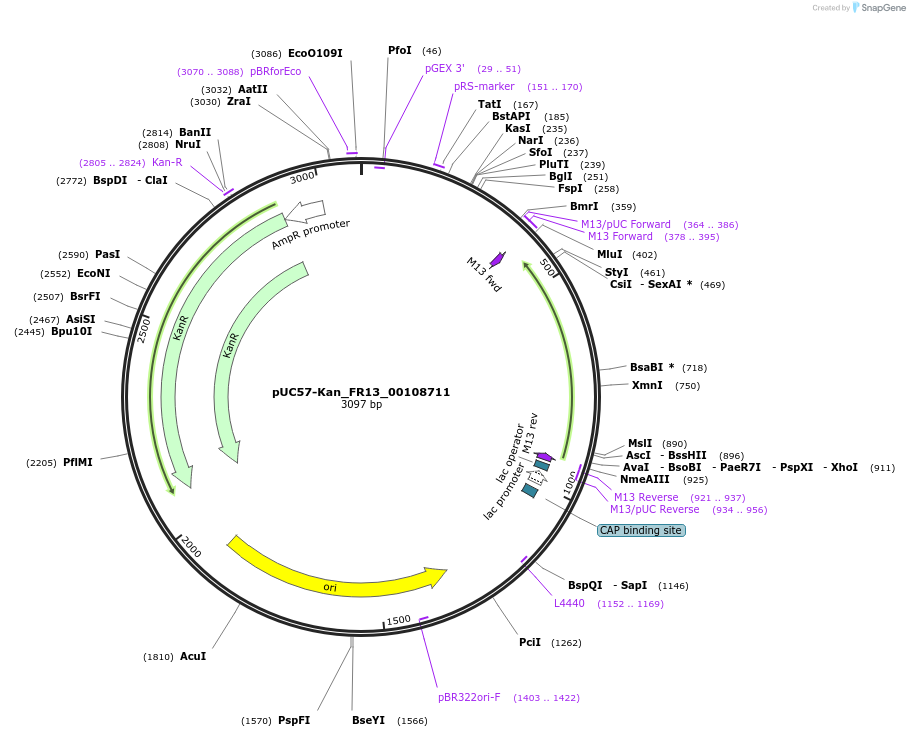
pUC57-Kan_FR13_00108711
Plasmid#121323PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertFR13_00108711
UseGolden gateMutationMagnaporthe oryzaeAvailable SinceMarch 14, 2019AvailabilityAcademic Institutions and Nonprofits only -
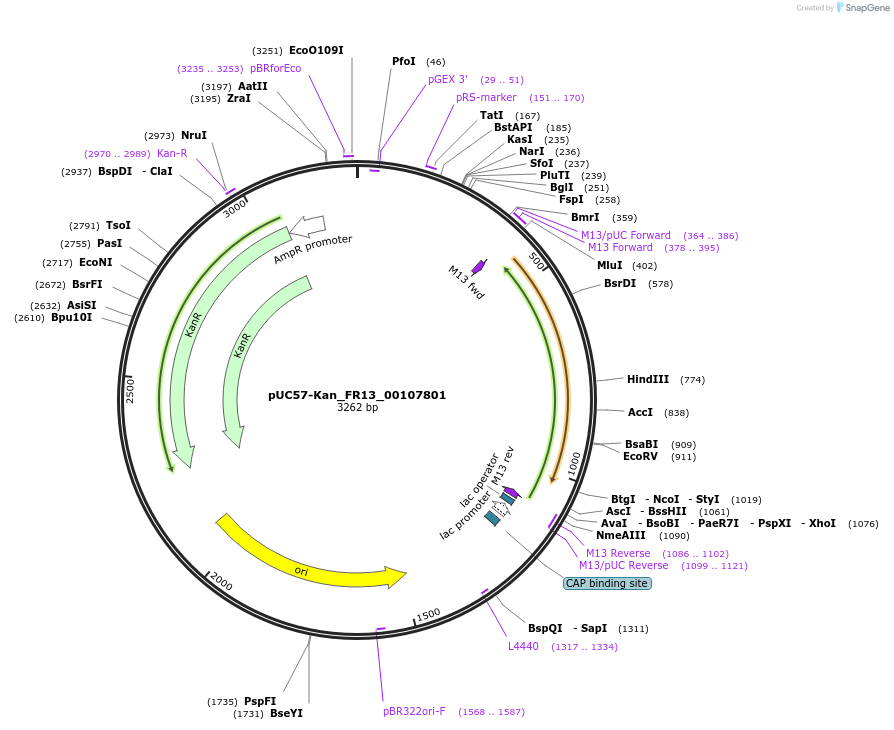
pUC57-Kan_FR13_00107801
Plasmid#121322PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertFR13_00107801
