We narrowed to 7,089 results for: CAD
-
Plasmid#48746PurposeExpression of luciferase driven by mPer2 promoter region (bases -1128 to +2129 with respect to transcription start site at +1)DepositorInsertmPer2 promoter/enhancer region (-1128 to +2129)
UseLuciferaseAvailable SinceDec. 19, 2013AvailabilityAcademic Institutions and Nonprofits only -
pHL1575
Plasmid#53028PurposePconNoHind:csrB, PLtetO-1:RBS(st3):csrDDepositorInsertscsrB
csrD
UseSynthetic BiologyTagsRBS (st3)ExpressionBacterialPromoterPconNoHind and pLtetO-1Available SinceMay 14, 2014AvailabilityAcademic Institutions and Nonprofits only -
pHL1756
Plasmid#53036PurposePconNoHindM12:glgC::gfp, PconNoHind:RBS(st3):tetRDepositorInsertsGFP
tetR
UseSynthetic BiologyTagsRBS(st3) and glgC leaderExpressionBacterialPromoterpConNoHind and pConNoHindM12Available SinceMay 14, 2014AvailabilityAcademic Institutions and Nonprofits only -
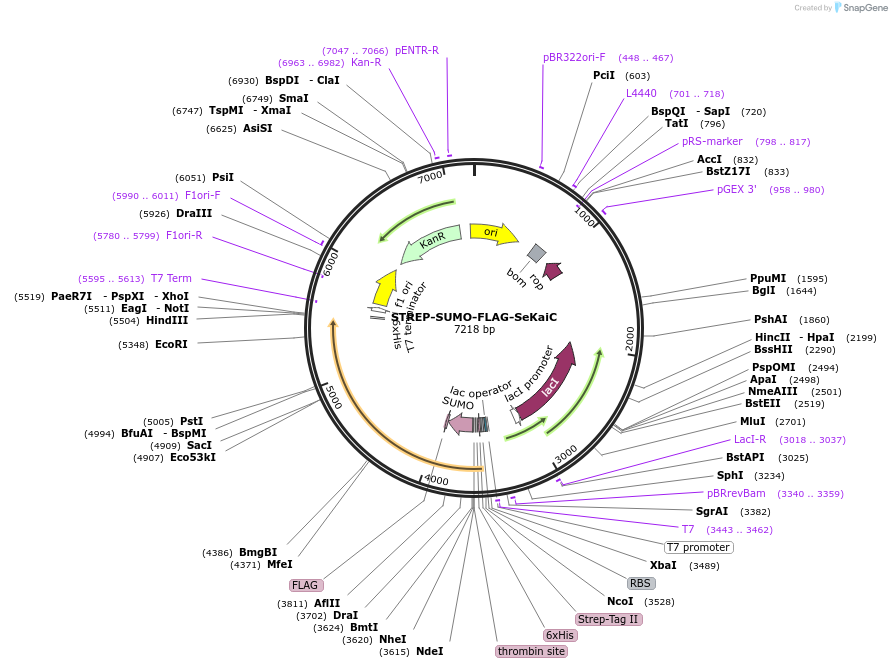
STREP-SUMO-FLAG-SeKaiC
Plasmid#218179PurposeWT SeKaiC for phosphorylation/dephosphorylation studyDepositorInsertSTREP-SUMO-FLAG-SeKaiC
TagsSTREP-TagExpressionBacterialAvailable SinceJune 3, 2024AvailabilityAcademic Institutions and Nonprofits only -
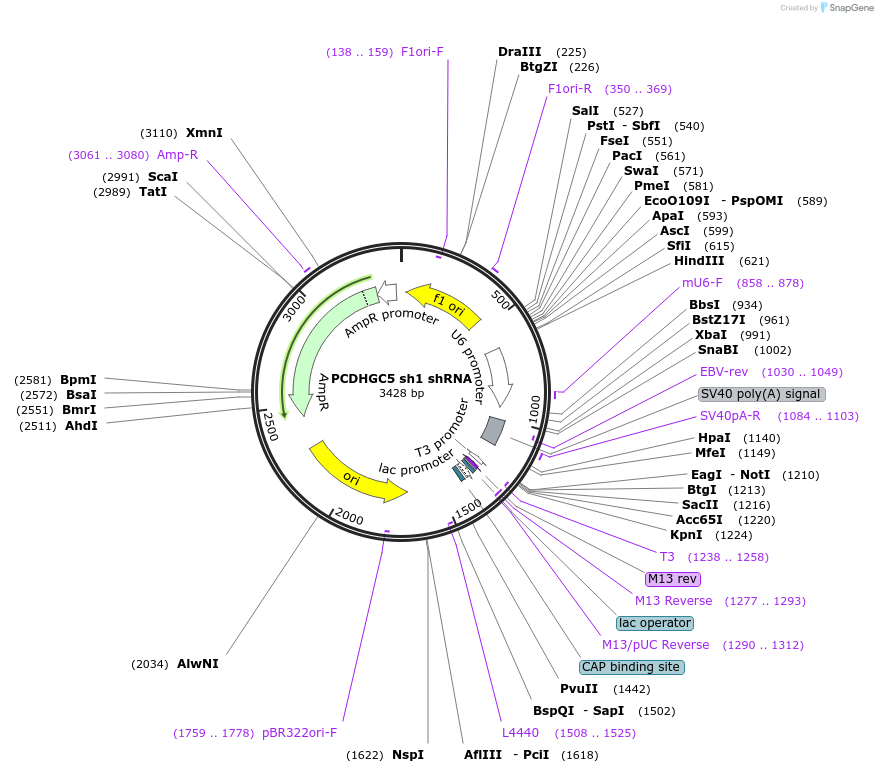
PCDHGC5 sh1 shRNA
Plasmid#122227PurposeKnock Down Protocadherin gamma C5. It targets Protocadherin gamma C5 mRNA (nucleotides 851–871 in the protein coding region).DepositorInsertPCDHGC5 shRNA (Pcdhgc5 Rat)
UseRNAiAvailable SinceMarch 8, 2019AvailabilityAcademic Institutions and Nonprofits only -
pHL1757
Plasmid#53037PurposePconNoHindM2:glgC::gfp, PconNoHind:RBS(st3):tetRDepositorInsertsGFP
tetR
