We narrowed to 13,335 results for: ache
-
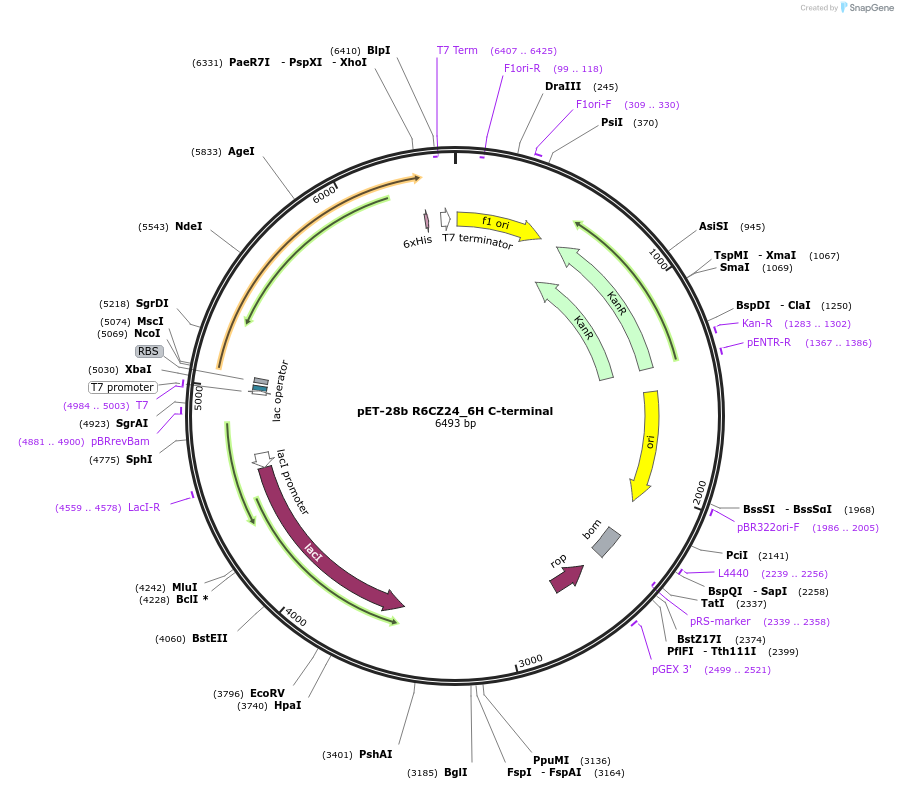
Plasmid#223435PurposeR6CZ24 Thiolase expression vector with C-terminal His6 tagDepositorInsertR6CZ24 Thiolase
TagsHis6Available SinceAug. 29, 2024AvailabilityAcademic Institutions and Nonprofits only -
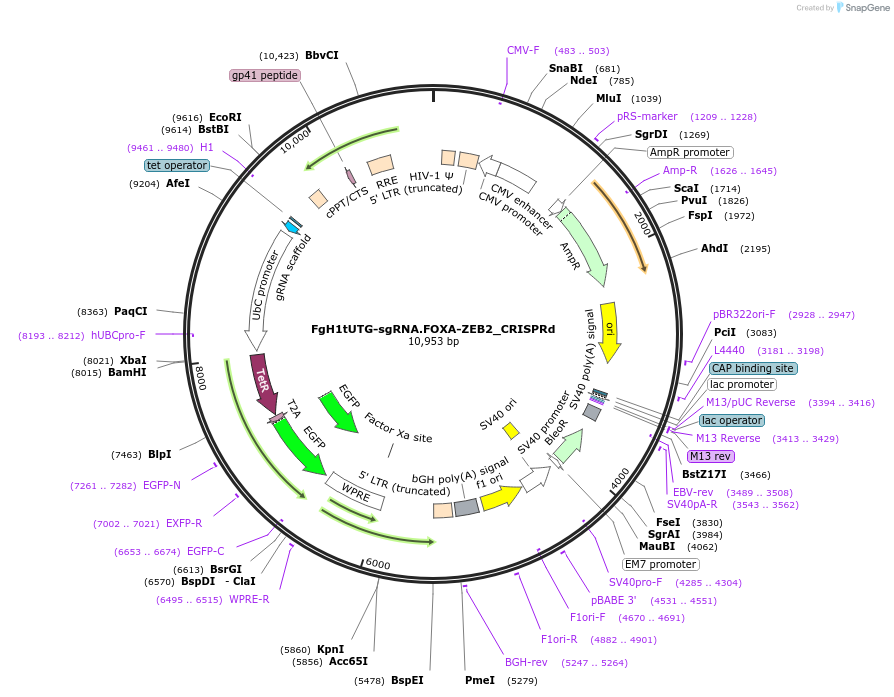
FgH1tUTG-sgRNA.FOXA-ZEB2_CRISPRd
Plasmid#216170PurposeExpress the gRNA targeting the FOXA-binding site in the human ZEB2 locus for the CRISPRd experimentDepositorInsertsgRNA- FOXA.bs- ZEB2.locus
UseLentiviralTagsEGFPAvailable SinceJuly 29, 2024AvailabilityAcademic Institutions and Nonprofits only -
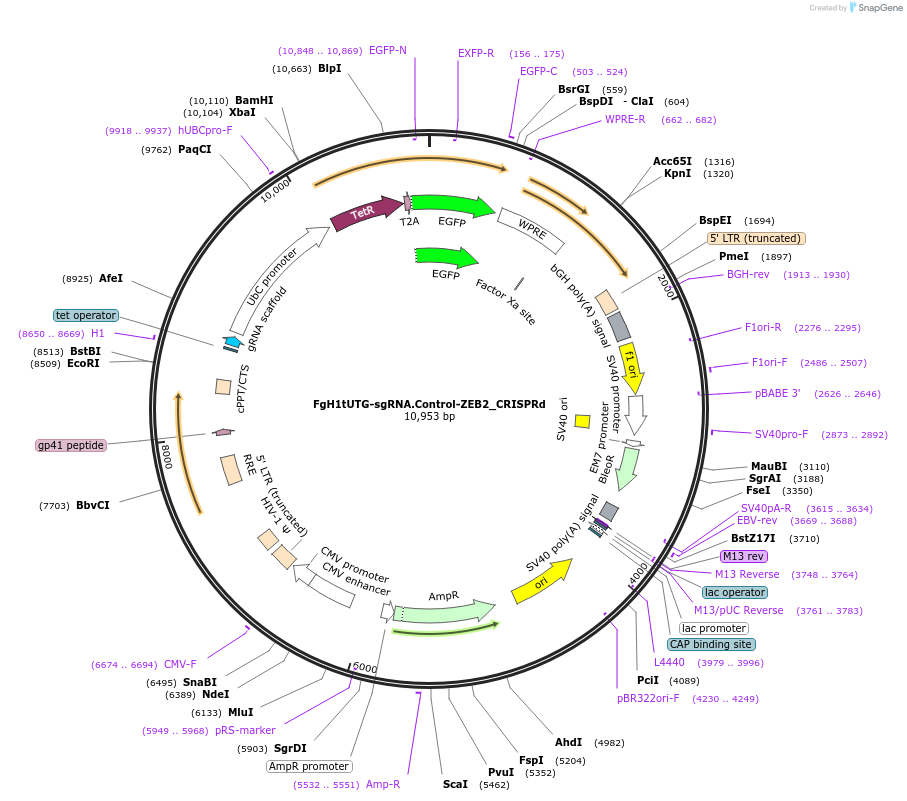
FgH1tUTG-sgRNA.Control-ZEB2_CRISPRd
Plasmid#216172PurposeExpress the gRNA targeting the control site in the human ZEB2 locus for the CRISPRd experimentDepositorInsertsgRNA- Control- ZEB2.locus
UseLentiviralTagsEGFPAvailable SinceJuly 29, 2024AvailabilityAcademic Institutions and Nonprofits only -
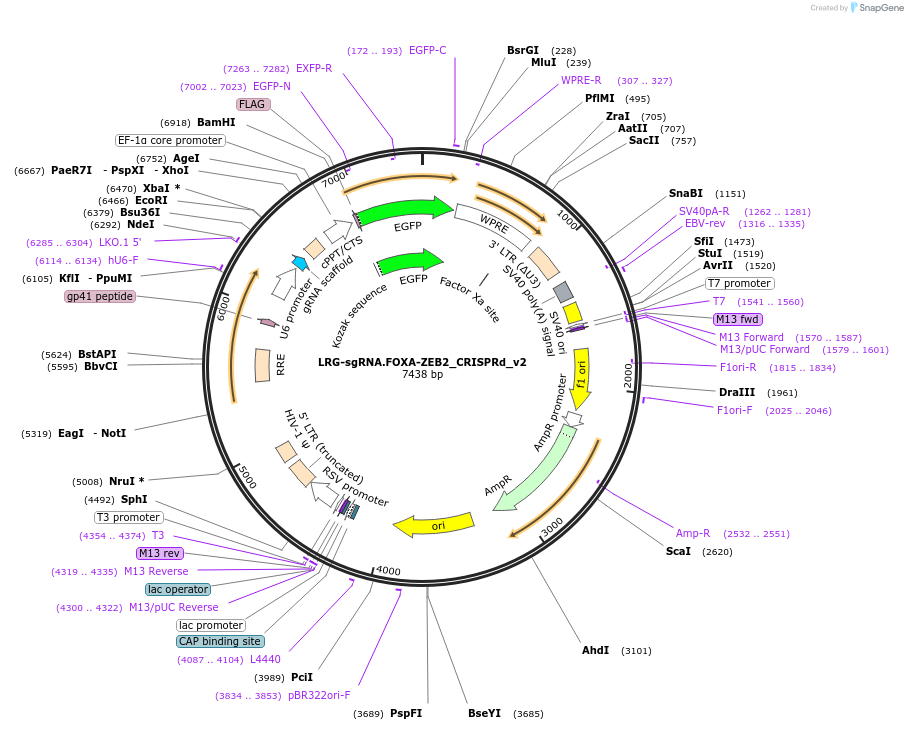
LRG-sgRNA.FOXA-ZEB2_CRISPRd_v2
Plasmid#216173PurposeExpress the gRNA targeting the FOXA-binding site in the human ZEB2 locus for the CRISPRd experimentDepositorInsertsgRNA- FOXA.bs- ZEB2.locus
UseLentiviralTagsEGFPAvailable SinceJuly 29, 2024AvailabilityAcademic Institutions and Nonprofits only -
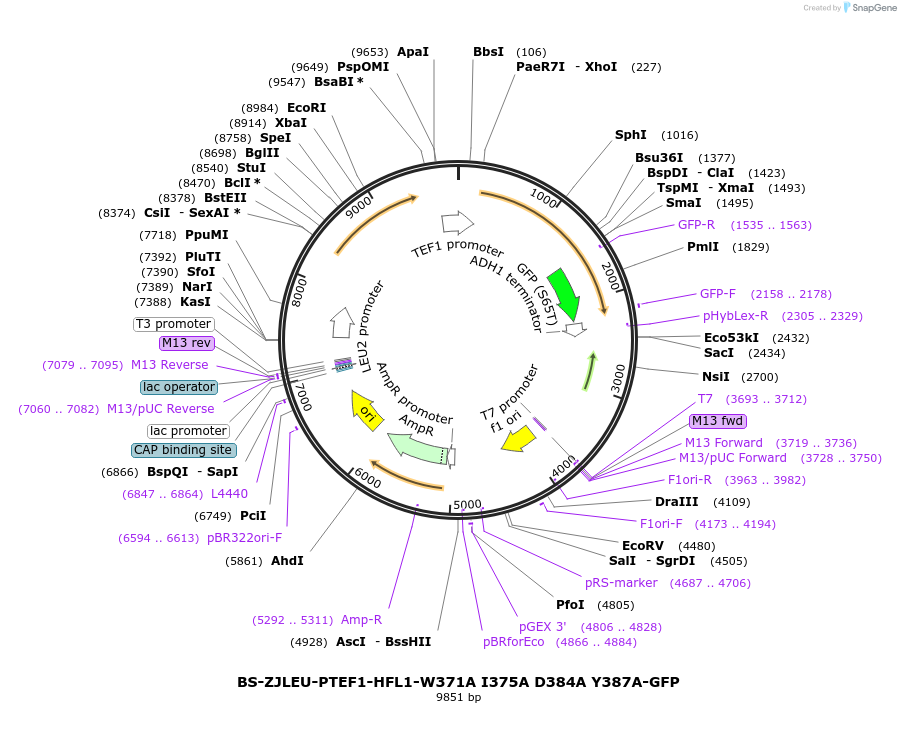
BS-ZJLEU-PTEF1-HFL1-W371A I375A D384A Y387A-GFP
Plasmid#207025PurposeExpression of Hfl1 point mutant. Uses auxotrophic marker LEU2 (Saccharomyces cerevisiae).DepositorInsertHFL1
TagsGFPExpressionYeastMutationW371A I375A D384A Y387APromoterpTEF1Available SinceJune 4, 2024AvailabilityAcademic Institutions and Nonprofits only -
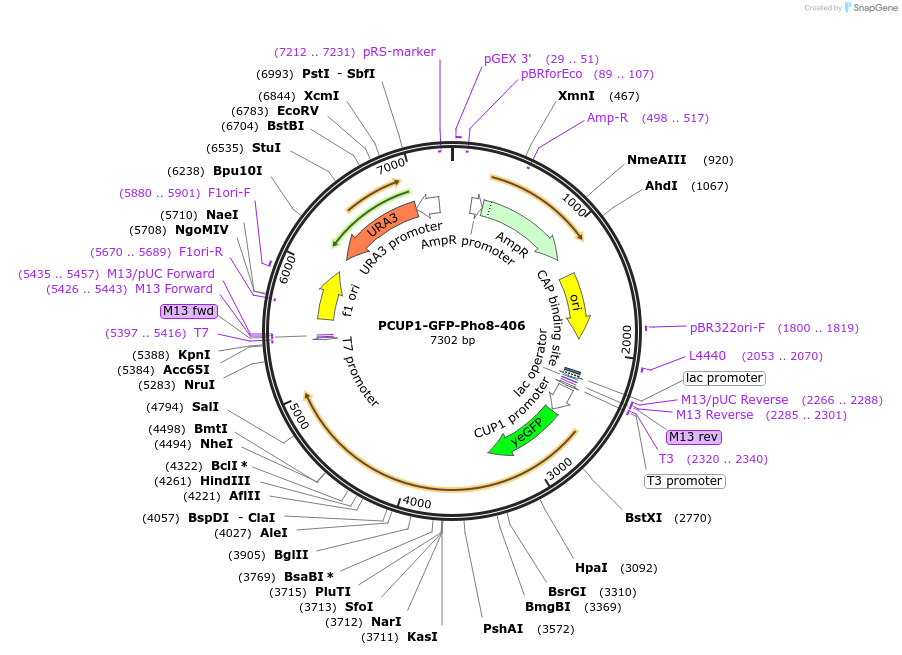
PCUP1-GFP-Pho8-406
Plasmid#207006PurposeIntegration of GFP-Pho8 as an additional copy in genome, expressed under CUP1 promoter. Uses auxotrophic marker URA3 (Saccharomyces cerevisiae) for selection. Marker for yeast vacuolar membrane.DepositorInsertPHO8
TagsGFPExpressionYeastAvailable SinceJune 3, 2024AvailabilityAcademic Institutions and Nonprofits only -
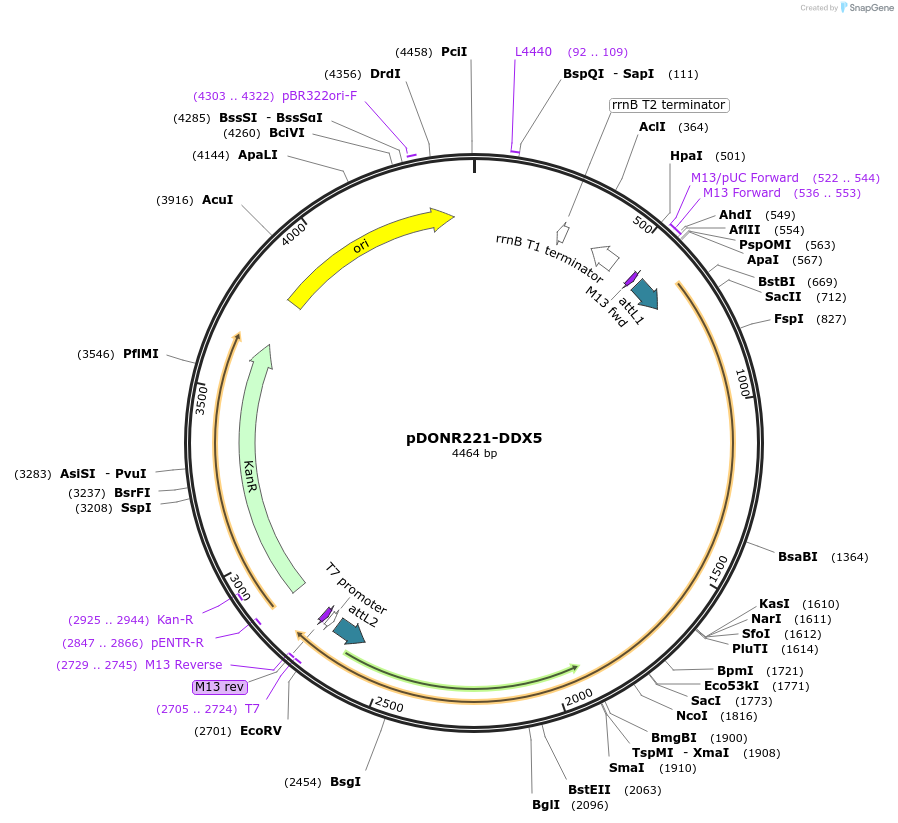
pDONR221-DDX5
