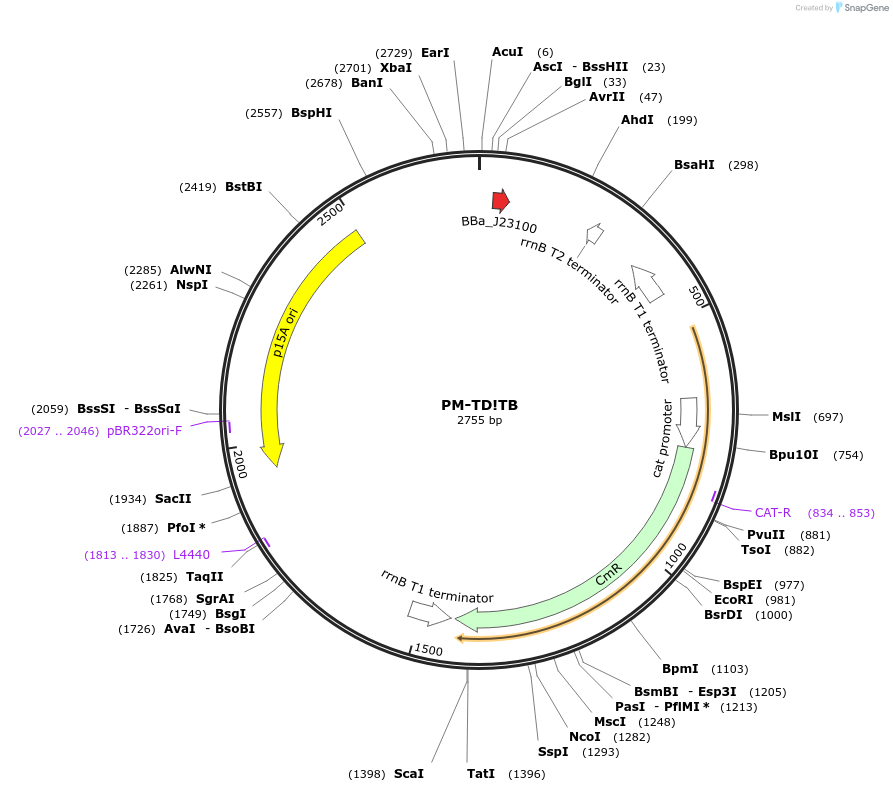
We narrowed to 11,701 results for: phen
-
Plasmid#48656PurposeBacterial TD crRNA expression: targets TD to protospacer B (ACTTTAAAAGTATTCGCCAT), p15A/chloramphenicolDepositorInsertBacterial TD crRNA to prototspacer B
UseCRISPRPromoterJ23100 promoterAvailable SinceOct. 17, 2013AvailabilityAcademic Institutions and Nonprofits only -
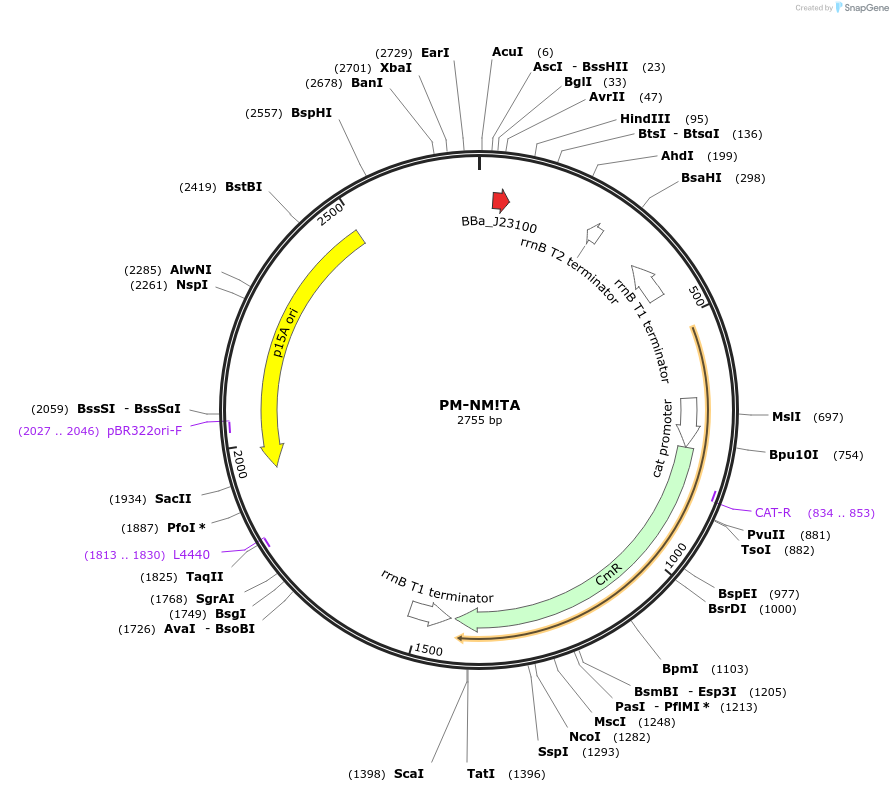
PM-NM!TA
Plasmid#48651PurposeBacterial NM crRNA expression: targets NM to protospacer A (TACCATCTCAAGCTTGTTGA), p15A/chloramphenicolDepositorInsertBacterial NM crRNA to prototspacer A
UseCRISPRPromoterJ23100 promoterAvailable SinceOct. 17, 2013AvailabilityAcademic Institutions and Nonprofits only -
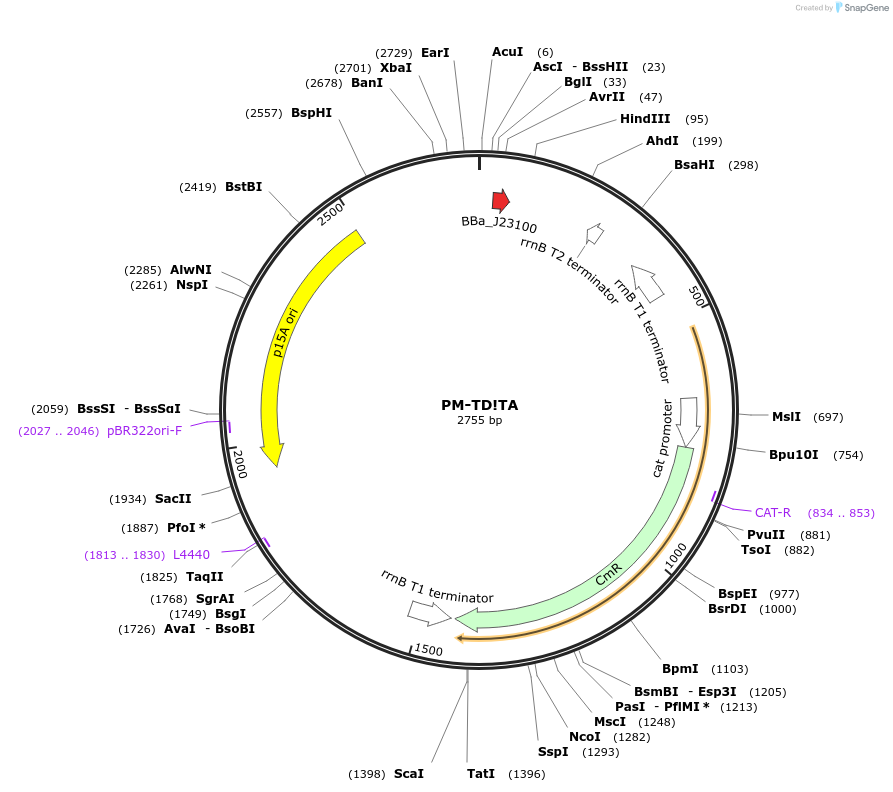
PM-TD!TA
Plasmid#48655PurposeBacterial TD crRNA expression: targets TD to protospacer A (TACCATCTCAAGCTTGTTGA), p15A/chloramphenicolDepositorInsertBacterial TD crRNA to prototspacer A
UseCRISPRPromoterJ23100 promoterAvailable SinceOct. 17, 2013AvailabilityAcademic Institutions and Nonprofits only -
G1S-HA-pCB6
Plasmid#74373PurposeMammalian expression of a G1S mutant HA from the X:31 strain of influenza virusDepositorInsertHA
ExpressionMammalianMutationG1SPromoterCMVAvailable SinceApril 13, 2016AvailabilityAcademic Institutions and Nonprofits only -
G1V-HA-pCB6
Plasmid#74372PurposeMammalian expression of a G1V mutant HA from the X:31 strain of influenza virusDepositorInsertHA
ExpressionMammalianMutationG1VPromoterCMVAvailable SinceApril 13, 2016AvailabilityAcademic Institutions and Nonprofits only -
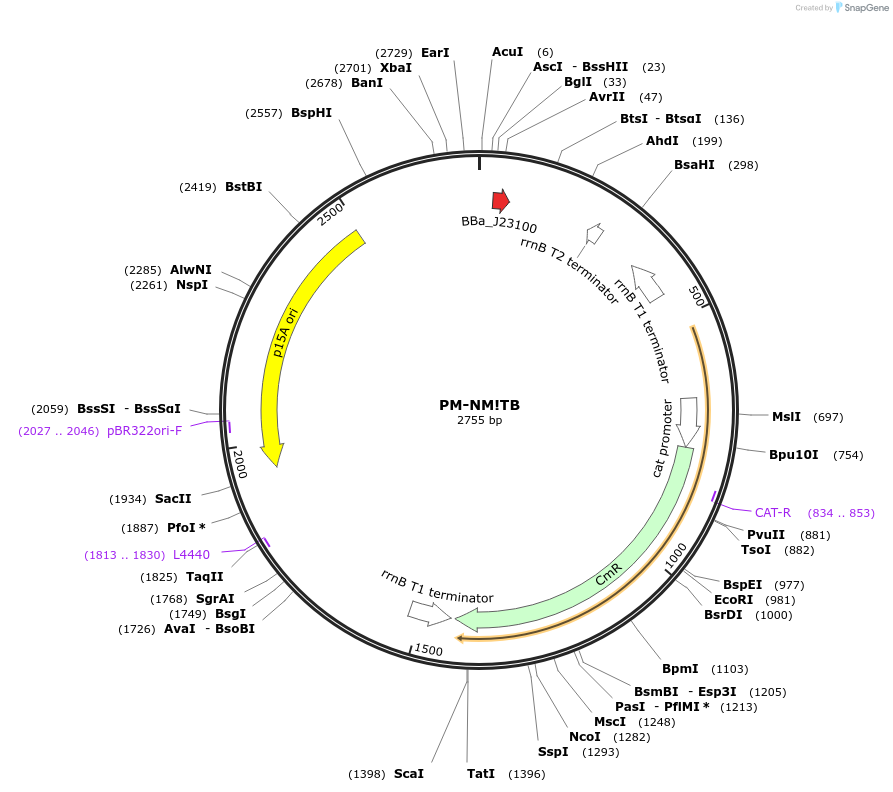
PM-NM!TB
Plasmid#48652PurposeBacterial NM crRNA expression: targets NM to protospacer B (ACTTTAAAAGTATTCGCCAT), p15A/chloramphenicolDepositorInsertBacterial NM crRNA to prototspacer B
