We narrowed to 974 results for: ggct
-
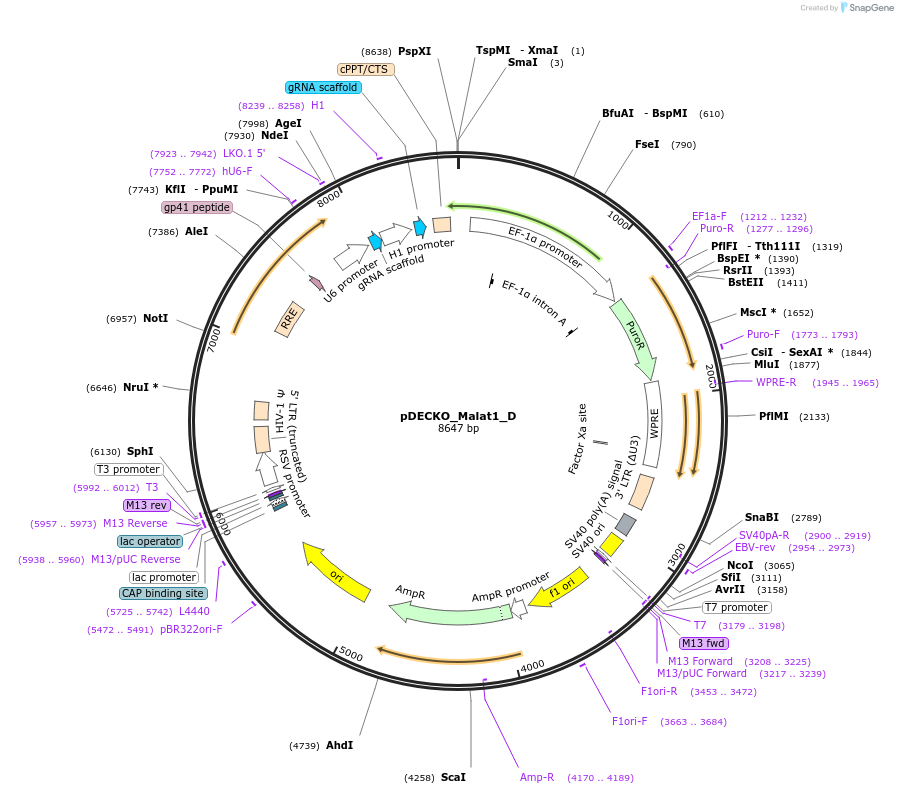
Plasmid#72623PurposeExpresses two gRNAs targeting MALAT1 promoterDepositorInsertgRNAs toward Malat1
ExpressionMammalianPromoterU6 (gRNA1) and H1 (gRNA2)Available SinceJune 14, 2016AvailabilityAcademic Institutions and Nonprofits only -
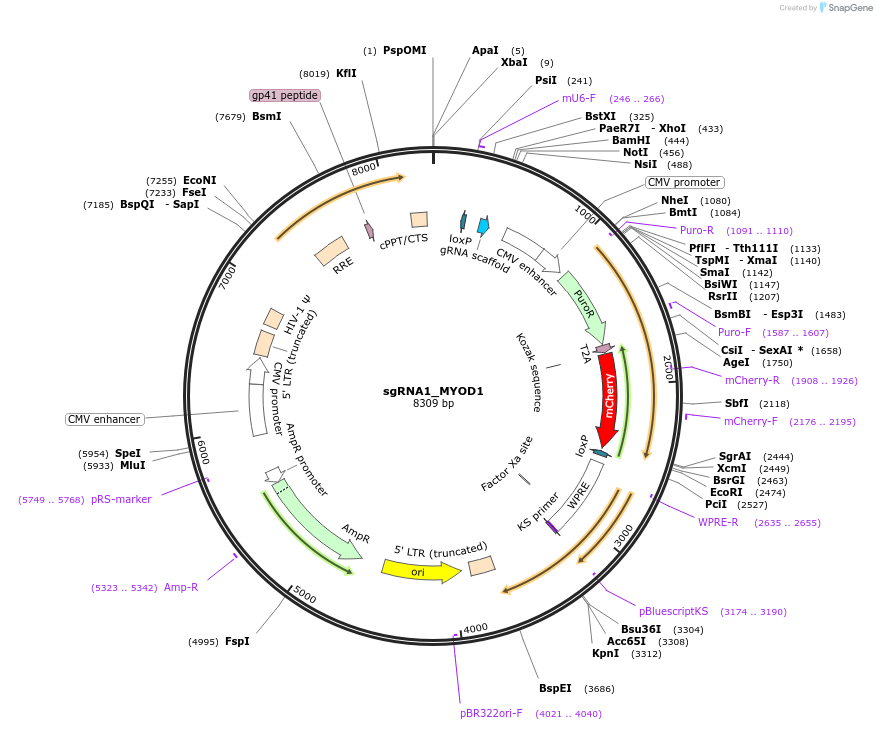
sgRNA1_MYOD1
Plasmid#64136PurposePhotoactivatable transcription system. Lentiviral expression of MYOD1 sgRNA1. Also contains a CMV-puro-t2A-mCherry expression cassette.DepositorAvailable SinceApril 15, 2015AvailabilityAcademic Institutions and Nonprofits only -
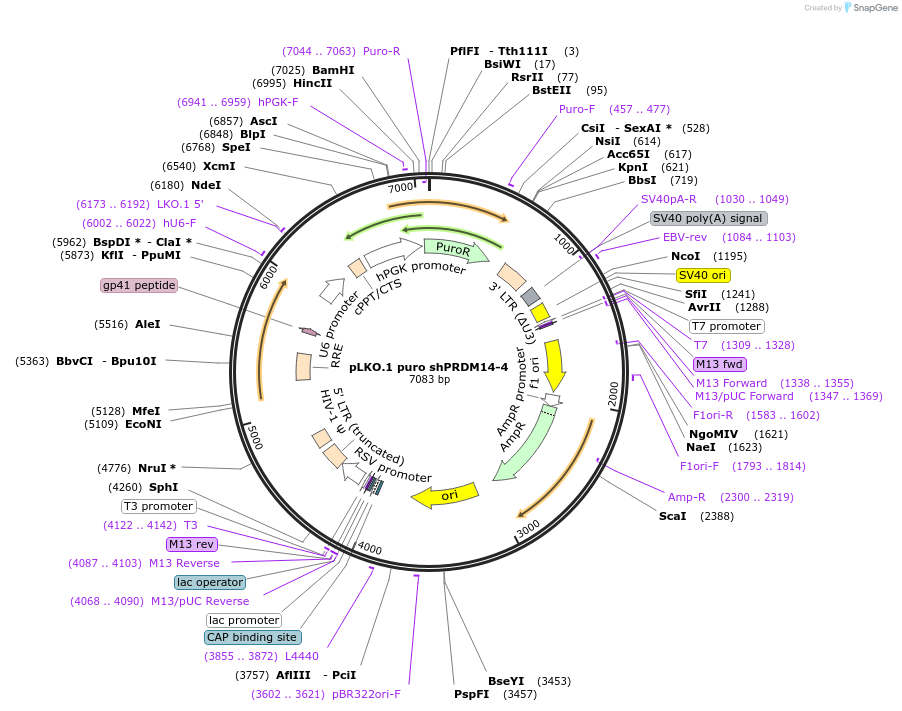
pLKO.1 puro shPRDM14-4
Plasmid#193697PurposeConstitutive lentiviral expression of PRDM14 shRNADepositorInsertPRDM14 (PRDM14 Human)
UseLentiviralAvailable SinceDec. 19, 2022AvailabilityAcademic Institutions and Nonprofits only -