Plasmid#166163PurposeGateway entry vector for N. furzeriDDX5DepositorInsertDDX5
UseGateway entry vectorAvailable SinceMay 20, 2024AvailabilityAcademic Institutions and Nonprofits only -
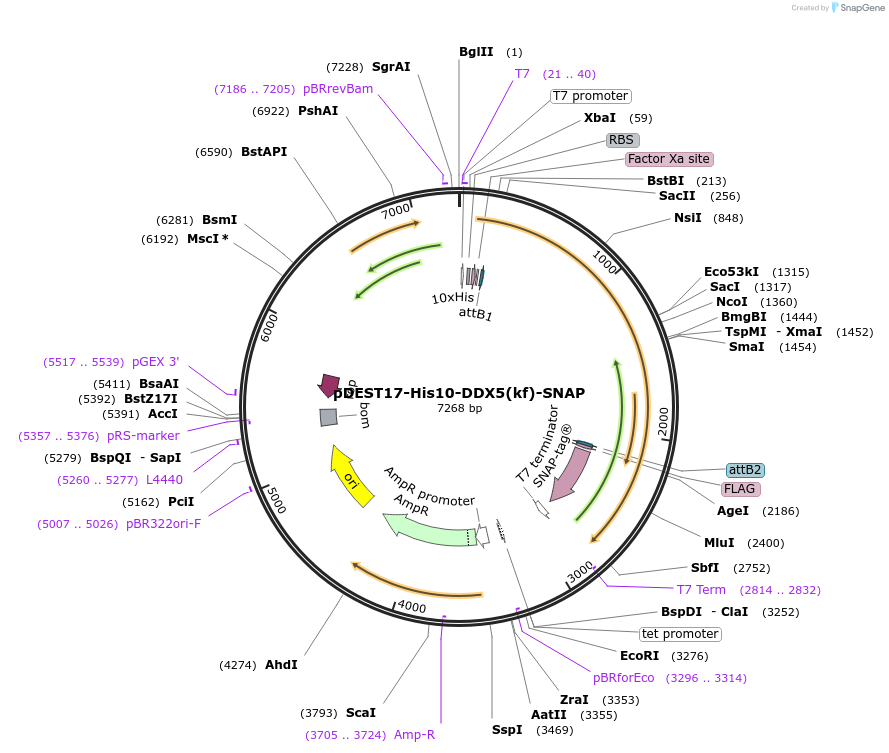
pDEST17-His10-DDX5(kf)-SNAP
Plasmid#166152PurposeExpression of N. furzeri DDX5 in E. coliDepositorInsertDDX5
TagsHis10 and SNAPExpressionBacterialPromoterT7Available SinceMay 20, 2024AvailabilityAcademic Institutions and Nonprofits only -
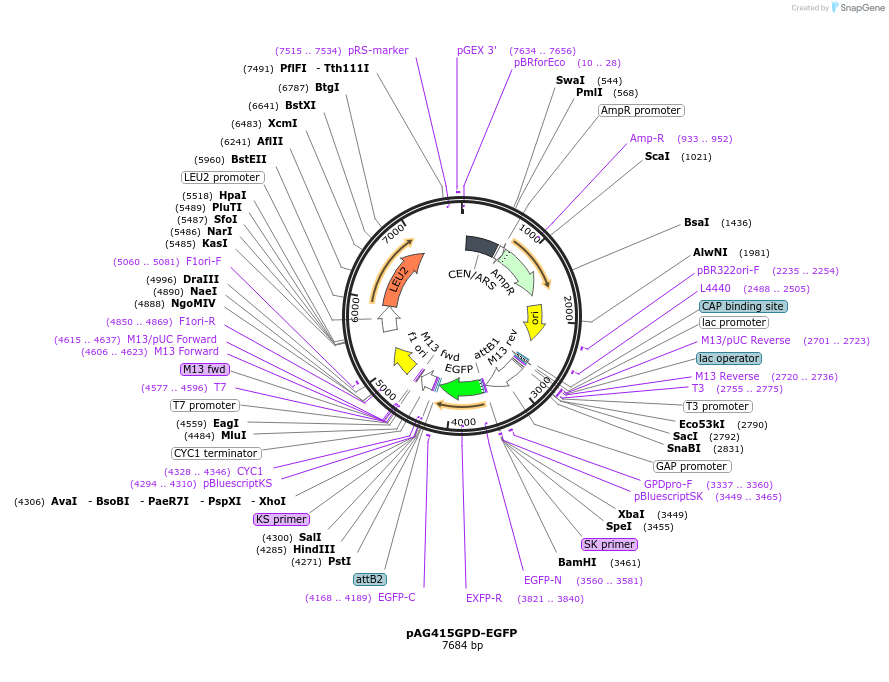
pAG415GPD-EGFP
Plasmid#166137PurposeEGFP expression under the control of the GPD promoter yeastDepositorInsertEGFP
ExpressionYeastPromoterGDPAvailable SinceMay 20, 2024AvailabilityAcademic Institutions and Nonprofits only -
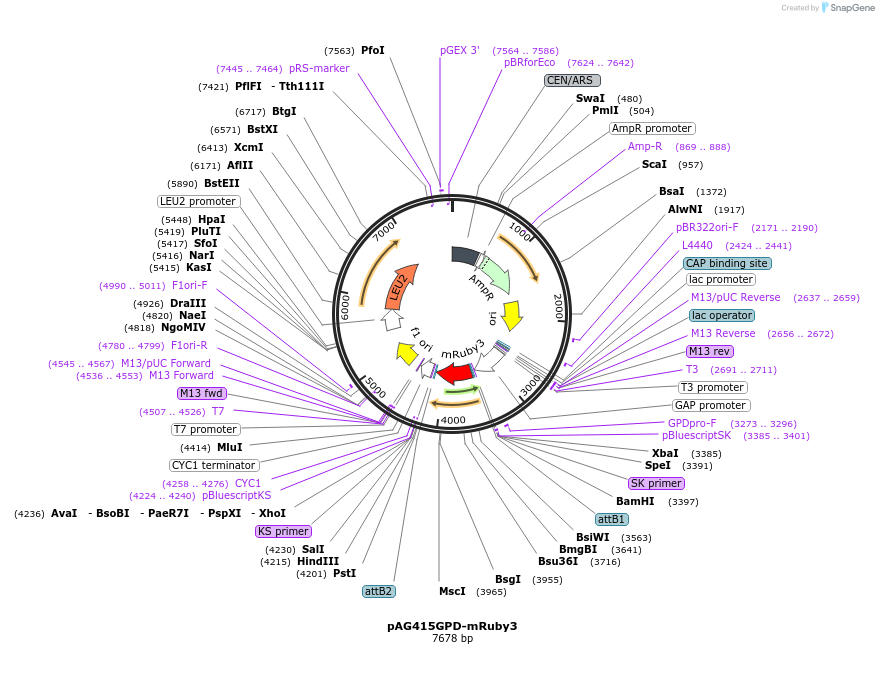
pAG415GPD-mRuby3
Plasmid#166138PurposemRuby3 expression under control of the GPD promoter in yeastDepositorInsertmRuby3
ExpressionYeastPromoterGDPAvailable SinceMay 20, 2024AvailabilityAcademic Institutions and Nonprofits only -
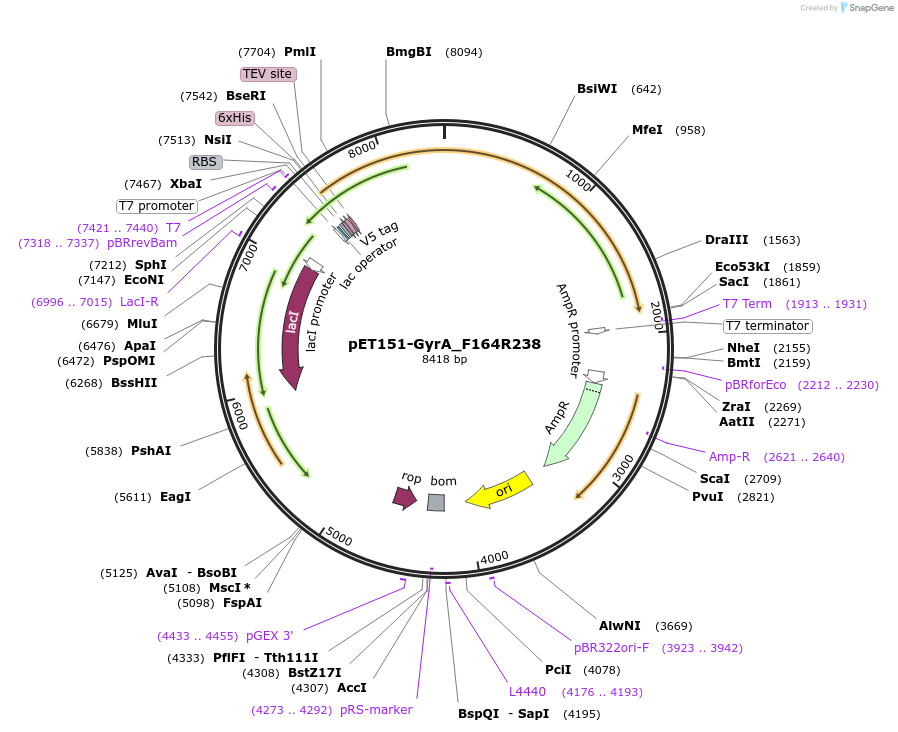
pET151-GyrA_F164R238
Plasmid#216230PurposeS. aureus USA300 GyrA mutant F164 R238 residues codon-optimized for E. coliDepositorInsertDNA gyrase
Tags6xHisExpressionBacterialMutationChanged arginine 238 to serine, phenylalanine 168…PromoterT7Available SinceApril 19, 2024AvailabilityAcademic Institutions and Nonprofits only -
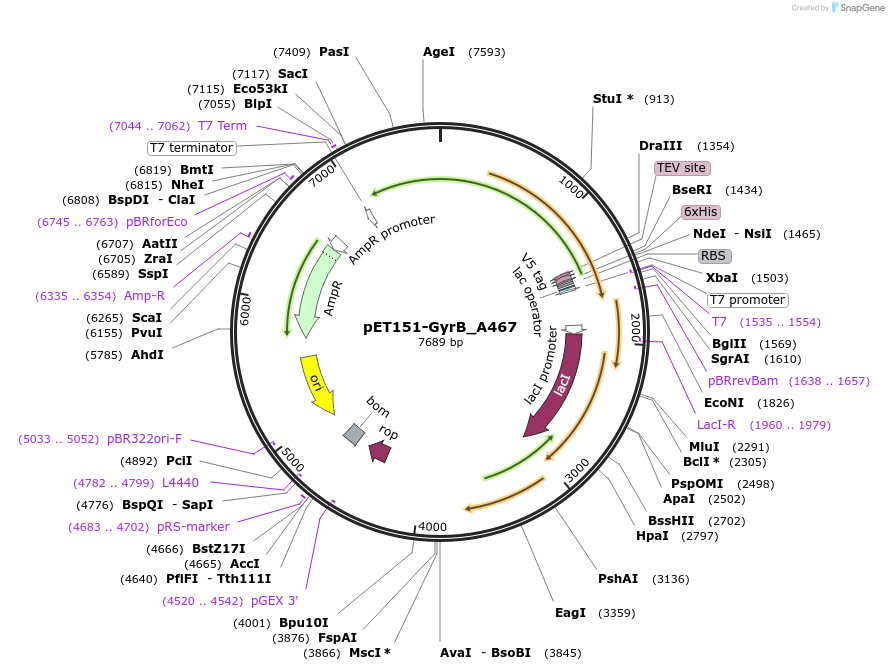
pET151-GyrB_A467
Plasmid#216234PurposeS. aureus USA300 GyrB mutant A467 residue codon-optimized for E. coliDepositorInsertDNA gyrase
Tags6xHisExpressionBacterialMutationChanged alanine 467 to valinePromoterT7Available SinceApril 19, 2024AvailabilityAcademic Institutions and Nonprofits only -
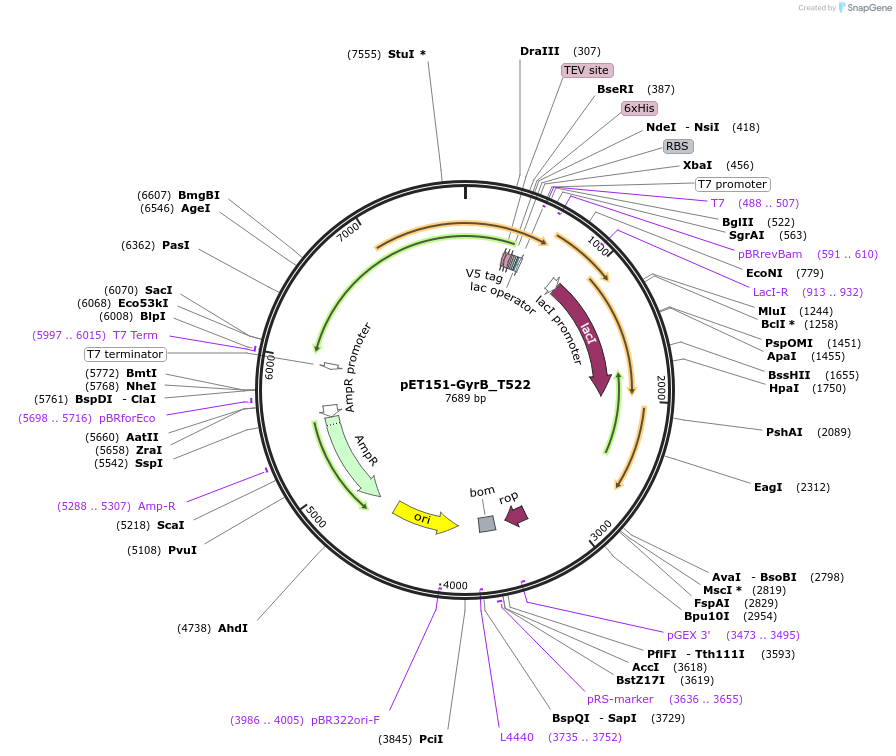
pET151-GyrB_T522
Plasmid#216235PurposeS. aureus USA300 GyrB mutant T522 residue codon-optimized for E. coliDepositorInsertDNA gyrase
Tags6xHisExpressionBacterialMutationChanged threonine 522 to isoleucinePromoterT7Available SinceApril 19, 2024AvailabilityAcademic Institutions and Nonprofits only -
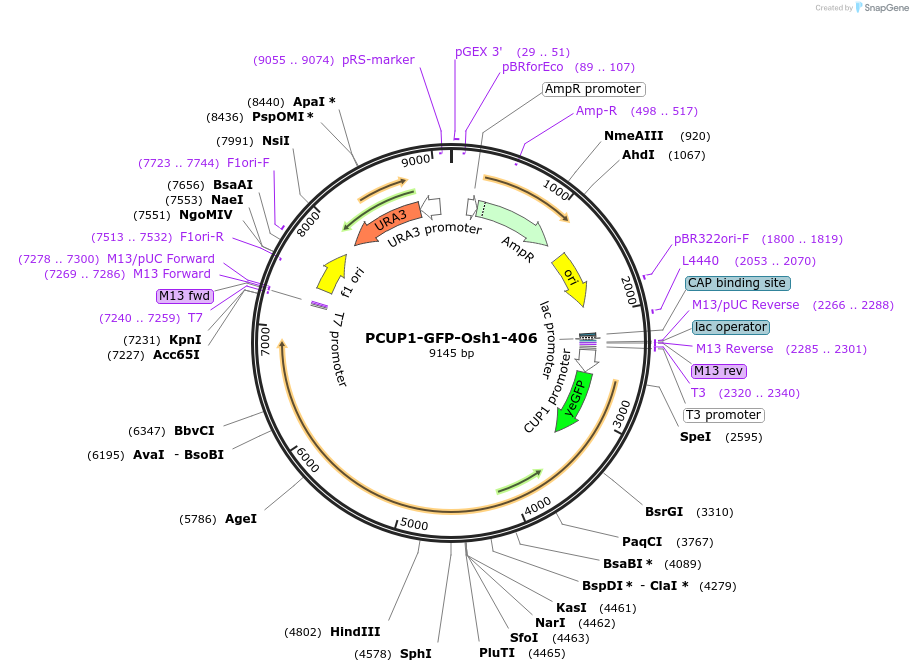
PCUP1-GFP-Osh1-406