UseSynthetic BiologyTagsRBS(st3) and glgC leaderExpressionBacterialPromoterPconNoHindM2 and pConNoHindAvailable SinceMay 14, 2014AvailabilityAcademic Institutions and Nonprofits only -
pHL1561
Plasmid#37571DepositorAvailable SinceAug. 14, 2012AvailabilityAcademic Institutions and Nonprofits only -
pHL1559
Plasmid#37570DepositorAvailable SinceAug. 14, 2012AvailabilityAcademic Institutions and Nonprofits only -
pHL1324
Plasmid#37638DepositorInsertsGFP
TetR
CsrA
UseSynthetic BiologyExpressionBacterialPromoterpCon and pConNoHindAvailable SinceJuly 27, 2012AvailabilityAcademic Institutions and Nonprofits only -
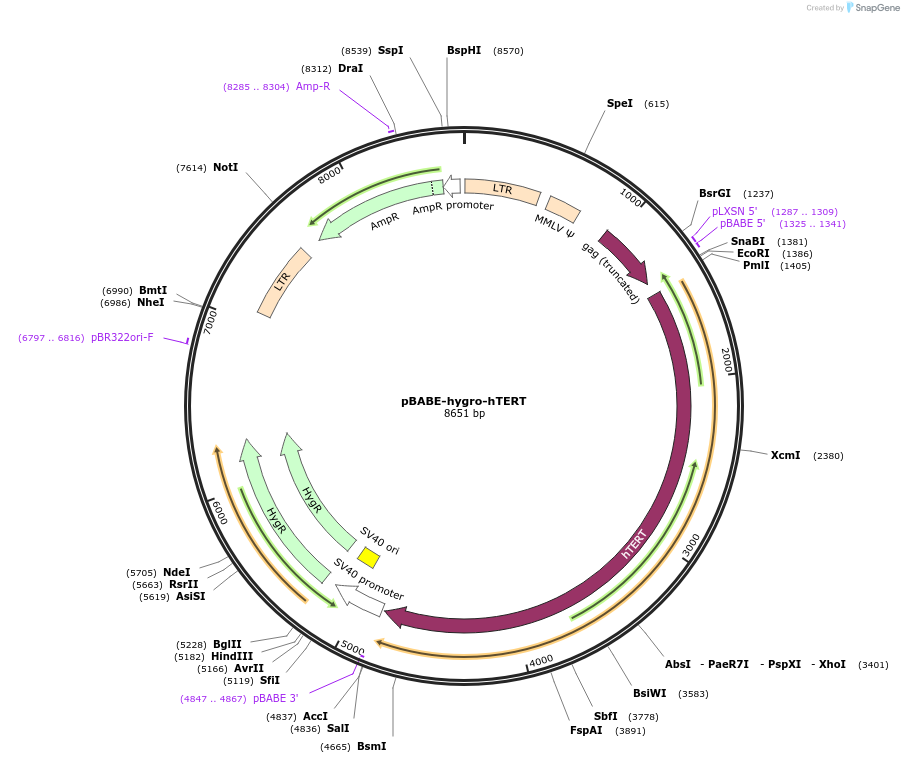
pBABE-hygro-hTERT
Plasmid#1773PurposeRetroviral expression of hTERT. Used to create immortalized cell lines.DepositorAvailable SinceJuly 18, 2006AvailabilityAcademic Institutions and Nonprofits only -
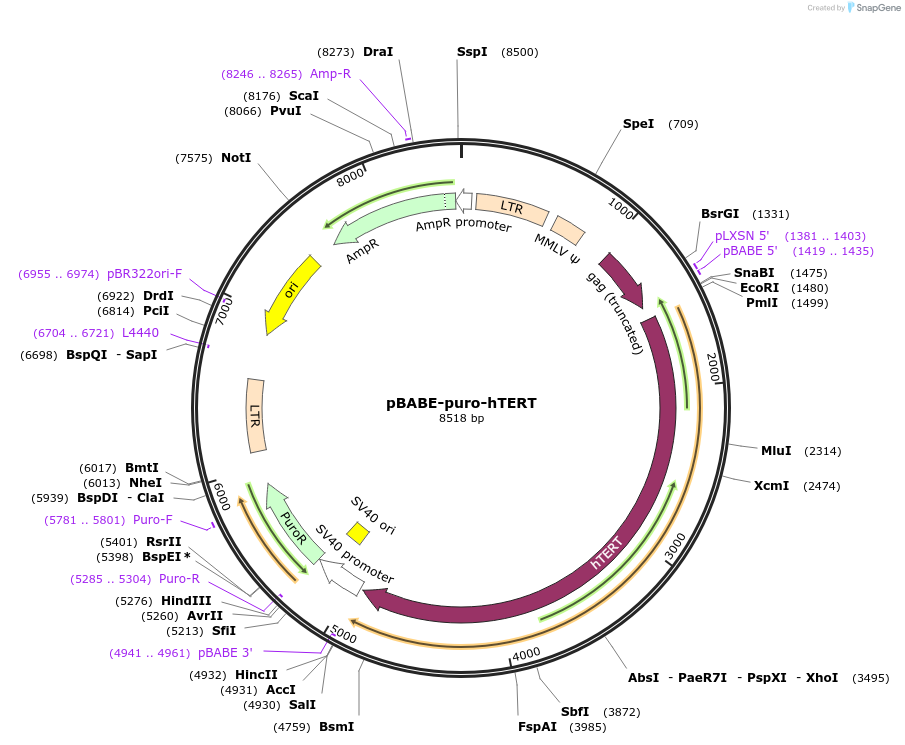
pBABE-puro-hTERT