UseCRISPRPromoterJ23100 promoterAvailable SinceOct. 17, 2013AvailabilityAcademic Institutions and Nonprofits only -
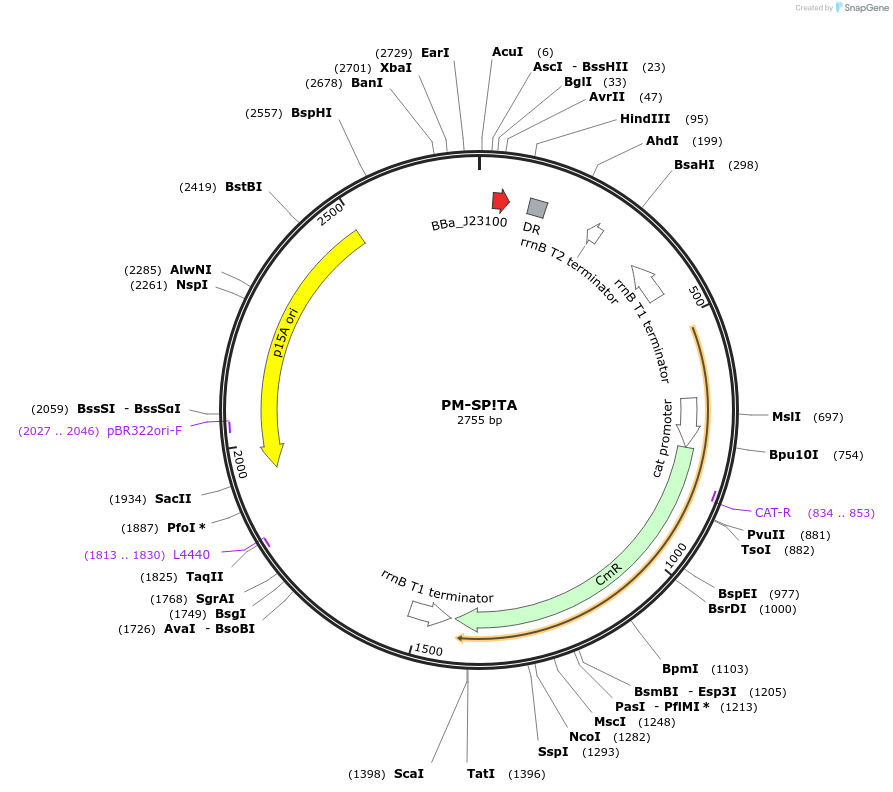
PM-SP!TA
Plasmid#48649PurposeBacterial SP crRNA expression: targets SP to protospacer A (TACCATCTCAAGCTTGTTGA), p15A/chloramphenicolDepositorInsertBacterial SP crRNA to prototspacer A
UseCRISPRPromoterJ23100 promoterAvailable SinceOct. 17, 2013AvailabilityAcademic Institutions and Nonprofits only -
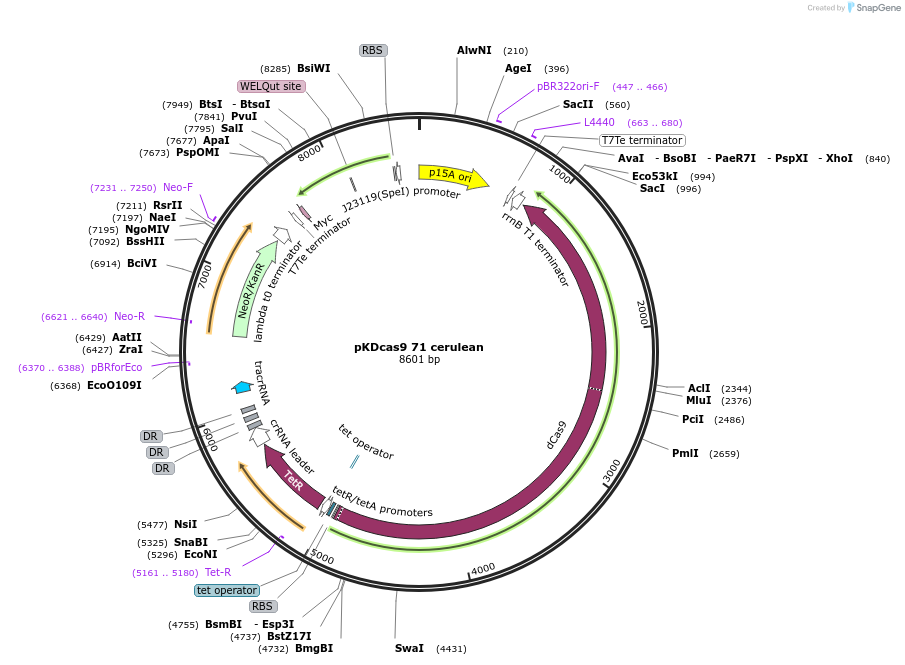
pKDcas9 71 cerulean
Plasmid#230954PurposeaTc inducible knockdown of FliZ and CRPDepositorInsertdCas9
UseCRISPRMutationadditional tetOAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
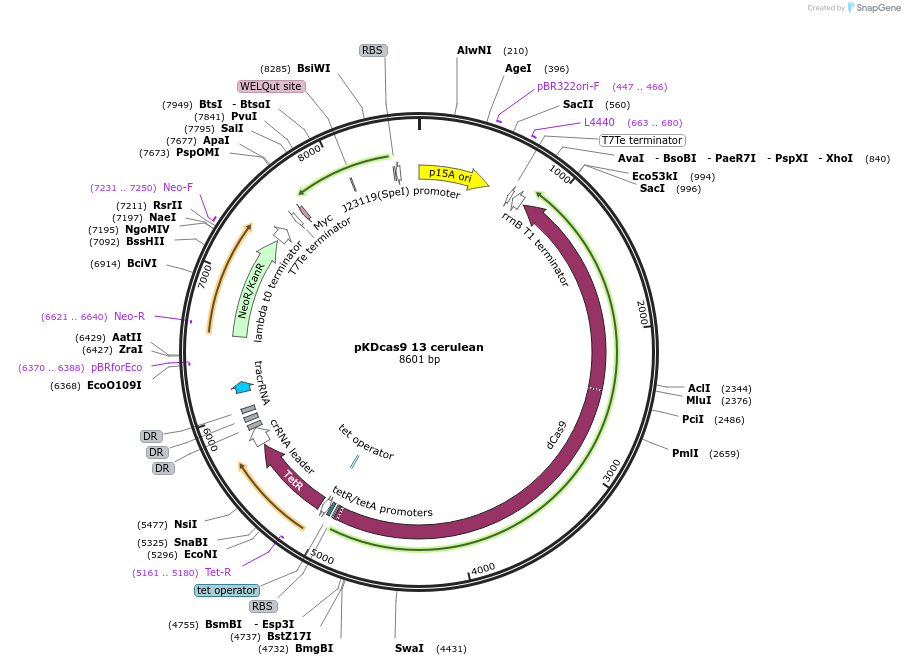
pKDcas9 13 cerulean
Plasmid#230955PurposeaTc inducible knockdown of CRP and HNSDepositorInsertdCas9
UseCRISPRMutationadditional tetOAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
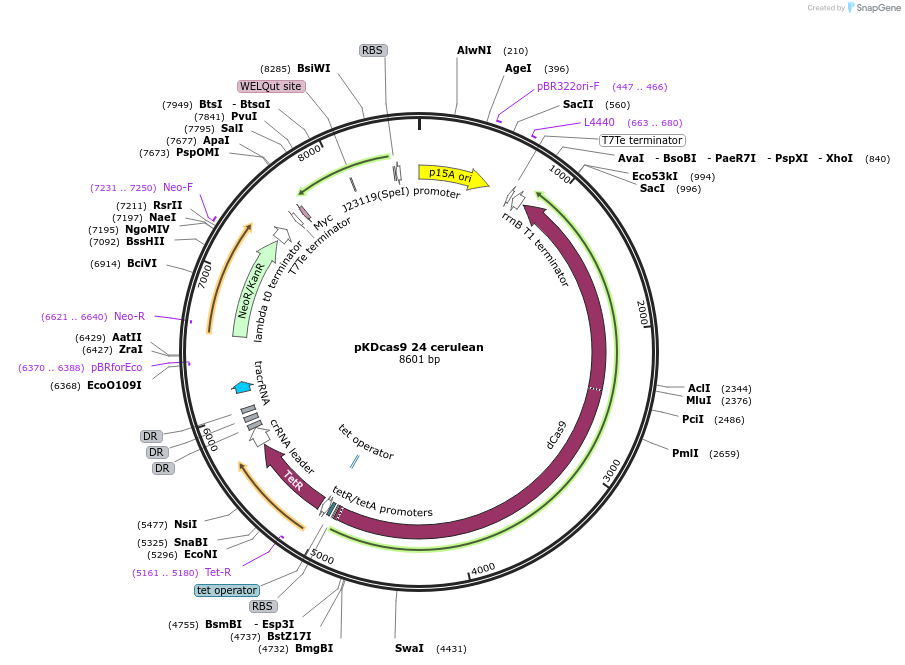
pKDcas9 24 cerulean
Plasmid#230956PurposeaTc inducible knockdown of Fis and mcaSDepositorInsertdCas9
UseCRISPRMutationadditional tetOAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
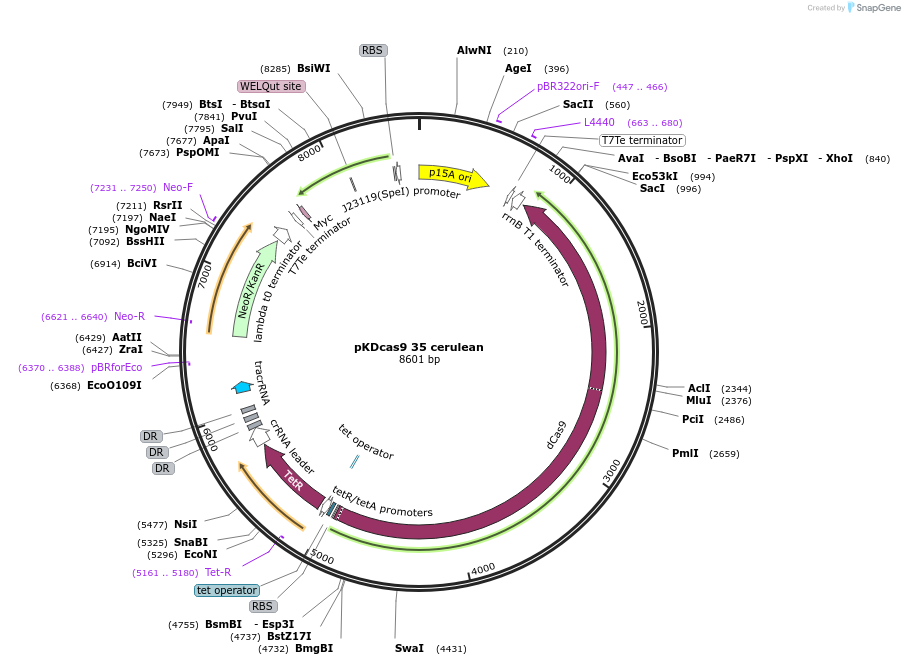
pKDcas9 35 cerulean
Plasmid#230957PurposeaTc inducible knockdown of HNS and CsgDDepositorInsertdCas9
UseCRISPRMutationadditional tetOAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
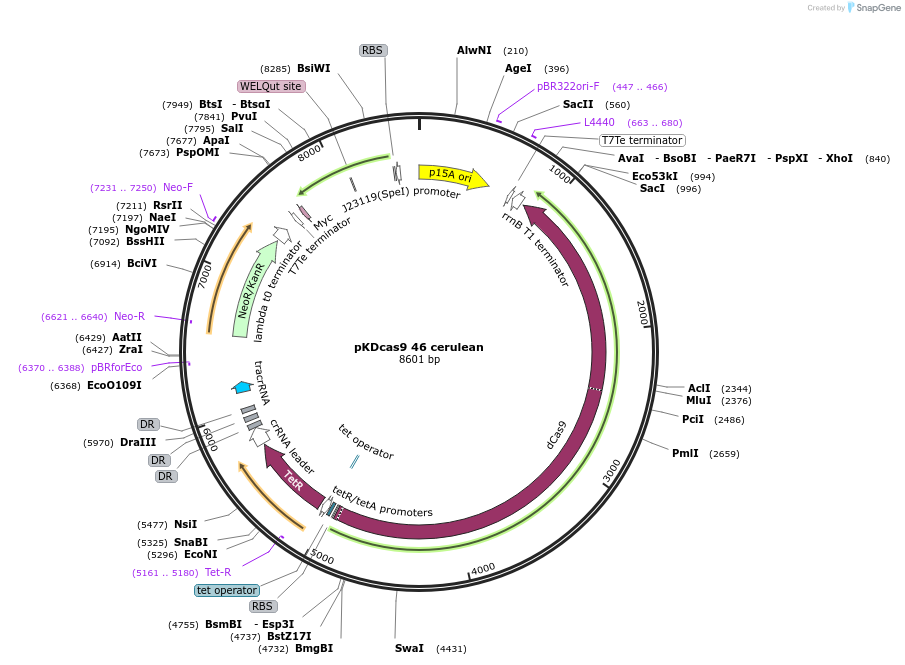
pKDcas9 46 cerulean
Plasmid#230958PurposeaTc inducible knockdown of mcaS and FlhDCDepositorInsertdCas9
UseCRISPRMutationadditional tetOAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
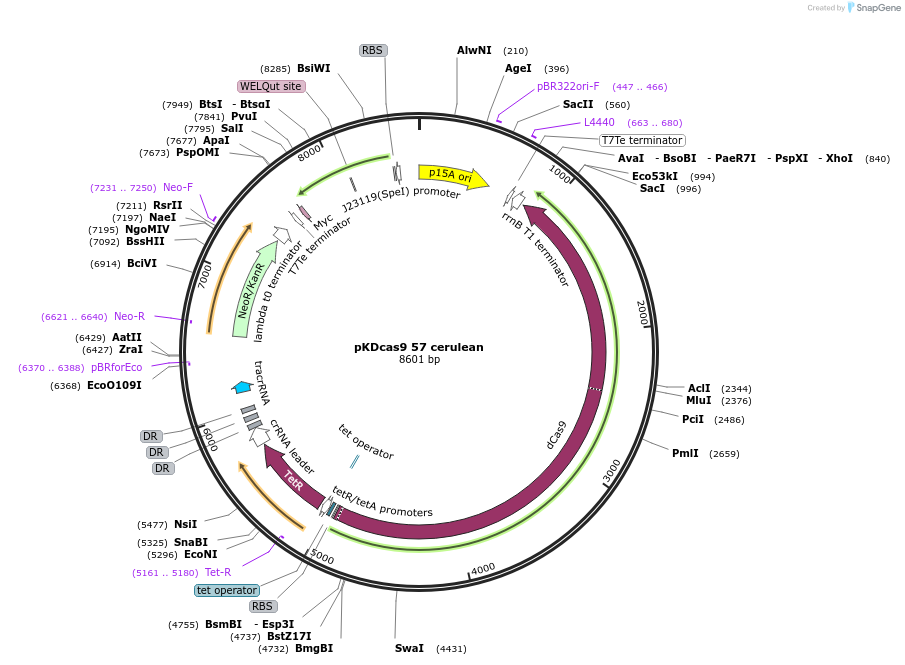
pKDcas9 57 cerulean
Plasmid#230959PurposeaTc inducible knockdown of CsgD and FliZDepositorInsertdCas9