UseGolden gateMutationMagnaporthe oryzaeAvailable SinceMarch 14, 2019AvailabilityAcademic Institutions and Nonprofits only -
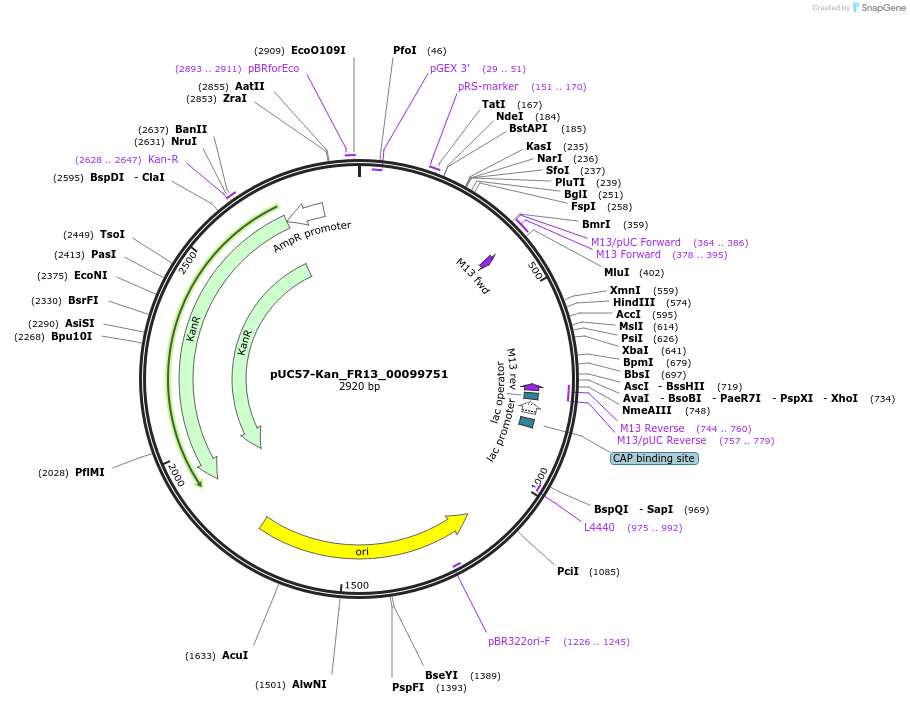
pUC57-Kan_FR13_00099751
Plasmid#121317PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertFR13_00099751
UseGolden gateMutationMagnaporthe oryzaeAvailable SinceMarch 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
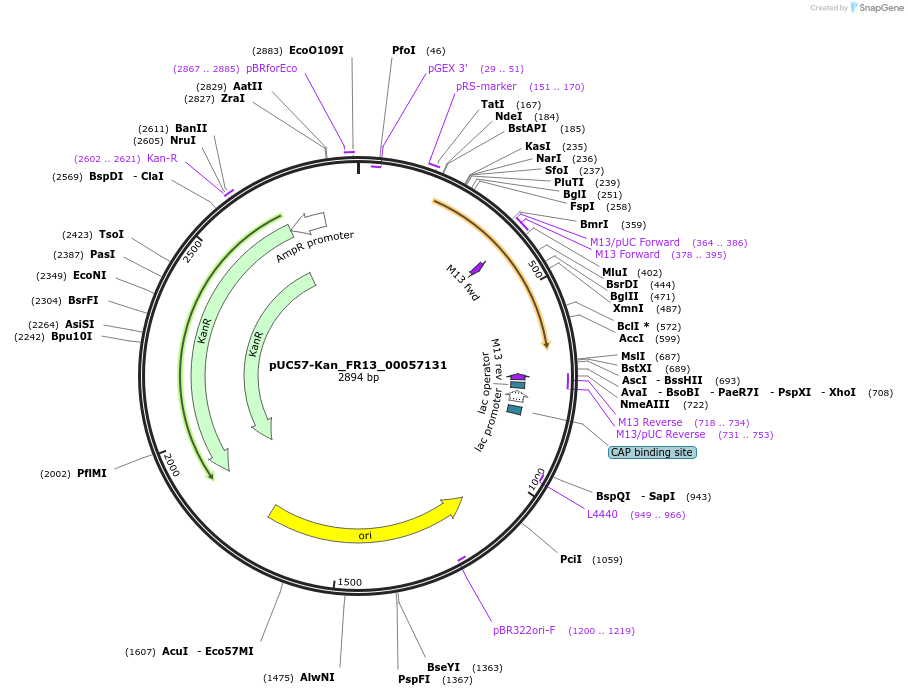
pUC57-Kan_FR13_00057131
Plasmid#121300PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertFR13_00057131
