We narrowed to 10,373 results for: epo
-
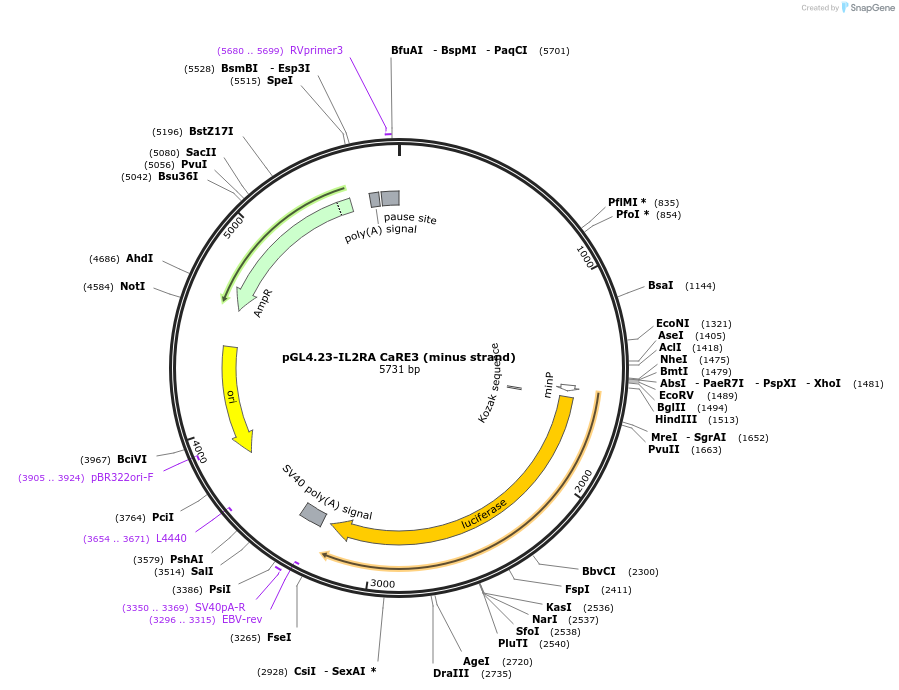
Plasmid#91835PurposeLuciferase reporter for IL2RA enhancer (IGI-P0612)DepositorInsertIL2RA CaRE3 (minus strand) (IL2RA Human)
UseLuciferaseExpressionMammalianMutationrs7893324 A>G, rs7090530 G>T, rs7090512 A&g…Available SinceJune 8, 2017AvailabilityAcademic Institutions and Nonprofits only -
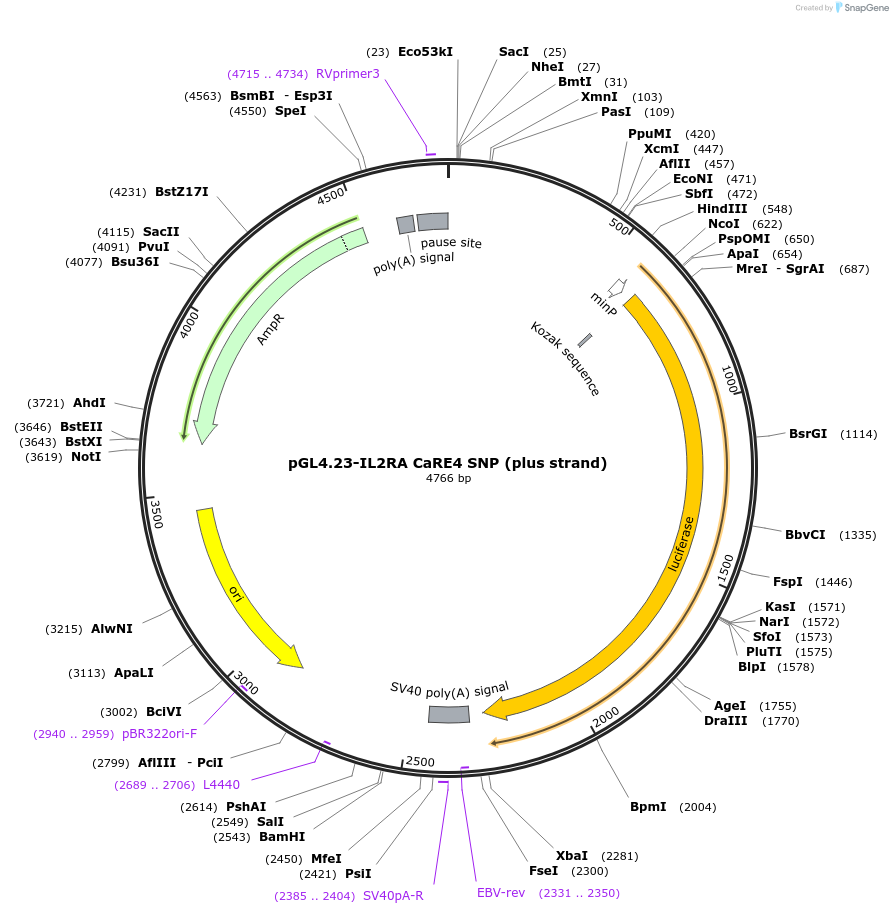
pGL4.23-IL2RA CaRE4 SNP (plus strand)
Plasmid#91851PurposeLuciferase reporter for IL2RA enhancer (IGI-P0346)DepositorInsertIL2RA CaRE4 SNP (plus strand) (IL2RA Human)
UseLuciferaseExpressionMammalianMutationrs61839660 C>TAvailable SinceMay 31, 2017AvailabilityAcademic Institutions and Nonprofits only -
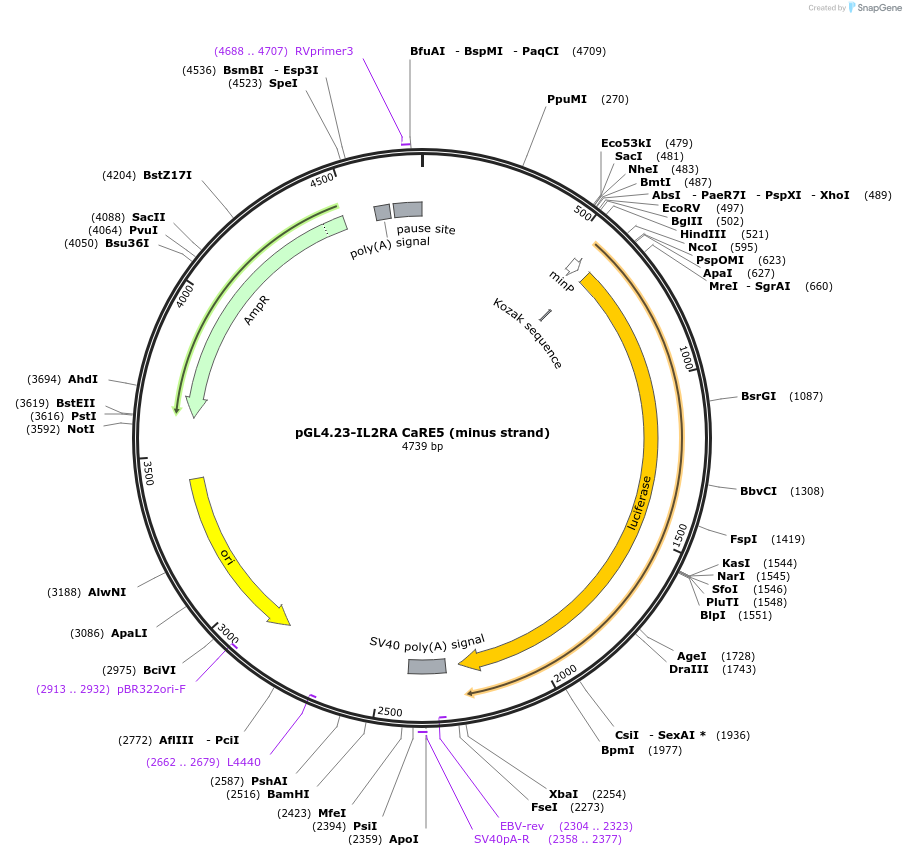
pGL4.23-IL2RA CaRE5 (minus strand)
Plasmid#91836PurposeLuciferase reporter for IL2RA enhancer (IGI-P0613)DepositorInsertIL2RA CaRE5 (minus strand) (IL2RA Human)
