We narrowed to 766 results for: HL
-
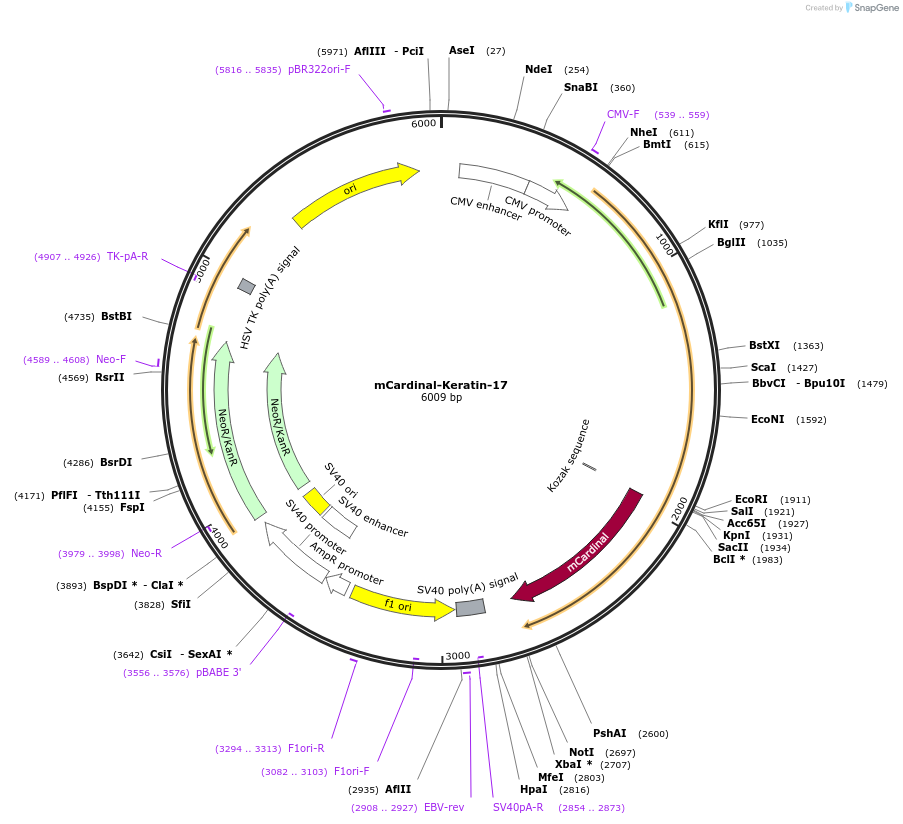
Plasmid#56164PurposeNote- This plasmid contains Keratin 18. Localization: Cytokeratin, Excitation: 604, Emission: 659DepositorAvailable SinceMarch 4, 2015AvailabilityIndustry, Academic Institutions, and Nonprofits
-
mCardinal-H1-10
Plasmid#56161PurposeLocalization: Nucleus/Histones, Excitation: 604, Emission: 659DepositorAvailable SinceFeb. 27, 2015AvailabilityIndustry, Academic Institutions, and Nonprofits -
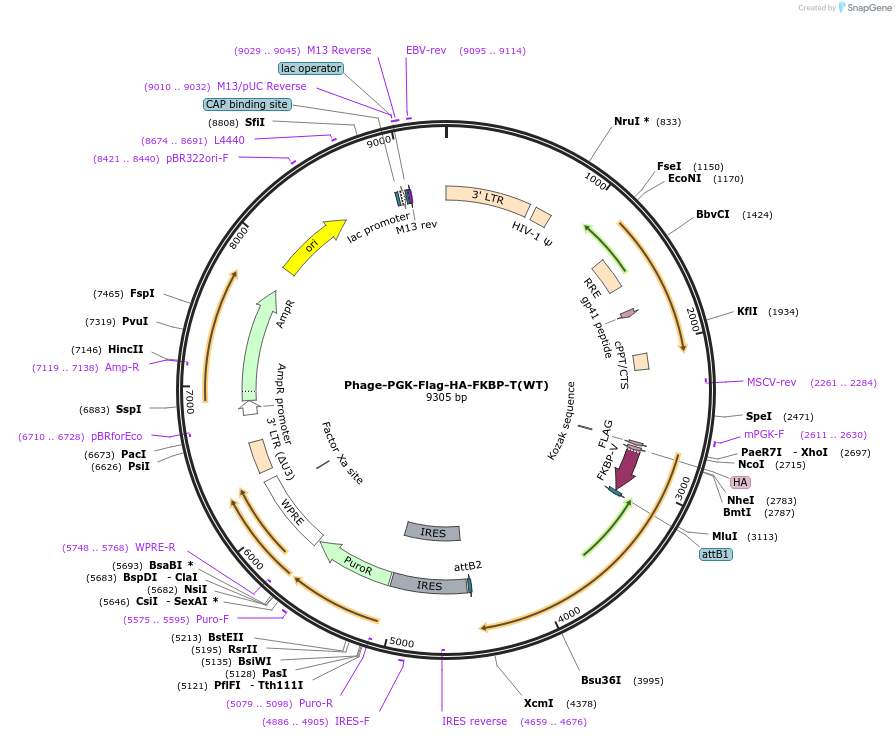
Phage-PGK-Flag-HA-FKBP-T(WT)
Plasmid#181734Purposefor lentiviral expression of brachyury ORF in mammalian cellsDepositorInsertT (brachyury)
UseLentiviralMutationa silent mutation in the PAM site of an sgRNAt ta…Available SinceMarch 21, 2022AvailabilityIndustry, Academic Institutions, and Nonprofits -