UseGolden gateMutationMagnaporthe oryzaeAvailable SinceMarch 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
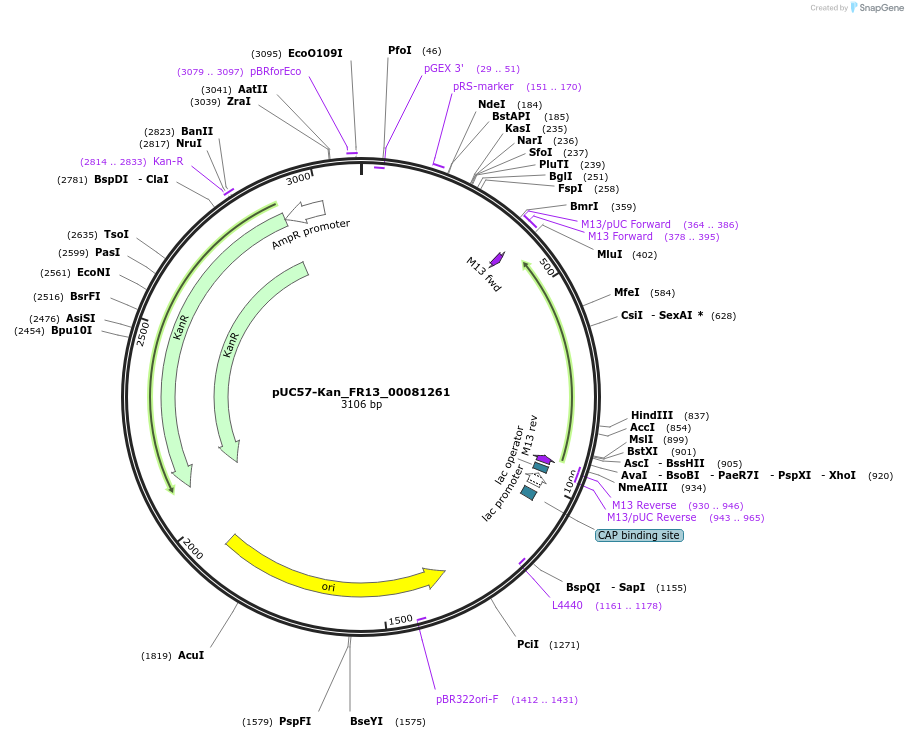
pUC57-Kan_FR13_00081261
Plasmid#121308PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertFR13_00081261
UseGolden gateMutationMagnaporthe oryzaeAvailable SinceMarch 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
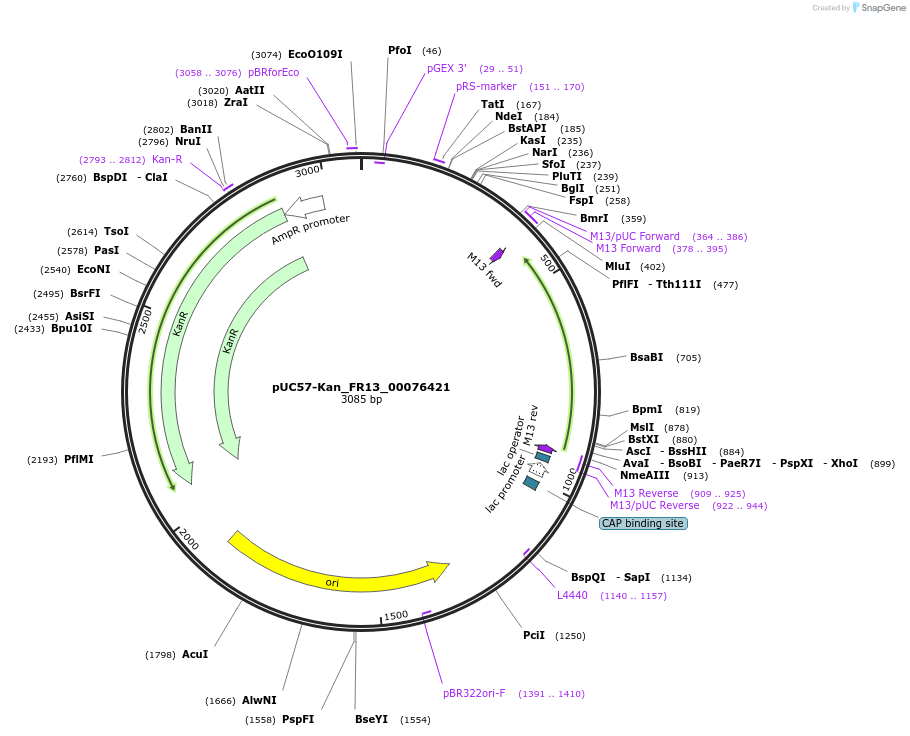
pUC57-Kan_FR13_00076421