UseLuciferaseExpressionMammalianMutationPerfect match to reference sequenceAvailable SinceMay 30, 2017AvailabilityAcademic Institutions and Nonprofits only -
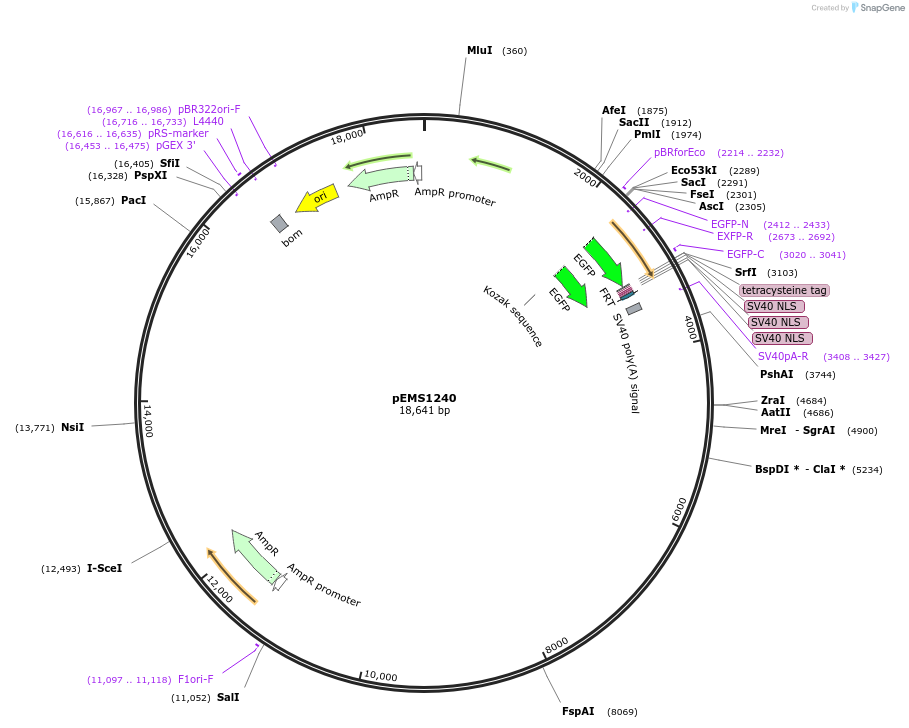
pEMS1240
Plasmid#29122PurposeExpression of the reporter gene under the control of the MiniPromoter in this plasmid has not been testedDepositorInsertPle11
TagsEGFP-NLSAvailable SinceNov. 4, 2011AvailabilityAcademic Institutions and Nonprofits only -
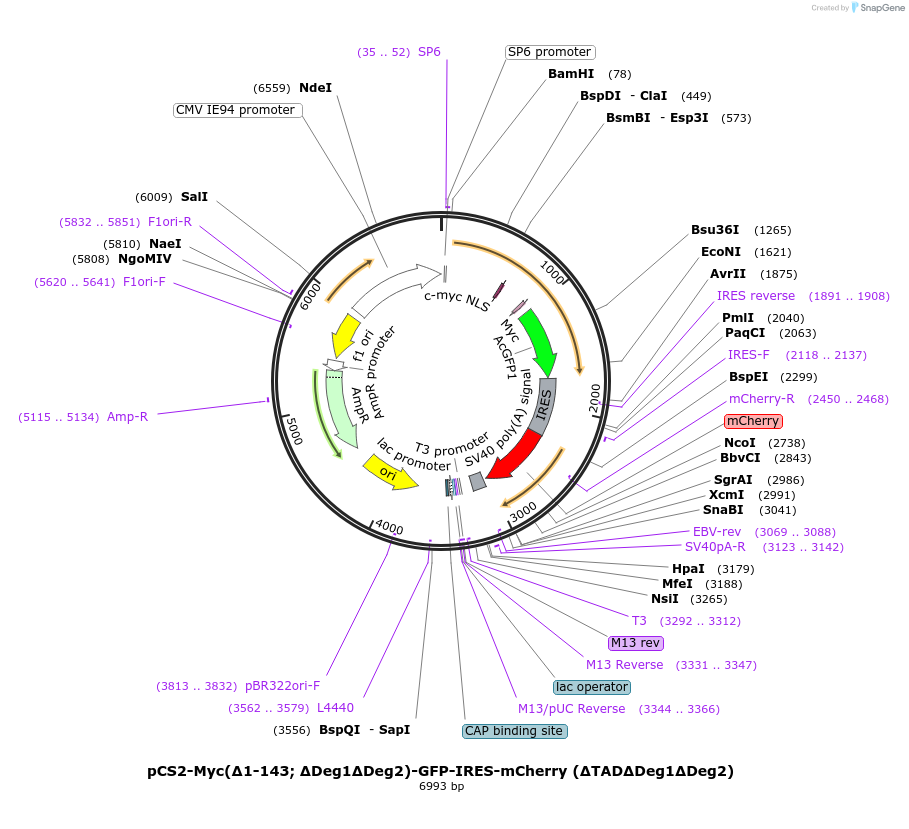
pCS2-Myc(Δ1-143; ΔDeg1ΔDeg2)-GFP-IRES-mCherry (ΔTADΔDeg1ΔDeg2)
Plasmid#231234PurposeMYC stability reporter construct with deletion of the N-terminal region (residues 1-143) and point mutations corresponding to the deletion of degron 1 and 2 for transient expression in mammalian cellsDepositorAvailable SinceJan. 24, 2025AvailabilityAcademic Institutions and Nonprofits only -
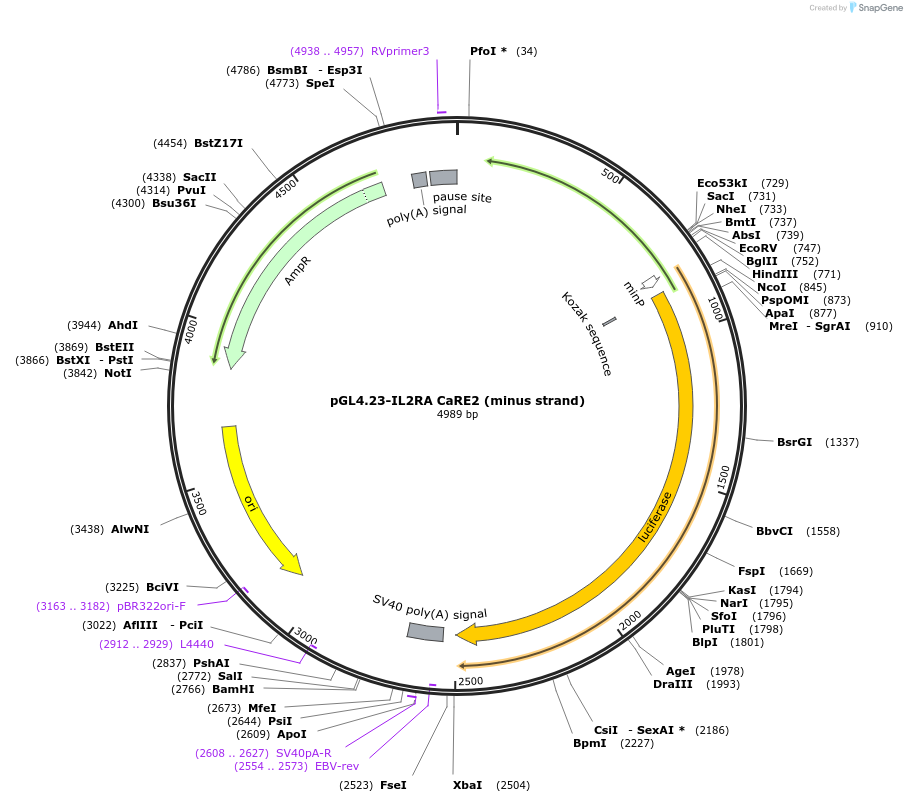
