We narrowed to 2,008 results for: rigi
-
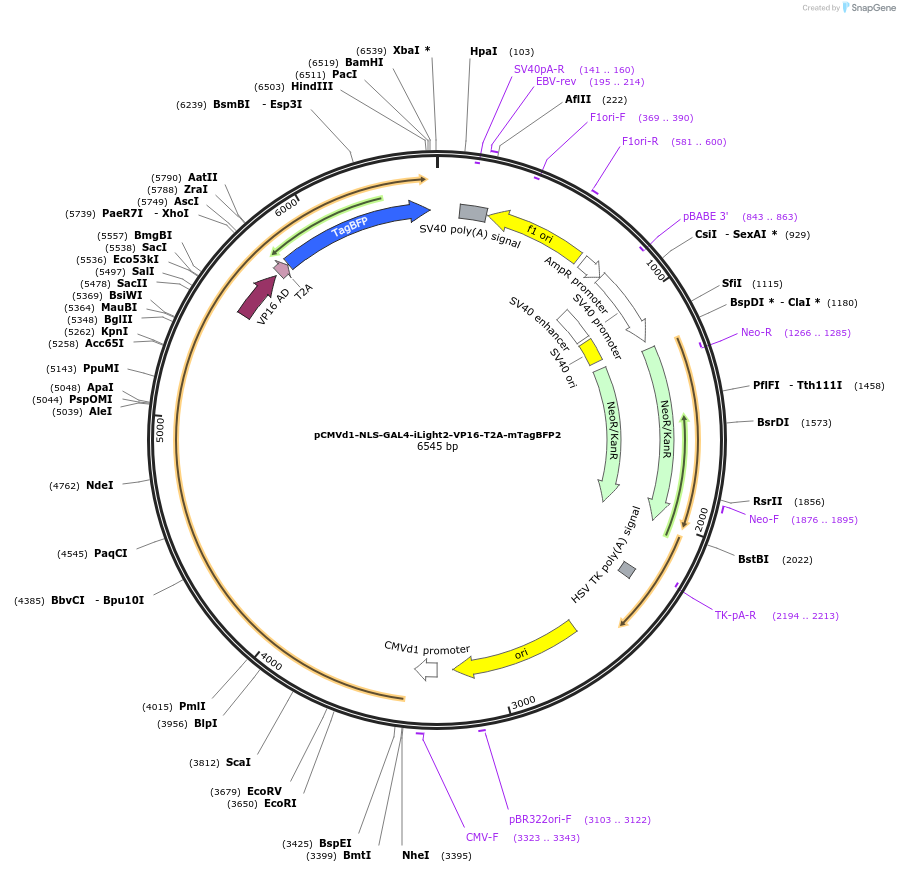
Plasmid#219548PurposeNLS-Gal4(1-65)-iLight2-VP16-T2A-mTagBFP2 construct expression in mammalian cells.DepositorInsertNLS-Gal4(1-65)-iLight2-VP16-T2A-mTagBFP2
ExpressionMammalianMutationThe (SAGG)x4 linker between GAL4(1-65) and the PC…PromoterCMVd1Available SinceMay 23, 2024AvailabilityAcademic Institutions and Nonprofits only -
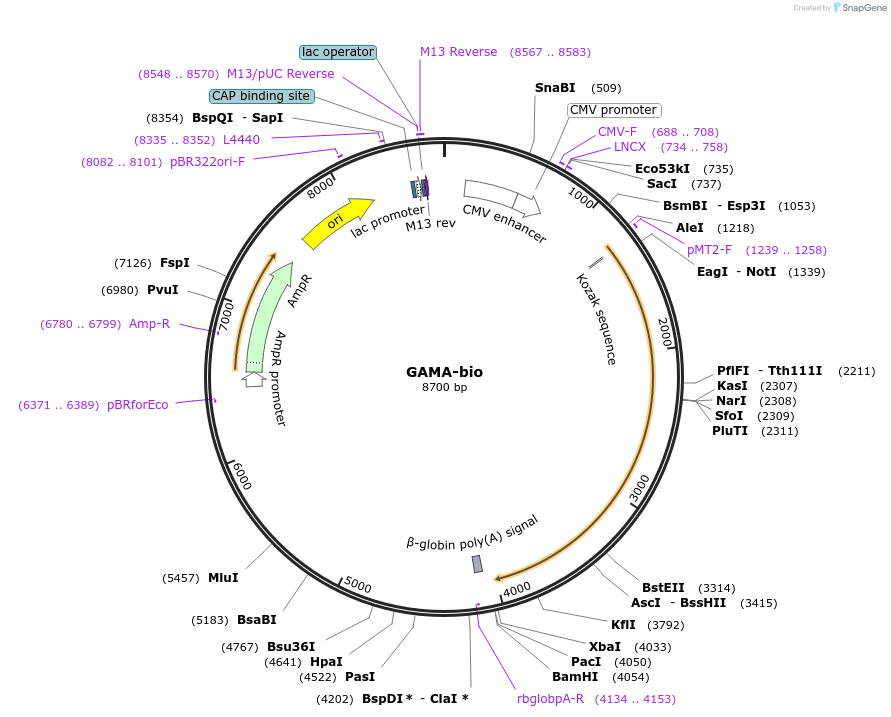
GAMA-bio
Plasmid#47747PurposeExpresses enzymatically monobiotinylated full-length GAMA ectodomain with no N-linked glycans upon cotransfection with BirA in mammalian cells. Contains a C-terminal rat Cd4 d3+4 tag.DepositorInsertCodon-optimised GAMA
Tagsenzymatic biotinylation sequence and rat Cd4 d3+4ExpressionMammalianMutationExogenous signal peptide of mouse origin, changed…PromoterCMVAvailable SinceFeb. 4, 2014AvailabilityAcademic Institutions and Nonprofits only -
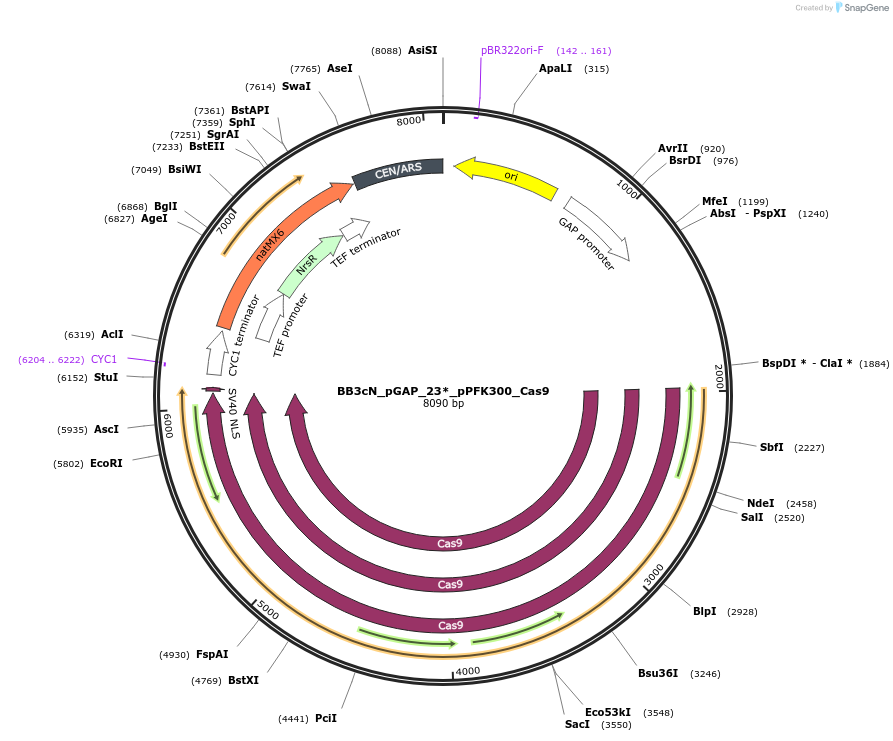
BB3cN_pGAP_23*_pPFK300_Cas9
Plasmid#104911PurposehCas9 under control of pPFK300 for direct cloning of HH-sgRNA-HDV PCR products for episomal expression in P. pastoris and selection on NTCDepositorInsertsHH - FS23 linker - HDV