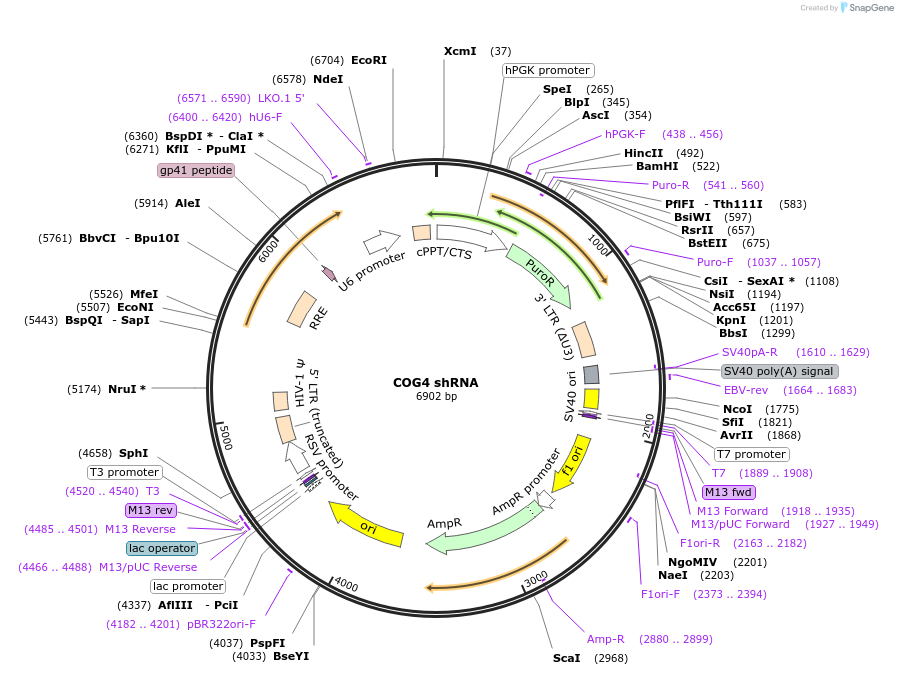
COG4 shRNA
Plasmid#163861PurposeshRNA targeting the coding sequences of COG4DepositorInsertcomponent of oligomeric golgi complex 4 (COG4 Human)
ExpressionMammalianAvailable SinceNov. 24, 2021AvailabilityAcademic Institutions and Nonprofits only -
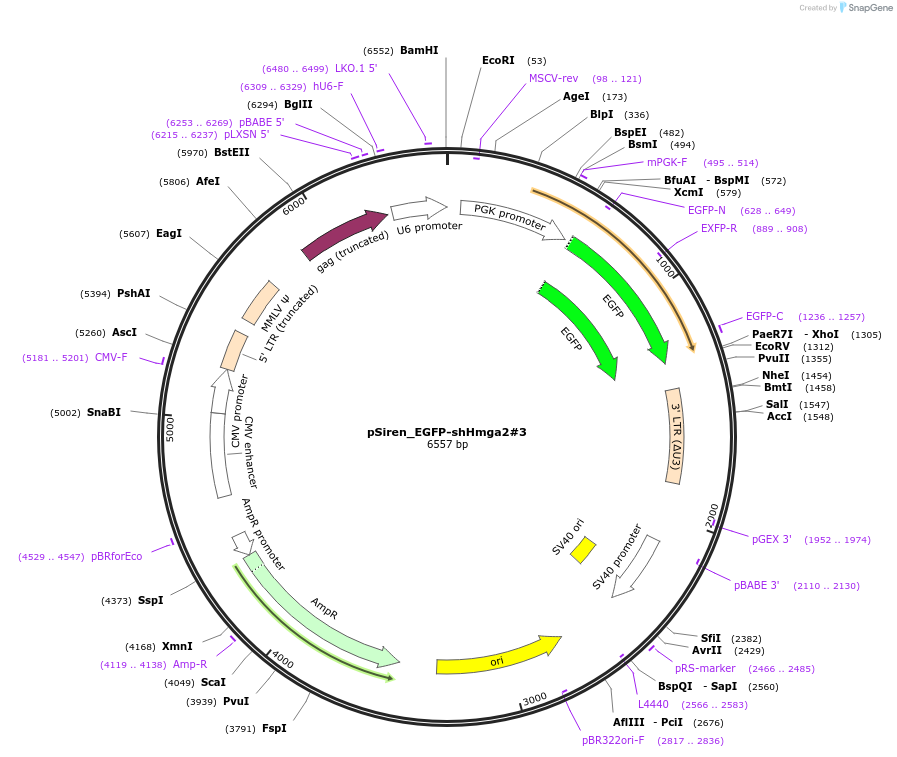
pSiren_EGFP-shHmga2#3
Plasmid#122291PurposeKnockdown for mouse Hmga2DepositorAvailable SinceMarch 22, 2019AvailabilityAcademic Institutions and Nonprofits only -
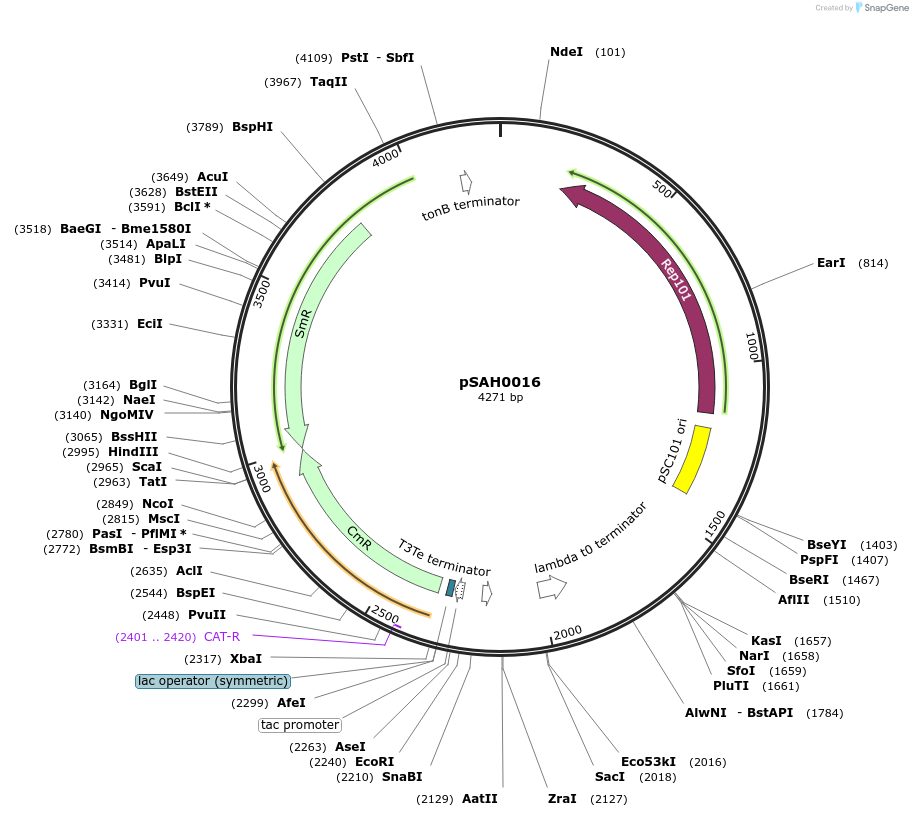
pSAH0016
Plasmid#79824PurposeDual selection plasmid with lac operator, selection for binding with spectinomycin, for non-binding with chloramphenicolDepositorInsertscat
aadA
PromoterAATTCAAAAAAATGCTTGACAATTAATCATCGGCTCGTATAATGTGTGG…Available SinceAug. 2, 2016AvailabilityAcademic Institutions and Nonprofits only -