UseCRISPRMutationadditional tetOAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
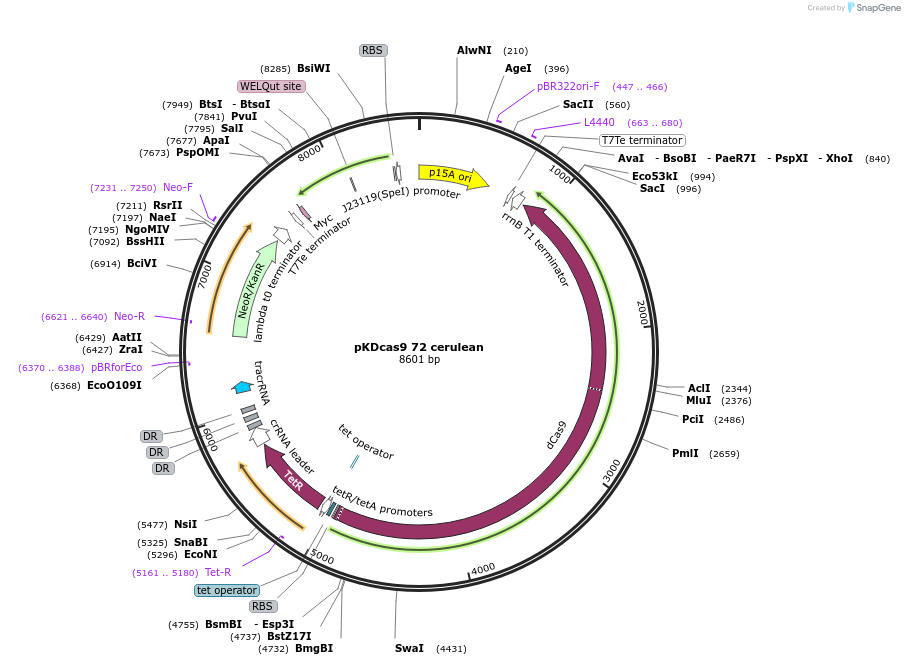
pKDcas9 72 cerulean
Plasmid#230961PurposeaTc inducible knockdown of FliZ and FisDepositorInsertdCas9
UseCRISPRMutationadditional tetOAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
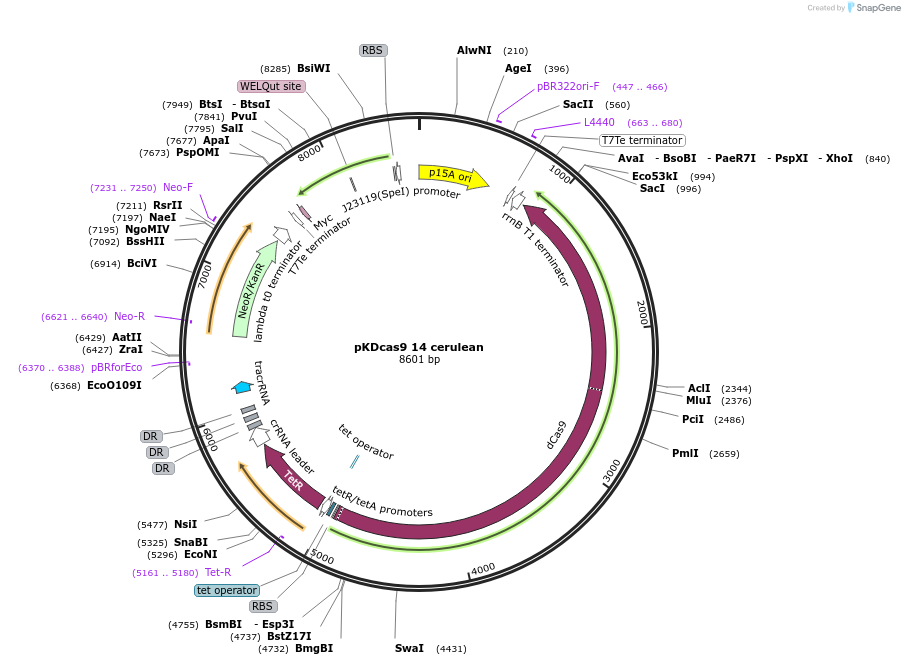
pKDcas9 14 cerulean
Plasmid#230962PurposeaTc inducible knockdown of CRP and mcaSDepositorInsertdCas9
UseCRISPRMutationadditional tetOAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
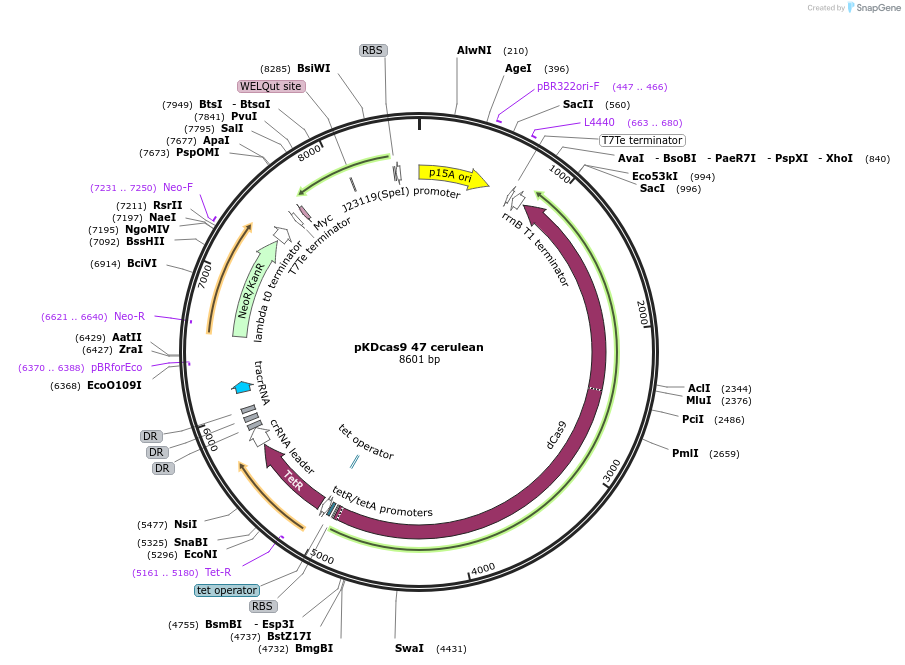
pKDcas9 47 cerulean
Plasmid#230965PurposeaTc inducible knockdown of mcaS and FliZDepositorInsertdCas9
UseCRISPRMutationadditional tetOAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
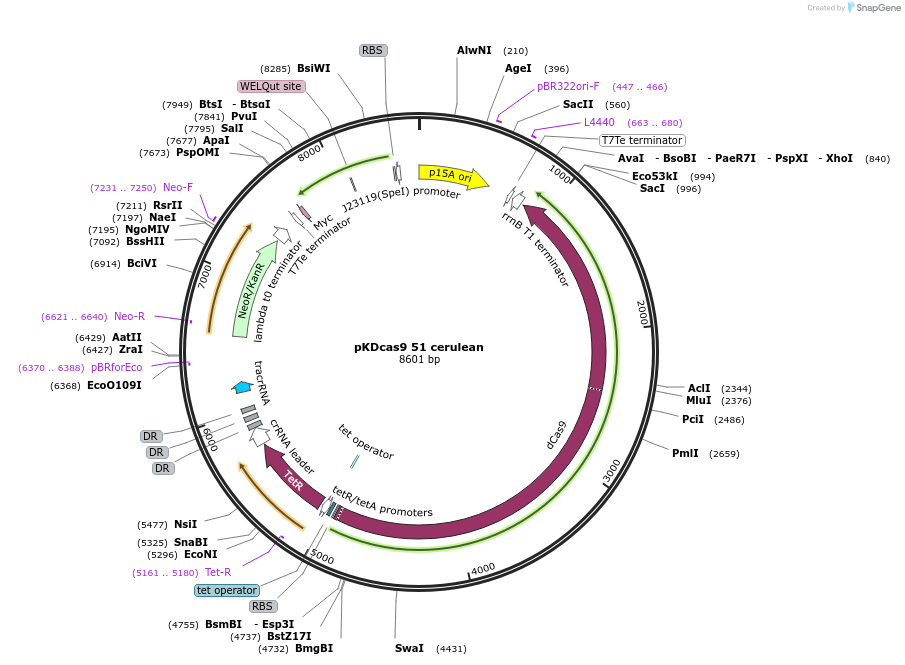
pKDcas9 51 cerulean
Plasmid#230966PurposeaTc inducible knockdown of CsgD and CRPDepositorInsertdCas9
UseCRISPRMutationadditional tetOAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
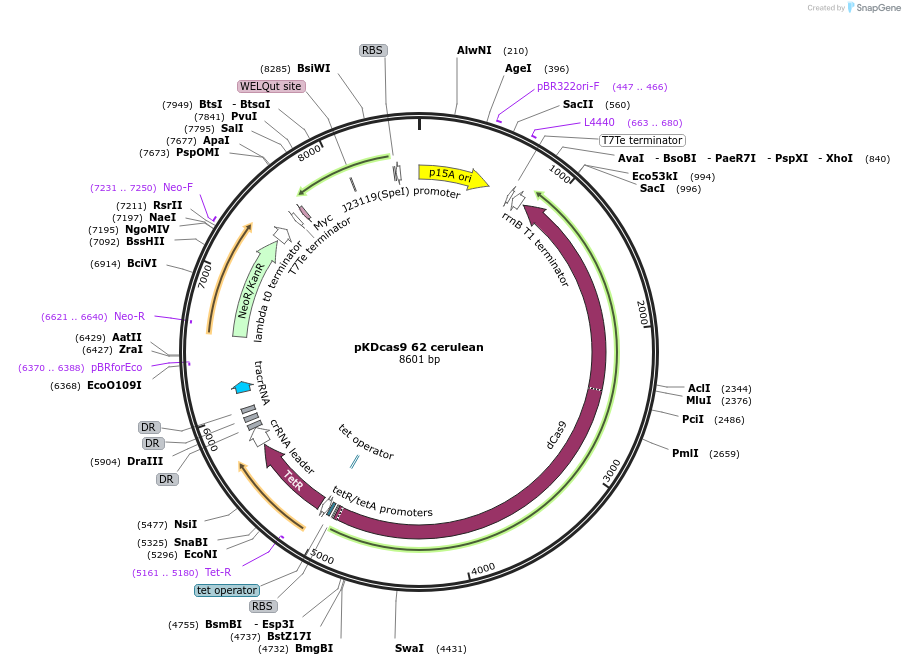
pKDcas9 62 cerulean
Plasmid#230967PurposeaTc inducible knockdown of FlhDC and FisDepositorInsertdCas9
UseCRISPRMutationadditional tetOAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
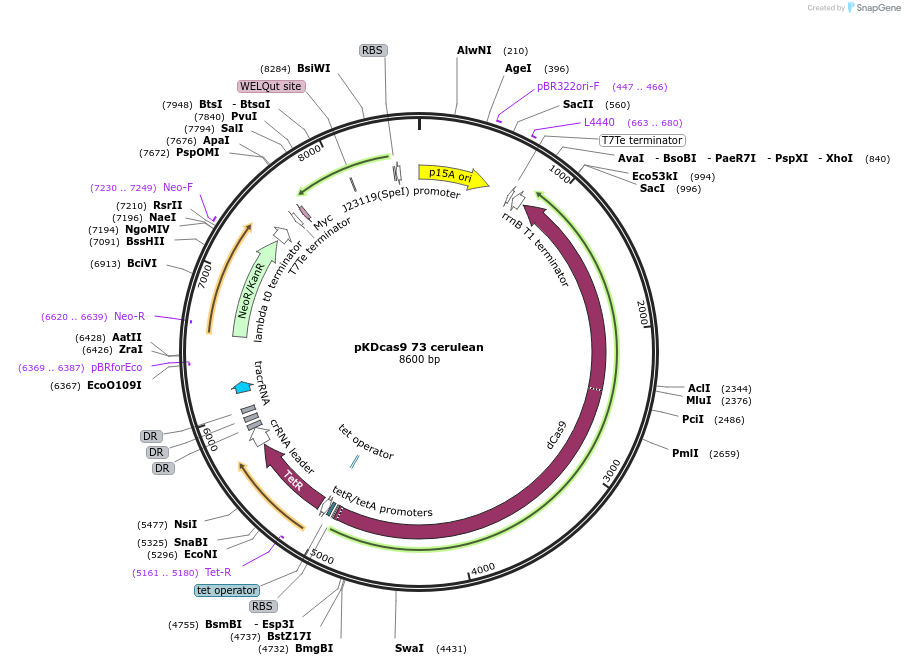
pKDcas9 73 cerulean
Plasmid#230968PurposeaTc inducible knockdown of FliZ and HNSDepositorInsertdCas9
UseCRISPRMutationadditional tetOAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
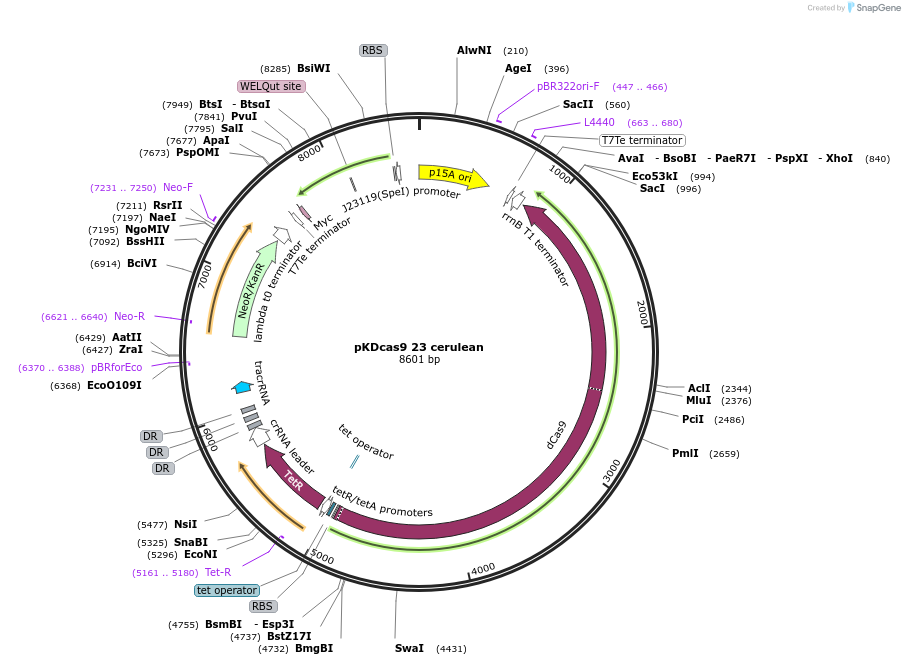
pKDcas9 23 cerulean
Plasmid#230949PurposeaTc inducible knockdown of Fis and HNSDepositorInsertdCas9
UseCRISPRMutationadditional tetOAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
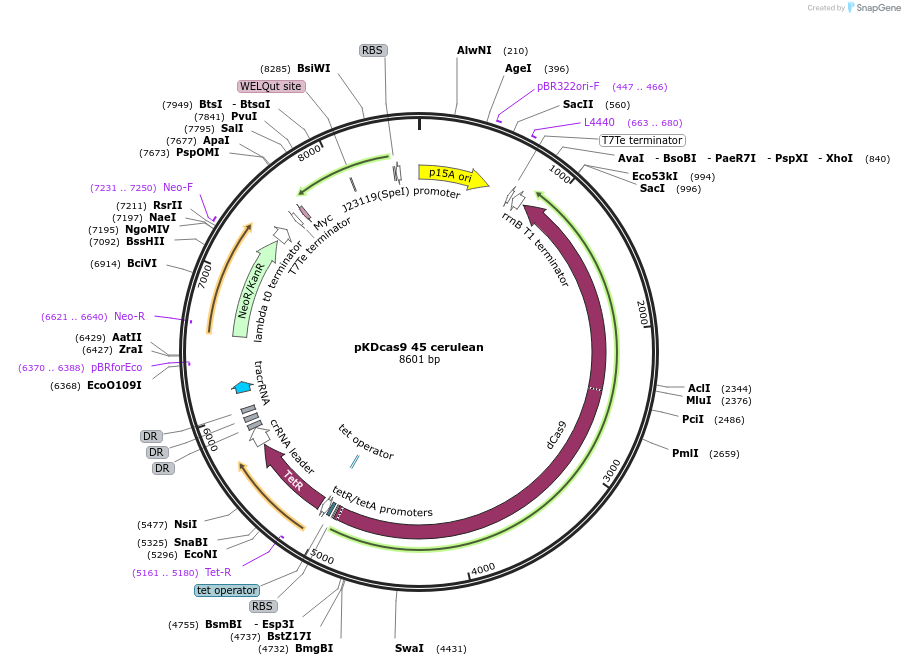
pKDcas9 45 cerulean