Plasmid#121306PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertFR13_00076421
UseGolden gateMutationMagnaporthe oryzaeAvailable SinceMarch 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
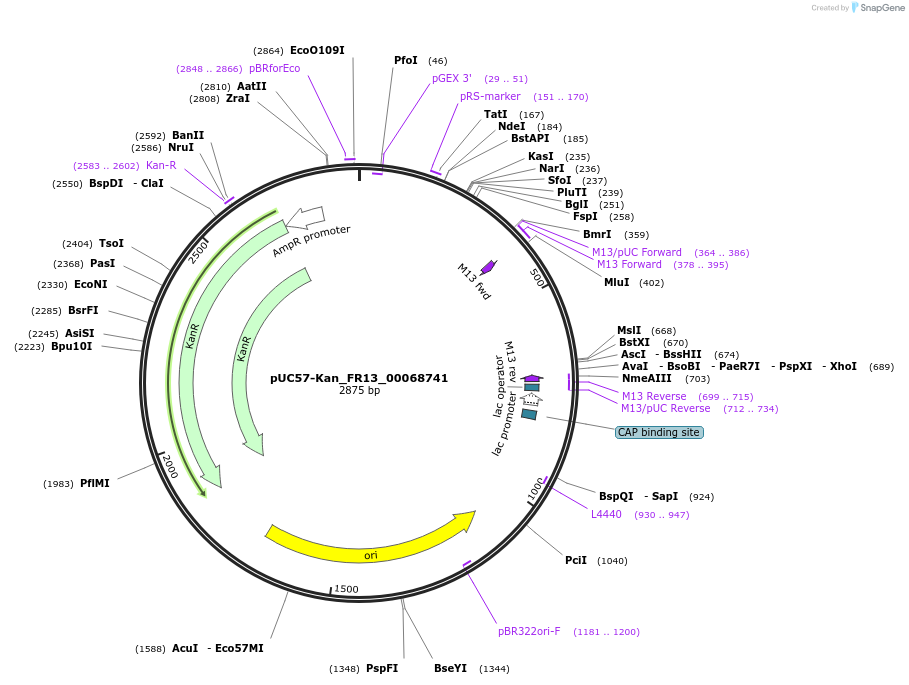
pUC57-Kan_FR13_00068741
Plasmid#121303PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertFR13_00068741
UseGolden gateMutationMagnaporthe oryzaeAvailable SinceMarch 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