-
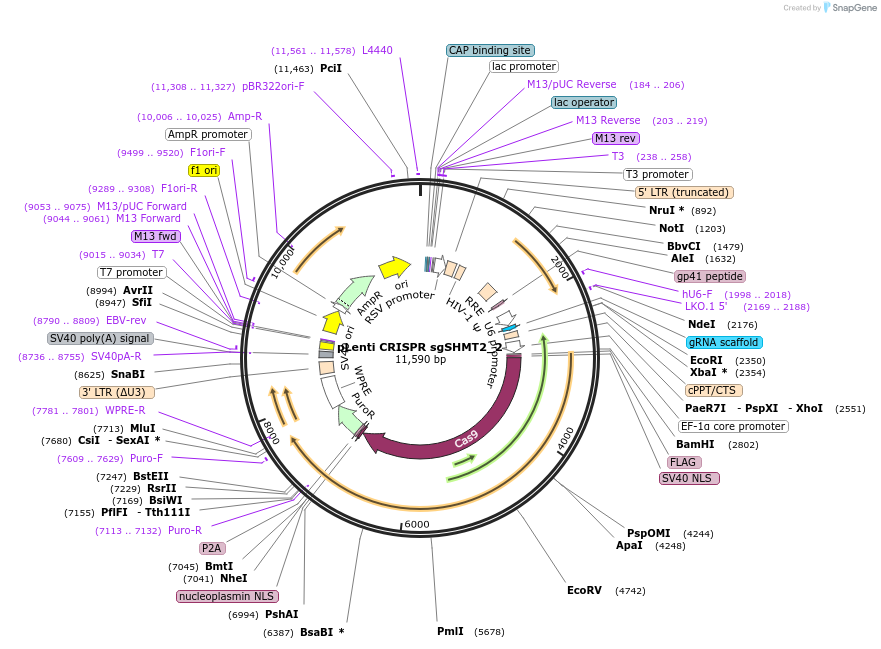
pLenti CRISPR sgSHMT2_2
Plasmid#106310PurposeExpress Cas9 and sgRNA targeting SHMT2DepositorInsertsgRNA targeting SHMT2
UseCRISPR and LentiviralExpressionMammalianAvailable SinceDec. 6, 2018AvailabilityAcademic Institutions and Nonprofits only -
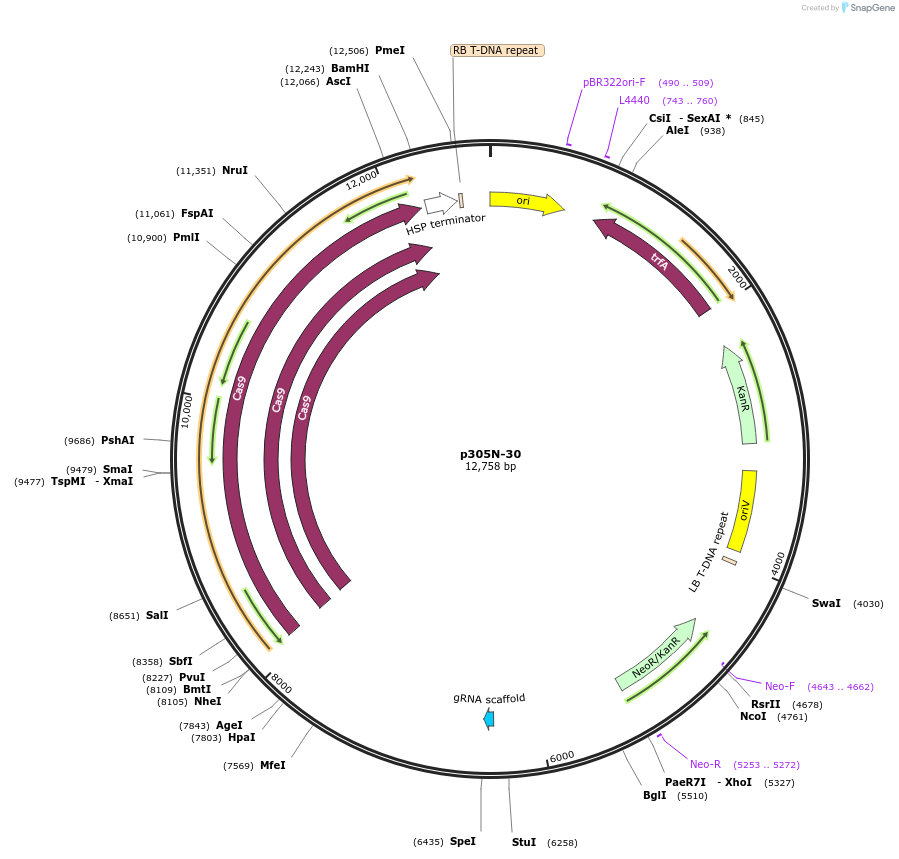
p305N-30
Plasmid#246313PurposeEvaluation of PtU6.2c3 promoter (Pol III promoter) sequence variation for CRISPR-Cas9 editing of mEGFP in a Nicotiana benthamiana reporter lineDepositorInsertPtU6.2c3 promoter
UseCRISPRExpressionPlantMutationdeletion: -T (-29 from TSS) within TATAPromoterPtU6.2c3 promoterAvailable SinceOct. 30, 2025AvailabilityAcademic Institutions and Nonprofits only -
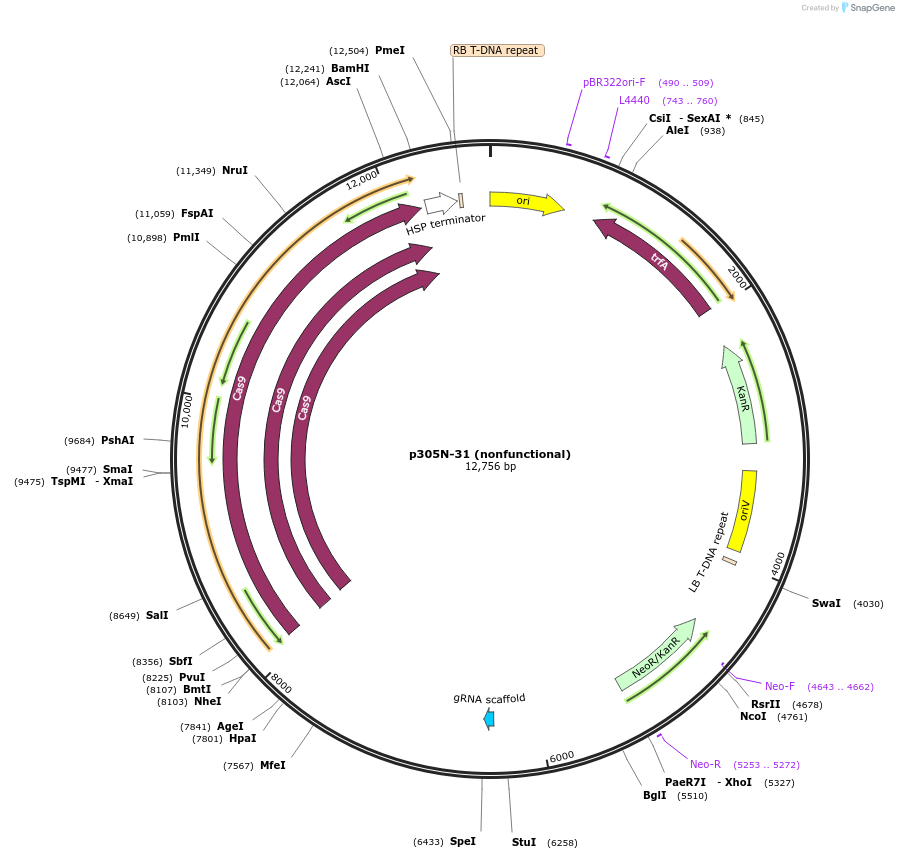
p305N-31 (nonfunctional)