Plasmid#207007PurposeIntegration of Osh1-GFP as an additional copy in genome, expressed under CUP1 promoter. Uses auxotrophic marker URA3 (Saccharomyces cerevisiae) for selection. Yeast PMN pathway substrate.DepositorInsertOSH1
TagsGFPExpressionYeastAvailable SinceApril 19, 2024AvailabilityAcademic Institutions and Nonprofits only -
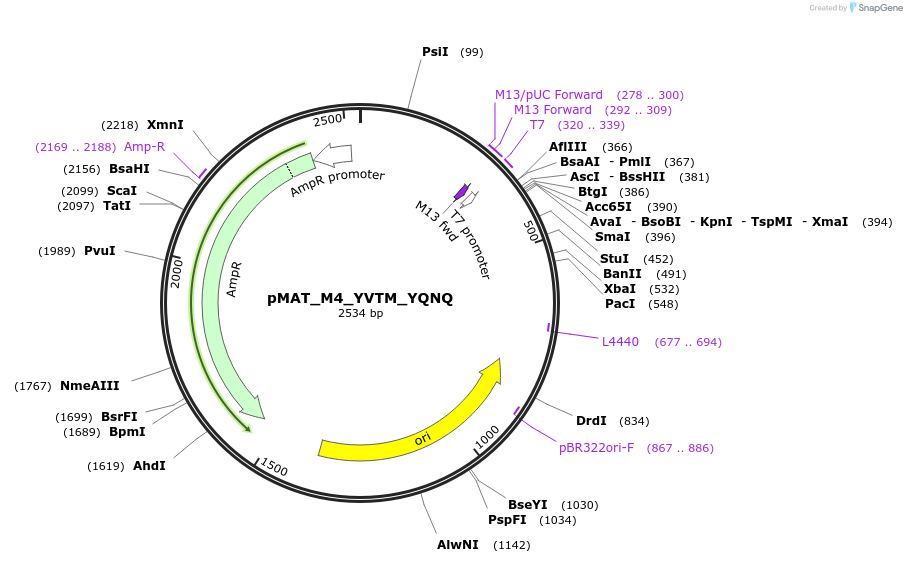
pMAT_M4_YVTM_YQNQ
Plasmid#197084PurposeThis plasmid contains the coding sequence for signaling peptide M4, which contains the YVTM and YQNQ signaling motifs. Digest this insert and add it to pHR_pSFFV_scFVCD19_CD8a_CD3zP2AEGFPDepositorInsertThis plasmid contains the coding sequence for signaling peptide M4, which contains the YVTM and YQNQ signaling motifs.
ExpressionMammalianAvailable SinceApril 3, 2024AvailabilityAcademic Institutions and Nonprofits only -
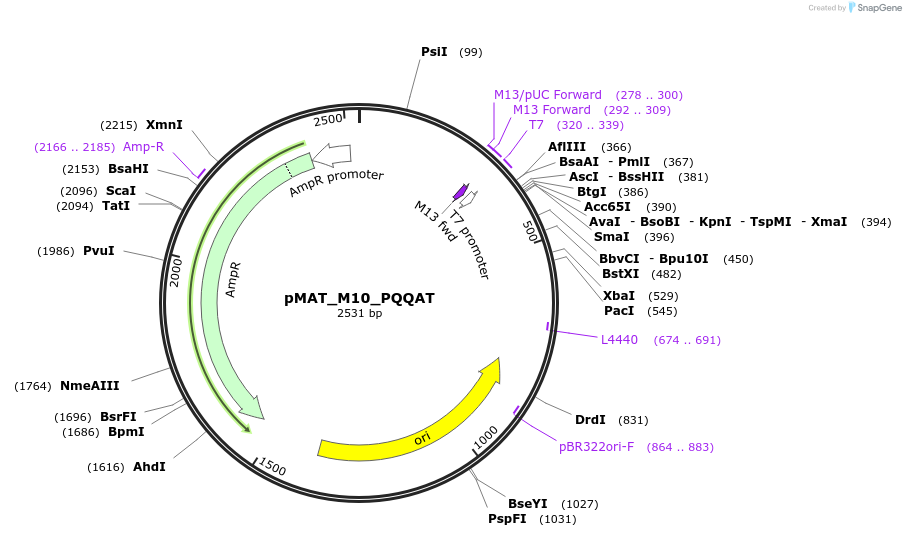
pMAT_M10_PQQAT
Plasmid#197093PurposeThis plasmid contains the coding sequence for signaling peptide M10, which contains the PQQAT signaling motif. Digest this insert and add it to pHR_pSFFV_scFVCD19_CD8a_CD3zP2AEGFPDepositorInsertThis plasmid contains the coding sequence for signaling peptide M10, which contains the PQQAT signaling motif.
ExpressionMammalianAvailable SinceApril 3, 2024AvailabilityAcademic Institutions and Nonprofits only -
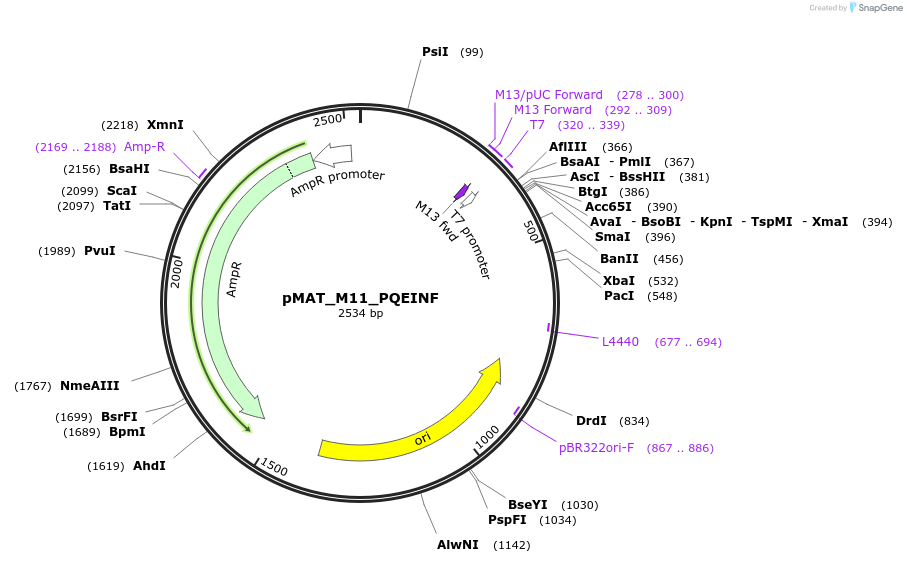
pMAT_M11_PQEINF
Plasmid#197094PurposeThis plasmid contains the coding sequence for signaling peptide M11, which contains the PQEINF signaling motif. Digest this insert and add it to pHR_pSFFV_scFVCD19_CD8a_CD3zP2AEGFPDepositorInsertThis plasmid contains the coding sequence for signaling peptide M11, which contains the PQEINF signaling motif.
ExpressionMammalianAvailable SinceApril 3, 2024AvailabilityAcademic Institutions and Nonprofits only -
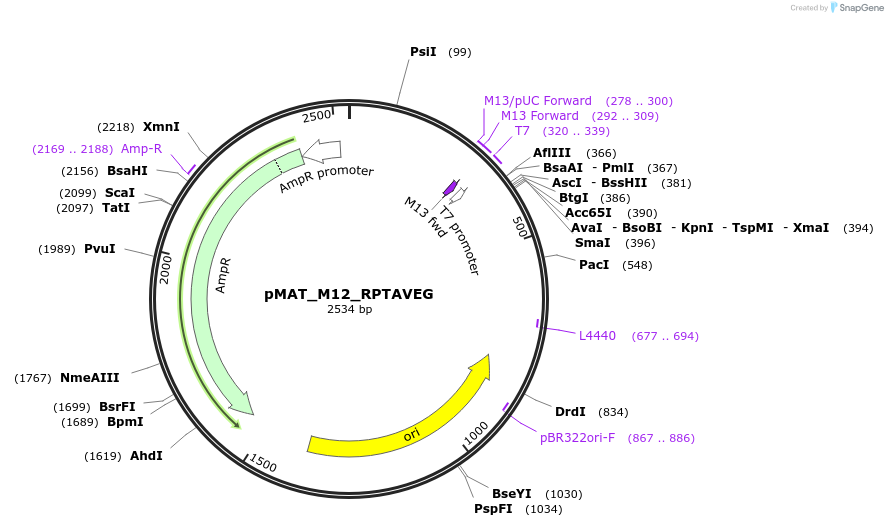
pMAT_M12_RPTAVEG
Plasmid#197095PurposeThis plasmid contains the coding sequence for signaling peptide M12, which contains the RPTAVEG signaling motif. Digest this insert and add it to pHR_pSFFV_scFVCD19_CD8a_CD3zP2AEGFPDepositorInsertThis plasmid contains the coding sequence for signaling peptide M12, which contains the RPTAVEG signaling motif.