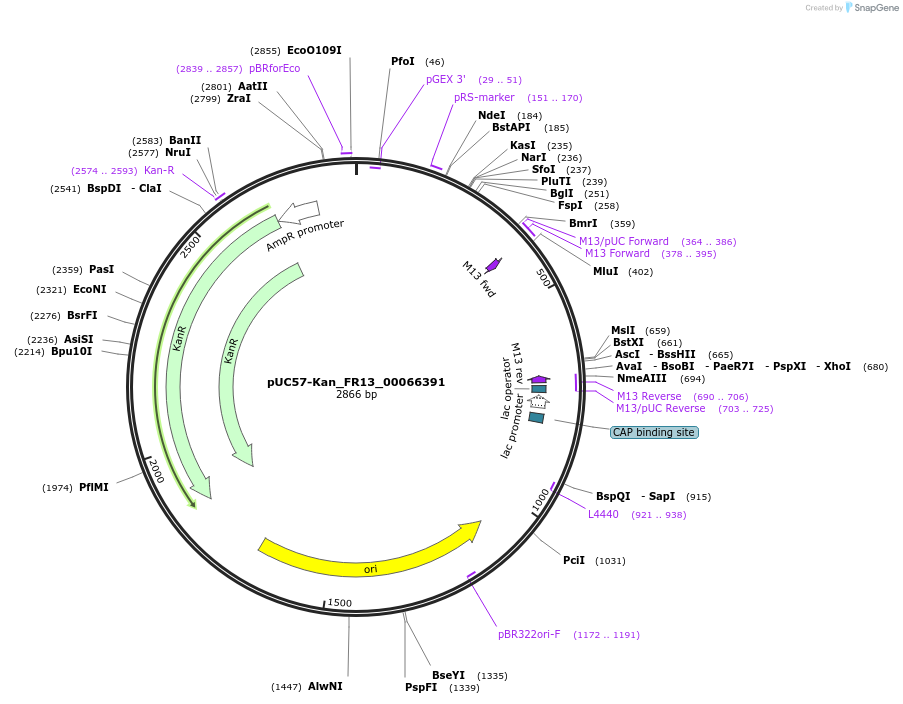
pUC57-Kan_FR13_00066391
Plasmid#121301PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertFR13_00066391
UseGolden gateMutationMagnaporthe oryzaeAvailable SinceMarch 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
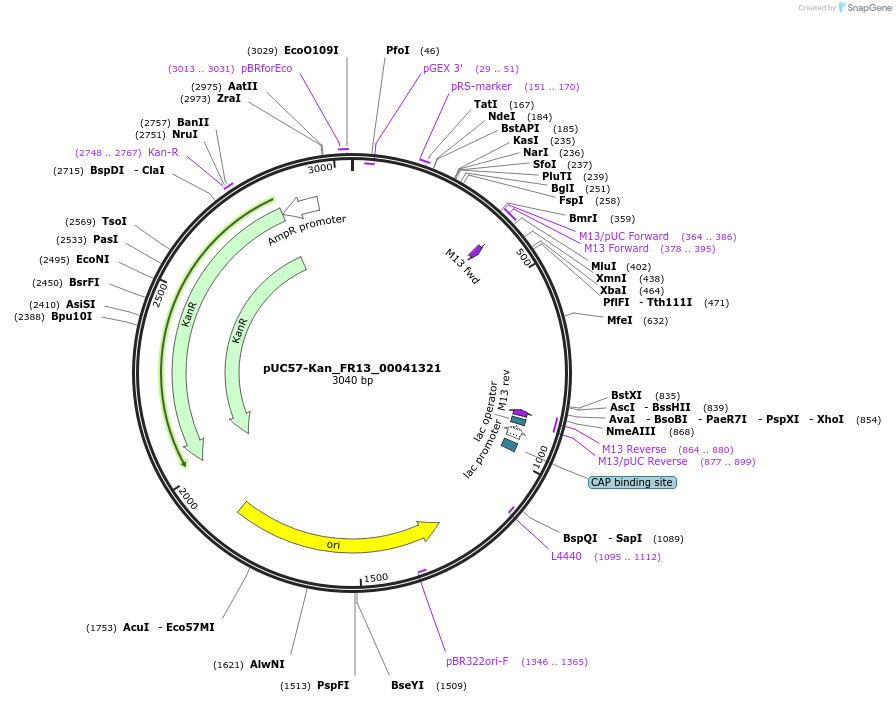
pUC57-Kan_FR13_00041321
Plasmid#121298PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertFR13_00041321
UseGolden gateMutationMagnaporthe oryzaeAvailable SinceMarch 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