Plasmid#1771PurposeRetroviral expression of hTERT. Used to create immortalized cell lines.DepositorAvailable SinceJune 3, 2005AvailabilityAcademic Institutions and Nonprofits only -
pGL4
Plasmid#48744PurposeExpression of luciferase driven by truncated mPer2 promoter region (bases -1128 to +1536 with respect to transcription start site at +1)DepositorInserttruncated mPer2 promoter/enhancer region (-1128 to +1536)
UseLuciferaseMutationtruncated promoter region (see pGL6)Available SinceJan. 31, 2014AvailabilityAcademic Institutions and Nonprofits only -
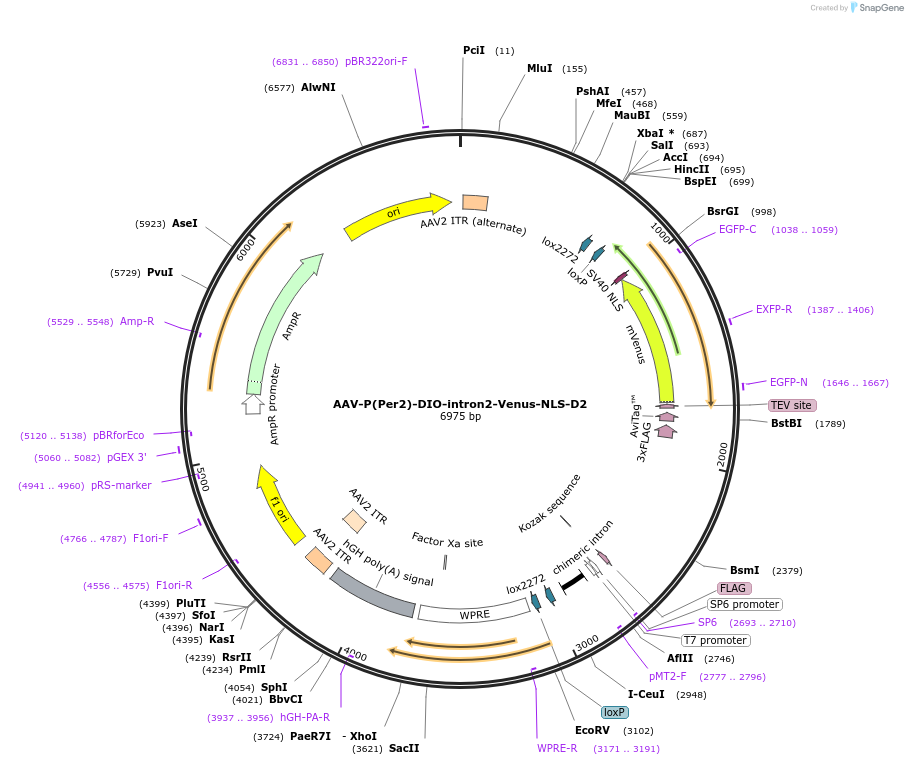
AAV-P(Per2)-DIO-intron2-Venus-NLS-D2
Plasmid#110057PurposePer2 transcription fluorescence reporterDepositorInsertPer2 promoter and Venus (Per2 Mouse)
UseAAV and Cre/LoxTagsNLS-D2ExpressionMammalianPromoterPer2Available SinceMay 30, 2018AvailabilityAcademic Institutions and Nonprofits only -