human codon-optimized hcas9 sequence fused to a nuclear localization signal (NLS)
UseCRISPRExpressionYeastAvailable SinceJan. 14, 2020AvailabilityAcademic Institutions and Nonprofits only -
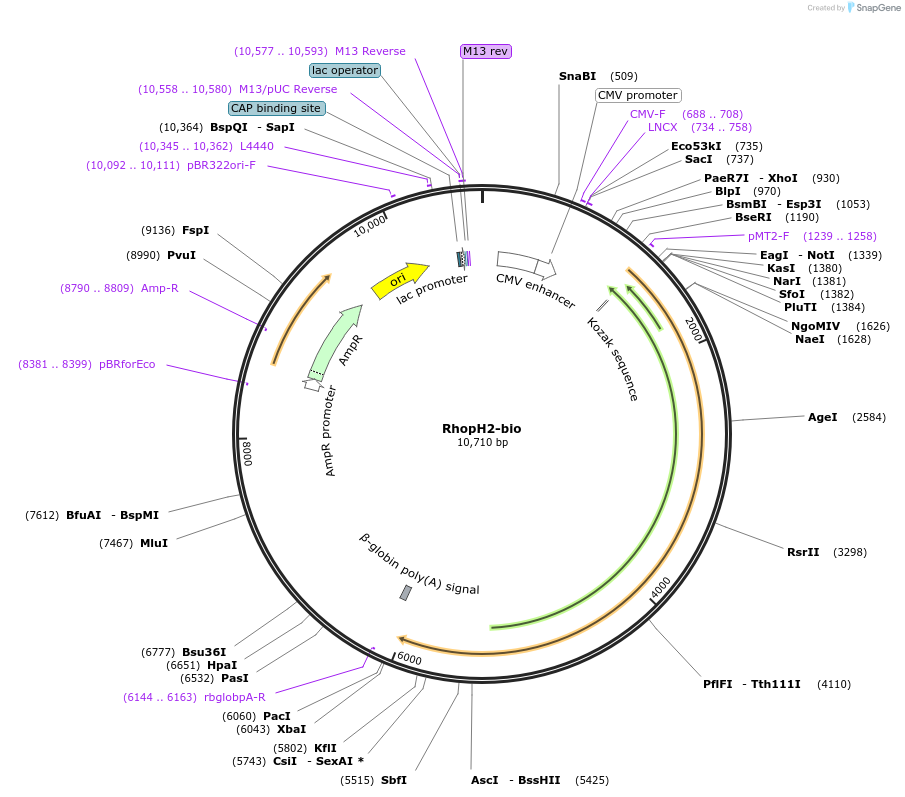
RhopH2-bio
Plasmid#47798PurposeExpresses enzymatically monobiotinylated full-length RhopH2 with no N-linked glycans upon cotransfection with BirA in mammalian cells. Contains a C-terminal rat Cd4 d3+4 tag.DepositorInsertCodon-optimised RhopH2
Tagsenzymatic biotinylation sequence and rat Cd4 d3+4ExpressionMammalianMutationExogenous signal peptide of mouse origin, changed…PromoterCMVAvailable SinceFeb. 4, 2014AvailabilityAcademic Institutions and Nonprofits only -
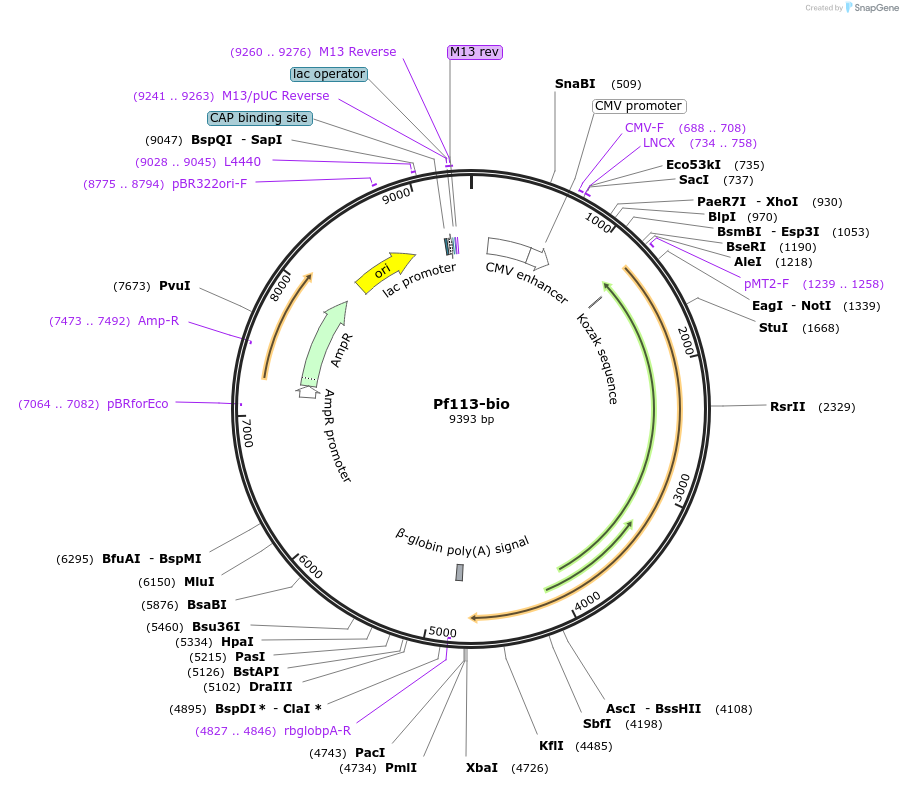
Pf113-bio
Plasmid#47729PurposeExpresses enzymatically monobiotinylated full-length Pf113 ectodomain with no N-linked glycans upon cotransfection with BirA in mammalian cells. Contains a C-terminal rat Cd4 d3+4 tag.DepositorInsertCodon-optimised Pf113