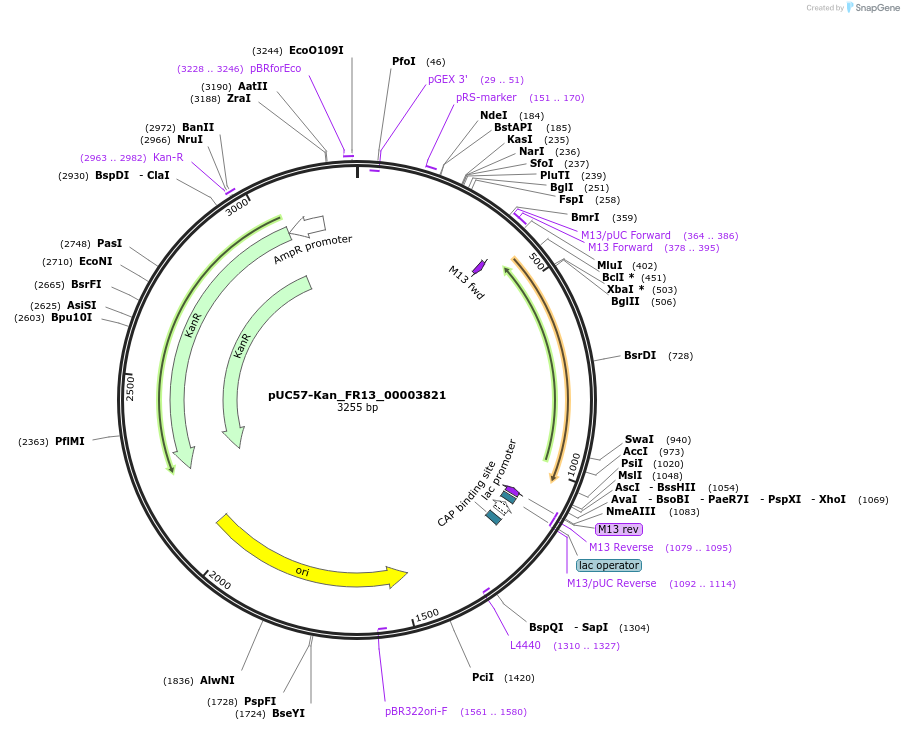
pUC57-Kan_FR13_00003821
Plasmid#121291PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertFR13_00003821
UseGolden gateMutationMagnaporthe oryzaeAvailable SinceMarch 12, 2019AvailabilityAcademic Institutions and Nonprofits only -
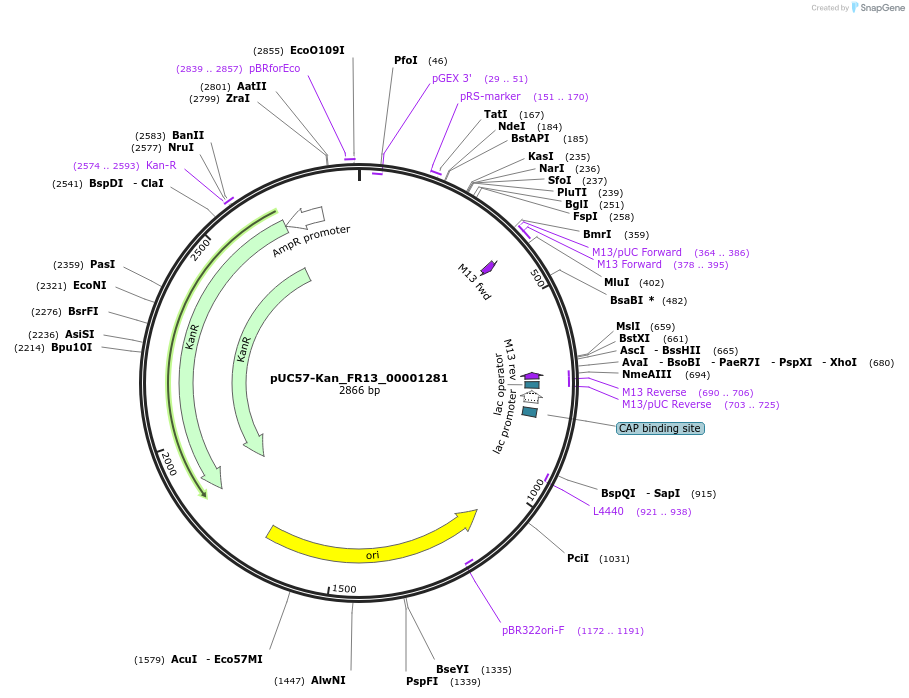
pUC57-Kan_FR13_00001281
Plasmid#121290PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertFR13_00001281