ExpressionMammalianAvailable SinceApril 3, 2024AvailabilityAcademic Institutions and Nonprofits only -
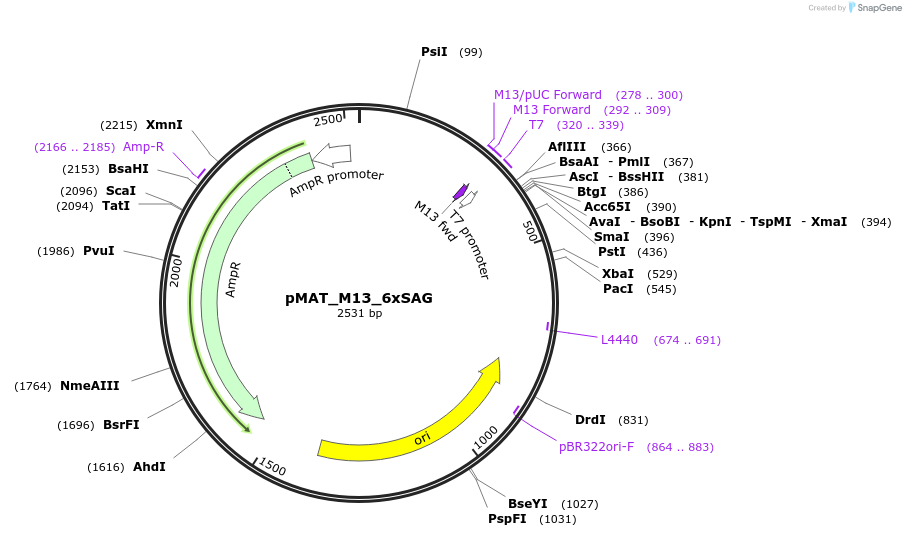
pMAT_M13_6xSAG
Plasmid#197096PurposeThis plasmid contains the coding sequence for the control peptide M13, which contains a linker with six repeats of serine-alanine-glycine. Digest this insert and add it to pHR_pSFFV_scFVCD19_CD8a_CDepositorInsertControl peptide M13, which contains a linker with six repeats of serine-alanine-glycine (SAG)
ExpressionMammalianAvailable SinceApril 3, 2024AvailabilityAcademic Institutions and Nonprofits only -
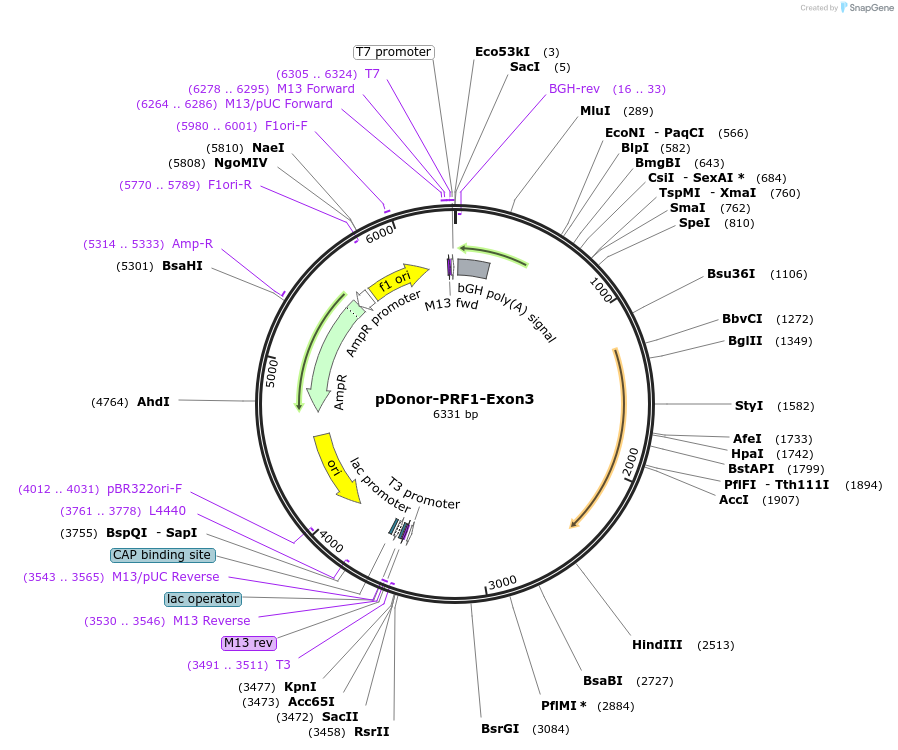
pDonor-PRF1-Exon3
Plasmid#209075PurposeTargeting vector for the human PRF1 locus to replace exon 3 with repaired exon 3DepositorInsertPRF1-Exon3 HDRT (PRF1 Human)
UseCRISPRAvailable SinceMarch 15, 2024AvailabilityAcademic Institutions and Nonprofits only -
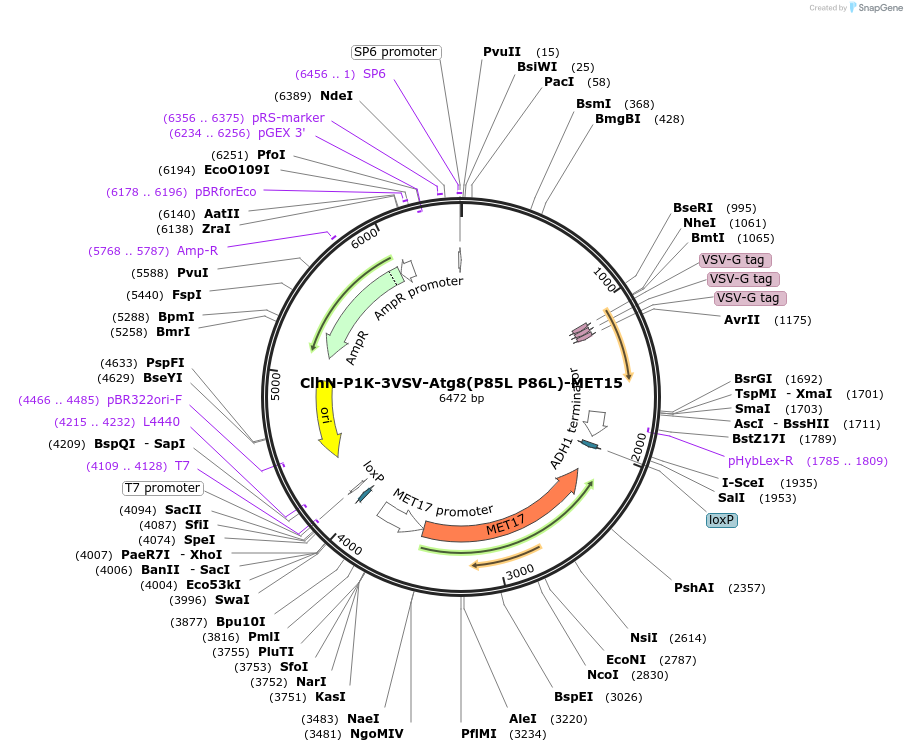
ClhN-P1K-3VSV-Atg8(P85L P86L)-MET15
Plasmid#207050PurposeExpression of 3VSV-Atg8 point mutant. Uses auxotrophic marker MET15(Saccharomyces cerevisiae).DepositorInsertATG8
Tags3xVSVExpressionYeastMutationP85L P86LAvailable SinceMarch 4, 2024AvailabilityAcademic Institutions and Nonprofits only -
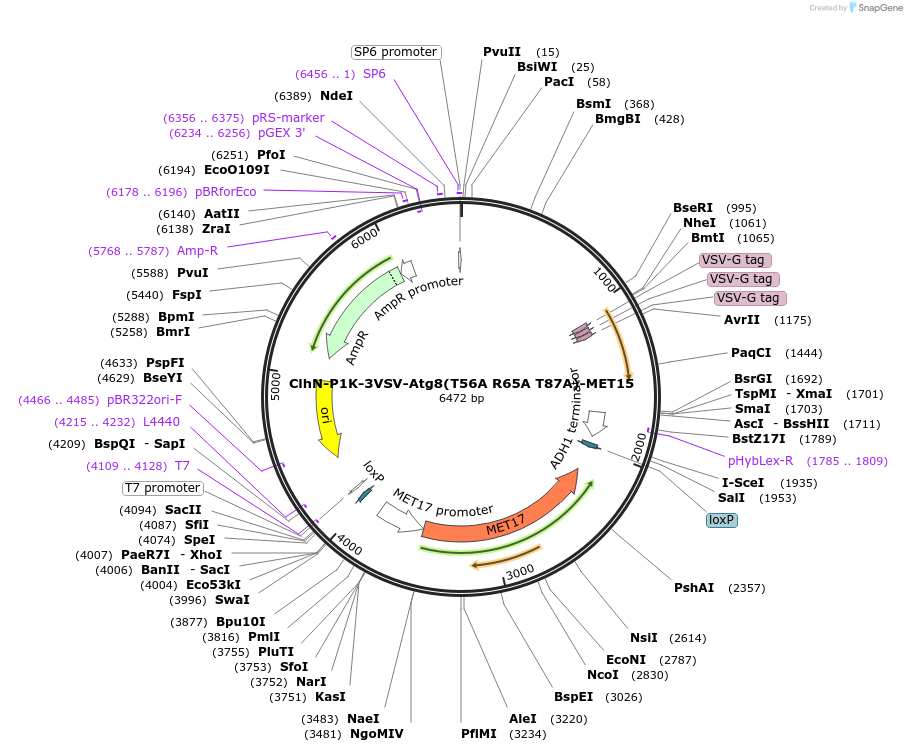
ClhN-P1K-3VSV-Atg8(T56A R65A T87A)-MET15
Plasmid#207051PurposeExpression of 3VSV-Atg8 point mutant. Uses auxotrophic marker MET15(Saccharomyces cerevisiae).DepositorInsertATG8
Tags3xVSVExpressionYeastMutationT56A R65A T87AAvailable SinceMarch 4, 2024AvailabilityAcademic Institutions and Nonprofits only -
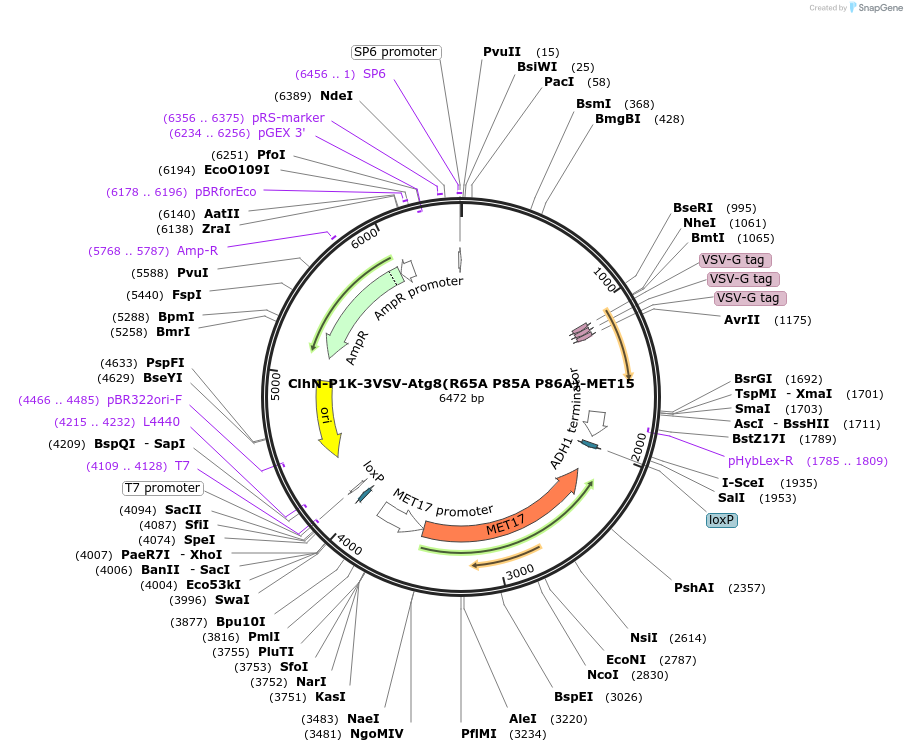
ClhN-P1K-3VSV-Atg8(R65A P85A P86A)-MET15
Plasmid#207052PurposeExpression of 3VSV-Atg8 point mutant. Uses auxotrophic marker MET15(Saccharomyces cerevisiae).DepositorInsertATG8
Tags3xVSVExpressionYeastMutationR65A P85A P86AAvailable SinceMarch 4, 2024AvailabilityAcademic Institutions and Nonprofits only -
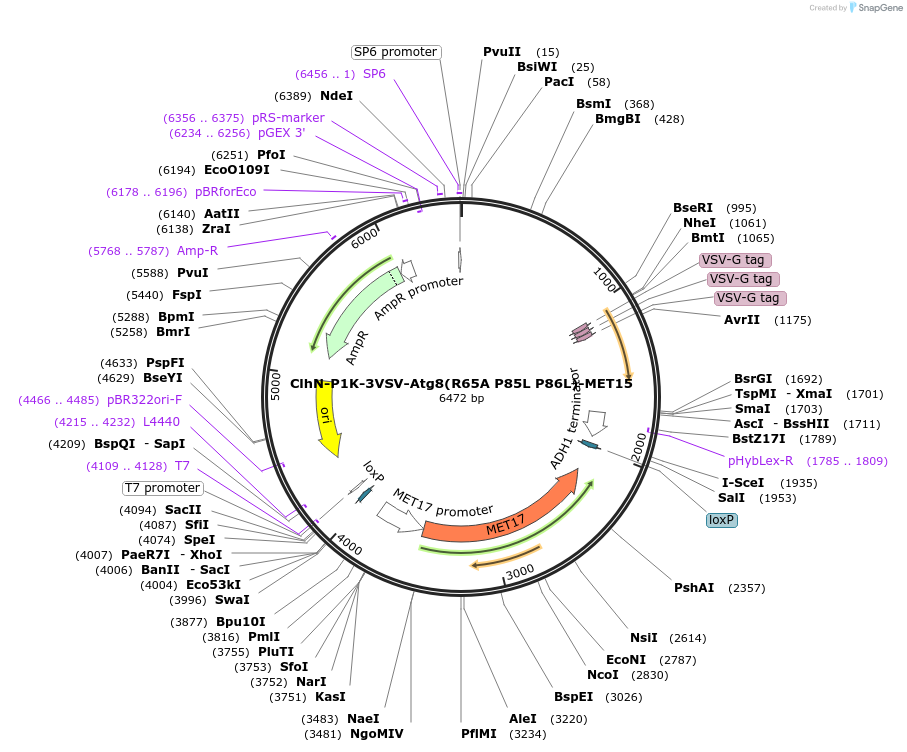
ClhN-P1K-3VSV-Atg8(R65A P85L P86L)-MET15
Plasmid#207053PurposeExpression of 3VSV-Atg8 point mutant. Uses auxotrophic marker MET15(Saccharomyces cerevisiae).DepositorInsertATG8
Tags3xVSVExpressionYeastMutationR65A P85L P86LAvailable SinceMarch 4, 2024AvailabilityAcademic Institutions and Nonprofits only -