Plasmid#230951PurposeaTc inducible knockdown of mcaS and CsgDDepositorInsertdCas9
UseCRISPRMutationadditional tetOAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
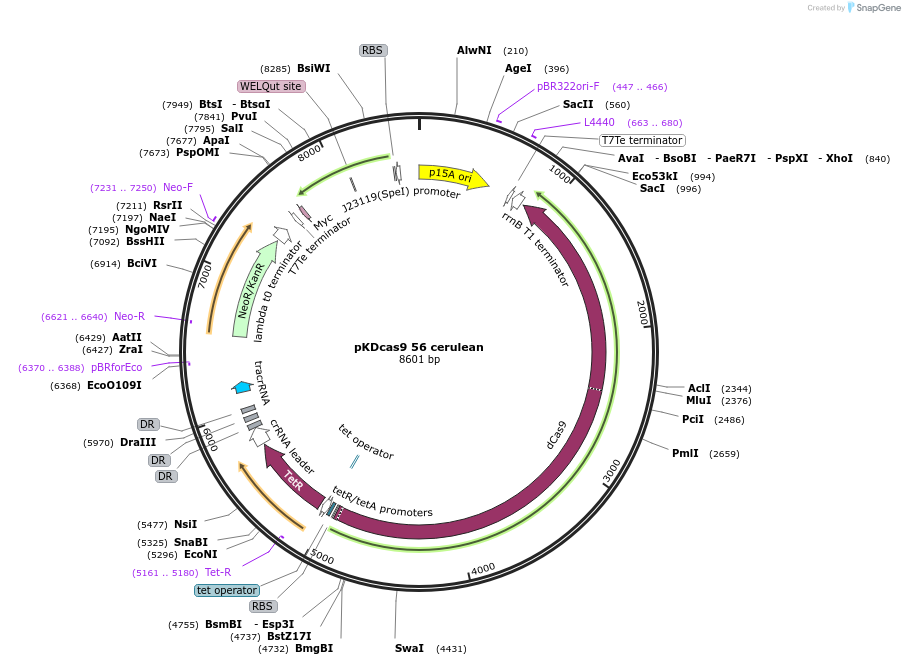
pKDcas9 56 cerulean
Plasmid#230952PurposeaTc inducible knockdown of CsgD and FlhDCDepositorInsertdCas9
UseCRISPRMutationadditional tetOAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
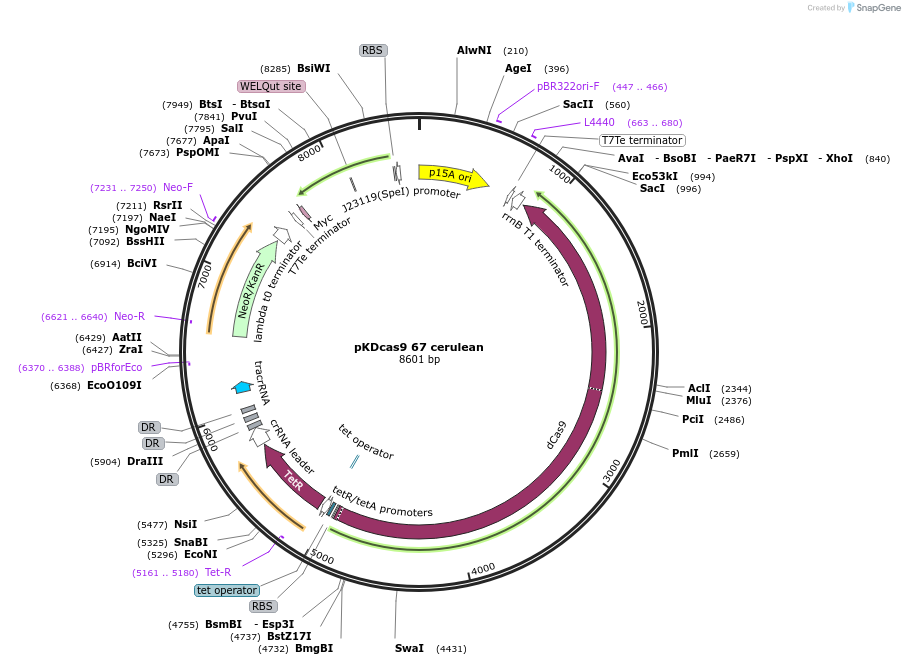
pKDcas9 67 cerulean
Plasmid#230953PurposeaTc inducible knockdown of FlhDC and FliZDepositorInsertdCas9
UseCRISPRMutationadditional tetOAvailable SinceAug. 20, 2025AvailabilityAcademic Institutions and Nonprofits only -
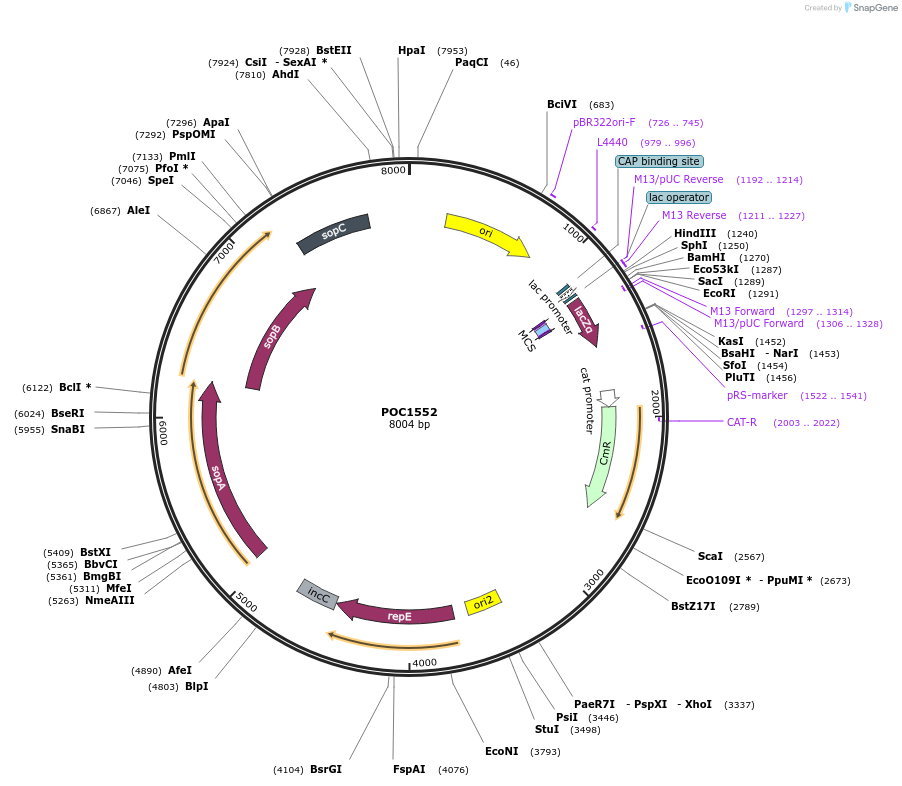
POC1552
Plasmid#226499PurposepMOBC360_StandVC: Standard Vector (Chloramphenicol R) to be methylated by the M.Osp807II BsaI-associated switch methylase. Used for the assembly using the methylation protectionDepositorTypeEmpty backboneUseSynthetic BiologyAvailable SinceJune 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
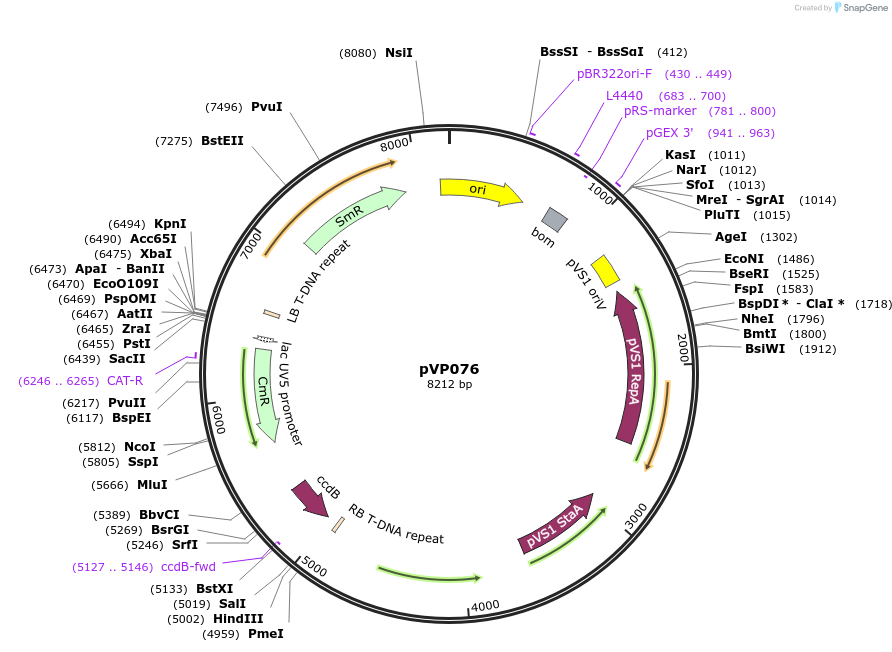
pVP076
Plasmid#222028PurposeEmpty spectinomycin resistant GreenGate destination vector based on pGGP-AG. Domesticated of Esp3I sites in the backbone and chloramphenicol resistance gene.DepositorTypeEmpty backboneUseSynthetic Biology; Greengate compatible cloning v…MutationAll Esp3i sites domesticatedAvailable SinceDec. 16, 2024AvailabilityIndustry, Academic Institutions, and Nonprofits -
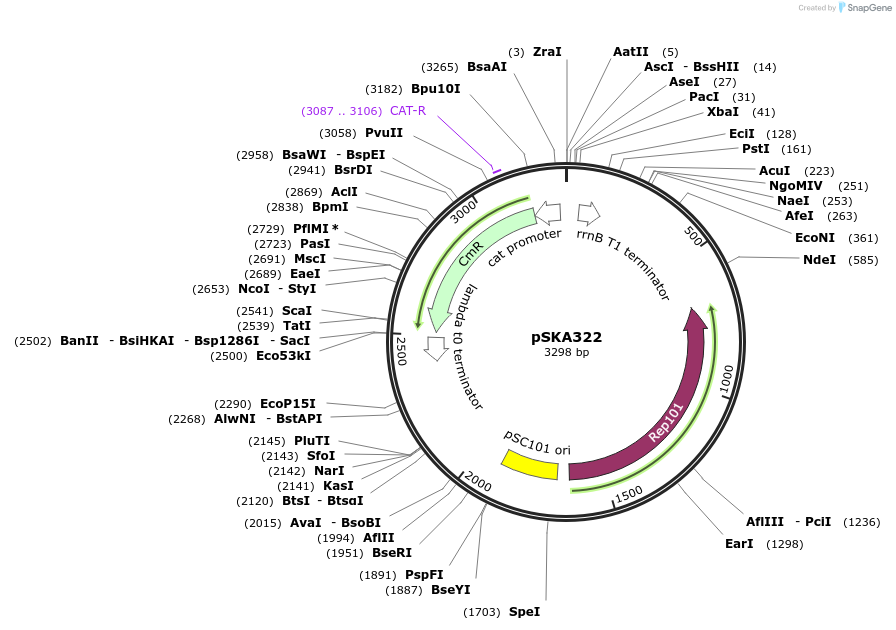
pSKA322
Plasmid#101676Purposeempty backbone with chloramphenicol resistanceDepositorTypeEmpty backboneUseSynthetic BiologyExpressionBacterialAvailable SinceSept. 14, 2022AvailabilityAcademic Institutions and Nonprofits only -
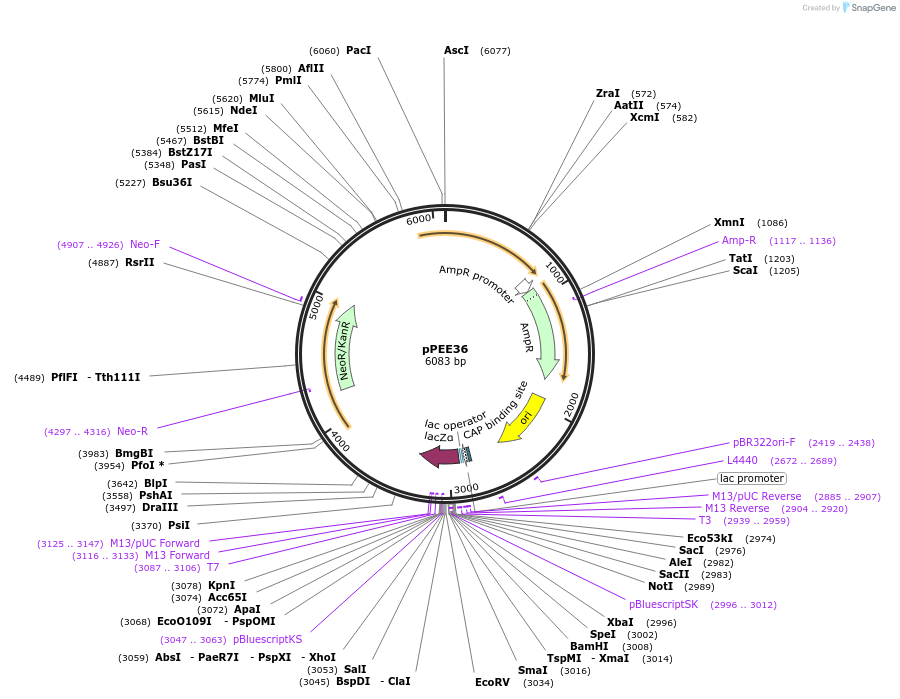