Tagsenzymatic biotinylation sequence and rat Cd4 d3+4ExpressionMammalianMutationExogenous signal peptide of mouse origin, changed…PromoterCMVAvailable SinceFeb. 4, 2014AvailabilityAcademic Institutions and Nonprofits only -
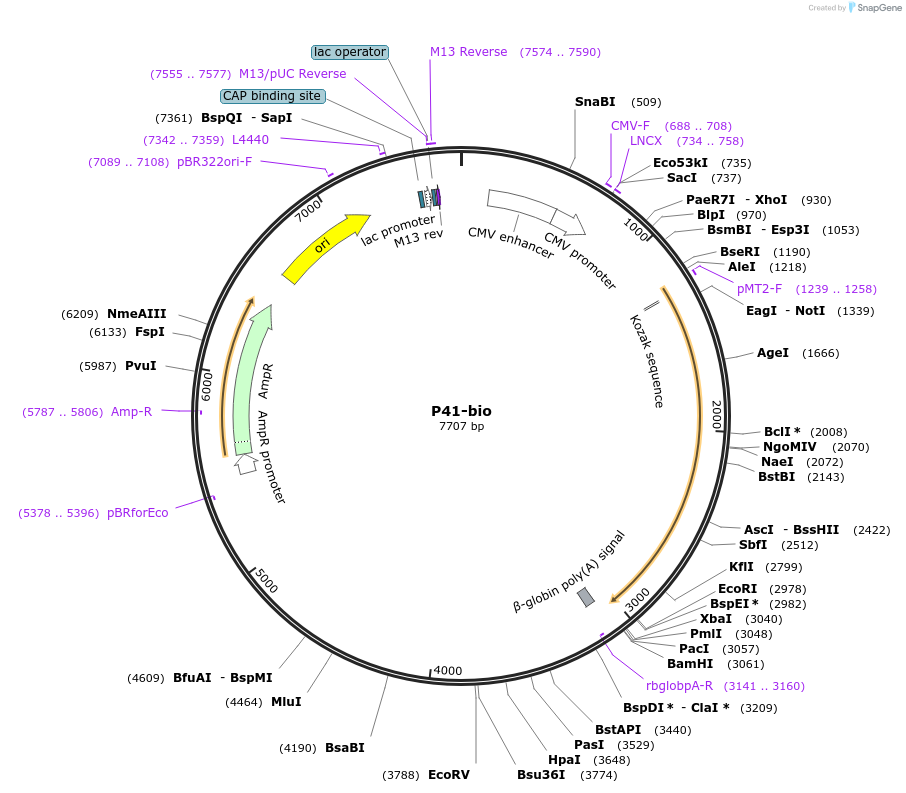
P41-bio
Plasmid#47739PurposeExpresses enzymatically monobiotinylated full-length P41 with no N-linked glycans upon cotransfection with BirA in mammalian cells. Contains a C-terminal rat Cd4 d3+4 tag.DepositorInsertCodon-optimised P41
Tagsenzymatic biotinylation sequence and rat Cd4 d3+4ExpressionMammalianMutationExogenous signal peptide of mouse origin, changed…PromoterCMVAvailable SinceFeb. 4, 2014AvailabilityAcademic Institutions and Nonprofits only -
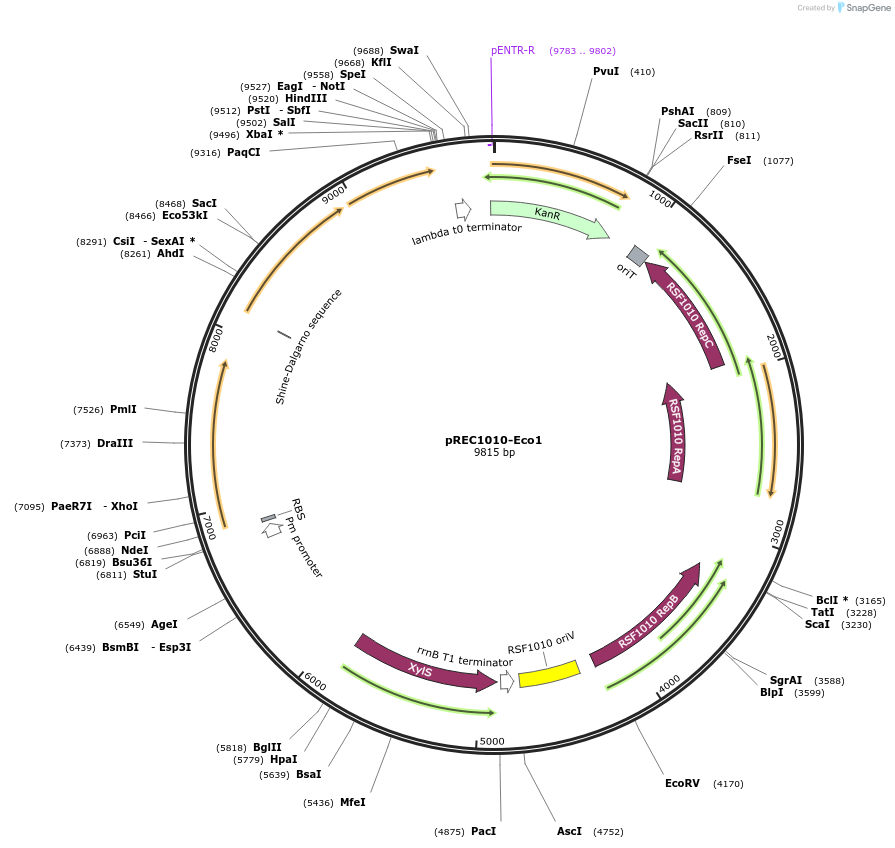
pREC1010-Eco1
Plasmid#240680PurposeRSF1010 origin of replication plasmid containing Eco1 recombitron with a SapI flanked stuffer in the ncRNA expressed by Pm promoterDepositorInsertEco1 RT, Eco1 ncRNA, CspRecT and EcSSB
ExpressionBacterialMutationWTAvailable SinceSept. 10, 2025AvailabilityAcademic Institutions and Nonprofits only -
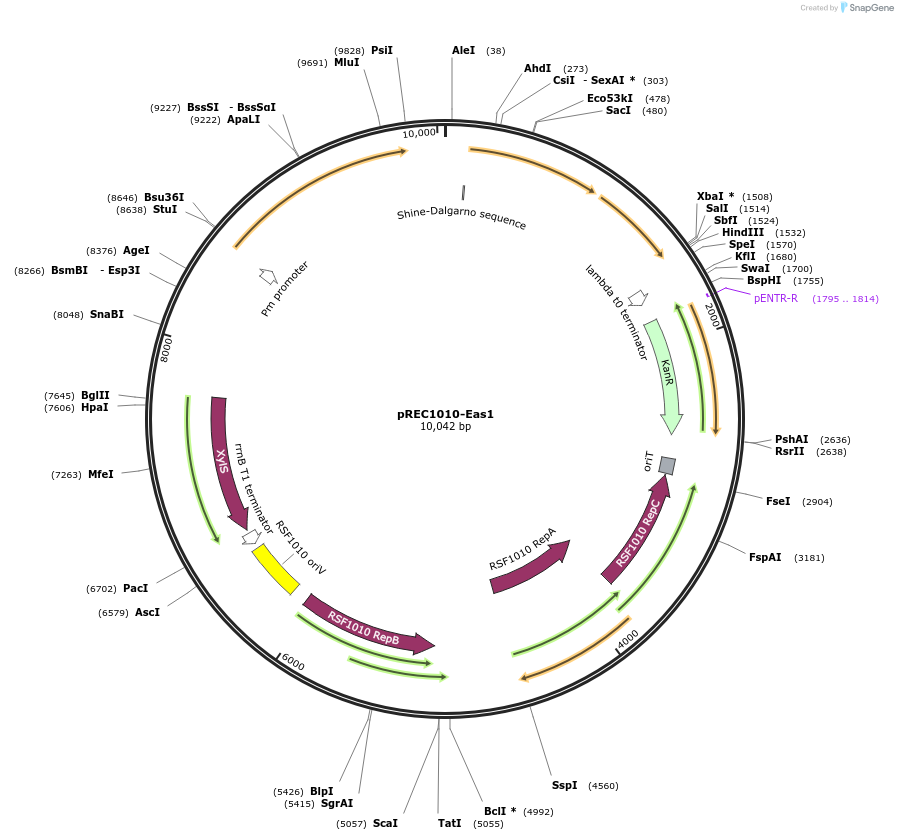