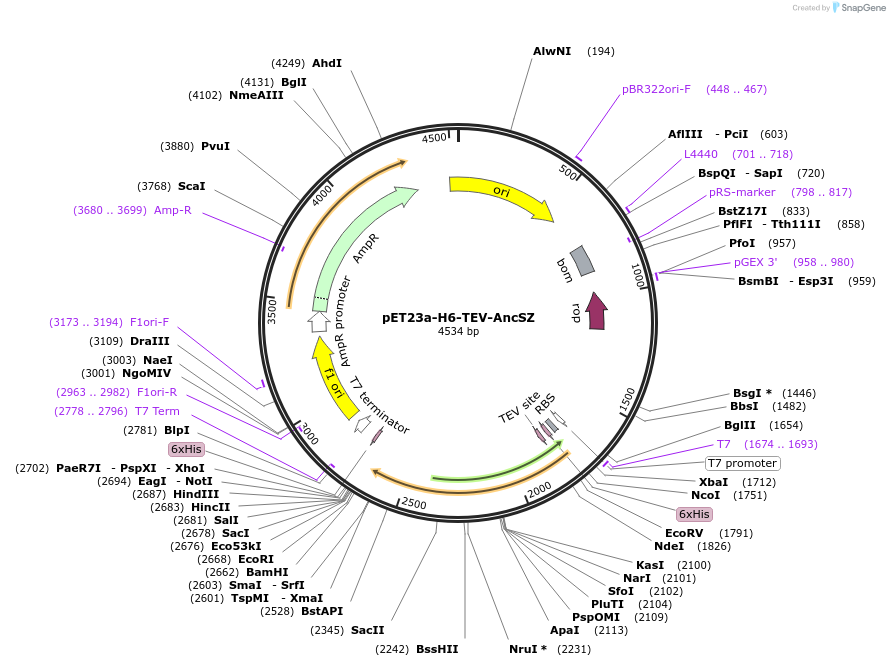
pET23a-H6-TEV-AncSZ
Plasmid#214235PurposeBacterial expression construct of AncSZ, an kinase domain engineered through ancestorial reconstruction of the kinase domains of both SYK and ZAP-70. For in vitro protein tyrosine phosphorylationDepositorInsertSyk kinase (SYK Human)
Tags6xHisExpressionBacterialMutationSequence gained through ancestral reconstruction …PromoterT7Available SinceFeb. 15, 2024AvailabilityAcademic Institutions and Nonprofits only -
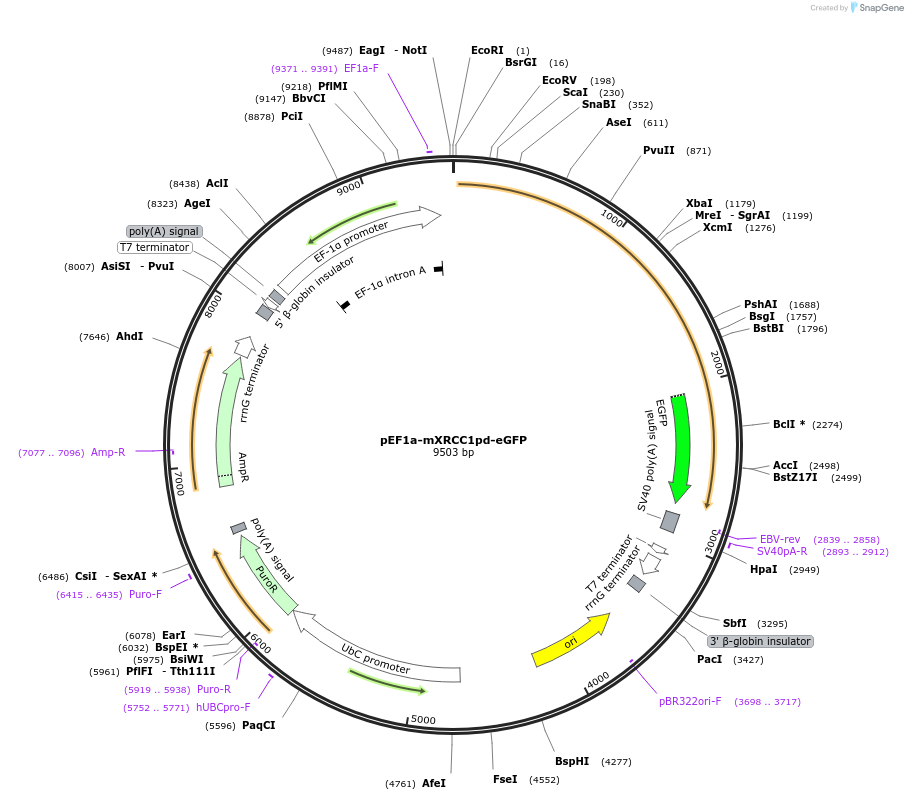
pEF1a-mXRCC1pd-eGFP
Plasmid#206035PurposeMammalian expression of PAR-deficient mXRCC1pd coupled to eGFP under the control of a EF1a promoterDepositorInsertmXRCC1pd
TagseGFPExpressionMammalianMutationS105A, S186A, K188A, S195A, S221A, S222A,S238A, S…PromoterEf1aAvailable SinceJan. 3, 2024AvailabilityAcademic Institutions and Nonprofits only -
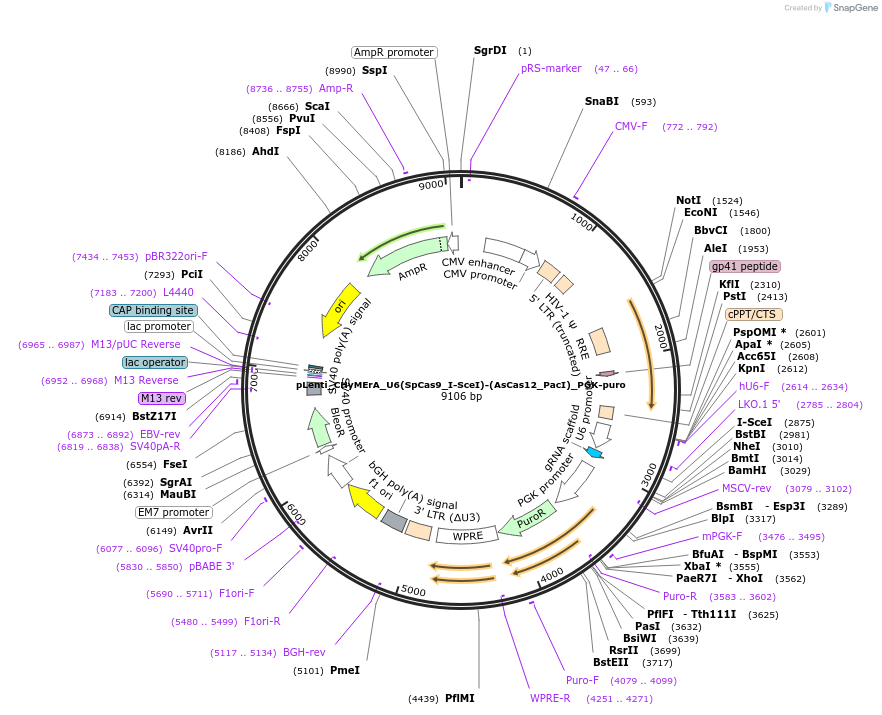
pLenti_CHyMErA_U6(SpCas9_I-SceI)-(AsCas12_PacI)_PGK-puro
Plasmid#189634PurposeLentiviral expression of single SpCas9 and AsCas12a gRNAs for generating combinatorial CHyMErA 3Cs librariesDepositorInserthU6 Cas9-Cas12a gRNA cassette, PGK puro cassette
UseCRISPR and LentiviralExpressionMammalianAvailable SinceNov. 16, 2023AvailabilityAcademic Institutions and Nonprofits only -
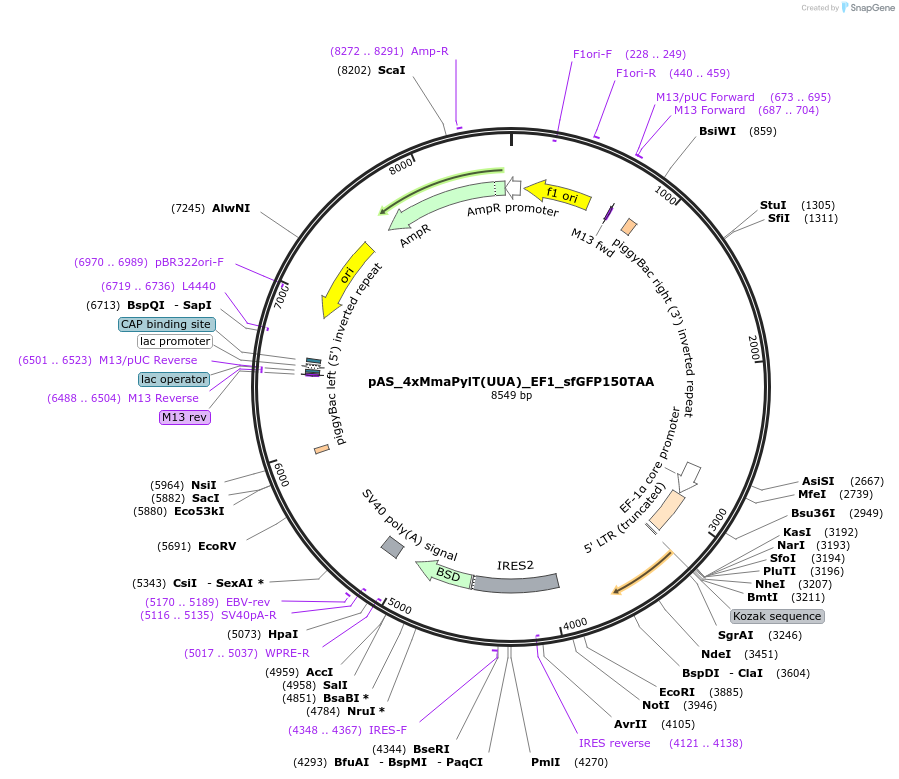
pAS_4xMmaPylT(UUA)_EF1_sfGFP150TAA
Plasmid#174897Purposeochre suppression reporter sfGFP150TAA expression, with MmaPylT(UUA) ochre suppressor tRNA cassetteDepositorInsertsf GFP
ExpressionMammalianMutation150TAA in sfGFPPromoterEF1Available SinceNov. 7, 2023AvailabilityAcademic Institutions and Nonprofits only -
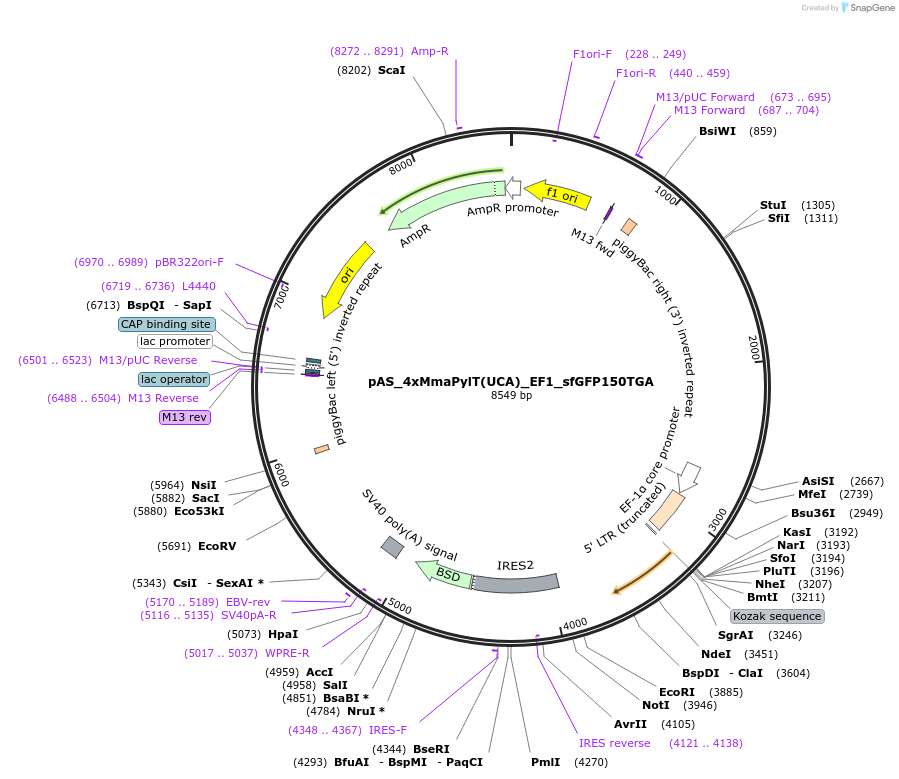
pAS_4xMmaPylT(UCA)_EF1_sfGFP150TGA
Plasmid#174898Purposeopal suppression reporter sfGFP150TGA expression, with MmaPylT(UCA) opal suppressor tRNA cassetteDepositorInsertsfGFP
ExpressionMammalianMutation150TGA in sfGFPPromoterEF1Available SinceNov. 7, 2023AvailabilityAcademic Institutions and Nonprofits only -
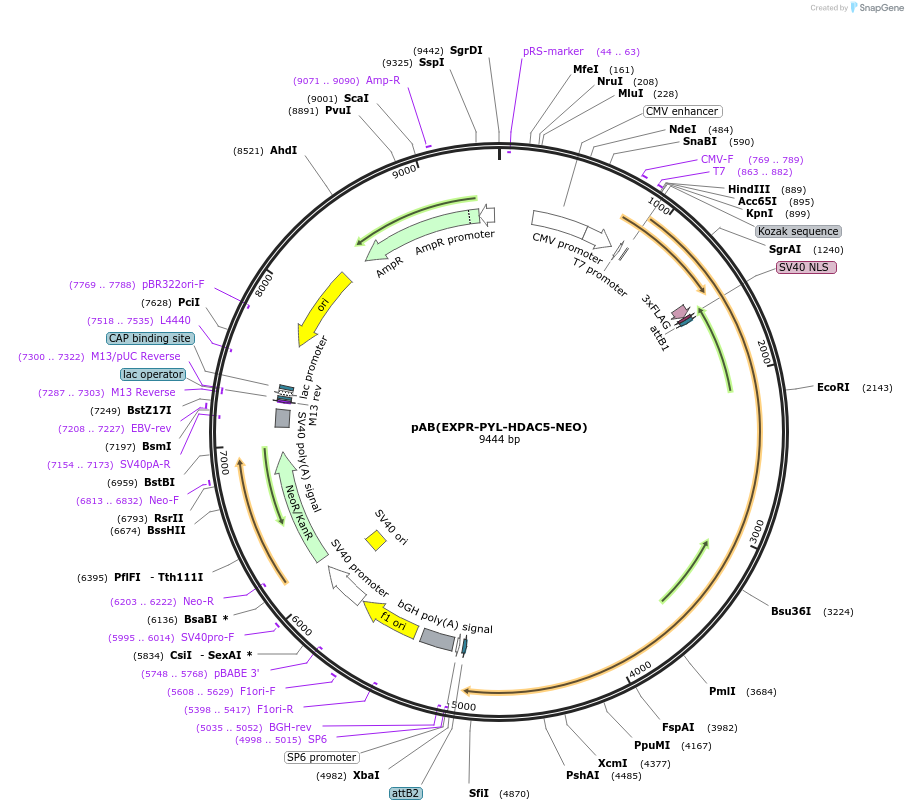
pAB(EXPR-PYL-HDAC5-NEO)
Plasmid#114395PurposeFor PYL-HDAC5 expressionDepositorInsertPYL-HDAC5
Tags3xFlag-NLS (internal)ExpressionMammalianMutationD593E compared with NCBI reference NP_005465.2PromoterCMVAvailable SinceAug. 22, 2023AvailabilityAcademic Institutions and Nonprofits only -