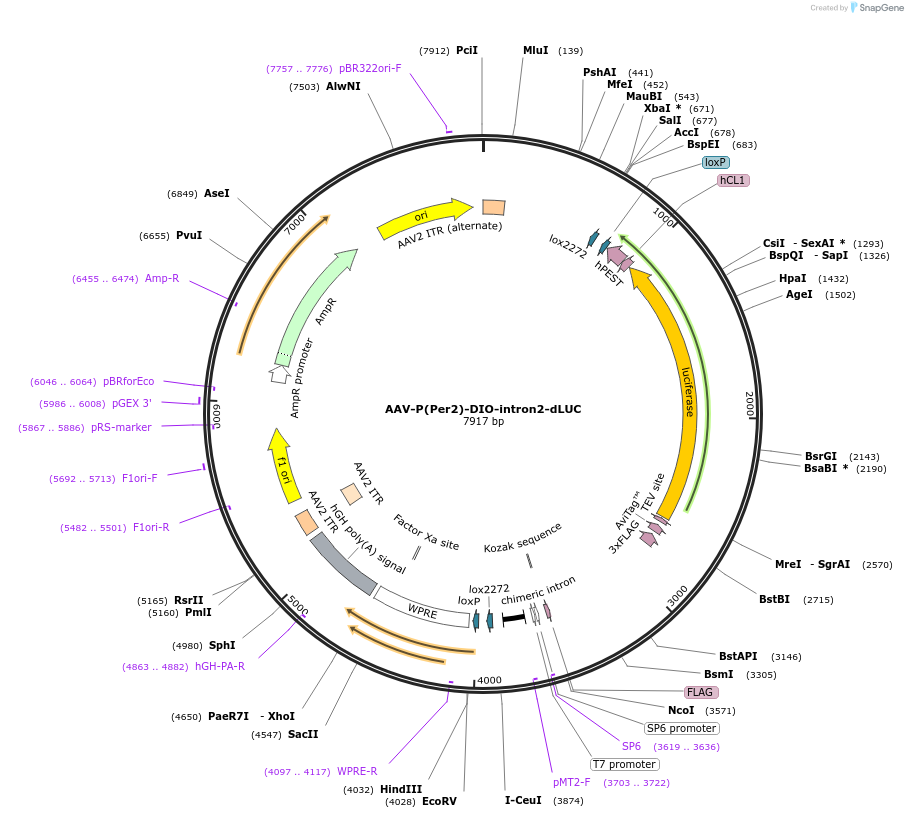
AAV-P(Per2)-DIO-intron2-dLUC
Plasmid#110059PurposePer2 transcription luciferase reporterDepositorInsertPer2 promoter and luciferase (Per2 Mouse)
UseAAV and Cre/LoxExpressionMammalianPromoterPer2Available SinceMay 30, 2018AvailabilityAcademic Institutions and Nonprofits only -
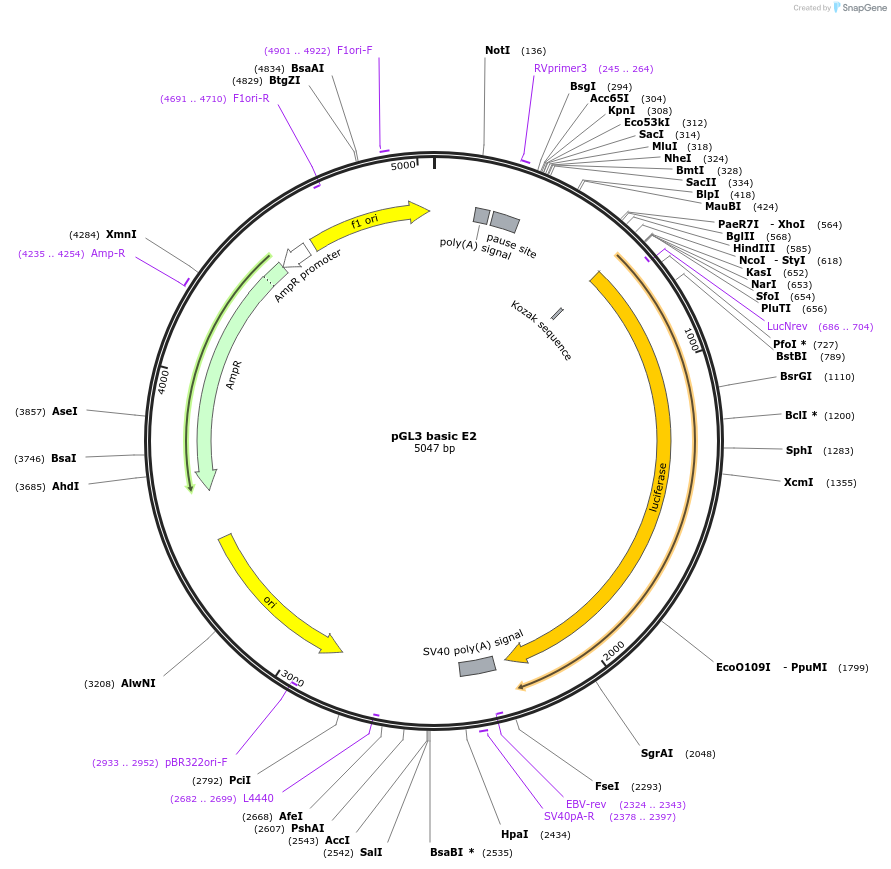
pGL3 basic E2
Plasmid#48747PurposeExpression of luciferase driven by mPer2 promoter fragment containing E-box2 (bases -112 to +98 with respect to transcription start site at +1)DepositorInsertmPer2 promoter/enhancer region containing E-box2 (-112 to +98)
UseLuciferaseAvailable SinceDec. 5, 2013AvailabilityAcademic Institutions and Nonprofits only -
pGL3
