We narrowed to 17,209 results for: Emb;
-
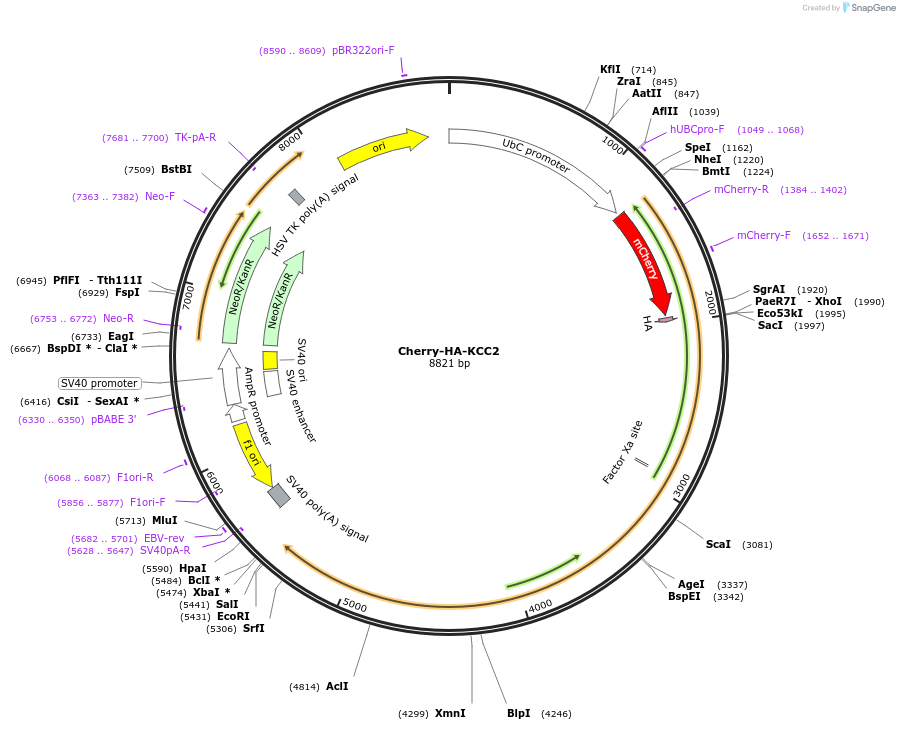
Plasmid#104077PurposeTo visualize the expression and localization of potassium-chloride cotransporter type 2 (KCC2)DepositorInsertKCC2 (b isoform) (Slc12a5 Rat)
TagsmCherry-HA tagExpressionMammalianMutationEcoR1 site in rat KCC2 was removed by silent muta…PromoterUbCAvailable SinceFeb. 20, 2018AvailabilityAcademic Institutions and Nonprofits only -
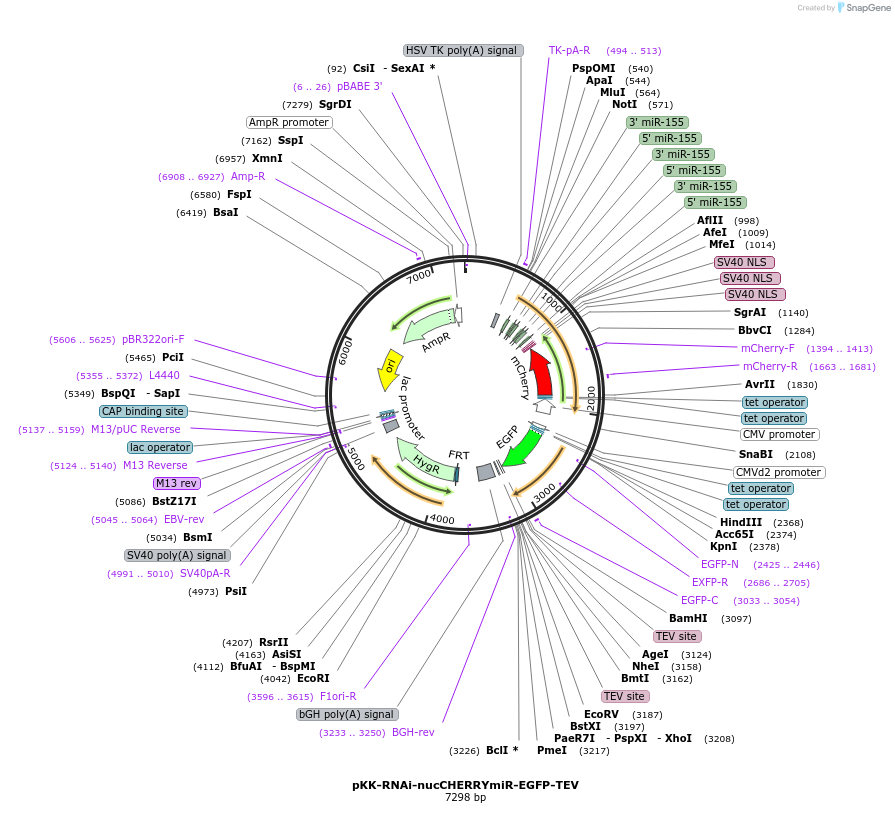
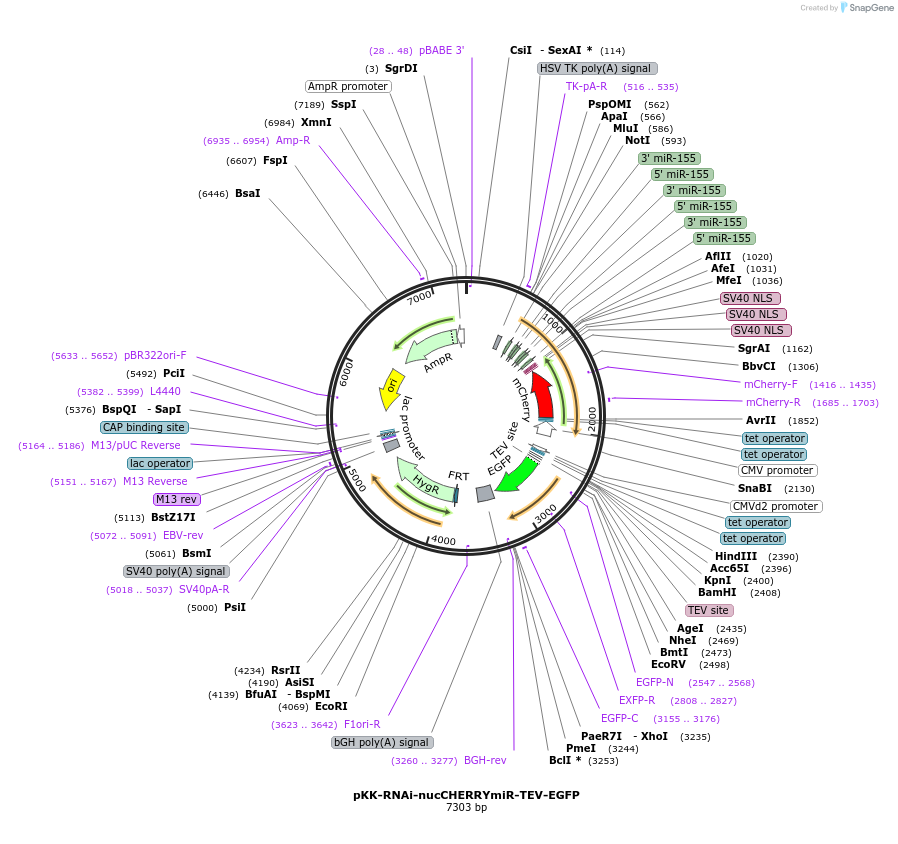
pKK-RNAi-nucCHERRYmiR-EGFP-TEV
Plasmid#105810PurposeFunctional studies using RNAi; rescue experiments; substituting protein for its different form. miRNA cassette with nuclear mCherry expression marker; your protein of interest with N-terminal EGFP.DepositorTypeEmpty backboneUseRNAi; Flp-in competentTagsEGFP-TEVExpressionMammalianAvailable SinceMarch 5, 2018AvailabilityAcademic Institutions and Nonprofits only -