mCardinal-LaminB1-10
Plasmid#56165PurposeLocalization: Nuclear Envelope, Excitation: 604, Emission: 659DepositorAvailable SinceMarch 4, 2015AvailabilityIndustry, Academic Institutions, and Nonprofits -
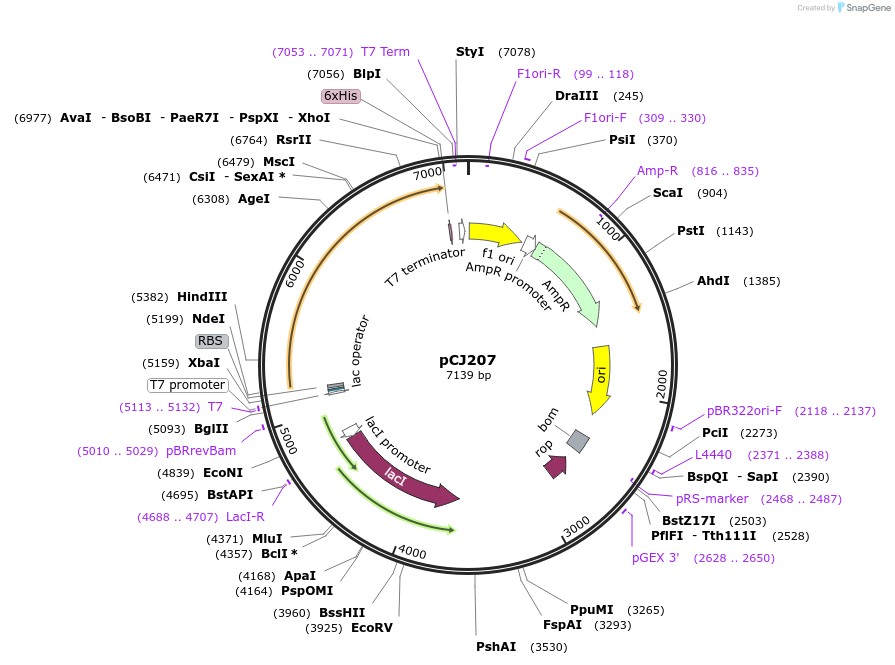
pCJ207
Plasmid#162680PurposepET-21b(+) based plasmid for expression of the putative MHETase from Hydrogenophaga sp. PML113 (Genbank WP_083293388.1) with C-terminal His tag, codon optimized for expression in E. coli K12.DepositorInsertPutative MHETase from Hydrogenophaga sp. PML113 (Genbank WP_083293388.1) with signal peptide
TagsHisExpressionBacterialMutationcodon optimized for expression in E. coli K12PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
mCardinal-Actin-C-18
