We narrowed to 7,225 results for: CAD;
-
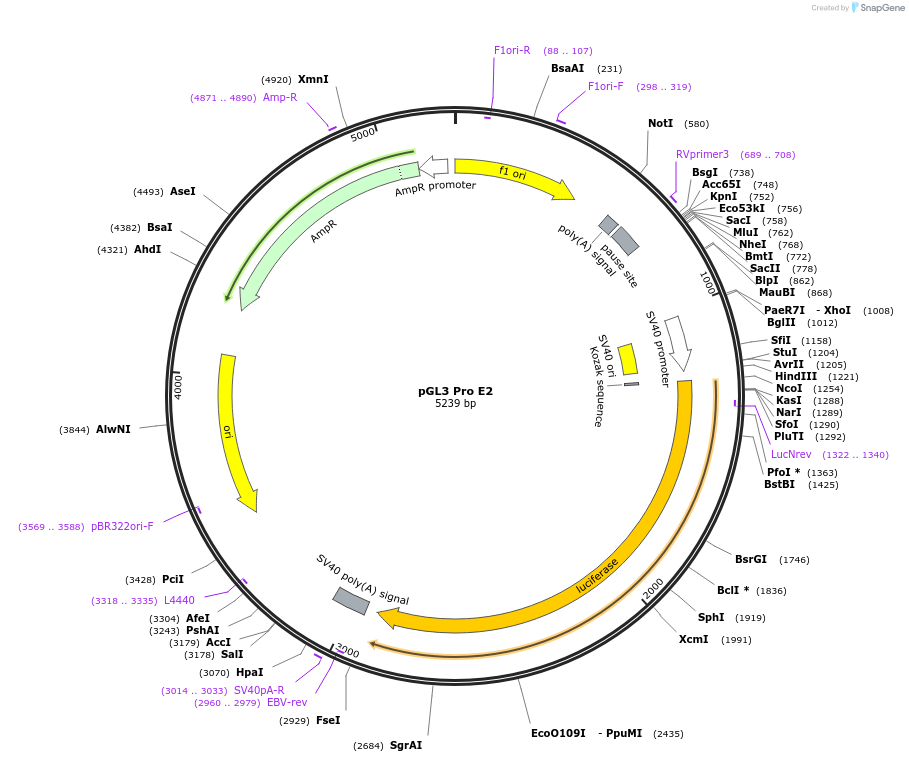
Plasmid#48749PurposeExpression of luciferase driven by mPer2 promoter fragment containing E-box2 (bases -112 to +98 with respect to transcription start site at +1) and SV40 promoter (backbone)DepositorInsertmPer2 promoter/enhancer region containing E-box2 (-112 to +98)
UseLuciferaseAvailable SinceDec. 5, 2013AvailabilityAcademic Institutions and Nonprofits only -
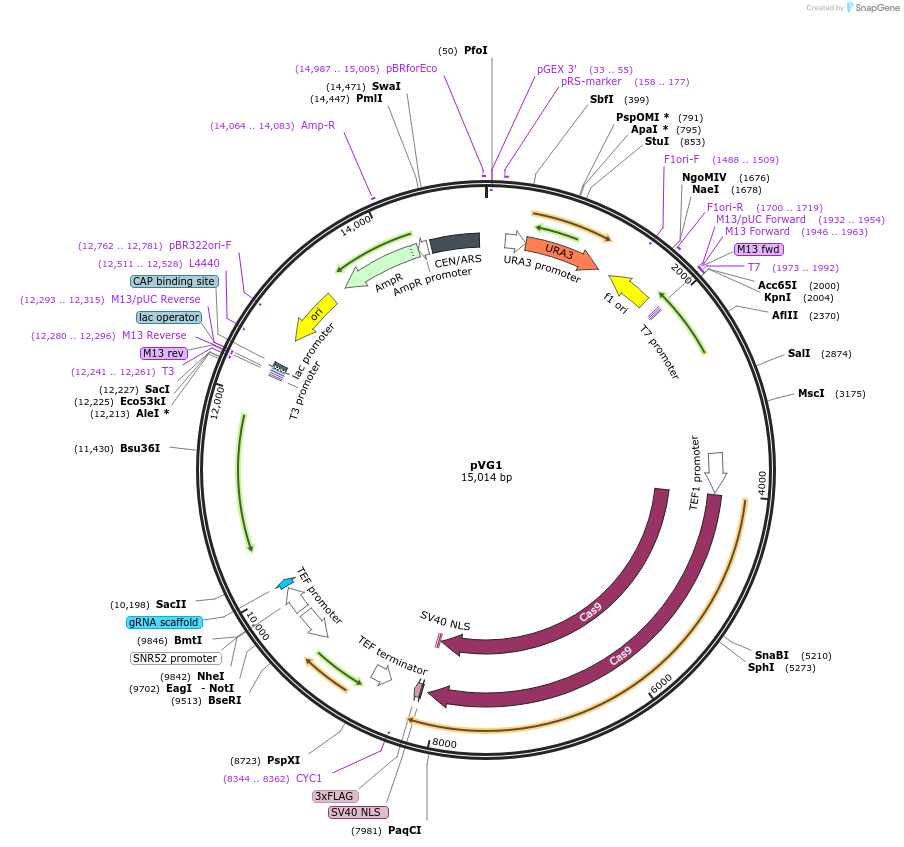
pVG1
Plasmid#111444PurposeUnified Solo vector pV1382 + sgScADE2 + ScADE2 stop codon repair tempateDepositorInsertCaCas9/sgScADE2/stop-codon repair
UseCRISPRExpressionYeastAvailable SinceDec. 7, 2018AvailabilityAcademic Institutions and Nonprofits only -
pGL2
Plasmid#48742PurposeExpression of luciferase driven by truncated mPer2 promoter region (bases -1128 to +386 with respect to transcription start site at +1)DepositorInserttruncated mPer2 promoter/enhancer region (-1128 to +386)