pPEE36
Plasmid#128376PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 7, a small gene-free region. Contains the NEO resistance markerDepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceApril 28, 2020AvailabilityAcademic Institutions and Nonprofits only -
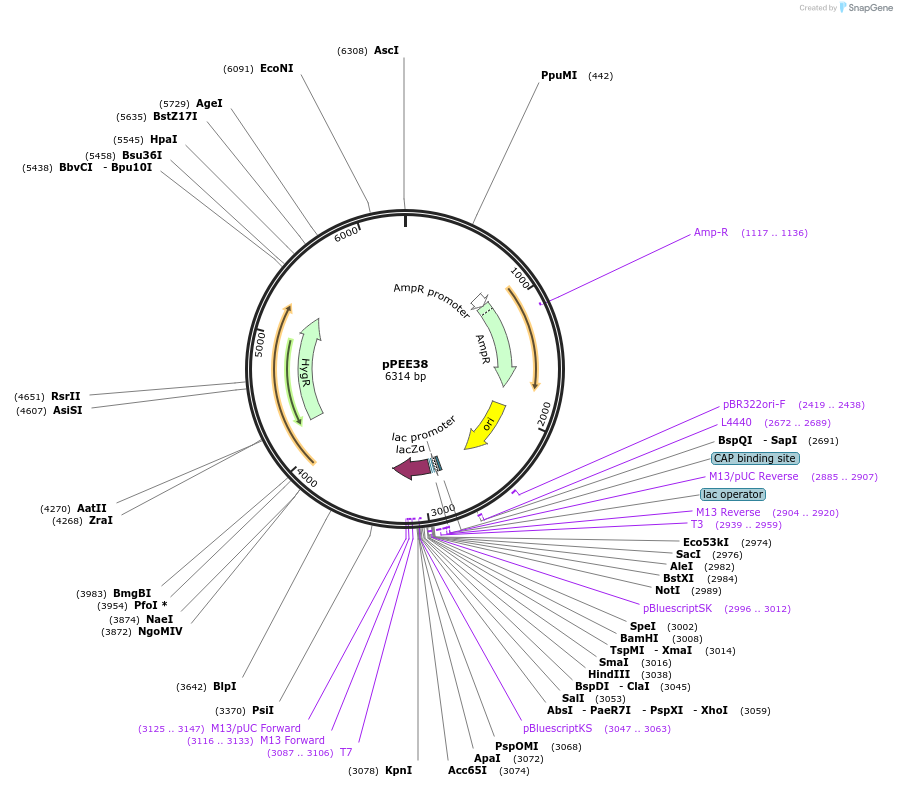
pPEE38
Plasmid#128378PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 4, a small gene-free region. Contains the HYG resistance marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceJan. 8, 2020AvailabilityAcademic Institutions and Nonprofits only -
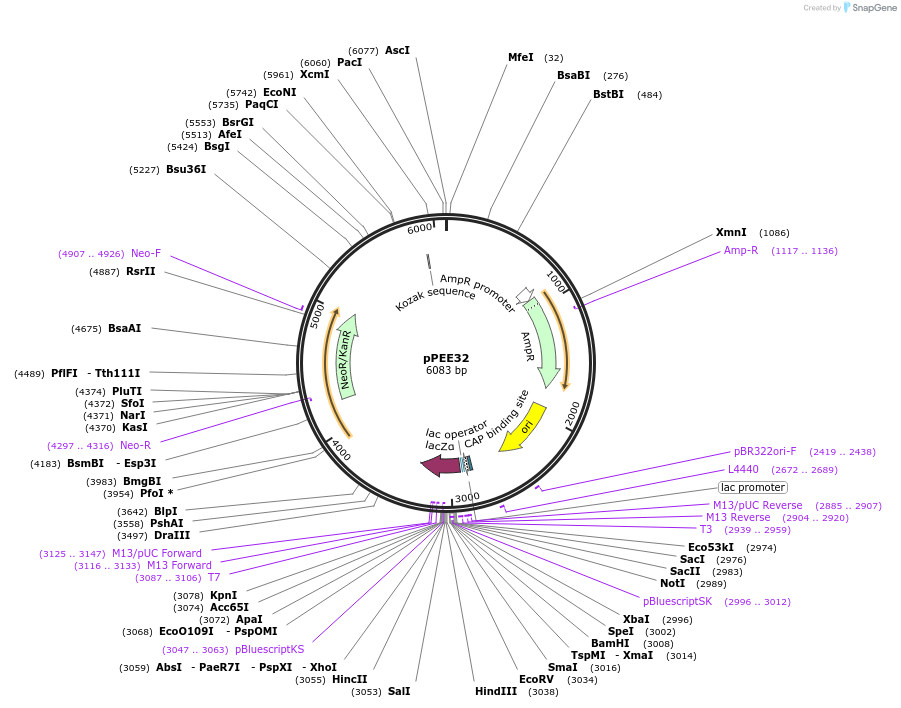
pPEE32
Plasmid#128372PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 3, a small gene-free region. Contains the NEO resistance markerDepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceJan. 8, 2020AvailabilityAcademic Institutions and Nonprofits only -
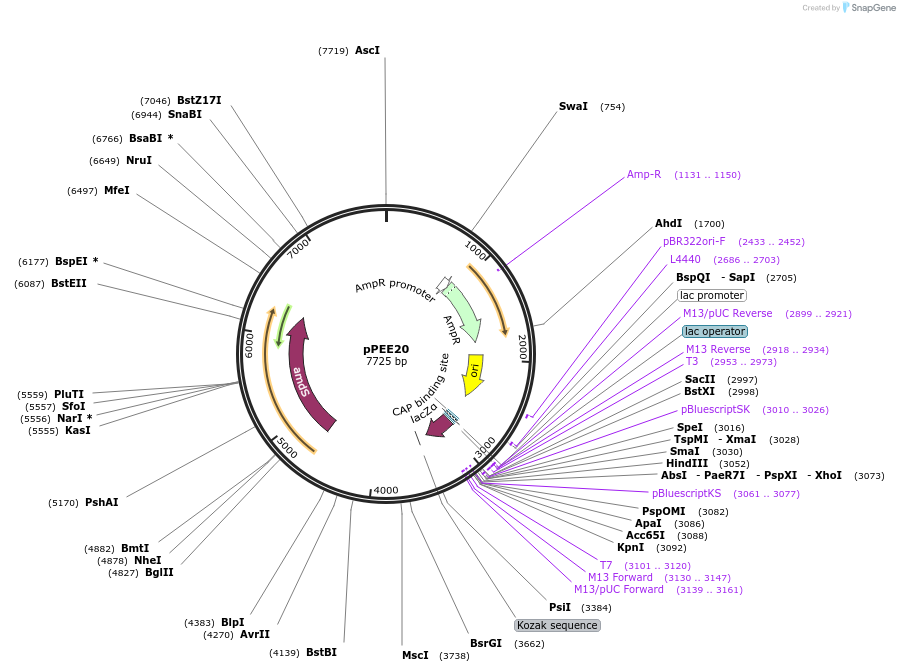
pPEE20
Plasmid#128363PurposeDominant marker in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 4, a small gene-free region. Contains the amdS2 counterselectable marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterTEF1Available SinceOct. 11, 2019AvailabilityAcademic Institutions and Nonprofits only -
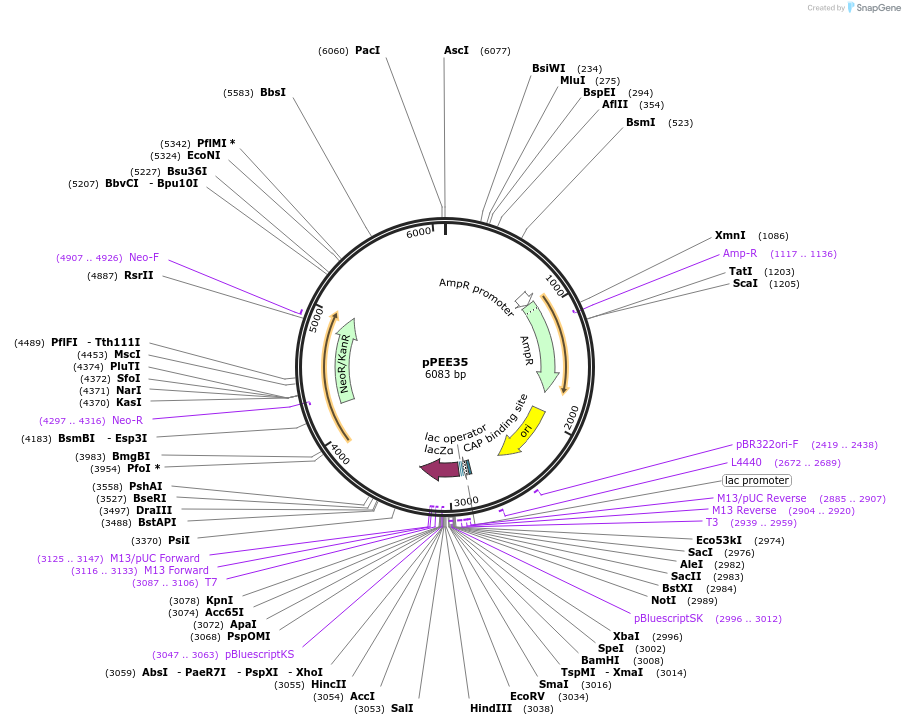
pPEE35
Plasmid#128375PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 6, a small gene-free region. Contains the NEO resistance markerDepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
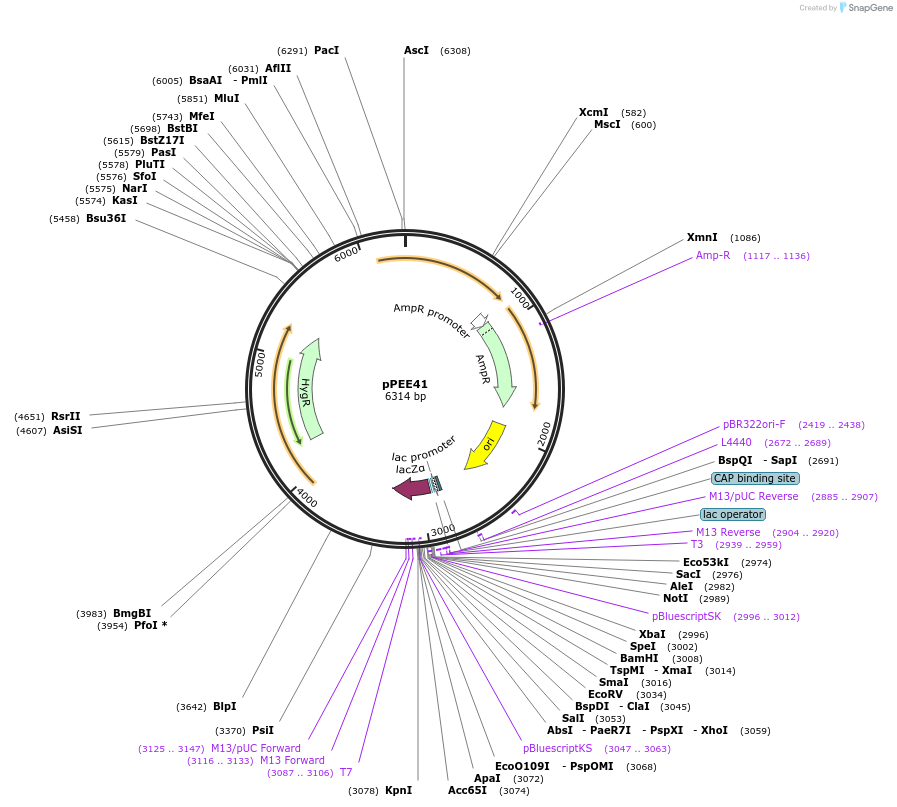
pPEE41
Plasmid#128381PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 7, a small gene-free region. Contains the HYG resistance marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
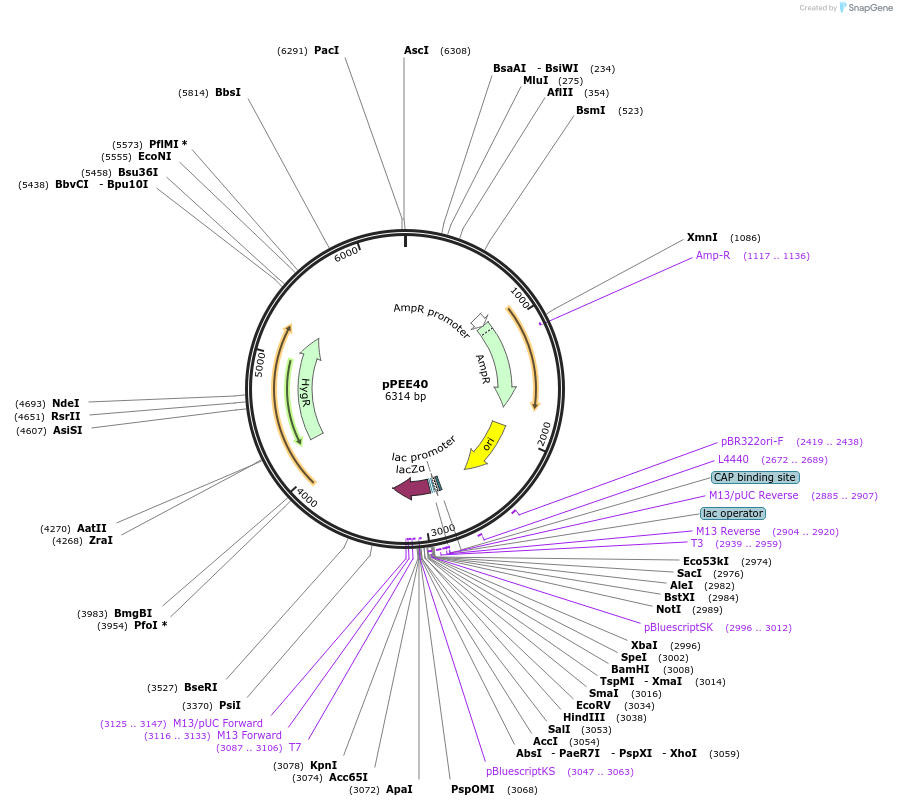
pPEE40