pREC1010-Eas1
Plasmid#240685PurposeRSF1010 origin of replication plasmid containing Eas1 recombitron with a SapI flanked stuffer in the ncRNA expressed by Pm promoterDepositorInsertEas1 RT, Eas1 ncRNA, CspRecT and EcSSB
ExpressionBacterialMutationWTAvailable SinceAug. 7, 2025AvailabilityAcademic Institutions and Nonprofits only -
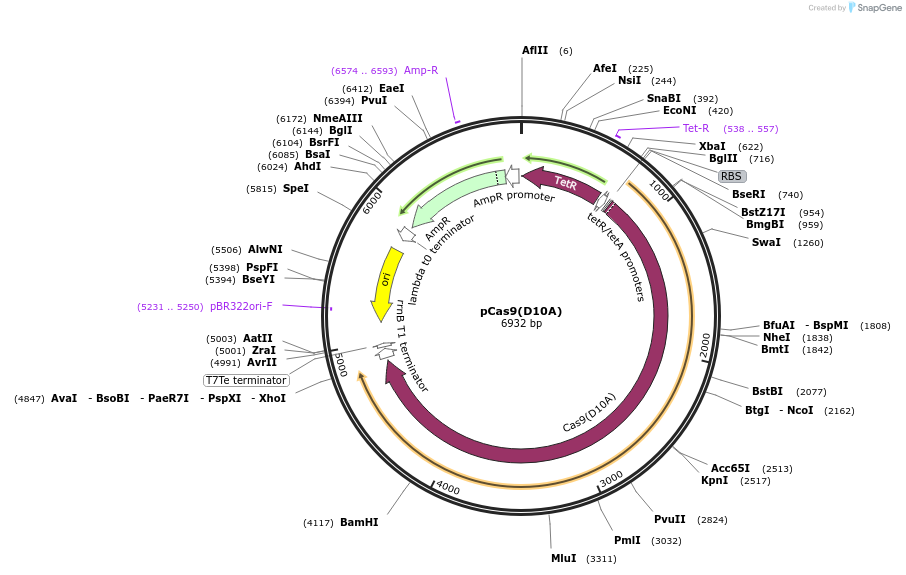
pCas9(D10A)
Plasmid#74495PurposeNicking Cas9 derived from pWT Cas9 (Addgene #44250). Ampicillin resistance marker, the tetracycline repressor (TetR), a ColE1 origin of replication and Streptococcus pyogenes Cas9 D10ADepositorInsertCas9 D10A
UseSynthetic BiologyExpressionBacterialMutationD10APromoterpTetoAvailable SinceMay 16, 2016AvailabilityAcademic Institutions and Nonprofits only -
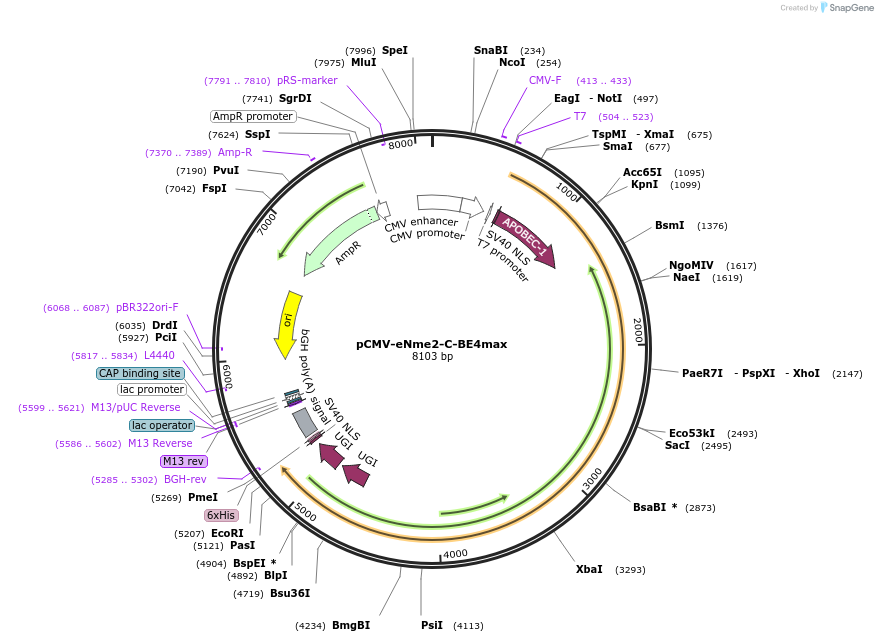
pCMV-eNme2-C-BE4max
Plasmid#183679PurposeExpresses eNme2-C-BE4 in mammalian cellsDepositorInsertrAPOBEC1-neNme2-C-2xUGI
UseCRISPRExpressionMammalianMutationNme2Cas9 P6S/D16A/E33G/K104T/D152A/F260L/A263T/A3…Available SinceOct. 14, 2022AvailabilityAcademic Institutions and Nonprofits only -