Plasmid#56151PurposeLocalization: Actin, Excitation: 604, Emission: 659DepositorAvailable SinceFeb. 27, 2015AvailabilityIndustry, Academic Institutions, and Nonprofits -
mCardinal-MCAK-C-7
Plasmid#56168PurposeLocalization: Microtubules/Kinesin, Excitation: 604, Emission: 659DepositorAvailable SinceMarch 4, 2015AvailabilityIndustry, Academic Institutions, and Nonprofits -
p-miR-Report + WT miR-181 BS in GATA6 3UTR
Plasmid#22794DepositorInsertWild-type miR-181 Binding Site in GATA binding protein 6 3 Untranslated Region (GATA6 Human)
UseLuciferaseAvailable SinceMarch 24, 2010AvailabilityAcademic Institutions and Nonprofits only -
p-miR-Report + MT miR-181 BS in GATA6 3UTR
Plasmid#22795DepositorInsertMutant miR-181 Binding Site in GATA binding protein 6 3 Untranslated Region (GATA6 Human)
UseLuciferaseAvailable SinceMarch 24, 2010AvailabilityAcademic Institutions and Nonprofits only -
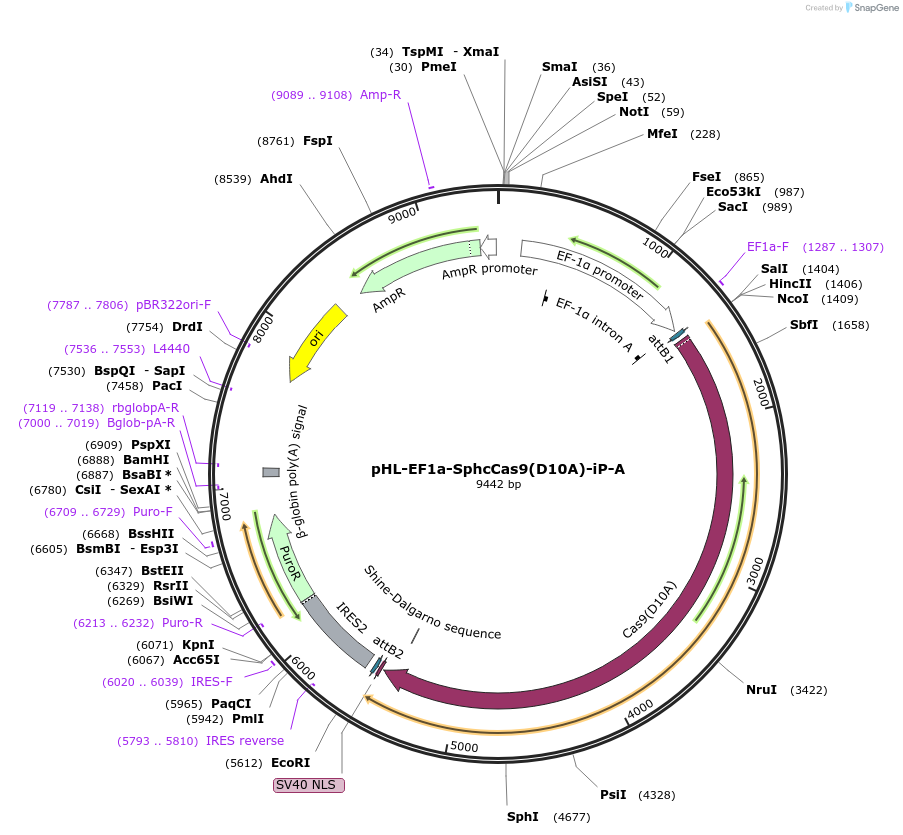
pHL-EF1a-SphcCas9(D10A)-iP-A
Plasmid#60600PurposeExpresses D10A mutant (nickase) of human codon-optimized Cas9 (derived from Streptococcus pyogenes) and pruomycin resistance gene.DepositorInsertsCRISPR Cas9 D10A
puromycin resistance gene
UseCRISPRExpressionMammalianMutationChanged Asp 10 to Ala, Codon usage optimized for …Available SinceDec. 11, 2014AvailabilityAcademic Institutions and Nonprofits only -
mCardinal-Calnexin-N-14
Plasmid#56154PurposeLocalization: Endoplasmic Reticulum, Excitation: 604, Emission: 659DepositorInsertCalnexin (CANX Human)
TagsmCardinalExpressionMammalianMutationI474N in CalnexinPromoterCMVAvailable SinceFeb. 27, 2015AvailabilityIndustry, Academic Institutions, and Nonprofits -
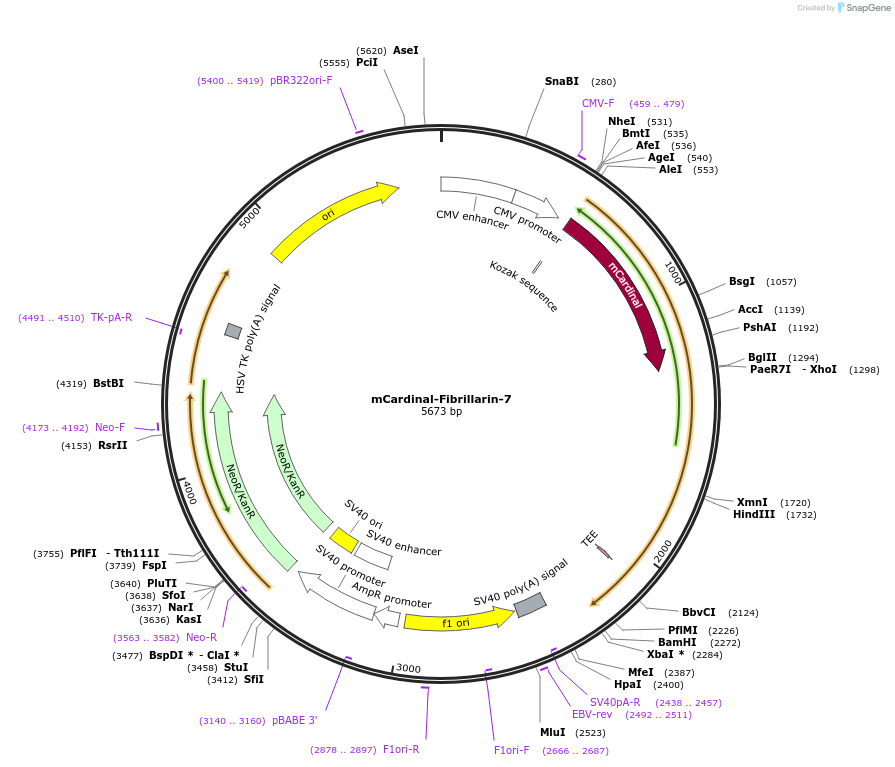