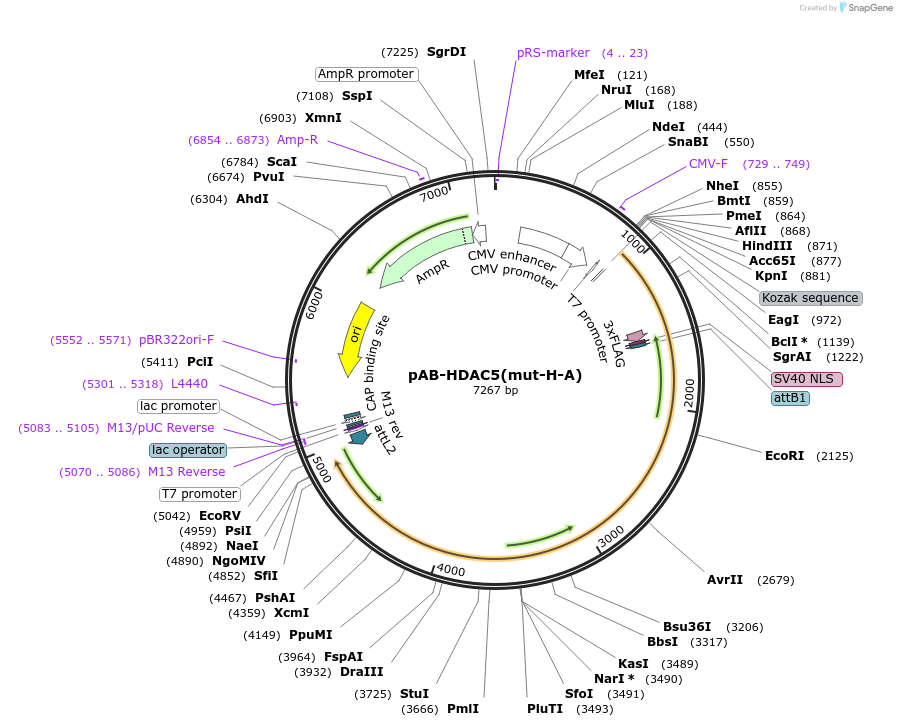
pAB-HDAC5(mut-H-A)
Plasmid#114397PurposeFor PYL-HDAC5 H1006A expressionDepositorInsertPYL-HDAC5 H1006A
Tags3xFlag-NLS (internal)ExpressionMammalianMutationD593E compared to NCBI ref NP_005465.2PromoterCMVAvailable SinceAug. 22, 2023AvailabilityAcademic Institutions and Nonprofits only -
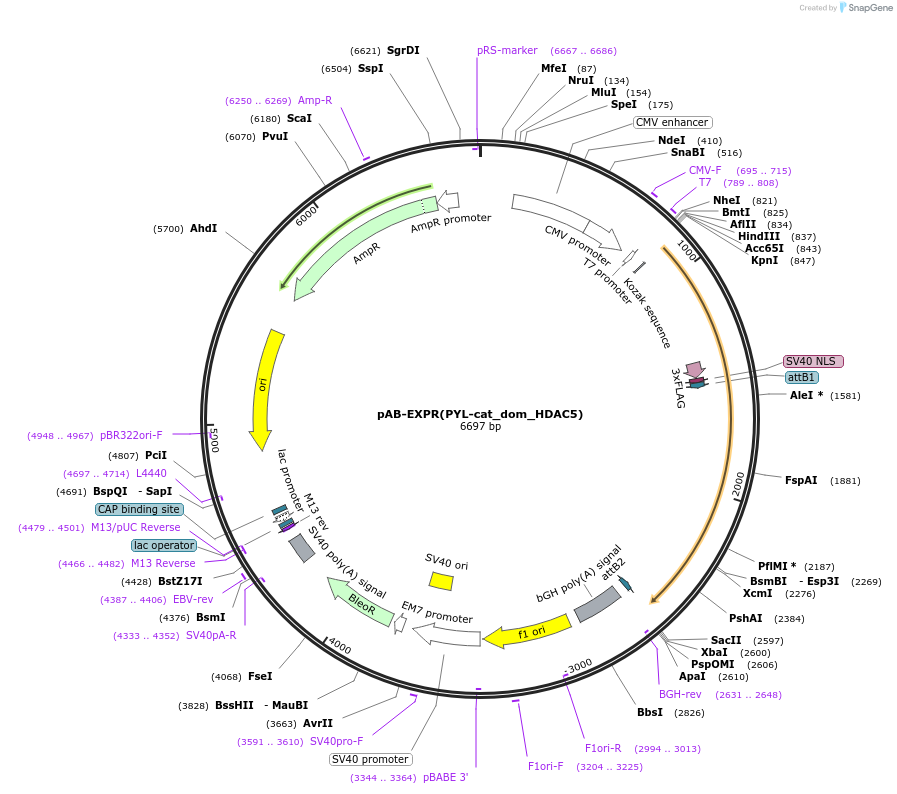
pAB-EXPR(PYL-cat_dom_HDAC5)
Plasmid#114401PurposeFor PYL-HDAC5 catalytic domain expressionDepositorInsertPYL-HDAC5 catalytic domain
Tags3xFlag-NLS (internal)ExpressionMammalianMutationS220F and the deletion of M221PromoterCMVAvailable SinceAug. 22, 2023AvailabilityAcademic Institutions and Nonprofits only -
pAB-EXPR(PYL-Nterm_dom_HDAC5)
Plasmid#114402PurposeFor PYL-HDAC5 N terminal domain expressionDepositorInsertPYL-HDAC5 N terminal domain
Tags3xFlag-NLS (internal)ExpressionMammalianMutationD593E compared to NCBI ref NP_005465.2PromoterCMVAvailable SinceAug. 22, 2023AvailabilityAcademic Institutions and Nonprofits only -
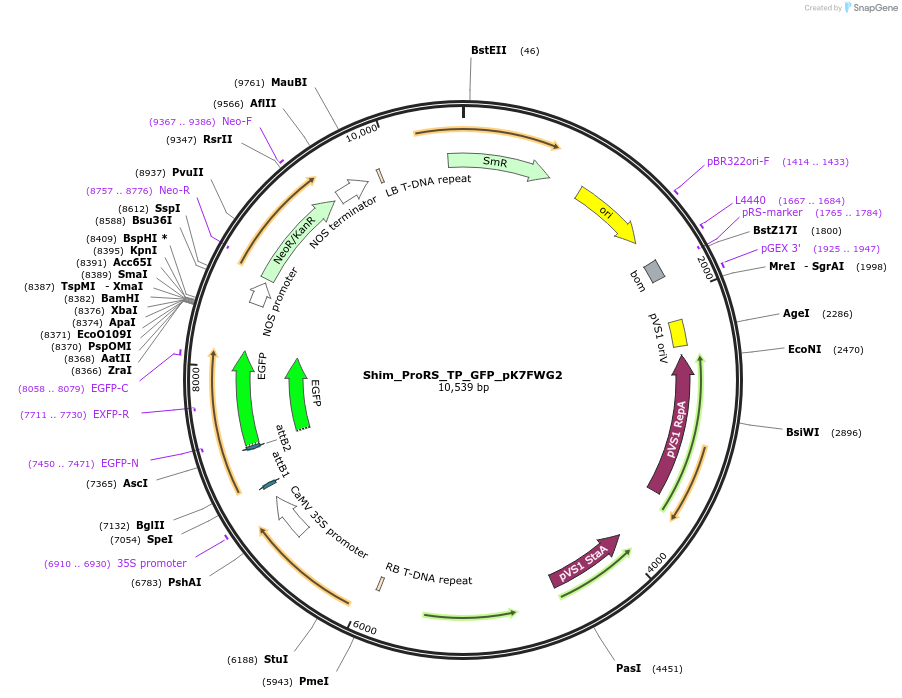
Shim_ProRS_TP_GFP_pK7FWG2
Plasmid#202651PurposeDetermine where transit peptide targets GFP in N. benthamianaDepositorInsertN-terminal transit peptide from putative cytosolic ProRS
TagsGreen Florescent ProtienExpressionPlantAvailable SinceAug. 10, 2023AvailabilityIndustry, Academic Institutions, and Nonprofits -
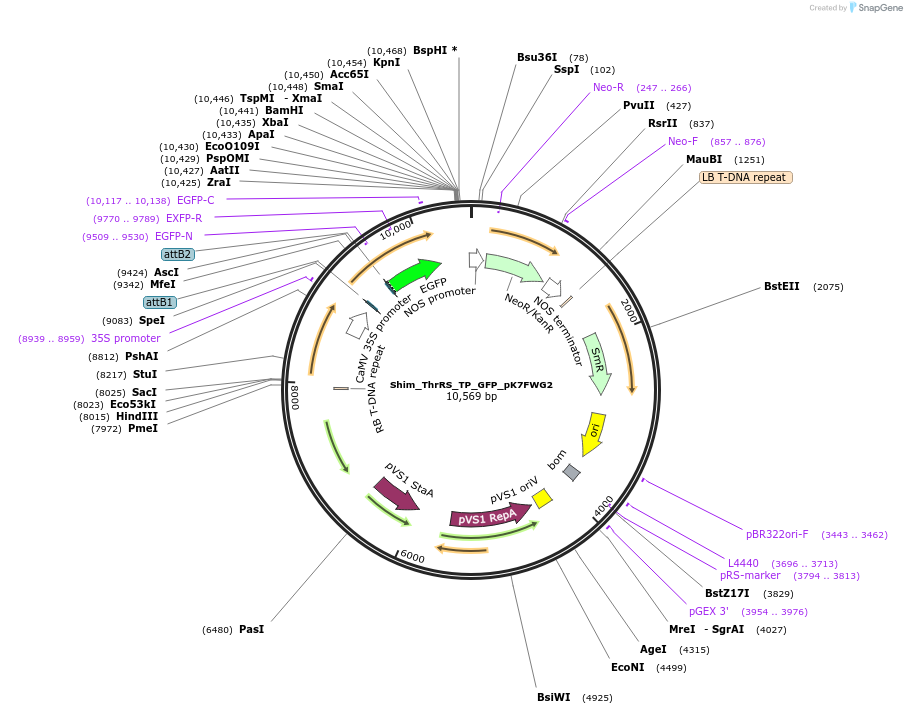
Shim_ThrRS_TP_GFP_pK7FWG2
Plasmid#202652PurposeDetermine where transit peptide targets GFP in N. benthamianaDepositorInsertN-terminal transit peptide from putative cytosolic ThrRS
TagsGreen Florescent ProtienExpressionPlantAvailable SinceAug. 10, 2023AvailabilityIndustry, Academic Institutions, and Nonprofits -
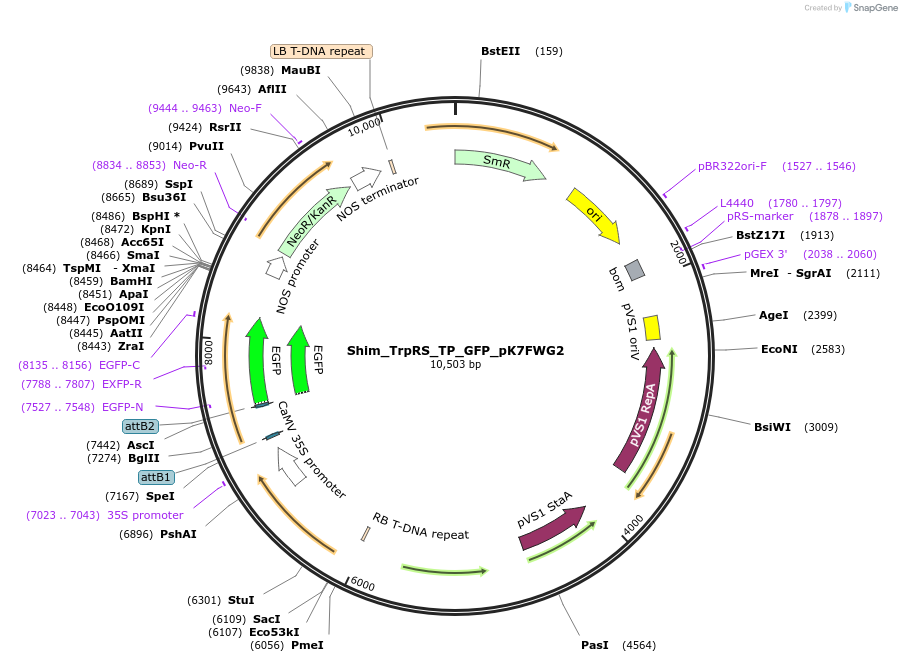
Shim_TrpRS_TP_GFP_pK7FWG2
Plasmid#202653PurposeDetermine where transit peptide targets GFP in N. benthamianaDepositorInsertN-terminal transit peptide from putative cytosolic TrpRS
TagsGreen Florescent ProtienExpressionPlantAvailable SinceAug. 10, 2023AvailabilityIndustry, Academic Institutions, and Nonprofits -
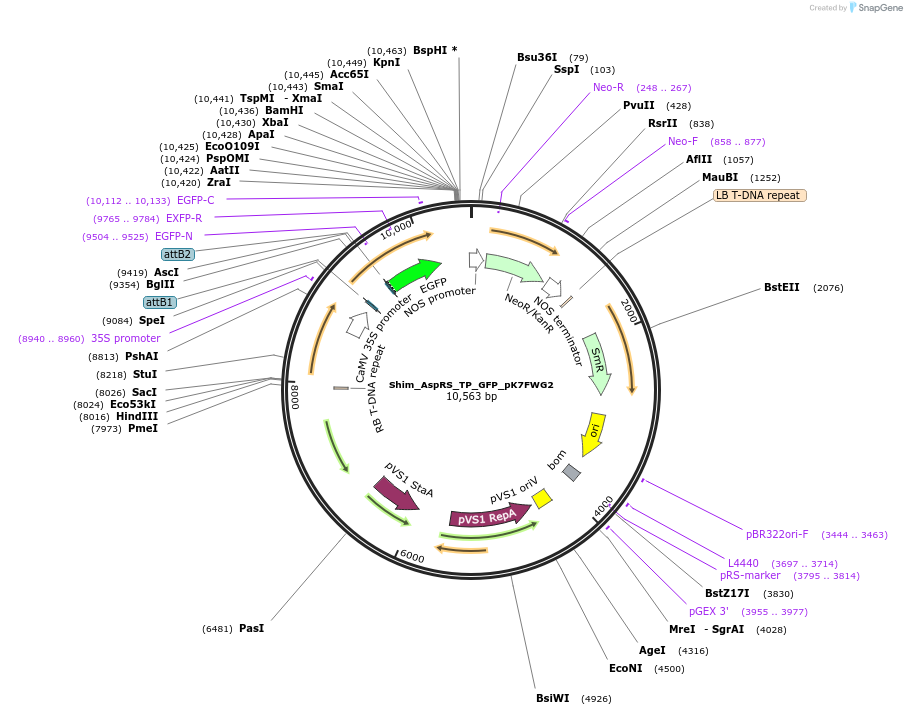
Shim_AspRS_TP_GFP_pK7FWG2