Plasmid#128380PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 6, a small gene-free region. Contains the HYG resistance marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
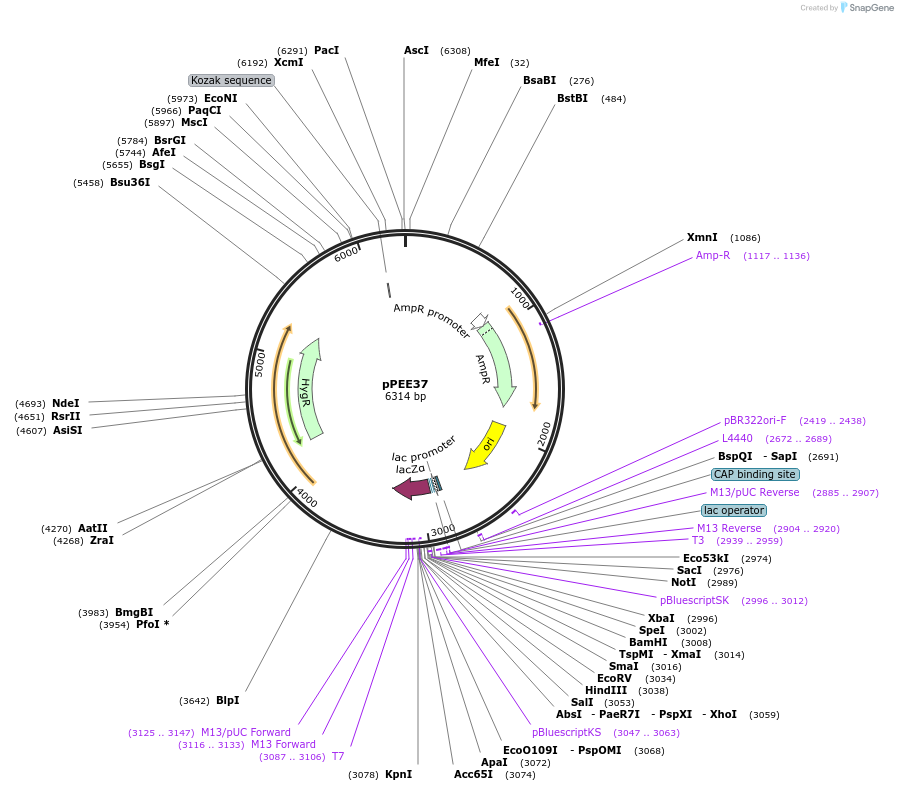
pPEE37
Plasmid#128377PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 3, a small gene-free region. Contains the HYG resistance marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
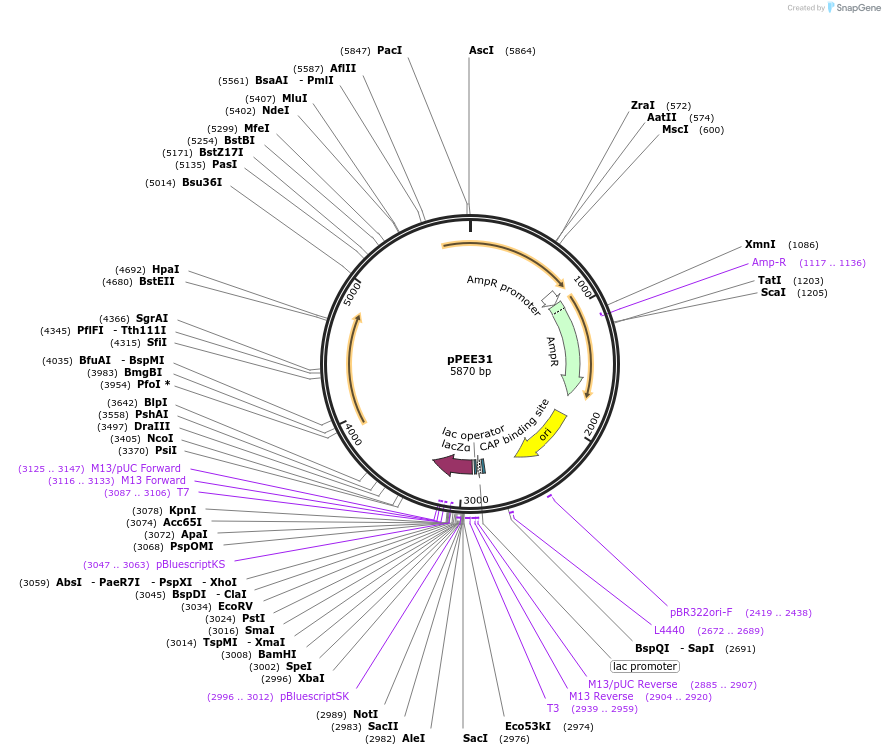
pPEE31
Plasmid#128371PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 7, a small gene-free region. Contains the nourseothricin resistance marker (NAT)DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
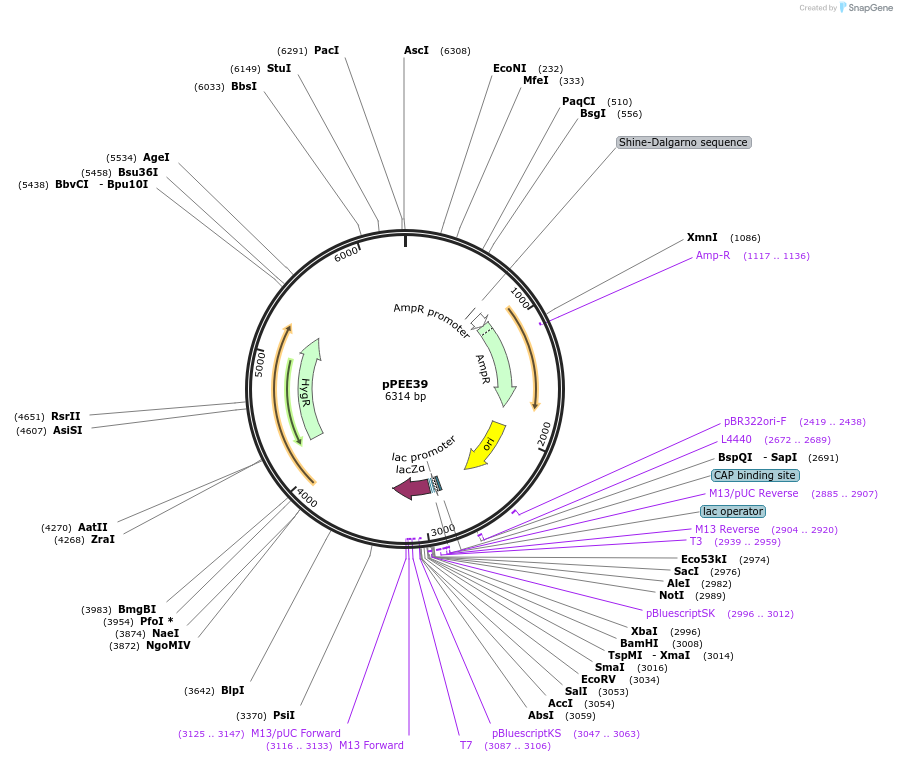
pPEE39
Plasmid#128379PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 5, a small gene-free region. Contains the HYG resistance marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
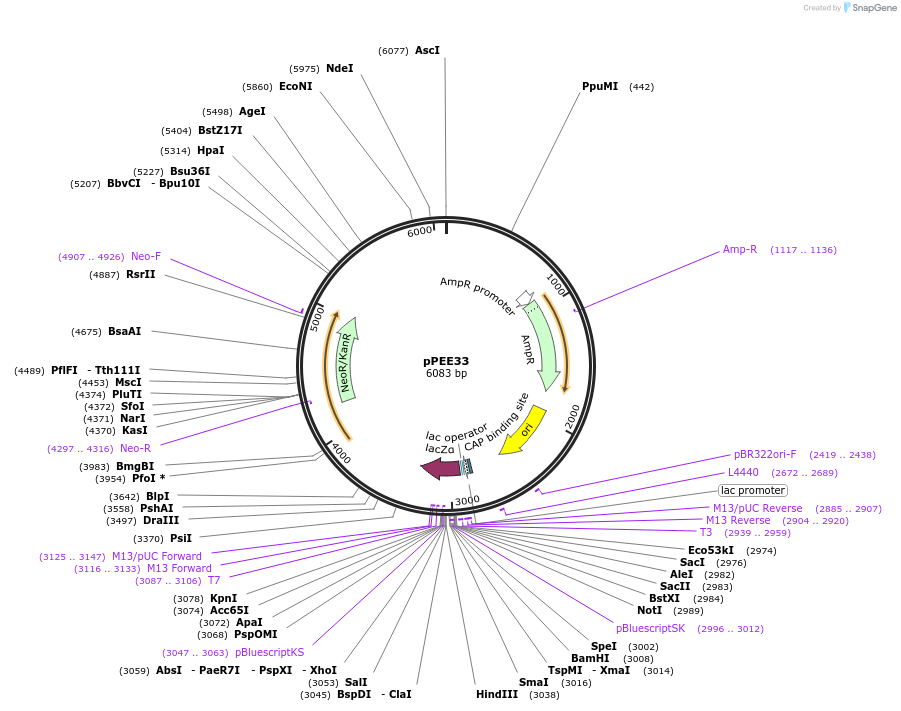
pPEE33
Plasmid#128373PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 4, a small gene-free region. Contains the NEO resistance markerDepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
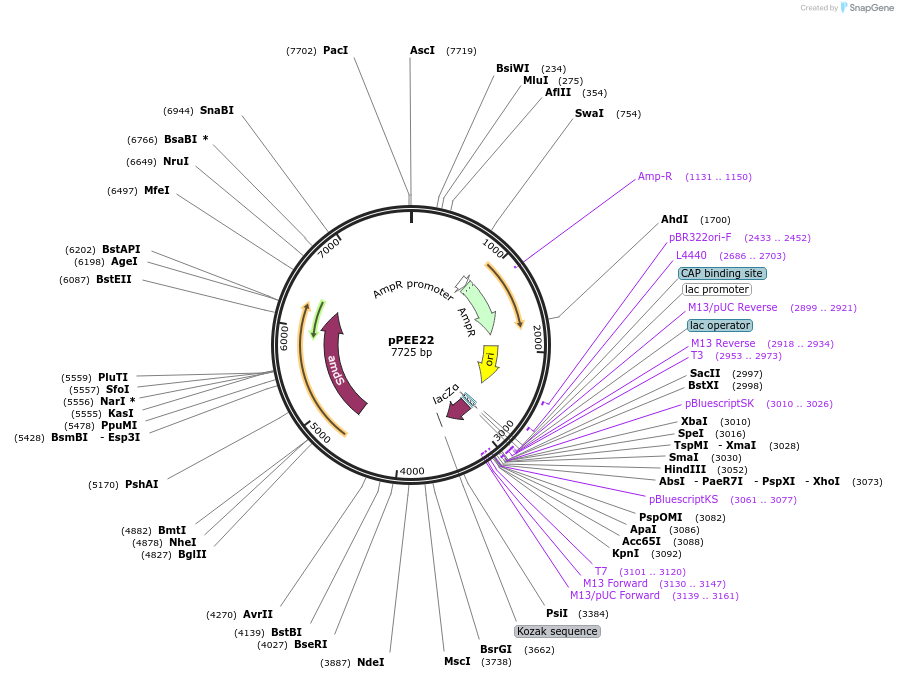
pPEE22
Plasmid#128365PurposeDominant marker in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 6, a small gene-free region. Contains the amdS2 counterselectable marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterTEF1Available SinceAug. 20, 2019AvailabilityAcademic Institutions and Nonprofits only -
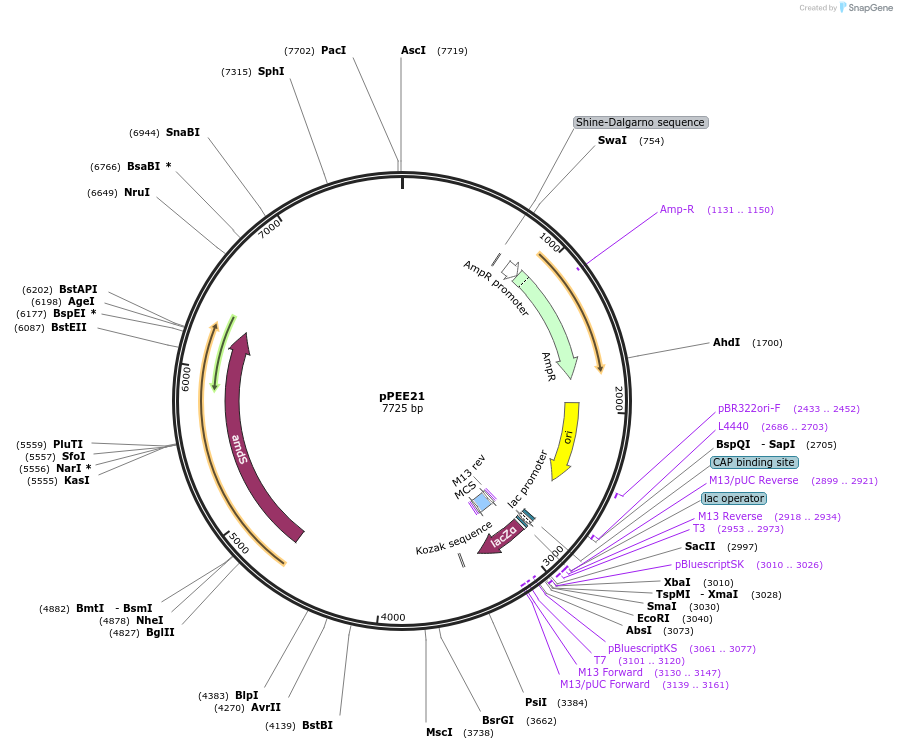
pPEE21
Plasmid#128364PurposeDominant marker in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 5, a small gene-free region. Contains the amdS2 counterselectable marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterTEF1Available SinceAug. 19, 2019AvailabilityAcademic Institutions and Nonprofits only -