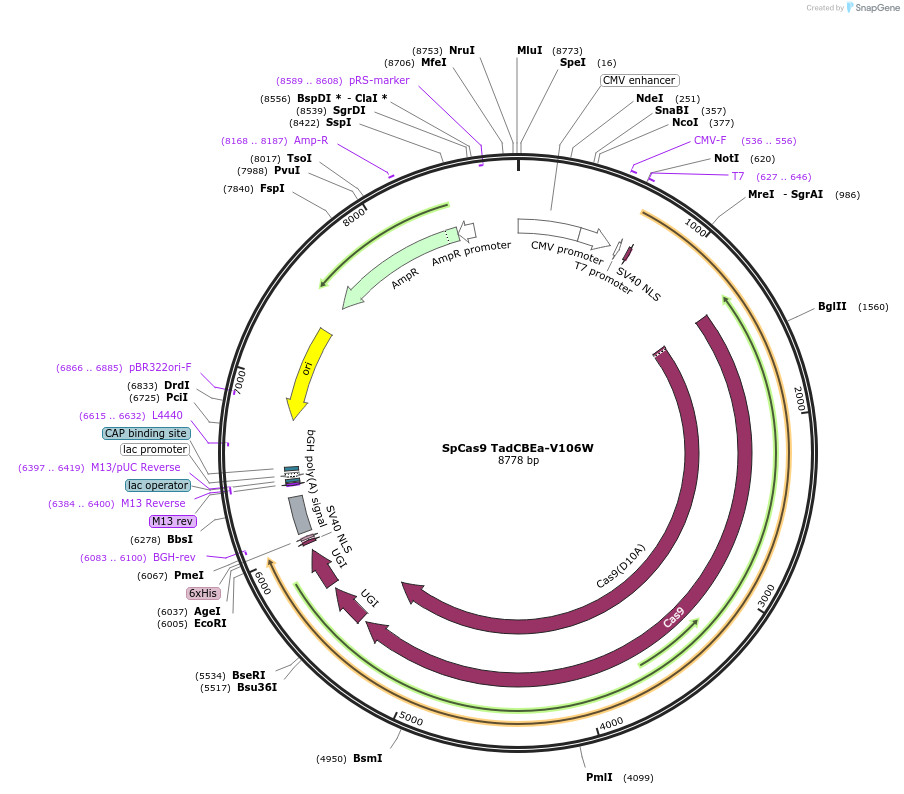
SpCas9 TadCBEa-V106W
Plasmid#193838PurposeExpress TadCBEa-V106W (with SpCas9) in mammalian cellsDepositorInsertTadA-CDa V106W-SpCas9 D10A
ExpressionMammalianAvailable SinceDec. 15, 2022AvailabilityAcademic Institutions and Nonprofits only -
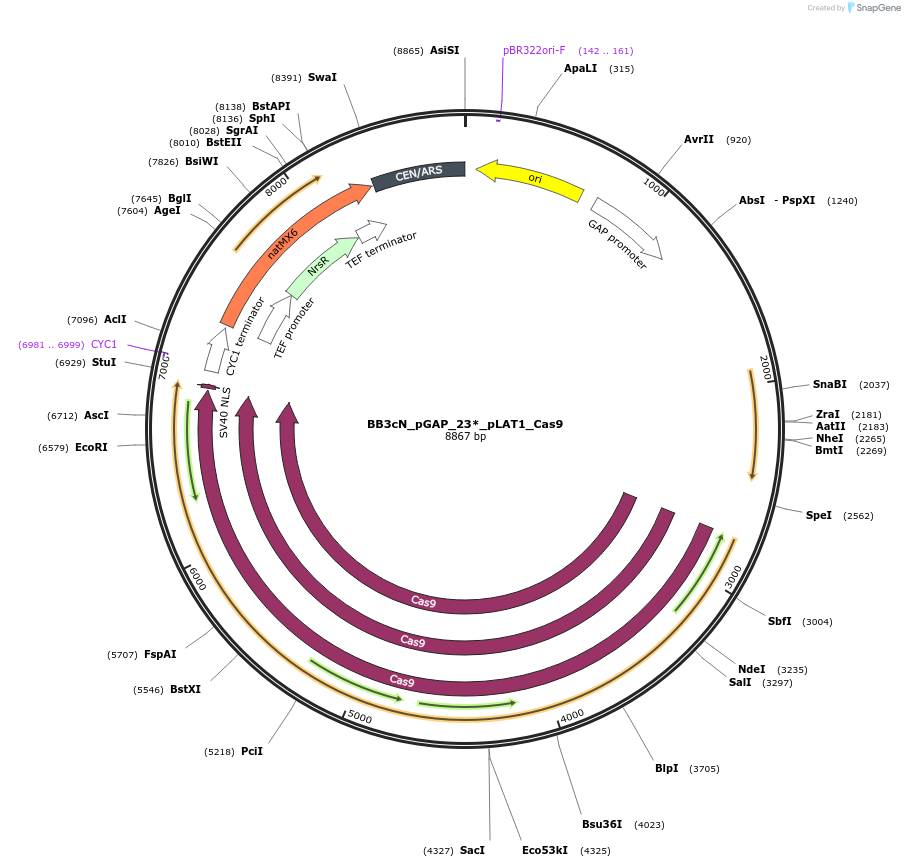
BB3cN_pGAP_23*_pLAT1_Cas9
Plasmid#104906PurposehCas9 under control of LAT1 for direct cloning of HH-sgRNA-HDV PCR products for episomal expression in P. pastoris and selection on NTCDepositorInsertsHH - FS23 linker - HDV
human codon-optimized hcas9 sequence fused to a nuclear localization signal (NLS)
UseCRISPRExpressionYeastAvailable SinceFeb. 23, 2019AvailabilityAcademic Institutions and Nonprofits only -
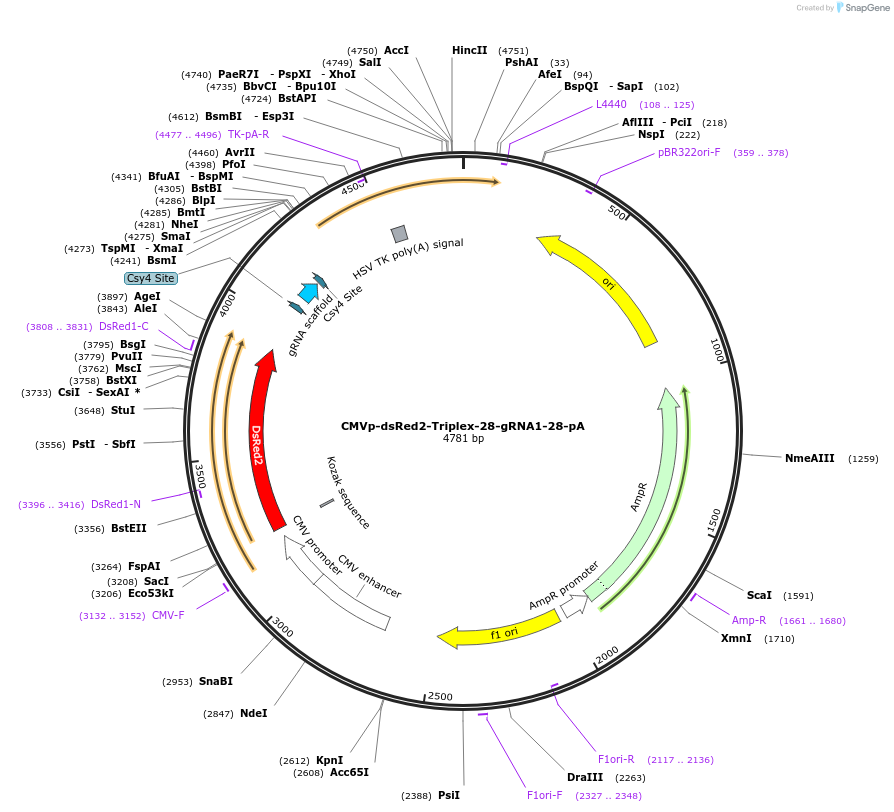
CMVp-dsRed2-Triplex-28-gRNA1-28-pA
Plasmid#55200PurposePlasmid encoding the Triplex/gRNA architecture.This is a modified form of the original plasmid described in the paper (Construct 3). mKate2 was replaced with dsRed2 because of distribution issues.DepositorInsertdsRed2