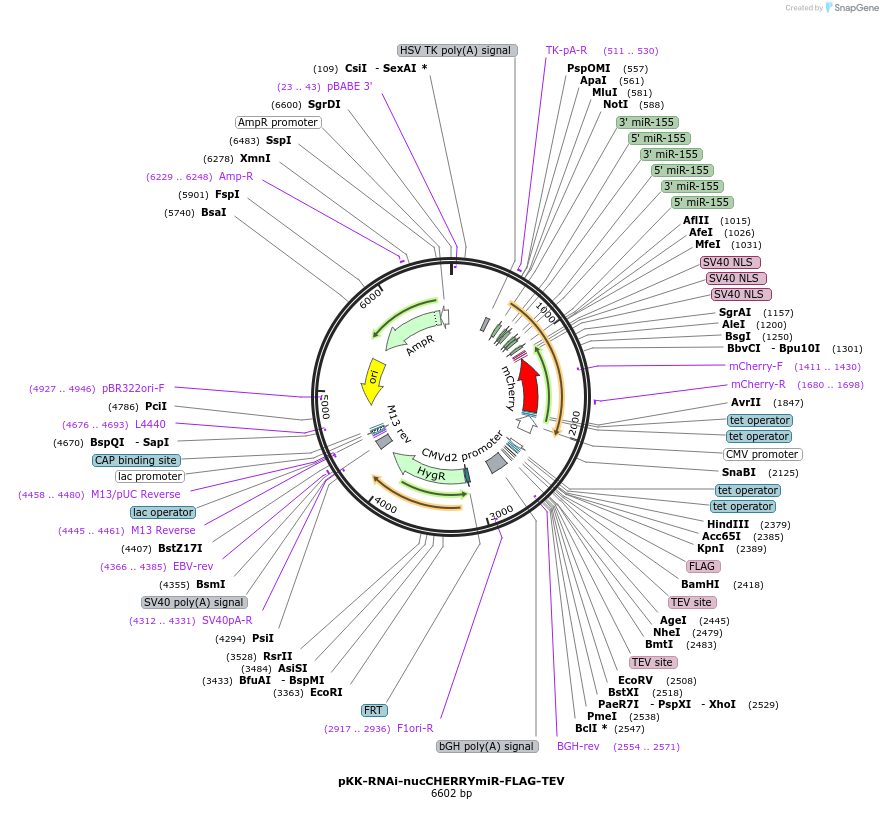
pKK-RNAi-nucCHERRYmiR-FLAG-TEV
Plasmid#105809PurposeFunctional studies using RNAi; rescue experiments; substituting protein for its different form. miRNA cassette with nuclear mCherry expression marker; your protein of interest with N-terminal FLAG.DepositorTypeEmpty backboneUseRNAi; Flp-in competentTagsFLAG-TEVExpressionMammalianAvailable SinceFeb. 13, 2018AvailabilityAcademic Institutions and Nonprofits only -
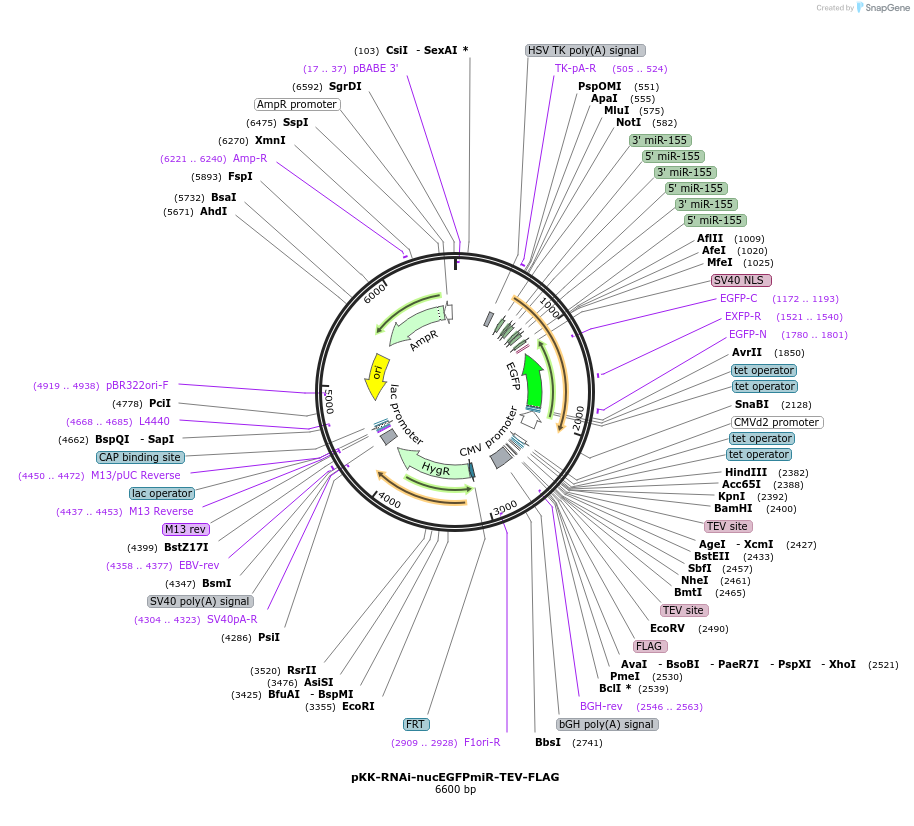
pKK-RNAi-nucEGFPmiR-TEV-FLAG
Plasmid#105811PurposeFunctional studies using RNAi; rescue experiments; substituting protein for its different form. miRNA cassette with nuclear EGFP expression marker; your protein of interest with C-terminal FLAG.DepositorTypeEmpty backboneUseRNAi; Flp-in competentTagsTEV-FLAGExpressionMammalianAvailable SinceMarch 5, 2018AvailabilityAcademic Institutions and Nonprofits only -
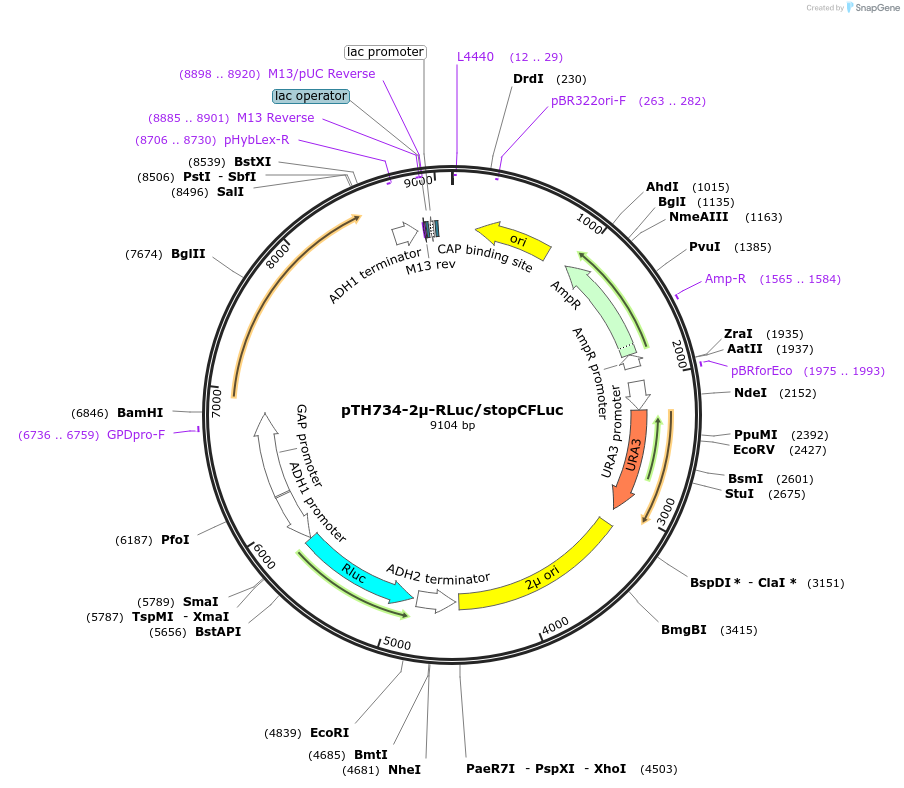