Plasmid#246314PurposeEvaluation of PtU6.2c4 promoter (nonfunctional) (Pol III promoter) sequence variation for CRISPR-Cas9 editing of mEGFP in a Nicotiana benthamiana reporter lineDepositorInsertPtU6.2c4 promoter (nonfunctional)
UseCRISPRExpressionPlantMutationdeletions: -A (-41 from TSS), -T (-30), -T (-29);…PromoterPtU6.2c4 promoter (nonfunctional)Available SinceOct. 30, 2025AvailabilityAcademic Institutions and Nonprofits only -
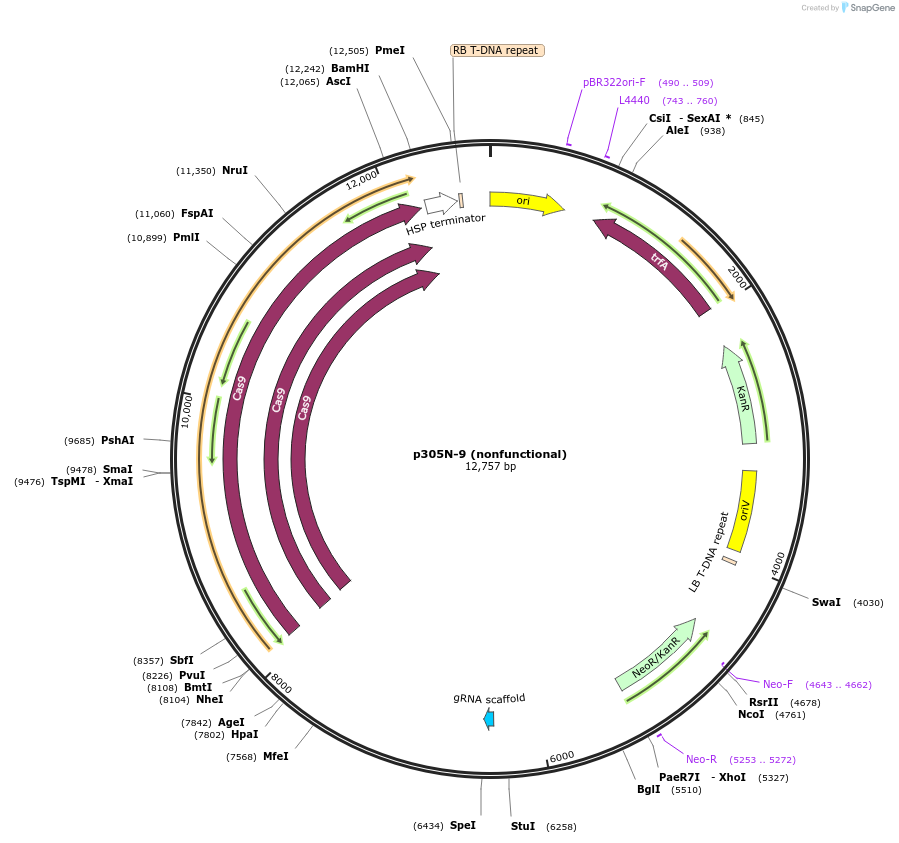
p305N-9 (nonfunctional)
Plasmid#246292PurposeEvaluation of AtU6.29c13 promoter (nonfunctional) (Pol III promoter) sequence variation for CRISPR-Cas9 editing of mEGFP in a Nicotiana benthamiana reporter lineDepositorInsertAtU6.29c13 promoter (nonfunctional)
UseCRISPRExpressionPlantMutationdeletions: -A (-58 from TSS), -T (-57) within USEPromoterAtU6.29c13 promoter (nonfunctional)Available SinceOct. 30, 2025AvailabilityAcademic Institutions and Nonprofits only -