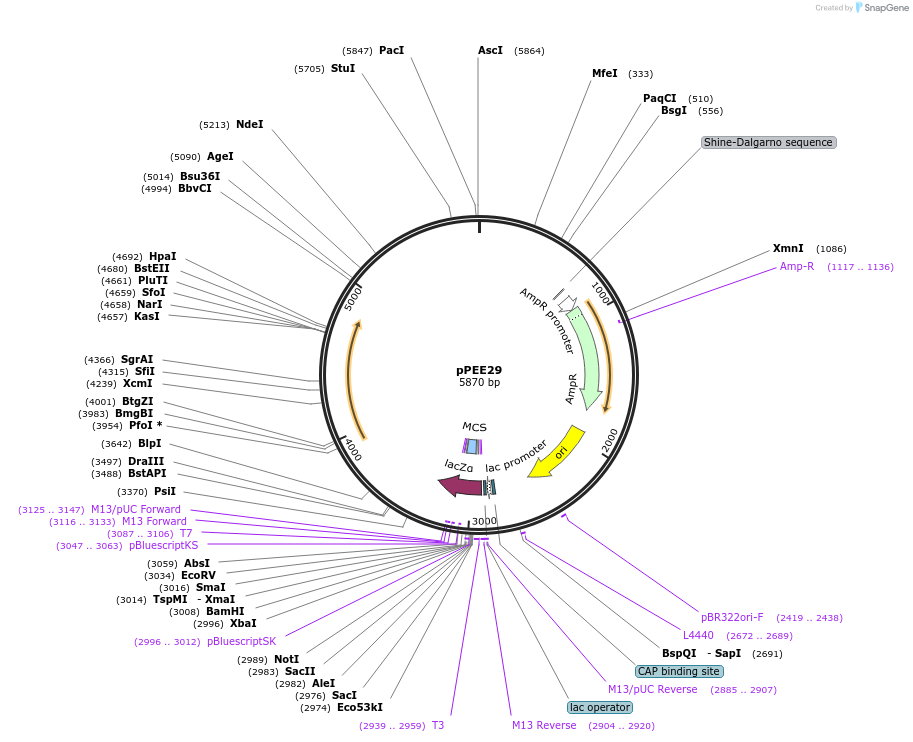
pPEE29
Plasmid#128369PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 5, a small gene-free region. Contains the nourseothricin resistance marker (NAT)DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
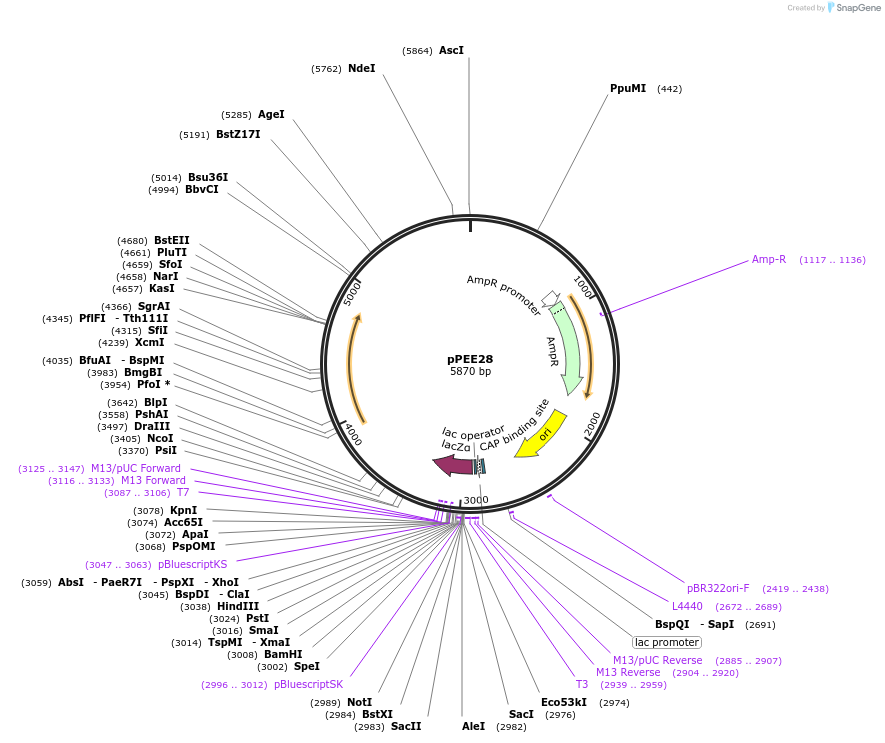
pPEE28
Plasmid#128368PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 4, a small gene-free region. Contains the nourseothricin resistance marker (NAT)DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
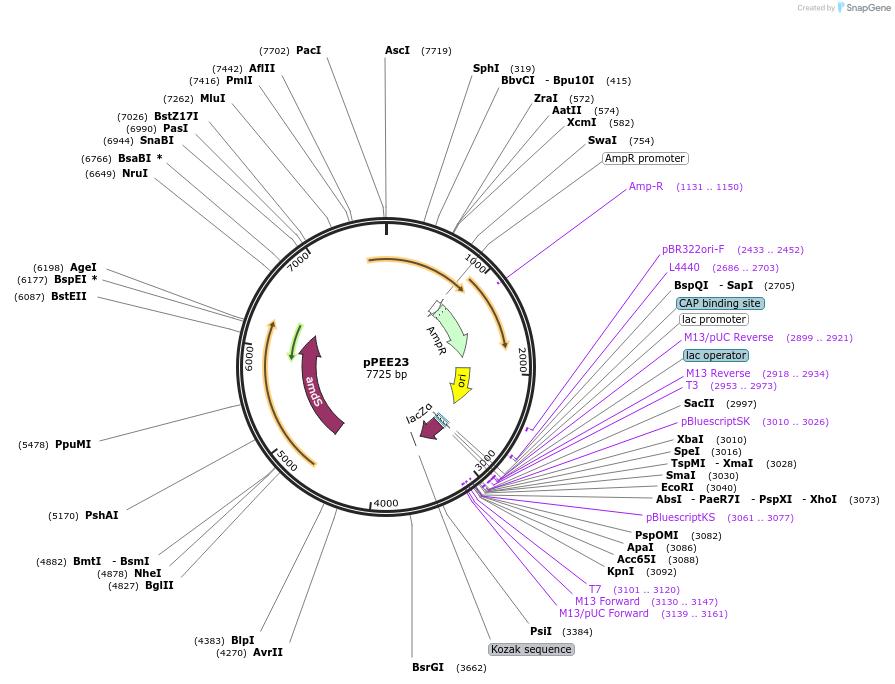
pPEE23
Plasmid#128366PurposeDominant marker in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 7, a small gene-free region. Contains the amdS2 counterselectable marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterTEF1Available SinceAug. 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
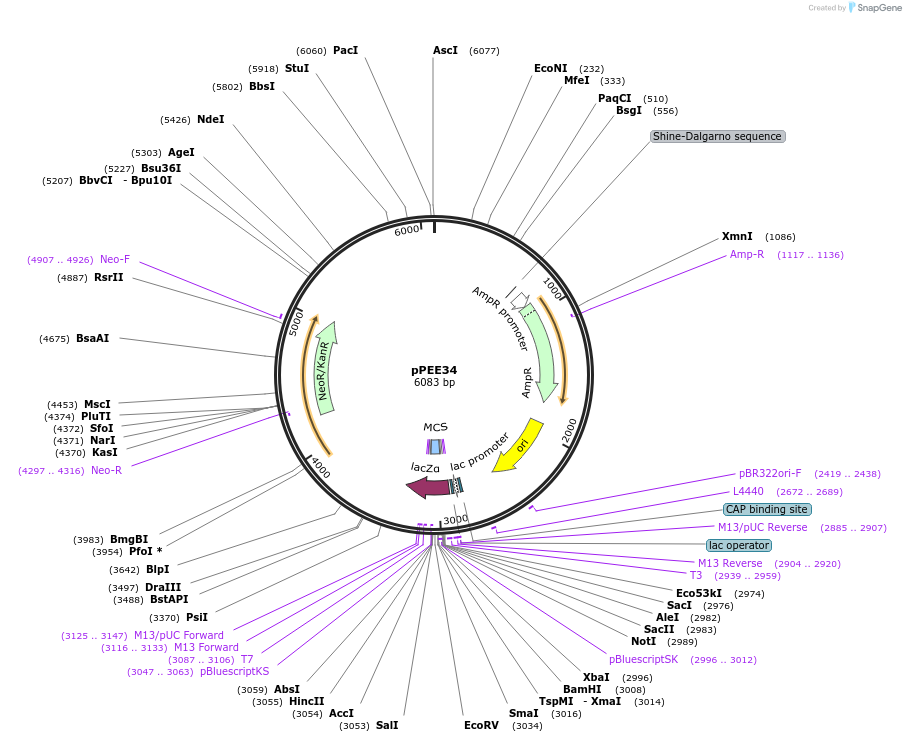
pPEE34
Plasmid#128374PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 5, a small gene-free region. Contains the NEO resistance markerDepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
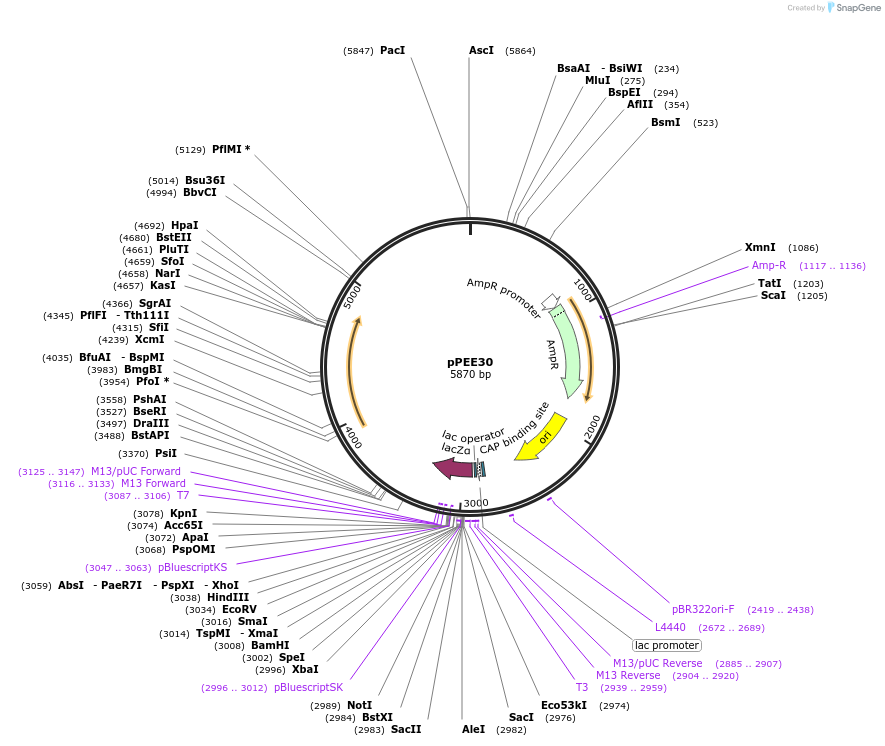
pPEE30
Plasmid#128370PurposeComplementation in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 6, a small gene-free region. Contains the nourseothricin resistance marker (NAT)DepositorTypeEmpty backboneUseUnspecifiedPromoterACT1Available SinceAug. 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
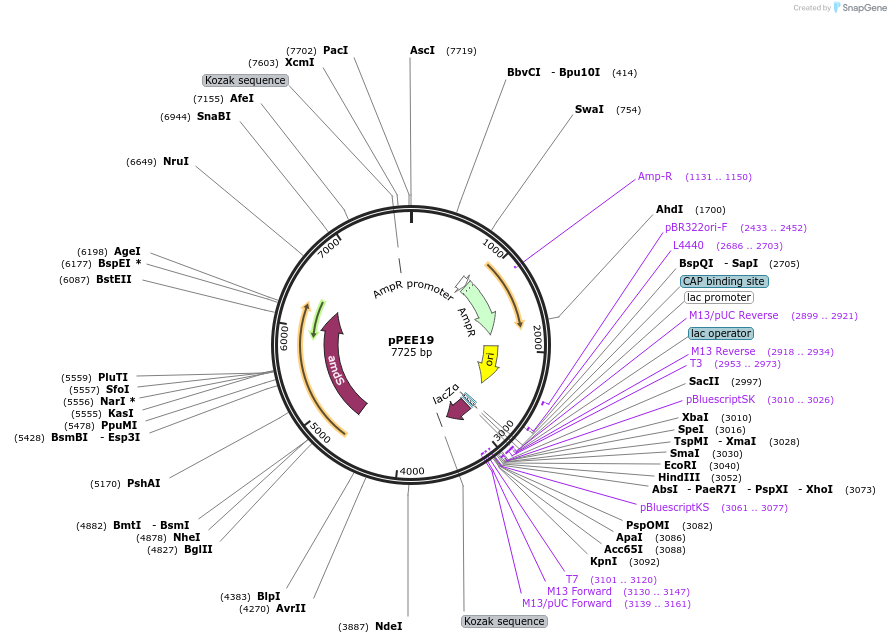
pPEE19
Plasmid#128362PurposeDominant marker in Cryptococcus neoformans. Targets DNA constructs to a Safe Haven Site 3, a small gene-free region. Contains the amdS2 counterselectable marker.DepositorTypeEmpty backboneUseUnspecifiedPromoterTEF1Available SinceAug. 13, 2019AvailabilityAcademic Institutions and Nonprofits only -