UseGolden gateMutationMagnaporthe oryzaeAvailable SinceMarch 12, 2019AvailabilityAcademic Institutions and Nonprofits only -
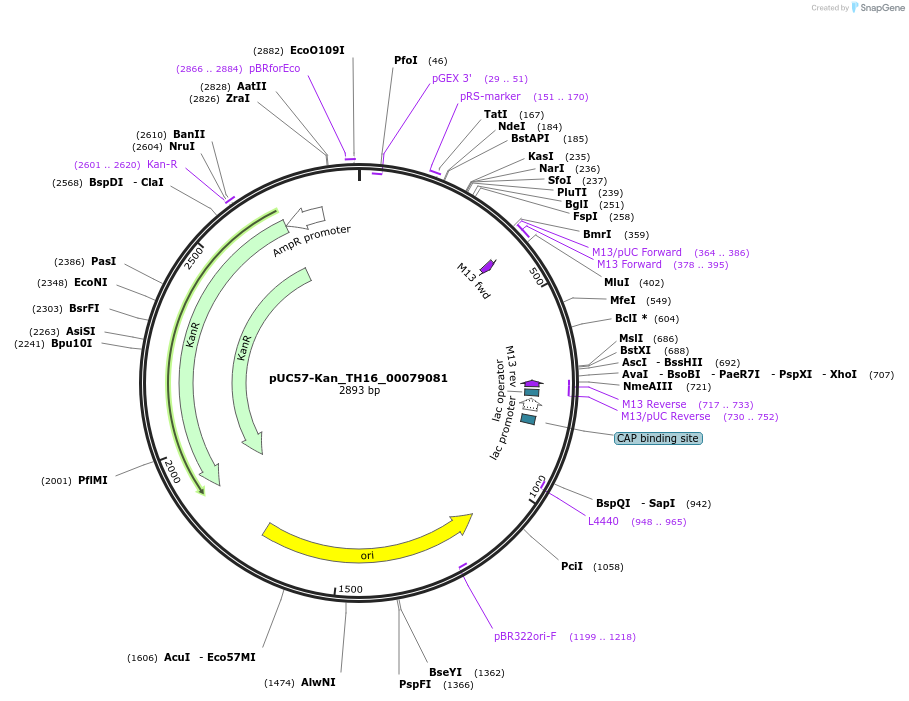
pUC57-Kan_TH16_00079081
Plasmid#121372PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertTH16_00079081
UseGolden gateMutationMagnaporthe oryzaeAvailable SinceSept. 4, 2019AvailabilityAcademic Institutions and Nonprofits only -
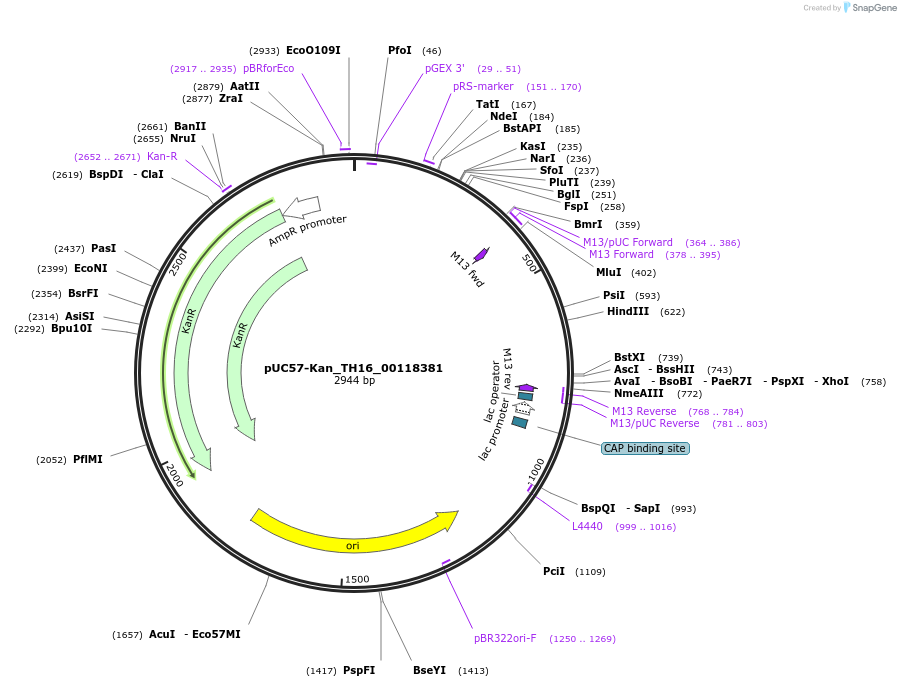
pUC57-Kan_TH16_00118381