pGL4.23-IL2RA CaRE2 (minus strand)
Plasmid#91847PurposeLuciferase reporter for IL2RA enhancer (IGI-P0624)DepositorInsertIL2RA CaRE2 (minus strand) (IL2RA Human)
UseLuciferaseExpressionMammalianMutationrs6602425 A>C, G>A mutation at chr10:611706…Available SinceJuly 24, 2017AvailabilityAcademic Institutions and Nonprofits only -
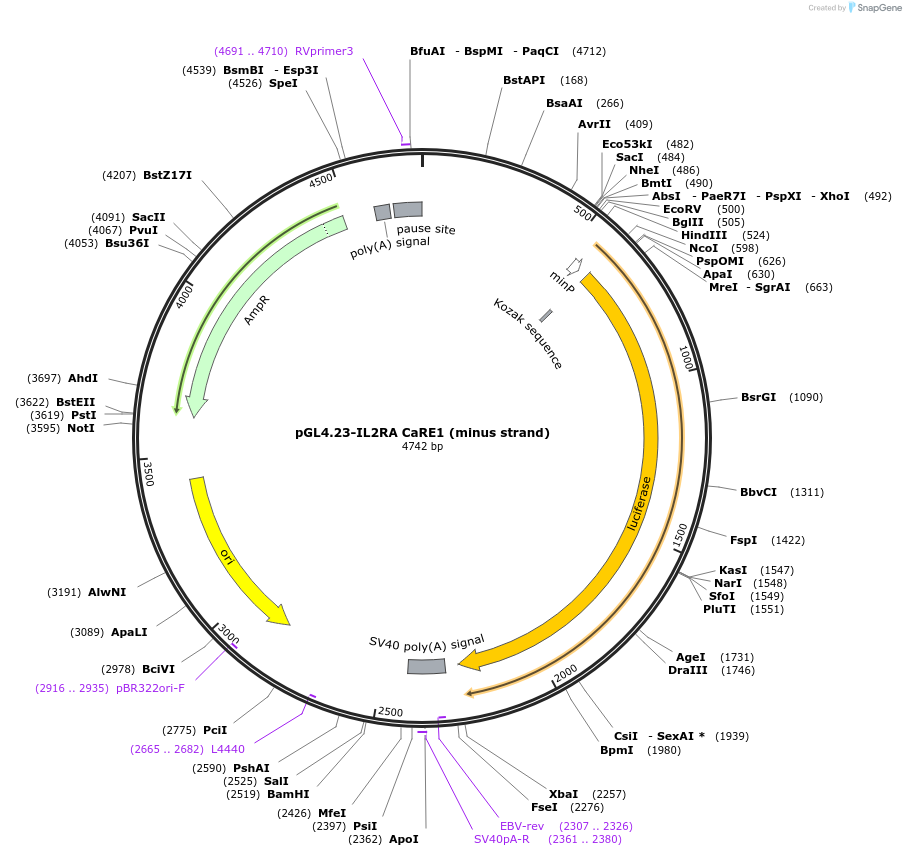
pGL4.23-IL2RA CaRE1 (minus strand)
Plasmid#91838PurposeLuciferase reporter for IL2RA enhancer (IGI-P0615)DepositorInsertIL2RA CaRE1 (minus strand) (IL2RA Human)
UseLuciferaseExpressionMammalianMutationPerfect match to reference sequenceAvailable SinceMay 30, 2017AvailabilityAcademic Institutions and Nonprofits only -
UQ6211
Bacterial Strain#37180DepositorBacterial ResistanceAmpicillin and KanamycinAvailable SinceAug. 9, 2012AvailabilityAcademic Institutions and Nonprofits only -
E. coli K-12 MG1655_rLP5(nth-ydgR)