UseLuciferaseMutationtruncated promoter region (see pGL6)Available SinceDec. 19, 2013AvailabilityAcademic Institutions and Nonprofits only -
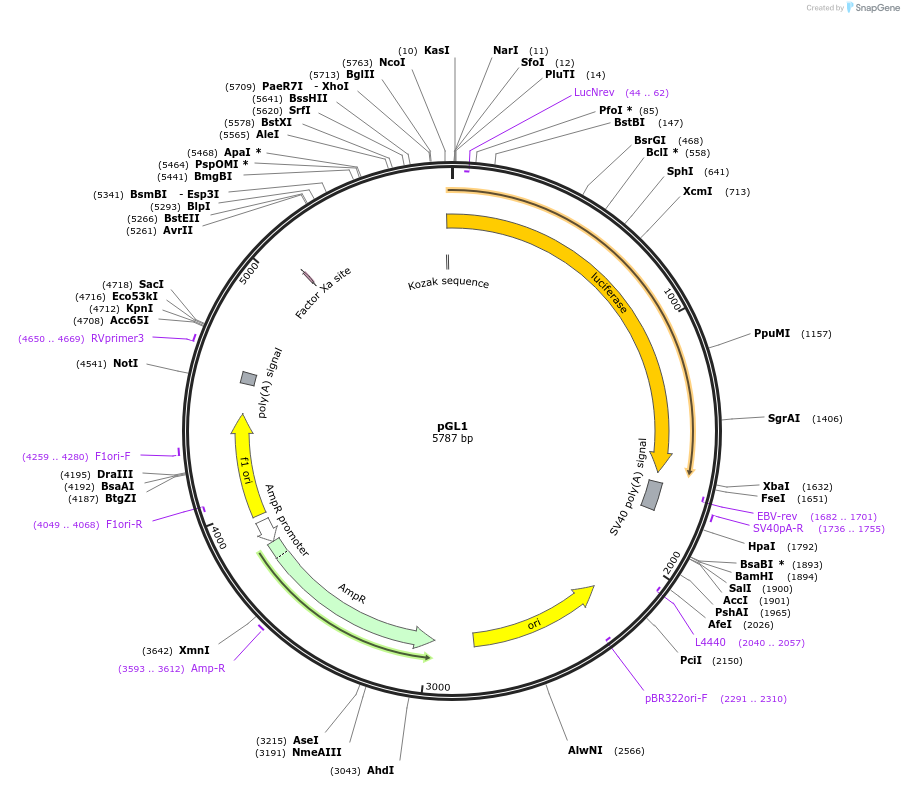
pGL1
Plasmid#48741PurposeExpression of luciferase driven by truncated mPer2 promoter region (bases -1128 to -141 with respect to transcription start site at +1)DepositorInserttruncated mPer2 promoter/enhancer region (-1128 to -141)
UseLuciferaseMutationtruncated promoter region (see pGL6)Available SinceDec. 5, 2013AvailabilityAcademic Institutions and Nonprofits only -
pGL5
Plasmid#48745PurposeExpression of luciferase driven by truncated mPer2 promoter region (bases -1128 to +2064 with respect to transcription start site at +1)DepositorInserttruncated mPer2 promoter/enhancer region (-1128 to +2064)