mCardinal-Fibrillarin-7
Plasmid#56160PurposeLocalization: Nuclear - Nucleolus, Excitation: 604, Emission: 659DepositorAvailable SinceMarch 4, 2015AvailabilityIndustry, Academic Institutions, and Nonprofits -
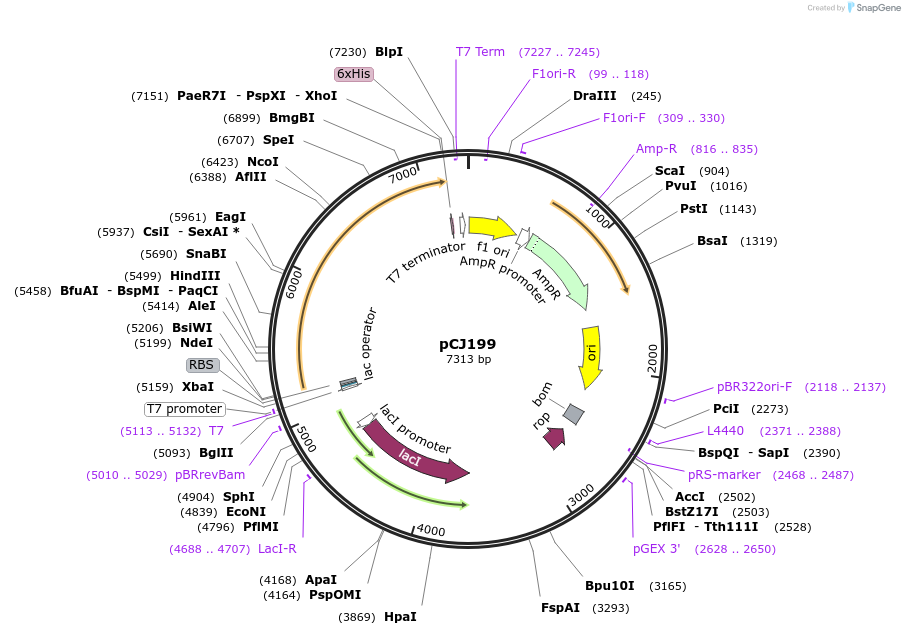
pCJ199
Plasmid#162672PurposepET-21b(+) based plasmid for expression of the putative MHET hydrolase from Comamonas thiooxydans (Genbank WP_080747404.1) with C-terminal His tag, codon optimized for expression in E. coli K12.DepositorInsertPutative MHET hydrolase from Comamonas thiooxydans (Genbank WP_080747404.1) with signal peptide
TagsHisExpressionBacterialMutationcodon optimized for expression in E. coli K12PromoterT7/lacAvailable SinceJan. 7, 2021AvailabilityAcademic Institutions and Nonprofits only -
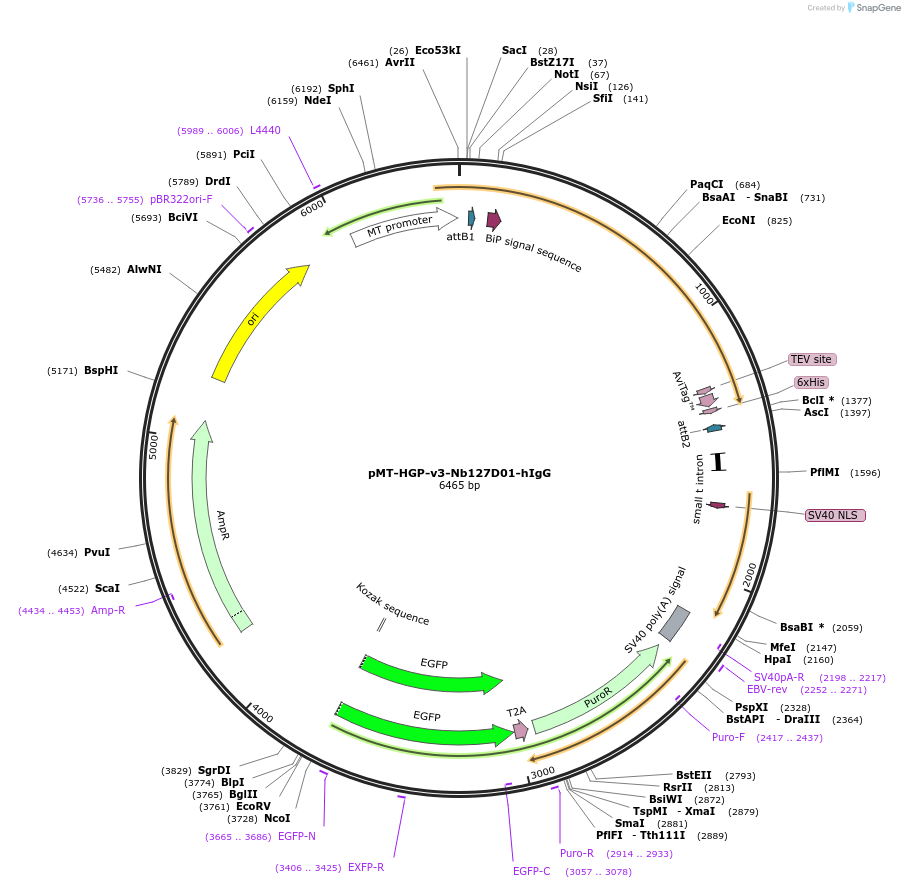
pMT-HGP-v3-Nb127D01-hIgG