Bacterial Strain#110245PurposeE. coli K-12 MG1655 with Landing Pad engineered into the nth-ydgR intergenic region. The Landing Pad consists of floxed mCherry,Chlor that are cotranscribed.DepositorBacterial ResistanceChloramphenicolAvailable SinceMarch 5, 2019AvailabilityAcademic Institutions and Nonprofits only -
E. coli K-12 MG1655_fLP4(essQ-cspB)
Bacterial Strain#110244PurposeE. coli K-12 MG1655 with Landing Pad engineered into the essQ-cspB intergenic region. The Landing Pad consists of floxed mCherry,Chlor that are cotranscribed.DepositorBacterial ResistanceChloramphenicolAvailable SinceMarch 5, 2019AvailabilityAcademic Institutions and Nonprofits only -
E. coli K-12 MG1655_rLP6(ygcE-ygcF)
Bacterial Strain#110246PurposeE. coli K-12 MG1655 with Landing Pad engineered into the ygcE-ygcF intergenic region. The Landing Pad consists of floxed mCherry,Chlor that are cotranscribed.DepositorBacterial ResistanceChloramphenicolAvailable SinceMarch 5, 2019AvailabilityAcademic Institutions and Nonprofits only -
E. coli K-12 MG1655_rLP1(atpI-gidB)
Bacterial Strain#110241PurposeE. coli K-12 MG1655 with Landing Pad engineered into the atpl-gidB intergenic region. The Landing Pad consists of floxed mCherry,Chlor that are cotranscribed.DepositorBacterial ResistanceChloramphenicolAvailable SinceMarch 5, 2019AvailabilityAcademic Institutions and Nonprofits only -
E. coli K-12 MG1655_fLP3(ybbD-ylbG)
Bacterial Strain#110243PurposeE. coli K-12 MG1655 with Landing Pad engineered into the ybbD-ylbG intergenic region. The Landing Pad consists of floxed mCherry,Chlor that are cotranscribed.DepositorBacterial ResistanceChloramphenicolAvailable SinceMarch 5, 2019AvailabilityAcademic Institutions and Nonprofits only -
E. coli K-12 MG1655_fLP2(yieN-trkD)