UseLuciferaseMutationtruncated promoter region (see pGL6)Available SinceDec. 19, 2013AvailabilityAcademic Institutions and Nonprofits only -
pHL1855
Plasmid#53041PurposePLlacO-1:glgC::gfp::LVA, PconNoHind:RBS(st3):tetR, PconNoHind:RBS(st3):csrADepositorInsertsGFP
tetR
csrA
UseSynthetic BiologyTagsLVA, RBS(st3), and glgC leaderExpressionBacterialPromoterpConNoHind and pLlac0-1Available SinceMay 16, 2014AvailabilityAcademic Institutions and Nonprofits only -
pHL600
Plasmid#37563DepositorUseSynthetic BiologyExpressionBacterialPromoterpLlacO-1 and pLtetO-1Available SinceAug. 14, 2012AvailabilityAcademic Institutions and Nonprofits only -
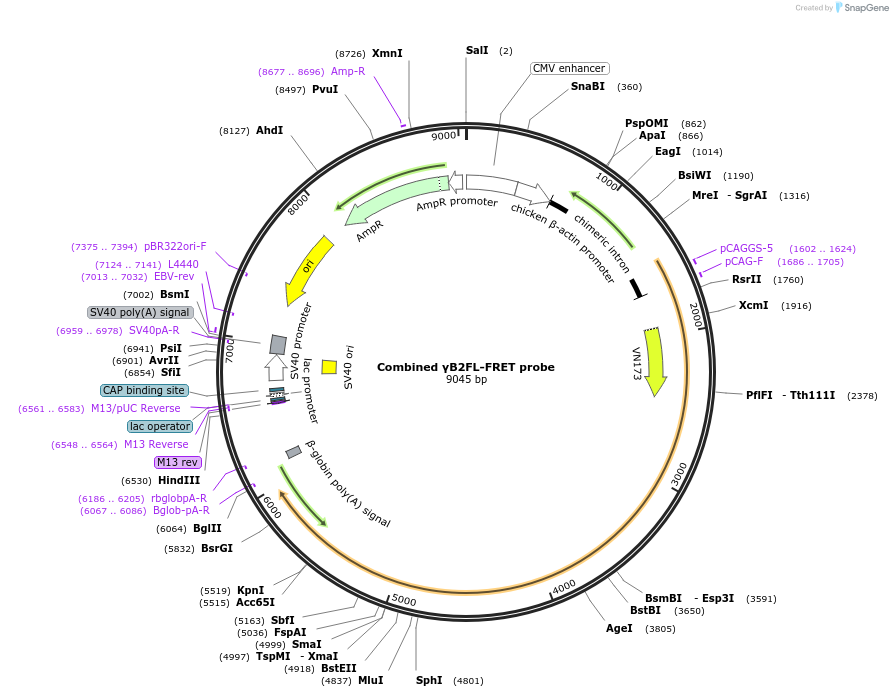
Combined γB2FL-FRET probe
Plasmid#209094PurposemTurquoise2- and mVenus-inserted PcdhgB2 full length (FL)DepositorAvailable SinceDec. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
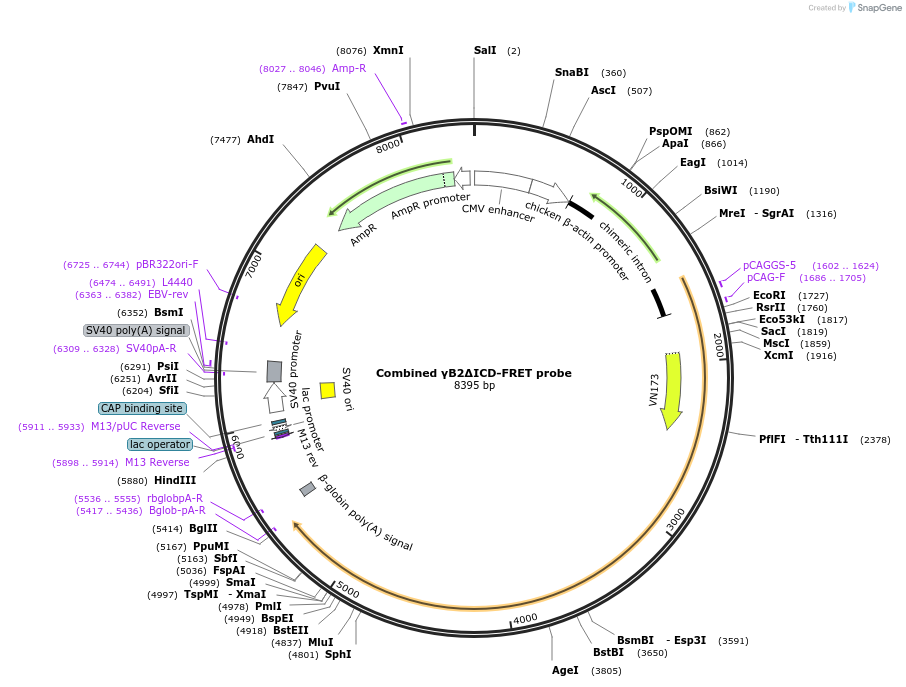