UseSynthetic BiologyExpressionMammalianPromoterCMVAvailable SinceJune 16, 2014AvailabilityAcademic Institutions and Nonprofits only -
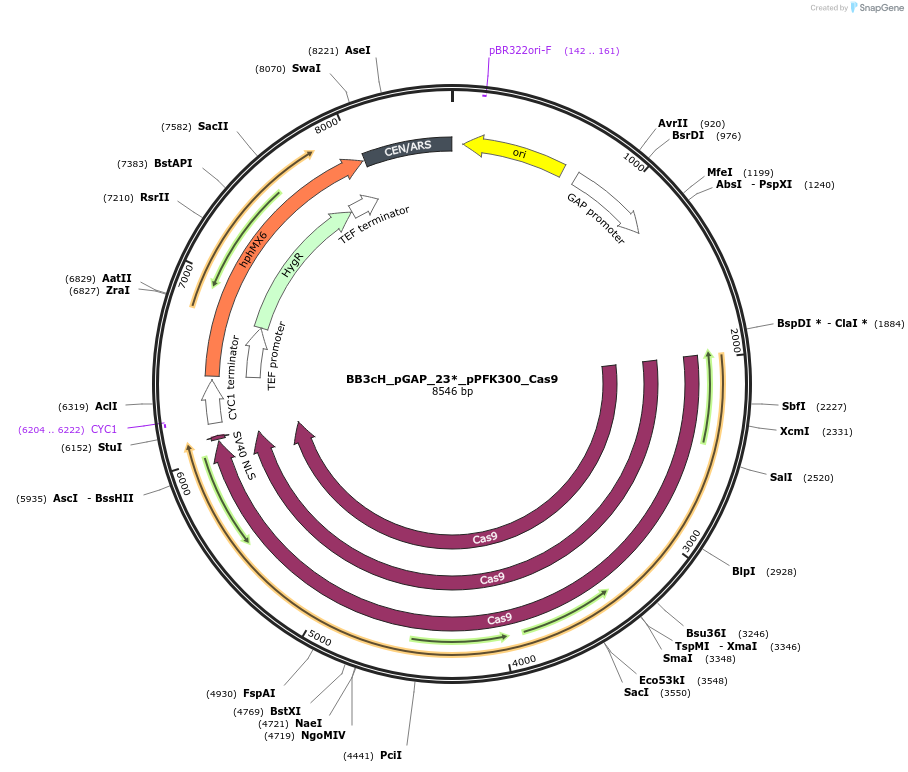
BB3cH_pGAP_23*_pPFK300_Cas9
Plasmid#104912PurposehCas9 under control of pPFK300 for direct cloning of HH-sgRNA-HDV PCR products for episomal expression in P. pastoris and selection on HygDepositorInsertsHH - FS23 linker - HDV
human codon-optimized hcas9 sequence fused to a nuclear localization signal (NLS)
UseCRISPRExpressionYeastAvailable SinceJan. 29, 2019AvailabilityAcademic Institutions and Nonprofits only -
pDual-eGFP(H66)
Plasmid#63558PurposeExpression of the Azure (blue) spectral variant of eGFP in bacteria and in mammalian cells. Could be used as a reporter for quantifying gene targeting and recombineering in mammalian cells.DepositorInsertHis-T7-eGFP(H66)
UseLentiviralTagsHis6 and T7ExpressionBacterial and MammalianMutationChanged tyrosine at position 66 with respect to t…PromoterCMV-EF1α hybrid (CEF)Available SinceApril 15, 2015AvailabilityAcademic Institutions and Nonprofits only -
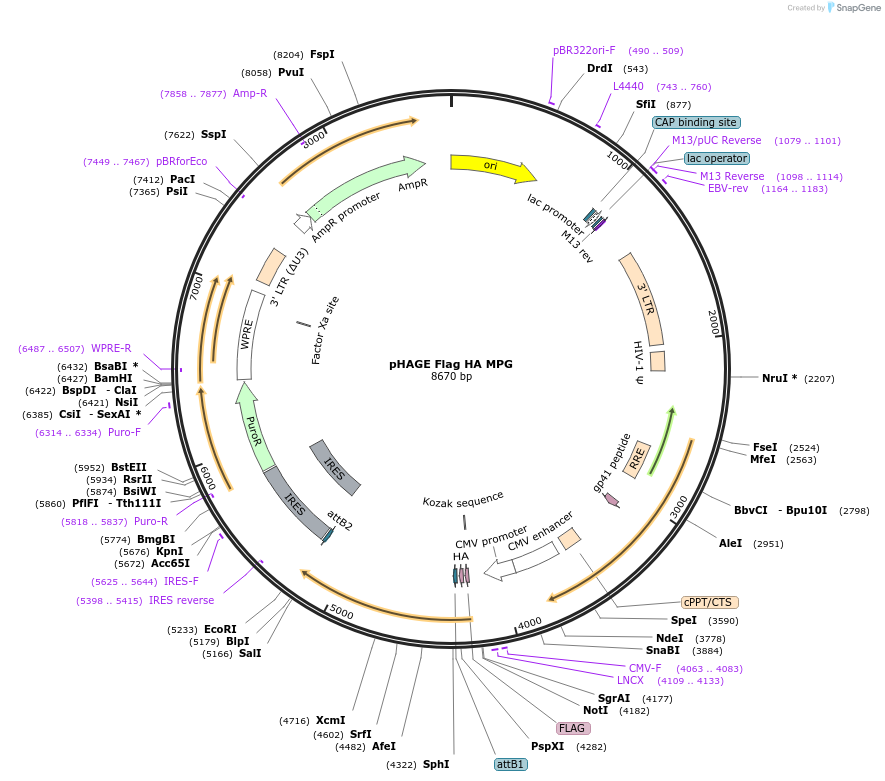
pHAGE Flag HA MPG