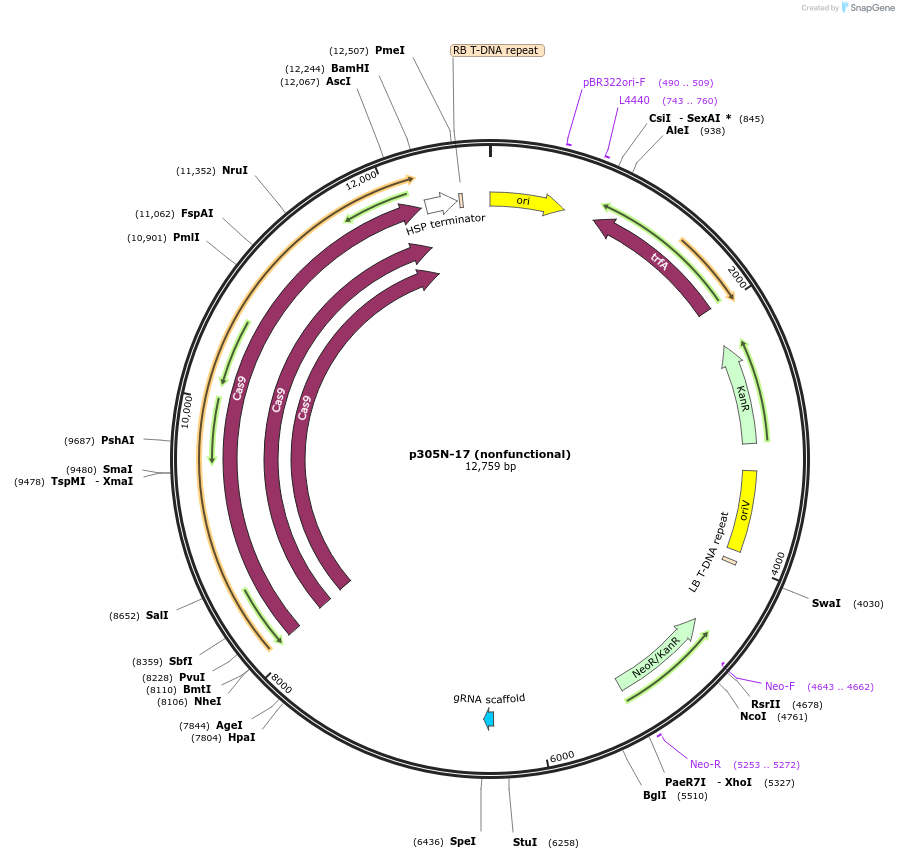
p305N-17 (nonfunctional)
Plasmid#246300PurposeEvaluation of HbU6.2m3 promoter (nonfunctional) (Pol III promoter) sequence variation for CRISPR-Cas9 editing of mEGFP in a Nicotiana benthamiana reporter lineDepositorInsertHbU6.2m3 promoter (nonfunctional)
UseCRISPRExpressionPlantMutationC-to-T (-29 from TSS), G-to-A (-24)PromoterHbU6.2m3 promoter (nonfunctional)Available SinceOct. 30, 2025AvailabilityAcademic Institutions and Nonprofits only -
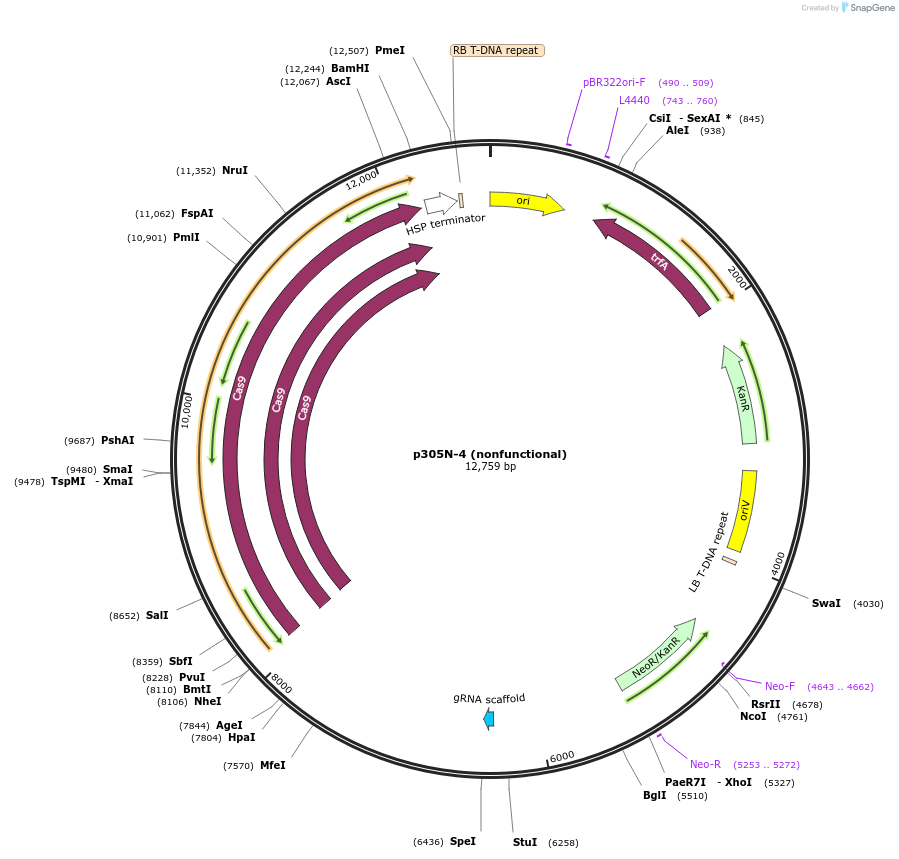
p305N-4 (nonfunctional)
Plasmid#246287PurposeEvaluation of AtU6.1c1 promoter (nonfunctional) (Pol III promoter) sequence variation for CRISPR-Cas9 editing of mEGFP in a Nicotiana benthamiana reporter lineDepositorInsertAtU6.1c1 promoter (nonfunctional)
UseCRISPRExpressionPlantMutationG-to-C at -15 from TSSPromoterAtU6.1c1 promoter (nonfunctional)Available SinceOct. 30, 2025AvailabilityAcademic Institutions and Nonprofits only -