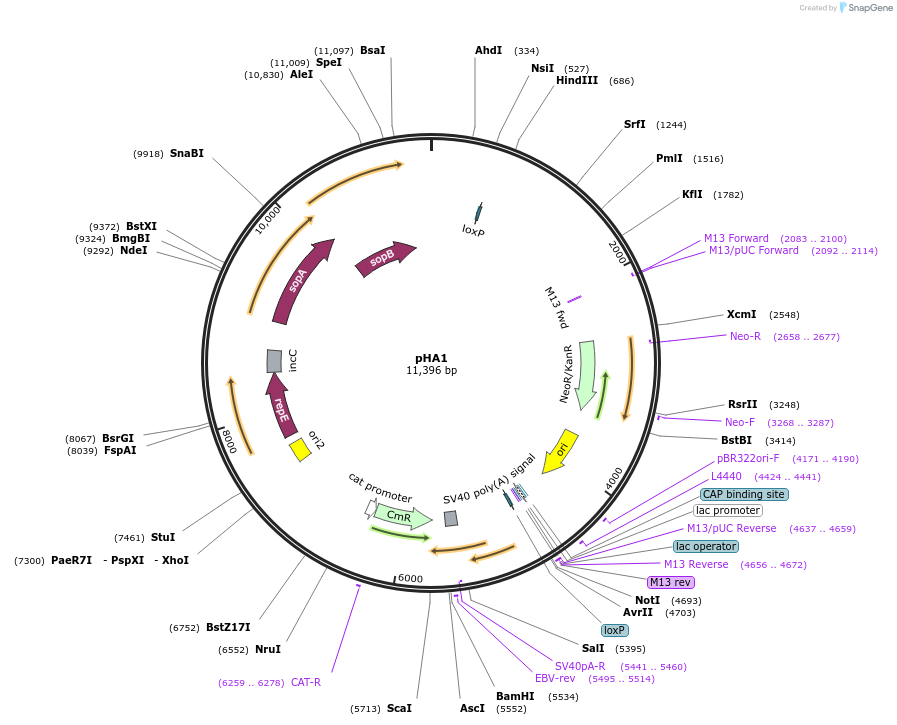
pHA1
Plasmid#85709PurposeThis plasmid provides a BAC vector backbone, which can be used for cloning of BAC inserts, for instance, herpesvirus genomesDepositorTypeEmpty backboneUseCre/Lox; Herpesvirus cloningExpressionBacterialAvailable SinceJune 28, 2019AvailabilityAcademic Institutions and Nonprofits only -
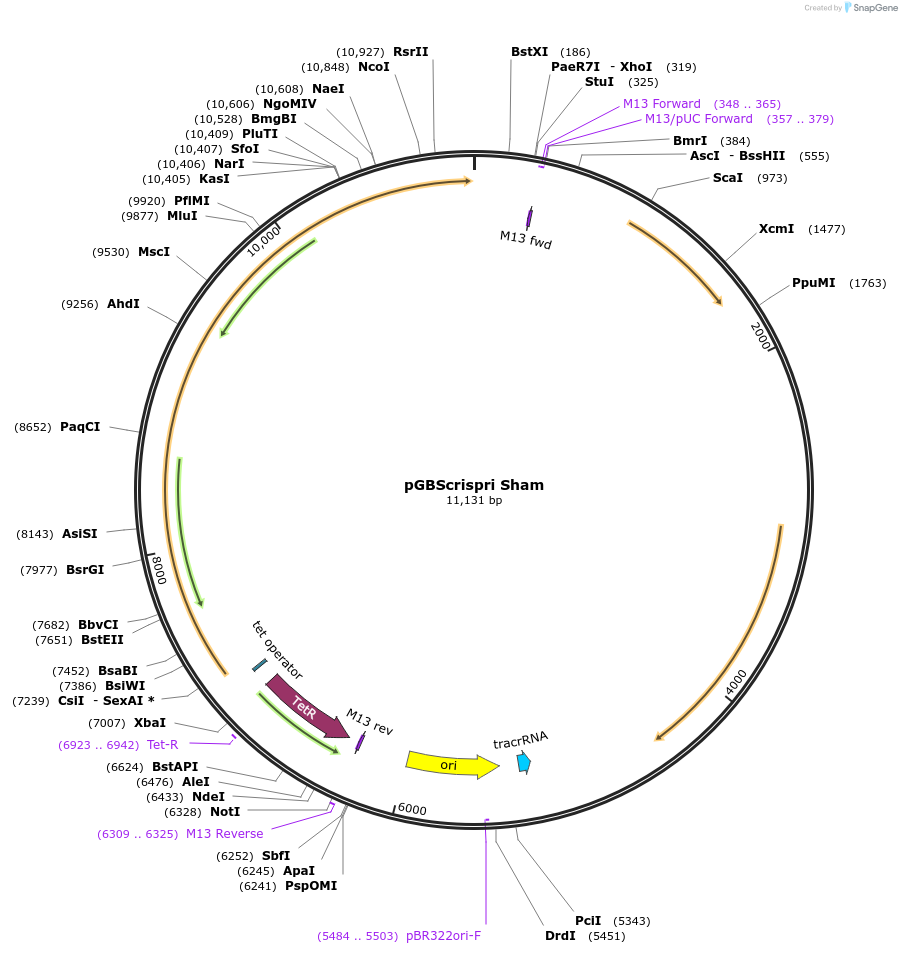
pGBScrispri Sham
Plasmid#223201PurposeGroup B Streptococcus inducible dCas12 CRISPRi with Modular crRNADepositorInsertN/A
UseCRISPRAvailable SinceDec. 19, 2024AvailabilityAcademic Institutions and Nonprofits only -
pSAM_Cam
Plasmid#91570PurposeMariner transposon mutagenesis & sequencing vector - Transposon contains KanR and Illumina sequencing adapters. Lac promoter expresses transposase. Chloramphenicol resistance on backbone.DepositorTypeEmpty backboneExpressionBacterialAvailable SinceAug. 10, 2017AvailabilityAcademic Institutions and Nonprofits only -
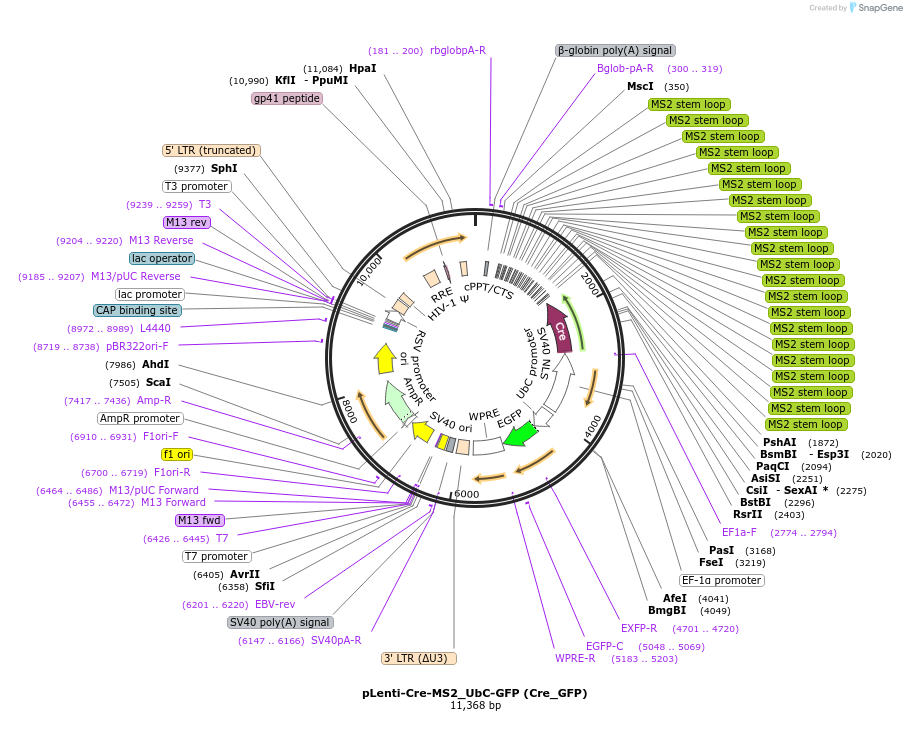
pLenti-Cre-MS2_UbC-GFP (Cre_GFP)
Plasmid#219536Purposea. Expresses Cre recombinase tagged with 21x repeats of MS2 binding sequence under EF1A promotor and b. GFP reporter under UbC promotorDepositorAvailable SinceNov. 11, 2024AvailabilityIndustry, Academic Institutions, and Nonprofits -
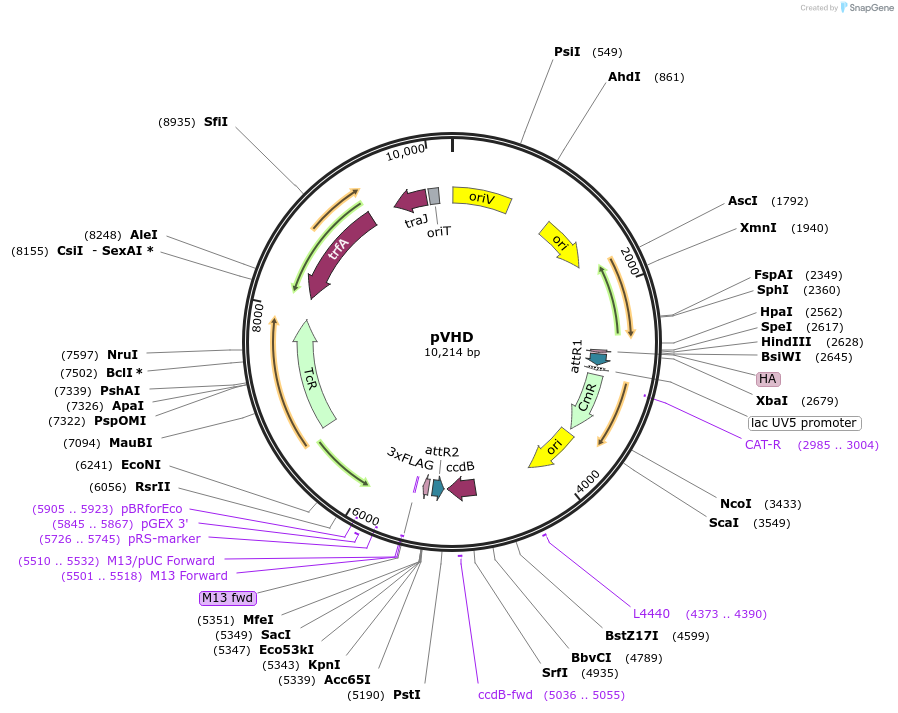
pVHD
Plasmid#61303PurposeVanillate-inducible gene expression plasmid for Sphingomonas melonis Fr1; destination vector for Gateway cloning, allowing C-terminal fusion of proteins to HA tagDepositorTypeEmpty backboneTagsHAExpressionBacterialPromoterPV10Available SinceAug. 3, 2015AvailabilityAcademic Institutions and Nonprofits only