Plasmid#121374PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertTH16_00118381
UseGolden gateMutationMagnaporthe oryzaeAvailable SinceMarch 15, 2019AvailabilityAcademic Institutions and Nonprofits only -
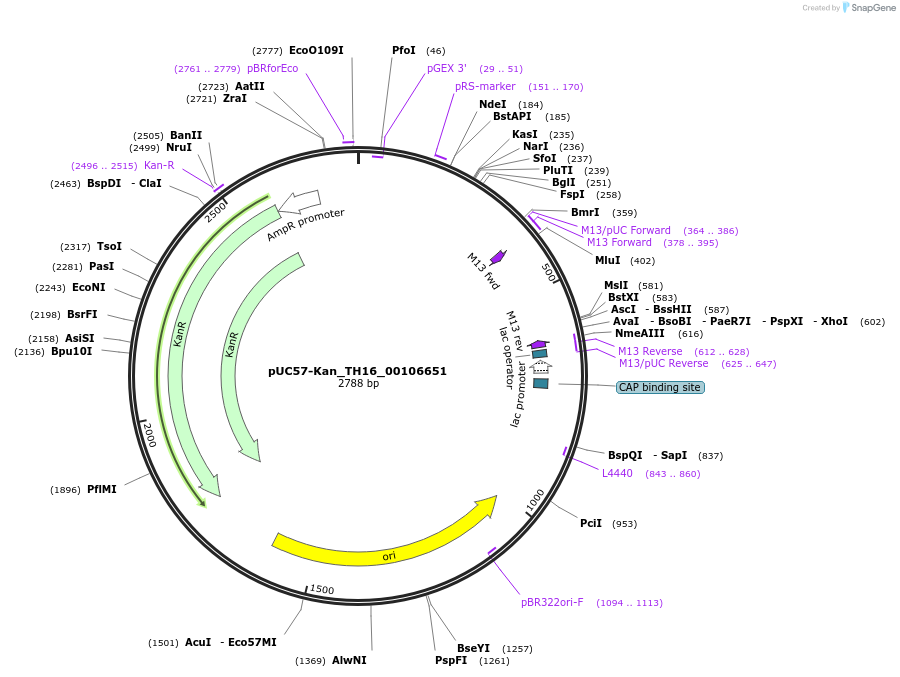
pUC57-Kan_TH16_00106651
Plasmid#121373PurposeLevel 0 Golden-Gate compatible vectorDepositorInsertTH16_00106651
UseGolden gateMutationMagnaporthe oryzaeAvailable SinceMarch 15, 2019AvailabilityAcademic Institutions and Nonprofits only