pTH734-2µ-RLuc/stopCFLuc
Plasmid#40609DepositorInsertsFirefly Luciferase (nonsense mutant)
Renilla luciferase
ExpressionYeastMutationAll codons of the original FLuc gene were exchang…PromoterADH1 and TDH3Available SinceOct. 30, 2012AvailabilityAcademic Institutions and Nonprofits only -
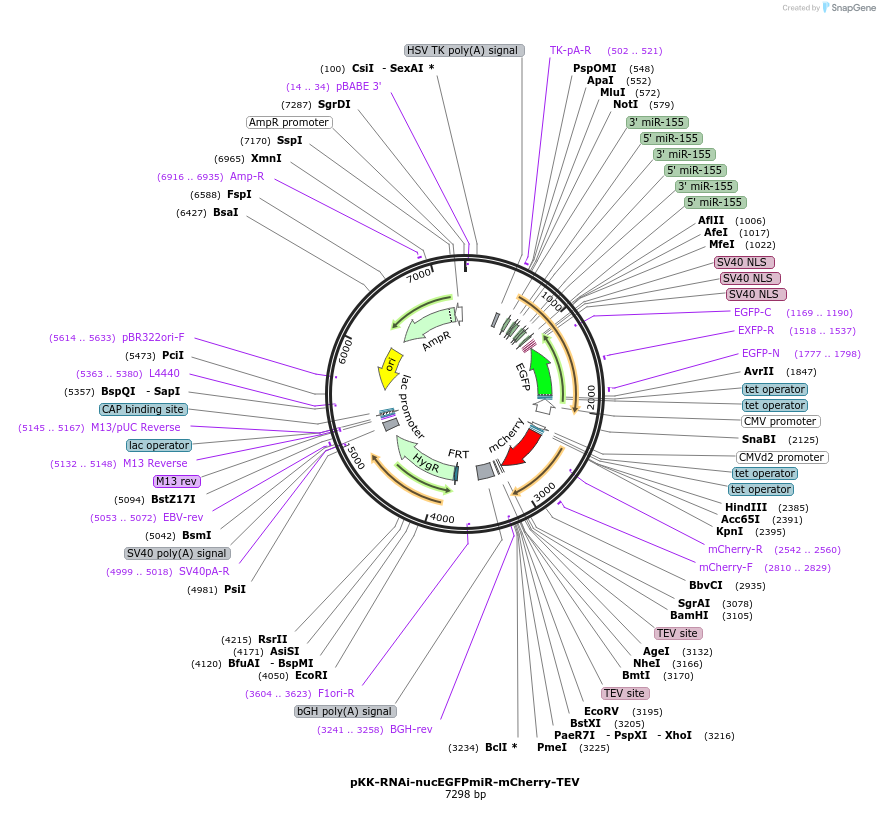
pKK-RNAi-nucEGFPmiR-mCherry-TEV
Plasmid#105808PurposeFunctional studies using RNAi; rescue experiments; substituting protein for its different form. miRNA cassette with nuclear EGFP expression marker; your protein of interest with N-terminal mCherry.DepositorTypeEmpty backboneUseRNAi; Flp-in competentTagsmCherry-TEVExpressionMammalianAvailable SinceMay 29, 2018AvailabilityAcademic Institutions and Nonprofits only -
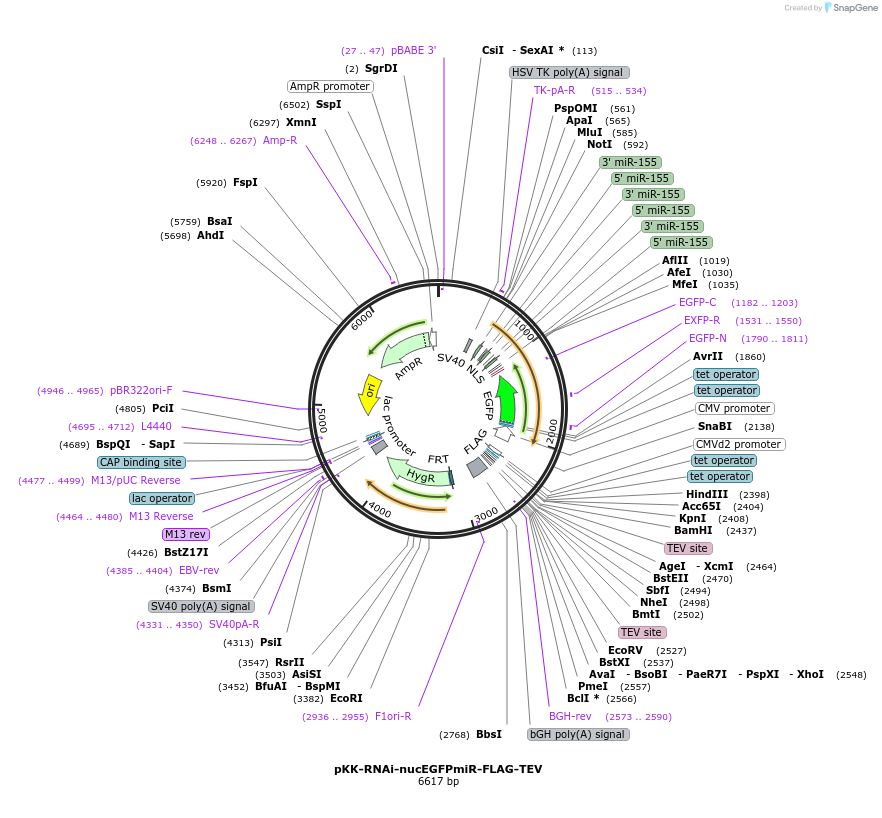
pKK-RNAi-nucEGFPmiR-FLAG-TEV