Plasmid#171564PurposeExpresses hIgG-tagged anti-CXCR2 (human) recombinant llama nanobody (Nb127D01-human IgG-TEV-Avi-His) in fly cellsDepositorInsertNb127D01-human IgG-TEV-Avi-His (CXCR2 Human)
TagsBiP signal peptide and hIgG1-hinge-N297A-TEV site…ExpressionInsectAvailable SinceJune 29, 2021AvailabilityAcademic Institutions and Nonprofits only -
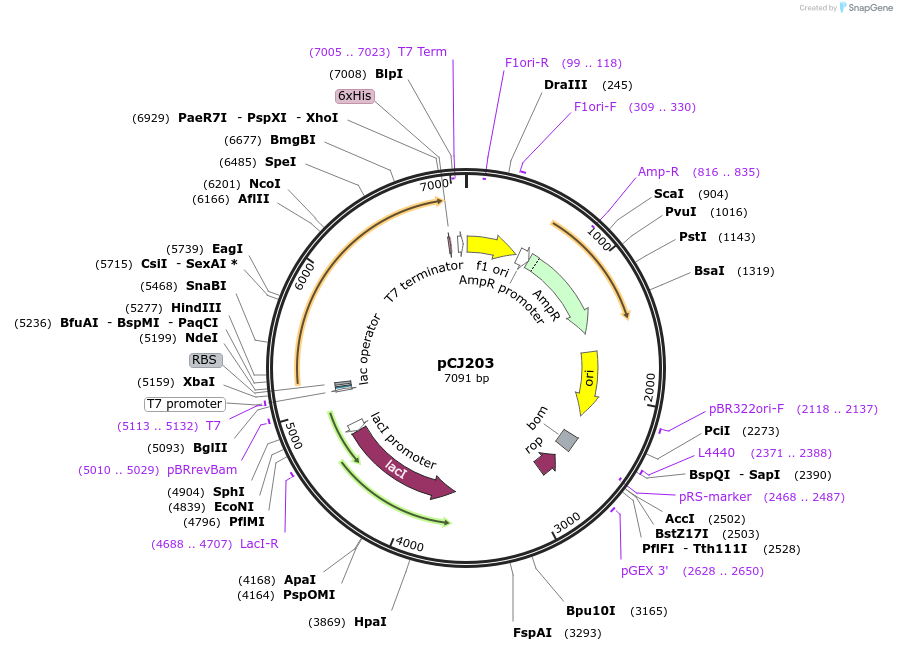
pCJ203
Plasmid#162678PurposepET-21b(+) based plasmid for expression of the putative MHET hydrolase from Comomonas thiooxydans (Genbank WP_080747404.1) with deletion of the predicted signal peptide (Phe2:Ala75)DepositorInsertPutative MHETase from Comomonas thiooxydans (Genbank WP_080747404.1) without 75 residue signal peptide