Combined γB2ΔICD-FRET probe
Plasmid#209093PurposemTurquoise2- and mVenus-inserted PcdhgB2 lacking an intracellular domain (ICD)DepositorAvailable SinceDec. 21, 2023AvailabilityAcademic Institutions and Nonprofits only -
pHL1853
Plasmid#53040PurposePLlacO-1:glgC::gfp::LVA, PconNoHind:RBS(st3):tetR, PconNoHind:RBS(st2):csrADepositorInsertsGFP
tetR
csrA
UseSynthetic BiologyTagsLVA, RBS(st2), RBS(st3), and glgC leaderExpressionBacterialPromoterpConNoHind and pLlac0-1Available SinceAug. 21, 2014AvailabilityAcademic Institutions and Nonprofits only -
pHL1670
Plasmid#37634DepositorInsertsUseSynthetic BiologyTagsASV degradation tag and glgC leader regionExpressionBacterialPromoterpConM12NoHind and pConNoHindAvailable SinceAug. 14, 2012AvailabilityAcademic Institutions and Nonprofits only -
pHL1668
Plasmid#37633DepositorInsertsUseSynthetic BiologyTagsASV degradation tag and glgC leader regionExpressionBacterialPromoterpConM12NoHind and pConNoHindAvailable SinceAug. 10, 2012AvailabilityAcademic Institutions and Nonprofits only -
pHL1530
Plasmid#37569DepositorInsertsGFP
TetR
CsrA
UseSynthetic BiologyTagsglgC leader regionExpressionBacterialMutationstart codon removedPromoterpConNoHind and pLlacO-1Available SinceAug. 10, 2012AvailabilityAcademic Institutions and Nonprofits only -
pHL601
Plasmid#37564DepositorUseSynthetic BiologyExpressionBacterialPromoterpLlacO-1 and pLtetO-1Available SinceAug. 10, 2012AvailabilityAcademic Institutions and Nonprofits only -
pHL1355
Plasmid#37566DepositorInsertsGFP
TetR
CsrA
UseSynthetic BiologyTagsglgC leader regionExpressionBacterialMutationstart codon removedPromoterpConNoHind and pConNoHindM12Available SinceJuly 26, 2012AvailabilityAcademic Institutions and Nonprofits only -
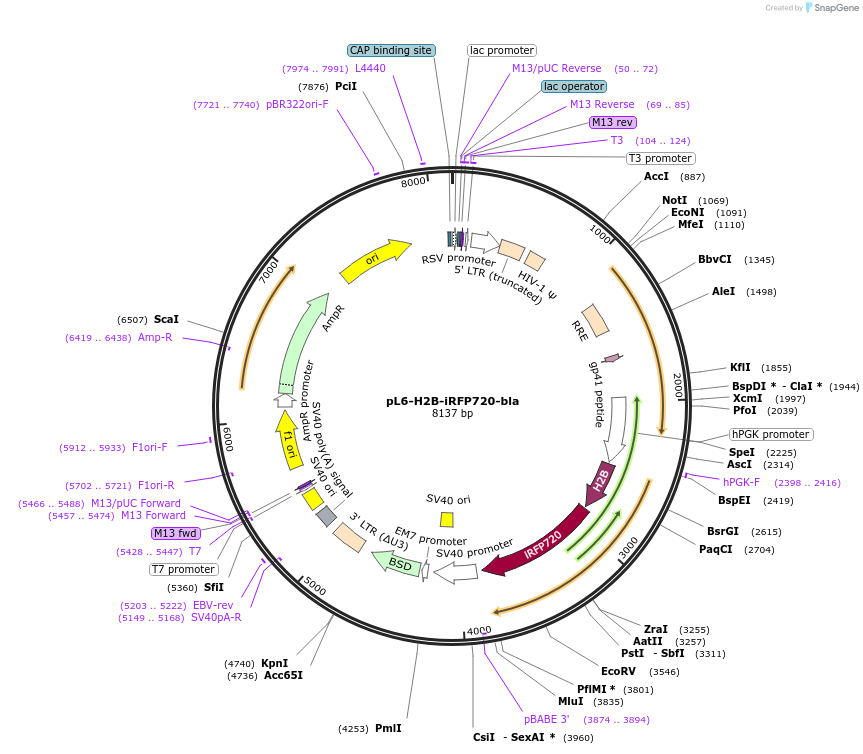
pL6-H2B-iRFP720-bla