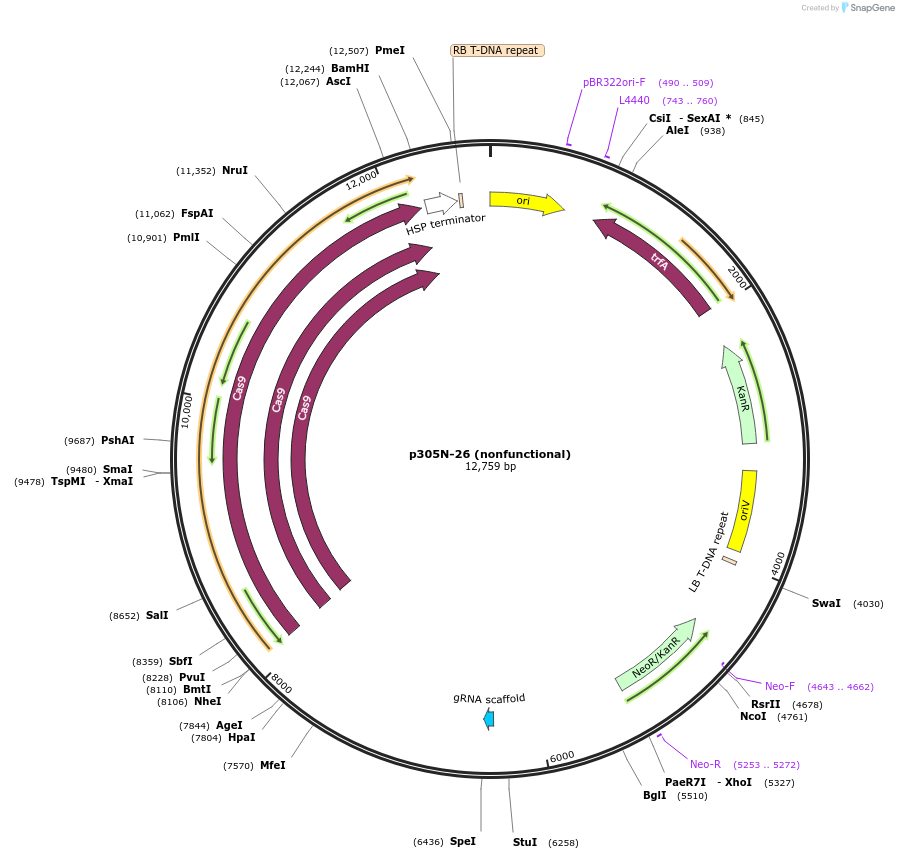
p305N-26 (nonfunctional)
Plasmid#246309PurposeEvaluation of MtU6.6m7 promoter (nonfunctional) (Pol III promoter) sequence variation for CRISPR-Cas9 editing of mEGFP in a Nicotiana benthamiana reporter lineDepositorInsertMtU6.6m7 promoter (nonfunctional)
UseCRISPRExpressionPlantMutationC-to-A (-63 from TSS), T-to-G (-57)PromoterMtU6.6m7 promoter (nonfunctional)Available SinceOct. 30, 2025AvailabilityAcademic Institutions and Nonprofits only -
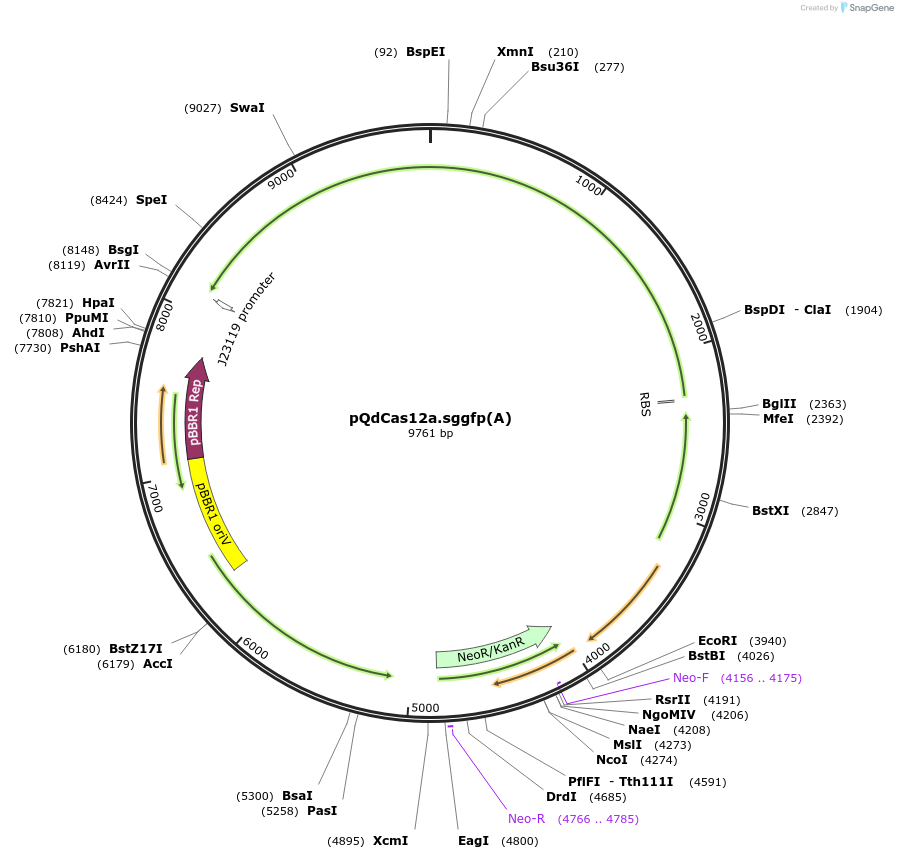
pQdCas12a.sggfp(A)
Plasmid#236038PurposeThe plasmid pQdCas12a.sggfp(A) expresses the dCas12a endonuclease and the sgRNA (design A) targeting the gfp gene.DepositorInsertquorum sensing cassette luxI, luxR, and the LuxR-dependent promoter pLuxI , dCas12a and sgRNA design (A) of gfp gene