TagsHisExpressionBacterialMutationdeltaF2-A75; codon optimized for expression in E.…PromoterT7/lacAvailable SinceFeb. 3, 2021AvailabilityAcademic Institutions and Nonprofits only -
p-miR-Report + WT miR-181 BS in CDX2 3UTR
Plasmid#22792DepositorInsertWild-type miR-181 Binding Site in Caudal type homeobox transcription factor 2 3 Untranslated Region (CDX2 Human)
UseLuciferaseAvailable SinceMarch 24, 2010AvailabilityAcademic Institutions and Nonprofits only -
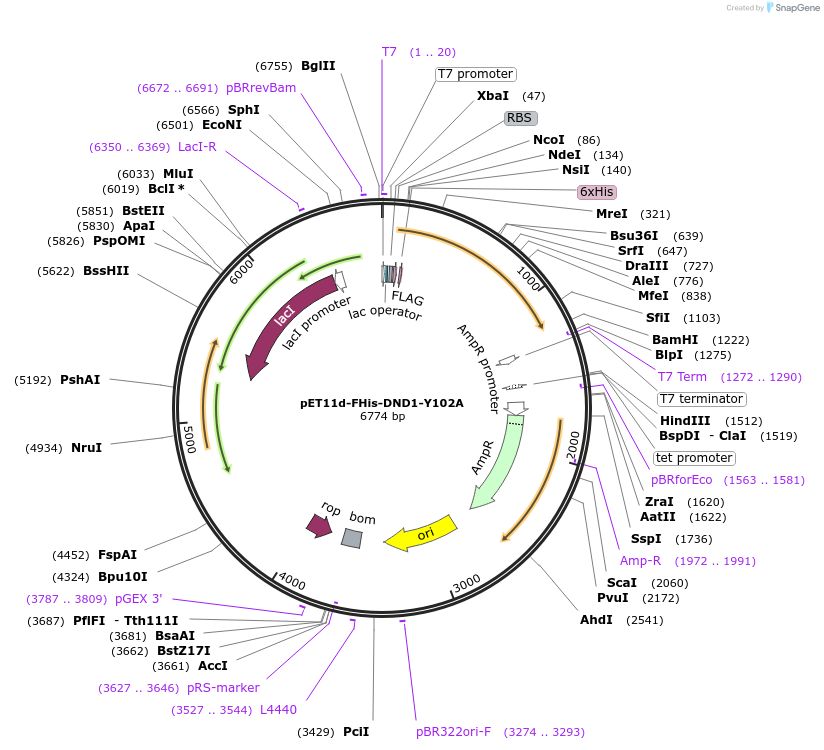
pET11d-FHis-DND1-Y102A
Plasmid#70082PurposeBacterial expression of FLAG-and 6xHis-tagged human DND1 protein with defective RRM (Y102A). Codons are optimized for bacteria expression.DepositorInsertDnd1 (DND1 Human)
Tags6xHIS and FLAGExpressionBacterialMutationY102A. Codon optimized for bacterial expressionAvailable SinceNov. 22, 2017AvailabilityAcademic Institutions and Nonprofits only -