Plasmid#48743PurposeExpression of luciferase driven by truncated mPer2 promoter region (bases -1128 to +1060 with respect to transcription start site at +1)DepositorInserttruncated mPer2 promoter/enhancer region (-1128 to +1060)
UseLuciferaseMutationtruncated promoter region (see pGL6)Available SinceDec. 19, 2013AvailabilityAcademic Institutions and Nonprofits only -
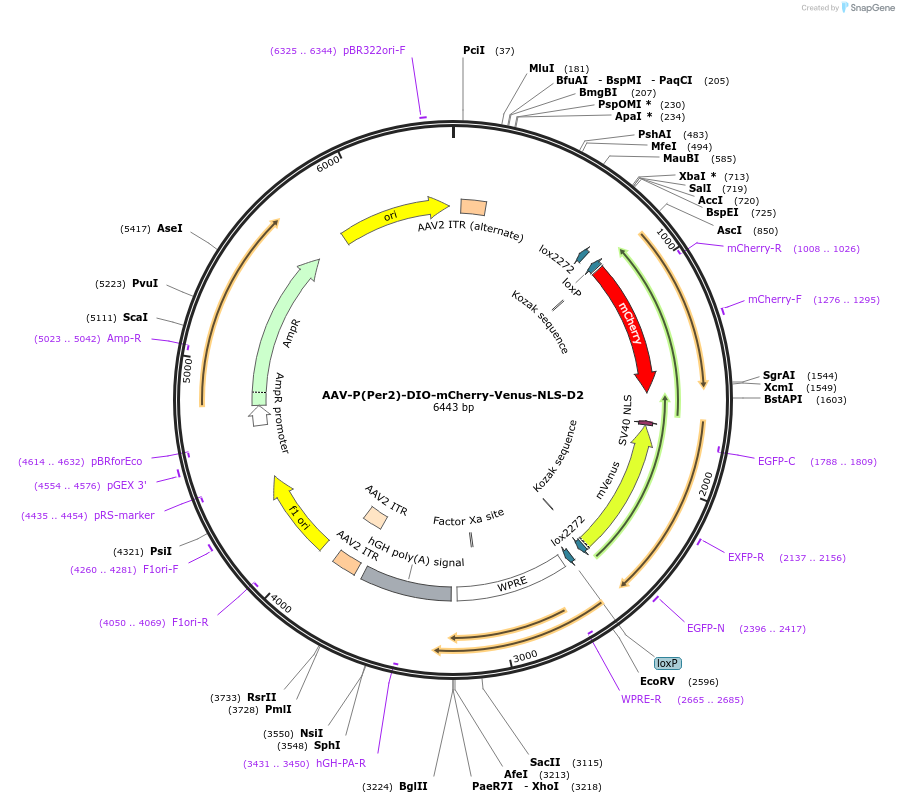
AAV-P(Per2)-DIO-mCherry-Venus-NLS-D2
Plasmid#110058PurposePer2 transcription fluorescence reporterDepositorInsertPer2 promoter and mCherry and Venus (Per2 Mouse)
UseAAV and Cre/LoxTagsNLS-D2ExpressionMammalianPromoterPer2Available SinceMay 30, 2018AvailabilityAcademic Institutions and Nonprofits only -
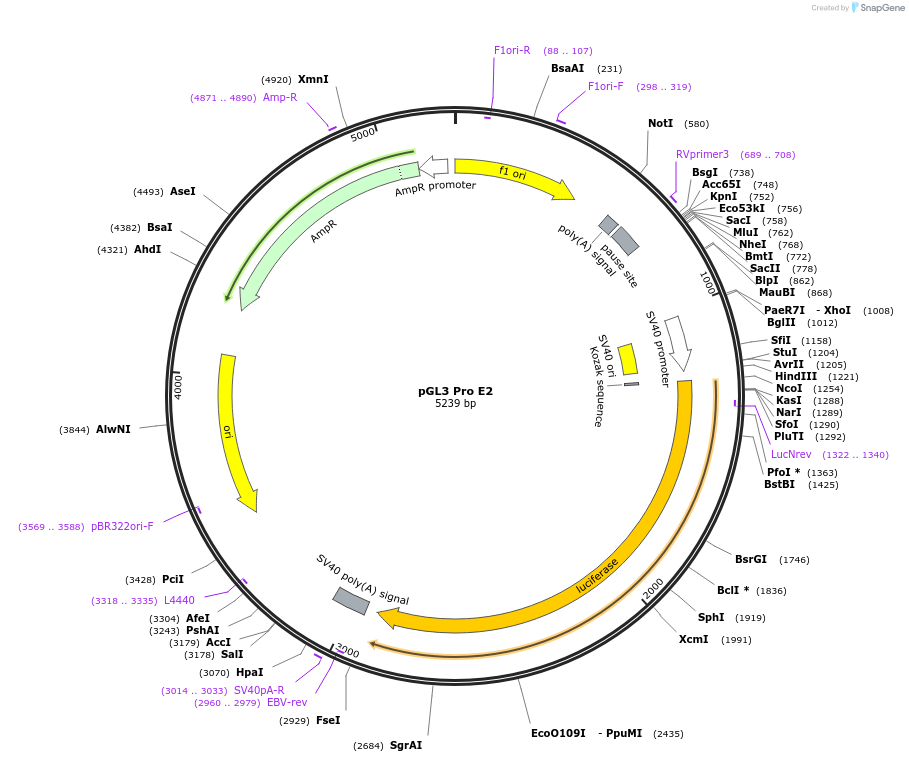
pGL3 Pro E2
Plasmid#48749PurposeExpression of luciferase driven by mPer2 promoter fragment containing E-box2 (bases -112 to +98 with respect to transcription start site at +1) and SV40 promoter (backbone)DepositorInsertmPer2 promoter/enhancer region containing E-box2 (-112 to +98)