Plasmid#105807PurposeFunctional studies using RNAi; rescue experiments; substituting protein for its different form. miRNA cassette with nuclear EGFP expression marker; your protein of interest with N-terminal FLAG tag.DepositorTypeEmpty backboneUseRNAi; Flp-in competentTagsFLAG-TEVExpressionMammalianAvailable SinceFeb. 12, 2018AvailabilityAcademic Institutions and Nonprofits only -
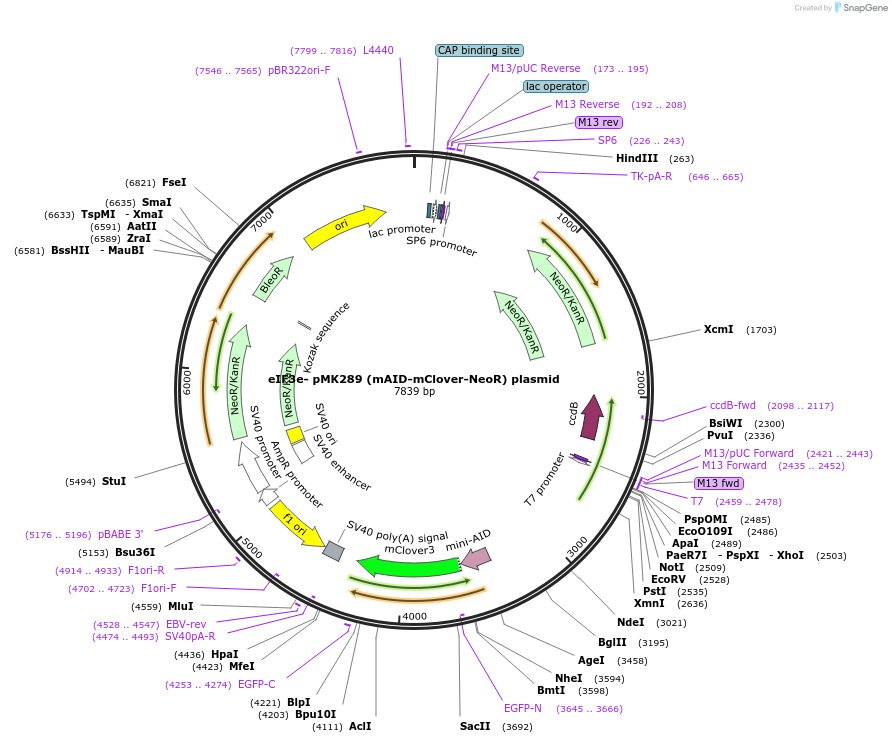
eIF3e- pMK289 (mAID-mClover-NeoR) plasmid
Plasmid#192239Purposeknockin donor vector of mAID-mClover-NeoR to C-terminus of endogenous human eIF3eDepositorInserteIF3e-knockin-homology arm-mAID-mClover-NeoR (EIF3E Human)
ExpressionMammalianAvailable SinceJan. 6, 2023AvailabilityAcademic Institutions and Nonprofits only -
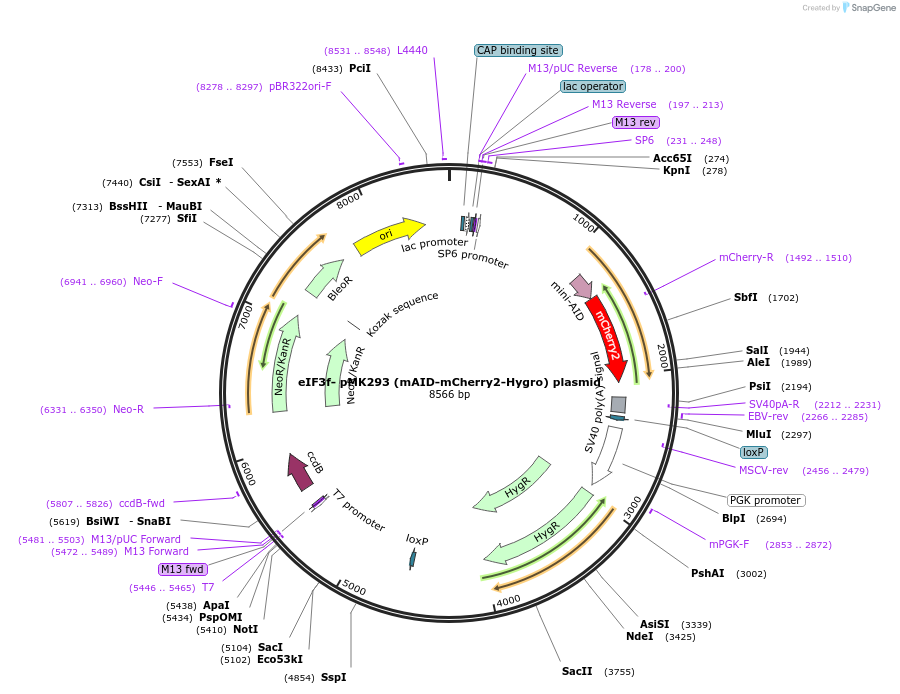
eIF3f- pMK293 (mAID-mCherry2-Hygro) plasmid
Plasmid#192242Purposeknockin donor vector of mAID-mCherry2-Hygro to C-terminus of endogenous human eIF3fDepositorInserteIF3f-knockin-homology arm-mAID-mCherry2-Hygro (EIF3F Human)
ExpressionMammalianAvailable SinceJan. 5, 2023AvailabilityAcademic Institutions and Nonprofits only -