Plasmid#189991PurposeLentivirial plasmid for nuclear iRFP-720 expression with blasticidin resistanceDepositorInsertHuman histone H2B
UseLentiviralTagsiRFP720ExpressionMammalianAvailable SinceFeb. 9, 2024AvailabilityAcademic Institutions and Nonprofits only -
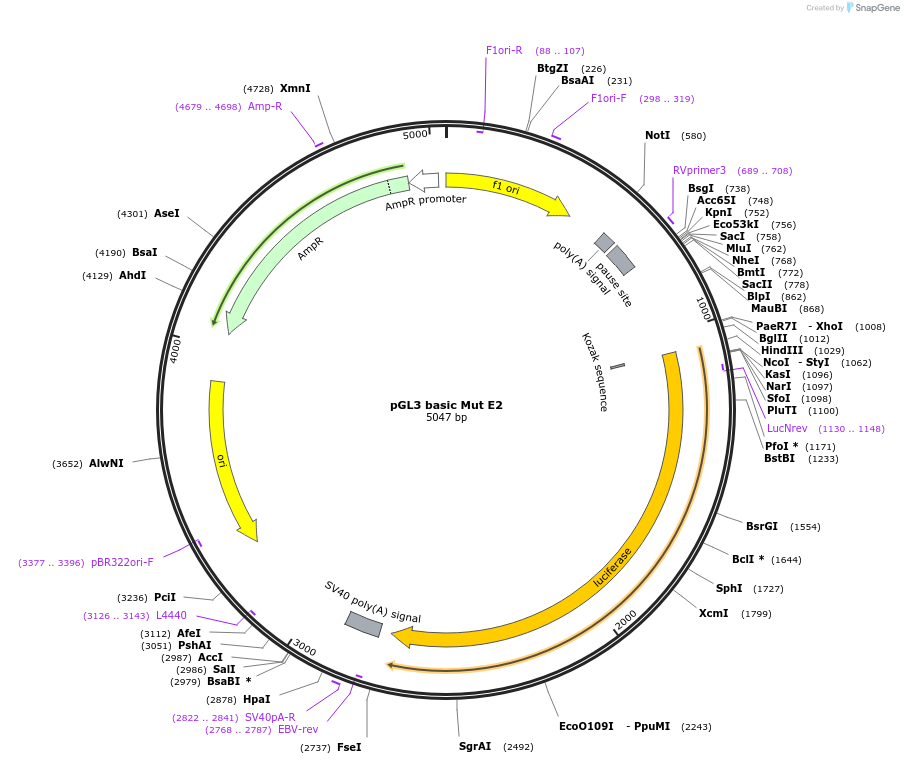
pGL3 basic Mut E2
Plasmid#48748PurposeExpression of luciferase driven by mPer2 promoter fragment containing mutated E-box2 (bases -112 to +98 with respect to transcription start site at +1)DepositorInsertmPer2 promoter/enhancer region containing mutated E-box2 (-112 to +98)
UseLuciferaseMutationMutated E-box2 sequence from CACGTT to GCTAGTAvailable SinceDec. 5, 2013AvailabilityAcademic Institutions and Nonprofits only -
lentiviral shRNA GEF-H1
Plasmid#21477DepositorInsertrho/rac guanine nucleotide exchange factor (GEF) 2 (Arhgef2 Rat)