UseCRISPRPromoterquorum sensing promoterAvailable SinceApril 24, 2025AvailabilityAcademic Institutions and Nonprofits only -
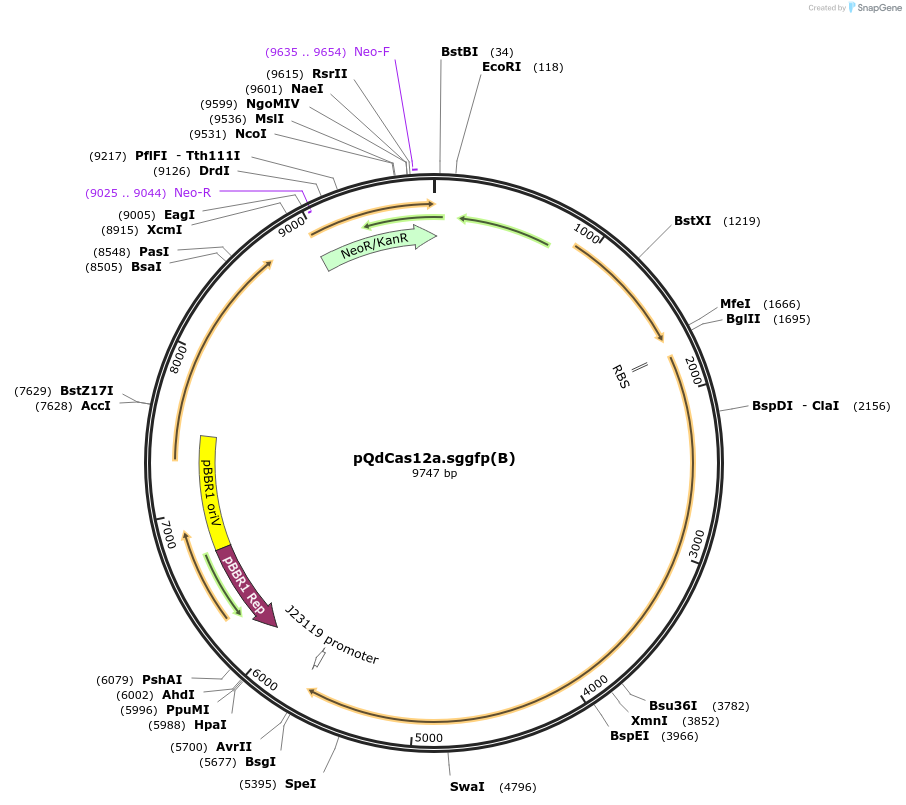
pQdCas12a.sggfp(B)
Plasmid#236040PurposeThe plasmid pQdCas12a.sggfp(B) expresses the dCas12a endonuclease and the sgRNA (design B) targeting the gfp gene.DepositorInsertquorum sensing cassette luxI, mutated luxR, and the LuxR-dependent promoter pLuxI , dCas12a and sgRNA design (B) of gfp gene
UseCRISPRPromoterquorum sensing promoterAvailable SinceApril 24, 2025AvailabilityAcademic Institutions and Nonprofits only -
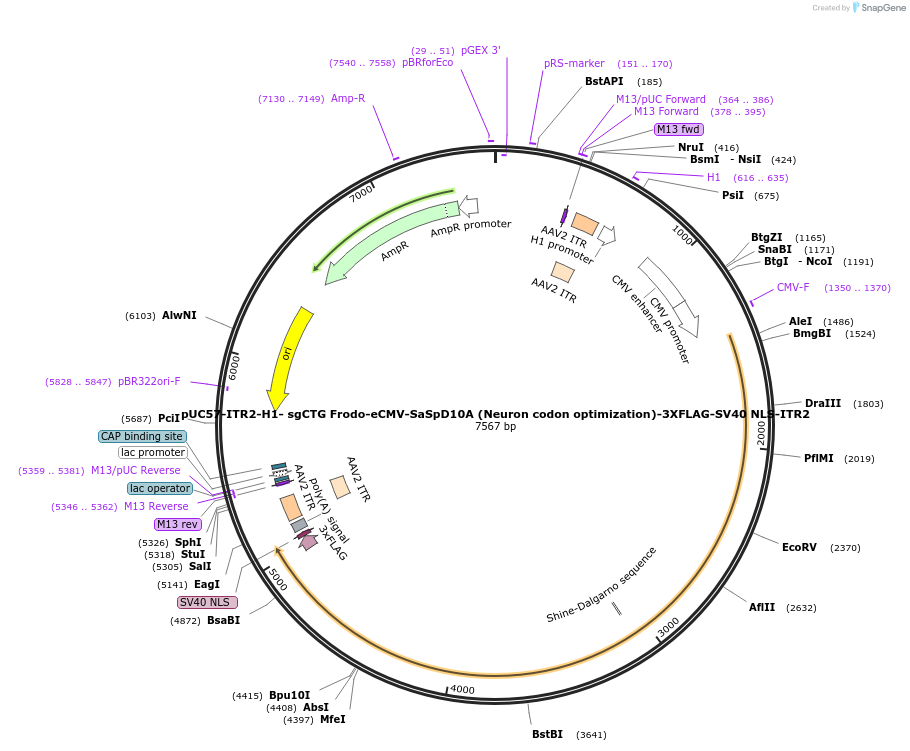
pUC57-ITR2-H1- sgCTG Frodo-eCMV-SaSpD10A (Neuron codon optimization)-3XFLAG-SV40 NLS-ITR2
Plasmid#210734PurposeCoding for SaSp D10A Cas9 alongside Frodo sgRNA targeting CAG repeatsDepositorInsertsSaSp D10A Cas9