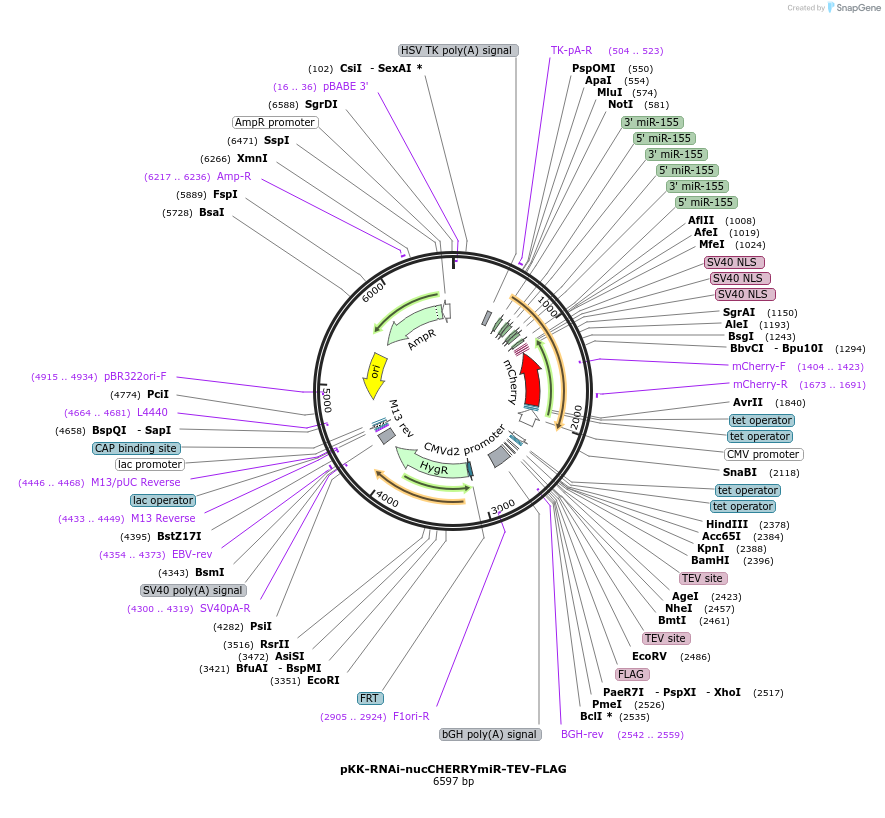
pKK-RNAi-nucCHERRYmiR-TEV-FLAG
Plasmid#105813PurposeFunctional studies using RNAi; rescue experiments; substituting protein for its different form. miRNA cassette with nuclear mCherry expression marker; your protein of interest with C-terminal FLAG.DepositorTypeEmpty backboneUseRNAi; Flp-in competentTagsTEV-FLAGExpressionMammalianAvailable SinceMarch 5, 2018AvailabilityAcademic Institutions and Nonprofits only -
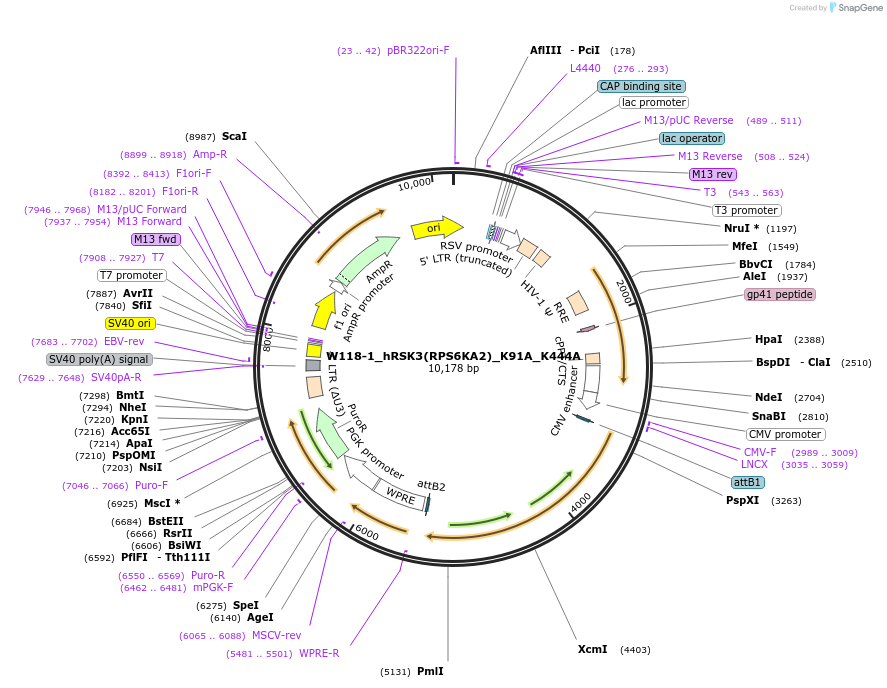
W118-1_hRSK3(RPS6KA2)_K91A_K444A
Plasmid#180385Purposelentiviral transduction of human RSK3 (RPS6KA2) gene with point mutations (K91A and K444A) that inactivate kinase activityDepositorInsertRPS6KA2_K91A/K444A (RPS6KA2 Human)
UseLentiviralExpressionMammalianMutationK91A/K444APromoterCMVAvailable SinceFeb. 9, 2022AvailabilityAcademic Institutions and Nonprofits only -
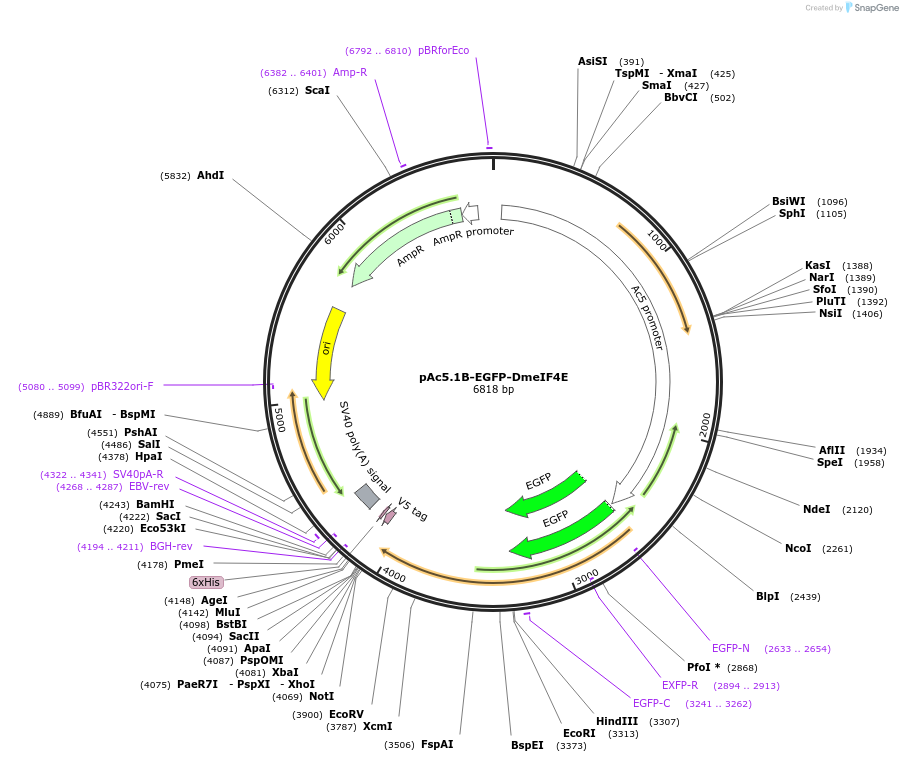