UseLentiviralTagseGFPExpressionMammalianAvailable SinceJuly 22, 2009AvailabilityAcademic Institutions and Nonprofits only -
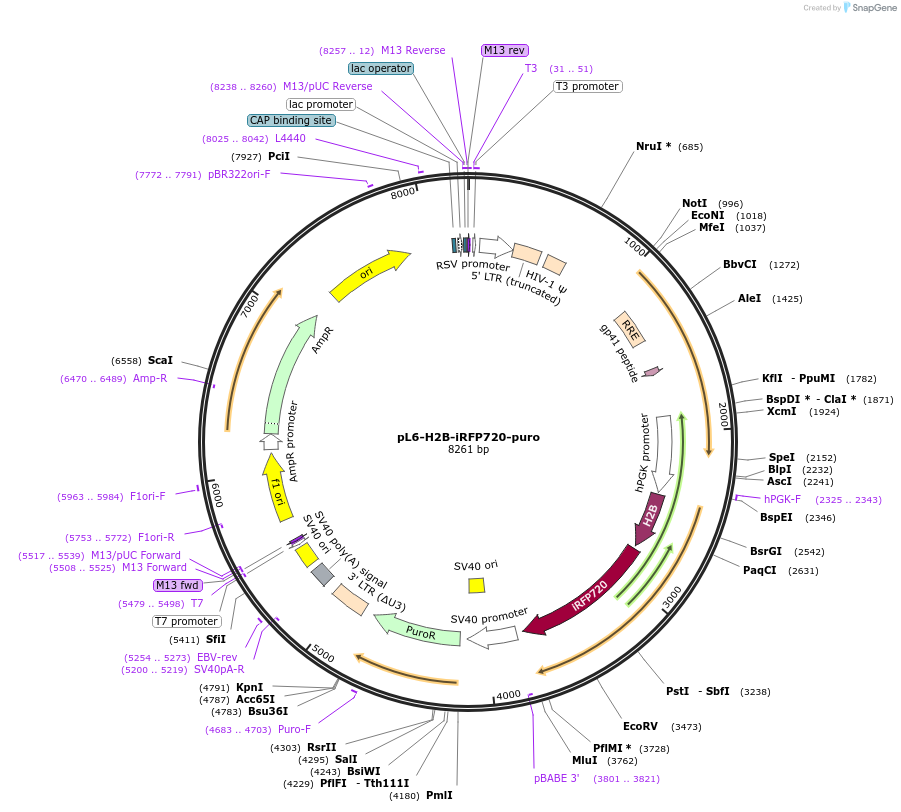
pL6-H2B-iRFP720-puro
Plasmid#189992PurposeLentivirial plasmid for nuclear iRFP-720 expression with puromycin resistanceDepositorInsertHuman histone H2B
UseLentiviralTagsiRFP720ExpressionMammalianAvailable SinceFeb. 9, 2024AvailabilityAcademic Institutions and Nonprofits only -
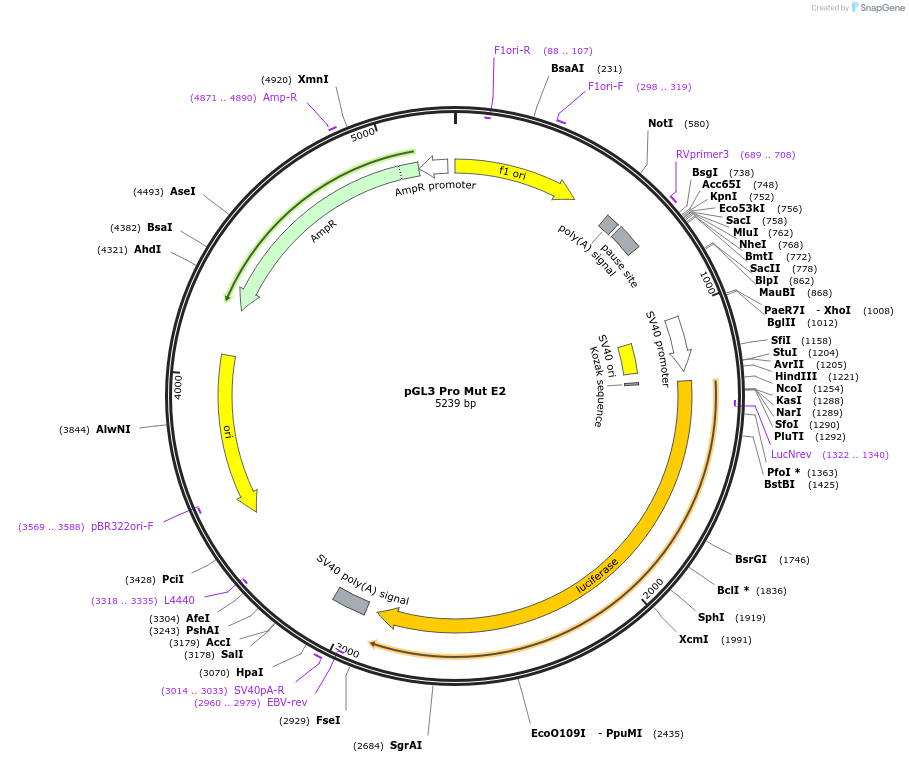
pGL3 Pro Mut E2
Plasmid#48750PurposeExpression of luciferase driven by mPer2 promoter fragment containing mutated E-box2 (bases -112 to +98 with respect to transcription start site at +1) and SV40 promoter (backbone)DepositorInsertmPer2 promoter/enhancer region containing mutated E-box2 (-112 to +98)
UseLuciferaseMutationMutated E-box2 sequence from CACGTT to GCTAGTAvailable SinceDec. 5, 2013AvailabilityAcademic Institutions and Nonprofits only