Bacterial Strain#110242PurposeE. coli K-12 MG1655 with Landing Pad engineered into the yieN-trkD intergenic region. The Landing Pad consists of floxed mCherry,Chlor that are cotranscribed.DepositorBacterial ResistanceChloramphenicolAvailable SinceMarch 5, 2019AvailabilityAcademic Institutions and Nonprofits only -
UQ5875
Bacterial Strain#37167DepositorBacterial ResistanceAmpicillin and KanamycinAvailable SinceAug. 3, 2012AvailabilityAcademic Institutions and Nonprofits only -
CSH130+pRCP
Bacterial Strain#195860PurposeWT E. coli strain transformed with plasmid carrying WT chromogenic protein (CP). Serves as control for suppressor strains. Grows purple when CP is induced with rhamnose.DepositorBacterial ResistanceChloramphenicolSpeciesAcropora milleporaAvailable SinceFeb. 8, 2023AvailabilityAcademic Institutions and Nonprofits only -
CSH130+pRCPam
Bacterial Strain#195859PurposeWT E. coli strain transformed with plasmid carrying chromogenic protein (CP) with amber nonsense mutation in chromophore. Serves as control for suppressor strains. No color when grown with or without rhamnose.DepositorBacterial ResistanceChloramphenicolSpeciesAcropora milleporaAvailable SinceFeb. 