Plasmid#202647PurposeDetermine where transit peptide targets GFP in N. benthamianaDepositorInsertN-terminal transit peptide from putative cytosolic AspRS
TagsGreen Florescent ProtienExpressionPlantAvailable SinceAug. 10, 2023AvailabilityIndustry, Academic Institutions, and Nonprofits -
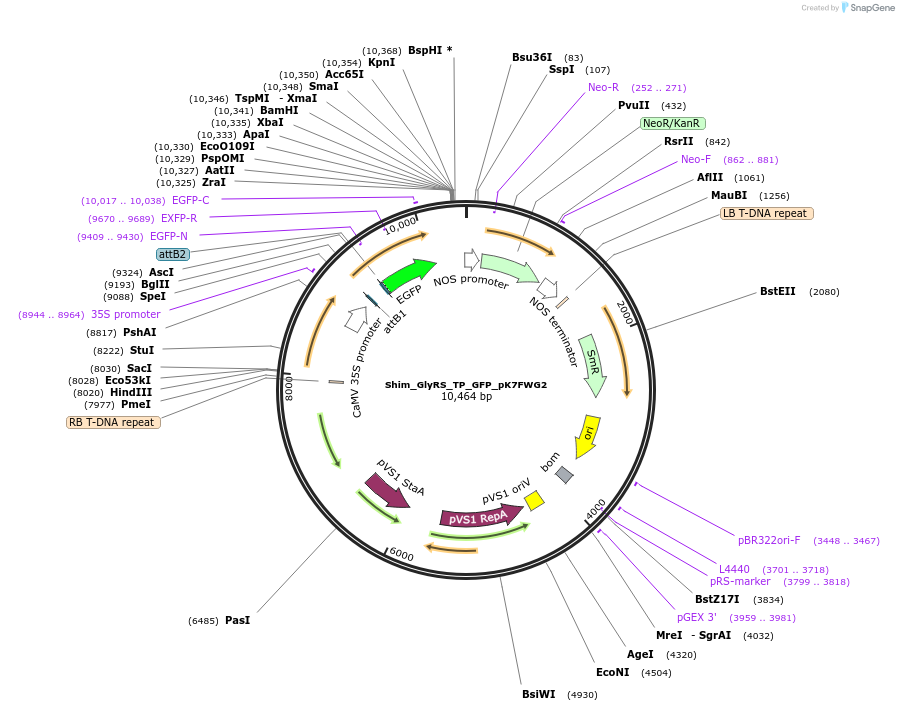
Shim_GlyRS_TP_GFP_pK7FWG2
Plasmid#202648PurposeDetermine where transit peptide targets GFP in N. benthamianaDepositorInsertN-terminal transit peptide from putative cytosolic GlyRS
TagsGreen Florescent ProtienExpressionPlantAvailable SinceAug. 10, 2023AvailabilityIndustry, Academic Institutions, and Nonprofits -
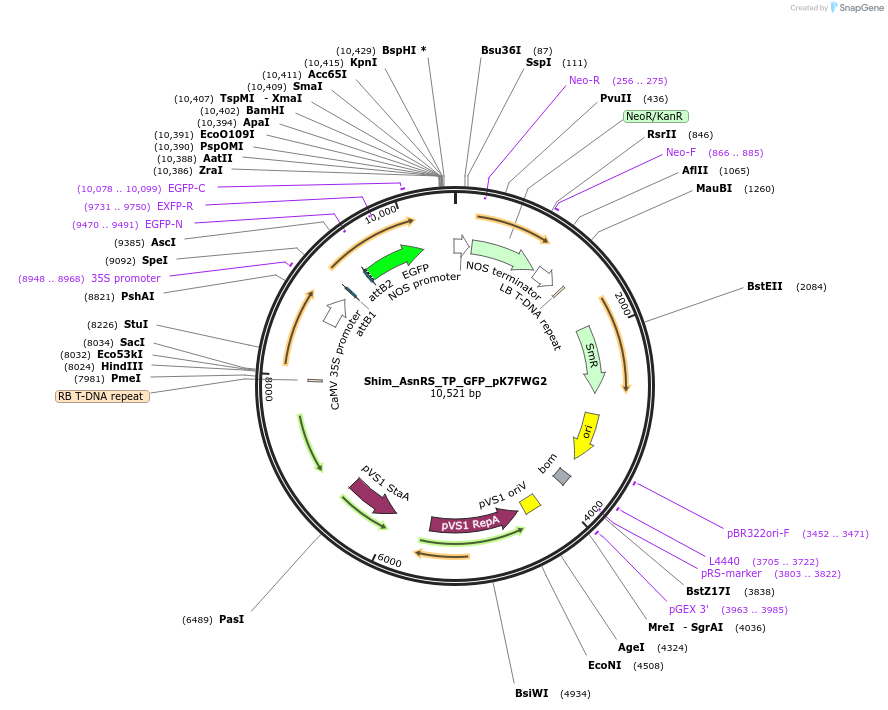
Shim_AsnRS_TP_GFP_pK7FWG2
Plasmid#202649PurposeDetermine where transit peptide targets GFP in N. benthamianaDepositorInsertN-terminal transit peptide from putative cytosolic AsnRS
TagsGreen Florescent ProtienExpressionPlantAvailable SinceAug. 10, 2023AvailabilityIndustry, Academic Institutions, and Nonprofits -
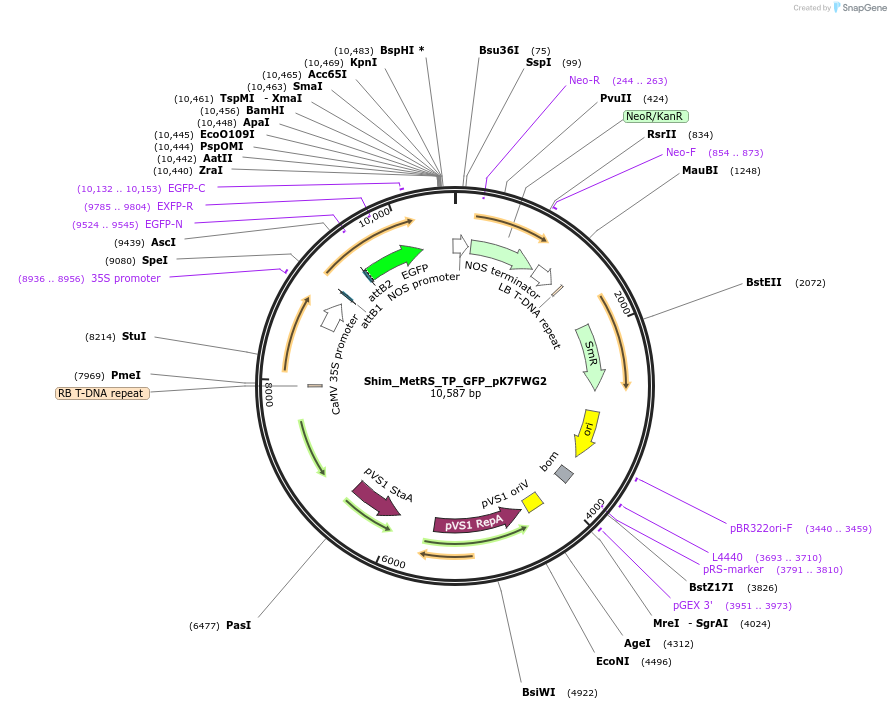
Shim_MetRS_TP_GFP_pK7FWG2
Plasmid#202650PurposeDetermine where transit peptide targets GFP in N. benthamianaDepositorInsertN-terminal transit peptide from putative cytosolic MetRS
TagsGreen Florescent ProtienExpressionPlantAvailable SinceAug. 10, 2023AvailabilityIndustry, Academic Institutions, and Nonprofits -
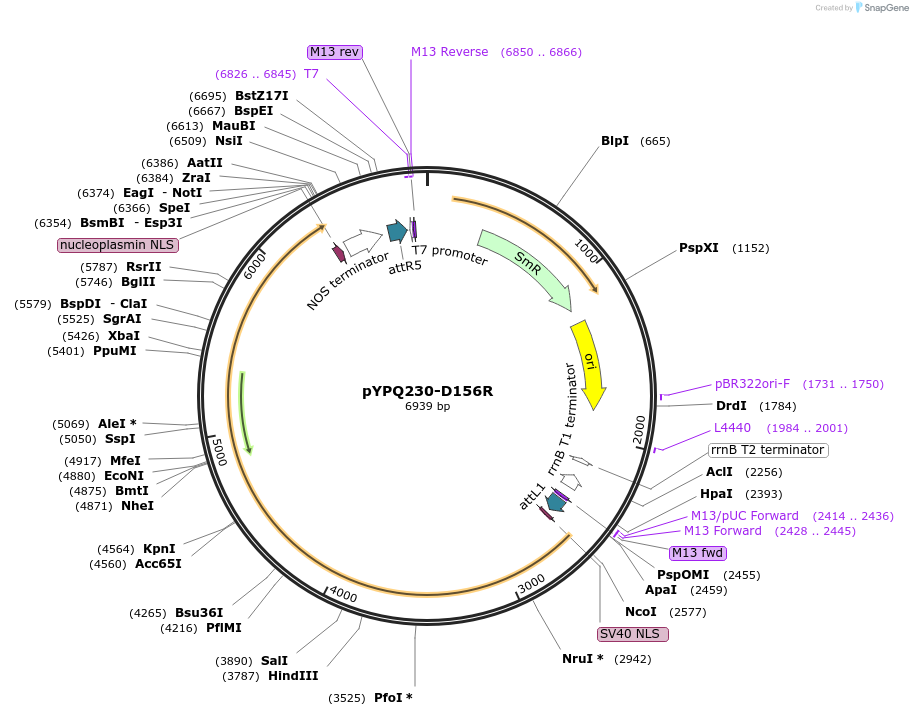
pYPQ230-D156R
Plasmid#188542PurposeGateway entry clone (attL1 & attR5) for LbCas12a-D156R mediated genome editingDepositorInsertLbCas12a-D156R
UseCRISPR; Gateway compatible lbcas12a-d156r entry c…TagsNLSExpressionPlantMutationD156RAvailable SinceMay 17, 2023AvailabilityAcademic Institutions and Nonprofits only -
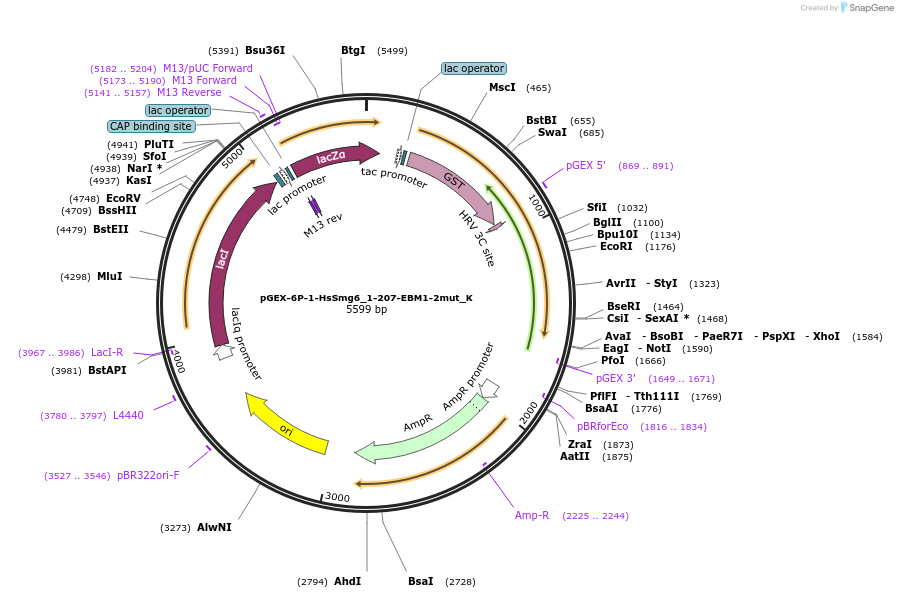
pGEX-6P-1-HsSmg6_1-207-EBM1-2mut_K
Plasmid#146756PurposeBacterial Expression of HsSmg6_1-207-EBM1-2mutDepositorInsertHsSmg6_1-207-EBM1-2mut (SMG6 Human)
ExpressionBacterialMutationone silent mutation compared to the sequence give…Available SinceMarch 16, 2023AvailabilityAcademic Institutions and Nonprofits only -
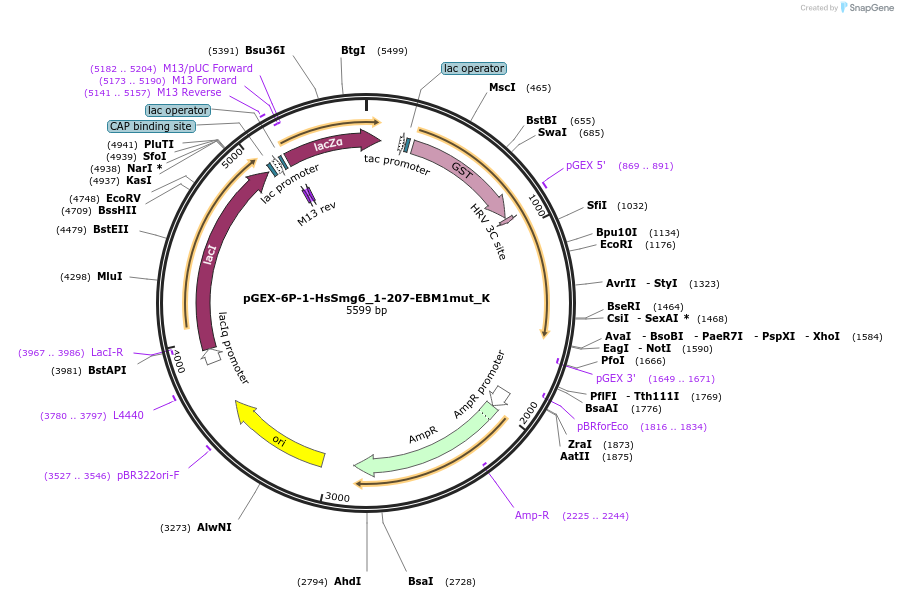
pGEX-6P-1-HsSmg6_1-207-EBM1mut_K
Plasmid#146757PurposeBacterial Expression of HsSmg6_1-207-EBM1mutDepositorInsertHsSmg6_1-207-EBM1mut (SMG6 Human)
ExpressionBacterialMutationone silent mutation compared to the sequence give…Available SinceMarch 16, 2023AvailabilityAcademic Institutions and Nonprofits only -