pAc5.1B-EGFP-DmeIF4E
Plasmid#79258PurposeExpresses EGFP-tagged DmeIF4E in S2 cellsDepositorAvailable SinceJuly 19, 2016AvailabilityAcademic Institutions and Nonprofits only -
NET23 pEGFP-N2 (1174)
Plasmid#62037Purposemammalian expression of nuclear envelope transmembrane proteinDepositorAvailable SinceAug. 17, 2015AvailabilityAcademic Institutions and Nonprofits only -
pKK-RNAi-nucCHERRYmiR-TEV-EGFP
Plasmid#105814PurposeFunctional studies using RNAi; rescue experiments; substituting protein for its different form. miRNA cassette with nuclear mCherry expression marker; your protein of interest with C-terminal EGFP.DepositorTypeEmpty backboneUseRNAi; Flp-in competentTagsTEV-EGFPExpressionMammalianAvailable SinceFeb. 12, 2018AvailabilityAcademic Institutions and Nonprofits only -
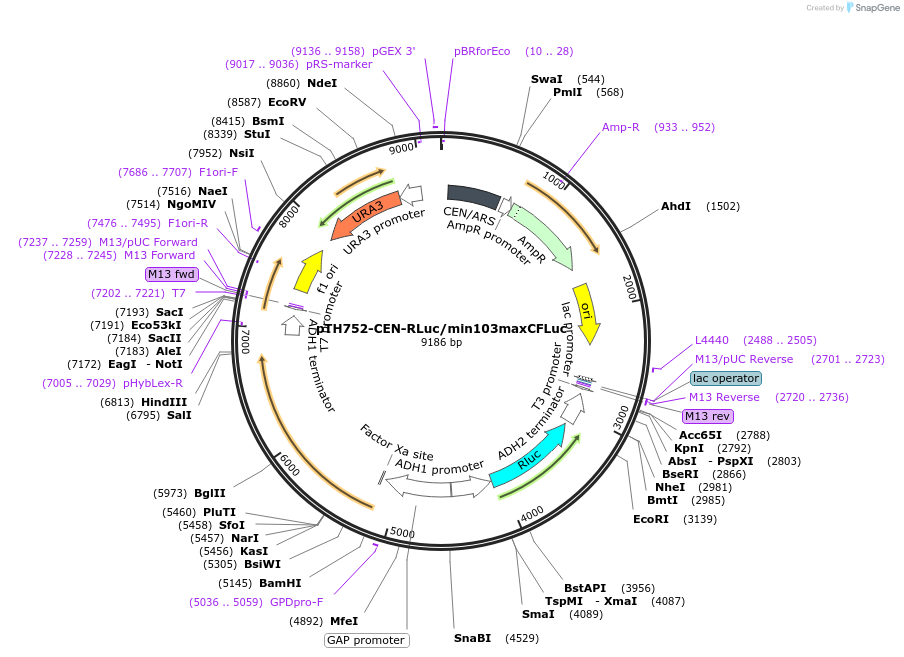
pTH752-CEN-RLuc/min103maxCFLuc
Plasmid#38218DepositorInsertsFirefly Luciferase
Renilla luciferase