8, 2023AvailabilityAcademic Institutions and Nonprofits only -
UQ6139
Bacterial Strain#37177DepositorBacterial ResistanceAmpicillin and KanamycinSpeciesBurkholderia xenovoransAvailable SinceAug. 9, 2012AvailabilityAcademic Institutions and Nonprofits only -
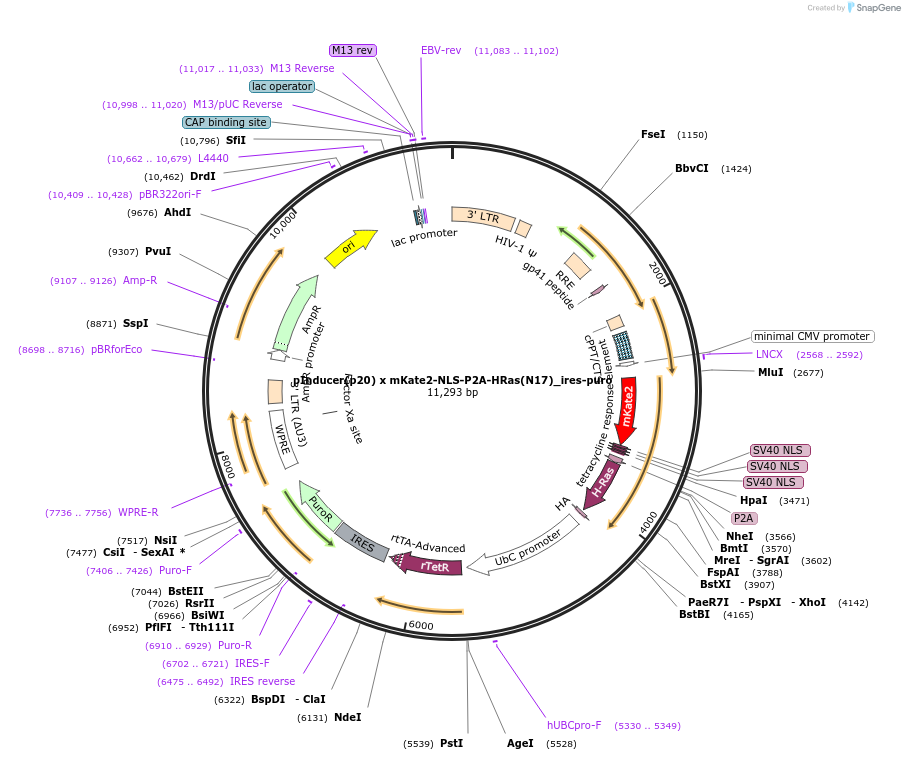
pInducer(p20) x mKate2-NLS-P2A-HRas(N17)_ires-puro
Plasmid#167832PurposeInducible expression of dominant negative HRas(N17) and P2A-coupled reporter mKate2-NLSDepositorAvailable SinceApril 20, 2021AvailabilityAcademic Institutions and Nonprofits only -
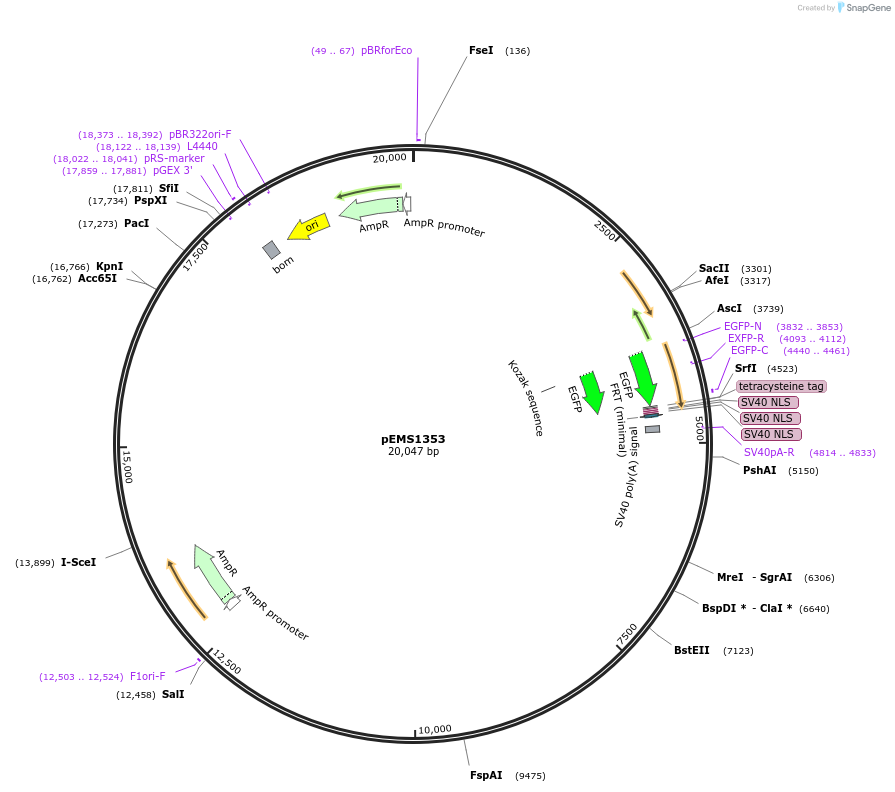
pEMS1353
Plasmid#29155PurposeExpression of the eGFP reporter gene was not detected in the brain and eye.DepositorAvailable SinceNov. 7, 2011AvailabilityAcademic Institutions and Nonprofits only