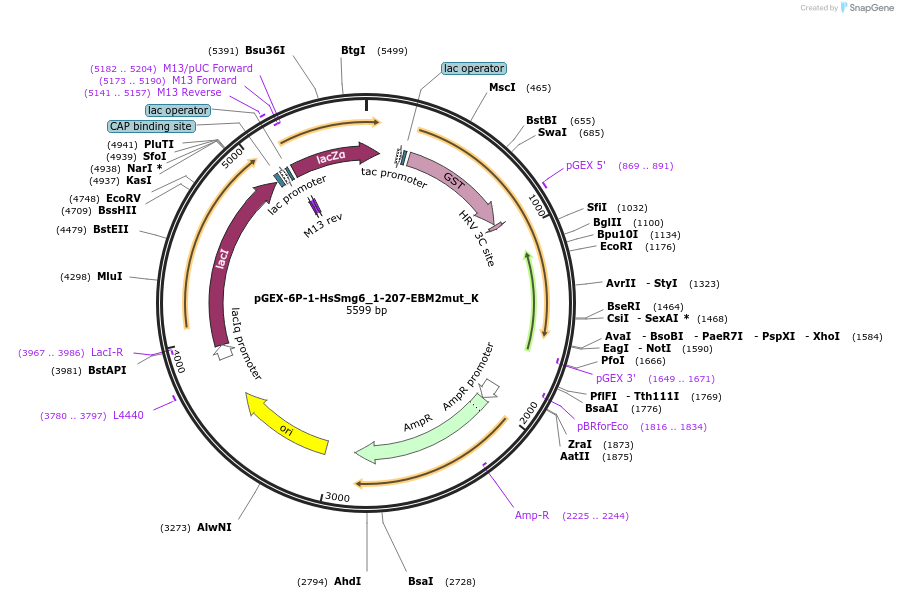
pGEX-6P-1-HsSmg6_1-207-EBM2mut_K
Plasmid#146758PurposeBacterial Expression of HsSmg6_1-207-EBM2mutDepositorInsertHsSmg6_1-207-EBM2mut (SMG6 Human)
ExpressionBacterialMutationone silent mutation compared to the sequence give…Available SinceMarch 16, 2023AvailabilityAcademic Institutions and Nonprofits only -
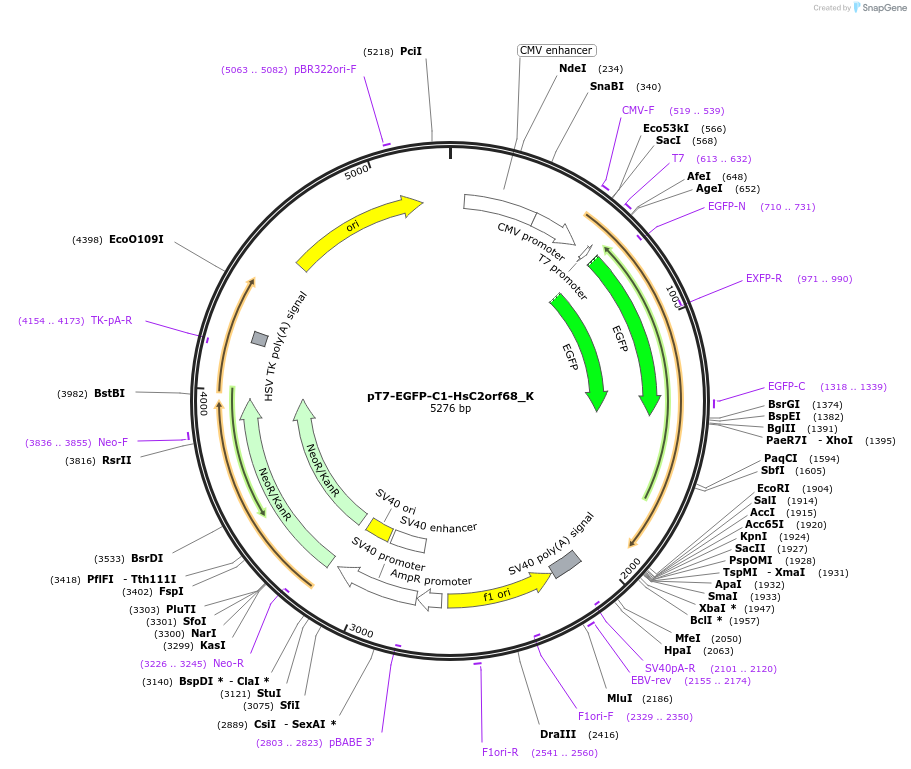
pT7-EGFP-C1-HsC2orf68_K
Plasmid#146760PurposeMammalian Expression of HsC2orf68DepositorInsertHsC2orf68 (C2orf68 Human)
ExpressionMammalianAvailable SinceMarch 16, 2023AvailabilityAcademic Institutions and Nonprofits only -
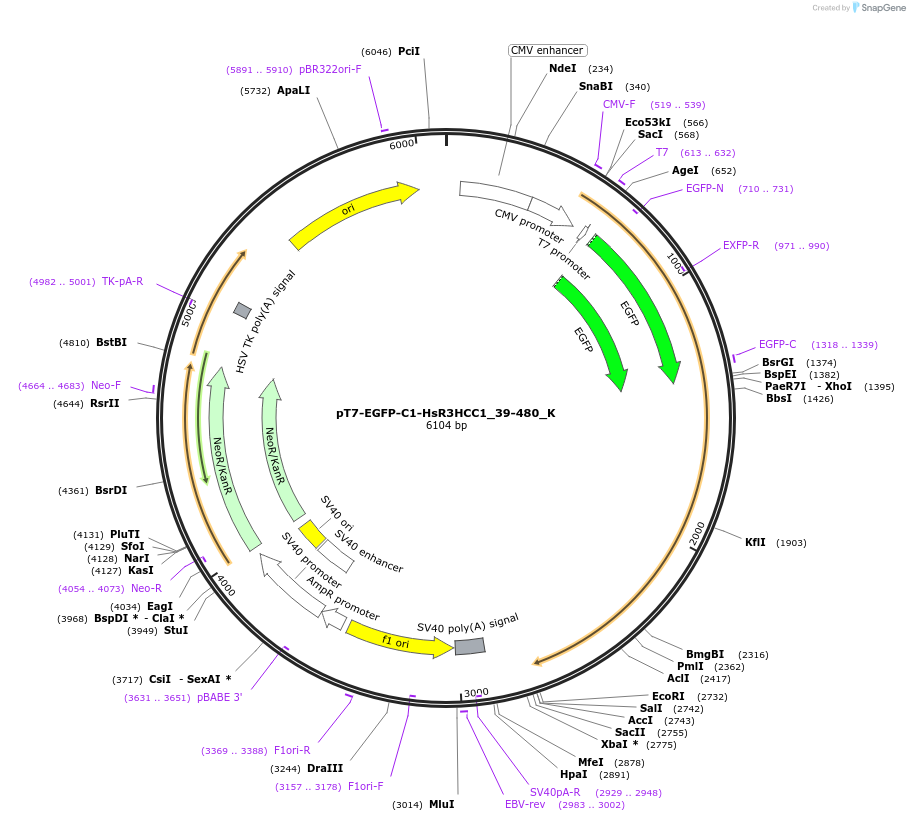
pT7-EGFP-C1-HsR3HCC1_39-480_K
Plasmid#146764PurposeMammalian Expression of HsR3HCC1_39-480DepositorInsertHsR3HCC1_39-480 (R3HCC1 Human)
ExpressionMammalianMutationsequence extracted from BC050572.1; 39-480 trunca…Available SinceMarch 16, 2023AvailabilityAcademic Institutions and Nonprofits only -
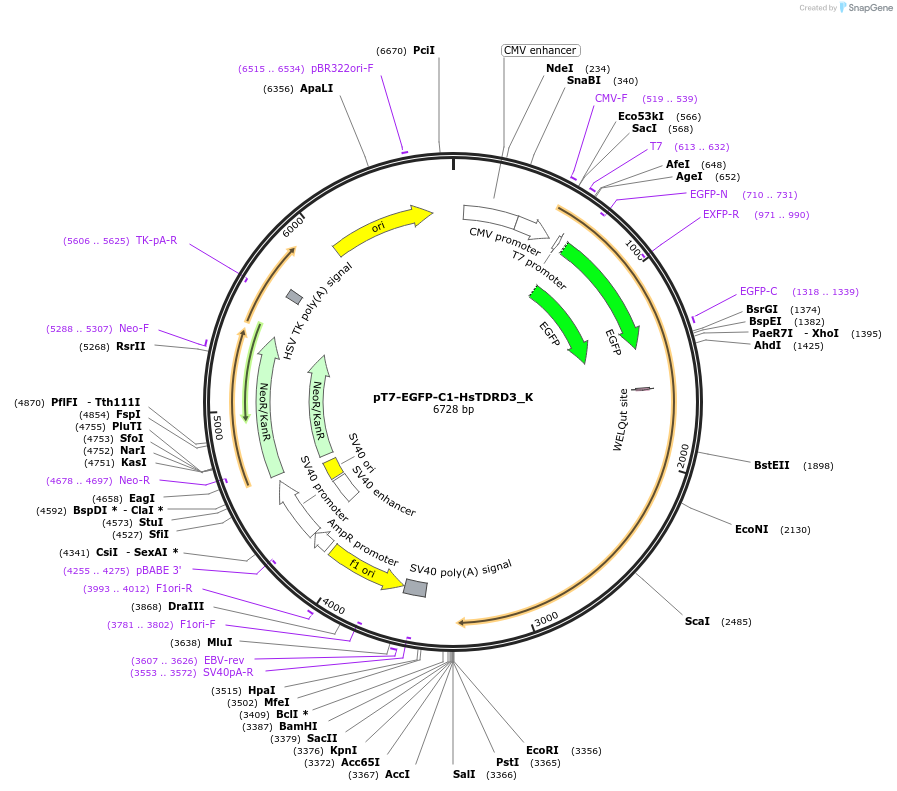
pT7-EGFP-C1-HsTDRD3_K
Plasmid#146768PurposeMammalian Expression of HsTDRD3DepositorInsertHsTDRD3 (TDRD3 Human)
ExpressionMammalianMutationtwo non silent mutations H100R and P508H compared…Available SinceMarch 16, 2023AvailabilityAcademic Institutions and Nonprofits only -
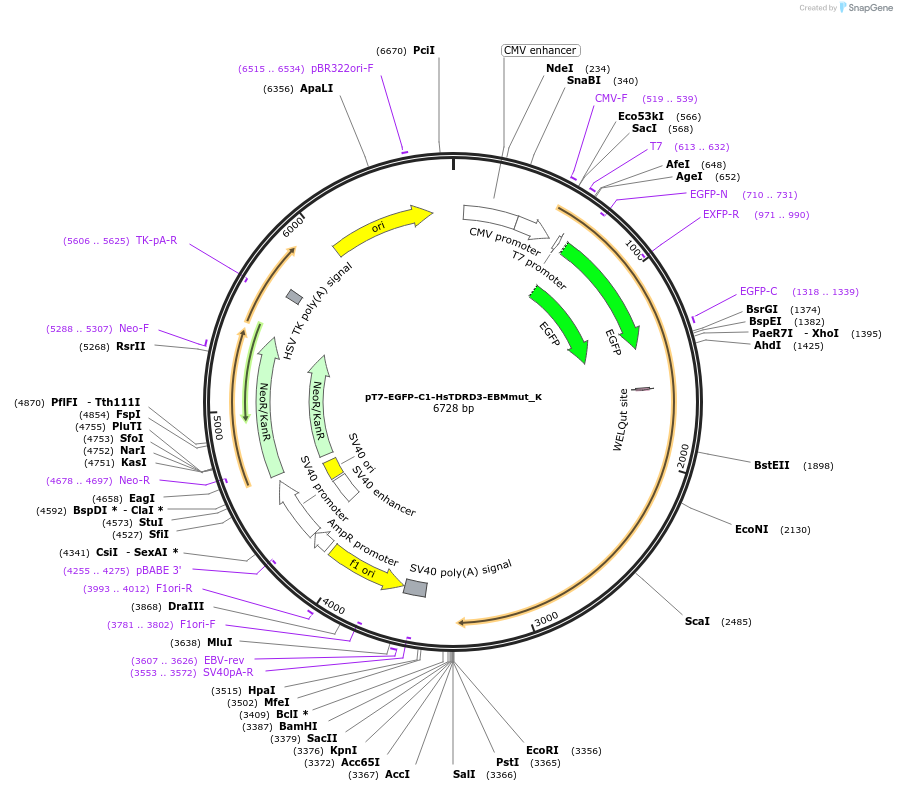
pT7-EGFP-C1-HsTDRD3-EBMmut_K
Plasmid#146769PurposeMammalian Expression of HsTDRD3-EBMmutDepositorInsertHsTDRD3-EBMmut (TDRD3 Human)
ExpressionMammalianMutationtwo non silent mutations H100R and P508H compare…Available SinceMarch 16, 2023AvailabilityAcademic Institutions and Nonprofits only -
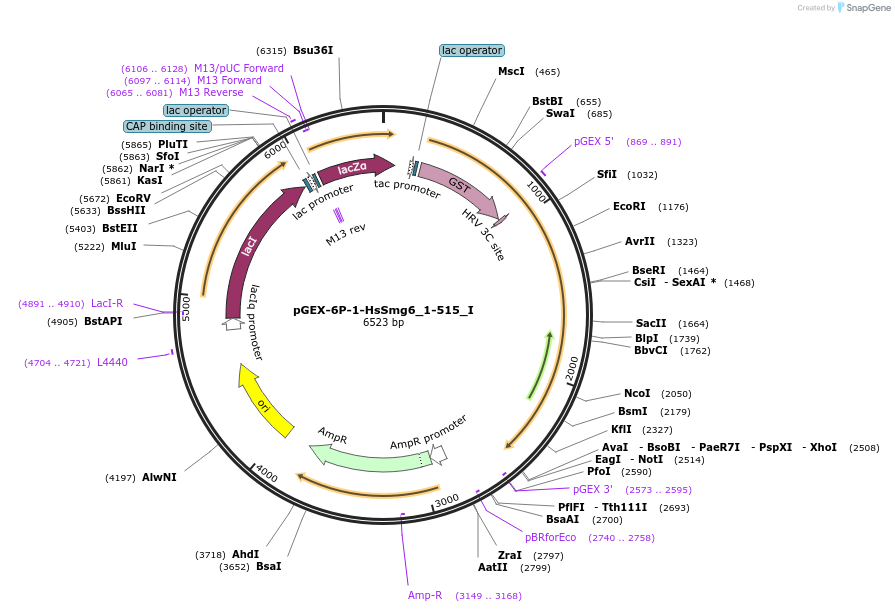
pGEX-6P-1-HsSmg6_1-515_I
Plasmid#146566PurposeBacterial Expression of HsSmg6_1-515DepositorInsertHsSmg6_1-515 (SMG6 Human)
ExpressionBacterialMutationtwo silent mutations and two non silent mutation …Available SinceMarch 16, 2023AvailabilityAcademic Institutions and Nonprofits only -
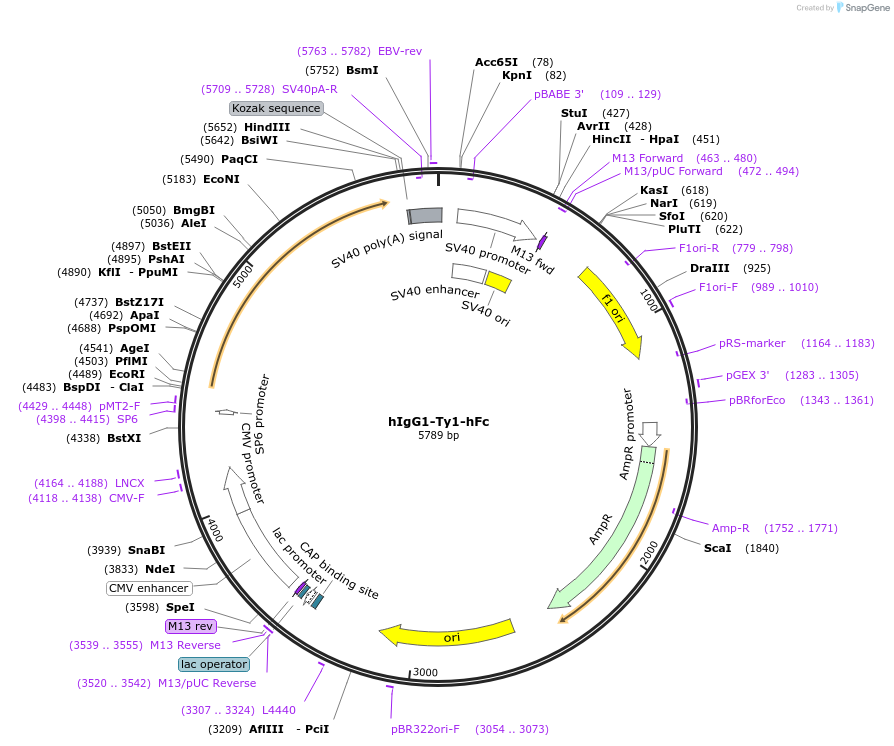
hIgG1-Ty1-hFc
Plasmid#194593PurposeMammalian expression of Ty1 nanobody upstream of human IgG1DepositorInsertTy1-hFc (S )
ExpressionMammalianAvailable SinceFeb. 1, 2023AvailabilityAcademic Institutions and Nonprofits only