ExpressionYeastMutationThe first 103 codons of the original FLuc gene we…PromoterADH1 and TDH3 (=GPD)Available SinceOct. 22, 2012AvailabilityAcademic Institutions and Nonprofits only -
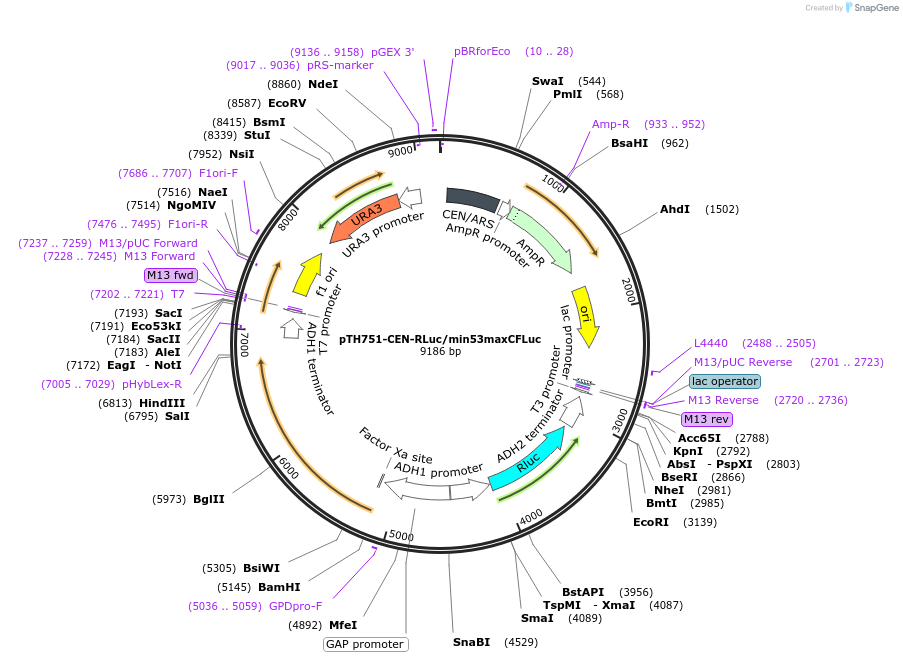
pTH751-CEN-RLuc/min53maxCFLuc
Plasmid#38217DepositorInsertsFirefly Luciferase
Renilla luciferase
ExpressionYeastMutationThe first 53 codons of the original FLuc gene wer…PromoterADH1 and TDH3 (=GPD)Available SinceOct. 22, 2012AvailabilityAcademic Institutions and Nonprofits only -
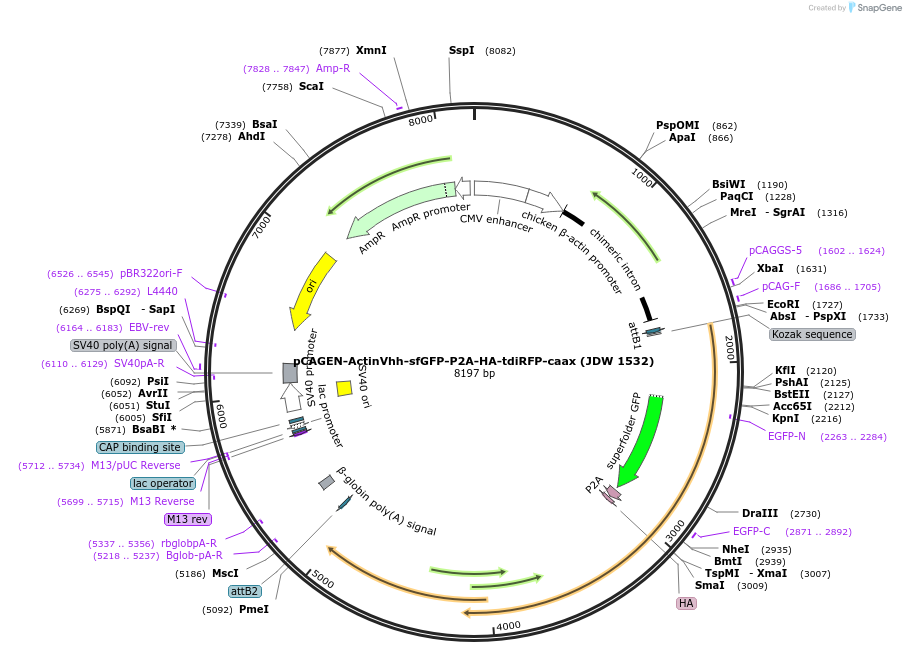
pCAGEN-ActinVhh-sfGFP-P2A-HA-tdiRFP-caax (JDW 1532)
Plasmid#242606PurposeA CAGGS driven expression vector containing an actin nanobody fused to sfGFP to label actin followed by a HA tagged tdiRFP fused to a caax tag to label the cell membrane.DepositorInsertActin Vhh-sfGFP-P2A-HA-tdiRFP-caax
ExpressionMammalianPromoterCAGGSAvailable SinceDec. 16, 2025AvailabilityAcademic Institutions and Nonprofits only -