UseLuciferaseAvailable SinceDec. 5, 2013AvailabilityAcademic Institutions and Nonprofits only -
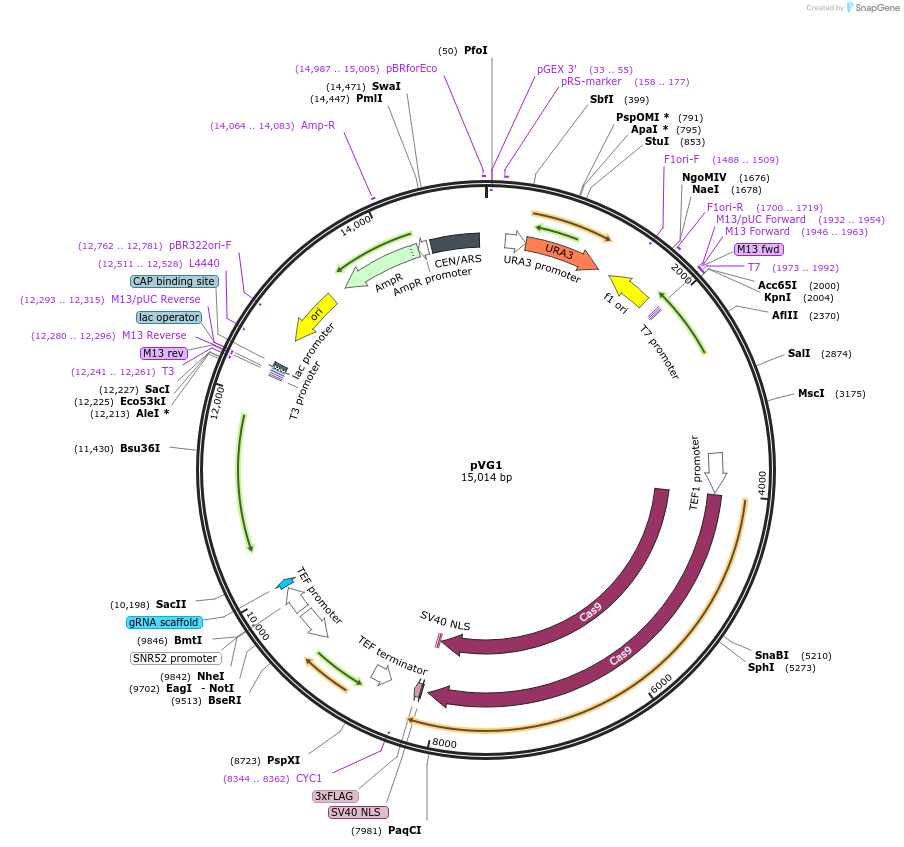
pVG1
Plasmid#111444PurposeUnified Solo vector pV1382 + sgScADE2 + ScADE2 stop codon repair tempateDepositorInsertCaCas9/sgScADE2/stop-codon repair
UseCRISPRExpressionYeastAvailable SinceDec. 7, 2018AvailabilityAcademic Institutions and Nonprofits only -
pGL2
Plasmid#48742PurposeExpression of luciferase driven by truncated mPer2 promoter region (bases -1128 to +386 with respect to transcription start site at +1)DepositorInserttruncated mPer2 promoter/enhancer region (-1128 to +386)
UseLuciferaseMutationtruncated promoter region (see pGL6)Available SinceDec. 19, 2013AvailabilityAcademic Institutions and Nonprofits only