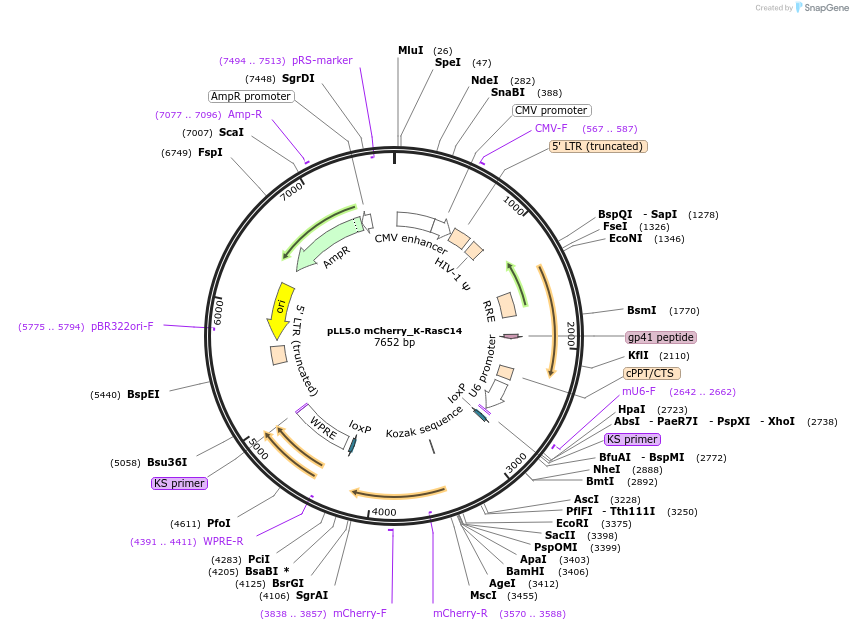
pLL5.0 mCherry_K-RasC14
Plasmid#101286PurposeLentiviral expression of K-Ras C14DepositorAvailable SinceMarch 28, 2019AvailabilityAcademic Institutions and Nonprofits only -
p-miR-Report + MT miR-181 BS in CDX2 3UTR
Plasmid#22793DepositorInsertMutant miR-181 Binding Site in Caudal type homeobox transcription factor 2 3 Untranslated Region (CDX2 Human)
UseLuciferaseAvailable SinceMarch 30, 2010AvailabilityAcademic Institutions and Nonprofits only -
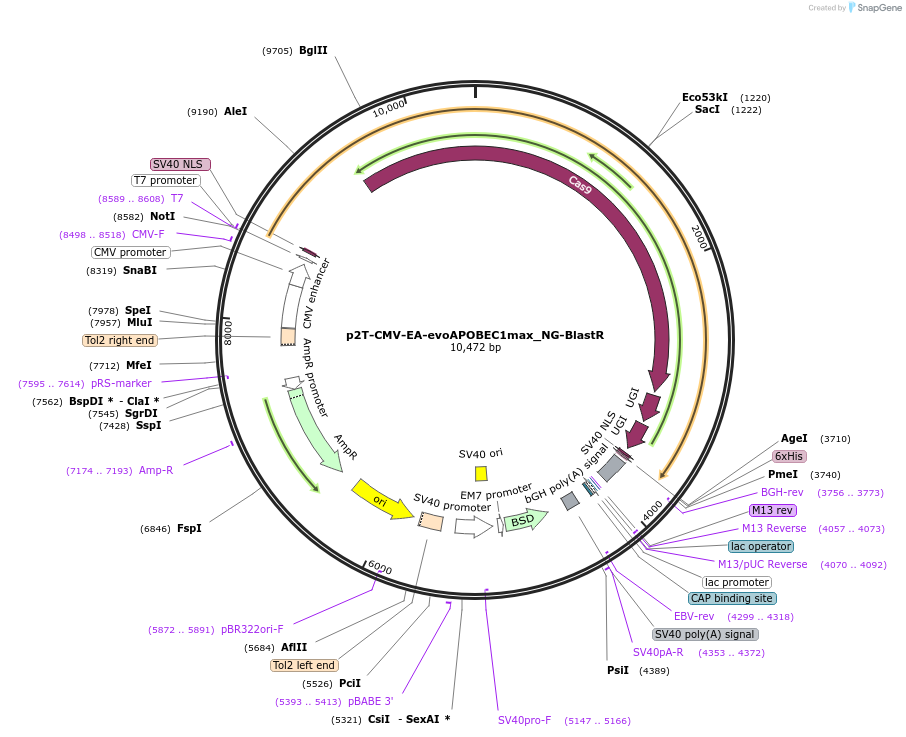
p2T-CMV-EA-evoAPOBEC1max_NG-BlastR
Plasmid#232721PurposeC-to-T base editorDepositorInsertevoAPOBEC1max H47E+S48A-nCas9(NG)-UGI-UGI
ExpressionMammalianMutationwithin evoAPOBEC1: H47E+S48A; within Cas9 D10AAvailable SinceMarch 3, 2025AvailabilityAcademic Institutions and Nonprofits only