Plasmid#237804PurposeLentiviral vector expressing WT MPGDepositorAvailable SinceApril 24, 2025AvailabilityAcademic Institutions and Nonprofits only -
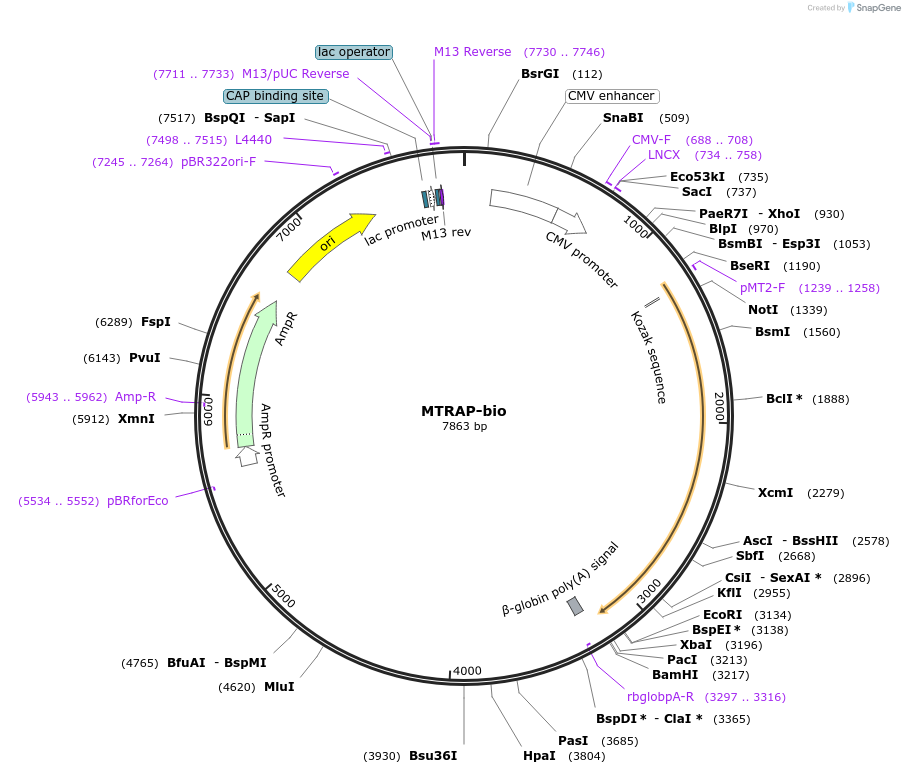
MTRAP-bio
Plasmid#47746PurposeExpresses enzymatically monobiotinylated full-length MTRAP ectodomain with no N-linked glycans upon cotransfection with BirA in mammalian cells. Contains a C-terminal rat Cd4 d3+4 tag.DepositorInsertCodon-optimised MTRAP
Tagsenzymatic biotinylation sequence and rat Cd4 d3+4ExpressionMammalianMutationExogenous signal peptide of mouse origin, changed…PromoterCMVAvailable SinceFeb. 4, 2014AvailabilityAcademic Institutions and Nonprofits only -
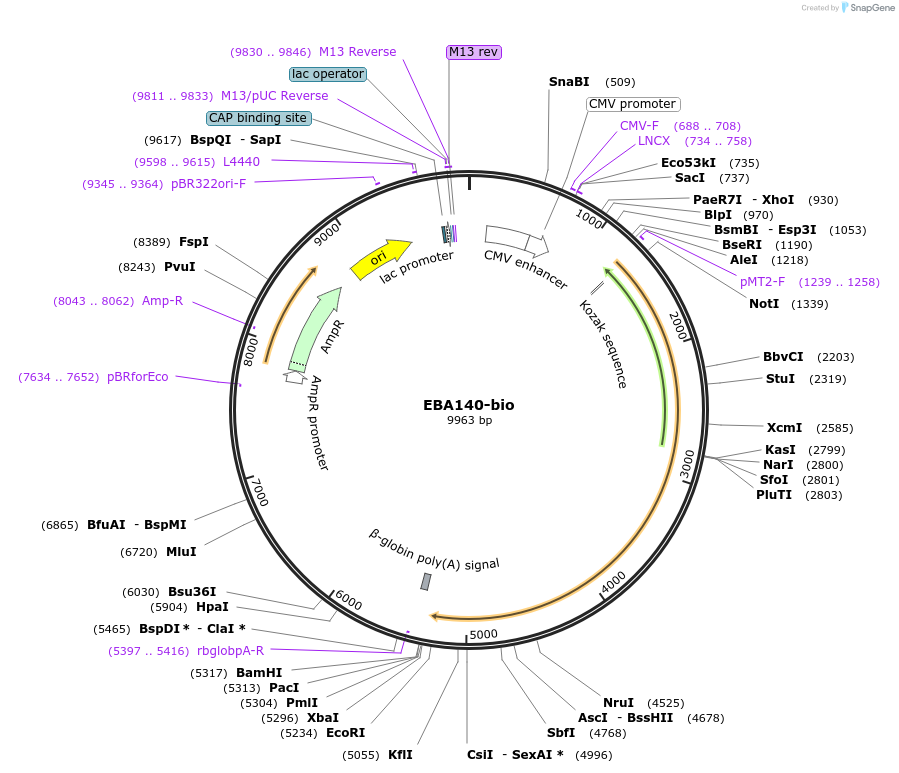
EBA140-bio
Plasmid#47742PurposeExpresses enzymatically monobiotinylated full-length EBA140 ectodomain with no N-linked glycans upon cotransfection with BirA in mammalian cells. Contains a C-terminal rat Cd4 d3+4 tag.DepositorInsertCodon-optimised EBA140
Tagsenzymatic biotinylation sequence and rat Cd4 d3+4ExpressionMammalianMutationExogenous signal peptide of mouse origin, changed…PromoterCMVAvailable SinceFeb. 4, 2014AvailabilityAcademic Institutions and Nonprofits only -