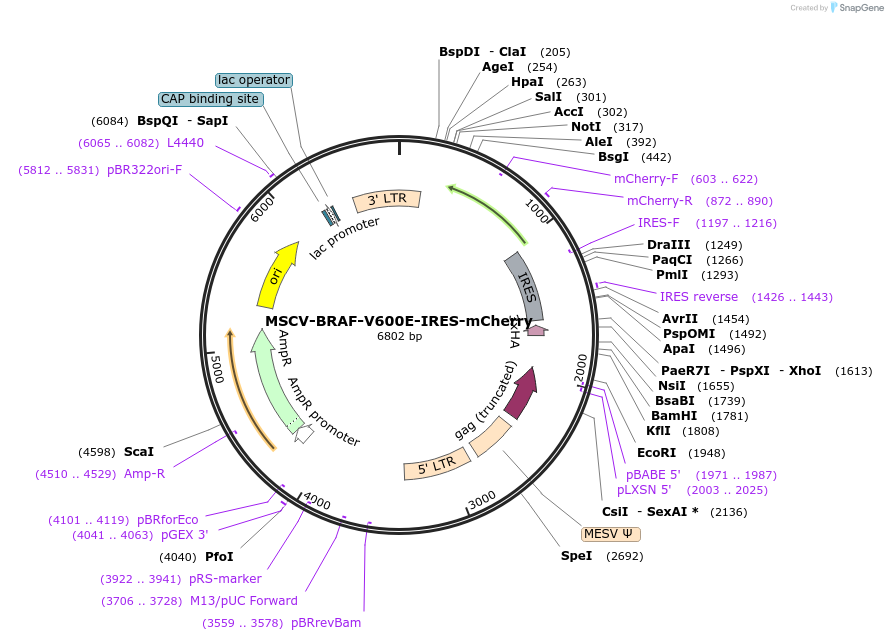
MSCV-BRAF-V600E-IRES-mCherry
Plasmid#221026PurposeFluorescent reporter for expressing a segment of BRAF-V600E CDSDepositorAvailable SinceAug. 13, 2025AvailabilityAcademic Institutions and Nonprofits only -
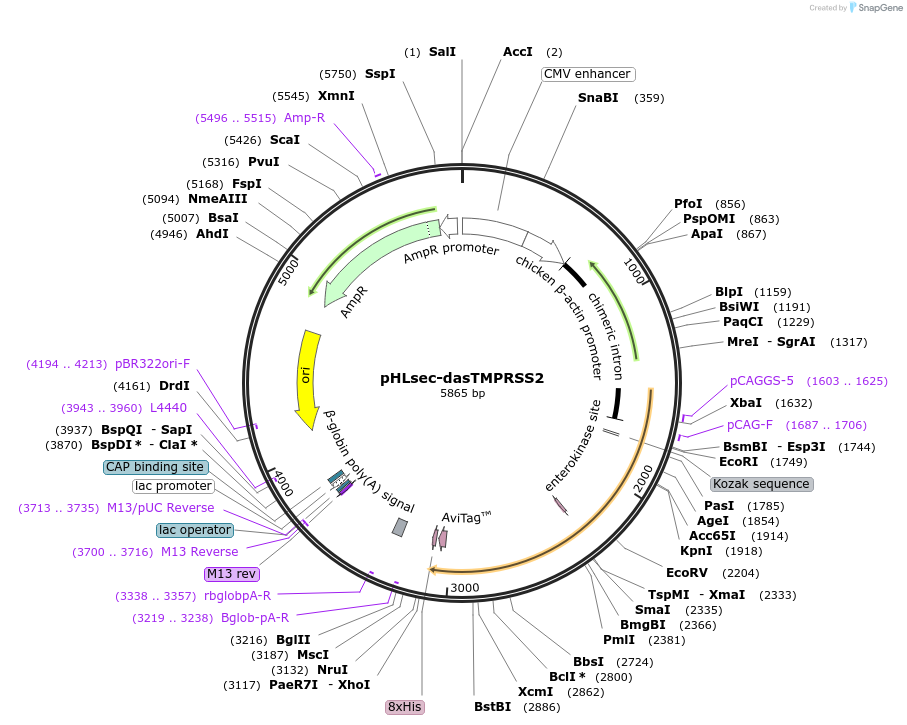
pHLsec-dasTMPRSS2
Plasmid#207850PurposeExpresses directed activation strategy TMPRSS2 in mammalian cells, such as HEK293F- or CHO-cellsDepositorInserttransmembrane serine protease 2 (TMPRSS2 Human)
TagsAviTag, 8xHIS tagExpressionMammalianMutationdeleted amino acid 1-108, S251D R252D Q253D S254D…PromoterCAGAvailable SinceMarch 25, 2025AvailabilityAcademic Institutions and Nonprofits only -
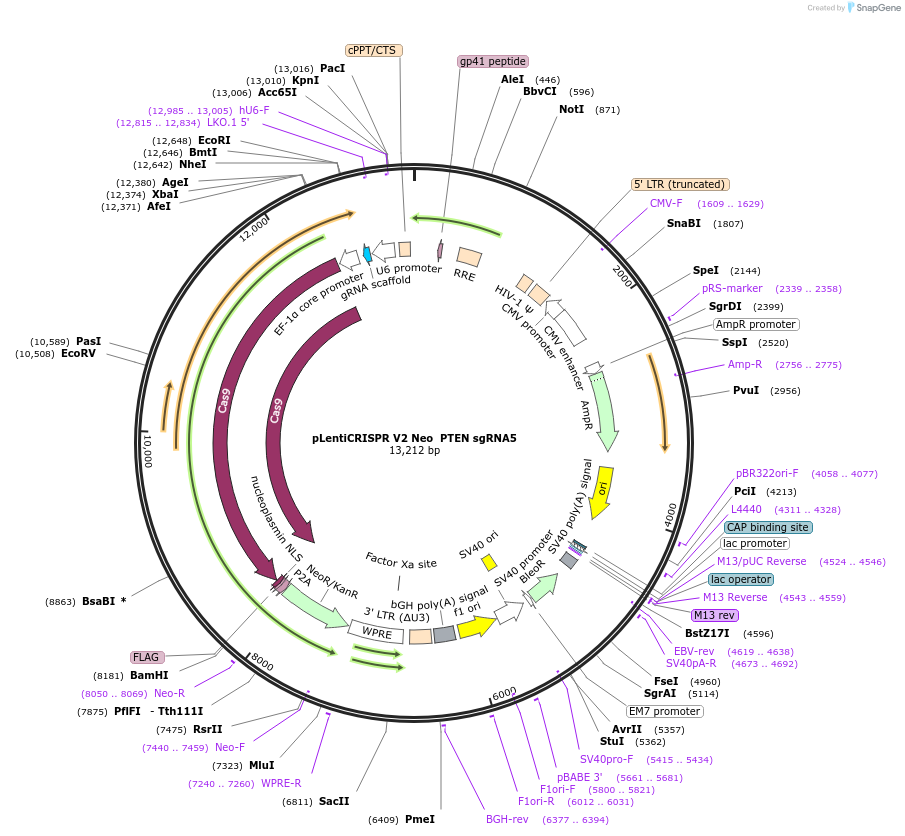
pLentiCRISPR V2 Neo PTEN sgRNA5
Plasmid#233246PurposeKnock out of murine PTENDepositorInsertPTEN (Pten Mouse)
UseLentiviralAvailable SinceMarch 20, 2025AvailabilityAcademic Institutions and Nonprofits only