Frodo sgCAG
UseCRISPRTags3xFLAG, SV40 NLS, PolyA signalExpressionMammalianMutationD10A, N-terminal Sa Cas9, C-terminal Sp Cas9PromoterH1 and eCMVAvailable SinceMarch 11, 2025AvailabilityAcademic Institutions and Nonprofits only -
-
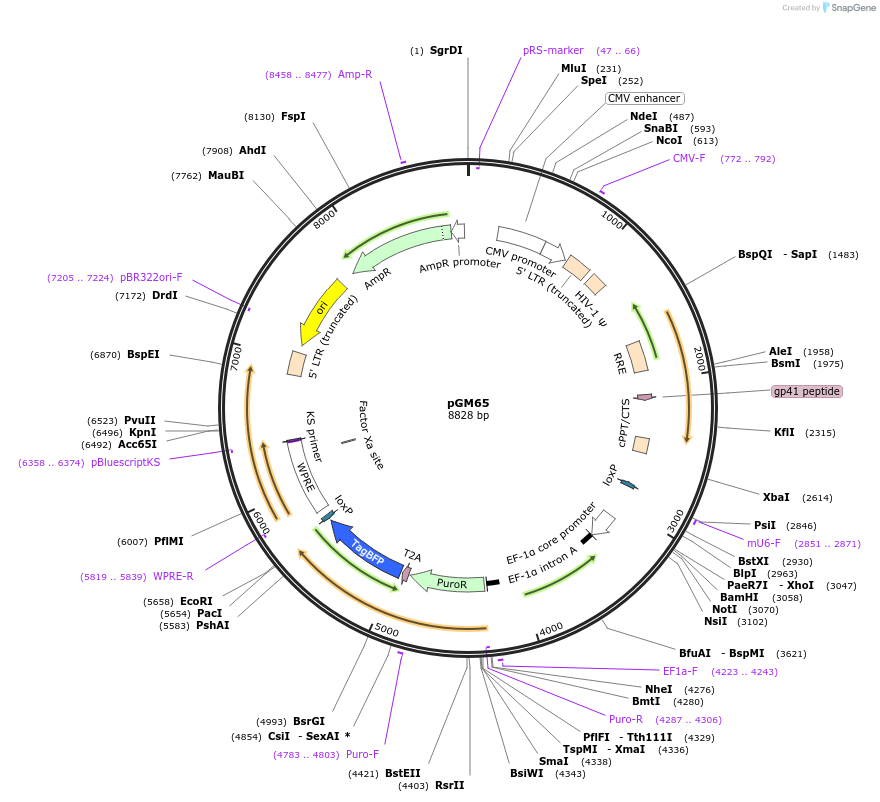
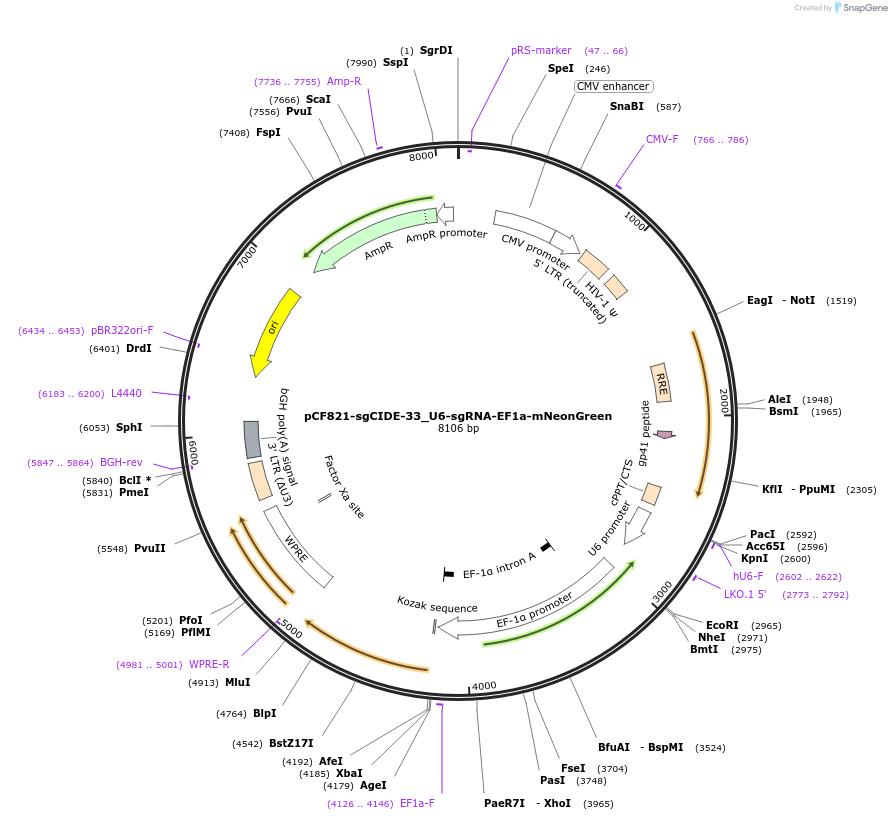
pCF821-sgCIDE-33_U6-sgRNA-EF1a-mNeonGreen
Plasmid#211677PurposeU6-sgRNA-EF1a-mNeonGreenDepositorInsertSpyCas9 and sgCIDE-33 guide RNA vector
UseCRISPR and LentiviralExpressionMammalianAvailable SinceFeb. 1, 2024AvailabilityAcademic Institutions and Nonprofits only -
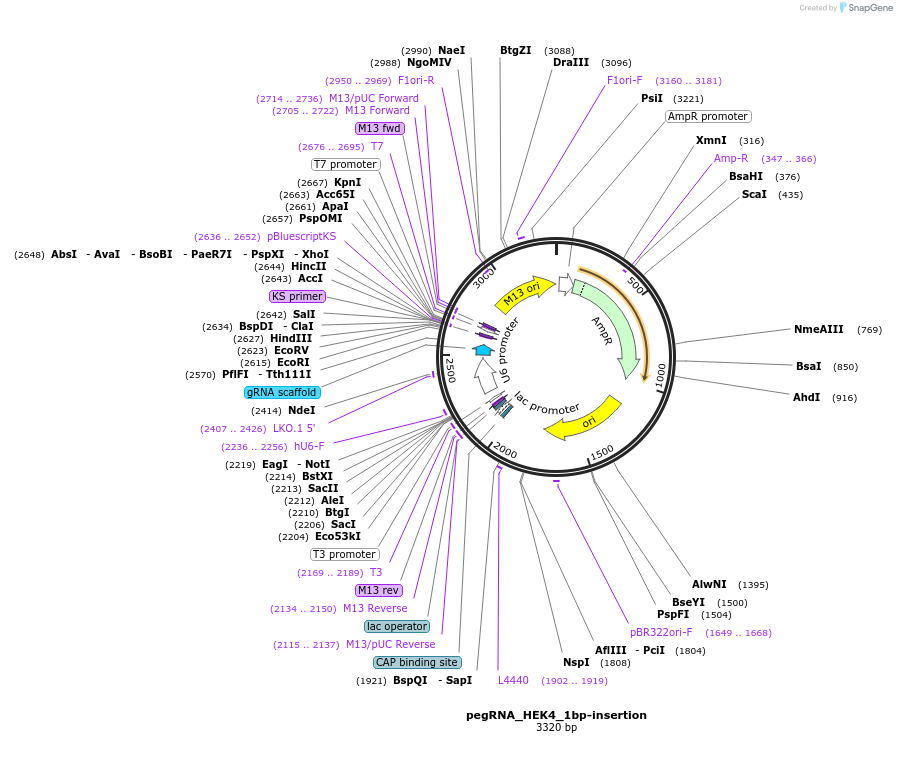
pegRNA_HEK4_1bp-insertion
Plasmid#201977PurposeExpress a pegRNA used for 1bp insertion at HEK4 siteDepositorInsertpegRNA_HEK4_1bp-insertion
ExpressionMammalianAvailable SinceJune 5, 2023AvailabilityAcademic Institutions and Nonprofits only