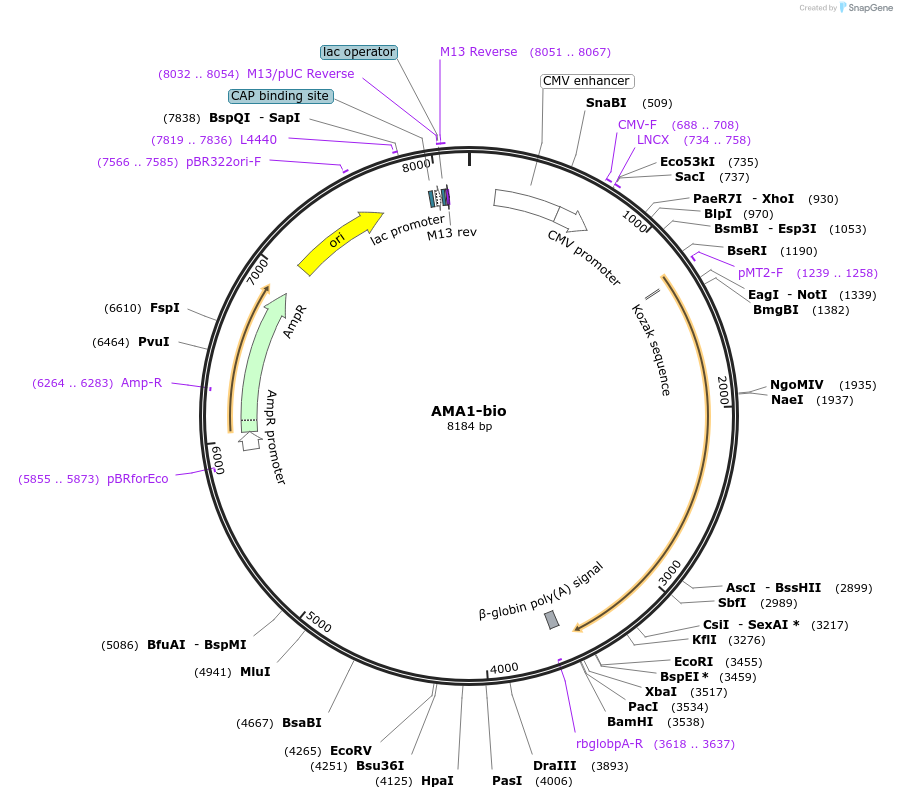
AMA1-bio
Plasmid#47741PurposeExpresses enzymatically monobiotinylated full-length AMA1 ectodomain with no N-linked glycans upon cotransfection with BirA in mammalian cells. Contains a C-terminal rat Cd4 d3+4 tag.DepositorInsertCodon-optimised AMA1
Tagsenzymatic biotinylation sequence and rat Cd4 d3+4ExpressionMammalianMutationExogenous signal peptide of mouse origin, changed…PromoterCMVAvailable SinceFeb. 4, 2014AvailabilityAcademic Institutions and Nonprofits only -
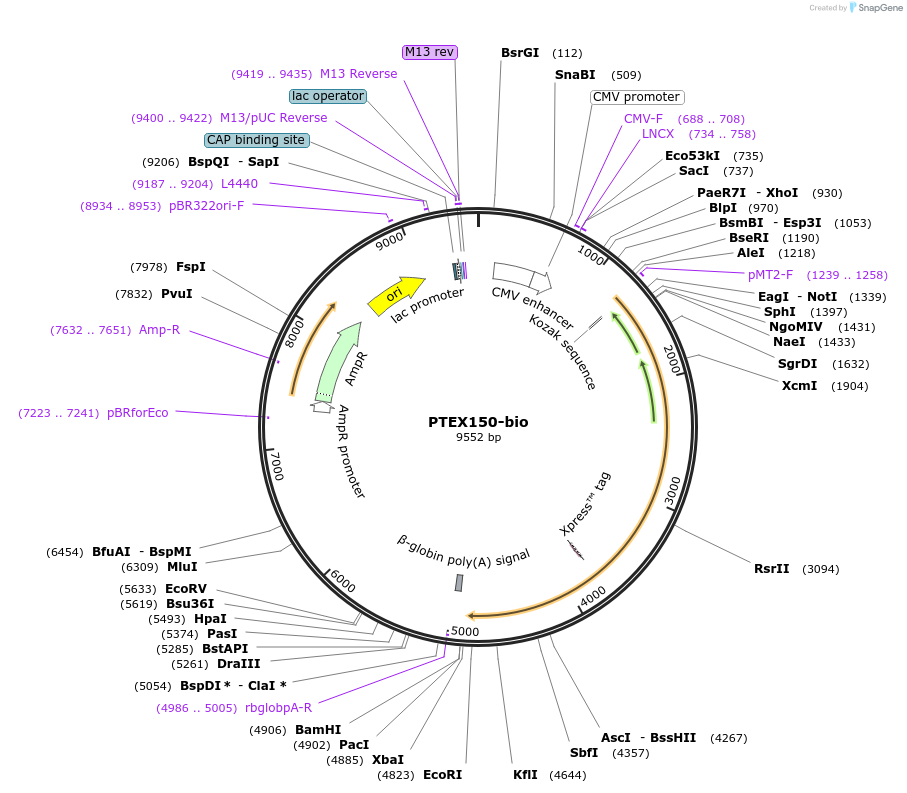
PTEX150-bio
Plasmid#47788PurposeExpresses enzymatically monobiotinylated full-length PTEX150 with no N-linked glycans upon cotransfection with BirA in mammalian cells. Contains a C-terminal rat Cd4 d3+4 tag.DepositorInsertCodon-optimised PTEX150
Tagsenzymatic biotinylation sequence and rat Cd4 d3+4ExpressionMammalianMutationExogenous signal peptide of mouse origin, changed…PromoterCMVAvailable SinceFeb. 4, 2014AvailabilityAcademic Institutions and Nonprofits only